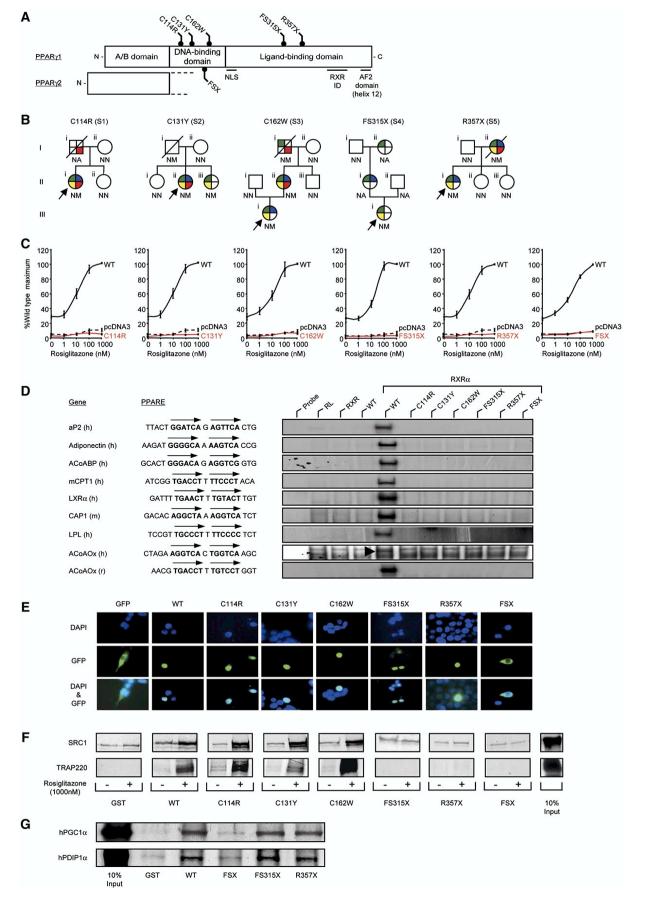
Figure 1.
Identification and characterization of loss-of-function mutations in human PPARγ
A) Schematic representation of the three major domains of PPARγ, showing the locations of the five mutations (C114R, C131Y, C162W, FS315X, and R357X-PPARγ1 nomenclature) and the previously reported FSX mutation. NLS, nuclear localisation signal; RXR ID, retinoid X receptor interaction domain; AF2, activation function 2 domain.
B) Family pedigrees showing genotypes (N, wild-type allele; M, mutant allele; NA, not available) and phenotypes (colored segments denote the presence of specific traits: green, type 2 diabetes mellitus/impaired glucose tolerance/hyperinsulinaemia; yellow, hypertriglyceridaemia; blue, hypertension; red, ischemic heart disease). Squares and circles represent male and female family members; slashed symbols denote deceased family members and arrows denote probands.
C) PPARγ mutants are unable to mediate ligand-dependent transactivation. 293EBNA cells were transfected with 100 ng of wild-type (WT), mutant, or empty (pcDNA3) expression vectors, together with 500 ng of (PPARE)3TKLUC reporter construct and 100 ng of Bos-β-gal internal control plasmid, and increasing concentrations of rosiglitazone. Results are expressed as a percentage of the maximum activation with WT PPARγ1 and represent the mean ± SEM of at least three independent experiments in triplicate.
D) PPARγ mutants are unable to bind to DNA. EMSA with in vitro translated wild-type (WT) or mutant PPARγ1 (C114R, C131Y, C162W, FS315X, R357X, or FSX) and RXR proteins coincubated with oligonucleotide duplexes corresponding to various natural PPAREs. aP2, adipocyte protein 2; ACoABP, acyl coenzyme A binding protein; mCPT1, muscle carnitine palmitoyl transferase 1; LXRα, liver X receptor a; CAP, cbl-associated protein; LPL, lipoprotein lipase, ACoAOx, acyl coenzyme A oxidase; h, human; m, mouse; r, rat; RL, reticulocyte lysate.
E) The C114R, C131Y, C162W, FS315X, and R357X mutants translocate to the nucleus whereas the FSX mutant remains cytoplasmic. 293EBNA cells were transfected as described. Top panels show DAPI-staining (blue) of nuclei, middle panels the cellular localisation of GFP-tagged receptors, and lower panels merged images.
F) The DBD PPARγ mutants recruit SRC1 and TRAP220 coactivators, whereas the FS315X, R357X, and FSX truncation mutants do not interact. GST alone or WT and mutant GST-PPARγ fusion proteins were tested with 35S-labeled in vitro translated SRC1 (upper panel) or TRAP220 (lower panel) in the absence or presence of rosiglitazone. Coomassie-stained gels confirmed comparable protein loading (data not shown). G) The LBD truncation mutants (FS315X, R357X) recruit PGC1α and PDIP1α coactivators, whereas the FSX mutant fails to interact. GST alone or WT and mutant GST-PPARγ fusion proteins were tested with 35S-labeled in vitro translated human PGC1α and human PDIP1α in the absence of ligand. Coomassie-stained gels confirmed comparable protein loading (data not shown).