Blood, Vol. 114, Issue 11, 2299-2306, September 10, 2009
Cytokine-mediated increases in fetal hemoglobin are associated with globin gene histone modification and transcription factor reprogramming
Blood Sripichai et al. 114: 2299
Supplemental materials for: Sripichai et al
Files in this Data Supplement:
- Table S1. Primer and probe sequences used for the quantitative real-time PCR amplification of globin genes and selected differentially expressed genes obtained from microarray (PDF, 23 KB)
-
The data showed normalized intensities of each array chip from both low-HbF and high-HbF culture conditions of 40 candidate regulators of globin gene expression selected from general literature reviews. Average (Ave) and SD are showed for comparison. - Table S2. Microarray analysis of candidate globin gene regulators expression in human adult erythroblasts (PDF, 30.4 KB) -
The data showed normalized intensities of each array chip from both low-HbF and high-HbF culture conditions of 40 candidate regulators of globin gene expression selected from general literature reviews. Average (Ave) and SD are showed for comparison. - Table S3. Microarray analysis of transcription factors expression in human adult erythroblasts (PDF, 89.5 KB) -
The data showed normalized intensities of each array chip from both low-HbF and high-HbF culture conditions of 534 probe sets representing 369 transcription factor genes which filtered by raw intensity >1000. Average (Ave) and SD are showed for comparison. The boldface represents the probe sets with greater than 2.5-fold change compared average intensities from low-HbF and high-HbF conditions. - Figure S1. Flow cytometric analysis. Appearance of erythroid progenitors cultures in low-HbF and high-HbF conditions on days 7 and 14 (JPG, 261 KB) -
Flow cytometric analysis of donor matched (A, B) low-HbF and (C, D) high-HbF erythroblasts cultured day 7 (A, C) and day 14 (B, D) stained with anti-transferrin receptor (CD71) and anti-glycophorin A (GPA) antibodies.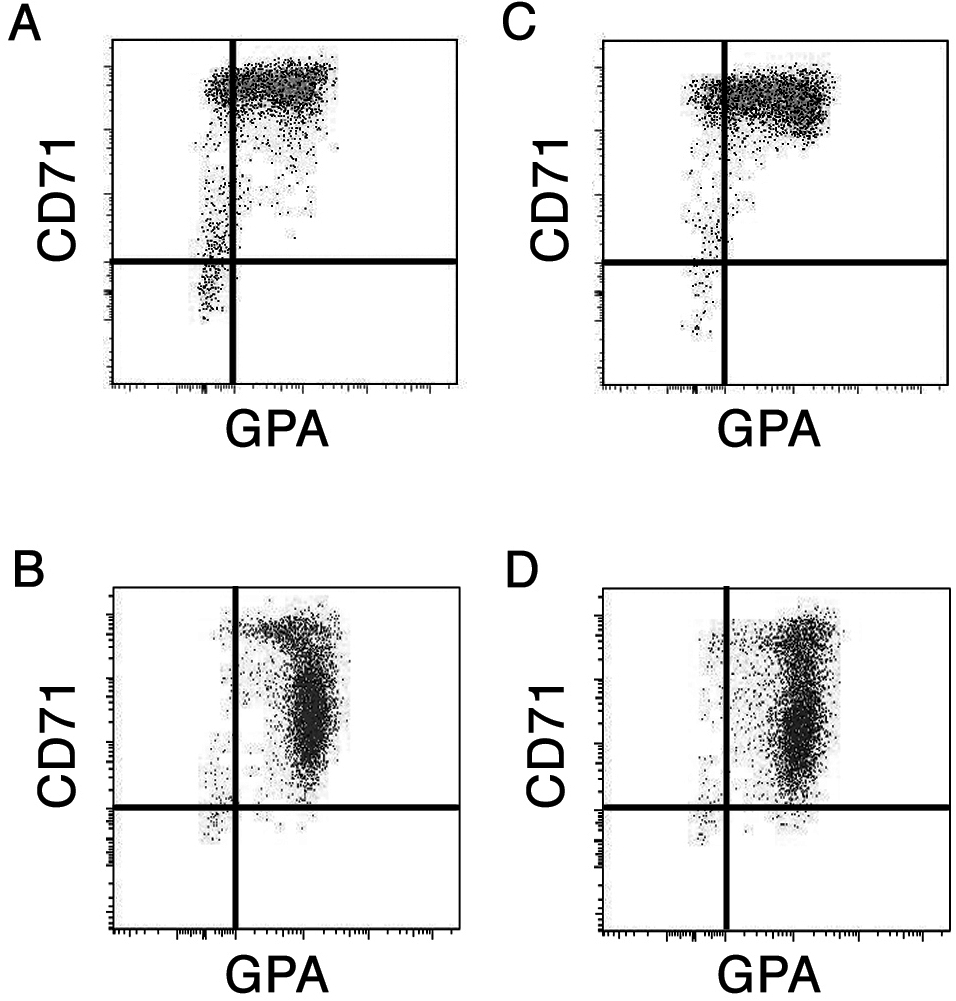
- Figure S2. Changes in histone methylation upon cytokine signal transduction (JPG, 392 KB) -
The ChIP results for histone H3 lysine 4 di-methylation (H3K4me2, red line) and histone H3 lysine 9 di-methylation (H3K9me2, blue line) are compared across the β-like globin genes cluster under (A) low-HbF and (B) high-HbF culture conditions. Relative enrichment was calculated and normalized by setting the highest value obtained for each antibody to one to account for differences in antibody efficiencies.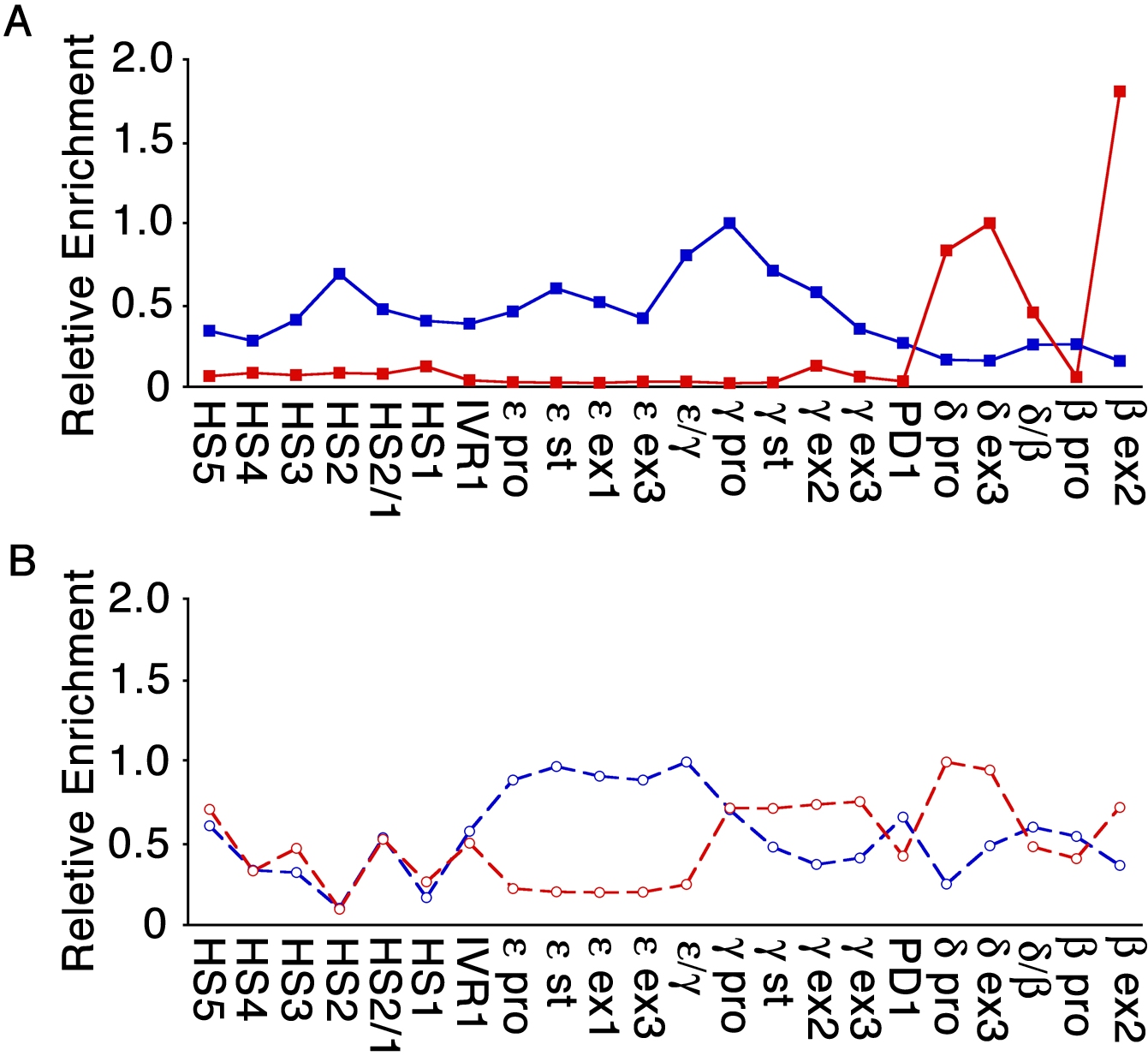
- Figure S3. Microarray experimental design (JPG, 494 KB) -
Primary CD34+ cells from 15 donors were cultured and processed according to the scheme shown in the Panel A. On day 14, HbF expression was validated by HPLC using samples from (B) low-HbF versus (C) high-HbF culture conditions. Percentages of HbF/HbF+HbA ratio (y-axis) calculated from HPLC for individual donors’ (grey bars) and pooled (black bars) erythroblasts are shown. Asterisks signify P
- Figure S4. Microarray and Western blotting analyses of BCL11A gene expression (JPG, 207 KB) -
(A) The figure displays the structure of the BCL11A gene and the Affymetrix HG-U133 Plus 2.0 probe sets binding regions (data from UCSC genome browser; http://genome.ucsc.edu). (B) Expression profiles of BCL11A gene identified by 5 different Affymetrix probe sets. The solid line represents the expression in the low-Hb culture condition and the dotted line represents the expression in the high-HbF condition. (C) Western analyses in 3 different nuclear extracts from low-HbF (L) and high-HbF (H) culture conditions using antibody against residues 1–171 of BCL11A (ab18688, Abcam), detected BCL11A isoforms 1–5. K562 (K) and Jurket (J) whole cell lysates (Santa Cruz Biotechnologies) were used for BCL11A negative and positive controls, respectively. (D) Blotting performed in cytoplasmic (C) and nuclear (N) extract samples from low-HbF and high-HbF culture conditions, showed no detectable signal of BCL11A in cytoplasmic extract.