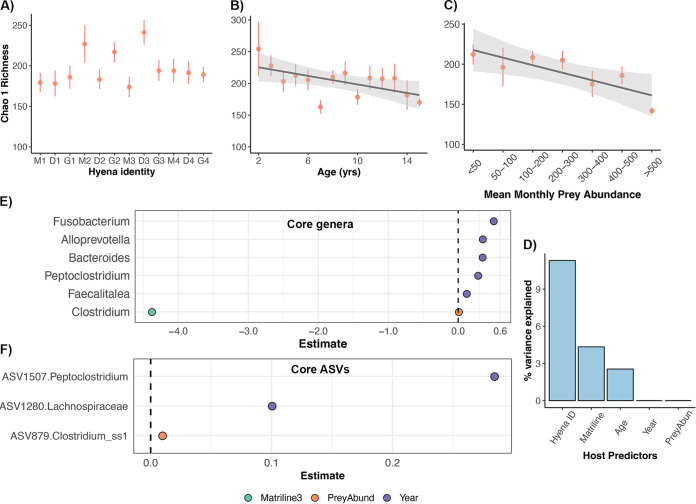
FIG 2.
Socioecological predictors of the gut microbiome in wild spotted hyenas. (A to C) Plots of microbiome Chao 1 richness (mean ± SE) for each hyena individual (A), age category (B), or mean monthly prey abundance (C) for 16S rRNA gene data. The shaded lines with a 95% confidence interval (CI) represent the relationship between x and y, estimated as a linear regression. See Data Set S1 in the supplemental material for statistical output from the linear models. (D) Plots of R2 values from PERMANOVA models testing whether host individual identity, matriline, age, mean monthly prey abundance, and sample year predicted gut microbiome beta-diversity (for 16S rRNA gene data). See Table 2 for exact model output. (E and F) Plots of beta-coefficients from linear mixed models regressing the abundance of core genera (E) or ASVs (F) against the five host predictors listed above. Each point represents a beta-coefficient and is color coded by host predictor. If a beta-coefficient is positive, the abundances of that bacterial taxon were positively correlated with the host predictor. Only beta-coefficients with P values of <0.05 after correcting for multiple comparisons are displayed.