Figure 3.
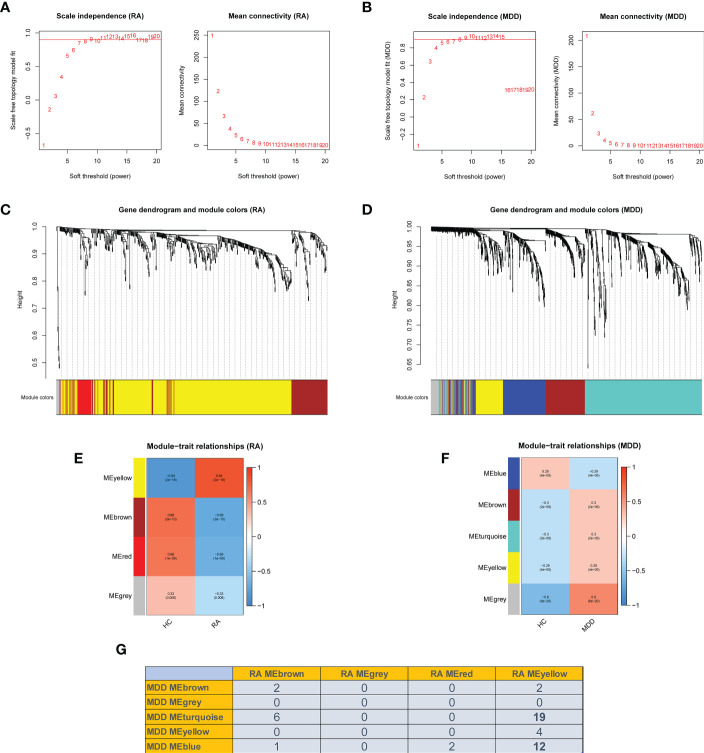
Weighted co-expression network related dataset construction and identification of related key modules in RA and MDD. (A, B) Analysis of network topology for various soft thresholds (β). The left panel shows the scale-free fit index (scale independence, y-axis) as a function of the soft threshold power (x-axis); the right panel displays the mean connectivity (degree, y-axis) as a function of the soft threshold power (x-axis). (C, D) Gene dendrograms were obtained by average linkage hierarchical clustering. The colored row underneath the dendrogram shows the module assignment determined by the dynamic tree cut method. (E, F) Module-trait relationships. Each row in the heatmap corresponds to an ME and each column to a clinical trait. Each cell contains the corresponding correlation and p-value. (G) Number of intersecting genes in related key modules in RA and MDD.