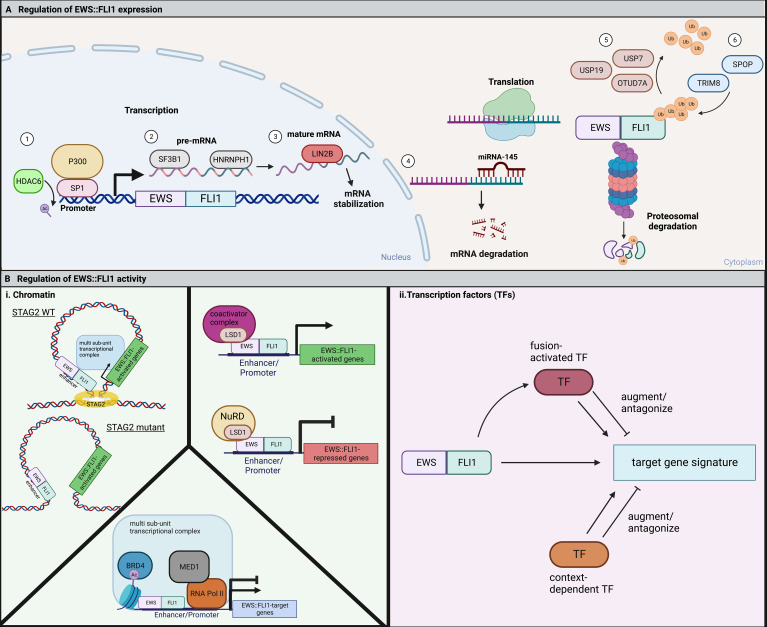
Figure 2.
(A) EWS::FLI1 expression and activity are highly regulated through many cell-intrinsic processes. 1) The EWS::FLI1 and endogenous EWSR1 promoter are positively regulated by SP1 in complex with the histone acetyltransferase, P300. HDAC6 keeps SP1 deacetylated, allowing for continued binding and expression of the fusion (58). 2) SF3B1 is a member of the spliceosome that is critical to for proper pre-mRNA splicing of EWS::FLI1. HNRNPH1 is a RNA binding protein that facilitates the splicing of EWSR1 exon 8-containing fusions (59). 3) The RNA-binding protein LIN28B can bind to and stabilize EWS::FLI1 mRNA (60). 4) In opposition, the microRNA miRNA-145 can bind the 3’ UTR of FLI1 and cause mRNA degradation of the fusion (64). EWS::FLI1 protein is degraded by the proteasome and regulated by the competing action of deubiquitinases and E3 ubiquitin ligases. 5) USP7, USP19, and OTUD7A are three deubiquitinases that have been shown to promote EWS::FLI1 expression (67–69). 6) TRIM8 and SPOP are two E3 ubiquitin ligases that have been reported to promote EWS::FLI1 degradation (19, 69). (B) (i) EWS::FLI1 activity is regulated by chromatin topologies and chromatin modifying complexes. In Ewing sarcoma cells with mutant STAG2, the EWS::FLI1 transcriptional signature is disrupted by alterations in chromatin looping (20, 21). LSD1 functions to regulate fusion activity in Ewing sarcoma cells through interaction with the fusion itself and either coactivator or repressive (NuRD) complexes resulting in either gene activation or repression (35, 49, 70–72). BRD4 regulates both the fusions activating and repressive activity through indirect binding with EWS::FLI1 in a large multi-subunit transcriptional complex (73). ii. EWS::FLI1 activity is also regulated by fusion-activated TFs or context-dependent TFs. These TFs then can either augment or antagonize expression of different subsets of the EWS::FLI1 target gene signature. Figure created with BioRender.com.