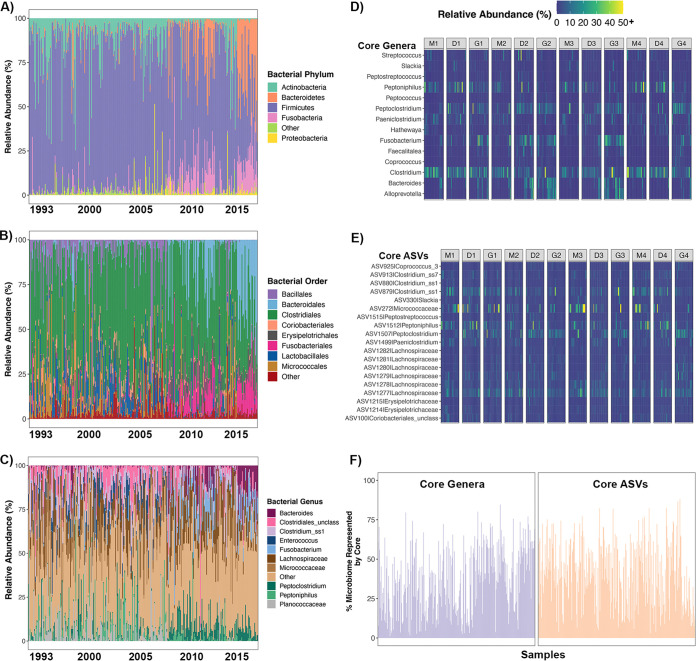
FIG 1.
Amid global and temporal shifts in gut microbiome composition, a taxonomic core was present in the guts of all studied hyenas. (A to C) Stacked bar plots show the relative frequencies of 16S rRNA gene sequences assigned to each bacterial phylum (A), order (B), and genus (C) across samples. Samples are ordered by sampling date, and each color represents a bacterial phylum, order, or genus. (D and E) Heatmap of the relative abundances of 14 core bacterial genera (D) or 19 core bacterial ASVs (E) across samples. These bacterial taxa were found in 85% of samples. Not all sequences were classified to genus or species level, and in those scenarios, the last known classification (e.g., family) was used. (F) Proportion of the microbiome represented by core genera (purple) or ASVs (orange) in each sample.