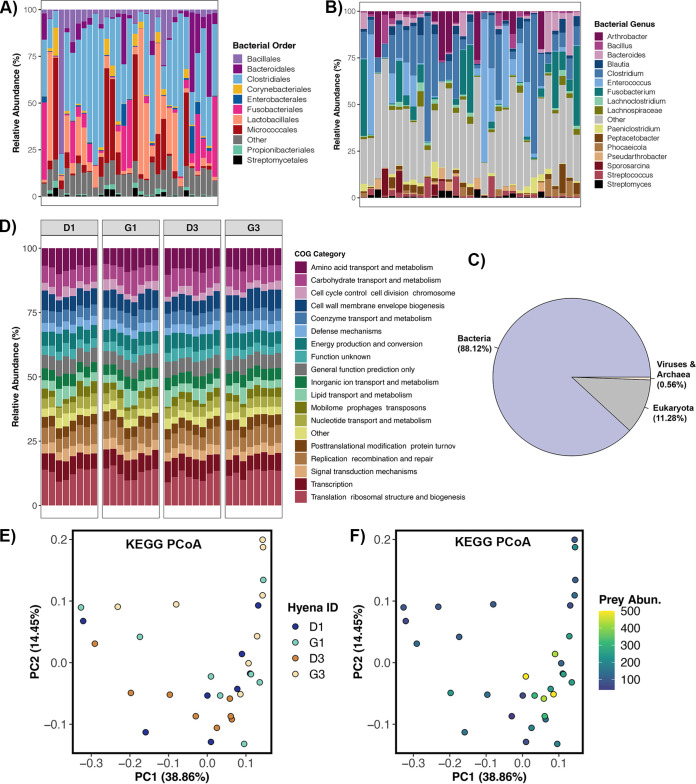
FIG 3.
Composition and predicted function of gut metagenomes obtained from shotgun metagenomic data. (A and B) Stacked bar plots show the relative frequencies of shotgun metagenome sequences assigned to each bacterial order (A) and genus (B) as determined by Kraken2. Samples are organized by hyena individual. (C) Proportion of metagenome sequences assigned to bacteria, eukaryota, and archaea and viruses as calculated by Kraken2. (D) Relative abundances of the most represented COG functional categories across samples. (E and F) PCoA ordinations based on KEGG relative abundances, color-coded by host individual identity (E) or mean monthly prey abundance (F). Contigs were assembled from metagenomic data and imported into Anvi’o for gene prediction and functional annotation. Salmon was used to calculate the relative abundances of genes in each sample (in TPM), and these values were converted to proportions (i.e., relative abundances).