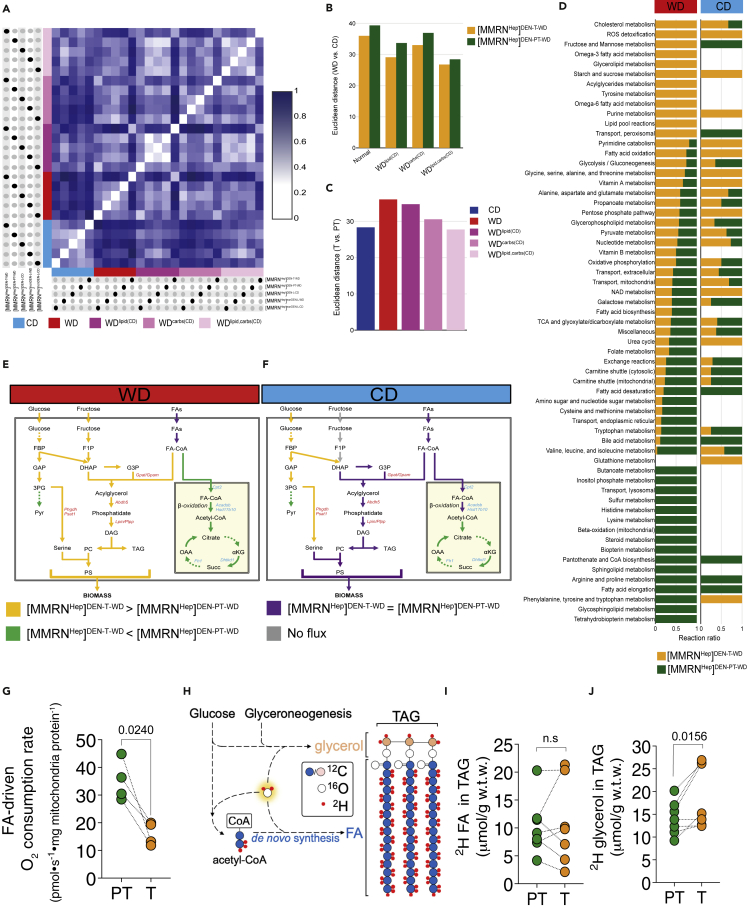
Figure 4.
Gene expression together with dietary nutrient availability dictate differential fate of FAs in tumors and peritumoral liver
(A) Effects of diet composition on flux distribution differences between csGSMMs assessed by SyDiCoS. FBA was used to calculate the flux distribution for each csGSMM provided with CD, WD, WDlipid(CD), WDcarbs(CD) and WDlipid,carbs(CD). The color scale represents the Euclidean distance values calculated in a pairwise manner between each of the flux distributions and plotted relative to the maximum distance value across all comparisons.
(B) Relative response to changes in diet composition of the flux distributions of tumoral or peritumoral models. Absolute Euclidean distances (from panel A) for either [MMRNHep]DEN−T−WD or [MMRNHep]DEN−PT−WD under different SyDiCoS conditions are plotted.
(C) Effect of changes in diet composition on the flux distributions differences between tumoral and peritumoral models. Absolute Euclidean distances (from panel A) between [MMRNHep]DEN−T−WD or [MMRNHep]DEN−PT−WD under different SyDiCoS conditions are plotted.
(D) Subsystems that include at least one reaction that carries flux in [MMRNHep]DEN−T−WD or [MMRNHep]DEN−PT−WD on either WD or CD. In each of these subsystems, the proportion of reactions with higher flux in [MMRNHep]DEN−T−WD compared to [MMRNHep]DEN−PT−WD on either WD or CD is plotted. For each diet, a reaction ratio = 1 for [MMRNHep]DEN−T−WD in a given subsystem indicates that all reactions in that subsystem have higher flux compared to [MMRNHep]DEN−PT−WD.
(E and F) Flux differences between tumoral and peritumoral models fed either WD (E) or CD (F). These two networks are schematic representations of the metabolic network shown in Data S1, which comprises all the reactions of subsystems from panel (D) that have a reaction ratio = 1 (either all reactions that carry higher flux in [MMRNHep]DEN−T−WD or in [MMRNHep]DEN−PT−WD) and partake in lipid and carbohydrate metabolism. Differential fluxes for T and PT are colored according to the legend at the bottom of these panels. FBP: fructose 1,6-bisphosphate; GAP: glyceraldehyde 3-phosphate; 3 PG: 3-phosphoglycerate; Pyr: pyruvate; F1P: fructose 1-phosphate; DHAP: dihydroxyacetone phosphate; G3P: glycerol 3-phosphate; DAG: diacylglycerol; TAG: triacylglycerol; PC: phosphatidylcholine; PS: phosphatidylserine; FAs: fatty acids; FA-CoA: fatty acyl-CoA; αKG: α-ketoglutarate; Succ: succinate; OAA: oxaloacetate; Phgdh: Phosphoglycerate dehydrogenase; Psat1: Phosphoserine aminotransferase 1; Gpat: Glycerol 3-phosphate acyltransferase; Gpam: Glycerol 3-phosphate acyltransferase 1, mitochondrial; Abdh5: 1-acylglycerol 3-phosphate O-acyltransferase; Lpin: Phosphatidate phosphatase; Plpp: Pyridoxal phosphate phosphatase; Mogat1: Monoacylglycerol O-acyltransferase 1; Cpt2: Carnitine palmitoyltransferase 2; Acadsb: Acyl-CoA dehydrogenase short/branched chain; Hsd17b10: Hydroxysteroid 17-β dehydrogenase 10; Dhtkd1: Dehydrogenase E1 and transketolase domain containing 1; Fh1: Fumarate hydratase 1.
(G) Comparison of FA-driven oxygen consumption rates in mitochondria isolated from liver tumors (T) or peritumoral (PT) tissues of mice treated as described in Figure 1A. Statistical significance determined by two-tailed paired t-test (n = 4 different mice, each providing a paired T and PT tissue sample from which mitochondria were isolated; oxygen consumption was measured in parallel for each T/PT sample pair).
(H) Schematic showing metabolic routes of 2H incorporation into the glycerol backbone and fatty-acyl chains in a triglyceride (TAG) molecule after administration of 2H2O to mice.
(I) Measurement of de novo synthesized fatty-acids (as outlined in H) in TAGs extracted from tumor (T) and peritumoural (PT) tissues.
(J) Measurement of de novo synthesized glycerol (as outlined in H) in TAGs extracted from tumor (T) and peritumoral (PT) tissues.
Statistical significance in (I) and (J) determined by Wilcoxon matched-pairs signed rank test (n = 7 different mice, each providing a paired T and PT tissue sample).