Figure 2.
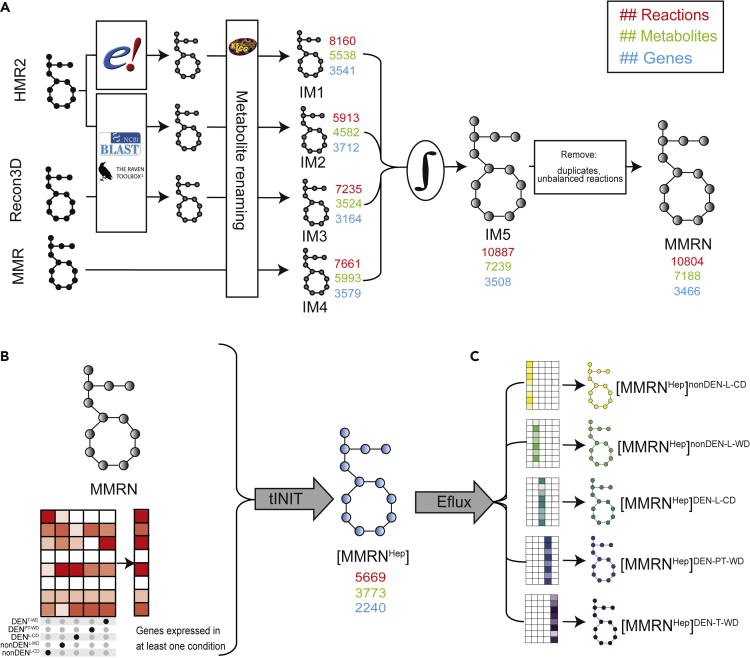
Outline of strategy for the construction of MMRN and derivative context-specific hepatic tissue models
(A) Construction of Mouse Metabolic Reaction Network (MMRN) from four intermediate networks (IM1-4). For IM1, proteins in HMR2 were replaced with their known mouse orthologs from the Ensembl database. The RAVEN 2.0 Toolbox was used to generate IM2 and IM3 based on the sequence homology of mouse proteins to the protein sequences encoded by genes in HMR2 and Recon3D, respectively. Metabolites in IM1-3 were renamed to their corresponding KEGG identifiers (Table S2); similarly renaming metabolites in MMR resulted in IM4. IM1-4 were integrated into a single network, IM5. Duplicate and elementally unbalanced reactions were removed from IM5 to obtain MMRN. The effect of each step on key model attributes is shown by the number of reactions (red), metabolites (green) and genes (blue) adjacent to each model.
(B) RNA-sequencing data were used to identify all genes that were expressed in at least one of the experimental conditions shown in Figure 1A. This list of genes, together with growth tasks and MMRN were used as input for the tINIT algorithm45 to reconstruct a generic hepatic GSMM, [MMRNHep]. MMRN and [MMRNHep] in MATLAB, SMBL and Excel format with corresponding code for simulations are available at https://github.com/sysbiomelab/MMRNHep.
(C) [MMRNHep] was further constrained using an adapted Eflux method (see STAR Methods) and gene expression data for each experimental condition to produce five condition-specific GSMMs (csGSMMs). The Eflux method46 imposes flux boundaries on individual reactions based on TPM expression of all genes predicted to catalyze each reaction.