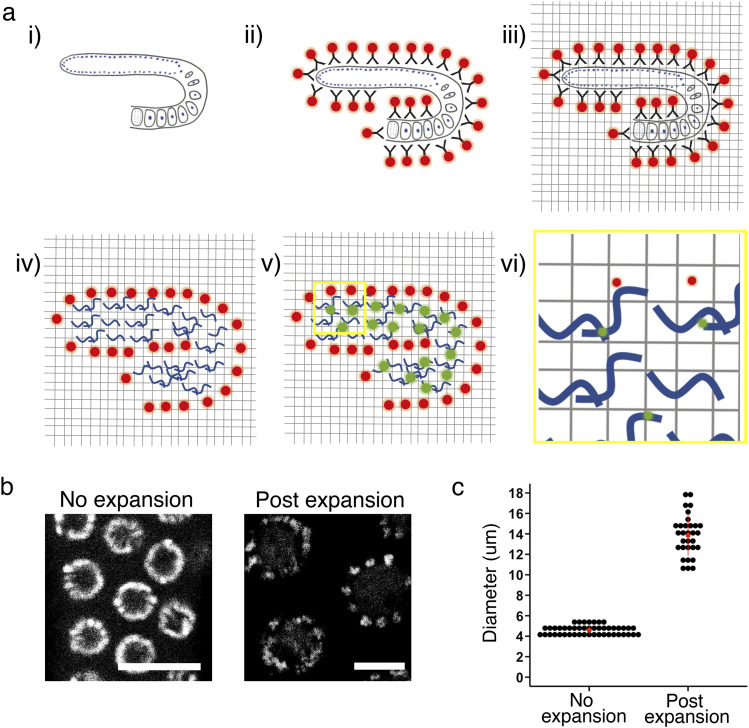
Figure 1. Workflow of expansion microscopy on C. elegans germline.
(A) Schematic of workflow. 3× EExM of immune-stained C. elegans germline with pan-protein stain. (i, ii) Dissected C. elegans germline tissues are stained with primary antibodies followed by fluorescently labelled secondary antibodies. (iii) Tissue is then chemically cross-linked to hydrogel which forms a mold of the tissue. (iv) Proteins within the tissue are digested before (v) pan-protein staining, whereby free amines are labelled with fluorescent dye. (vi) Hydrogel is expanded 3× and imaged. Resulting resolutions are xy ∼40 nm and z ∼100 nm. (B) DAPI staining of the pachytene nuclei in C. elegans germline pre- and post-expansion. Scale bar = 10 µm. (C) Nucleus diameter measured via DAPI staining to determine expansion factor. The average expansion factor across experiments is 3×. 31 expanded nuclei were measured from six independent experiments, and 47 non-expanded nuclei were measured from three independent experiments.
Source data are available for this figure.