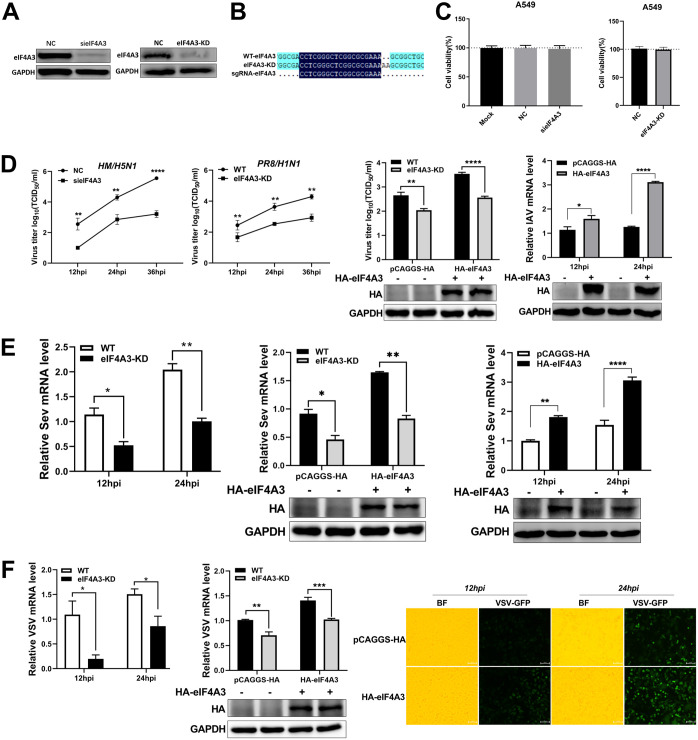
FIG 1.
The positive regulation of eIF4A3 on IAV, SeV, and VSV replication. (A to D) eIF4A3 promotes the replication of IAV. (A) The silencing efficiency of eIF4A3 or CRISPR-mediated knockdown efficiency of eIF4A3 was measured by Western blot assay. (B) The Sanger sequencing result indicated that a frameshift occurred in the eIF4A3 knockdown cells (eIF4A3-KD). (C) The cell viability of sieIF4A3 cells or eIF4A3 knockdown cells was measured by CCK-8 assay (mean ± SD of three independent experiments; *, P < 0.05; **, P < 0.01; ***, P < 0.001; ****, P < 0.0001; two-tailed Student’s t test). (D) The effect of eIF4A3 on HM and PR8 replication. A549 cells were transfected with sieIF4A3 or NC for 24 h, and were infected with HM virus (MOI, 0.01). The cell supernatants were harvested at 12 hpi, 24 hpi, and 36 hpi. Virus titers were determined by TCID50 assay on MDCK cells. eIF4A3 knockdown A549 cells or WT A549 cells were infected with PR8 (MOI, 0.1). Cell supernatants were collected at 12 hpi, 24 hpi, and 36 hpi, and virus titers were determined by TCID50 assay on MDCK cells. eIF4A3 knockdown A549 cells and WT A549 cells were transfected with HA-eIF4A3 or vector for 24 h and were infected with PR8 virus (MOI, 0.1). Cell supernatants were collected at 24 hpi, and virus titers were determined by TCID50 assay on MDCK cells. A549 cells were transfected with HA-eIF4A3 or vector for 24 h and were infected with PR8 at an MOI of 0.1. Samples were collected at 12 hpi and 24 hpi. The mRNA level of NP was detected by qRT-PCR. The viral RNAs levels were normalized to the GAPDH level (mean ± SD of three independent experiments *, P < 0.05; **, P < 0.01; ***, P < 0.001; ****, P < 0.0001; two-tailed Student’s t test). (E) The effect of eIF4A3 on SeV replication. eIF4A3 knockdown A549 cells or WT A549 cells were infected with SeV virus at an MOI of 1.0. Samples were collected at 12 hpi and 24 hpi. The mRNA level of SeV-N was detected by qRT-PCR, and the viral RNAs levels were normalized to the GAPDH level. eIF4A3 knockdown A549 cells and WT A549 cells were transfected with HA-eIF4A3 or vector for 24 h and were infected with SeV virus at an MOI of 1.0. Samples were collected at 24 hpi. The mRNA level of SeV-N was detected by qRT-PCR, and the viral RNAs levels were normalized to the GAPDH level. A549 cells were transfected with HA-eIF4A3 or vector for 24 h and were infected with SeV virus at an MOI of 1.0. Samples were collected at 12 hpi and 24 hpi. The mRNA level of SeV-N was detected by qRT-PCR. The viral RNA levels were normalized to the GAPDH level (mean ± SD of three independent experiments; *, P < 0.05; **, P < 0.01; ***, P < 0.001; ****, P < 0.0001; two-tailed Student’s t test). (F) The effect of eIF4A3 on VSV-GFP replication. eIF4A3 knockdown A549 cells or WT A549 cells were infected with VSV-GFP (MOI, 1.0). Samples were collected at 12 hpi and 24 hpi. The VSV-GFP viral RNA level was determined by qRT-PCR. The viral RNA levels were normalized to the GAPDH level. eIF4A3 knockdown A549 cells or WT A549 cells were transfected with HA-eIF4A3 or vector for 24 h and infected with VSV-GFP (MOI, 1.0). Samples were collected at 24 hpi. The VSV-GFP viral RNA level was determined by qRT-PCR. The viral RNA levels were normalized to the GAPDH level. A549 cells were transfected with HA-eIF4A3 or vector for 24 h and infected with VSV-GFP (MOI, 1.0). The VSV-GFP was visualized by fluorescence microscopy at 12 hpi and 24 hpi (×100 magnification) (mean ± SD of three independent experiments; *, P < 0.05; **, P < 0.01; ***, P < 0.001; ****, P < 0.0001; two-tailed Student’s t test).