Figure 3.
Structure guided optimisation
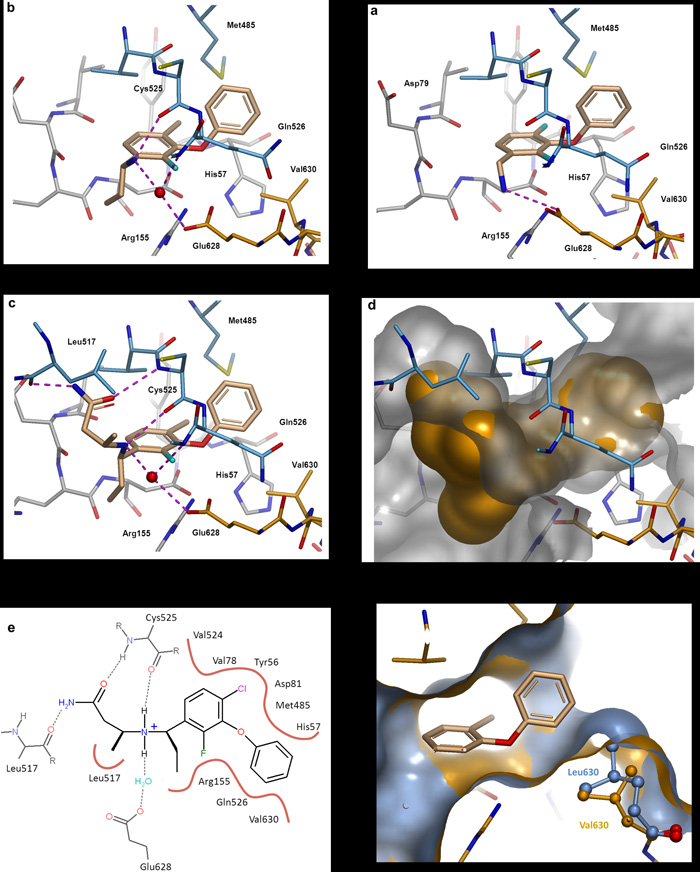
Protein-ligand co-crystal structures of compounds 3 to 5.
a. Compound 3. Flanking fluorine substituents were introduced to stabilise the bound conformation of 2 and also improve hydrophobic interactions. The aminomethyl group was moved to the meta position allowing the positively charged NH3 group to interact with the acidic side chain of Glu628.
b. Compound 4. The 4-fluoro was replaced with a larger chlorine atom to maximize van der Waals interactions. An ethyl group was introduced at the benzylic position to restrict the rotation of the aminomethyl side chain, which now interacts with Glu628 via a water molecule and also forms a hydrogen bond to the backbone carbonyl of Cys525.
c. The X-ray crystal structure of 4 was used to design a small set of compounds to probe the pocket formed by Tyr516, Leu517, Val524 and Cys525. This resulted in the identification of compound 5 which makes key interactions with residues from the protease domain (Arg155, His57) and the helicase domain (Leu517, Cys525, Gln526, Met485) including the C-terminus (Glu628, Val630). H-bonds = dashed lines, conserved water molecules = red spheres.
d. Surface representation of compound 5 and the suface of the allosteric pocket.
e. 2D representation of compound 5 bound in the allosteric site. H-bonds = dashed lines, hydrophobic contacts = solid lines.
f. Surface representation of the wild type protein (gold) complexed with compound 5, overlaid with the crystal structure of the Val630Leu mutant protein (grey). The larger protein side-chain constricts the allosteric binding site.