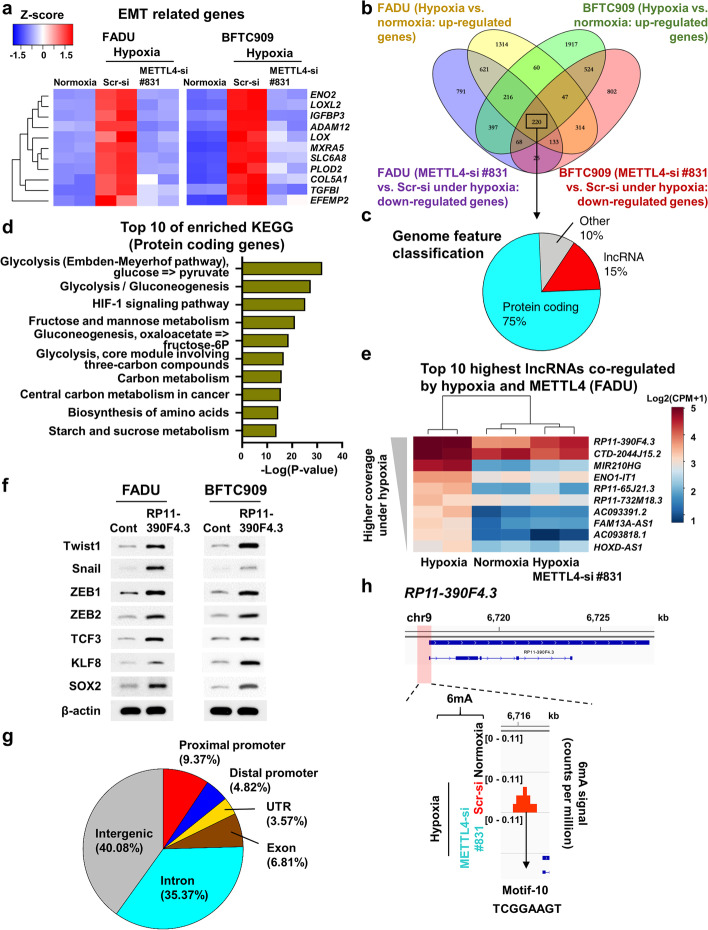
Fig. 4.
Analysis of RNA-seq and 6mA-ChIP-exo-seq datasets shows hypoxia/METTL4 co-regulated genes and 6mA signals-regulated genes. a The expression heatmap of EMT-related genes in FADU and BFTC909 cells, whose expression increased under hypoxia and decreased in the hypoxic status undergoing METTL4 knockdown. b Venn diagram shows the overlapping set of genes (n=220) co-regulated by hypoxia and METTL4 through overlapping of upregulated genes under hypoxia and downregulated genes in the hypoxic status undergoing METTL4 knockdown in both FADU and BFTC909 cells. c Pie chart shows the different percentage of hypoxia and METTL4 co-regulated genes according to the classification of gene feature (protein coding, 75%; lncRNA, 15%; others, 10%). d KEGG analysis shows the top 10 enriched pathways in the class of protein-coding genes co-regulated by hypoxia and METTL4. e Heatmap shows the expression of top ten lncRNAs co-regulated by hypoxia and METTL4 using RNA-seq datasets from FADU cells. f Western blot analysis shows that overexpression of lncRNA RP11-390F4.3 activated a set of EMT transcriptional regulators. g Pie chart shows the annotation of hypoxia-induced/METTL4 dependent gain-of-6mA regions located in different genomic regions. h LncRNA RP11-390F4.3 was used as an example of hypoxia-induced/METTL4-dependent 6mA regulated gene that contained hypoxia-induced/METTL4-dependent 6mA signals on its promoter region. Different 6mA motifs were calculated by HOMER. Only motif-10 was indicated on the lncRNA RP11-390F4.3 promoter. A magnified window around the motif-10 area from 3 gene tracks is shown. Y-axis denotes the scale of the number of reads