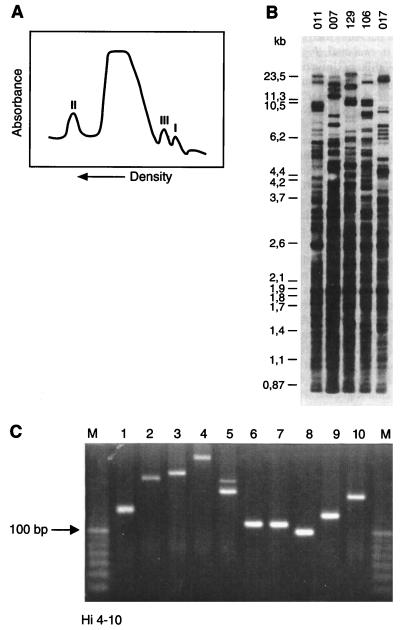
FIG. 3.
Molecular identification of SSR-type DNA. (A) Microdensitometer tracing of human leukocyte native DNA centrifuged to equilibrium in density gradients with an analytical ultracentrifuge. The satellite peaks representing repetitive DNA fractions displaying aberrant densities are indicated (I to III). (B) DNA fingerprints of human individuals generated by probing with repetitive DNA. DNA from five individuals (identified by numerals above the lanes) was digested with a restriction enzyme, the fragments were separated by electrophoresis, and after blotting, the resulting Southern blot was probed with a synthetic oligonucleotide SSR consisting of 10 units of a TTAGG motif. The autoradiograph shows that this SSR is widely dispersed throughout the human genome and clearly depicts the hypervariability in the observed banding patterns. (C) PCR amplification of SSR regions. A specific SSR was identified in the genome of H. influenzae, and primers bordering the repetitive motif were synthesized. When DNA from bacterial strains 1 to 10 was used as template, various amplicons were generated, most of them showing clear differences in length (related to the number of repeat units present). Lanes M contain 10-bp molecular size markers.