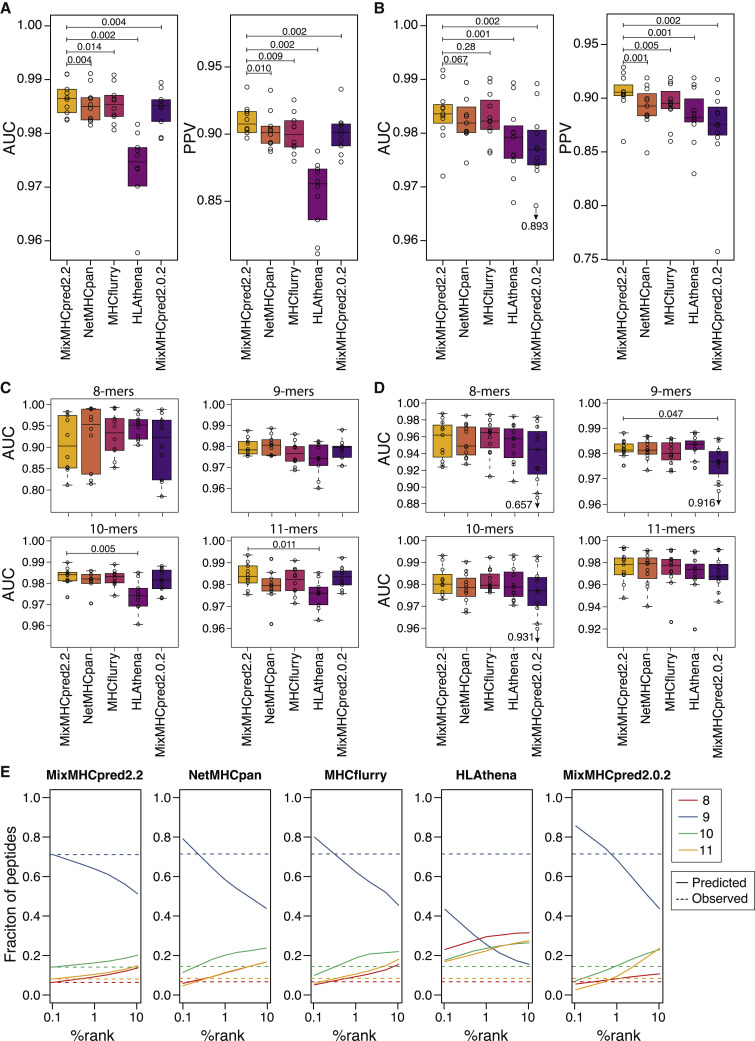
Figure 2.
Models of HLA-I binding specificities and peptide length distributions improve predictions of naturally presented HLA-I ligands
(A) Boxplot of AUC and PPV values for the different predictors considered in this study applied on the 10 HLA-I peptidomics samples from Gfeller et al.18
(B) AUC and PPV values obtained for HLA-I peptidomics samples from Pyke et al.20
(C) AUC values computed separately for each peptide length for the ten samples from Gfeller et al.18 Only p values between MixMHCpred2.2 and other tools and smaller than 0.05 are indicated.
(D) AUC values computed separately for each peptide length for eleven samples from Pyke et al.20 Only p values between MixMHCpred2.2 and other tools and smaller than 0.05 are indicated.
(E) Predicted peptide length distributions at different %rank thresholds for each HLA-I ligand predictor (average over alleles available in all predictors). Dashed lines show the peptide length distributions observed in naturally presented HLA-I ligands (average over all alleles). Boxplots in (A)–(D) represent the median and lower/upper quartiles. p values were computed with paired Wilcoxon test.