Figure 3.
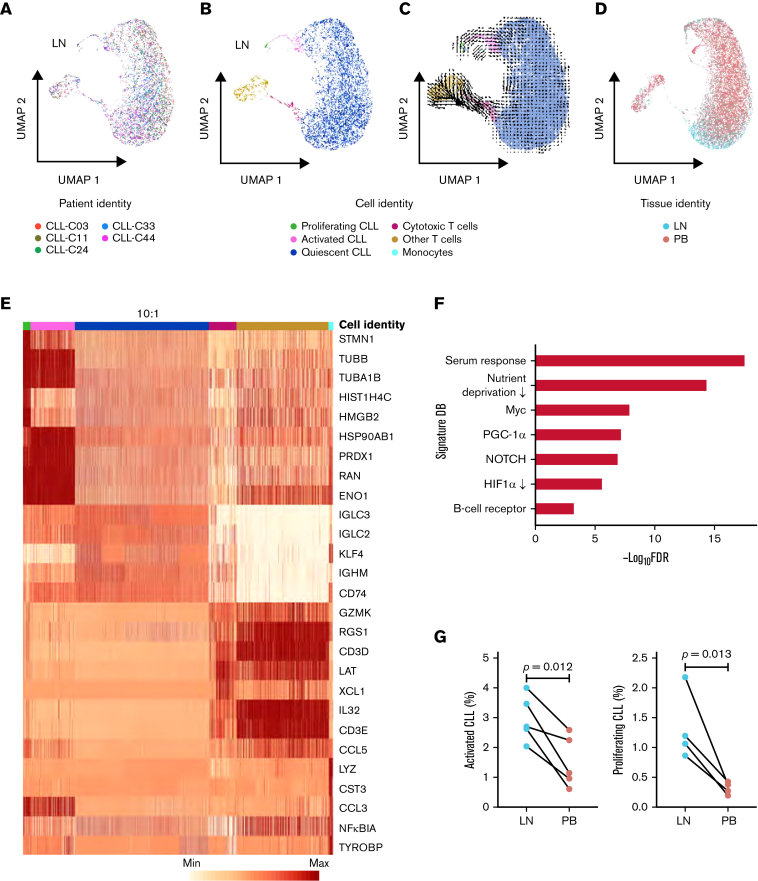
Single-cell transcriptomic analysis reveals intratumoral heterogeneity in LNs. (A) Uniform manifold approximation and projection (UMAP) of integrated scRNA-seq data from 15 107 single cells across 5 LN samples. Each color represents a different LN sample. (B) Clustering of LN single cells into major cell lineages and subpopulations. Each color represents a different cell identity cluster. CLL cells clustered into 3 subpopulations (proliferating, activated, quiescent) based on differences in transcriptional profiles. (C) RNA velocities derived from deterministic modeling projected on a UMAP of LN samples. CLL cells begin in the proliferating state, transition to the activated state, and end in the quiescent state. (D) UMAP of single cells from PB and LN (n = 5 pairs). Cell rendering after integration of PB and LN data is slightly different from that of LN data alone, as expected when computed from combined data. (E) Heatmap of the 5 most differentially expressed genes in each cell identity cluster. The color scale represents the number of reads mapping to the indicated gene per 10 000 reads. The size of the quiescent CLL population has been downscaled to improve the visualization of smaller clusters. Each column representing a quiescent CLL cell is 0.1× the width of columns representing other cells. (F) Significant overlap (hypergeometric test, FDR <0.05) between genes upregulated in activated CLL cells in LN and the indicated gene signatures. (G) Percentage of activated and proliferating CLL cells in LN and PB. Connecting lines indicate paired samples from each patient.