Fig. 1.
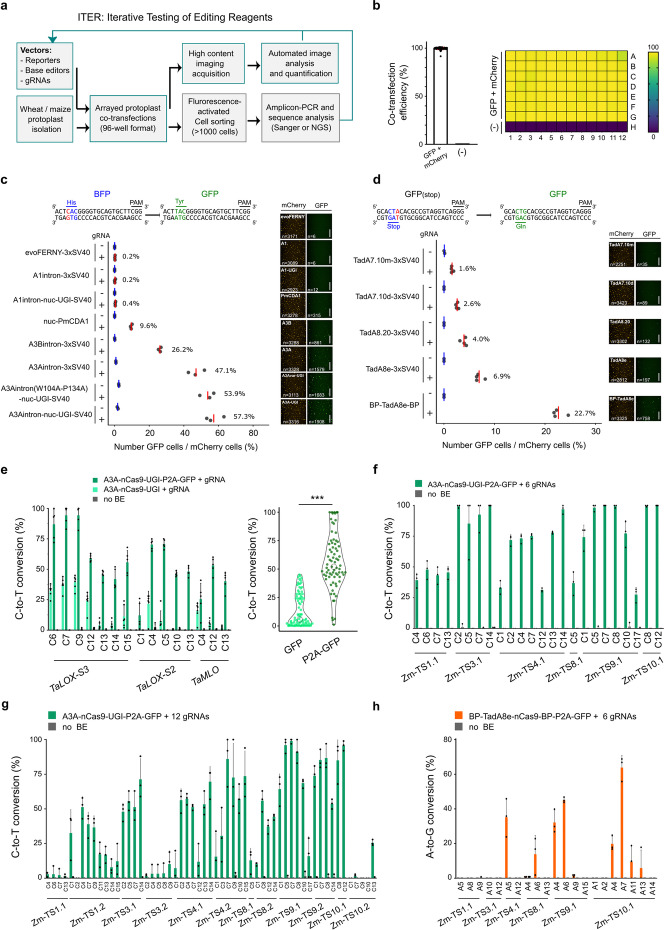
ITER is an efficient platform for testing base editing in wheat and maize protoplasts. a Schematic illustration of ITER. Wheat or maize protoplasts are co-transfected in 96-well format. Cells are imaged and analyzed by automated high-content imaging. Resulting data are interpreted and inform the subsequent optimization cycle. Validation of base editor activity at endogenous targets is performed by genotyping after fluorescence-activated cell sorting (FACS). b Co-transfection efficiency as measured by the number of GFP over the number of mCherry positive cells in wheat protoplasts. Negative controls (−) do not contain vector DNA. The grid shows the 96-well plate positions and the gradient colors coding represents co-transfection efficiencies. c, d Base editing efficiencies for multiple nCas9-CBE (c) or nCas9-ABE (d) configurations in wheat protoplasts. Efficiencies were measured as the percentage of cells showing BFP to GFP conversion for C-to-T editing (c) or GFP recovery for A-to-G editing (d). Red and blue lines represent mean efficiency of cells transfected with or without gRNA, respectively. Representative pictures show mCherry or GFP fluorescence in protoplasts. n indicates the total number of cells measured after image segmentation in the mCherry or GFP channel. Scale bars: 200 μm. e Multiplex C-to-T base editing of A3A-nCas9-UGI or A3A-nCas9-UGI-P2A-GFP at three wheat targets. One sub-genome was analyzed by Sanger sequencing and EditR analysis. Asterisks represents significant difference in efficiency as measured by Student’s t test (***: p<0.001). f, g Multiplex C-to-T base editing of A3A-nCas9-UGI at 6 (f) or 12 (g) maize targets. h Multiplex A-to-G base editing efficiency of BP-TadA8e-nCas9-BP ABE at 6 maize targets. Transfection of the cells with a GFP expressing vector (no BE) was used as a negative control in e–h. x-axis in e–h indicates targeted Cytosines (e, f, g) or Adenines (h) at different positions along the protospacer, with the PAM-distal base being position 1. Editing rates were calculated from three to 6 independent biological replicates that are depicted as dots on plots (c–h)