Figure 2.
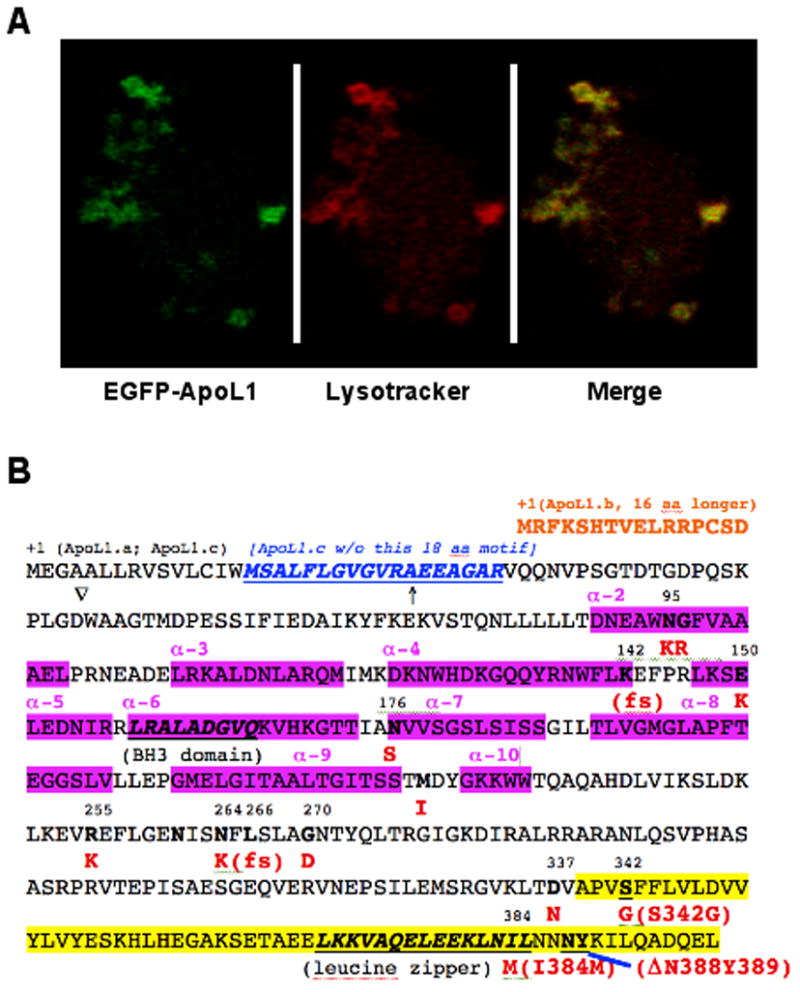
(A). Human ApoL1 localizes in the cytosol and lysosomes. Confocal microscopy analysis showed that EGFP-ApoL1 fusion protein (in Green) and the lysotracker (in Red) were co-localized (in yellow) almost perfectly in human fibroblast cells, suggesting that human ApoL1 is mainly lysosomal. (B). Non-synonymous SNPs, domains and motifs of human ApoL1 isoforms. Three ApoL1 isoforms can be generated by alternative splicing. ApoL1.a, a 398 amino-acid (aa) polypeptide, is shown in its complete sequence. ApoL1.b is 16 aa N-ter longer than ApoL1.a (Shown in orange), while ApoL1.c is missing a 18 aa motif (shown in blue and underlined) close to the N-ter of ApoL1.a. Interestingly, fourteen naturally occurring, non-synonymous variant/alleles of APOL1 have been identified thus far, based on the residue numbers of ApoL1.a, N95K, G96R, K142fs, E150K, N176S, M218I, R255K, N264K, L266fs, G270D, D337N, S342G, I384M and DN388Y389 (amino acid changes are in red). The BH3 domain (codons 158-166) and the leucine zipper domain (codons 365-392) are in bold and underlined. The two risk alleles of APOL1 in ESKD in African Americans, S342G/I384M and DN388Y389, are in the C-ter domain (highlighted in yellow; codons 342 to 398), which interacts with trypanosomal SRA (see text). Interestingly, I384M mutation is present in the leucine zipper domain, which may affect binding of ApoL1 with other proteins in human cells. K142fs and L266fs alleles were identified in an Indian patient whose serum lacked the TLF activity [101].↑, indicates the putative signal peptide, an N-ter 27 aa leader sequence of ApoL1.a); ∇, putative proteolytic cleavage site (between D and W) to generate 39 kDa form of ApoL1. N261, a potential N-glycosylation site. The alpha helixes (α-2 to 10) in the protein are also indicated (in pink).
