Abstract
Trypsin and chymotrypsin-like serine proteases from family S1 (clan PA) constitute the largest protease group in humans and more generally in vertebrates. The prototypes chymotrypsin, trypsin and elastase represent simple digestive proteases in the gut, where they cleave nearly any protein. Multidomain trypsin-like proteases are key players in the tightly controlled blood coagulation and complement systems, as well as related proteases that are secreted from diverse immune cells. Some serine proteases are expressed in nearly all tissues and fluids of the human body, such as the human kallikreins and kallikrein-related peptidases with specialization for often unique substrates and accurate timing of activity. HtrA and membrane-anchored serine protease fulfill important physiological tasks with emerging roles in cancer. The high diversity of all family members, which share the tandem β-barrel architecture of the chymotrypsin-fold in the catalytic domain, is conferred by the large differences of eight surface loops, surrounding the active site. The length of these loops alters with insertions and deletions, resulting in remarkably different three-dimensional arrangements. In addition, metal binding sites for Na+, Ca2+ and Zn2+ serve as regulatory elements, as do N-glycosylation sites. Depending on the individual tasks of the protease, the surface loops determine substrate specificity, control the turnover and allow regulation of activation, activity and degradation by other proteins, which are often serine proteases themselves. Most intriguingly, in some serine proteases, the surface loops interact as allosteric network, partially tuned by protein co-factors. Knowledge of these subtle and complicated molecular motions may allow nowadays for new and specific pharmaceutical or medical approaches.
Abbreviations
- CARASIL
cerebral autosomal recessive arteriopathy with subcortical infarcts and leukoencephalopathy
- CUB
complement C1r/C1s/Uegf/Bmp1
- DESC
differentially expressed in squamous cell carcinoma
- EGF-like
epidermal growth factor-like
- FDA
Food and Drug Administration
- FIMAC
Factor I (CI) membrane attack complex
- Gla
γ- carboxyglutamate
- HAP
haemagglutinin protease
- HAT
human airway trypsin
- LDLR
low density lipoprotein receptor
- MAM
meprin/A-5 protein/receptor protein-tyrosine phosphatase mu
- MAPK
mitogen-activated protein kinase
- PAR
proteinase activated receptor
- PDZ
post synaptic density protein (PSD95)/Drosophila disc large tumor suppressor (Dlg1)/zonula occludens-1 protein (zo-1)
- PSA
prostate specific antigen
- TMPRSS
Transmembrane protease, serine
- SCRC
scavenger receptor cysteine rich
- SEA
sea urchin sperm protein/enterokinase/agrin
1. Introduction
Among the known proteases the family S1 from the clan PA stands out with the largest number of members and highly diverse endopeptidase functions, according to its classification in the early 1990s [1]. Bovine chymotrypsin was the first protease, whose structure was determined at atomic resolution (Fig. 1) [2]. Based on the research history, the prototypic chymotrypsin A serves as reference for the numbering scheme proposed by Hartley in 1970 for the catalytic domain [3]. Thus, the catalytic triad residues are designated His57, Asp102 and Ser195 as in chymotrypsinogen A, which are present in all members of the subclan PA(S) [4]. The chymotrypsin fold is even preserved in several viral proteases of subclan PA(C), with His-Asp-Cys or His-Glu-Cys catalytic triads and His-Cys dyads [5]. Most of them are picornains, such as the poliovirus gene 3C product [6], while bacterial and viral serine proteases of 13 other families from S3 to S75 possess His-Asp-Ser triads [7].
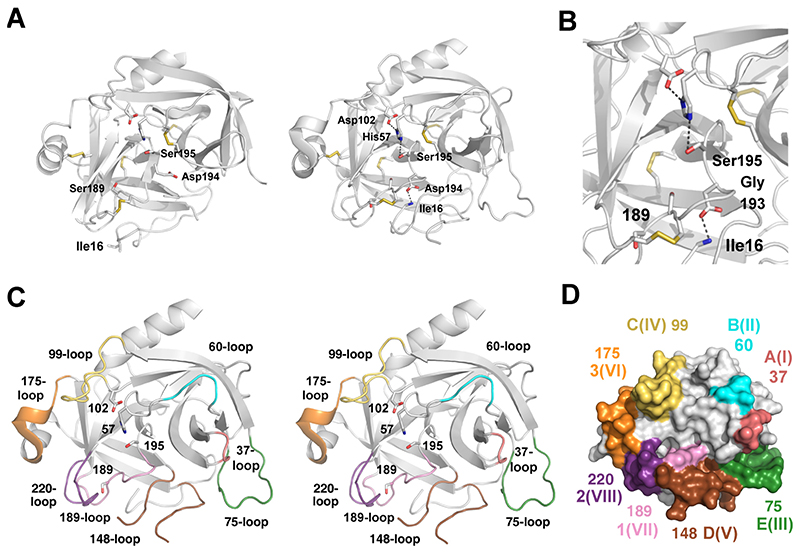
Figure 1. The chymotrypsin/trypsin fold consisting of two β-barrels and eight surface loops.
1A: The left panel shows the crystal structure of the proform or zymogen of bovine chymotrypsinogen A (PDB code 1CHG [288]) and the right panel displays the active form (2CHA [289]), both in ribbon representation. The catalytic triad residues Ser195, His57, and Asp102 are depicted as stick models, as well as the specificity determining Ser189 and the five disulfide bridges. Upon activation with formation of a salt bridge between the new N-terminus Ile16 and Asp194, large rearrangements are seen around the active site, which rigidifies and shapes the S1 pocket. 1B: Close-up view of the active site. Hydrogen bonds are shown as dotted lines. Residue 189 at the bottom of the S1 subsite is the crucial residue that determines the primary specificity of serine proteases, while the oxyanion hole, based on the N-H groups of Ser195 and Gly193 stabilizes the transition state of substrates at the scissile bond. 1C: Stereo representation of chymotrypsin with the eight surface loops in color around the active site. The numbering scheme is mainly based on the central residue of the loop, as derived from bovine chymotrypsinogen A (bCTRA) or by the most common designations. All these loops show significant variations among the S1 family proteases and can serve as regulatory elements. 1D: Molecular surface of bCTRA with the eight colored active site loops and alternative numbering schemes, e.g. I-VIII, or as proposed by Perona and Craik (A–E and 1-3) [14].
A multiple gene duplication encoding a 5-6 amino acid proto-strand has been suggested for the early evolution of trypsin-like proteins, which eventually assembled in two six-stranded antiparallel β-barrel domains [8]. According to the high diversification of functional trypsin-like proteases and their distribution from bacteria to vertebrates, a family tree was assumed that is more than one billion years old [9]. Since most trypsin-like serine proteases genes have the codon TCN (N = A/G/C/T) for the catalytic Ser195 and most blood coagulation factors genes have the corresponding serine codon AGY (Y = C/T), it was hypothesized that the latter ones evolved from proteases with cysteine (codon TGY) and threonine nucleophiles (codon ACN), respectively [7, 10]. Also, in some non-peptidase homologs of trypsin-like proteases the catalytic residues have been replaced by various amino acids residues, such as Arg57 or Asn195 in examples from snake venom [11].
After protein synthesis precursors of the S1 family are translocated to the endoplasmic reticulum with an N-terminal signal peptide that is cleaved by the Sec transmembrane complex. Some of these proteases can already be active, while others are transported as zymogens via the Golgi to secretory vesicles, undergoing posttranslational modifications [12]. Proteolytic cleavage of the propeptide activates the protease after secretion to the extracellular matrix, blood plasma and other body fluids. Usually, the N-terminal Ile16 or Val16 residues in the chymotrypsin numbering scheme insert into the so-called activation pocket (Fig. 1A). Thereby the N-terminal amino group forms a salt-bridge with Asp194, which rearranges and stabilizes the active site, in particular the catalytic triad, the oxyanion hole and the S1 specificity pockets (Figs. 1A, B). The general catalytic mechanism of serine peptidases is described in a concise overview by Polgar in the Handbook of Proteolytic enzymes [13]. Eight surface loops around the active site are the major determinants of specificity and regulation (Figs. 1C, D) [14].
Although the family is termed trypsin-like according to their three-dimensional fold, their members display three major substrate preferences. True trypsin-like peptidases cleave amide bonds after Arg or Lys in the P1 position, chymotrypsin-like ones prefer Phe, Tyr and other large hydrophobic P1 residues, while elastase-like ones accept only small hydrophobic P1 residues, such as Ala, Val or Thr. However, some S1 family peptidase exhibit mixed or more specialized P1 preferences. In addition, the extended specificity up to the P4 and P4′ positions and sometimes further results in an enormous diversity with respect to protease substrates. This specificity depends to a great deal on the eight surface loops around the active site, which confer most of the functional characteristics to individual proteases [15]. Moreover, the focus of this review is on the chymo/trypsin-like serine proteases of the family S1, in particular for the 176 human members, with essential physiological and disease-related functions.
2. Functional classes of serine proteases in the family S1
2.1. Digestive serine proteases from the pancreas, their activators and variants
After proteolytic cleavage in the stomach by the aspartic protease pepsin and its isoforms, three major types of serine proteases cleave proteins into smaller fragments, which are eventually degraded to free amino acids by exopeptidases, such as various amino- and carboxypeptdidases. The activation cascade of pancreatic enzymes is not fully understood, but it comprises more additional pathways and feedback loops as in current textbook schemes (Fig. 2A) [16].
Figure 2. Trypsin-like serine proteases in the digestive tract of the small intestine.
2A: Activation scheme. The membrane anchored, mosaic enteropeptidase (EK, trypsin-like serine protease domain in red, with additional domains) activates trypsin (TRY) by cleavage of the N-terminal propeptide, whereby the neo-N-terminus inserts in the activation pocket, depicted as N within the active protease domain in red. Trypsin activates chymotrypsins (CTR), elastases (ELA), carboxypeptidases (CBP) and tissue kallikrein 1 (KLK1). 2B: Overlay of bovine chymotrypsin B (green, PDB code 1CBW [290]), human trypsin-1 (red, 1TRN [291]) and porcine elastase 2A (yellow, 1BRU). These proteases are highly conserved in mammals, whereby trypsin is seven residues shorter than chymotrypsin, while elastase is eighth residues longer. The most distinguishing features are the primary specificity determining Asp189 of trypsin and Ser189 in chymotrypsin and elastase. The latter possesses a Ser226 instead of Gly226, which excludes the binding of large S1 side-chains. In addition, trypsin exhibits three Glu residues in the 75-loop, which bind the activity-enhancing Ca2+. 2C: Molecular surface of chymotrypsin, trypsin and elastase with electrostatic potential. Depending on the overall short surface loops, most specificity pockets, except the S1 subsite are not well defined and allow interaction with various substrate side-chains. Trypsin has a strong negative potential (red) in the S1 pocket and binds basic P1-Arg and Lys residues. Chymotrypsin prefers hydrophobic, aromatic P1-residues, such as Phe, Tyr and Trp, whereas elastase accepts shorter side-chains, such as Ala, Val, Ile, and Thr.
2.1.1. Trypsins
The prototype of all trypsin-like serine proteases in family S1 is the eponymous trypsin (MEROPS classification S01.151, enzyme catalogue number EC 3.4.21.4) from pancreas, which secretes a mixture of trypsinogen-1 (~65%), -2 (~33%) and -3 (<2%) into the duodenum, the first section of the small intestine [17]. Also with respect to structural data, trypsin is the prototype of family S1 proteases besides chymotrypsin, since the crystal structure of bovine trypsin was determined in 1974 [18, 19].
Enteropeptidase or enterokinase from the hepsin/TMPRSS family (see section 2.6.) is a heavily glycosylated, membrane-anchored “mosaic” protease in the brush border of the duodenal mucosa with tryptic specificity [20]. While the activator of enteropeptidase is still not well characterized, it generates active trypsin-1, trypsin-2 and trypsin-3 by cleavage of the propeptide VDDDDK from Ile16. Then, trypsin activates chymotrypsinogen, proelastase and procarboxypeptidase, a metalloprotease zymogen, before these proteases can degrade proteins from food in a rather promiscuous manner. The relatively low overall specificity of trypsin except for P1-Arg and -Lys residues can be explained by the wide specificity pockets and adjacent short surface loops around the active site (Figs. 2B, C). Small differences in the sequence and structure of the three trypsins confer a slightly altered preference not only for substrates, but also in the interaction with natural inhibitors [17]. Trypsin-3 or “mesotrypsin” is up-regulated in various aggressive cancer types, with an impact on metastasis formation and tumor growth, as it can cleave and inactivate protease inhibitors [21].
Pancreatitis leads in 40% of the patients to pancreatic cancer. Mutations in the trypsin-1 (PRSS1) gene as well as in the serine protease inhibitor Kazal type 1 encoding gene SPINK1 and in the cystic fibrosis transmembrane conductance regulator (CFTR) are implicated in hereditary pancreatitis [22]. Pathogenic mutations in the SPINK1 gene result in a decreased trypsin inhibitor expression, whereas the two trypsin-1 missense variants Arg117His (chymotrypsinogen numbering) and Asn21Ile account for almost 90% of all pancreatitis cases and both are associated with a gain of function, i.e. an increased enzymatic activity, due to enhanced trypsinogen auto-activation [23]. Since the Ala-Arg117 bond is the primary and physiologically relevant autolysis site in trypsin-1, it prevents premature activation in the pancreas and, consequently, the Arg117His mutation with the disfavored P1-His increases the stability of the active enzyme [24]. Other rare mutations lie either within the propeptide of trypsinogen (Ala8Val, Asp11Ala, Asp14Gly, Lys15Arg) or within the mature enzyme (Asn29Thr, Asn29Ile) and tend to an increased auto-activation [25].
2.1.2. Chymotrypsins
Similar to trypsin, chymotrypsin can be considered as the second prototype of the family S1 proteases, in particular, since the first crystal structure of a protease came from bovine chymotrypsin A [2]. However, chymotrypsin A (S01.001, EC 3.4.21.1),) is restricted to cattle, while chymotrypsin B (S01.152) is found in most vertebrates (Fig. 2C). Both share a distinct specificity for P1-Phe, -Tyr, -Trp and other large hydrophobic side-chains, while chymotrypsin A cleaves P1-Trp substrates significantly faster than the B type. Chymotrypsin C (S01.157), with a pronounced preference for P1-Leu residues, adopts an intermediate position between chymotrypsin A/B and elastases, which do not accept large hydrophobic residues [26]. Seemingly, properly active chymotrypsin C (CTRC) can degrade trypsins, which reduces the risk of pancreatitis, and it exhibits, in contrast to most other digestive proteases, N-glycosylation at three Asn residues, which is important for folding and stability. The major function is degradation of trypsin-1 in the pancreas, which is triggered by cleavage between Leu81 and Glu82 in the Ca2+ binding loop of trypsin, however, in addition, an autolytic cleavage of trypsin between the Arg–Val118 peptide bond is required [27]. A secondary function of CTRC is cleavage at the Phe-Asp11 bond of trypsinogen, resulting in the removal of a tripeptide which is accompanied by a 3-fold stimulation of trypsinogen activation and a positive feedback loop, since trypsin activates CTRC as well [25].
2.1.3. Elastases
The name elastase derives from their substrate, the elastic fiber protein elastin with a high content of Ala and Val. Although the specificity of some elastases may overlap with chymotrypsins, they prefer small hydrophobic side-chains, such as Ala, Val or Ile and accept Thr and Asn (Fig. 2C). Despite the name pancreatic elastase I (PE I, S01.153, EC 3.4.21.36), it is mainly expressed in human skin, in contrast to its expression in the pancreas of other mammals, while the major human pancreatic elastases are PE II and PE III, both present as A and B isoforms [28]. Overall, the digestive trypsin-like serine proteases appear to be compact, with short and rigid loops around the active site, which in case of trypsin is further stabilized by Ca2+ binding in the 75 or 70-80 loop, resulting in rather unspecific, promiscuous binding of substrates and rapid turnover (Fig. 2C).
2.2. Blood coagulation factors and fibrinolysis
Enzyme cascades for the activation of serine proteases have been found in insect embryogenesis, in the primordial coagulation system of arthropods, and in the complement system and blood coagulation of vertebrates [29]. Early in evolution a system existed similar to the hemolymph coagulation of invertebrates and diverged about 450 million years ago into the blood coagulation cascade and the innate immune response or complement system [30]. During blood coagulation injured blood vessels are sealed by the intrinsic “contact activation” pathway and the extrinsic “tissue factor” pathway resulting in the prothrombinase complex of the common pathway. This complex activates thrombin, which cleaves fibrinogen to fibrin that forms clots around platelet plugs [31]. In contrast to most digestive enzymes, coagulation factors comprise additional domains in a mosaic-like architecture and are highly specific for a very small number of physiological substrates, with a tight control of activation and activity by cofactors and inhibitors, including complex regulatory feedback loops (Fig. 3A).
Figure 3. Trypsin-like proteases in blood coagulation and fibrinolysis.
3A: Scheme of the blood coagulation system. Intrinsic and extrinsic pathways lead to the common pathway with thrombin as central protease that cleaves fibrinogen monomers, which assemble to fibrin polymers. The protease activation cascades include active tryptic proteases (red), non-protease regulators participate (white), and the crosslinking transglutaminase fXIIIa (grey). Plasmin removes fibrin clots for restoring blood vessels. 3B: Regulation of blood coagulation depends on distinct conformational changes of the active site loops, such as in fXII, whose zymogen conformation involves allostery of the N-terminus with the undefined 189- and 220- loops (dots) (4XE4 [34]). The color scheme for the loops corresponds to Figs. 1C and 1D, e.g. the 99-loop is depicted yellow, the 75-loop is green, etc. 3C: A comparison of fIXa with and without Ca2+ bound in the 75-loop (green) confirmed an allosteric communication line (arrow) to the N-terminus and the 189-loop (pink) (2WPI [40]); same color scheme as in Fig. 3B. 3D: A complex of snake venom homologs is a good model of the fVa-fXa complex or prothrombinase, and resembles the Xase complex of fVIIIa-fIXa (4BXS [47]). The crucial interaction is between the A2 domain (green) and the c175-loop (orange), resulting in an open “unlocked” conformation of the protease. 3E: Bovine thrombin depicted with electrostatic surface potential and fibrinogen peptide bound to the active site (1UCY [292]). Two anion binding exosites mediate interactions with various substrates and regulators. 3F: The structure of full-length plasminogen type II (4DUR [79]) reveals the arrangement of the five kringle domains (various colors), which contribute to fibrin clot and receptor binding. The N-terminal PAp domain (blue) maintains the closed and inactive conformation, as well as the unprocessed propeptide (black), while the catalytic domain (white) is in the zymogen state (catalytic triad residues shown as spheres).
2.2.1. Factors XII and XI
Most coagulation factors are produced in the liver, such as the proenzyme factor XII (fXII, Hageman factor), which is activated to the trypsin-like protease fXIIa by contact with neutral or negatively charged surfaces of natural or artificial origin, in particular during injury of blood vessels [32]. Thus, mature fXIIa starts the enzyme cascade of the intrinsic pathway, enhanced by binding to high molecular weight kininogen and plasma kallikrein, which can activate fXII as well [33]. Factor XII comprises 596 residues with the so-called N-terminal heavy chain and the light chain of the catalytic domain, in which the new N-terminus Val354 (c16 in chymotrypsinogen numbering for the catalytic domain) forms a salt bridge to Asp543 (c194) next to the catalytic Ser544 (c195), whereby both chains remain connected via a disulfide. The heavy chain contains fibronectin II, epidermal-growth factor-like (EGF-like), fibronectin I, another EGF-like, kringle and Pro-rich domains, which mediate binding of fXI, Zn2+, fibrin and heparin, as well as of artificial surfaces. To date, the structure of the mature active catalytic domain is unknown, whereas a construct with the two additional N-terminal residues Ser-Arg353 resulted in a zymogen conformation, lacking a properly formed oxyanion hole and disordered 189- and 220-loops around a distorted S1 pocket (Fig. 3B) [34].
Factor XI is the major substrate of fXII with a different multidomain “mosaic” pattern more similar to plasma kallikrein, exhibiting four apple domains (A1-4) of about 90 residues in the N-terminal heavy chain, whereby A4 provides interacting regions within a disulfide linked homodimer. Cleavage of the Arg-Ile370 bond by fXIIa, thrombin and auto-activation generates a trypsin-like N-terminus [35]. The catalytic domain exhibits significant insertions in the 37- and 60-loops, with Arg37d and Tyr59a contributing to specificity for residues P2´ to P5´, such as in the interaction with the physiological inhibitor protease nexin 2 [36]. Factor XI is encoded on chromosome 4, and genetic deficiencies lead to the recessive bleeding disorder hemophilia C, also known as Rosenthal syndrome, which for gynecological reasons affects more women than men. Hemophilia C does usually not lead to spontaneous bleeding, but can result in excessive blood loss by physical traumata or surgery [37].
2.2.2. Factors IX, X, and VII
The architecture of factors IX and X from the intrinsic pathway differs significantly from fXII and fXI, comprising two EGF-like domains and an N-terminal Gla domain, which contains up to 12 γ-carboxyglutamates, forming four Ca2+ and four Mg2+ sites that allow for binding phospholipids of vascular cell membranes [38]. Gla domain mediated binding of coagulation factors is essential for their localized and highly efficient activity [39]. Full activation of fIX requires the stepwise cleavage at the Arg-Ala146 and Arg-Val181 (c16) bonds, with a complete excision of the stretch Ala146-Arg180. In the catalytic domain of fIX, Glu388 (c219) distorts the S1 pocket and reduces the activity with respect to most other trypsin-like proteases that possess a Gly219, while mutants, such as Glu388Thr are more than 1000-fold active [40, 41]. The stimulatory effect of Ca2+ binding for the activity of the catalytic fIX domain was explained by a conformational communication line from the 75-loop via the Ile16-Asp194 salt bridge to the Cys191-Cys220 stretch, including the catalytic Ser195 and the oxyanion hole (Fig. 3C).
Activation of fX by fIX depends on Ca2+, phospholipids and fVIIIa, a non-protease enhancer of fIXa activity, which can increase Vmax more than 10,000-fold, once the “tenase (Xase)” complex has formed. Most likely, binding of fVIIIa rearranges the 99-loop with Tyr266 (c99) as a particularly critical residue, which can rotate from the S2 into the S4 subsite upon binding of larger P2 side-chains from substrates [42]. Binding of fVIIIa to fIXa opens the “locked” conformation of Tyr345 (c177) on the non-prime side, while the blocking Lys265 (c98) side-chain has to be pushed away by the substrate segment of fX [43]. The hereditary disease hemophilia centers on fIX and its cofactor VIII. The genetic deficiency of fVIII was termed hemophilia A (HA), while the corresponding deficiency of fIX was termed hemophilia B (HB) or “Christmas disease”, which can be both treated by regular injections of the recombinant factors VIII or IX [44]. As both genes are located on the X chromosome, the frequency in males is about 0.02% (HA) and 0.003% (HB), while both are extremely rare in women and heterozygous female carriers show usually only mild symptoms [45].
Resembling its activator fIX from the intrinsic pathway, fX (Stuart-Prower factor) is activated by excision of an activation peptide at the Arg-Ile195 (c16) bond, in order to form the salt bridge with Asp378 (c194), requiring Ca2+, phospholipids, and the non-protease cofactor fVa, to form the so-called prothrombinase complex, which converts prothrombin into active thrombin [46]. Typically, the phospholipids are located on vesicles from activated platelets, whereby in positive feedback loops, fX is capable of activating fVIII, fV, and fVII. First insights into the architecture of the prothrombinase complex were gained by a fX-fV complex from the venom of the Australian Eastern Brown Snake (Pseudonaja textilis) [47]. Besides interactions of the EGF2 domain of fX and the A3 domain of fV, a major interaction site is the 175-loop of the catalytic domain with the A2 domain that might be a parallel to the “unlocking” of fIX (Fig. 3D).
In addition, fVIIa in complex with the non-protease, membrane-anchored tissue factor (TF, thromboplastin) can generate fXa by cleavage of the Arg-Ile153 bond [48]. Essentially, the overall architecture of fVII is the same as in fIX and fX, with one Gla- and two EGF-like domains, whereas its activation does require only a single cut at Arg-Ile195. Factor VII can auto-activate and activate factors IX, V and VIII, while hyaluron binding protease (HABP, FASP) activates fVII as well and cleaves the tissue factor pathway inhibitor (TFPI), thus, accelerating the coagulation cascade [49, 50]. Apparently, fVIIa is the only coagulation factor that is inhibited by Zn2+, which binds to the otherwise Ca2+ binding 75-loop [51]. However, this loop of fVII is capable to bind Ca2+, while two Zn2+ ions bind Lys24, Asp79 and His117 or His76 and Glu80 [52].
2.2.3. Thrombin and protein C
Thrombin (α-thrombin, fibrinogenase, fIIa), the central trypsin-like protease of blood coagulation, cleaves fibrinogen, resulting in fibrin clots, which stop blood loss through injured vessels (Figs. 3A, E). The specificity for the fibrinogen cleavage sites has been structurally elucidated with GVR-GPR as one of the preferred P3 to P3′ sequences [53]. Furthermore thrombin activates fXIII, a transglutaminase, which cross-links Glu and Lys side-chains, in order to render fibrin clots more stable. In addition, thrombin cleaves protease-activated receptor 1 (PAR-1), resulting in the aggregation of platelets (thrombocytes), which clump at vessel injuries supported by the von Willebrand factor (VWF) [54]. Furthermore, thrombin amplifies prothrombin activation in a positive feedback loop by activation of the fIXa and fXa cofactors fVIII and fV [55].
Prothrombin is membrane-bound via an N-terminal Gla domain, followed by two kringle domains. Cleavage at Arg-Ile321 (c16) by prothrombinase (fXa/Va) generates the intermediate meizothrombin, while a second cleavage at Arg-Thr272 releases active α-thrombin, fIIa [56]. Besides the A-glycosylation at cAsn61, the remarkable insertion of nine residues in the 60-loop covers the active site like a lid and shields it from less specific protein substrates [57, 58]. A unique Na+ binding site in the 220-loop enables the transition from “slow” to “fast” thrombin, with formation of the salt bridge cAsp222-cArg187 and the ideal conformation of cAsp189 and cSer195 for substrate recognition and turnover [59, 60]. In contrast to most other coagulation factor proteases, which require Ca2+ binding in the 75-loop for maximum activity, cLys70 precludes such additional stimulation. Regarding the molecular mechanism of substrate binding and turnover, thrombin has served as paragon for the conformational selection mechanism, with an open E and a closed E* conformation, depending mostly on movements of the 220-loop [61]. The anticoagulant effect of the glycosamine heparin is achieved by binding of sulfonic acid groups to the positively charged exosite II of thrombin, which facilitates complex formation with the serpin inhibitor antithrombin III [62]. The dual procoagulant and anticoagulant functions of thrombin were attributed to distinct interactions with either exosite I or II, whereby the electrostatic interactions were only required for the anticoagulant effects [63]. Infections with bacteria from the genus Staphylococus can induce blood clotting by insertion of the N-terminal segment IVTK of staphylocoagulase into the activation pocket of prethrombin-2, with formation of the salt-bridge to cAsp194 and a conformational switch to an active zymogen thrombin species [64].
Activation of protein C, the terminator of blood coagulation, depends on Ca2+ binding in the 75-loop and a specificity switch in thrombin induced by thrombomodulin binding, via the anion binding exosite I in the 37-, 60- and 75-loops [65]. This switch changes the substrate specificity of thrombin from rather basic residues in P3 and P3′ to acceptance of Glu in these positions [66]. Protein C (APC) is a trypsin-like serine protease with an N-terminal Gla domain and two EGF-like domains, whose activity is enhanced by the non-protease protein S and intact fV, while its 60-loop seems to partially block access to the active site [67, 68]. Its exosite I is more extended than the one of thrombin, comprising the 148-loop as important binding region for fVIII [69]. The anticoagulant effects of protein C are achieved by multiple cleavages in fVIIIa and fVa, which suffices to shut down blood coagulation, however, natural mutations of fV, such as Arg506Gln, confer resistance to cleavage by protein C [70].
2.2.4. The plasminogen activators uPA, tPA and plasmin
According to its function in the lysis of fibrin clots, plasmin was originally called fibrinolysin, requiring urokinase-type and tissue-type plasminogen activators (uPA and tPA), which are both highly specific trypsin-like serine proteases as their substrate [71]. Originally isolated from urine as “urokinase”, uPA circulates mostly in blood plasma as zymogen pro-uPA, which is also termed single-chain uPA (sc-uPA) to distinguish it from the active two-chain form (tc-uPA). Various serine proteases, including plasmin, can cleave at the Lys-Ile159 bond of uPA, which has an N-terminal kringle and an EGF-like and domain. By contrast, a FN1-like domain precedes the EGF-like, followed by two kringle domains in tPA. The plasminogen cleavage site CPGR-V(c16)VGG does not represent an optimal recognition sequence for uPA or tPA, which prefer SGR-SA or even YGR-SA, respectively [72]. EGF-like domain mediated binding of uPA to the membrane-anchored uPA receptor (uPAR) is an important activation and regulation cycle, which allows binding of inactive uPA forms or fragments, facilitated by increasing encounters when plasminogen is bound to fibrin at cell surfaces [73]. In addition, activation of plasminogen and inhibition of uPA in the receptor bound state play a role in pathophysiological conditions, such as tumor cell migration.
Both uPA and tPA exhibit insertions in nearly all surface loops with respect to chymotrypsin, such as 37a-d, 60a-c, 110a-d, and 186a-b or 186a-h, which contribute to a limited acceptance of other substrates, but increase the specificity for plasminogen [74]. Remarkably, the zymogen sc-tPA can adopt an active state, enhanced by fibrin binding, when Lys429 (c156) forms a salt bridge to Asp477 (c194), although the properly activated variant with Ile276 (c16) is much more active [75]. The loop extensions of uPA and tPA influence mainly the prime side region and confer the high specificity for the only substrate plasminogen, whereas the extended 37- and 60-loop act as special mediators of inhibition by the serpin PAI-1 [76]. Arg399 (c70) replaces a conserved Glu70 of most trypsin-like proteases and prevents very efficiently Ca2+ binding [77].
Plasminogen features a unique zymogen conformation with a hydrogen-bond from His586 (c40) to Asp740 (c194), precluding formation of the oxyanion hole and the S1 pocket, which is occupied by Trp761 (c215) (Fig. 3F) [78]. Mature plasmin comprises nearly 800 residues, after cleavage by either uPA or tPA at the Arg-Val562 bond and a second cleavage of the Lys-Lys78 bond by plasmin itself [79]. The heavy chain (78-561) comprises five kringle domains (80 residues) and remains covalently linked by two disulfides to the protease domain with the catalytic Ser741 (c195). Varying cleavage events generate other variants of plasmin, e.g. the “microplasmin” protease without kringle domains. Plasmin degrades fibrin clots by cleaving preferentially after P1-Lys and -Arg, exhibiting the unique feature of the so-called 94-shunt, a six residue deletion of the 99-loop, which is the basis of a high substrate promiscuity in the large S2 pocket [80]. The physiological inhibitor of plasmin is the serpin α2-antiplasmin, whereas aprotinin (bovine pancreatic trypsin inhibitor or “Trasylol”) had been used to prevent bleeding during surgery until around 2008 [81]. Additional functions of plasmin have been reported in inflammation, angiogenesis, wound healing, and in tumor growth. In order to enhance blood clot degradation by plasminogen activation, bacterial streptokinase or staphylokinase are employed in medical applications, displaying a similar activation mode as staphylocoagulase with thrombin [64, 82].
2.3. Serine proteases in the immune response
2.3.1. Complement system pathway overview
The complement system is a more than 550 million years old part of the innate immune response, resulting in pathogen lysis or phagocytosis [83]. Primordial organisms like horse shoe crabs with a hemolymph possess a simple version that combines blood coagulation and innate immune response [84]. The mammalian complement comprises eight trypsin-like proteases, various cofactors and pore forming proteins of the membrane attack complex (MAC), which are mostly expressed in the liver and interact with blood coagulation factors and kallikreins [85, 86]. Besides the classical and alternative pathways, additional pathways require lectins, macrophages and polymorphonuclear leukocytes or activated thrombin, that all result in the opsonization, a marking of target cells, and their recognition by complement receptors of immune cells for phagocytosis as well as in MAC formation for cytotoxic processes [87]. A simplified overview of the complicated network is displayed in Fig. 4A [88].
Figure 4. Trypsin-like proteases of the complement system.
4A: Scheme of the complement cascade and the three major pathways. The classical pathway involves proteases C1r and C1s, which activate the C2a component of the C3 convertase. C2a forms the protease component of the C5 convertase as well. The lectin pathway is initiated by mannan binding lectin (MBL) or ficolin, whereby the complex with the MASP proteases generates the C3 convertase C4bC2a. The alternative pathway generates the C5 convertase with factors CD and CB corresponding functionally to C1s and C2. Proteases are displayed red, thioesterase complexes black, and other protein cofactors white. All pathways are more complicated due to complement receptors, additional functions of cleavage products and various interconnections [88]. 4B: The MASP2-C4 Michaelis complex as example of the initiating step of the cascade (4FXG [175]). The trypsin-like protease domain with additional domains is shown in red shades, while the substrate C4 domains are shown in green and yellow. The active site bound segment and the catalytic triad residues are shown as spheres. Except for the alternative pathway, interaction of the serine protease with pathogen binding molecules is required. 4C: Factor C2a, the protease component of the C3 and C5 convertase of the classical pathway, consists of a serine protease domain (white), with many alterations compared to trypsin, and a regulatory VWA domain (green). The regulator helix α7 (dark green) sits at the domain interface with an N-glycan (spheres) at the linker (black) and can together with helix α1 switch to an active, open conformation (2I6Q and 2I6S [99]). Active C2a is still in a partial zymogen-like state, due to the flipped peptide of Lys656-Gly657 of the oxyanion hole, albeit activated by the unusual salt bridge of Arg696 (c224) in the 220-loop (purple) to Glu658 (c194). 4D: Factor CI, a major control element of the complement cascades circulates as active protease, although in a zymogen-like state, as corroborated by a largely disordered N-terminal segment up to residue 325 (c19), including the 75- (green), 148- (brown), 189- (pink), and 220-loops (magenta), which are depicted with their defined residues as spheres, like His57 (2XRC [103]). By contrast, the N-terminal FIMAC, LDLRA1/2 and SRCR domains, which mediate contact to the major substrate C3b, are well defined. 4E: Factor CB structure with three N-terminal sushi or CCP domains (labeled 1 to 3), which require cleavage at the Arg234-Lys235 bond for activation (2OK5 [108]). Active CBb of the alternative pathway resembles largely factor C2a, with a regulatory VWA domain and similar characteristics of the active site, including the unusual salt bridge (c224-c194) and the same oxyanion hole conformation.
The “classical” pathway is initiated by pathogen antigens binding to IgG or IgM antibodies and subsequent recognition by complement factor C1q, which activates the serine protease C1r, followed by activation of the related protease C1s [89]. C1s cleaves factor C4 into the anaphylatoxins C4a/b, as well as C2 into C2a/b, with the protease component C2a, which form the C3 convertase complex C4b2a that cleaves factor C3 into C3a/b, subsequently forming the C5 convertase C4b2a3b. Similar to C4b, the C5 convertase subunit C3b binds covalently to hydroxyl groups of carbohydrates on pathogen surfaces by a thiol nucleophile from a thioester, a central event of complement dependent opsonization [90]. Eventually C4b2a3b cleaves C5, resulting in C5b/6/7/8/poly-9, the pore-forming MAC, which lyzes pathogen cells [89].
The lectin pathway, depends on the mannan-binding lectin-associated serine proteases (MASP), resembling largely the C1r/s proteases. MASP-1 and MASP-2 are mainly activated by complex formation with mannose bound mannan-binding lectin (MBL) on pathogens and generate the C3 convertase C4b2a similar to the active C1 complex (Fig. 4B). In addition, the three human ficolins M, L and H are capable of MASP activation. In the alternative pathway, complement factor B replaces the homologous protease component of C2 in the C3 and C5 convertase complexes, while factor D corresponds to protease C1s of the classical pathway and cleaves factor B in complex with factor C3b, resulting in the C3bBb complex and subsequently in the C3 convertase. Complement protease CI controls all these pathways, since it is able to inactivate the essential C3 and C5 convertase components C3b and C4b to C3i and C4i in the presence of cofactors, such as factor H, C4b-binding protein, complement receptor 1 or CD46 [91].
2.3.2. Complement proteases of the classical pathway
The initiator protease of the classical pathway, C1r, forms a homodimer and comprises 688 amino acids, with two N-terminal CUB domains, an EGF-like domain and two complement control protein (CCP) or sushi domains [92, 93]. Upon IgG/M binding in the inactive, Ca2+ stabilized C1qC1r2C1s2complex, C1r auto-activates by liberation of the N-terminal Ile446 (c16) and subsequently cleaves the zymogen C1s [94]. C1s has the same modular domain structure as C1r and remains in the C1 complex with C1q and C1r once it is activated by release of Ile423 (c16), cleaving the substrates C2 and C4 at Arg-Lys and Arg-Ala bonds, which form the complex C4b2a protease. Recent structural studies indicate that the C1 complex is activated via intermolecular events and that six Ca2+ binding sites in the CUB-EGF-CUB domain segment contribute to the stabilization of the C1r-C1s heterodimer subunits [95, 96]. Homozygous genetic deficiency in C1q is strongly associated with developing the autoimmune disease systemic lupus erythematosus, while C1r or C1s-deficienct patients experience recurrent bacterial, viral, and fungal infections combined with severe cutaneous lesions [97].
The C3 convertase (C4b2a) cleaves C3 into C3a and C3b, which assembles with the C3 convertase to the C5 convertase C4b2a3b [89]. Factor C2 is composed of C2a with the catalytic domain and one von Willebrand factor A-type domain (VWA), whereas the three N-terminal sushi (CCP) domains are cleaved off for activation by C1r [98]. Active C2a exhibits some remarkable differences to the common trypsin-like proteases. Apart from the Mg2+ requiring complex formation with C4b, mediated by the metal ion-dependent adhesion site (MIDAS) of the VWA domain in an open conformation, the activation domain is stabilized by a salt bridge of the side-chain of Arg696 in a loop 2 (220-loop) insertion to Glu658 (c194) [99]. This salt bridge is important for the oxyanion hole conformation, resembling partially a zymogen state, as the peptide plane Lys656-Gly657 (c192-193) is flipped by 180° with respect to the conformation of active trypsin (Fig. 4C). Moreover, insertions of 9, 13, and 15 residues in loops C (99), 3 (175), and 2 (220), as well as deletions of 4 and 12 residues in loops E (75) and D (148) virtually abolish the S2 and S3 specificity pockets. Thus, C2a cleaves in a highly specific manner only in the C3/5 convertase complex at Arg-Ser727 (C3) and Arg-Leu75 (C5) bonds. Individuals with homozygous complement C2 gene deficiency often do not display any obvious disease, very likely due to a factor B-mediated bypass activation of the complement. Still, about 40% of the individuals develop systemic lupus erythematosus or a lupus-like disease [100]. Furthermore, molecular variants of genes encoding C2, C3 and CI, are associated with a high risk for age-related macular degeneration [101].
CI is an atypical trypsin-like serine protease that circulates as active protease and cannot be inhibited by natural inhibitors, whereas it cleaves small synthetic substrates [102]. It exhibits a unique architecture with the FIMAC domain that enables membrane insertion, one SCRC and two LDL-receptor class A domains preceding the protease domain, which exhibits a zymogen-like active site with disordered loops 148, 191 and 220, albeit with a free N-terminus at Ile322 (c16) [103]. It has been suggested that an allosteric auto-inhibition occurs by contacts of the N-terminal heavy chain to the C-terminal helix and opposite to the active site, while activity might be substrate-induced as observed for thrombin (Fig. 4D). CI cleaves at Arg-Thr/Ser/Asn/Glu bonds, whereby changes in specificity depend on the cofactors [104]. Under certain conditions CI activity can result in phagocytosis instead of lysis of pathogens.
2.3.3. Complement proteases of the lectin and the alternative pathway
For MASP-1 some parallels to thrombin have been described, in particular heparin binding and a more accessible active site than in the self-inhibited factor D, despite a similar salt bridge between Asp640 (c189) and Arg677 (c224) [105]. As thrombin, pro-MASP-2 displays a significant zymogen activity, which may depend on the similarly extended 60-loop, while its substrate specificity is nearly identical to that of the C1 complex [106, 107]. In the alternative pathway, complement factor B essentially replaces the highly homologous C2 in the C3 and C5 convertase complexes. A full-length structure of CB comprising the three CCP or sushi domains explained the zymogen state by a closed conformation of the MIDAS motif in the VWA domain, which shields the Arg234-Lys235 bond (Fig. 4E) [108]. In addition, Arg705 as equivalent of Arg696 in factor C2a, is located more than 10 Å away from Asp673 (194), preventing an active conformation of the catalytic domain as present in factor Bb [109]. Recently, MASP-3 has been characterized as link between the lectin and alternative pathway, as it is able to activate pro-factor D, which circulates mostly as active protease in blood [110].
In the alternative pathway, factor D corresponds to C1s of the classical pathway. Pro-factor D exhibits an intact catalytic triad, but a typically distorted activation domain as observed for other zymogens [111]. Factor D is highly specific for cleaving only the Arg-Lys235 bond to activate protease factor B. The active C3bBD* complex structure revealed that the residues c213-c218 of the 220-loop adopt a self-inhibitory conformation, where Ser215 forces the catalytic His57 out of the triad [112]. In addition, Arg218 forms a salt-bridge to Asp189 in the self-inhibited state. Binding to the substrate C3bB is mediated by a positively charged exosite of factor D, consisting of loops D and 1 to 3, which essentially correspond to the 148-, 175-, 191-, and 220-loops. Very low blood plasma-concentrations of factor D define its role as rate-limiting enzyme of the alternative pathway and the loss of complement activation in CD deficiency increases the susceptibility to invasive meningococcal disease [113]. By contrast, increased levels of CD in age-related macular degeneration make it a potential therapeutic target for the treatment of this disease and its advanced form geographic atrophy [114].
2.3.4. Neutrophil serine proteases
Neutrophil elastase or leukocyte elastase (HNE or HLE) is present as mature protease in storage “azurophil” granules of polymorphonuclear neutrophils (blood), together with myeloblastin/proteinase 3 [115]. Their major task is killing pathogens, such as phagocytosed bacteria, and regulating inflammatory processes together with other neutrophil serine proteases. HNE and proteinase 3 (PR3), also named myeloblastin, participate with histones, antimicrobial peptides and DNA fibers in so-called neutrophil extracellular traps (NETs), which catch and kill bacteria and fungi. Both proteases prefer Val, Thr, Ala or Met in P1 position, which can be explained by the unusual Gly189/Val190 or Ile190 combinations, whereby variations in the loops, such as Asp61 and Lys99 confer a distinct substrate specificity for basic P1′ and acidic P2 residues [116]. An inhibitor complex structure of HNE showed that only two of the three known N-glycosylation sites at Asn72, 109, and 159 were occupied, which, nevertheless are important for the interaction with the physiological inhibitor α1-antitrypsin [117]. PR3 can cleave cathelicidin and thereby generate the antimicrobial peptide LL-37, activates tumor necrosis factor alpha (TNF-α), interleukin-β (IL-β) and transforming growth factor β (TGF- β). A non-protease homolog of PR3 is azurocidin, without catalytic triad, due to the mutations His57Ser and Ser201Gly, while it alarms and enhances the immune system and its response, respectively [118, 119].
Like HNE and PR3, cathepsin G (CG) from neutrophils contributes to killing microbes in phagosomes. CG is a chymotryptic protease with a preference for P1-Phe, while P1-Lys can be turned over due to the presence of Glu226, similar as in chymase from rat mast cells [120]. An unusual structural feature is the missing disulfide Cys191-Cys220, which is otherwise highly conserved, and the rather hydrophobic combination of Ala189/Ala190. Regarding the loops, HNE exhibits a three residue insertion in the 37-loop, whereas CG has such an insertion in the 60-loop. According to the architecture of the S1 subsite, HNE and PR3 are more chymotrypsin-like, whereas CG has a mixed specificity cleaving after large, hydrophobic and basic amino acids [115]. The fourth member of the neutrophil serine protease family, NSP4, displays similar to HNE and PR3 a Gly189/Phe190 combination in the S1 pocket [121]. Although NSP4 shows an elastase-like active site, it is trypsin-like with a preference for P1-Arg. Its low abundance in neutrophils suggests a more regulatory role in the innate immune reactions, instead of digesting mostly bacterial proteins [121, 122]. The physiological activity of neutrophil serine proteases is tightly controlled by α1-protease inhibitor, α2-macroglobulin, secretory leucoprotease inhibitor (SLPI), and elafin to maintain tissue homeostasis. However, unbalanced protease activity can cause serious inflammatory lung diseases, such as chronic obstructive pulmonary disease (COPD) and cystic fibrosis [115].
2.3.5. Serine proteases from mast cells
Human mast cells belong to the immune system, defending the host against intruding pathogens, such as bacteria and parasites, with chymotryptic chymases and tryptic tryptases as prominent serine proteases, and other proteases that are stored as zymogens in secretory granules, which release their content when immune signals initiate degranulation [123]. Chymase, an α-chymase, is the only human member of these specialized serine proteases, which are more diversified and better characterized in rodents, with up to five different α- and β-chymases [124].
Four human tryptases are known: α-tryptase (tryptase alpha 1) prefers P1-Lys over P1-Arg, like β-tryptase (tryptase beta 1/2/3), γ-tryptase (tryptase gamma 1) favors P1-Arg, while δ-tryptase (tryptase delta 1) is inactive [125, 126]. Interestingly, α-tryptase is a constitutively secreted protease with very low activity, due to the substitution of Gly216 by Asp, whose absence in some humans has not effects on health [127]. β-tryptase is known in three isoforms, e.g. βII, which all form ring-like tetramers with the active sites located inside the water filled cavity. Tetramerization of β-tryptase is a prerequisite for its enzymatic activity and most likely the cause for resisting all known physiological inhibitors [128, 129]. Six surface loop stretches bind neighbor molecules, whereby the 60-loop and the 175-loop are extended by five and nine residues. Only γ-tryptase is anchored with a C-terminal peptide in membranes, whereas a truncation of 40 C-terminal residues renders δ-tryptase inactive [130]. Tryptases and other mast cell proteases have been implicated in a plethora of diseases, including cardio-vascular, respiratory, digestive, reproductive, nervous system disorders, and oncogenesis [123, 131].
2.3.6. Granzymes of cytotoxic T lymphocytes
The name “granzymes” derives from granules of T lymphocytes and trypsin-like enzymes, which play a role in the destruction of pathogens in blood together with the pore forming perforin [132]. The formation of large membrane pores, allows granzymes to enter the cytosol of target cells, where they initiate apoptosis, which is important in killing tumor cells [133]. To date twelve granzymes (GZM) are known, of which GZMA, -B, -H, -K, and – M are present in humans, which are usually activated by cathepsin C (DPPI) [134].
Granzyme A was described in natural killer cells (NK), lymphokine activated killer cells (LAK) as major executor of apoptosis, although its various roles in human have not been fully revealed. GZMA prefers Arg over Lys in the P1 position and forms a unique disulfide-linked homodimer via Cys93 in the 99-loop. Originally classified as chymotrypsin homolog due to the presence of Thr189, Granzyme B turned out to be one of the few trypsin-related peptidases with a primary preference for acidic residues, such as P1-Asp [135]. The different architecture of the wider S1 subsite in GZMB, which lacks the highly conserved disulfide Cys191-Cys220, results in the formation of a salt bridge from Arg226 to P1-Asp residues [136].
In contrast to GZMA and B, GZMH possesses the unique Glu14-Glu15 pro-dipeptide and exhibits a distinct chymotryptic specificity for P1-Phe, -Tyr and –Met, which can be explained by Thr189 at the bottom of the S1 pocket and a Gly226 instead of an Arg226 as in GZMB [137]. The major task of GZMH is killing target cells upon entering the cytosol and the cleavage of proteins that are critical in viral infections, e.g. in hepatitis B [138].
Besides GZMA and based on the presence of an Asp189, GZMK is the only tryptic granzyme, which equally accepts Lys and Arg in P1 position according to various scanning approaches, which related the extended specificity to the natural substrate tubulin, as present in the microtubulin network of tumor cells [139]. The fifth human granzyme, GZMM displays a preference for hydrophobic P1-residues without Cβ-branching, such as Met and Leu. This special preference can be explained by the unusual combination of Ala189-Pro190 and the presence of Ser216 instead of Gly216. Overall, the humans granzymes show no striking aberrations in the surface loops, although individual GZMs have significant insertions in the 60-loop (GZMK/M), deletions in the 75-loop (GZMM) and insertions in the 175-loop (GZMA/K). GZMA and GZMB are the major granzymes implicated in cell death processes, whereby GZMB represents the most powerful pro-apoptotic granzyme which is most likely based on the unusual preference for P1-Asp substrates, similar to caspases. GZMB-mediated cell death is rapid in contrast to a slower form of cell death, which is initiated by the tryptic GZMA in a caspase-independent pathway, called athetosis, depending on an intact actin cytoskeleton as shown in a mouse model [140].
2.4. Kallikreins and kallikrein-related peptidases (KLKs)
Basically, kallikreins and KLKs are expressed in all tissues and fluids of the human body, however, their individual tissue expression varies extremely with development, healthy or pathological conditions [141–144]. They are closely related to trypsin and the historical distinction of the classical kallikreins 1-3 and the new kallikreins 4-15 is justified by the peculiar 11 residue insertion in the 99-loop, giving rise to the name “kallikrein loop”. It is preferable to classify them according to their functional tasks, which are associated with proteolytic activation cascades (Fig. 5A) [145–147]. KLK3/PSA is the best known member of this group, since it is employed as biomarker and therapeutic target for prostate cancer. Moreover, most KLKs are implicated in cancer and other diseases, which is described in detail by several comprehensive reviews with respect to pathophysiological roles and clinical relevance [148–152]. Similarly, structure and functions have been reviewed by various research groups [153–155].
Figure 5. Kallikreins and kallikrein-related peptidases.
5A: Tissue kallikrein and plasma kallikrein KLKB are kininogen cleaving regulators of blood pressure, while the prostatic KLKs 2, 3, 4, 5, and 11 are either part of an activation cascade and/or degrade gel forming proteins to enhance sperm motility for impregnation. In brain, KLK6 and 8 participate in the regulation of glia cell and synaptic remodeling, whereas the skin related KLKs 5, 7, and 14 activate themselves and cleave cell connecting molecules for skin cell shedding. 5B: KLK2 is glycosylated at Asn95, depicted as core glycan, which favors the closed conformation of the 99-loop (yellow) with an 11-residue insertion with respect to bCTRA. KLK3 has an N-glycan linked to Asn61 (not shown) and a 99-loop, which can cover the non-prime-side from S4 to S2 like a lid, as in KLK2 (lower panel). The eKLK3 structure represents an E* form, with closed 99- (yellow), 148- (brown), 189- (pink), and 220-loops (magenta), but intact catalytic triad and Ile16-Asp194 salt bridge (1GVZ [165]), color scheme as in Figs. 1C and 1D. 5C: KLK4 exhibits a unique Zn2+ binding site that connects the 75-loop with the N-terminal segment via the ligands Glu74 and His25, with an additional cation binding Glu74* ligand from a neighbor molecule (2BDH [174]). Zn2+ binding causes a conformational change to a zymogen-like conformation of the active site via an exposed N-terminus, perhaps mediated by the disulfide Cys22-Cys157, as suggested for fVIIa. 5D: KLK5, 7, and 8 possess inhibitory Zn2+ binding sites, involving His99 and other ligands, such as His96 (2PSX and 2PSY [203]). The inhibition process is depicted in three steps: The two His side chains of the 99-loop in KLK5 are ready to bind Zn2+ in solution (1), an intermediate state with His96 and His99 bound to Zn2+ was observed for KLK5 (2). Eventually, the inactive state with a disrupted catalytic triad (3) is represented by tonin, a KLK2 ortholog from rat, where Zn2+ is bound by His57 (1TON [180]). 5E: KLK8 exhibits both a stimulatory Ca2+ binding site in the 75-loop and the aforementioned Zn2+ inhibition site in the 99-loop. Structure based molecular dynamics simulation corroborated that these loops are connected by a variant of the communication line of the coagulation factors (5MS3 [193]). Significant concerted loop movements upon Ca2+ removal with respect to the crystal structure are depicted as spheres for the 37- (red), 75- (green) and 99-loops (olive).
2.4.1. Kallikreins
In humans, tissue and plasma kallikreins reduce blood pressure by processing low and high molecular weight kininogens (LMWK, HMWK) to the vasodilators lysyl-bradykinin (kallidin) and bradykinin, hormone-like deca- and nonapeptides with additional functions in inflammation and pain (Fig. 5A) [156]. Essentially, tissue kallikrein (KLK1) belongs to the 15 membered family of the kallikrein-related peptidases (KLKs), whereas plasma kallikrein (PK, KLKB1) is only distantly related and participates in blood coagulation [143, 157].
Plasma kallikrein is mostly expressed in liver and activated in blood by coagulation factor XII, where it cleaves HMWK at the Lys-Arg bond, facilitated by the four apple domains, which are also present in fXI [143, 158]. A comparison of a ligand-free catalytic domain of human PK with an inhibitor-bound murine PK demonstrated that the conserved Glu217 in the 220-loop adopts a closed conformation, blocking the active site by a hydrogen bond to the catalytic Ser195 [159]. The physiological functions of PK are hemostasis, e.g. by activation of fXII, which in turn activates PK, inflammation and immune response, whereas a pathological deficiency leads to various cardiovascular symptoms [157].
In contrast to PK, tissue kallikrein or KLK1 is produced mostly in pancreas or kidneys and can cleave kininogens at the Met-Lys and Lys-Arg bonds, releasing lysyl-bradykinin or bradykinin from LMWK and HMWK [157]. A distinguishing feature of KLK1 is the N- and O-glycosylation at six positions, three alone in the 99-loop [160]. The structure of a recombinant ligand-free KLK1 displayed a partially disordered 99-loop with only a single short glycan linked to Asn95, while the heavy physiological glycosylation may contribute to the control of substrate turnover as in KLK2 (see below) [161].
2.4.2. Prostatic KLKs in human reproduction
Remarkably, the expression of KLKs 2, 3, 4 and 11 in the male reproductive system, including prostate, seminal fluid, and testes, where KLK5 expression is high, appears to be complementary to the expression of other KLKs in the female reproductive system, such as uterus and cervico-vaginal fluid [144]. KLK2 seems to be an auto-activating tryptic protease and is most likely the physiological activator of KLK3/PSA in seminal fluid. Both KLKs exhibit a similar propeptide with the scissile bond Arg-Ile16. Their common task is the degradation of semenogelin-1 and -2, resulting in liquefaction of the seminal clot, which is required for sperm motility prior to fertilization (Fig. 5A). Zymogen-like and mature protease conformations in crystal structures of the chymotryptic KLK3 confirm the insertion mechanism of the N-terminal amino group into the activation pocket with formation of the salt bridge to Asp194 [162]. However, the flexible kallikrein/99-loop of KLK2 and KLK3 can adopt an open or a closed conformation, which completely shields the S2 to S4 specificity pockets [163, 164]. A concerted transition of the 99-, 148-, and 220-loops in KLK2 from the closed E* to the open E-form is the best explanation for a conformational selection mechanism, similar to the situation in thrombin [61]. Horse KLK3 (eKLK3) adopts the E* form with closed 99-, 148-, 189- and 220-loops, despite an intact N-terminal salt bridge and catalytic triad [165]. The eKLK3 structure suggests that the inhibitory Zn2+ binding site of KLK2 and KLK3 is formed by His91, His101 and His233. An N-glycan linked to Asn95 of KLK2 is the crucial element in regulating the substrate access and turnover by favoring the closed conformation and protecting from cleavage (Fig. 5B) [166].
According to their substrate profiles, the tryptic proteases KLK4 and KLK11 could be additional activators of pro-KLK2 and pro-KLK3 in a corresponding proteolytic cascade,, while KLK3 and KLK11 cleave the unusual Gln-Ile16 bond of pro-KLK4 [146, 167, 168]. However, glycosylated KLK2 was the most efficient pro-KLK4 activator, not KLK3 [169]. Activation of KLK4 in other body fluids and tissues, such as developing teeth, depends most likely on matrix metalloproteinases, in particular MMP-3 and MMP-20 [170, 171]. Based on its hippocampal expression, KLK11 was named hippostasin, while the prostatic isoform-3 exhibits a 25 residue long insertion in the 60-loop, which leaves the enzyme activity intact [172].
Except for KLK11, Zn2+ inhibition at micromolar concentrations has been confirmed for the prostatic KLKs, which is in the physiological range since Zn2+ in prostatic fluid reaches millimolar concentrations [173]. KLK4 has a distinct role in enamel formation, as corroborated by the genetic disease amelogenesis imperfecta, which is characterized by malformed teeth, due to an inactive version of KLK4. Mutational analyses localized the inhibitory Zn2+ binding site for KLK2 in the kallikrein/99-loop and in KLK4 at His25 near the N-terminus and Glu74 in the 75-loop (Fig. 5C) [164, 174]. In case of KLK4, the inhibition mechanism appears to be similar as in fVIIa, in which Zn2+ bound to the 75-loop switched the mature protease to a zymogenic state with an exposed N-terminus [51]. The distantly related, trypsin-like and Zn2+-regulated acrosin is stored in sperm heads for the acrosome reaction for fusion with the ovum, featuring a positively charged N-terminal domain with patches for membrane interaction, similar to coagulation factors [175].
2.4.3. Skin derived KLKs 5, 7, and 14
Early studies designated KLK5 and KLK7 as stratum corneum tryptic and chymotryptic enzymes, referring to the primary specificity of both enzymes, with a distinct preference of KLK7 for P1-Tyr [167]. Together with the tryptic KLK14, these KLKs form a proteolytic activation cascade, which can start with pro-KLK5 auto-activation, facilitated by its 37 residue long propeptide [176]. All three KLKs degrade cell adhesion molecules in the corneodesmosomes of the corneocytes for desquamation, i.e. skin cell shedding (Fig. 5A). Dysregulation of KLK activity occurs in in Netherton syndrome and atopic dermatitis, involving mutations in the natural lymphoepithelial Kazal inhibitor-1 (LEKTI-1) [177, 178].
KLK5 possesses a specific Zn2+ binding site in the 99 loop, as observed by binding the ligands His96 and His99, similar to rat kallikrein tonin, with His57 as third ligand, explaining the inhibition by disruption of the catalytic triad (Fig. 5D) [179, 180]. Although KLK7 lacks a His96, the physiologically relevant cations Cu2+ and Zn2+ inhibit at the same position, as confirmed by the His99Ala mutant [181]. Both KLK5 and KLK7 possess shorter and more compact loops than the other KLKs, exhibiting a common positively charged area, corresponding to exosites I and II of thrombin [153]. A comparison of the free mature KLK7 (PDB 3BSQ) and an inhibitor bound structure (2QXH) suggests that both are virtually identical as in the lock-and-key model of enzymes, with a rigid, open E conformation that cannot adopt a closed E* form [181–183]. As most kallikrein-related peptidases, KLK14 is expressed in many tissues and body fluids, whereas its expression and function change in tumor tissue, such as becoming an activator of PAR-2 [184, 185]. Basically, all human KLKs have been implicated in PAR-1/2/3/4 signaling in inflammation and cancer processes [186].
2.4.4. Central nervous system KLKs 6 and 8
Mainly KLK6 and KLK8, also known as neurosin and neuropsin, are expressed in human brain. KLK6 resembles trypsin in many aspects and has a specific role in neuronal inflammation and multiple sclerosis, e.g. by degrading myelin basic protein of glial cells [187, 188]. Recently, it was demonstrated that KLK6 can remodel glial cells of the central nervous system, which can be complementary to the role of KLK8 in synaptic remodeling of hippocampus neurons (Fig. 5A)[189, 190].
Since rodent and human KLK6 and KLK8 are highly conserved, corresponding studies have contributed to the identification of neuregulin-1 or EphB as KLK8 substrates [191, 192]. KLK8 is rather unique with respect to binding stimulatory Ca2+ in the 75-loop like trypsin and inhibitory Zn2+ in the 99-loop, which was corroborated by crystal structures and a mutational analysis [193]. Additional features are N-glycosylation at Asn95 and the presence of Cys93 as in granzyme A, which dimerizes via a disulfide [194]. Similar to KLK4, molecular dynamics of KLK8 suggested that the surface loops of trypsin-like serine proteases are connected in an allosteric network (Fig. 5E) [193, 195]. Recently, an intriguing additional role of KLK8 in skin has been discovered, as elevated protease activity of KLK8 results in KLK6 activation and PAR-2 signaling, associated with rapid wound healing, while both KLK6 and 8 were coordinately expressed in ovarian cancer [196, 197]. Similar to KLK6, but in contrast to KLK8, trypsin-3 (mesotrypsin) targets PAR-1 with subsequent Ca2+ signaling in neurons, in particular in astrocytes [189, 198]. Another trypsin-like, membrane-bound “mosaic” protease, called neurotrypsin, is found from insect neurons to vertebrate brains, where it cleaves the proteoglycan agrin at motoneuron junctions for synapse formation [199, 200].
2.4.5. Other KLKs
Human KLKs 9, 10, and 12-15 have been functionally characterized to some extent, however, distinct physiological roles have not been determined for all of them, despite a very restricted tissue expression in some cases [144]. Some basic characteristics can be derived from their amino acid sequences in context with functional data: KLK9 and KLK15 may have an unusual substrate specificity, based on their S1 pocket residues Gly189 and Glu189, respectively, while KLK15 has an eight residue insertion in the 148-loop and in KLK13 the formation of a standard N-terminus requires cleavage after Phe15 or an alternative propeptide cleavage after Lys5 [173].
An eight residue insertion in the 99-loop of KLK10 resembles the classical KLKs 1 to 3, while the physiology of KLK10 is not well characterized. A role as tumor suppressor was suggested in breast, ovarian and prostate cancer, which remains controversial [201, 202]. Active human KLK10 requires a three residue longer N-terminus as the standard Ile/Val16 and cleaves with a mixed preference for basic and unbranched hydrophobic P1-side-chains [167]. Structures of ligand-free and Zn2+ bound zymogen-like KLK10 explained the inhibition by disruption of the catalytic triad, while the disordered N-terminus and 75-, 148-, and the 99-loops are connected in an allosteric network [203]. Most likely, substrate binding favors the productive active site conformation as in the induced fit model, which seems to be only a special case of the conformational selection model [204].
2.5. HtrA proteases
The high temperature requirement A (HtrA) proteases were discovered in E. coli and termed DegP, DegQ and DegS, displaying a modified elastase-like preference for P1-Val and Ile [205]. In general, DegP degrades misfolded proteins in the bacterial periplasm with the help of two PDZ domains at elevated temperatures from 37 °C on, but acts as chaperone at lower temperatures, whereby the resting state is a double trimer and the active state a 24-mer [206].
All four human HtrAs are secreted and possess a C-terminal PDZ domain, and except HtrA2, N-terminal IGFBP (insulin-like growth factor binding protein) and Kazal domains [207]. None of the additional domains is required for protease activity and both the IGFBP and Kazal domains are not functional, although HtrA4 is hardly characterized compared to the other members of the group [208]. The basic HtrA assembly is a trimer, while higher oligomers show increasing activity, in particular HtrA1 with an elastase-like specificity for P1-Thr, Val, Ala and Leu [209, 210]. Based on the N-terminal domain insertions, the catalytic triad residues of HtrA1 are His220 (c57), Asp250 (c102) and Ser328 (c195), whereby a stabilizing salt-bridge to the N-terminus is missing. PDZ-mediated substrate binding induces an active conformation of the triad, requiring the concerted opening of the three disordered loops L1 (323-327, 191-loop), L2 (348-351, 220-loop) and L3 (307-316, 175-loop), a sensor for allosteric activation (Fig. 6) [14, 211].
Figure 6. Various trypsin-like serine proteases with allosteric loop networks. Significant loops are labeled.
6A: Human HtrA1 displays a zymogen-like conformation with a highly disordered region around the 175-loop (L3), since it misses an N-terminal activating salt bridge. Trimerization and substrate binding induce conformational changes, including the 189- and 220-loops (L1 and L2) and the catalytic triad (3NWU, 3NZI [211]). 6B: Matriptase of the TMPRSS family is a membrane anchored protease related to enterokinase. A locked zymogen mutant Arg614Ala (c15) displays 3% activity, due to a missing activating salt bridge to the N-terminus, as well as closed 148-, 175-, and 220-loops (transparent surfaces), while the mature protease adopts standard conformations (cartoon representation) (1EAX, 5LYO [227, 293]). 6C: Streptogrisin B from a fungus represents a minimal version of trypsin-like serine proteases with only 186 residues. It contains extremely short loops with respect to trypsin, while only the 189- and 220-loops reach standard lengths (1DS2 [294]).
Mitochondrial HtrA2 is elastase-like, but favors P1-Leu and disfavors P1-Ala in degradome analyses, which lead to the identification of actin and tubulin as substrates [212].
Access to the active site is hindered by the L1 (191) and LD (148) loops, the latter being the major regulator through interaction with the PDZ domain [213]. Secreted HtrA3 exists in two isoforms, the shorter one lacks the PDZ domain and presumably remodels the extracellular matrix, with loop B (LB, c60) as the dominating regulator [214, 215]. HtrA1 and HtrA3 are involved in embryo implantation, but dysregulation of HtrA1, 2 or 3 is associated with many diseases, such as cancer, arthritis, neurodegeneration, and age-related macular degeneration [216]. For example, biallelic mutations in the HtrA1 gene cause the cerebral small vessel disease CARASIL and both HtrA1 and HtrA2 participate in programmed cell death, either in apoptosis or in anoikis, which is induced in anchorage-dependent cells upon detachment from the surrounding matrix, whereas dysfunctional HtrA2 has been linked to Parkinson's and Alzheimer's disease [217–219]. HtrA3 was identified as pregnancy-related serine protease, but is also expressed in heart, ovary and testis, and is an indicator of early-onset preeclampsia, similar to HtrA4, which is restricted to the placenta [220, 221].
2.6. Membrane anchored serine proteases
About 20 membrane anchored serine proteases fulfill a variety of important tasks in the human body [222, 223]. They have gained interest due to their physiological functions, and emerging roles in protease-mediated signaling of cancer [224]. The prominent example of enteropeptidase, belonging to the hepsin/TMPRSS subfamily, has already been described in the context of digestive enzyme activation cascades (section 2.1.). Hepsin and the other members of the subfamily, such as epitheliasin (TMPRSS2) consist of an N-terminal membrane-spanning domain, a LDLRA domain and a protease domain. Hepsin can activate several proteases from the blood coagulation cascade and seems to play a special role in prostate cancer [225]. Corin is expressed in heart and uterus; it regulates salt levels and blood pressure, with mutated forms contributing to preeclampsia, and it exhibits a complicated mosaic architecture with TM, Fz1 (frizzle-like), LDLR1-5, Fz2, LDLR6-8, and protease domains [226].
Matriptase is a tryptic transmembrane mosaic protease that is important for epidermal barrier function. The “zymogen locked” mutant Arg614Ala (c15) displays a zymogen activity around 3= of the mature protease, corresponding to a zymogenicity factor of 33, which was explained by a distorted oxyanion hole and S1 subsite, as well as by closed conformations of the 148-, 175-, and 220-loops (Fig. 6B) [227]. The closely related matriptase-2 controls the iron levels in blood, but is largely uncharacterized as matriptase-3 and the polyserases with up to three protease domains within a single polypeptide [228, 229].
DESC1-4, HAT-like 1, 4, and 5 are related trypsin-like transmembrane proteases, as exemplified by the structure of the catalytic domain of DESC1, resembling trypsin, enteropeptidase, hepsin and matriptase, with a typical S1 subsite that was also compared to active thrombin [230]. Most of the membrane-anchored serine proteases are type II transmembrane proteases, whereas γ-tryptase is the only member encompassing a C-terminal type I transmembrane domain; testisin and prostasin are membrane-anchored via C-terminal glycophospatidylinositol (GPI) segments, with a restricted expression in testes and potential roles in prostate cancer involving PAR signaling [231, 232]. Prostasin and the soluble hepatocyte growth factor activator (HGFA) can switch between open and closed states, depending mostly on the 220-loop, similar to the E and E* forms of α-thrombin [233].
Dysregulated expression of several membrane-anchored proteases, especially hepsin, TMPRSS2, matriptase, testisin and prostasin, is linked to carcinogenesis, since increased levels of them in relation to their endogenous inhibitors are associated with poor patient prognosis [224]. The oncogenic activity mediated by these proteases may depend on their role as proteolytic modifiers of growth factors and protease-activated receptors, especially PAR-2, rather than representing extracellular matrix protein degrading enzymes [234].
2.7. Serine proteases as toxins and allergens
2.7.1. Reptilian and mammalian trypsin-like protease toxins
Many snake venoms contain metalloproteases (SVMPs), which can be the dominating toxin, or even more frequently serine protease toxins (SVSPs), which often interfere with the blood coagulation system [235, 236]. Several pit vipers produce venoms that contain thrombin-like serine proteases, such as venombin A and venombin AB (gabonase), which cleave fibrinogen and even activate the cross-linking transglutaminase fXIII [237]. Russell's viper venom factor V activator (RVV-V) from the South Asian Daboia russelii is another example of a thrombin-like protease, in which the Ser-Trp-Gly216 stretch is replaced by Ala-Gly-Gly199 (c216). Scutelarin from the Australian taipan (Oxyuranus scutellatus) is a multidomain activator of prothrombin with fXa/fVa-like domains, whereas the plateletaggregating endopeptidase from the South American pit viper Bothrops jararaca activates PAR-1 on platelets like thrombin and its protease A degrades fibrin and fibrinogen [238, 239].
By contrast, mammalian toxins are rare, with the notable exception of the short-tailed shrew (Blarina brevicauda), which produces blarina toxin (BLTX) in the submaxillary and sublingual glands, a venom that is lethal for mice and share 53% identical residues with human tissue kallikrein [240]. Similar to the gila toxin from lizards of the genus Heloderma, blarina toxin cleaves like a kininogenase LMWK and HMWK, exhibiting more positively charged residues around the active site and a unique 21 residue insertion in the 99-loop compared with the 11 residue insertion of KLK1 [241].
2.7.2. Allergenic trypsin-like proteases from mites and cockroaches
Aerosolized proteins from animal sources, e.g. arthropod feces, are a worldwide health problem, affecting around 10% of humans with symptoms of allergy or atopic dermatitis. Among these allergens are several proteases, in particular trypsin-like serine proteases [242]. House dust mites, such as Dermatophagoides pteronyssinus or farinae and Euroglyphus maynei produce the cysteine protease Der p 1 (or Der f 3 and Eur m 3), which activates the serine proteases Der p 3, 6 and 9 [243]. Der p 3 exhibits tryptic and Der p 6 chymotryptic specificity, while the collagenolytic Der p 9 is mixed chymotryptic/elastase-like, due to the presence of Ala189 in the S1 pocket [244, 245]. Structural differences to the mammalian counterparts are small, such as a Thr15 or Gly15 preceding the propeptide, only three disulfides and a shortened 75-loop. It has been demonstrated that Der p 3 and 9 can activate PAR-2 and that serpins inhibit them [246]. Other mites from tropical areas exhibit related allergenic proteases, such as Blo t 3, Tyr p 3 and Sar s 3 [245].
Also, the American cockroach (Periplaneta americana) produces a trypsin-like serine protease allergen, Per a 10, which causes allergic bronchial inflammation via cleavage of several cellular receptor proteins n [247]. Recently, three Per a 10 related enzymes from the German cockroach (Blatella germanica), were identified, which activate MAP kinase and enhance the allergic response via cleavage of PAR-2 [248].
2.8. Serine proteases from fungi, bacteria and viruses
Alpha-lytic protease from Lysobacter enzymogenes, with elastase-like specificity, has gained some interest as model protease for structure-functional studies, e.g. crystallography in the subångström range [249, 250]. Streptogrisin B is a fungal P1-Lys specific serine protease from Streptomyces griseus, which serves as model protease for structure-based mechanistic studies, despite considerable deletions, resulting in a 186 residue molecule (Fig. 6C) [251]. By contrast, streptogrisin E (glutamyl endopeptidase II) and other serine proteases produced by bacteria from the genus Staphylococcus and Bacillus, such as glutamyl endopeptidase I, display an unusual specificity for P1-Glu [252]. The related glutamyl epidermolysin A and B or so-called exfoliative toxins from Staphylococcus aureus and similar species destroy components of the human epidermis, resulting in blisters and progression to deeper layers of the skin [253]. They cleave cadherins, such as desmoglein-1 after P1-Glu residues with high specificity. IgA1-specific serine proteases are present in Neisseria bacteria and the virus Haemophilus influenza, and are related to the HAP protease, an unusual autotransporter of S6 family with a chymotrypsin-like protease domain [254].
3. The loop system of family S1 serine proteases and its modulators
3.1. The N-terminal activation loop
Although this crucial segment starting with residue c16 is limited in length and variability, several modulatory phenomena were observed. For this loop-like segment and the eight active site surrounding surface loops, the major conformational states, such as zymogen, closed and open forms are listed in Table 1 with the most relevant examples. Some trypsin-like protease show aberrations from the standard pattern of residue c16 inserted in the activation pocket and forming the crucial salt bridge with the N-terminal α-ammonium group to the side-chain of Asp194. A classic example is single chain tPA, in which the side-chain of Lys156 inserts in the activation pocket, resulting in a protease with up to 20% activity of the fully processed, mature protease [75]. Two examples of the KLKs show further deviations from standard N-termini (Fig. 7). First, KLK10 is only active with a three residue extension starting with Leu13 and not Glu16 at the N-terminus, despite a zymogen-like conformation of the active variant [203]. This phenomenon can be explained by an induced fit model upon substrate binding, which is now considered as special case of the conformational selection model [204]. Most unusual among the family S1 is KLK13, which is equally active with a standard Leu16 residue and with a Lys5 (Fig. 7, upper panel). One can only speculate that either the α-ammonium group of Lys5 inserts into the activation pocket or its side-chain, similar to Lys156 of sc-tPA.
Table 1. Loop states of selected family S1 serine proteases with structural examples. Effectors and inhibitors are depicted in bold italic font.
| Loop | Zymogen | Active zymogen | Closed (E*) | Open (E) | Clipped active |
|---|---|---|---|---|---|
| N-term | Chymotrypsinogen | Sc-tPA, HtrA1 | Chymotrypsin | ||
| 37 (LA) | pro-KLK6 | APC, uPA, tPA | Trypsin | ||
| 60 (LB) | pro-KLK6 | HtrA3 | Thrombin, APC uPA, tPA | Thrombin | |
| 75(LE) | Trypsinogen | KLK10 | fIX, CI, APC | Trypsin Ca2+ | |
| 99(LC) | Plasminogen | KLK10 | eKLK3, fIXa, fXa | Plasmin (94-shunt) | KLK2 (active) |
| 148 (LD) | Chymotrypsinogen | HtrA2 | eKLK3, CI, APC | Trypsin Ca2 | δ-Chymotrypsin |
| 189 (L1) | fXII | HtrA1, HtrA2 | eKLK3, C2a, CI | Trypsin | |
| 175 (L3) | Plasminogen | HtrA1 | fIXa, fXa | fXa fVa | |
| 220(L2) | Thrombin, fXII | HtrA1 | Thrombin (slow) eKLK3, C2a, CI | Thrombin (fast) Na+ |
Figure 7. Sequence alignment for stretches of the loops in selected trypsin-like serine protease domains.
Human trypsin-1 (TRY1) is included with bovine chymotrypsinogen (bCTRA) as numbering reference (underlined residues), while a gap is left for residues 107 to 130. Residues in bold font define the loops according to the reference chymotrypsin or have special functions, such as the catalytic triad residue His57, Asp102 and Ser195 or the specificity determining c189 und c190 residues. Negatively charged residues are shown in red, positively charged ones in blue. Glycosylation sites or sequons are shown in green, while additional metal binding residues for Na+, Ca2+ and Zn2+ are displayed magenta. Most mature trypsin-like protease domains possess the standard N-terminus with a hydrophobic residue (c16), such as Ile or Val. Blood coagulation factors fVII, fIX, fX, fXI, fXII and thrombin (Thrb) are partially aligned. The nine residue insertion with an N-glycosylation site in the 60-loop makes thrombin unique among the blood coagulation proteases, since it considerably shields the active site. Insertions in the usually short 37-loop influence substrate binding in the prime side and can enhance the specificity for plasmin (Plasm) as in the plasminogen activators uPA and tPA, which is accompanied by extended 60-loops. Partial protease domain segments of complement factors C1s, C2a, human neutrophil elastase (ELNE), and the human pancreatic elastase PE IIA (CEL2A) are displayed as well as of KLKs 2, 3, 4, 5, 8 and 10. Apart from zymogen forms, KLK10 is only active with a three residue extension, beginning with Leu13, while KLK13 can be active with a standard c16 N-terminus or with a Leu5.
3.2. The 37-loop
The shortest of the eight surface loops around the active site can be a distinct modulatory unit as demonstrated for fIXa and fXa with the serpin Z-dependent protease inhibitor (ZPI) and tissue factor pathway inhibitor (TFPI). Both strongly inhibit fXa, due the presence of three glutamate residues (Glu36, Glu37, and Glu39), which hold this loop most likely in an open conformation, in contrast to fIXa with a Lys36 that forms a salt bridge to Glu60 [255]. Also, the physiologically relevant interaction of fXa with the serpin anti-thrombin depends largely on this negatively charged patch [256]. Insertions in the usually short 37-loop of trypsin-like proteases, which are often accompanied by insertions in 60-loop, have a marked influence on substrate and inhibitor interaction in the prime side, as corroborated for fXI as well as for uPA and tPA, which gain a nearly exclusive selectivity for their substrate plasminogen [36]. Similar functions can be expected for the insertions of HNE and KLK14 in the 37-loop, which displays a certain lack for ligand binding or posttranslational modifications. Recently, studies with engineered nanobodies directed against mouse uPA, demonstrated that the 37-loop participates in an allosteric loop network, which can even be driven to adopt the closed E* conformation [257].
3.3. The 60-loop
Some of the unique characteristics of thrombin depend on the insertion of nine residues after residue 60 compared to chymotrypsin and most other serine proteases (Fig. 7, upper panel). In trypsin and most KLKs, however, this loop is further shortened by a single residue deletion. In thrombin, this loop shields the active site considerably and is N-glycosylated at the sequon Asn-Xaa-Ser, which may contribute to a closed conformation (E*) that opens only upon binding of specific substrates, such as fibrinogen or heparin-like stimulating molecules at the nearby anion binding exosite I. The N-glycan appears to play a distinct role in the arrangement of heparin in the ternary thrombin complex with antithrombin III, whereby N-glycans of the serpin serve as additional heparin binding moieties [62]. Similar to thrombin KLK3/PSA exhibits an N-glycosylation in the 60-loop, whose function close to anion exosite I seems to have some parallels. At least in recombinant glycosylated KLK3, the N-glycan seems to provide some protection against proteolytic cleavage [169].
3.4. The calcium binding 75- or 70-80 loop
The paragon of the calcium binding 70-80 loop (III, E) is found in trypsin, with the three side-chains of Glu70, Glu77 and Glu80 as ligand for Ca2+, and the two backbone carbonyl groups of Asn72 and Leu75 (Fig. 7, middle panel). Early studies showed that the long term stability of the protease depends on the Ca2+ complex, as well as an increase of the enzymatic activity of about 4-fold [258]. The stimulatory effect by Ca2+ is surpassed in some coagulation factors, such as fIXa and fXa, reaching more than 30-fold higher activity, which has been attributed to an overall increase of the rigidity of the active site region, accompanied by an optimized active conformation around the S1 pocket [38, 259]. The Ca2+ ligands of this loop can be Asp residues as well, while Glu80 is most conserved. Only KLK8 exhibits a similar Ca2+ binding site, which seems to have a similar function as in trypsin, as a 4-fold stimulation of activity was reported as well [193].
Interestingly, this loop can bind Zn2+ in a few examples, such as fVII and KLK4 with His71, Glu74 and other ligands, resulting in inhibition or at least significant reduction of the activity. However, this loop of fVII is capable to bind Ca2+, while two Zn2+ ions bind Lys24, Asp79 and His117 or His76 and Glu80 [52]. Other examples comprise shorter 75-loops without clear functions, while the disulfide Cys417-Cys420 (c77-c80) of fXIIa certainly stabilizes this loop, which is otherwise sensitive to clipping and auto-inactivation, as shown for KLK2 [166]. Thus, N-glycosylation of the 75-loop as observed for a few serine proteases might also protect against auto-cleavage or against cleavage by other proteases.
3.5. The 99-loop
Standard length 99-loops often allow acceptance of large P2 side-chains, such as Arg or even Trp, as in KLK6. However, small insertions of only two residues as in fIXa with corresponding conformational rearrangements may suffice to accept only small P2 residues and reduce the catalytic activity by several orders of magnitude [42]. Apparently, residue 99 plays a particular role in determining the P2 specificity, as corroborated by Tyr99 (fIXa, fXa and TPA) preferring P2-Gly and by the smaller Leu99 or Thr99 of thrombin and protein C, respectively, favoring P2-Pro (Fig. 7, middle panel). In blood coagulation factors, the 99-loop is an important regulatory region, which can lock the active site completely. Mutations of fIXa with unlocking character demonstrated that the protease activity can be enhanced 7,000-fold in the absence of fVa [43]. By contrast, the trypsin-like hepatocyte growth factor activator (HGFA) was completely inhibited by an antibody, which bound at the 99-loop and altered its conformation, resulting essentially in closure of the S2 and S4 pockets [260]. A comparison with the substrate-like HGFA inhibitor-1, a two Kunitz domain inhibitor, showed that also the 220-loop is involved in the allosteric changes of HGFA [233].
As already mentioned, since tissue kallikrein, KLK1, and the so-called classical KLKs 2 and 3 exhibit an 11 residue insertion in this loop, the extended 99-loop with respect to chymotrypsin has been termed “kallikrein loop”. Despite, this extraordinary long and active site shielding segment, P2-Phe and -Tyr are among the most favored residues in substrates of KLKs 1, 2 and 3. In fact, the composition and three-dimensional architecture of the 99-loop are additional parameters that determine its substrate specificity. Posttranslational modifications, such as N-glycosylation at the conserved Asn95 influence the catalytic activity in a distinct manner, in particular to enhance the substrate specificity [12]. Glycosylation of the 99-loop appears to be a special feature of KLKs, with KLK1 being heavily N- and O-glycosylated, but also in most other human family members [169, 261, 262]. Zn2+ binding to this loop in KLKs 5, 7, 8, and 10, involving His99 and His96 or Asp99 has been confirmed as mechanism of inhibition by mutational and structural studies [173, 193, 203]. The structures of rat tonin and an engineered trypsin showed that His57 rotates from its position in the catalytic triad, rendering the enzyme inactive [180, 263].
3.6. The autolysis or 148-loop
The autolysis of chymotrypsin with cleavages after residues 13, 15, and notably 146 and 148 generates the active π, δ, κ and α forms (Fig. 7, middle panel) [264]. By contrast a clipped 148-loop, due to autolysis or proteolytic activity of other proteases, often results in complete inactivation, as demonstrated for fXa with plasmin [265]. In line with these findings, Ca2+ binding at physiological concentrations in the 75-loop prevented such cleavage events and stabilized fXa significantly [266]. The cross-talk between the Ca2+ binding loop and the 148-loop has been explained by a communication line via the activation domain as in fIXa, although the direction of this communication from the 75-loop may be different in KLK8, where it seems to reach the 99-loop [40, 193].
In active protein C (APC), a loop chimera with a trypsin segment c143–c154 did not alter the proteolysis of fVa, but increased plasma inhibitor binding, which confirms the regulatory variability of this loop [267]. Similarly, the 148-loop is an important regulatory element of catalytic activity in thrombin. Human thrombin exhibits a five residue insertion relative to trypsin and chymotrypsin, namely A149aNVGK149e, and the deletion 146 to 149e turns the procoagulant thrombin into an anticoagulant, while a mouse chimera variant rigidified this loop and enhanced both Na+ binding and the activity about 10-fold [268]. A comparative analysis of structures from the KLK family confirmed that the 148-loop can adopt a closed conformation, together with the 99- and 220-loops, in an E* form, as initially found for thrombin [164]. Also, in HtrA2 the 148-loop (LD) appears to be a major regulatory element via its interaction with the PDZ domain (section 2.5).
3.7. The 170/175 loop
Due to its position at the rim of the active site, the role of this in most cases relatively short loop may have been neglected, as it starts sometimes with a helical turn as in chymotrypsin and is consequently more rigid than other loops. In HtrA1, this loop (L3) has been identified as sensor of allosteric activation (section 2.5). For fIXa binding at the 175-loop by fVIIIa is opening the “locked” conformation of Tyr177, which has an influence on the nearby 99-loop, in particular Lys98, in order to allow substrate binding (section 2.3) (Fig. 7, lower panel). Also, in prothrombinase, the 175-loop is the major contact area of the catalytic domain of fXa and fV [47]. Studies of loop chimeras of fXa and trypsin discovered a special role of the 175-loop in conformations of the 99- and 189/191-loops, conferring significant plasticity in an allosteric interaction network [269].
3.8. The S1 forming or 189/191-loop
In general, the most significant regulatory step in the 189-loop is the critical event of the salt bridge formation between the N-terminal residue c16 and residue c194, preceding the catalytic Ser195. However, numerous examples of trypsin-like proteases exist, where zymogen activity occurs, often leading to auto-activation, as well as unusual variants of salt-bridge formation with side-chains, as Lys156(c) in tPA or the complement factors C2a and CBb with Arg224(c), see sections 2.2 and 2.3. Also, KLKs 10 and 13 seem to be active with non-standard N-termini, perhaps even 10 residues longer than usual (section 2.4). The variations of substrate recognition, specificity determinants and factors influencing the turnover are too numerous, to be discussed in this context, although single amino acid changes at position c189 have strong effects on structure and activity [270]. Nevertheless, the 189-loop serves as a highly variable element of additional regulation.
Thus, it is not surprising that the extended 189-loops of tPA and uPA can be disordered in parts (section 2.2), while it is astonishing that in complement factor I the region including the catalytic Ser is completely flexible, despite its virtually active, mature state (Fig. 7, lower panel). At least for fXa, the role of the solvent-exposed residues Asp185, Lys186, and Glu188 has been elucidated, as they have differential effects on catalytic activity and Na+ binding, as well as they represent a distinct interaction region for fVa [271]. However, the prothrombinase complex from a reptile, comprising the fXa and fVa did not reveal how this interaction might look like in detail, since the 189-loop is largely disordered, although the construct lacked the Gla- and the EGF1 domain [47]. If the disulfide Cys191-Cys220 is present and intact, it represents a strong allosteric link between the 189-loop and the 220-loop, as shown for mutants of thrombin, explaining the high activity of the Na+ bound form by preventing solvent exposure of Asp189 and disorder around the S1 pocket [272].
3.9. The Na+ binding or 220-loop
Although the 220-loops of the zymogen forms trypsinogen and chymotrypsinogen are not disordered, other proteases, such as fXII exhibit considerable disorder in this loop, extending to the 189-loop, including the oxyanion hole [34]. Undoubtedly, thrombin is the protease with an outstanding role of the 220-loop, since its capacity to bind Na+ and K+ as activity enhancing regulators is remarkable. This topic with respect to allostery and regulation of activity has been extensively studied and described in comprehensive reviews [273, 274]. The direct structural link to the 189-loop is the disulfide Cys191-Cys220, while the bridging Na+ coordination by the backbone carbonyl oxygen atoms of Arg221a and Lys224 extends to the water stabilizing carbonyl of Tyr184a (Fig. 7, lower panel) [275]. Most digestive S1 family proteases with the TCN codon for the catalytic Ser195 possess a Pro225, which excludes binding of monovalent cations, whereas most coagulation factors with the AGY codon for Ser195 possess Tyr/Phe225 residues, which favor Na+/K+ binding [276]. Noteworthy, a recent structure of fIXa showed that Na+ is coordinated by the backbone carbonyl O-atoms of residues 184a, 185, 221a, and 224, which resembles the corresponding binding site of fXa and establishes a strong allosteric link to the 189-loop and the S1 subsite, respectively [277].
Despite the special role of the 220-loop in several blood coagulation factors, its importance as regulatory element in other proteases should not be forgotten. It may be a very unusual feature, but in complement factor C2a (and CBb), the Arg696 (around c224) side-chain is the critical activating element of the mature protease, instead of an N-terminal c16 (section 2.4). For KLKs it has been shown that the 220-loop can most likely open and close in a concerted manner with the 99- and 148-loops.
4. The allosteric dynamic loop network of trypsin-like serine proteases
An overall scheme of the allosteric loop network with various trypsin-like serine proteases is depicted in figure 8 and accompanied by a cross-talk matrix in Table 2. The most obvious allosteric interaction in the vast majority of trypsin-like serine proteases is the one of the N-terminal loop with the 189-loop upon salt bridge formation of the N-terminus of c16 with Asp194. In particular, the S1 pocket conformation around the 189-loop, including formation of the catalytic triad and the oxyanion hole depend directly on this major rearrangement after cleavage of the propeptides of the zymogens. The best known examples with structural proof come from chymotrypsinogen and trypsinogen, which exhibits a largely disordered activation domain. Usually the 148-loop participates in these conformational changes to a large extent, as clipping in this “autolysis” loop often inactivates serine proteases.
Figure 8. Overall scheme of the allosteric loop network in trypsin-like serine proteases.
The allosterically interacting N-terminus and loops are colored and labeled for the respective proteases, as well as proteinaceous and ionic cofactors. Several examples of crosstalk between distinct loops have been discovered and confirm that serine proteases are dynamic and flexible molecular machines, with adaptations to their specific tasks. For example in thrombin, Na+ binding changes the 220- and 189-loop conformations to the fast form, while thrombomodulin binding at the 37-, 60,- and 75-loops alters the substrate specificity (section 2.2.3.). In case of KLK10 a zymogen-like conformation of the N-terminus, the 75-, and 148-loops, involving the active site is capable to bind substrates and most likely rearranges to an active state. (see section 2.4.5.) Detailed knowledge of the individual allosteric network will facilitate the development of new modulatory compounds, biologics or synthetic molecules, which can be exploited in novel therapeutic approaches.
Table 2. Cross-talk matrix for the surface loops including the N-terminal segment. Effectors and inhibitors are indicated in bold and italic font.
| Effect | N-term | 37/LA | 60/LB | 75/LE | 99/LC | 148/LD | 175/L3 | 189/L1 | 220/L2 |
|---|---|---|---|---|---|---|---|---|---|
| Cause | |||||||||
| N-term | Trypsin, KLK10 | KLK10 | bCTRA Trypsin KLK10 CI | bCTRA Trypsin KLK4/10 CI | KLK10 CI | ||||
| 37/LA | muPA ab | muPA ab | muPA ab | muPA ab | muPA ab | ||||
| 60/LB | uPA tPA | Thrombin HtrA3 | Thrombin | ||||||
| 75/LE | KLK4 Zn2 fVII Zn2 muPA ab | KLK8 Ca2+ | KLK8 Ca2+ | Trypsin/ fVIIa/fIXa/ KLK8 Ca2+ muPA ab | muPA ab | Trypsin/ fVIIa/fIXa/ KLK8 Ca2+ KLK4 Zn2 muPA ab | muPA ab | ||
| 99/LC | eKLK3 | eKLK3 KLKs 5/7/ 8/10 Zn2+ | eKLK3 | ||||||
| 148/LD | eKLK3 | eKLK3 Sc-tPA HtrA2 | eKLK3 | ||||||
| 175/L3 | fXa fVa fIXa fVIIIa | fXa fVa fIXa fVIIIa HtrA1 | HtrA1 | ||||||
| 189/L1 | KLK4, fIXa | uPA, tPA | uPA, tPA | eKLK3 | eKLK3 | HtrA1 | eKLK3 | eKLK3 fXII HtrA1 | |
| 220/L2 | eKLK3 | eKLK3 | Thrombin/ fIXa Na+ eKLK3 C2a | eKLK3 |
In most coagulation factors, as demonstrated for factor IXa, Ca2+ binding in the 75-loop has an allosteric effect via the N-terminal salt bridge and the 148-loop to the S1 subsite and the 189-loop, which is modified in KLK8, with an additional communication line via the nearby 37- and 60-loops to the 99-loop [193]. Molecular dynamics simulations indicated that a Ca2+-free 75-loop closes together with the 60-loop a region of the substrate binding cleft in the prime side, and even the 99-loop blocks access for P2 larger side-chains in the S2 pocket. A corresponding locked conformation was observed in fIX by an interaction with Tyr177 from the 175-loop, which can be opened by complex formation and interaction in this region by the natural activator fVIIla.
An additional allosteric long range interaction has been characterized for factor IXa with the Na+-binding 220-loop, although this particular interaction cannot take place in thrombin, it might be structurally equivalent to the Lys70 side-chain substituting the cation binding in the 75 loop (Fig. 8) [278]. The structural comparison of most KLKs suggested a concerted transition of the 99-, 148-, and 220-loops from the closed E* to the open E form in line with conformational selection, which includes the 189-loop as corroborated by the horse eKLK3 structure in the E* state [164, 165]. For KLK4 an alternative communication channel was described in a crystal structure based molecular dynamics approach, where the 75-loop was connected via its Zn2+ binding site to the N-terminus and the nearby 37-loop in the prime side region, as well as to the 189- and 220–loops in the non-prime side [195].
Regarding the accumulated data for all trypsin-like serine proteases it is not unlikely that all eight surface loops and the N-terminal segment are intrinsically connected in an allosteric machinery. The short loops seem to participate to a lower extent, while the longer ones, with modifications, such as glycosylation and ion binding sites, are capable of stronger allosteric effects and interactions and offer protein-protein and other molecular interaction sites, e.g. for heparin, with stimulating and inhibiting functions.
5. Conclusions and Outlook
Therapeutic exploitation of the surface loop sequences of serine proteases offer the most specific interaction that is conceivable for antibodies (abs), Fab fragments and nanobodies (nbs) of the camelid type. Recently, targeting of serine proteases, such as KLK2, by radionucleotide carrying monoclonal antibodies (Mabs) has been used to deliver the α-emitting Ac to tumor tissue [279]. In this approach the exact binding site of the protease is not known, while the usual location of the metal chelator at the N- or C-terminus of the antibody is not a drawback and may be extended to click chemistry approaches, even with small affibodies (affi-bs) [280]. Click chemistry offers numerous possibilities to attach active site inhibitors of any size to an ab, ideally in a conformation, in which a complementarity determining region (CDR) of the ab binds a unique protease loop and positions the active site inhibitor properly [281]. The potentially very high specificity is based on the large differences of the surface loops regarding length and primary amino acid sequence. In cases of a unique three-dimensional arrangement or additional secondary structural elements, such as helical turns, e.g. in the 75-loop of KLK4 or the 99-loop of KLK2/3, targeting should be nearly 100% specific without any cross-reactivity. Therefore, no unwanted side-effects due to binding of related serine protease are possible. All these immune system-derived biomolecules can be humanized either by mutation or post-translational modifications in suitable expression systems, which will improve bioavailability. In order to bind with the highest specificity, distinct loop conformations, known from atomic structures or perhaps calculated by molecular dynamics, can be employed to target only these versions of a target protease that are really involved in pathological processes. In particular, bifunctional/chimeric full length abs will be able to bind two different target regions, e.g. a distinct surface loop and post-translational modifications, such as N-glycosylation, which is only found on cancer-derived serine protease variants.
Combinations of antibody therapeutic and organic compounds, which could be either generated by enzymatic reactions, e.g. modified amino acids with ligases or lysine modifiers similar to the ubiquitination or sumoylation process, could be envisioned. Another route to such hybrid molecules are the established click reactions and their novel variants [281]. Thus, the ab or nb moiety could confer the maximum specificity for a target protease as the demyelinating KLK6 in multiple sclerosis, while a covalently linked specific inhibitor binds only the active site of KLK6 molecules [282].
A new generation of small molecules inhibitors based on rational design, which does not only bind the specificity pockets of a protease, but also the unique and distinguishing features of the individual surface loops, could have several advantages over the relatively large ab or nb derived therapeutics. As long as they largely obey the “rule of 5”, they can reach parts of organs or tissue regions, which are too distant or separated from blood vessels [283]. In addition, they might have a better chance to cross the blood brain barrier and membranes. Potential targets could be neutrophil elastase, cathepsin G, and granzymes, which are all stored in in cytosolic vesicles. It should be not be forgotten that all mentioned antigen binding molecules (abs, nbs, Fab fragments, affi-bs) can act as stimulators as well, in particular when they stabilize a highly active E conformation according to the conformational selection model, which includes essentially induced fit situations [204]. Recent progress with engineered nbs demonstrated a site-directed targeting of the allosteric loop network in uPA, with an efficient switch from the active E form to the inactive closed E* form [284].
Serine proteases are used as FDA approved drugs since a long time, however, they are mostly restricted to supplementing deficiencies in blood coagulation or enhancing fibrinolysis [285]. Similarly, the cysteine proteases papain and bromelain, as well as clostridial collagenases have been applied in debridement of wounds [286, 287]. However, the potential of proteases as drugs seems much higher, when their ability to degrade target proteins in a highly specific manner could be exploited, e.g. by engineering granzymes and related pathogen-attacking proteases, combined with substrate specificity tuning. Their stability can be greatly enhanced by abolishing sensitive clipping sites in their surface loops or by introducing new N-glycosylation sites. Further advantages of such approaches are the acceptance of human proteases by the immune system and slower clearance from circulation compared with small molecule drugs.
Highlight.
Trypsin-like serine proteases are the largest protease group in higher organisms
They are connected in activation networks in physiology and many diseases
Eight loops near the active site and the N-terminus mainly regulate the activity
The loops interact allosterically and with regulatory ions or molecules
Pharmacological drugs can target several serine proteases via allosteric effects
Acknowledgements
We are grateful to Associate Prof. Jonathan Harris and the Institute of Health and Biomedical Innovation (IHBI) at the Queensland University of Technology in Brisbane (Australia) for their encouragement and support. The present study was supported (VM) by grants from the Wilhelm Sander-Stiftung, grant 2016-024.1, and the Deutsche Forschungsgemeinschaft, MA 1236/10-1, as well as by the Austrian Science Fund (FWF) project I3877-B21 (PG).
Footnotes
Author Contributions
The role of the authors: Peter Goettig designed the review, wrote most parts based on the currently available literature and created the figures. Hans Brandstetter contributed by writing some sections, in particular in the blood coagulation chapter, as well as with critical reading. Viktor Magdolen corrected the whole manuscript by rewriting numerous sections or sentences and did critical reading.
References
- [1].Rawlings ND, Barrett AJ. Evolutionary families of peptidases. Biochem J. 1993;290:205–218. doi: 10.1042/bj2900205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [2].Matthews BW, Sigler PB, Henderson R, Blow DM. Three-dimensional Structure of Tosyl-α-chymotrypsin. Nature. 1967;214:652–656. doi: 10.1038/214652a0. [DOI] [PubMed] [Google Scholar]
- [3].Hartley BS, Neurath H. Homologies in Serine Proteinases [and Discussion] Phil Trans R Soc B. 1970;257:77–87. doi: 10.1098/rstb.1970.0010. [DOI] [PubMed] [Google Scholar]
- [4].Rawlings ND, Barrett AJ, Thomas PD, Huang X, Bateman A, Finn RD. The MEROPS database of proteolytic enzymes, their substrates and inhibitors in 2017 and a comparison with peptidases in the PANTHER database. Nucleic Acids Res. 2018;46:D624–D632. doi: 10.1093/nar/gkx1134. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [5].Gorbalenya AE, Donchenko AP, Blinov VM, Koonin EV. Cysteine proteases of positive strand RNA viruses and chymotrypsin-like serine proteases. Febs Lett. 1989;243:103–114. doi: 10.1016/0014-5793(89)80109-7. [DOI] [PubMed] [Google Scholar]
- [6].Mosimann SC, Cherney MM, Sia S, Plotch S, James MNG. Refined X-ray crystallographic structure of the poliovirus 3C gene product. J Mol Biol. 1997;273:1032–1047. doi: 10.1006/jmbi.1997.1306. [DOI] [PubMed] [Google Scholar]
- [7].Rawlings ND, Barrett AJ. In: Handbook of Proteolytic Enzymes. Third Edition. Rawlings ND, Salvesen G, editors. Academic Press; London: 2013. Chapter 559 - Introduction: Serine Peptidases and Their Clans; pp. 2491–2523. [Google Scholar]
- [8].Baptista AM, Jonson PH, Hough E, Petersen SB. The Origin of Trypsin: Evidence for Multiple Gene Duplications in Trypsins. J Mol Evol. 1998;47:353–362. doi: 10.1007/pl00006393. [DOI] [PubMed] [Google Scholar]
- [9].Krem MM, Rose T, Di Cera E. Sequence Determinants of Function and Evolution in Serine Proteases. Trends Cardiovasc Med. 2000;10:171–176. doi: 10.1016/s1050-1738(00)00068-2. [DOI] [PubMed] [Google Scholar]
- [10].Brenner S. The molecular evolution of genes and proteins: a tale of two serines. Nature. 1988;334:528–530. doi: 10.1038/334528a0. [DOI] [PubMed] [Google Scholar]
- [11].Wu J, Jin Y, Zhong S, Chen R, Zhu S, Wang W, Lu Q, Xiong Y. A unique group of inactive serine protease homologues from snake venom. Toxicon. 2008;52:277–284. doi: 10.1016/j.toxicon.2008.05.013. [DOI] [PubMed] [Google Scholar]
- [12].Goettig P. Effects of Glycosylation on the Enzymatic Activity and Mechanisms of Proteases. Int J Mol Sci. 2016;17:E1969. doi: 10.3390/ijms17121969. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [13].Polgár L. In: Handbook of Proteolytic Enzymes. Third Edition. Rawlings ND, Salvesen G, editors. Academic Press; London: 2013. Chapter 560 - Catalytic Mechanisms of Serine and Threonine Peptidases; pp. 2524–2534. [Google Scholar]
- [14].Perona JJ, Craik CS. Evolutionary Divergence of Substrate Specificity within the Chymotrypsin-like Serine Protease Fold. J Biol Chem. 1997;272:29987–29990. doi: 10.1074/jbc.272.48.29987. [DOI] [PubMed] [Google Scholar]
- [15].Page MJ, Di Cera E. Serine peptidases: Classification, structure and function. Cell Mol Life Sci. 2008;65:1220–1236. doi: 10.1007/s00018-008-7565-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [16].Szabó A, Pilsak C, Bence M, Witt H, Sahin-Tóth M. Complex Formation of Human Proelastases with Procarboxypeptidases A1 and A2. J Biol Chem. 2016;291:17706–17716. doi: 10.1074/jbc.M116.743237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [17].Schilling O, Biniossek ML, Mayer B, Elsässer B, Brandstetter H, Goettig P, Stenman UH, Koistinen H. Specificity profiling of human trypsin-isoenzymes. Biol Chem. 2018;399:997–1007. doi: 10.1515/hsz-2018-0107. [DOI] [PubMed] [Google Scholar]
- [18].Stroud RM, Kay LM, Dickerson RE. The structure of bovine trypsin : Electron density maps of the inhibited enzyme at 5 Å and at 2·7 Å resolution. J Mol Biol. 1974;83:185–208. doi: 10.1016/0022-2836(74)90387-8. [DOI] [PubMed] [Google Scholar]
- [19].Huber R, Kukla D, Bode W, Schwager P, Bartels K, Deisenhofer J, Steigemann W. Structure of the complex formed by bovine trypsin and bovine pancreatic trypsin inhibitor : II. Crystallographic refinement at 1.9 Å resolution. J Mol Biol. 1974;89:73–101. doi: 10.1016/0022-2836(74)90163-6. [DOI] [PubMed] [Google Scholar]
- [20].Zheng XL, Kitamoto Y, Sadler JE. Enteropeptidase, a type II transmembrane serine protease. Front Biosci E. 2009;1:242–249. doi: 10.2741/E23. [DOI] [PubMed] [Google Scholar]
- [21].Cohen I, Kayode O, Hockla A, Sankaran B, Radisky DC, Radisky ES, Papo N. Combinatorial protein engineering of proteolytically resistant mesotrypsin inhibitors as candidates for cancer therapy. Biochem J. 2016;473:1329–1341. doi: 10.1042/BJ20151410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [22].Keiles S, Kammesheidt A. Identification of CFTR, PRSS1, and SPINK1 mutations in 381 patients with pancreatitis. Pancreas. 2006;33:221–227. doi: 10.1097/01.mpa.0000232014.94974.75. [DOI] [PubMed] [Google Scholar]
- [23].Hasan A, Moscoso DI, Kastrinos F. The Role of Genetics in Pancreatitis. Gastrointest Endosc Clin N Am. 2018;28:587–603. doi: 10.1016/j.giec.2018.06.001. [DOI] [PubMed] [Google Scholar]
- [24].Whitcomb DC, Gorry MC, Preston RA, Furey W, Sossenheimer MJ, Ulrich CD, Martin SP, Gates LK, Amann ST, Toskes PP, Liddle R, et al. Hereditary pancreatitis is caused by a mutation in the cationic trypsinogen gene. Nat Genet. 1996;14:141–145. doi: 10.1038/ng1096-141. [DOI] [PubMed] [Google Scholar]
- [25].Nemoda Z, Sahin-Tóth M. Chymotrypsin C (Caldecrin) Stimulates Autoactivation of Human Cationic Trypsinogen. J Biol Chem. 2006;281:11879–11886. doi: 10.1074/jbc.M600124200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [26].Bence M, Sahin-Tóth M. Asparagine-linked glycosylation of human chymotrypsin C (CTRC) is required for folding and secretion but not for enzyme activity. FEBS J. 2011;278:4338–4350. doi: 10.1111/j.1742-4658.2011.08351.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [27].Szmola R, Sahin-Tóth M. Chymotrypsin C (caldecrin) promotes degradation of human cationic trypsin: Identity with Rinderknecht’s enzyme Y. Proc Natl Acad Sci USA. 2007;104:11227–11232. doi: 10.1073/pnas.0703714104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [28].de Oliveira EB, Salgado MCO. In: Handbook of Proteolytic Enzymes. Third Edition. Rawlings ND, Salvesen G, editors. Academic Press; London: 2013. Chapter 584 - Pancreatic Elastases; pp. 2639–2645. [Google Scholar]
- [29].Krem MM, Cera ED. Evolution of enzyme cascades from embryonic development to blood coagulation. Trends Biochem Sci. 2002;27:67–74. doi: 10.1016/s0968-0004(01)02007-2. [DOI] [PubMed] [Google Scholar]
- [30].Arneth B. Coevolution of the coagulation and immune systems. Inflamm Res. 2019;68:117–123. doi: 10.1007/s00011-018-01210-y. [DOI] [PubMed] [Google Scholar]
- [31].Davidson CJ, Tuddenham EG, McVey JH. 450 million years of hemostasis. J Thromb Haemost. 2003;1:1487–1494. doi: 10.1046/j.1538-7836.2003.00334.x. [DOI] [PubMed] [Google Scholar]
- [32].Renné T, Schmaier AH, Nickel KF, Blombäck M, Maas C. In vivo roles of factor XII. Blood. 2012;120:4296–4303. doi: 10.1182/blood-2012-07-292094. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [33].Schmaier AH. The contact activation and kallikrein/kinin systems: pathophysiologic and physiologic activities. J Thromb Haemost. 2016;14:28–39. doi: 10.1111/jth.13194. [DOI] [PubMed] [Google Scholar]
- [34].Pathak M, Wilmann P, Awford J, Li C, Hamad BK, Fischer PM, Dreveny I, Dekker LV, Emsley J. Coagulation factor XII protease domain crystal structure. J Thromb Haemost. 2015;13:580–591. doi: 10.1111/jth.12849. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [35].Mohammed BM, Matafonov A, Ivanov I, Sun M-f, Cheng Q, Dickeson SK, Li C, Sun D, Verhamme IM, Emsley J, Gailani D. An update on factor XI structure and function. Thromb Res. 2018;161:94–105. doi: 10.1016/j.thromres.2017.10.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [36].Navaneetham D, Jin L, Pandey P, Strickler JE, Babine RE, Abdel-Meguid SS, Walsh PN. Structural and mutational analyses of the molecular interactions between the catalytic domain of factor XIa and the Kunitz protease inhibitor domain of protease nexin 2. J Biol Chem. 2005;280:36165–36175. doi: 10.1074/jbc.M504990200. [DOI] [PubMed] [Google Scholar]
- [37].Bolton-Maggs PHB. 10 Factor XI deficiency. Baillieres Clin Haematol. 1996;9:355–368. doi: 10.1016/s0950-3536(96)80068-0. [DOI] [PubMed] [Google Scholar]
- [38].Zögg T, Brandstetter H. In: Progress in Molecular Biology and Translational Science. Di Cera E, editor. Academic Press; London: 2011. Chapter 2 - Complex Assemblies of Factors IX and X Regulate the Initiation, Maintenance, and Shutdown of Blood Coagulation; pp. 51–103. [DOI] [PubMed] [Google Scholar]
- [39].Morrissey JH, Tajkhorshid E, Sligar SG, Rienstra CM. Tissue Factor/Factor VIIa Complex: Role of the Membrane Surface. Thromb Res. 2012;129:S8–S10. doi: 10.1016/j.thromres.2012.02.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [40].Zögg T, Brandstetter H. Structural Basis of the Cofactor- and Substrate-Assisted Activation of Human Coagulation Factor IXa. Structure. 2009;17:1669–1678. doi: 10.1016/j.str.2009.10.011. [DOI] [PubMed] [Google Scholar]
- [41].Brandstetter H, Bauer M, Huber R, Lollar P, Bode W. X-ray structure of clotting factor IXa: active site and module structure related to Xase activity and hemophilia B. Proc Natl Acad Sci USA. 1995;92:9796–9800. doi: 10.1073/pnas.92.21.9796. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [42].Hopfner K-P, Lang A, Karcher A, Sichler K, Kopetzki E, Brandstetter H, Huber R, Bode W, Engh RA. Coagulation factor IXa: the relaxed conformation of Tyr99 blocks substrate binding. Structure. 1999;7:989–996. doi: 10.1016/s0969-2126(99)80125-7. [DOI] [PubMed] [Google Scholar]
- [43].Sichler K, Kopetzki E, Huber R, Bode W, Hopfner K-P, Brandstetter H. Physiological fIXa Activation Involves a Cooperative Conformational Rearrangement of the 99-Loop. J Biol Chem. 2003;278:4121–4126. doi: 10.1074/jbc.M210722200. [DOI] [PubMed] [Google Scholar]
- [44].Bolton-Maggs PHB, Pasi KJ. Haemophilias A and B. Lancet. 2003;361:1801–1809. doi: 10.1016/S0140-6736(03)13405-8. [DOI] [PubMed] [Google Scholar]
- [45].Castaman G, Matino D. Hemophilia A and B: molecular and clinical characteristics of similar, but different diseases. Haematologica. 2019:haematol.2019.221093 [Google Scholar]
- [46].Bode W, Brandstetter H, Mather T, Stubbs MT. Comparative analysis of haemostatic proteinases: structural aspects of thrombin, factor Xa, factor IXa and protein C. Thromb Haemost. 1997;78:501–511. [PubMed] [Google Scholar]
- [47].Lechtenberg BC, Murray-Rust TA, Johnson DJD, Adams TE, Krishnaswamy S, Camire RM, Huntington JA. Crystal structure of the prothrombinase complex from the venom of Pseudonaja textilis . Blood. 2013;122:2777–2783. doi: 10.1182/blood-2013-06-511733. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [48].Sichler K, Banner DW, D’Arcy A, Hopfner K-P, Huber R, Bode W, Kresse G-B, Kopetzki E, Brandstetter H. Crystal Structures of Uninhibited Factor VIIa Link its Cofactor and Substrate-assisted Activation to Specific Interactions. J Mol Biol. 2002;322:591–603. doi: 10.1016/s0022-2836(02)00747-7. [DOI] [PubMed] [Google Scholar]
- [49].Etscheid M, Hunfeld A, König H, Seitz R, Dodt J. Activation of proPHBSP, the Zymogen of a Plasma Hyaluronan Binding Serine Protease, by an Intermolecular Autocatalytic Mechanism. Biol Chem. 2000;381:1223–1231. doi: 10.1515/BC.2000.150. [DOI] [PubMed] [Google Scholar]
- [50].Etscheid M, Muhl L, Pons D, Jukema JW, König H, Kanse SM. The Marburg I polymorphism of factor VII activating protease is associated with low proteolytic and low pro-coagulant activity. Thromb Res. 2012;130:935–941. doi: 10.1016/j.thromres.2012.07.023. [DOI] [PubMed] [Google Scholar]
- [51].Petersen LC, Olsen OH, Nielsen LS, Freskgard PO, Persson E. Binding of Zn2+ to a Ca2+ loop allosterically attenuates the activity of factor VIIa and reduces its affinity for tissue factor. Protein Sci. 2000;9:859–866. doi: 10.1110/ps.9.5.859. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [52].Bajaj SP, Schmidt AE, Agah S, Bajaj MS, Padmanabhan K. High Resolution Structures of p-Aminobenzamidine- and Benzamidine-VIIa/Soluble Tissue Factor: Unpredicted Conformation of the 192-193 Peptide Bond and Mapping of Ca2+, Mg2+, Na+, and Zn2+ Sites in Factor VIIa. J Biol Chem. 2006;281:24873–24888. doi: 10.1074/jbc.M509971200. [DOI] [PubMed] [Google Scholar]
- [53].Stubbs MT, Oschkinat H, Mayr I, Huber R, Angliker H, Stone SR, Bode W. The interaction of thrombin with fibrinogen. Eur J Biochem. 1992;206:187–195. doi: 10.1111/j.1432-1033.1992.tb16916.x. [DOI] [PubMed] [Google Scholar]
- [54].Lundblad RL, White GC. The interaction of thrombin with blood platelets. Platelets. 2005;16:373–385. doi: 10.1080/09537100500123568. [DOI] [PubMed] [Google Scholar]
- [55].Le Bonniec BF. In: Handbook of Proteolytic Enzymes. Third Edition. Rawlings ND, Salvesen G, editors. Academic Press; London: 2013. Chapter 643 - Thrombin; pp. 2915–2932. [Google Scholar]
- [56].Papaconstantinou ME, Gandhi PS, Chen Z, Bah A, Di Cera E. Na+ binding to meizothrombin desF1. Cell Mol Life Sci. 2008;65:3688–3697. doi: 10.1007/s00018-008-8502-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [57].Rezaie AR. Contribution of the 60 Loop To Substrate and Inhibitor Specificity of Thrombin. Blood. 2004;104:1734. [Google Scholar]
- [58].Liu T, Qian W-J, Gritsenko MA, Camp DG, Monroe ME, Moore RJ, Smith RD. Human Plasma N-Glycoproteome Analysis by Immunoaffinity Subtraction, Hydrazide Chemistry, and Mass Spectrometry. J Proteome Res. 2005;4:2070–2080. doi: 10.1021/pr0502065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [59].Pineda AO, Carrell CJ, Bush LA, Prasad S, Caccia S, Chen ZW, Mathews FS, Di Cera E. Molecular dissection of Na+ binding to thrombin. J Biol Chem. 2004;279:31842–31853. doi: 10.1074/jbc.M401756200. [DOI] [PubMed] [Google Scholar]
- [60].Di Cera E, Guinto ER, Vindigni A, Dang QD, Ayala YM, Wuyi M, Tulinsky A. The Na+ Binding Site of Thrombin. J Biol Chem. 1995;270:22089–22092. doi: 10.1074/jbc.270.38.22089. [DOI] [PubMed] [Google Scholar]
- [61].Niu W, Chen Z, Gandhi PS, Vogt AD, Pozzi N, Pelc LA, Zapata F, Di Cera E. Crystallographic and Kinetic Evidence of Allostery in a Trypsin-like Protease. Biochemistry. 2011;50:6301–6307. doi: 10.1021/bi200878c. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [62].Li W, Johnson DJD, Esmon CT, Huntington JA. Structure of the antithrombin-thrombin-heparin ternary complex reveals the antithrombotic mechanism of heparin. Nat Struct Mol Biol. 2004;11:857–862. doi: 10.1038/nsmb811. [DOI] [PubMed] [Google Scholar]
- [63].Chahal G, Thorpe M, Hellman L. The Importance of Exosite Interactions for Substrate Cleavage by Human Thrombin. PLOS ONE. 2015;10:e0129511. doi: 10.1371/journal.pone.0129511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [64].Friedrich R, Panizzi P, Fuentes-Prior P, Richter K, Verhamme I, Anderson PJ, Kawabata S-I, Huber R, Bode W, Bock PE. Staphylocoagulase is a prototype for the mechanism of cofactor-induced zymogen activation. Nature. 2003;425:535–539. doi: 10.1038/nature01962. [DOI] [PubMed] [Google Scholar]
- [65].Rezaie AR. Regulation of the Protein C Anticoagulant and Antiinflammatory Pathways. Curr Med Chem. 2010;17:2059–2069. doi: 10.2174/092986710791233706. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [66].Fuentes-Prior P, Iwanaga Y, Huber R, Pagila R, Rumennik G, Seto M, Morser J, Light DR, Bode W. Structural basis for the anticoagulant activity of the thrombin-thrombomodulin complex. Nature. 2000;404:518–525. doi: 10.1038/35006683. [DOI] [PubMed] [Google Scholar]
- [67].Dahlbäck B, Villoutreix BO. The anticoagulant protein C pathway. Febs Lett. 2005;579:3310–3316. doi: 10.1016/j.febslet.2005.03.001. [DOI] [PubMed] [Google Scholar]
- [68].Mather T, Oganessyan V, Hof P, Huber R, Foundling S, Esmon C, Bode W. The 2.8 A crystal structure of Gla-domainless activated protein C. EMBO J. 1996;15:6822–6831. [PMC free article] [PubMed] [Google Scholar]
- [69].Cramer TJ, Gale AJ. Function of the activated protein C (APC) autolysis loop in activated FVIII inactivation. Br J Haematol. 2011;153:644–654. doi: 10.1111/j.1365-2141.2011.08657.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [70].Pezeshkpoor B, Castoldi E, Mahler A, Hanel D, Müller J, Hamedani NS, Biswas A, Oldenburg J, Pavlova A. Identification and functional characterization of a novel F5 mutation (Ala512Val, FVBonn) associated with activated protein C resistance. J Thromb Haemost. 2016;14:1353–1363. doi: 10.1111/jth.13339. [DOI] [PubMed] [Google Scholar]
- [71].Mayer M. Biochemical and biological aspects of the plasminogen activation system. Clin Biochem. 1990;23:197–211. doi: 10.1016/0009-9120(90)90601-p. [DOI] [PubMed] [Google Scholar]
- [72].Ke S-H, Coombs GS, Tachias K, Corey DR, Madison EL. Optimal Subsite Occupancy and Design of a Selective Inhibitor of Urokinase. J Biol Chem. 1997;272:20456–20462. doi: 10.1074/jbc.272.33.20456. [DOI] [PubMed] [Google Scholar]
- [73].Mahmood N, Mihalcioiu C, Rabbani SA. Multifaceted Role of the Urokinase-Type Plasminogen Activator (uPA) and Its Receptor (uPAR): Diagnostic, Prognostic, and Therapeutic Applications. Front Oncol. 2018;8:24. doi: 10.3389/fonc.2018.00024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [74].Spraggon G, Phillips C, Nowak UK, Ponting CP, Saunders D, Dobson CM, Stuart DI, Jones EY. The crystal structure of the catalytic domain of human urokinase-type plasminogen activator. Structure. 1995;3:681–691. doi: 10.1016/s0969-2126(01)00203-9. [DOI] [PubMed] [Google Scholar]
- [75].Renatus M, Engh RA, Stubbs MT, Huber R, Fischer S, Kohnert U, Bode W. Lysine 156 promotes the anomalous proenzyme activity of tPA: X-ray crystal structure of single-chain human tPA. EMBO J. 1997;16:4797–4805. doi: 10.1093/emboj/16.16.4797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [76].Lamba D, Bauer M, Huber R, Fischer S, Rudolph R, Kohnert U, Bode W. The 2.3 Å Crystal Structure of the Catalytic Domain of Recombinant Two-chain Human Tissue-type Plasminogen Activator. J Mol Biol. 1996;258:117–135. doi: 10.1006/jmbi.1996.0238. [DOI] [PubMed] [Google Scholar]
- [77].Melchor JP, Strickland S. Tissue plasminogen activator in central nervous system physiology and pathology. Thromb Haemost. 2005;93:655–660. doi: 10.1160/TH04-12-0838. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [78].Peisach E, Wang J, de los Santos T, Reich E, Ringe D. Crystal Structure of the Proenzyme Domain of Plasminogen. Biochemistry. 1999;38:11180–11188. doi: 10.1021/bi991130r. [DOI] [PubMed] [Google Scholar]
- [79].Law Ruby HP, Caradoc-Davies T, Cowieson N, Horvath Anita J, Quek Adam J, Encarnacao Joanna A, Steer D, Cowan A, Zhang Q, Lu Bernadine GC, Pike Robert N, et al. The X-ray Crystal Structure of Full-Length Human Plasminogen. Cell Rep. 2012;1:185–190. doi: 10.1016/j.celrep.2012.02.012. [DOI] [PubMed] [Google Scholar]
- [80].Wu G, Quek AJ, Caradoc-Davies TT, Ekkel SM, Mazzitelli B, Whisstock JC, Law RHP. Structural studies of plasmin inhibition. Biochem Soc Trans. 2019:BST20180211. doi: 10.1042/BST20180211. [DOI] [PubMed] [Google Scholar]
- [81].Swedberg JE, Harris JM. Plasmin Substrate Binding Site Cooperativity Guides the Design of Potent Peptide Aldehyde Inhibitors. Biochemistry. 2011;50:8454–8462. doi: 10.1021/bi201203y. [DOI] [PubMed] [Google Scholar]
- [82].Wang X, Lin X, Loy JA, Tang J, Zhang XC. Crystal Structure of the Catalytic Domain of Human Plasmin Complexed with Streptokinase. Science. 1998;281:1662–1665. doi: 10.1126/science.281.5383.1662. [DOI] [PubMed] [Google Scholar]
- [83].Dzik JM. The ancestry and cumulative evolution of immune reactions. Acta Biochim Pol. 2010;57:443–466. [PubMed] [Google Scholar]
- [84].Kairies N, Beisel H-G, Fuentes-Prior P, Tsuda R, Muta T, Iwanaga S, Bode W, Huber R, Kawabata S-i. The 2.0-Å crystal structure of tachylectin 5A provides evidence for the common origin of the innate immunity and the blood coagulation systems. Proc Natl Acad Sci USA. 2001;98:13519–13524. doi: 10.1073/pnas.201523798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [85].Huber-Lang M, Ekdahl KN, Wiegner R, Fromell K, Nilsson B. Auxiliary activation of the complement system and its importance for the pathophysiology of clinical conditions. Semin Immunopathol. 2018;40:87–102. doi: 10.1007/s00281-017-0646-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [86].Conway EM. Reincarnation of ancient links between coagulation and complement. J Thromb Haemost. 2015;13:S121–S132. doi: 10.1111/jth.12950. [DOI] [PubMed] [Google Scholar]
- [87].Sim RB, Tsiftsoglou SA. Proteases of the complement system. Biochem Soc Trans. 2004;32:21–27. doi: 10.1042/bst0320021. [DOI] [PubMed] [Google Scholar]
- [88].Zewde N, Morikis D. A computational model for the evaluation of complement system regulation under homeostasis, disease, and drug intervention. PLOS ONE. 2018;13:e0198644. doi: 10.1371/journal.pone.0198644. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [89].Ehrnthaller C, Ignatius A, Gebhard F, Huber-Lang M. New insights of an old defense system: structure, function, and clinical relevance of the complement system. Mol Med. 2011;17:317–329. doi: 10.2119/molmed.2010.00149. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [90].Law SK, Dodds AW. The internal thioester and the covalent binding properties of the complement proteins C3 and C4. Protein Sci. 1997;6:263–274. doi: 10.1002/pro.5560060201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [91].Nilsson SC, Sim RB, Lea SM, Fremeaux-Bacchi V, Blom AM. Complement factor I in health and disease. Mol Immunol. 2011;48:1611–1620. doi: 10.1016/j.molimm.2011.04.004. [DOI] [PubMed] [Google Scholar]
- [92].Budayova-Spano M, Lacroix M, Thielens NM, Arlaud GJ, Fontecilla-Camps JC, Gaboriaud C. The crystal structure of the zymogen catalytic domain of complement protease C1r reveals that a disruptive mechanical stress is required to trigger activation of the C1 complex. EMBO J. 2002;21:231–239. doi: 10.1093/emboj/21.3.231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [93].Makou E, Herbert Andrew P, Barlow Paul N. Creating functional sophistication from simple protein building blocks, exemplified by factor H and the regulators of complement activation. Biochem Soc Trans. 2015;43:812–818. doi: 10.1042/BST20150074. [DOI] [PubMed] [Google Scholar]
- [94].Thielens NM, Tedesco F, Bohlson SS, Gaboriaud C, Tenner AJ. C1q: A fresh look upon an old molecule. Mol Immunol. 2017;89:73–83. doi: 10.1016/j.molimm.2017.05.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [95].Mortensen SA, Sander B, Jensen RK, Pedersen JS, Golas MM, Jensenius JC, Hansen AG, Thiel S, Andersen GR. Structure and activation of C1, the complex initiating the classical pathway of the complement cascade. Proc Natl Acad Sci USA. 2017;114:986–991. doi: 10.1073/pnas.1616998114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [96].Almitairi JOM, Venkatraman Girija U, Furze CM, Simpson-Gray X, Badakshi F, Marshall JE, Schwaeble WJ, Mitchell DA, Moody PCE, Wallis R. Structure of the C1r–C1s interaction of the C1 complex of complement activation. Proc Natl Acad Sci USA. 2018;115:768–773. doi: 10.1073/pnas.1718709115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [97].Lintner KE, Wu YL, Yang Y, Spencer CH, Hauptmann G, Hebert LA, Atkinson JP, Yu CY. Early Components of the Complement Classical Activation Pathway in Human Systemic Autoimmune Diseases. Front Immunol. 2016;7:36. doi: 10.3389/fimmu.2016.00036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [98].Krishnan V, Xu Y, Macon K, Volanakis JE, Narayana SVL. The structure of C2b, a fragment of complement component C2 produced during C3 convertase formation. Acta Cryst D. 2009;65:266–274. doi: 10.1107/S0907444909000389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [99].Milder FJ, Raaijmakers HCA, Vandeputte MDAA, Schouten A, Huizinga EG, Romijn RA, Hemrika W, Roos A, Daha MR, Gros P. Structure of Complement Component C2a: Implications for Convertase Formation and Substrate Binding. Structure. 2006;14:1587–1597. doi: 10.1016/j.str.2006.08.008. [DOI] [PubMed] [Google Scholar]
- [100].Mayilyan KR. Complement genetics, deficiencies, and disease associations. Protein Cell. 2012;3:487–496. doi: 10.1007/s13238-012-2924-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [101].Cascella R, Ragazzo M, Strafella C, Missiroli F, Borgiani P, Angelucci F, Marsella LT, Cusumano A, Novelli G, Ricci F, Giardina E. Age-Related Macular Degeneration: Insights into Inflammatory Genes. J Ophthalmol. 2014;2014:582842. doi: 10.1155/2014/582842. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [102].Tsiftsoglou SA, Sim RB. Human Complement Factor I Does Not Require Cofactors for Cleavage of Synthetic Substrates. J Immunol. 2004;173:367–375. doi: 10.4049/jimmunol.173.1.367. [DOI] [PubMed] [Google Scholar]
- [103].Roversi P, Johnson S, Caesar JJE, McLean F, Leath KJ, Tsiftsoglou SA, Morgan BP, Harris CL, Sim RB, Lea SM. Structural basis for complement factor I control and its disease-associated sequence polymorphisms. Proc Natl Acad Sci U S A. 2011;108:12839–12844. doi: 10.1073/pnas.1102167108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [104].Sim RB. In: Handbook of Proteolytic Enzymes. Third Edition. Rawlings ND, Salvesen G, editors. Academic Press; London: 2013. Chapter 636 - Complement Factor I; pp. 2875–2880. [Google Scholar]
- [105].Dobó J, Harmat V, Beinrohr L, Sebestyén E, Závodszky P, Gál P. MASP-1, a Promiscuous Complement Protease: Structure of Its Catalytic Region Reveals the Basis of Its Broad Specificity. J Immunol. 2009;183:1207–1214. doi: 10.4049/jimmunol.0901141. [DOI] [PubMed] [Google Scholar]
- [106].Harmat V, Gál P, Kardos J, Szilágyi K, Ambrus G, Végh B, Náray-Szabó G, Závodszky P. The Structure of MBL-associated Serine Protease-2 Reveals that Identical Substrate Specificities of C1s and MASP-2 are Realized Through Different Sets of Enzyme–Substrate Interactions. J Mol Biol. 2004;342:1533–1546. doi: 10.1016/j.jmb.2004.07.014. [DOI] [PubMed] [Google Scholar]
- [107].Gál P, Harmat V, Kocsis A, Bián T, Barna L, Ambrus G, Végh B, Balczer J, Sim RB, Náray-Szabó G, Závodszky P. A True Autoactivating Enzyme: STRUCTURAL INSIGHT INTO MANNOSE-BINDING LECTIN-ASSOCIATED SERINE PROTEASE-2 ACTIVATIONS. J Biol Chem. 2005;280:33435–33444. doi: 10.1074/jbc.M506051200. [DOI] [PubMed] [Google Scholar]
- [108].Milder FJ, Gomes L, Schouten A, Janssen BJC, Huizinga EG, Romijn RA, Hemrika W, Roos A, Daha MR, Gros P. Factor B structure provides insights into activation of the central protease of the complement system. Nat Struct Mol Biol. 2007;14:224–228. doi: 10.1038/nsmb1210. [DOI] [PubMed] [Google Scholar]
- [109].Ponnuraj K, Xu Y, Macon K, Moore D, Volanakis JE, Narayana SVL. Structural Analysis of Engineered Bb Fragment of Complement Factor B: Insights into the Activation Mechanism of the Alternative Pathway C3-Convertase. Mol Cell. 2004;14:17–28. doi: 10.1016/s1097-2765(04)00160-1. [DOI] [PubMed] [Google Scholar]
- [110].Dobó J, Szakács D, Oroszlán G, Kortvely E, Kiss B, Boros E, Szász R, Závodszky P, Gál P, Pál G. MASP-3 is the exclusive pro-factor D activator in resting blood: the lectin and the alternative complement pathways are fundamentally linked. Sci Rep. 2016;6:31877. doi: 10.1038/srep31877. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [111].Jing H, Macon KJ, Moore D, DeLucas LJ, Volanakis JE, Narayana SVL. Structural basis of profactor D activation: from a highly flexible zymogen to a novel self-inhibited serine protease, complement factor D. EMBO J. 1999;18:804–814. doi: 10.1093/emboj/18.4.804. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [112].Forneris F, Ricklin D, Wu J, Tzekou A, Wallace RS, Lambris JD, Gros P. Structures of C3b in Complex with Factors B and D Give Insight into Complement Convertase Formation. Science. 2010;330:1816–1820. doi: 10.1126/science.1195821. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [113].Forneris F, Gros P. In: Handbook of Proteolytic Enzymes. Third Edition. Rawlings ND, Salvesen G, editors. Academic Press; London: 2013. Chapter 630 - Complement Factor D; pp. 2841–2848. [Google Scholar]
- [114].Yang C-Y, Phillips JG, Stuckey JA, Bai L, Sun H, Delproposto J, Brown WC, Chinnaswamy K. Buried Hydrogen Bond Interactions Contribute to the High Potency of Complement Factor D Inhibitors. ACS Med Chem Lett. 2016;7:1092–1096. doi: 10.1021/acsmedchemlett.6b00299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [115].Korkmaz B, Horwitz MS, Jenne DE, Gauthier F. Neutrophil Elastase, Proteinase 3, and Cathepsin G as Therapeutic Targets in Human Diseases. Pharmacol Rev. 2010;62:726–759. doi: 10.1124/pr.110.002733. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [116].Hajjar E, Broemstrup T, Kantari C, Witko-Sarsat V, Reuter N. Structures of human proteinase3 and neutrophil elastase–so similar yet so different. FEBS J. 2010;277:2238–2254. doi: 10.1111/j.1742-4658.2010.07659.x. [DOI] [PubMed] [Google Scholar]
- [117].Hochscherf J, Pietsch M, Tieu W, Kuan K, Abell AD, Gutschow M, Niefind K. Crystal structure of highly glycosylated human leukocyte elastase in complex with an S2’ site binding inhibitor. Acta Cryst F. 2018;74:480–489. doi: 10.1107/S2053230X1800537X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [118].Soehnlein O, Lindbom L. Neutrophil-derived azurocidin alarms the immune system. J Leukoc Biol. 2009;85:344–351. doi: 10.1189/jlb.0808495. [DOI] [PubMed] [Google Scholar]
- [119].Olczak M, Indyk K, Olczak T. Reconstitution of human azurocidin catalytic triad and proteolytic activity by site-directed mutagenesis. Biol Chem. 2008;389:955–962. doi: 10.1515/BC.2008.101. [DOI] [PubMed] [Google Scholar]
- [120].Hof P, Mayr I, Huber R, Korzus E, Potempa J, Travis J, Powers JC, Bode W. The 1.8 A crystal structure of human cathepsin G in complex with Suc-Val-Pro-PheP-(OPh)2: a Janus-faced proteinase with two opposite specificities. EMBO J. 1996;15:5481–5491. [PMC free article] [PubMed] [Google Scholar]
- [121].Perera NC, Schilling O, Kittel H, Back W, Kremmer E, Jenne DE. NSP4, an elastase-related protease in human neutrophils with arginine specificity. Proc Natl Acad Sci USA. 2012;109:6229–6234. doi: 10.1073/pnas.1200470109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [122].Lin SJ, Dong Ken C, Eigenbrot C, van Lookeren Campagne M, Kirchhofer D. Structures of Neutrophil Serine Protease 4 Reveal an Unusual Mechanism of Substrate Recognition by a Trypsin-Fold Protease. Structure. 2014;22:1333–1340. doi: 10.1016/j.str.2014.07.008. [DOI] [PubMed] [Google Scholar]
- [123].Caughey GH. Mast cell proteases as pharmacological targets. Eur J Pharmacol. 2016;778:44–55. doi: 10.1016/j.ejphar.2015.04.045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [124].Caughey GH. In: Handbook of Proteolytic Enzymes. Third Edition. Rawlings ND, Salvesen G, editors. Academic Press; London: 2013. Chapter 590 - Chymases; pp. 2675–2683. [Google Scholar]
- [125].Trivedi NN, Caughey GH. In: Handbook of Proteolytic Enzymes. Third Edition. Rawlings ND, Salvesen G, editors. Academic Press; London: 2013. Chapter 591 - Human α-, β- and δ-Tryptases; pp. 2683–2693. [Google Scholar]
- [126].Trivedi NN, Caughey GH. In: Handbook of Proteolytic Enzymes. Third Edition. Rawlings ND, Salvesen G, editors. Academic Press; London: 2013. Chapter 592 - γ-Tryptase; pp. 2694–2697. [Google Scholar]
- [127].Selwood T, Wang Z-M, McCaslin DR, Schechter NM. Diverse Stability and Catalytic Properties of Human Tryptase α and β Isoforms are Mediated by Residue Differences at the S1 Pocket. Biochemistry. 2002;41:3329–3340. doi: 10.1021/bi015662v. [DOI] [PubMed] [Google Scholar]
- [128].Pereira PJB, Bergner A, Macedo-Ribeiro S, Huber R, Matschiner G, Fritz H, Sommerhoff CP, Bode W. Human β-tryptase is a ring-like tetramer with active sites facing a central pore. Nature. 1998;392:306–311. doi: 10.1038/32703. [DOI] [PubMed] [Google Scholar]
- [129].Paesen GC, Siebold C, Harlos K, Peacey MF, Nuttall PA, Stuart DI. A Tick Protein with a Modified Kunitz Fold Inhibits Human Tryptase. J Mol Biol. 2007;368:1172–1186. doi: 10.1016/j.jmb.2007.03.011. [DOI] [PubMed] [Google Scholar]
- [130].Trivedi NN, Raymond WW, Caughey GH. Chimerism, point mutation, and truncation dramatically transformed mast cell δ-tryptases during primate evolution. J Allergy Clin Immunol. 2008;121:1262–1268. doi: 10.1016/j.jaci.2008.01.019. [DOI] [PubMed] [Google Scholar]
- [131].Atiakshin D, Buchwalow I, Samoilova V, Tiemann MJH, Biology C. Tryptase as a polyfunctional component of mast cells. Histochem Cell Biol. 2018;149:461–477. doi: 10.1007/s00418-018-1659-8. [DOI] [PubMed] [Google Scholar]
- [132].Masson D, Tschopp J. A family of serine esterases in lytic granules of cytolytic T lymphocytes. Cell. 1987;49:679–685. doi: 10.1016/0092-8674(87)90544-7. [DOI] [PubMed] [Google Scholar]
- [133].Voskoboinik I, Whisstock JC, Trapani JA. Perforin and granzymes: function, dysfunction and human pathology. Nat Rev Immunol. 2015;15:388. doi: 10.1038/nri3839. [DOI] [PubMed] [Google Scholar]
- [134].Vahedi F, Fraleigh N, Vlasschaert C, McElhaney J, Hanifi-Moghaddam P. Human granzymes: Related but far apart. Med Hypotheses. 2014;83:688–693. doi: 10.1016/j.mehy.2014.09.019. [DOI] [PubMed] [Google Scholar]
- [135].Harris JL, Peterson EP, Hudig D, Thornberry NA, Craik CS. Definition and Redesign of the Extended Substrate Specificity of Granzyme B. J Biol Chem. 1998;273:27364–27373. doi: 10.1074/jbc.273.42.27364. [DOI] [PubMed] [Google Scholar]
- [136].Rotonda J, Garcia-Calvo M, Bull HG, Geissler WM, McKeever BM, Willoughby CA, Thornberry NA, Becker JW. The three-dimensional structure of human granzyme B compared to caspase-3, key mediators of cell death with cleavage specificity for aspartic acid in P1. Chem Biol. 2001;8:357–368. doi: 10.1016/s1074-5521(01)00018-7. [DOI] [PubMed] [Google Scholar]
- [137].Wang L, Zhang K, Wu L, Liu S, Zhang H, Zhou Q, Tong L, Sun F, Fan Z. Structural Insights into the Substrate Specificity of Human Granzyme H: The Functional Roles of a Novel RKR Motif. J Immunol. 2012;188:765–773. doi: 10.4049/jimmunol.1101381. [DOI] [PubMed] [Google Scholar]
- [138].Tang H, Li C, Wang L, Zhang H, Fan Z. Granzyme H of Cytotoxic Lymphocytes Is Required for Clearance of the Hepatitis B Virus through Cleavage of the Hepatitis B Virus X Protein. J Immunol. 2012;188:824–831. doi: 10.4049/jimmunol.1102205. [DOI] [PubMed] [Google Scholar]
- [139].Plasman K, Demol H, Bird PI, Gevaert K, Van Damme P. Substrate Specificities of the Granzyme Tryptases A and K. J Proteome Res. 2014;13:6067–6077. doi: 10.1021/pr500968d. [DOI] [PubMed] [Google Scholar]
- [140].Susanto O, Stewart SE, Voskoboinik I, Brasacchio D, Hagn M, Ellis S, Asquith S, Sedelies KA, Bird PI, Waterhouse NJ, Trapani JA. Mouse granzyme A induces a novel death with writhing morphology that is mechanistically distinct from granzyme B-induced apoptosis. Cell Death Differ. 2013;20:1183. doi: 10.1038/cdd.2013.59. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [141].Magdolen V, Sommerhoff Christian P, Fritz H, Schmitt M. Volume 1 Characterization, regulation, and interactions within the protease web. De Gruyter; Berlin: 2012. [Google Scholar]
- [142].Kalinska M, Meyer-Hoffert U, Kantyka T, Potempa J. Kallikreins – The melting pot of activity and function. Biochimie. 2016;122:270–282. doi: 10.1016/j.biochi.2015.09.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [143].Schmaier AH. In: Handbook of Proteolytic Enzymes. Third Edition. Rawlings ND, Salvesen G, editors. Academic Press; London: 2013. Chapter 638 - Prekallikrein and Plasma Kallikrein; pp. 2885–2892. [Google Scholar]
- [144].Shaw JLV, Diamandis EP. Distribution of 15 Human Kallikreins in Tissues and Biological Fluids. Clin Chem. 2007;53:1423–1432. doi: 10.1373/clinchem.2007.088104. [DOI] [PubMed] [Google Scholar]
- [145].Yoon H, Laxmikanthan G, Lee J, Blaber SI, Rodriguez A, Kogot JM, Scarisbrick IA, Blaber M. Activation Profiles and Regulatory Cascades of the Human Kallikrein-related Peptidases. J Biol Chem. 2007;282:31852–31864. doi: 10.1074/jbc.M705190200. [DOI] [PubMed] [Google Scholar]
- [146].Yoon H, Blaber SI, Debela M, Goettig P, Scarisbrick IA, Blaber M. A completed KLK activome profile: investigation of activation profiles of KLK9, 10 and 15. Biol Chem. 2009;390:373–377. doi: 10.1515/BC.2009.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [147].Sotiropoulou G, Pampalakis G, Diamandis EP. Functional Roles of Human Kallikrein-related Peptidases. J Biol Chem. 2009;284:32989–32994. doi: 10.1074/jbc.R109.027946. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [148].Dorn J, Beaufort N, Schmitt M, Diamandis EP, Goettig P, Magdolen V. Function and clinical relevance of kallikrein-related peptidases and other serine proteases in gynecological cancers. Crit Rev Clin Lab Sci. 2014;51:63–84. doi: 10.3109/10408363.2013.865701. [DOI] [PubMed] [Google Scholar]
- [149].Prassas I, Eissa A, Poda G, Diamandis EP. Unleashing the therapeutic potential of human kallikrein-related serine proteases. Nat Rev Drug Discov. 2015;14:183–202. doi: 10.1038/nrd4534. [DOI] [PubMed] [Google Scholar]
- [150].Kryza T, Silva ML, Loessner D, Heuzé-Vourc’h N, Clements JA. The kallikrein-related peptidase family: Dysregulation and functions during cancer progression. Biochimie. 2016;122:283–299. doi: 10.1016/j.biochi.2015.09.002. [DOI] [PubMed] [Google Scholar]
- [151].Loessner D, Goettig P, Preis S, Felber J, Bronger H, Clements JA, Dorn J, Magdolen V. Kallikrein-related peptidases represent attractive therapeutic targets for ovarian cancer. Expert Opin Ther Targets. 2018;22:745–763. doi: 10.1080/14728222.2018.1512587. [DOI] [PubMed] [Google Scholar]
- [152].Avgeris M, Scorilas A. Kallikrein-related peptidases (KLKs) as emerging therapeutic targets: focus on prostate cancer and skin pathologies. Expert Opin Ther Targets. 2016;20:801–818. doi: 10.1517/14728222.2016.1147560. [DOI] [PubMed] [Google Scholar]
- [153].Debela M, Beaufort N, Magdolen V, Schechter NM, Craik CS, Schmitt M, Bode W, Goettig P. Structures and specificity of the human kallikrein-related peptidases KLK 4, 5, 6, and 7. Biol Chem. 2008;389:623–632. doi: 10.1515/BC.2008.075. [DOI] [PubMed] [Google Scholar]
- [154].Swedberg JE, de Veer SJ, Harris JM. Natural and engineered kallikrein inhibitors: an emerging pharmacopoeia. Biol Chem. 2010;391:357–374. doi: 10.1515/BC.2010.037. [DOI] [PubMed] [Google Scholar]
- [155].Masurier N, Arama DP, El Amri C, Lisowski V. Inhibitors of kallikrein-related peptidases: An overview. Med Res Rev. 2018;38:655–683. doi: 10.1002/med.21451. [DOI] [PubMed] [Google Scholar]
- [156].Rhaleb NE, Yang XP, Carretero OA. The kallikrein-kinin system as a regulator of cardiovascular and renal function. Compr Physiol. 2011;1:971–993. doi: 10.1002/cphy.c100053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [157].Chao J. In: Handbook of Proteolytic Enzymes. Third Edition. Rawlings ND, Salvesen G, editors. Academic Press; London: 2013. Chapter 607 - Human Kallikrein 1,Tissue Kallikrein; pp. 2757–2761. [Google Scholar]
- [158].Pathak M, Shen Wong S, Dreveny I, Emsley J. Structure of plasma and tissue kallikreins. Thromb Haemost. 2013;110:423–433. doi: 10.1160/TH12-11-0840. [DOI] [PubMed] [Google Scholar]
- [159].Xu M, Chen Y, Xu P, Andreasen PA, Jiang L, Li J, Huang M. Crystal structure of plasma kallikrein reveals the unusual flexibility of the S1 pocket triggered by Glu217. Febs Lett. 2018;592:2658–2667. doi: 10.1002/1873-3468.13191. [DOI] [PubMed] [Google Scholar]
- [160].Kellermann J, Lottspeich F, Geiger R, Deutzmann R. Human urinary kallikrein--amino acid sequence and carbohydrate attachment sites. Protein Seq Data Anal. 1988;1:177–182. [PubMed] [Google Scholar]
- [161].Laxmikanthan G, Blaber SI, Bernett MJ, Scarisbrick IA, Juliano MA, Blaber M. 1.70 angstrom X-ray structure of human apo kallikrein 1: Structural changes upon peptide inhibitor/substrate binding. Proteins. 2005;58:802–814. doi: 10.1002/prot.20368. [DOI] [PubMed] [Google Scholar]
- [162].Stura EA, Muller BH, Bossus M, Michel S, Jolivet-Reynaud C, Ducancel F. Crystal structure of human prostate-specific antigen in a sandwich antibody complex. J Mol Biol. 2011;414:530–544. doi: 10.1016/j.jmb.2011.10.007. [DOI] [PubMed] [Google Scholar]
- [163].Menez R, Michel S, Muller BH, Bossus M, Ducancel F, Jolivet-Reynaud C, Stura EA. Crystal Structure of a Ternary Complex between Human Prostate-specific Antigen, Its Substrate Acyl Intermediate and an Activating Antibody. J Mol Biol. 2008;376:1021–1033. doi: 10.1016/j.jmb.2007.11.052. [DOI] [PubMed] [Google Scholar]
- [164].Skala W, Utzschneider DT, Magdolen V, Debela M, Guo S, Craik CS, Brandstetter H, Goettig P. Structure-Function Analyses of Human Kallikrein-Related Peptidase 2 Establish the 99-Loop as Master Regulator of Activity. J Biol Chem. 2014;289:34267–34283. doi: 10.1074/jbc.M114.598201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [165].Carvalho AL, Sanz L, Barettino D, Romero A, Calvete JJ, Romão MJ. Crystal Structure of a Prostate Kallikrein Isolated from Stallion Seminal Plasma: A Homologue of Human PSA. J Mol Biol. 2002;322:325–337. doi: 10.1016/s0022-2836(02)00705-2. [DOI] [PubMed] [Google Scholar]
- [166].Guo S, Skala W, Magdolen V, Briza P, Biniossek ML, Schilling O, Kellermann J, Brandstetter H, Goettig P. A Single Glycan at the 99-Loop of Human Kallikrein-related Peptidase 2 Regulates Activation and Enzymatic Activity. J Biol Chem. 2016;291:593–604. doi: 10.1074/jbc.M115.691097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [167].Debela M, Magdolen V, Schechter N, Valachova M, Lottspeich F, Craik CS, Choe Y, Bode W, Goettig P. Specificity Profiling of Seven Human Tissue Kallikreins Reveals Individual Subsite Preferences. J Biol Chem. 2006;281:25678–25688. doi: 10.1074/jbc.M602372200. [DOI] [PubMed] [Google Scholar]
- [168].LeBeau AM, Singh P, Isaacs JT, Denmeade SR. Prostate-Specific Antigen is a Chymotrypsin-like Serine Protease with Unique P1 Substrate Specificity. Biochemistry. 2009;48:3490–3496. doi: 10.1021/bi9001858. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [169].Guo S, Briza P, Magdolen V, Brandstetter H, Goettig P. Activation and activity of glycosylated KLKs 3, 4 and 11. Biol Chem. 2018;399:1009–1022. doi: 10.1515/hsz-2018-0148. [DOI] [PubMed] [Google Scholar]
- [170].Beaufort N, Plaza K, Utzschneider D, Schwarz A, Burkhart Julia M, Creutzburg S, Debela M, Schmitt M, Ries C, Magdolen V. Interdependence of kallikrein-related peptidases in proteolytic networks. Biol Chem. 2010;391:581–587. doi: 10.1515/BC.2010.055. [DOI] [PubMed] [Google Scholar]
- [171].Yamakoshi Y, Simmer JP, Bartlett JD, Karakida T, Oida S. MMP20 and KLK4 activation and inactivation interactions in vitro. Arch Oral Biol. 2013;58:1569–1577. doi: 10.1016/j.archoralbio.2013.08.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [172].Nakamura T, Mitsui S, Okui A, Miki T, Yamaguchi N. Molecular cloning and expression of a variant form of hippostasin/KLK11 in prostate. Prostate. 2003;54:299–305. doi: 10.1002/pros.10191. [DOI] [PubMed] [Google Scholar]
- [173].Goettig P, Magdolen V, Brandstetter H. Natural and synthetic inhibitors of kallikrein-related peptidases (KLKs) Biochimie. 2010;92:1546–1567. doi: 10.1016/j.biochi.2010.06.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [174].Debela M, Magdolen V, Grimminger V, Sommerhoff C, Messerschmidt A, Huber R, Friedrich R, Bode W, Goettig P. Crystal structures of human tissue kallikrein 4: activity modulation by a specific zinc binding site. J Mol Biol. 2006;362:1094–1107. doi: 10.1016/j.jmb.2006.08.003. [DOI] [PubMed] [Google Scholar]
- [175].Adham IM, Schlösser M, Engel W. In: Handbook of Proteolytic Enzymes. Third Edition. Rawlings ND, Salvesen G, editors. Academic Press; London: 2013. Chapter 602 - Acrosin; pp. 2731–2735. [Google Scholar]
- [176].Brattsand M, Stefansson K, Lundh C, Haasum Y, Egelrud T. A proteolytic cascade of kallikreins in the stratum corneum. J Invest Dermatol. 2005;124:198–203. doi: 10.1111/j.0022-202X.2004.23547.x. [DOI] [PubMed] [Google Scholar]
- [177].Deraison C, Bonnart C, Lopez F, Besson C, Robinson R, Jayakumar A, Wagberg F, Brattsand M, Hachem JP, Leonardsson G, Hovnanian A. LEKTI Fragments Specifically Inhibit KLK5, KLK7, and KLK14 and Control Desquamation through a pH-dependent Interaction. Mol Biol Cell. 2007;18:3607–3619. doi: 10.1091/mbc.E07-02-0124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [178].Igawa S, Kishibe M, Minami-Hori M, Honma M, Tsujimura H, Ishikawa J, Fujimura T, Murakami M, Ishida-Yamamoto A. Incomplete KLK7 Secretion and Upregulated LEKTI Expression Underlie Hyperkeratotic Stratum Corneum in Atopic Dermatitis. J Invest Dermatol. 2017;137:449–456. doi: 10.1016/j.jid.2016.10.015. [DOI] [PubMed] [Google Scholar]
- [179].Debela M, Goettig P, Magdolen V, Huber R, Schechter NM, Bode W. Structural Basis of the Zinc Inhibition of Human Tissue Kallikrein 5. J Mol Biol. 2007;373:1017–1031. doi: 10.1016/j.jmb.2007.08.042. [DOI] [PubMed] [Google Scholar]
- [180].Fujinaga M, James MN. Rat submaxillary gland serine protease, tonin. Structure solution and refinement at 1.8 A resolution. J Mol Biol. 1987;195:373–396. doi: 10.1016/0022-2836(87)90658-9. [DOI] [PubMed] [Google Scholar]
- [181].Debela M, Hess P, Magdolen V, Schechter NM, Steiner T, Huber R, Bode W, Goettig P. Chymotryptic specificity determinants in the 1.0 Å structure of the zinc-inhibited human tissue kallikrein 7. Proc Natl Acad Sci USA. 2007;104:16086–16091. doi: 10.1073/pnas.0707811104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [182].Fernández IS, Ständker L, Mägert H-J, Forssmann W-G, Giménez-Gallego G, Romero A. Crystal Structure of Human Epidermal Kallikrein 7 (hK7) Synthesized Directly in its Native State in E. coli: Insights into the Atomic Basis of its Inhibition by LEKTI Domain 6 (LD6) J Mol Biol. 2008;377:1488–1497. doi: 10.1016/j.jmb.2008.01.089. [DOI] [PubMed] [Google Scholar]
- [183].Vogt AD, Chakraborty P, Di Cera E. Kinetic Dissection of the Pre-existing Conformational Equilibrium in the Trypsin Fold. J Biol Chem. 2015;290:22435–22445. doi: 10.1074/jbc.M115.675538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [184].Emami N, Deperthes D, Malm J, Diamandis EP. Major Role of Human KLK14 in Seminal Clot Liquefaction. J Biol Chem. 2008;283:19561–19569. doi: 10.1074/jbc.M801194200. [DOI] [PubMed] [Google Scholar]
- [185].Gratio V, Loriot C, Virca GD, Oikonomopoulou K, Walker F, Diamandis EP, Hollenberg MD, Darmoul D. Kallikrein-Related Peptidase 14 Acts on Proteinase-Activated Receptor 2 to Induce Signaling Pathway in Colon Cancer Cells. Am J Pathol. 2011;179:2625–2636. doi: 10.1016/j.ajpath.2011.07.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [186].Hollenberg Morley D. KLKs and their hormone-like signaling actions: a new life for the PSA-KLK family. Biol Chem. 2014;395:915–929. doi: 10.1515/hsz-2014-0123. [DOI] [PubMed] [Google Scholar]
- [187].Andrea LOH, Virender B, Alexander DW, Craig SM, George SR. Human Kallikrein 6 Cerebrospinal Levels are Elevated in Multiple Sclerosis. Curr Drug Discov Technol. 2010;7:137–140. doi: 10.2174/157016310793180611. [DOI] [PubMed] [Google Scholar]
- [188].Angelo PF, Lima AR, Alves FM, Blaber SI, Scarisbrick IA, Blaber M, Juliano L, Juliano MA. Substrate specificity of human kallikrein 6: salt and glycosaminoglycan activation effects. J Biol Chem. 2006;281:3116–3126. doi: 10.1074/jbc.M510096200. [DOI] [PubMed] [Google Scholar]
- [189].Yoon H, Radulovic M, Scarisbrick Isobel A. Kallikrein-related peptidase 6 orchestrates astrocyte form and function through proteinase activated receptor-dependent mechanisms. Biol Chem. 2018;399:1041–1052. doi: 10.1515/hsz-2018-0122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [190].Shiosaka S, Ishikawa Y. Neuropsin - A possible modulator of synaptic plasticity. J Chem Neuroanat. 2011;42:24–29. doi: 10.1016/j.jchemneu.2011.05.014. [DOI] [PubMed] [Google Scholar]
- [191].Tamura H, Kawata M, Hamaguchi S, Ishikawa Y, Shiosaka S. Processing of neuregulin-1 by neuropsin regulates GABAergic neuron to control neural plasticity of the mouse hippocampus. J Neurosci. 2012;32:12657–12672. doi: 10.1523/JNEUROSCI.2542-12.2012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [192].Sonderegger P, Matsumoto-Miyai K. Activity-controlled proteolytic cleavage at the synapse. Trends Neurosci. 2014;37:413–423. doi: 10.1016/j.tins.2014.05.007. [DOI] [PubMed] [Google Scholar]
- [193].Debela M, Magdolen V, Skala W, Elsässer B, Schneider EL, Craik CS, Biniossek ML, Schilling O, Bode W, Brandstetter H, Goettig P. Structural determinants of specificity and regulation of activity in the allosteric loop network of human KLK8/neuropsin. Sci Rep. 2018;8:10705. doi: 10.1038/s41598-018-29058-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [194].Kishi T, Kato M, Shimizu T, Kato K, Matsumoto K, Yoshida S, Shiosaka S, Hakoshima T. Crystal structure of neuropsin, a hippocampal protease involved in kindling epileptogenesis. J Biol Chem. 1999;274:4220–4224. doi: 10.1074/jbc.274.7.4220. [DOI] [PubMed] [Google Scholar]
- [195].Riley BT, Ilyichova O, Costa MGS, Porebski BT, de Veer SJ, Swedberg JE, Kass I, Harris JM, Hoke DE, Buckle AM. Direct and indirect mechanisms of KLK4 inhibition revealed by structure and dynamics. Sci Rep. 2016;6:35385. doi: 10.1038/srep35385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [196].Kishibe M, Bando Y, Tanaka T, Ishida-Yamamoto A, Iizuka H, Yoshida S. Kallikrein-Related Peptidase 8-Dependent Skin Wound Healing Is Associated with Upregulation of Kallikrein-Related Peptidase 6 and PAR2. J Invest Dermatol. 2012;132:1717–1724. doi: 10.1038/jid.2012.18. [DOI] [PubMed] [Google Scholar]
- [197].Ahmed N, Dorn J, Napieralski R, Drecoll E, Kotzsch M, Goettig P, Zein E, Avril S, Kiechle M, Diamandis Eleftherios P, Schmitt M, et al. Clinical relevance of kallikrein-related peptidase 6 (KLK6) and 8 (KLK8) mRNA expression in advanced serous ovarian cancer. Biol Chem. 2016;397:1265–1276. doi: 10.1515/hsz-2016-0177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [198].Wang Y, Luo W, Wartmann T, Halangk W, Sahin-Tóth M, Reiser G. Mesotrypsin, a brain trypsin, activates selectively proteinase-activated receptor-1, but not proteinase-activated receptor-2, in rat astrocytes. J Neurochem. 2006;99:759–769. doi: 10.1111/j.1471-4159.2006.04105.x. [DOI] [PubMed] [Google Scholar]
- [199].Mitsui S, Osako Y, Yokoi F, Dang MT, Yuri K, Li Y, Yamaguchi N. A mental retardation gene, motopsin/neurotrypsin/prss12, modulates hippocampal function and social interaction. Eur J Neurosci. 2009;30:2368–2378. doi: 10.1111/j.1460-9568.2009.07029.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [200].Gisler C, Lüscher D, Schätzle P, Dürr S, Baici A, Galliciotti G, Reif R, Bolliger Marc F, Kunz B, Sonderegger P. Zymogen activation of neurotrypsin and neurotrypsin-dependent agrin cleavage on the cell surface are enhanced by glycosaminoglycans. Biochem J. 2013;453:83–100. doi: 10.1042/BJ20130166. [DOI] [PubMed] [Google Scholar]
- [201].Hu J, Lei H, Fei X, Liang S, Xu H, Qin D, Wang Y, Wu Y, Li B. NES1/KLK10 gene represses proliferation, enhances apoptosis and down-regulates glucose metabolism of PC3 prostate cancer cells. Sci Rep. 2015;5:17426. doi: 10.1038/srep17426. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [202].Geng X, Liu Y, Dreyer T, Bronger H, Drecoll E, Magdolen V, Dorn J. Elevated tumor tissue protein expression levels of kallikrein-related peptidases KLK10 and KLK11 are associated with a better prognosis in advanced high-grade serous ovarian cancer patients. Am J Cancer Res. 2018;8:1856–1864. [PMC free article] [PubMed] [Google Scholar]
- [203].Debela M, Magdolen V, Bode W, Brandstetter H, Goettig P. Structural basis for the Zn2+ inhibition of the zymogen-like kallikrein-related peptidase 10. Biol Chem. 2016;397:1251–1264. doi: 10.1515/hsz-2016-0205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [204].Chakraborty P, Di Cera E. Induced Fit Is a Special Case of Conformational Selection. Biochemistry. 2017;56:2853–2859. doi: 10.1021/acs.biochem.7b00340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [205].Kolmar H, Waller PR, Sauer RT. The DegP and DegQ periplasmic endoproteases of Escherichia coli: specificity for cleavage sites and substrate conformation. J Bacteriol. 1996;178:5925–5929. doi: 10.1128/jb.178.20.5925-5929.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [206].Weber N, Kolmar H. In: Handbook of Proteolytic Enzymes. Third Edition. Rawlings ND, Salvesen G, editors. Academic Press; London: 2013. Chapter 569 - DegP Protease; pp. 2567–2571. [Google Scholar]
- [207].Zurawa-Janicka D, Wenta T, Jarzab M, Skorko-Glonek J, Glaza P, Gieldon A, Ciarkowski J, Lipinska B. Structural insights into the activation mechanisms of human HtrA serine proteases. Arch Biochem Biophys. 2017;621:6–23. doi: 10.1016/j.abb.2017.04.004. [DOI] [PubMed] [Google Scholar]
- [208].Eigenbrot C, Ultsch M, Lipari Michael T, Moran P, Lin SJ, Ganesan R, Quan C, Tom J, Sandoval W, van Lookeren Campagne M, Kirchhofer D. Structural and Functional Analysis of HtrA1 and Its Subdomains. Structure. 2012;20:1040–1050. doi: 10.1016/j.str.2012.03.021. [DOI] [PubMed] [Google Scholar]
- [209].Chien J, Shridhar V. In: Handbook of Proteolytic Enzymes. Third Edition. Rawlings ND, Salvesen G, editors. Academic Press; London: 2013. Chapter 571 - HtrA1 Peptidase; pp. 2577–2584. [Google Scholar]
- [210].Chien J, He X, Shridhar V. Identification of tubulins as substrates of serine protease HtrA1 by mixture-based oriented peptide library screening. J Cell Biochem. 2009;107:253–263. doi: 10.1002/jcb.22121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [211].Truebestein L, Tennstaedt A, Mönig T, Krojer T, Canellas F, Kaiser M, Clausen T, Ehrmann M. Substrate-induced remodeling of the active site regulates human HTRA1 activity. Nat Struct Mol Biol. 2011;18:386. doi: 10.1038/nsmb.2013. [DOI] [PubMed] [Google Scholar]
- [212].Vande Walle L, Van Damme P, Lamkanfi M, Saelens X, Vandekerckhove J, Gevaert K, Vandenabeele P. Proteome-wide Identification of HtrA2/Omi Substrates. J Proteome Res. 2007;6:1006–1015. doi: 10.1021/pr060510d. [DOI] [PubMed] [Google Scholar]
- [213].Merski M, Moreira C, Abreu RMV, Ramos MJ, Fernandes PA, Martins LM, Pereira PJB, Macedo-Ribeiro S. Molecular motion regulates the activity of the Mitochondrial Serine Protease HtrA2. Cell Death Dis. 2017;8:e3119. doi: 10.1038/cddis.2017.487. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [214].Glaza P, Osipiuk J, Wenta T, Zurawa-Janicka D, Jarzab M, Lesner A, Banecki B, Skorko-Glonek J, Joachimiak A, Lipinska B. Structural and Functional Analysis of Human HtrA3 Protease and Its Subdomains. PLOS ONE. 2015;10:e0131142. doi: 10.1371/journal.pone.0131142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [215].Wenta T, Glaza P, Jarząb M, Zarzecka U, Żurawa-Janicka D, Lesner A, Skórko-Glonek J, Lipińska B. The role of the LB structural loop and its interactions with the PDZ domain of the human HtrA3 protease. Biochim Biophys Acta. 2017;1865:1141–1151. doi: 10.1016/j.bbapap.2017.06.013. [DOI] [PubMed] [Google Scholar]
- [216].Skorko-Glonek J, Zurawa-Janicka D, Koper T, Jarzab M, Figaj D, Glaza P, Lipinska B. HtrA Protease Family as Therapeutic Targets. Curr Pharm Des. 2013;19:977–1009. doi: 10.2174/1381612811319060003. [DOI] [PubMed] [Google Scholar]
- [217].Hara K, Shiga A, Fukutake T, Nozaki H, Miyashita A, Yokoseki A, Kawata H, Koyama A, Arima K, Takahashi T, Ikeda M, et al. Association of HTRA1 Mutations and Familial Ischemic Cerebral Small-Vessel Disease. N Engl J Med. 2009;360:1729–1739. doi: 10.1056/NEJMoa0801560. [DOI] [PubMed] [Google Scholar]
- [218].Zurawa-Janicka D, Skorko-Glonek J, Lipinska B. HtrA proteins as targets in therapy of cancer and other diseases. Expert Opin Ther Targets. 2010;14:665–679. doi: 10.1517/14728222.2010.487867. [DOI] [PubMed] [Google Scholar]
- [219].Bhuiyan MS, Fukunaga K. Mitochondrial Serine Protease HtrA2/Omi as a Potential Therapeutic Target. Curr Drug Targets. 2009;10:372–383. doi: 10.2174/138945009787846399. [DOI] [PubMed] [Google Scholar]
- [220].Singh H, Zhao M, Chen Q, Wang Y, Li Y, Kaitu’u-Lino TuJ, Tong S, Nie G. Human HtrA4 Expression Is Restricted to the Placenta, Is Significantly Up-Regulated in Early-Onset Preeclampsia, and High Levels of HtrA4 Cause Endothelial Dysfunction. J Clin Endocrinol Metab. 2015;100:E936–E945. doi: 10.1210/jc.2014-3969. [DOI] [PubMed] [Google Scholar]
- [221].Dynon K, Heng S, Puryer M, Li Y, Walton K, Endo Y, Nie G. HtrA3 as an Early Marker for Preeclampsia: Specific Monoclonal Antibodies and Sensitive High-Throughput Assays for Serum Screening. PLOS ONE. 2012;7:e45956. doi: 10.1371/journal.pone.0045956. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [222].Bugge TH, Antalis TM, Wu Q. Type II Transmembrane Serine Proteases. J Biol Chem. 2009;284:23177–23181. doi: 10.1074/jbc.R109.021006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [223].Szabo R, Bugge TH. Membrane-Anchored Serine Proteases in Vertebrate Cell and Developmental Biology. Annu Rev Cell Dev Biol. 2011;27:213–235. doi: 10.1146/annurev-cellbio-092910-154247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [224].Tanabe LM, List K. The role of type II transmembrane serine protease-mediated signaling in cancer. FEBS J. 2017;284:1421–1436. doi: 10.1111/febs.13971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [225].Wu Q, Parry G. Hepsin and prostate cancer. Front Biosci. 2007;12:5052–5059. doi: 10.2741/2447. [DOI] [PubMed] [Google Scholar]
- [226].Dong N, Zhou T, Zhang Y, Liu M, Li H, Huang X, Liu Z, Wu Y, Fukuda K, Qin J, Wu Q. Corin Mutations K317E and S472G from Preeclamptic Patients Alert Zymogen Activation and Cell Surface Targeting. J Biol Chem. 2014;289:17909–17916. doi: 10.1074/jbc.M114.551424. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [227].Tamberg T, Hong Z, De Schepper D, Skovbjerg S, Dupont DM, Vitved L, Schar CR, Skjoedt K, Vogel LK, Jensen JK. Blocking the proteolytic activity of zymogen matriptase with antibody-based inhibitors. J Biol Chem. 2019;294:314–326. doi: 10.1074/jbc.RA118.004126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [228].Ramsay AJ, Hooper JD, Folgueras AR, Velasco G, López-Otín C. Matriptase-2 (TMPRSS6): a proteolytic regulator of iron homeostasis. Haematologica. 2009;94:840–849. doi: 10.3324/haematol.2008.001867. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [229].Moncada-Pazos A, Cal S, López-Otín C. In: Handbook of Proteolytic Enzymes. Third Edition. Rawlings ND, Salvesen G, editors. Academic Press; London: 2013. Chapter 653 - Polyserases; pp. 2990–2994. [Google Scholar]
- [230].Kyrieleis OJP, Huber R, Ong E, Oehler R, Hunter M, Madison EL, Jacob U. Crystal structure of the catalytic domain of DESC1, a new member of the typeII transmembrane serine proteinase family. FEBS J. 2007;274:2148–2160. doi: 10.1111/j.1742-4658.2007.05756.x. [DOI] [PubMed] [Google Scholar]
- [231].Driesbaugh KH, Buzza MS, Martin EW, Conway GD, Kao JPY, Antalis TM. Proteolytic Activation of the Protease-activated Receptor (PAR)-2 by the Glycosylphosphatidylinositol-anchored Serine Protease Testisin. J Biol Chem. 2015;290:3529–3541. doi: 10.1074/jbc.M114.628560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [232].Chen L-M, Skinner ML, Kauffman SW, Chao J, Chao L, Thaler CD, Chai KX. Prostasin Is a Glycosylphosphatidylinositol-anchored Active Serine Protease. J Biol Chem. 2001;276:21434–21442. doi: 10.1074/jbc.M011423200. [DOI] [PubMed] [Google Scholar]
- [233].Eigenbrot C, Ganesan R, Kirchhofer D. Hepatocyte growth factor activator (HGFA): molecular structure and interactions with HGFA inhibitor-1 (HAI-1) FEBS J. 2010;277:2215–2222. doi: 10.1111/j.1742-4658.2010.07638.x. [DOI] [PubMed] [Google Scholar]
- [234].Pawar NR, Buzza MS, Antalis TM. Membrane-Anchored Serine Proteases and Protease-Activated Receptor-2–Mediated Signaling: Co-Conspirators in Cancer Progression. Cancer Res. 2019;79:301–310. doi: 10.1158/0008-5472.CAN-18-1745. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [235].Serrano SMT. The long road of research on snake venom serine proteinases. Toxicon. 2013;62:19–26. doi: 10.1016/j.toxicon.2012.09.003. [DOI] [PubMed] [Google Scholar]
- [236].Tasoulis T, Isbister GK. A Review and Database of Snake Venom Proteomes. Toxins. 2017;9:290. doi: 10.3390/toxins9090290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [237].Pirkle H. Thrombin-like enzymes from snake venoms: an updated inventory. Scientific and Standardization Committee’s Registry of Exogenous Hemostatic Factors. Thromb Haemost. 1998;79:675–683. [PubMed] [Google Scholar]
- [238].Paes Leme AF, Prezoto BC, Yamashiro ET, Bertholim L, Tashima AK, Klitzke CF, Camargo AC, Serrano SM. Bothrops protease A, a unique highly glycosylated serine proteinase, is a potent, specific fibrinogenolytic agent. J Thromb Haemost. 2008;6:1363–1372. doi: 10.1111/j.1538-7836.2008.02995.x. [DOI] [PubMed] [Google Scholar]
- [239].Yamashiro ET, Oliveira AK, Kitano ES, Menezes MC, Junqueira-de-Azevedo IL, Paes Leme AF, Serrano SMT. Proteoforms of the platelet-aggregating enzyme PA-BJ, a serine proteinase from Bothrops jararaca venom. Biochim Biophys Acta. 2014;1844:2068–2076. doi: 10.1016/j.bbapap.2014.09.012. [DOI] [PubMed] [Google Scholar]
- [240].Kita M, Nakamura Y, Okumura Y, Ohdachi SD, Oba Y, Yoshikuni M, Kido H, Uemura D. Blarina toxin, a mammalian lethal venom from the short-tailed shrew <em>Blarina brevicauda</em>: Isolation and characterization. Proc Natl Acad Sci USA. 2004;101:7542–7547. doi: 10.1073/pnas.0402517101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [241].Aminetzach YT, Srouji JR, Kong CY, Hoekstra HE. Convergent evolution of novel protein function in shrew and lizard venom. Curr Biol. 2009;19:1925–1931. doi: 10.1016/j.cub.2009.09.022. [DOI] [PubMed] [Google Scholar]
- [242].Miller JD. The Role of Dust Mites in Allergy. Clin Rev Allergy Immunol. 2018:1–18. doi: 10.1007/s12016-018-8693-0. [DOI] [PubMed] [Google Scholar]
- [243].Herman J, Thelen N, Smargiasso N, Mailleux A-C, Luxen A, Cloes M, De Pauw E, Chevigné A, Galleni M, Dumez M-E. Der p 1 is the primary activator of Der p 3, Der p 6 and Der p 9 the proteolytic allergens produced by the house dust mite Dermatophagoides pteronyssinus. Biochim Biophys Acta. 2014;1840:1117–1124. doi: 10.1016/j.bbagen.2013.11.017. [DOI] [PubMed] [Google Scholar]
- [244].Jacquet A, Campisi V, Szpakowska M, Dumez M-E, Galleni M, Chevigné A. Profiling the Extended Cleavage Specificity of the House Dust Mite Protease Allergens Der p 1, Der p 3 and Der p 6 for the Prediction of New Cell Surface Protein Substrates. Int J Mol Sci. 2017;18:E1373. doi: 10.3390/ijms18071373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [245].Takai T. In: Handbook of Proteolytic Enzymes. Third Edition. Rawlings ND, Salvesen G, editors. Academic Press; London: 2013. Chapter 671 - Serine Endopeptidase Allergens from Dermatophagoides Species; pp. 3055–3060. [Google Scholar]
- [246].Sun G, Stacey MA, Schmidt M, Mori L, Mattoli S. Interaction of Mite Allergens Der P3 and Der P9 with Protease-Activated Receptor-2 Expressed by Lung Epithelial Cells. J Immunol. 2001;167:1014–1021. doi: 10.4049/jimmunol.167.2.1014. [DOI] [PubMed] [Google Scholar]
- [247].Kale SL, Agrawal K, Gaur SN, Arora N. Cockroach protease allergen induces allergic airway inflammation via epithelial cell activation. Sci Rep. 2017;7:42341. doi: 10.1038/srep42341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [248].Polley DJ, Mihara K, Ramachandran R, Vliagoftis H, Renaux B, Saifeddine M, Daines MO, Boitano S, Hollenberg MD. Cockroach allergen serine proteinases: Isolation, sequencing and signalling via proteinase-activated receptor-2. Clin Exp Allergy. 2017;47:946–960. doi: 10.1111/cea.12921. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [249].Fuhrmann CN, Daugherty MD, Agard DA. Subangstrom Crystallography Reveals that Short Ionic Hydrogen Bonds, and Not a His-Asp Low-Barrier Hydrogen Bond, Stabilize the Transition State in Serine Protease Catalysis. J Am Chem Soc. 2006;128:9086–9102. doi: 10.1021/ja057721o. [DOI] [PubMed] [Google Scholar]
- [250].Ota N, Agard DA. Enzyme specificity under dynamic control II: Principal component analysis of α-lytic protease using global and local solvent boundary conditions. Protein Sci. 2001;10:1403–1414. doi: 10.1110/ps.800101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [251].Lee T-W, James MNG. 1.2Å-resolution crystal structures reveal the second tetrahedral intermediates of streptogrisin B (SGPB) Biochim Biophys Acta. 2008;1784:319–334. doi: 10.1016/j.bbapap.2007.11.012. [DOI] [PubMed] [Google Scholar]
- [252].Demidyuk IV, Chukhontseva KN, Kostrov SV. Glutamyl Endopeptidases: The Puzzle of Substrate Specificity. Acta Naturae. 2017;9:17–33. [PMC free article] [PubMed] [Google Scholar]
- [253].Prévost G. In: Handbook of Proteolytic Enzymes. Third Edition. Rawlings ND, Salvesen G, editors. Academic Press; London: 2013. Chapter 562 - Exfoliative Toxins – Staphylococcal Epidermolysins; pp. 2539–2543. [Google Scholar]
- [254].Spahich NA, St Geme JW., 3rd Structure and function of the Haemophilus influenzae autotransporters. Front Cell Infect Microbiol. 2011;1:1–9. doi: 10.3389/fcimb.2011.00005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [255].Yang L, Rezaie AR. Residues of the 39-Loop Restrict the Plasma Inhibitor Specificity of Factor IXa. J Biol Chem. 2013;288:12692–12698. doi: 10.1074/jbc.M113.459347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [256].Quinsey NS, Whisstock JC, Le Bonniec B, Louvain V, Bottomley SP, Pike RN. Molecular Determinants of the Mechanism Underlying Acceleration of the Interaction between Antithrombin and Factor Xa by Heparin Pentasaccharide. J Biol Chem. 2002;277:15971–15978. doi: 10.1074/jbc.M108131200. [DOI] [PubMed] [Google Scholar]
- [257].Kromann-Hansen T, Louise Lange E, Peter Sørensen H, Hassanzadeh-Ghassabeh G, Huang M, Jensen JK, Muyldermans S, Declerck PJ, Komives EA, Andreasen PA. Discovery of a novel conformational equilibrium in urokinase-type plasminogen activator. Sci Rep. 2017;7:3385. doi: 10.1038/s41598-017-03457-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [258].Sipos T, Merkel JR. Effect of calcium ions on the activity, heat stability, and structure of trypsin. Biochemistry. 1970;9:2766–2775. doi: 10.1021/bi00816a003. [DOI] [PubMed] [Google Scholar]
- [259].Rezaie AR, Esmon CT. Asp-70-->Lys mutant of factor X lacks high affinity Ca2+ binding site yet retains function. Journal of Biological Chemistry. 1994;269:21495–21499. [PubMed] [Google Scholar]
- [260].Ganesan R, Eigenbrot C, Wu Y, Liang W-C, Shia S, Lipari MT, Kirchhofer D. Unraveling the Allosteric Mechanism of Serine Protease Inhibition by an Antibody. Structure. 2009;17:1614–1624. doi: 10.1016/j.str.2009.09.014. [DOI] [PubMed] [Google Scholar]
- [261].Kellermann J, Lottspeich F, Geiger R, Deutzmann R. Human urinary kallikrein: amino acid sequence and carbohydrate attachment sites. Adv Exp Med Biol. 1989;247A:519–525. doi: 10.1007/978-1-4615-9543-4_80. [DOI] [PubMed] [Google Scholar]
- [262].Guo S, Skala W, Magdolen V, Brandstetter H, Goettig P. Sweetened kallikrein-related peptidases (KLKs): glycan trees as potential regulators of activation and activity. Biol Chem. 2014;395:959–976. doi: 10.1515/hsz-2014-0140. [DOI] [PubMed] [Google Scholar]
- [263].McGrath ME, Haymore BL, Summers NL, Craik CS, Fletterick RJ. Structure of an engineered, metal-actuated switch in trypsin. Biochemistry. 1993;32:1914–1919. doi: 10.1021/bi00059a005. [DOI] [PubMed] [Google Scholar]
- [264].Mac Sweeney A, Birrane G, Walsh MA, O’Connell T, Malthouse JPG, Higgins TM. Crystal structure of [delta]-chymotrypsin bound to a peptidyl chloromethyl ketone inhibitor. Acta Cryst D. 2000;56:280–286. doi: 10.1107/s0907444999016583. [DOI] [PubMed] [Google Scholar]
- [265].Pryzdial ELG, Kessler GE. Autoproteolysis or Plasmin-mediated Cleavage of Factor Xaα Exposes a Plasminogen Binding Site and Inhibits Coagulation. J Biol Chem. 1996;271:16614–16620. doi: 10.1074/jbc.271.28.16614. [DOI] [PubMed] [Google Scholar]
- [266].Sabharwal AK, Padmanabhan K, Tulinsky A, Mathur A, Gorka J, Bajaj SP. Interaction of Calcium with Native and Decarboxylated Human Factor X. Effect of proteolysis in the autolysis loop on catalytic efficiency and factor Va binding. J Biol Chem. 1997;272:22037–22045. doi: 10.1074/jbc.272.35.22037. [DOI] [PubMed] [Google Scholar]
- [267].Qureshi SH, Manithody C, Bae J-S, Yang L, Rezaie AR. Autolysis loop restricts the specificity of activated protein C: Analysis by FRET and functional assays. Biophys Chem. 2008;134:239–245. doi: 10.1016/j.bpc.2008.02.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [268].Pozzi N, Chen R, Chen Z, Bah A, Di Cera E. Rigidification of the autolysis loop enhances Na+ binding to thrombin. Biophys Chem. 2011;159:6–13. doi: 10.1016/j.bpc.2011.04.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [269].Reyda S, Sohn C, Klebe G, Rall K, Ullmann D, Jakubke H-D, Stubbs MT. Reconstructing the Binding Site of Factor Xa in Trypsin Reveals Ligand-induced Structural Plasticity. J Mol Biol. 2003;325:963–977. doi: 10.1016/s0022-2836(02)01337-2. [DOI] [PubMed] [Google Scholar]
- [270].Szabó E, Venekei I, Böcskei Z, Náray-Szabó G, Gráf L. Three Dimensional Structures of S189D Chymotrypsin and D189S Trypsin Mutants: The Effect of Polarity at Site 189 on a Proteasespecific Stabilization of the Substrate-binding Site. J Mol Biol. 2003;331:1121–1130. doi: 10.1016/s0022-2836(03)00849-0. [DOI] [PubMed] [Google Scholar]
- [271].Rezaie AR, Kittur FS. The Critical Role of the 185–189-Loop in the Factor Xa Interaction with Na+ and Factor Va in the Prothrombinase Complex. J Biol Chem. 2004;279:48262–48269. doi: 10.1074/jbc.M409964200. [DOI] [PubMed] [Google Scholar]
- [272].Bush-Pelc LA, Marino F, Chen Z, Pineda AO, Mathews FS, Di Cera E. Important Role of the Cys-191–Cys-220 Disulfide Bond in Thrombin Function and Allostery. J Biol Chem. 2007;282:27165–27170. doi: 10.1074/jbc.M703202200. [DOI] [PubMed] [Google Scholar]
- [273].Di Cera E. Thrombin: a paradigm for enzymes allosterically activated by monovalent cations. C R Biol. 2004;327:1065–1076. doi: 10.1016/j.crvi.2004.07.011. [DOI] [PubMed] [Google Scholar]
- [274].Gohara DW, Di Cera E. Molecular Mechanisms of Enzyme Activation by Monovalent Cations. J Biol Chem. 2016;291:20840–20848. doi: 10.1074/jbc.R116.737833. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [275].Huntington JA. How Na+ activates thrombin – a review of the functional and structural data. Biol Chem. 2008;389:1025–1035. doi: 10.1515/BC.2008.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [276].Dang QD, Di Cera E. Residue 225 determines the Na(+)-induced allosteric regulation of catalytic activity in serine proteases. Proc Natl Acad Sci USA. 1996;93:10653–10656. doi: 10.1073/pnas.93.20.10653. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [277].Vadivel K, Schreuder HA, Liesum A, Schmidt AE, Goldsmith G, Bajaj SP. Sodium-site in serine protease domain of human coagulation factor IXa: evidence from the crystal structure and molecular dynamics simulations study. J Thromb Haemost. 2019;17:574–584. doi: 10.1111/jth.14401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [278].Rezaie AR, He X. Sodium Binding Site of Factor Xa: Role of Sodium in the Prothrombinase Complex. Biochemistry. 2000;39:1817–1825. doi: 10.1021/bi992006a. [DOI] [PubMed] [Google Scholar]
- [279].Thorek DLJ, Ku AT, Mitsiades N, Veach D, Watson PA, Metha D, Strand S-E, Sharma SK, Lewis JS, Abou DS, Lilja HG, et al. Harnessing Androgen Receptor Pathway Activation for Targeted Alpha Particle Radioimmunotherapy of Breast Cancer. Clin Cancer Res. 2019;25:881–891. doi: 10.1158/1078-0432.CCR-18-1521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [280].De León-Rodríguez LM, Kovacs Z. The Synthesis and Chelation Chemistry of DOTA-Peptide Conjugates. Bioconjugate Chem. 2008;19:391–402. doi: 10.1021/bc700328s. [DOI] [PubMed] [Google Scholar]
- [281].Takayama Y, Kusamori K, Nishikawa M. Click Chemistry as a Tool for Cell Engineering and Drug Delivery. Molecules. 2019;24:E172. doi: 10.3390/molecules24010172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [282].Panos M, Christophi George P, Rodriguez M, Scarisbrick Isobel A. Differential expression of multiple kallikreins in a viral model of multiple sclerosis points to unique roles in the innate and adaptive immune response. Biol Chem. 2014:1063. doi: 10.1515/hsz-2014-0141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [283].Doak Bradley C, Over B, Giordanetto F, Kihlberg J. Oral Druggable Space beyond the Rule of 5: Insights from Drugs and Clinical Candidates. Chem Biol. 2014;21:1115–1142. doi: 10.1016/j.chembiol.2014.08.013. [DOI] [PubMed] [Google Scholar]
- [284].Kromann-Hansen T, Lange EL, Lund IK, Høyer-Hansen G, Andreasen PA, Komives EA. Ligand binding modulates the structural dynamics and activity of urokinase-type plasminogen activator: A possible mechanism of plasminogen activation. PLOS ONE. 2018;13:e0192661. doi: 10.1371/journal.pone.0192661. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [285].Craik Charles S, Page Michael J, Madison Edwin L. Proteases as therapeutics. Biochem J. 2011;435:1–16. doi: 10.1042/BJ20100965. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [286].Watters CM, Burton T, Kirui DK, Millenbaugh NJ. Enzymatic degradation of in vitro Staphylococcus aureus biofilms supplemented with human plasma. Infect Drug Resist. 2016;9:71–78. doi: 10.2147/IDR.S103101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [287].Das A, Datta S, Roche E, Chaffee S, Jose E, Shi L, Grover K, Khanna S, Sen CK, Roy S. Novel mechanisms of Collagenase Santyl Ointment (CSO) in wound macrophage polarization and resolution of wound inflammation. Sci Rep. 2018;8:1696. doi: 10.1038/s41598-018-19879-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [288].Freer ST, Kraut J, Robertus JD, Wright HT, Nguyen Huu X. Chymotrypsinogen: 2,5-Å crystal structure, comparison with α-chymotrypsin, and implications for zymogen activation. Biochemistry. 1970;9:1997–2009. doi: 10.1021/bi00811a022. [DOI] [PubMed] [Google Scholar]
- [289].Birktoft JJ, Blow DM. Structure of crystalline α-chymotrypsin: V. The atomic structure of tosyl-α-chymotrypsin at 2 Å resolution. J Mol Biol. 1972;68:187–240. doi: 10.1016/0022-2836(72)90210-0. [DOI] [PubMed] [Google Scholar]
- [290].Scheidig AJ, Hynes TR, Pelletier LA, Wells JA, Kossiakoff AA. Crystal structures of bovine chymotrypsin and trypsin complexed to the inhibitor domain of alzheimer’s amyloid β-protein precursor (APPI) and basic pancreatic trypsin inhibitor (BPTI): Engineering of inhibitors with altered specificities. Protein Sci. 1997;6:1806–1824. doi: 10.1002/pro.5560060902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [291].Gaboriaud C, Serre L, Guy-Crotte O, Forest E, Fontecilla-Camps J-C. Crystal Structure of Human Trypsin 1: Unexpected Phosphorylation of Tyr151. J Mol Biol. 1996;259:995–1010. doi: 10.1006/jmbi.1996.0376. [DOI] [PubMed] [Google Scholar]
- [292].Martin PD, Malkowski MG, DiMaio J, Konishi Y, Ni F, Edwards BFP. Bovine Thrombin Complexed with an Uncleavable Analog of Residues 7–19 of Fibrinogen Aα: Geometry of the Catalytic Triad and Interactions of the P1‘, P2‘, and P3‘ Substrate Residues. Biochemistry. 1996;35:13030–13039. doi: 10.1021/bi960656y. [DOI] [PubMed] [Google Scholar]
- [293].Friedrich R, Fuentes-Prior P, Ong E, Coombs G, Hunter M, Oehler R, Pierson D, Gonzalez R, Huber R, Bode W, Madison EL. Catalytic Domain Structures of MT-SP1/Matriptase, a Matrix-degrading Transmembrane Serine Proteinase. J Biol Chem. 2002;277:2160–2168. doi: 10.1074/jbc.M109830200. [DOI] [PubMed] [Google Scholar]
- [294].Bateman KS, Huang K, Anderson S, Lu W, Qasim MA, Laskowski M, James MNG. Contribution of peptide bonds to inhibitor-protease binding: crystal structures of the turkey ovomucoid third domain backbone variants OMTKY3-Pro18I and OMTKY3-Ψ[COO]-Leu18I in complex with Streptomyces griseus proteinase B (SGPB) and the structure of the free inhibitor, OMTKY3-Ψ[CH2NH2+]-Asp19I1 1. Rees D, editor. J Mol Biol. 2001;305:839–849. doi: 10.1006/jmbi.2000.4343. [DOI] [PubMed] [Google Scholar]