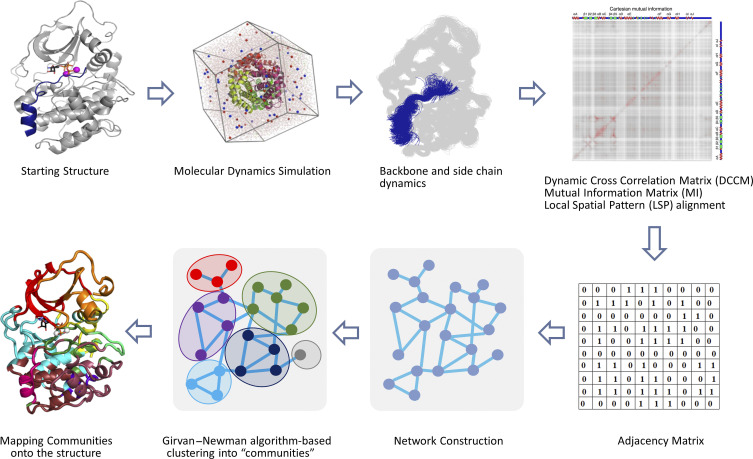
FIG. 4.
Schematic workflow of community map analysis. Amino acid dynamics is obtained by microsecond MD simulations and is assessed for entropic correlations by computing DCCM, MI, or LSP matrices. These matrices, in turn, are used to create an adjacency matrix to reveal a residue network. Girvan–Newman algorithm is used to hierarchically cluster the network into communities based on the property of Edge-Betweenness. Deduced communities are mapped onto the structure to gain a visual assessment of amino acid dynamics.