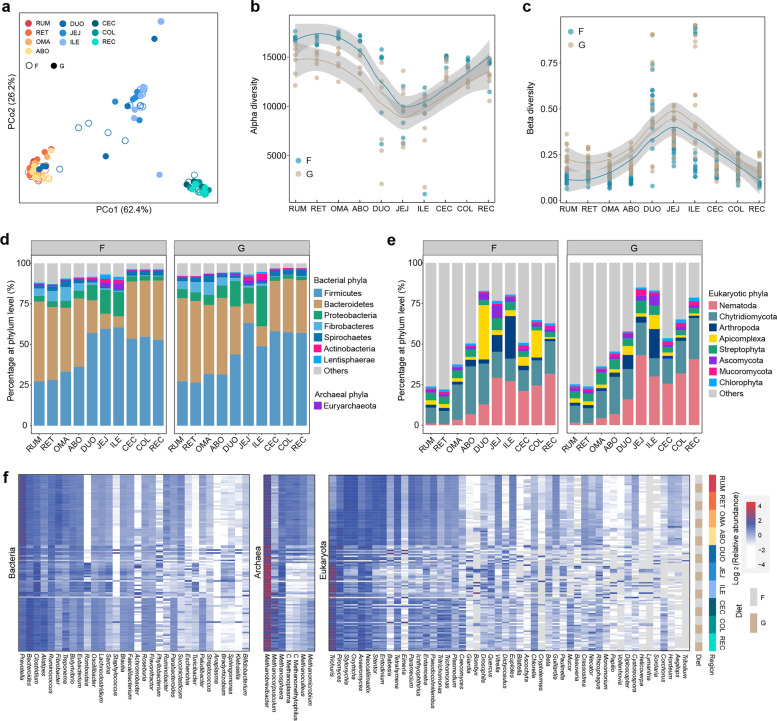
Fig. 1.
Region-specific distribution of distinct microbial populations from the proximal to distal gastrointestinal tract (GIT) of dairy cattle in the forage-based (F) and grain-based (G) diets, respectively. a Principal coordinate analysis (PCoA) profile among the 10 GIT regions. Alpha diversity (b richness index) and beta diversity (c Bray–Curtis) across the GIT regions. d Relative abundance (%) of dominant phyla, including the bacterial, archaeal, and eukaryotic (e) communities from the proximal to distal GIT. f Relative abundance of major genera across the GIT regions belonging to bacteria, archaea, and eukaryotes. C, Candidatus. Only the dominant phyla or genera with a mean relative abundance greater than 1% at one site were listed. RUM, rumen; RET, reticulum; OMA, omasum; ABO, abomasum; DUO, duodenum; JEJ, jejunum; ILE, ileum; CEC, cecum; COL, colon; REC, rectum