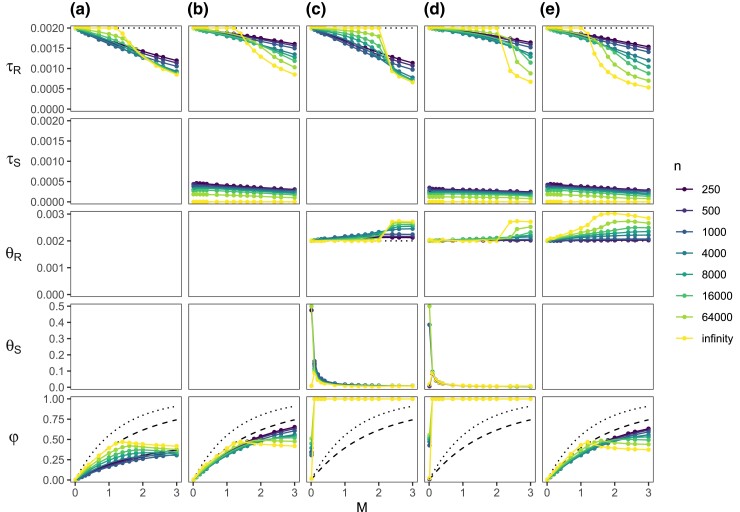
Fig. 2.
Best-fitting parameter values under the MSci model of figure 1d when data of two sequences per locus (one per species), each of n sites, are generated under the IM model of figure 1a. Five methods (a-e) are used to fit the MSci model, estimating 2, 3, 4, 5, and 4 parameters, respectively, whereas the other parameters are fixed. In (a), and are estimated, but and are fixed at their true values in the IM model, and the introgression time is fixed at . In (b), is estimated as well. In (c), is fixed, whereas the other four parameters are estimated. In (d), all five parameters are estimated. In (e), the constraint is enforced so that four free parameters are estimated. The dotted lines for indicate the true total amount of introgression of equation (10). The dashed lines indicate of equation (11). The true and best-fitting distributions of the coalescent time (t) are shown in supplementary figure S1, Supplementary Material online.