Abstract
Macroautophagy is characterized by the de novo formation of double-membrane vesicles termed autophagosomes. The precursor structure of autophagosomes is a membrane cistern called phagophore, which elongates through a massive acquisition of lipids until closure. The phagophore establishes membrane-contact sites (MCSs) with the endoplasmic reticulum (ER), where conserved ATG proteins belonging to the ATG9 lipid scramblase, ATG2 lipid transfer and Atg18/WIPI4 β-propeller families concentrate. Several recent in vivo and in vitro studies have uncovered the relevance of these proteins and MCSs in the lipid supply required for autophagosome formation. Although important conceptual advances have been reached, the functional interrelationship between ATG9, ATG2 and Atg18/WIPI4 proteins at the phagophore-ER MCSs and their role in the phagophore expansion are not completely understood. In this review, we describe the current knowledge about the structure, interactions, localizations, and molecular functions of these proteins, with a particular emphasis on the yeast Saccharomyces cerevisiae and mammalian systems.
Keywords: phagophore, autophagosome, autophagy, ATG protein, lipid transfer, tethering, scramblase, ATG2, ATG9, Atg18/WIPI4
Introduction
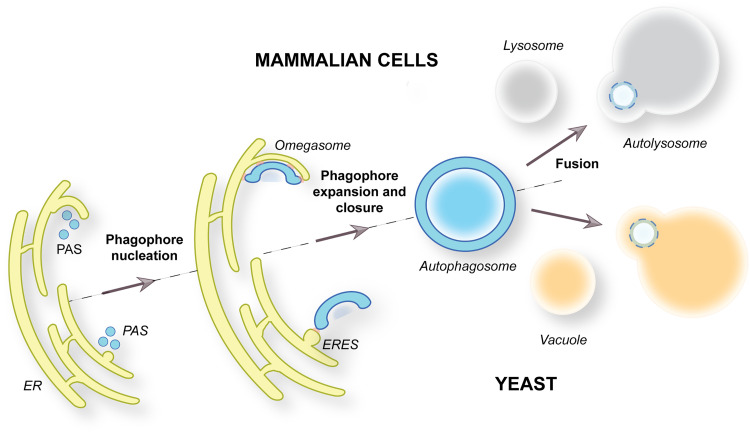
Macroautophagy, hereafter autophagy, is an evolutionary conserved catabolic pathway that allows eukaryotic cells to catabolize long-lived and unwanted proteins or protein complexes, but also excess or dysfunctional organelles as well as invading pathogens. The delivery of these cargoes to lysosomes/vacuoles for degradation is mediated by selective or non-selective autophagy (Nakatogawa, 2020; Gomez-Sanchez et al., 2021). The hallmark of autophagy is the de novo formation of double-membrane vesicles called autophagosomes, which sequester the material targeted for degradation (Nakatogawa, 2020; Gomez-Sanchez et al., 2021) (Figure 1). Autophagosomes biogenesis is mediated by a set of approximately twenty highly conserved proteins known as the autophagy-related (ATG) proteins, which are organized in six functional groups: the Atg1/ULK kinase complex, the Atg9/ATG9A-positive vesicles, the autophagy-specific phosphatidylinositol 3-kinase (PI3K) complex I, the Atg2/ATG2 proteins-Atg18/WIPI4 complex, the Atg12/ATG12 and the Atg8/LC3 conjugation systems (Nakatogawa, 2020; Gomez-Sanchez et al., 2021). Autophagosome formation is initiated by the orchestrated assembly of the ATG proteins and the concomitant nucleation of a membranous cistern known as phagophore or isolation membrane (Suzuki et al., 2007; Nakatogawa, 2020; Gomez-Sanchez et al., 2021) (Figure 1). The biogenesis of this structure occurs at specific locations within cells known as the phagophore assembly site (PAS) in yeast or omegasomes in mammalian cells, which are both in close proximity to the endoplasmic reticulum (ER) (Nakatogawa, 2020; Gomez-Sanchez et al., 2021; Zheng et al., 2022) (Figure 1). The phagophore subsequently expands, sequestering the targeted cargoes, and seals it into an autophagosome. Then autophagosomes fuse with vacuoles in yeast or with late endosomes and/or lysosomes in mammalian cells, in which the inner vesicle of autophagosomes and their cargo are degraded into basic metabolites by resident hydrolases (Figure 1). These metabolites are eventually transported into the cytoplasm and used as either a source of energy or building blocks for the biosynthesis of new macromolecules (Lahiri et al., 2019).
Figure 1.
Overview of the Autophagy Pathway. Autophagy is characterized by the de novo formation of double-membrane vesicles called autophagosomes. This process starts with the nucleation of the phagophore, which is probably characterized by the heterotypic fusion of vesicles. The generation of phagophores occurs at the PAS, which is located in close proximity of both the ER and vacuole in yeast (bottom panel), or within omegasomes, a specialized subdomain of the ER, in mammalian cells (upper panel). The phagophores forms MCSs with the ER-exit sites (ERES) in yeast or directly with the ER at the omegasomes in mammals. The phagophore elongation culminates when its extremities fuse together through fission, leading to the formation of an autophagosome. Complete autophagosomes then fuse with vacuoles in yeast and late endosomes/lysosomes in mammalian cells. The autophagosomal cargoes are degraded in the interior of these lytic compartment by resident hydrolases.
In eukaryotic cells, intracellular organelles are in constant exchange and communication with each other to synchronize cellular functions and metabolism (Jain and Holthuis, 2017; Bohnert, 2020; Prinz et al., 2020). In addition to vesicular trafficking, this exchange also takes place along membrane contact sites (MCSs) between organelles, which are specialized platforms in which organelle membranes are positioned in close proximity and anchored to each other by tethers factors. The principal function of MCSs is the transfer of lipids, but they can also mediate calcium signalling, cargo sorting and fission/fusion processes. Since lipid supply is critical for formation and expansion of the phagophore, this cisterna possesses MCSs with several organelles, including the ER, lipid droplets, mitochondria and additionally in yeast the vacuole (Bieber et al., 2022; Zwilling and Reggiori, 2022; Li et al., 2023). The MCSs between the phagophore and the ER, the main lipid synthesis factory in eukaryotic cells, appears to be particularly important for the generation of autophagosomes. In this review, we will focus on the proteins localized at these MCSs, more specifically on Atg9/ATG9A, Atg2/ATG2A/ATG2B and Atg18/WIPI4, and their function during phagophore expansion. The emphasis of this piece will be on the yeast Saccharomyces cerevisiae and mammalian systems.
ATG9 Scramblases and Their Role in Autophagy Initiation and Progression
Structure and Molecular Function
Atg9/ATG9A is the only transmembrane protein within the core ATG machinery (Noda et al., 2000; Young et al., 2006; Ungermann and Reggiori, 2018). The amino acid length of ATG9 proteins varies between 700 and 1000 residues among species (Figure 2A) and amino acid sequence alignments have shown that the common conserved region in these proteins is the central presence of approximately 500 residues distributed over six predicted intramembrane segments (Noda et al., 2000; Young et al., 2006; Lai et al., 2020; Maeda et al., 2020; Chumpen Ramirez et al., 2022). The two cytoplasmic termini, in contrast, are apparently divergent (Figure 2A), at least in sequence length, highlighting possible organism-specific differences in the regulation and function of these proteins, possibly also outside the context of autophagy. The overall Atg9/ATG9A structure consists as homotrimer with C3 symmetry, in which each protomer harbours four transmembrane domains and two helices that are partially inserted in the membrane, which match the six predicted intramembrane segments (Figure 2A and B) (Guardia et al., 2020; Maeda et al., 2020; Matoba et al., 2020). Additionally, each protomer possesses cytoplasmic helices which connect the transmembrane domain of the protein (Figure 2A and C) (Guardia et al., 2020; Matoba et al., 2020). The Atg9/ATG9A trimers are organized as an extensive domain-swapping in which a unit of two helices from one protomer crosses over and heaps parallelly with the next protomer. This conformation generates a planar surface radially projected from the centre of the trimer (Figure 2D) (Guardia et al., 2020; Maeda et al., 2020; Matoba et al., 2020). The N and C termini from the first and second partially membrane-embedded helices, respectively, contain highly conserved proline residues forming a kinked unit linking the planar surface and creating an elbow-like shape (Guardia et al., 2020; Maeda et al., 2020; Matoba et al., 2020). Transmembrane segments from neighbouring protomers are enriched in phenylalanine residues and they might both mediate protein-protein interactions and promote conformational changes (Guardia et al., 2020; Chumpen Ramirez et al., 2022). In fact, cryo-EM analyses have described two conformations of the ATG9A trimers, known as status A and B, which may represent two functional rearrangements (Guardia et al., 2020; Maeda et al., 2020; Matoba et al., 2020).
Figure 2.
Structural characteristics of Atg9 and ATG9A. (A) Topological representation of yeast Atg9 and mammalian ATG9A. Atg9 and ATG9A possess four transmembrane domains and two segments partially embedded into the membrane (brown cylinders), which are interconnected by loops (black lines) that differ in amino acid length between the two species. The N- and C-termini of the two proteins, which are coloured in green and red, respectively, are oriented toward the cytosol and they also differ in amino acid length quite remarkably between species. (B) AlphaFold (Jumper et al., 2021; Varadi et al., 2022) prediction of Atg9 and ATG9A protomers. The transmembrane domains of each monomer are coloured in brown, while the cytoplasmic and luminal parts connecting them are in black. N and C termini are coloured as in A. (C) Lateral view of the Atg9/ATG9A trimers. The sequences of the Atg9 and ATG9A were obtained from the UniProt database (https://www.uniprot.org/) and used as template. The modelling of the trimeric complex was performed using the online SWISS-MODEL software (https://swissmodel.expasy.org/interactive) and the resulting structure was coloured using the ChimeraX software (https://www.cgl.ucsf.edu/chimera/) (Pettersen et al., 2004). The structures cover the sequences between amino acids 296 and 783, and between 36 and 352 of Atg9 and ATG9A, respectively. The transmembrane domains of each protomer are highlighted in brown, yellow and orange, while the cytoplasmic domains are in different blue colours. On the right of ATG9A, the interaction between the transmembrane domains of adjacent monomers is highlighted through enlargement. (D) Top view of the homotrimeric structure of Atg9 and ATG9A, obtained by rotating 90 degrees the models presented in Panel C. Lateral pores (LP) are pointed out with dashed red lines, while the central pore (CP) is highlighted in the middle of both structures.
Lipid scramblases are proteins that can flip lipids bidirectionally between the leaflets of the membranes in an energy-independent manner (Daleke, 2003). Several studies have revealed that ATG9 proteins possess phospholipid scrambling activity in vitro (Maeda et al., 2020; Matoba et al., 2020; Ghanbarpour et al., 2021; Chumpen Ramirez et al., 2022). These results are based on a fluorescence-based lipid scramblase assay (Maeda et al., 2020; Chumpen Ramirez et al., 2022) or the quick-freezing and freeze-fracture replica labelling technique to measure phosphatidylinositol-3-phosphate (PtdIns3P) distribution between the inner and outer leaflets of the limiting membrane of liposomes (Matoba et al., 2020). The quick-freezing and freeze-fracture replica technique was subsequently used to also show that newly synthesized phospholipids such as phosphatidylcholine and phosphatidylserine are incorporated into nascent autophagosomes and symmetrically distributed between the two leaflets of the phagophore membrane in an Atg9-dependent manner (Matoba et al., 2020; Orii et al., 2021). The structural analyses have revealed that each protomer harbours a lateral pore (LP)/cavity that flows into the central pore (CP) and connects the exposed entrance to the cytosol with the cytosolic region of the bilayer (Figure 2) (Guardia et al., 2020; Maeda et al., 2020; Matoba et al., 2020). This leads to the formation of an internal network of branched cavities within the trimer. Mutations in amino acids located in the LP and CP have showed that these residues compromise the scramblase activity of ATG9 proteins and their involvement in autophagy (Maeda et al., 2020; Matoba et al., 2020). A possible explanation might be that these point mutations affect the lipid binding within the cavities of the trimer, interfering with ATG9 protein function (Maeda et al., 2020).
Interestingly, a recent study has identified a highly conserved phenylalanine residue at the interface between neighbouring protomers of ATG9 proteins (Chumpen Ramirez et al., 2022). Point mutations of this specific amino acid in both Atg9 and ATG9A block autophagy. However, trimerization as well as the interaction between Atg9 and the Atg2-Atg18 complex and the cellular localization of Atg9 are not affected by this mutation, respectively (Chumpen Ramirez et al., 2022). Surprisingly, this Atg9 point mutant has identical lipid scrambling activity as the wild-type protein, suggesting that the interaction between the transmembrane segments of the protomers provides the trimer with an additional function (Chumpen Ramirez et al., 2022). This additional function remains to be established, but it has been speculated that it might be required for the formation of Atg9 oligomers via the association of multiple Atg9 trimers, forming a proteinaceous subdomain (Chumpen Ramirez et al., 2022).
Localization and Role in Autophagosome Biogenesis
The first ATG9 protein to be identified was yeast Atg9 and its characterization revealed that this protein is distributed into punctate structures dispersed throughout the cytoplasm, with one of them corresponding to the PAS (Noda et al., 2000). Subsequent fluorescence and electron microscopy approaches show that the cytoplasmic Atg9-positive puncta are a tubovesicular compartment, often adjacent to mitochondria, which does not colocalize with other endomembrane marker proteins (Mari et al., 2010; Yamamoto et al., 2012). The induction of autophagosome biogenesis leads to the relocalization of at least one of the Atg9-positive puncta to the site where the PAS is formed (Mari et al., 2010; Yamamoto et al., 2012), in an Atg11 or Atg17-dependent manner depending on the nutritional conditions (He et al., 2006; Sekito et al., 2009). This potential role of the Atg9-positive structures for being the seed membrane for the nucleation of the phagophore has recently been confirmed with in vitro experiments (Sawa-Makarska et al., 2020). At the PAS, the Atg9-positive vesicles are probably participating in the nucleation of the phagophore through homo- or heterotypic fusion with other vesicles in a process that may involve the Atg1 kinase complex and/or SNARE proteins (Nair et al., 2011; Rao et al., 2016; Matscheko et al., 2019). Moreover, Atg9 may also act as an organizational hub for the Atg machinery not only by interacting with the components of the Atg1 kinase complex (Suzuki et al., 2015; Rao et al., 2016; Matscheko et al., 2019), but also by recruiting the PI3K complex I (Suzuki et al., 2015). In this regard, Atg9 localizes to the extremities of the phagophore (Graef et al., 2013; Suzuki et al., 2013), possibly because ATG9 proteins appear to be prone to localize to membranes with high curvature (Guardia et al., 2020), where it promotes the recruitment and assembly of the Atg2-Atg18 complex, thus participating in the establishment of the phagophore-ER MCSs (Gomez-Sanchez et al., 2018).
Similar to the yeast protein, mammalian ATG9A is concentrated in tubovesicular clusters that reside in post-Golgi compartments by continuously cycling between the trans-Golgi network and the endosomal system via vesicular transport (Young et al., 2006; Orsi et al., 2012; Imai et al., 2016). Upon autophagosome formation induction, ATG9A partially redistributes to recycling endosomes and then relocalizes adjacently to the ER, where it has been suggested that it promotes the phagophore nucleation (Longatti et al., 2012; Orsi et al., 2012; Soreng et al., 2018; Mailler et al., 2021). Interestingly, a new study has shown that although small ATG9A-positive vesicles are undetectable by fluorescence microscopy, these structures are required for the generation of phagophores confirming their crucial role as platform for autophagosome biogenesis (Broadbent et al., 2023). Moreover, ATG9A can also be found in ER-rich membranes positive for PAS marker proteins (Olivas et al., 2023). How ATG9A participates in the orchestration of the ATG machinery in mammalian cells, however, remains largely unclear. Nonetheless, it has been confirmed that ATG9A binds to ATG2A (Guardia et al., 2020; Ghanbarpour et al., 2021; Mailler et al., 2021; van Vliet et al., 2022).
The role of ATG9 proteins in phagophore expansion is not exclusively connected to their role in localizing and organizing the Atg2/ATG2-Atg18/WIPI4 complex at the phagophore-ER MCSs (see below). In fact, the lipid scrambling activity of ATG9 proteins is also essential for the phagophore expansion (Maeda et al., 2020; Matoba et al., 2020). That is, expression of Atg9 and ATG9A variants displaying a lipid scrambling activity defect in vitro in knockout cells, leads to block of autophagy progression and accumulation of smaller phagophores in yeast and mammalian cells, respectively (Maeda et al., 2020; Matoba et al., 2020).
ATG2 Proteins as Tethers and Lipid Transfer Proteins
Structure and Molecular Function
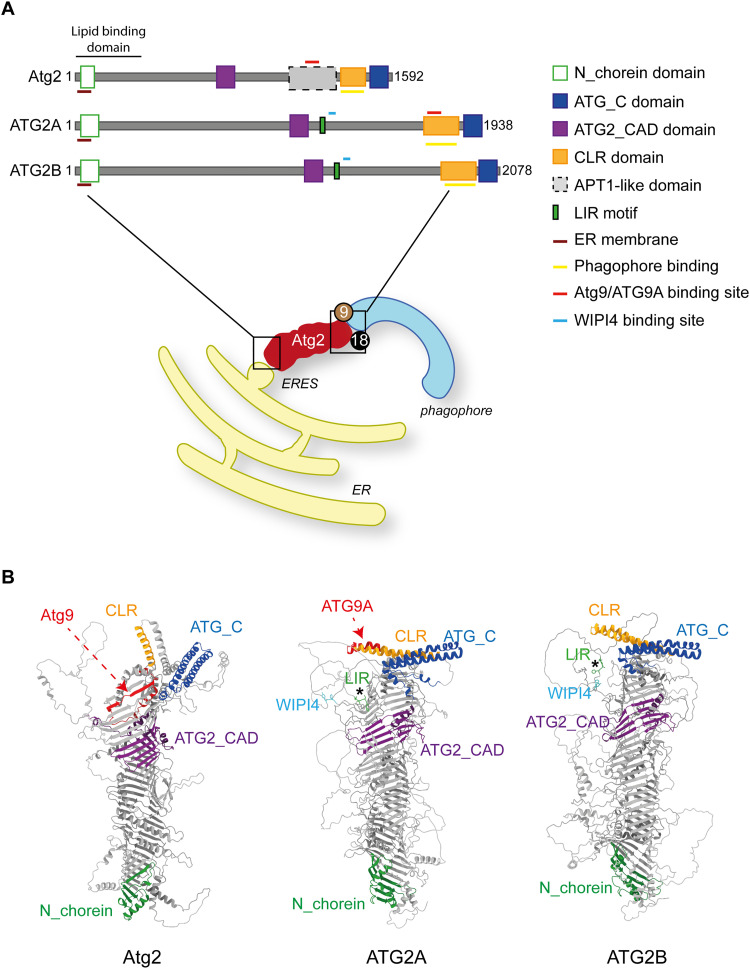
Yeast Atg2 and its mammalian homologs, ATG2A and ATG2B, are one of the key factors in the phagophore expansion and autophagosome formation, and thus they are essential for autophagy (Shintani et al., 2001; Wang et al., 2001; Velikkakath et al., 2012). ATG2A and ATG2B have redundant function since the depletion of the individual genes is not affecting autophagy progression (Velikkakath et al., 2012). ATG2 proteins are the largest ATG core proteins, with amino acid sequences between 1600 and 2300 residues that can be longer in some species such as Capsaspora owczarzaki (i.e., 3299 amino acids), which are organized in a rod-like structure with a length of approximately 20 nm (Chowdhury et al., 2018; Maeda et al., 2019; Osawa et al., 2019; Osawa and Noda, 2019; Valverde et al., 2019; van Vliet et al., 2022) (Figure 3). The Pfam database assigns evolutionary conserved domains such as the N_chorein, the ATG2 CAD (Atg2_CAD), and the ATG C terminal (ATG_C) domain, which are present at either the N- or the C-terminus of ATG2 proteins (Chowdhury et al., 2018) (Figure 3A). Although it has not been registered in the Pfam database, ATG2 proteins also contain a conserved short sequence at the C-terminal localization region (CLR), which consists to an amphipathic α-helixes allowing the membrane association (Kotani et al., 2018; Maeda et al., 2019) (Figure 3A). Moreover, an APT1 domain is found between the ATG_2_CAD and ATG2_C domain of the yeast protein (Chowdhury et al., 2018) (Figure 3A). Mammalian ATG2A is 44.5% identical in amino acid sequence to ATG2B, whereas yeast Atg2 shows 17% and 21% identity to ATG2A and ATG2B, respectively (Velikkakath et al., 2012; Bozic et al., 2020). ATG2A and ATG2B also possesses a highly conserved LC3-interacting region (LIR) between at positions 1362–1365 and 1491–1494, respectively (Figure 3A). LIRs consist in a short sequence of four amino acids with a consensus sequence of [E/D/S/T]-W/F/Y-X1-X2-L/I/V (in which X1/X2 can be any amino acid) and are involved in the interactions with members of the ATG8 protein family (Birgisdottir et al., 2013; Bozic et al., 2020). The ATG2A and ATG2B LIR mediates the interaction of these proteins with the members of the GABARAP subfamily, which are part of the mammalian ATG8 protein family (Bozic et al., 2020). As GABARAP proteins (Weidberg et al., 2010; Nguyen et al., 2016), ATG2 proteins also appear to function at the latest stages in the autophagosome formation in a LIR-dependent manner. In particular, ATG2 variants unable to bind GABARAP proteins do not sustain phagophore formation and/or autophagosome closure under starvation conditions (Bozic et al., 2020). It remains to be established whether Atg2 possess a functional LIR.
Figure 3.
Structural Characteristics of Atg2 and its ATG2A/ATG2B Homologs. (A) The domain organization of ATG2 protein consists of a N_Chorein domain located at their N-terminus (green rectangle), in which a Glycerophospholipid binding domain resides. In the middle, Atg2/ATG2 proteins contain a ATG2_CAD domain (purple rectangle), while the CLR region and the ATG_C domain (orange and blue rectangle, respectively) are located at the C-terminus. The membrane binding site responsible of the attachment with the phagophore is within the CLR domain and represented by a yellow line. In contrast to the mammalian ATG2 proteins, Atg2 possesses an APT1-like domain (grey), which is flanking the ATG2_CAD and CLR Motifs. The mammalian ATG2 proteins have one conserved LIR that is represented by a green rectangle in the diagram. As highlighted in the drawing, the N-terminus of the Atg2/ATG2 protein is required for their binding to the ER/ERES while their C-terminus is responsible for the localization at the end of the phagophore and interaction with the Atg9/ATG9A and Atg18/WIPI4 proteins. (B) The structures of Atg2, ATG2A and ATG2B predicted by AlphaFold and coloured based on the diagram in panel A using the ChimeraX Software. The location and name of the different domains is indicated across the structures. The conserved LIR domains in ATG2A and ATG2B are highlighted by asterisks and the amino acids are coloured in green. The Putative WIPI4 binding site in ATG2A and ATG2B is underlined by colouring the amino acids in light blue. The binding region of Atg9 in Atg2 and that of ATG9A in ATG2A are pointed with red dash lines and coloured in the same colour in the structures.
The overall structure of ATG2 proteins consists in a long groove containing hydrophobic residues, which allows the accommodation and transfer of lipids (Figure 3B) (Melia and Reinisch, 2022; Neuman et al., 2022; Dabrowski et al., 2023). Structural studies on the C-terminal regions have been difficult due to the flexibility of this part of the ATG2 proteins. Thus, most of the investigations have focused on their N-terminus. The N-termini are found at one of the extremities of the rod and possess a N_chorein domain (Figure 3), an evolutionary conserved region present also in other lipid transfer proteins such as the members of the VPS13 protein family, yeast Csf1 and mammalian KIAA0100 (Levine, 2022). Structural analysis of the N-terminal region of Schizosaccaromyces pombe Atg2 revealed the presence of five α-helices and a twisted β-sheet harbouring 6 β-strands (Figure 3B) (Osawa et al., 2019). This conformation forms a hydrophobic cavity in which phospholipid acyl chains are accommodated. This arrangement is consistent with a structural hallmark of the other lipid transport proteins, including VPS13 proteins, which are also located in specific MCSs (Osawa et al., 2019; Levine, 2022; Melia and Reinisch, 2022). Like VPS13 and KIAA100, ATG2 proteins also possess a repeating β-groove (RBG) domain, which consists of a repeated module that harbours five β-sheets followed by a loop, mostly composed of a single helix and a short disordered polypeptide chain (Figure 3B) (Neuman et al., 2022). These similarities have prompted studies on the possible function of Atg2/ATG2 proteins as lipid transfer proteins (see below).
The membrane binding sites of Atg2/ATG2 proteins appear to be located at both ends of the rod-like structure. While the N- terminus associates with the ER, the C-terminal parts are required for the binding to the phagophore (see below). In agreement with this notion, purified Atg2/ATG2 proteins can tether membranes with high curvature and large lipid-packing defects, in a PtdIns3P-independent manner (Chowdhury et al., 2018; Kotani et al., 2018; Osawa et al., 2019). However, the presence of Atg18/WIPI4 proteins facilitates the tethering activity of ATG2 proteins and provides directionality towards PtdIns3P-containing liposomes. That is, while the N-termini bind to the liposomes that do not contain PtdIns3P, Atg18/WIPI4 and the C-termini of the Atg2/ATG2 proteins associate with the PtdIns3P-positive membranes. The amphipathic helix located at the C-terminus of Atg2/ATG2 proteins might favour the binding of Atg18/WIPI4 to PtdIns3P (Chowdhury et al., 2018; Kotani et al., 2018). Nevertheless, there is a discrepancy regarding the role of the C-terminal parts of the Atg2/ATG2 proteins. While this domain is necessary for the tethering activity of Atg2, it is dispensable for the one of ATG2A (Maeda et al., 2019).
Several in vitro studies have demonstrated that Atg2/ATG2 proteins transfer lipids between adjacent liposomes (Tamura et al., 2017; Chowdhury et al., 2018; Maeda et al., 2019; Osawa et al., 2019; Osawa and Noda, 2019; Valverde et al., 2019; Chumpen Ramirez et al., 2022). Although the C-termini are involved in Atg2/ATG2 association with membranes, this region of ATG2A takes part in the lipid transfer activity moderately in comparison with the N-terminus (Maeda et al., 2019; van Vliet et al., 2022). Consistently, the key lipid transfer modules reside at the N-terminal of the Atg2/ATG2 proteins, between amino acids 7-228 in Atg2, amino acids 1-345 in ATG2A and amino acids 1-387 in ATG2B (Figure 3) (Valverde et al., 2019; Nishimura and Tooze, 2020). A couple of studies have reported that this domain is sufficient to rescue almost completely the autophagy defect of atg2Δ and atg2a−/− atg2b−/− cells, indicating that the lipid transfer activity of Atg2/ATG2 proteins is important for autophagy in vivo (Osawa and Noda, 2019; Valverde et al., 2019). Interestingly, experiments using a Vps13-Atg2 chimera also rescued the autophagy block of the atg2Δ knockout strain, suggesting the existence of specific and conserved mechanism in extracting phospholipids from donor membranes between lipid transfer proteins from different protein families (Osawa et al., 2019). Along this line, a recent study has suggested that Vps13 is located at the rim of phagophore and may cooperate with Atg2 in generating the phagophore-ER contact site and driving autophagosomes expansion (Dabrowski et al., 2023).
Localization and Role in Autophagosome Formation
The majority of ATG proteins, including Atg2/ATG2 proteins, are cytosolic. However, the initial characterization of Atg2/ATG2 proteins already revealed that they are peripherally associated with membranes at the site of autophagosome formation (Shintani et al., 2001; Suzuki et al., 2001; Wang et al., 2001; Kim et al., 2002; Velikkakath et al., 2012). More specifically, yeast Atg2 localizes in the extremities of the phagophore, where this cisterna forms MCSs with the ER (Graef et al., 2013; Suzuki et al., 2013; Gomez-Sanchez et al., 2018). Similarly, overexpressed GFP-ATG2A also localizes to the rim of the phagophore (Sakai et al., 2020) and the autophagosomal membrane-ER MCSs (Tang et al., 2019; Valverde et al., 2019). Atg2 association to the phagophore extremities requires coincidence binding to Atg9 and PtdIns3P (Gomez-Sanchez et al., 2018). While it is still unclear how Atg2 binds to PtdIns3P, although it has been suggested that it is via its APT1 domain (Kolakowski et al., 2020), an amino acid sequence present at amino acids 1234–1268 appears to be essential for the Atg2–Atg9 interaction (Gomez-Sanchez et al., 2018). A recent study in mammalian cells has revealed the existence of two sequences mediating the interaction between ATG2A and the ATG9A trimers in vitro, which were named ATG2A S1 and ATG2A S2 (van Vliet et al., 2022). The Atg9 binding site identified previously in yeast shares homology with ATG2A S1, but this site is not crucial for the interaction between ATG2A and ATG9A proteins in vivo (Gomez-Sanchez et al., 2018; van Vliet et al., 2022). This finding suggests that the interaction mechanisms between Atg2/ATG2 and Atg9/ATG9A proteins might differ between species. Indeed, deletion of the ATG2A S2, which is localized in the CLR domain between amino acid positions 1760 and 1779, shows a reduction in ATG2A–ATG9A binding in vivo and autophagy progression in vivo (Velikkakath et al., 2012; van Vliet et al., 2022). Moreover, an ATG2A variant lacking this domain displays a reduction in tethering activity in an ATG9A-independent manner, which could explain the observed impairment in both lipid transfer in vitro and autophagic flux in vivo (van Vliet et al., 2022). Deletion of CLR domain in Atg2 also leads to a reduction in its interaction with Atg9, suggesting that this binding site may be evolutionary conserved (Kotani et al., 2018; van Vliet et al., 2022).
The CLR domain of ATG2 proteins is also essential for autophagy (van Vliet et al., 2022). Although ATG2A S2 localizes to this domain, it is difficult to conclude whether the ATG2A variant lacking the CLR domain does not sustain autophagy because of either its inability to interact with ATG9A via this domain or another function of this region (van Vliet et al., 2022). The CLR domain of ATG2A, which contains an amphipathic helix, is required for the association of this protein with autophagosomal membranes in vivo (Velikkakath et al., 2012; Tamura et al., 2017). However, it seems that ATG2A can bind to ATG9A in the absence of membrane binding because the variant lacking the CLR domain still interacts with ATG9A, possibly via ATG2A S1 or another domain (van Vliet et al., 2022). Together, these observations suggest that both the ATG9A and the membrane-binding sites located in the ATG2A_CLR domain are required for the function of the ATG2A-ATG9A complex in autophagy.
The chorein-N domain of ATG2A is also important for its recruitment to membranes (Tamura et al., 2017; Kotani et al., 2018). This domain localizes to the N-terminal extremities of rod-like ATG2 proteins, which is required to bind the donor liposomes in the in vitro tethering assays (Kotani et al., 2018) and consistently, it is involved in Atg2 association with the ER/ER-exit sites (ERES) in vivo (Kotani et al., 2018). ATG2A is also found associated with lipid droplets (Velikkakath et al., 2012; Pfisterer et al., 2014). Interestingly, this localization depends on the chorein-N and CLR domains, but also on a motif at amino acids 1723–1829 that is highly conserved in ATG2B at residues 1863–1982 and appears to be also present among other species (Velikkakath et al., 2012; Tamura et al., 2017). Although this motif from ATG2B shows binding affinity to lipid droplets (Velikkakath et al., 2012), there is no evidence that ATG2B localizes to this organelle. Nonetheless, accumulation and enlargement of lipid droplets in cells in which ATG2A and ATG2B are depleted simultaneously suggest that those proteins also directly or indirectly function in lipid droplets morphology and dispersion (Velikkakath et al., 2012).
Atg18/WIPI4 as Regulators of ATG2 Proteins
Structure and Molecular Function
Yeast Atg18 and its paralogues Atg21 and Hsv2 belong to the β-propellers that bind polyphosphoinositides (PROPPINs) protein family, and have four mammalian homologues: WIPI1, WIPI2, WIPI3 and WIPI4 (Proikas-Cezanne et al., 2015; Lei et al., 2021). The hallmarks of this family are the β-propeller conformation and the presence of that conserved FRRG/LRRG motif that allow binding to PtdIns3P and PtdIns3,5P2 (Figure 4A) (Dove et al., 2004; Krick et al., 2006; Thumm et al., 2013; Proikas-Cezanne et al., 2015; Lei et al., 2021). Thus, Atg18/WIPI proteins are considered as conserved PtdIns3P effectors that, except for Hsv2, play an important role in autophagy (Dove et al., 2004; Watanabe et al., 2012; Thumm et al., 2013). The seven β-propeller blades are formed by four antiparallel β-strands (Figure 4B) (Watanabe et al., 2012; Thumm et al., 2013). Each blade is connected by an amino acid loop and contains WD40 repeats, which are characteristic of these proteins, and this is why they are also known as WD-repeat protein interacting with phosphoinositides (WIPI) (Proikas-Cezanne et al., 2004). WD40 repeats are composed of approximately 40 amino acids often ending with a dipeptide formed by Trp and Asp, and frequently fold into β-propellers (Li and Roberts, 2001). β-propellers are seen as scaffold proteins providing a platform for protein-protein interactions and protein complex assembly (Paoli, 2001).
Figure 4.
Structural Characteristics of Atg18 and WIPI4. (A) The domain organization of Atg18 and WIPI4 is characterized by seven β-propeller blades connected by loops, highlighted with different colours. These proteins function as PtdIns3P effectors and the conserved FRRG/LRRG PtdIns3P-binding motif is indicated. A recent study has shown the presence of the unique 7AB loop (light green) in Atg18. Moreover, the 6CD loop (light blue) is longer in Atg18 than WIPI4. (B) AlphaFold prediction of Atg18 and WIPI4 structures analysed using ChimeraX software are presented. Each blade and loop were coloured based on the domain diagram shown in Panel A. The binding sites for Atg2 and ATG2A/ATG2B are marked by a red arc line in Atg18 and WIPI4, respectively. Similarly, the FRRG and LRRG motif of Atg18 and WIPI4 are highlighted with a black arc line and a dashed arrow, respectively. The 6CD (light blue) 7AB (light green) loops are also indicated.
PROPPINs are also subunits of large molecular complexes (Baskaran et al., 2012a; Lei et al., 2021). The high-resolution structure for Atg18 is still unsolved, hindering the full understanding the function of the Atg2-Atg18 and Atg9-Atg2-Atg18 complexes (Lei et al., 2021) (see below). The predicted structure of Atg18 and WIPI proteins has been initially deducted by solving the structure of Hsv2 from Kluyveromyces lactis (Krick et al., 2006; Baskaran et al., 2012a; Baskaran et al., 2012b; Watanabe et al., 2012). Like Hsv2, the overall structure of Atg18 and WIPI proteins displays a highly conserved discoidal shape (Figure 4B). The structural investigations have confirmed the biochemical data (Krick et al., 2006) and showed that the FRRG/LRRG motif, which is localized to the loop 5, is indeed the phosphoinositide-binding site in Atg18 and WIPI proteins (Krick et al., 2006; Baskaran et al., 2012a; Lei et al., 2021) (Figure 4). The existence of a second phosphoinositide-binding site at the blade 6 of Atg18 has been postulated based on the structure of Hsv2 (Baskaran et al., 2012a). Mutational analysis has confirmed that Atg18 interacts with PtdIns3P also via this second binding site (Watanabe et al., 2012). Since blades 5 and 6 are highly conserved, the dual PtdIns3P binding mode is very likely conserved among all the WIPI proteins as well (Proikas-Cezanne et al., 2015).
Although the seven blades are well structured, bioinformatics and structural analysis have highlighted that Atg18 contains two long loops connecting the blades, i.e., the 6CD and the 7AB loop (Figure 4) (Lei et al., 2021). Alignment of the amino acid sequences has revealed that the 7AB loop (position 433–460) is not present in Atg21, Hsv2 and the WIPI proteins, suggesting a functional importance of this region (Lei et al., 2021). Indeed, it was shown that the 7AB loop is required for autophagy and may harbour a new binding site for Atg2 (Lei et al., 2021). Consistently, the 7AB loop appears to modulate Atg2 localization to the PAS (Lei et al., 2021). However, two different other studies reported that the Atg2 binding site in Atg18 is located at the blade and loop 2, and mutation of these sites affects autophagy progression (Watanabe et al., 2012; Rieter et al., 2013). This apparent discrepancy could reflect a dynamicity in the Atg2-Atg18 interaction. Although it appears that all the WIPI proteins can interact to different extend at least with ATG2B in vitro (Zheng et al., 2017), WIPI4 is clearly a binding partner of ATG2A and ATG2B also in vivo (Behrends et al., 2010; Zheng et al., 2017; Chowdhury et al., 2018; Maeda et al., 2019). In contrast, it remains to be established whether the interaction between ATG2A/ATG2B and the rest of the WIPI proteins occurs in vivo.
More specifically, WIPI4 interacts via its loop 3 with the conserved aromatic His/Phe/Tyr motif in ATG2A and ATG2B (Zheng et al., 2017; Chowdhury et al., 2018; Lei et al., 2021). Since the 7AB loop is absent in the WIPI proteins, this might point to a difference in the Atg2-Atg18 and ATG2A/ATG2B-WIPI4 interaction (Lei et al., 2021). However, it has been shown that three sites within blades 1 and 3 of WIPI3 recognize a conserved motif in ATG2 proteins that consists of β-strand-(X)n-Φ-X-Φ-X-X-X-Ψ-F, in which Φ and Ψ are an hydrophobic and aromatic residue, respectively (Ren et al., 2020). This interaction resembles the one described for Atg2 and Atg18 in yeast. Thus, although essential for autophagy, how ATG2 and Atg18/WIPI4 proteins interact remains to be understood.
Atg18 and WIPI4 have not an evident enzymatic activity. β-propellers are often a domain within a protein that as mentioned, is involved in protein–protein interactions and complex formation (Paoli, 2001). As a result, one hypothesis is that the regulated recruitment of Atg18 and WIPI4 onto PtdIns3P-positive autophagosomal membranes and interaction with ATG2 proteins, promote the assembly of a complex and/or a structural rearrangement that regulate the function of ATG2 proteins. In this context, structural studies in the methylotrophic yeast Pichia angusta have revealed that Atg18 rapidly forms oligomers upon membrane binding while it remains as a monomer in solution (Scacioc et al., 2017). This oligomerization, which is promoted by membrane curvature (Scacioc et al., 2017), might create a local microdomain that could be important to locally regulate the function and assembly of other factors. Interestingly, in vitro experiments have revealed that the presence of Atg18 and WIPI4 assists both the tethering and the lipid transfer activity of Atg2 and ATG2A in a PtdIns3P dependent manner (Otomo et al., 2018; Osawa and Noda, 2019; Chumpen Ramirez et al., 2022). Another postulated scenario is that Atg18, possibly together with Atg9, might define the direction of phospholipids that are transferred by Atg2 (Osawa and Noda, 2019). Despite all this information, the precise molecular function of Atg18 and WIPI4 is still a mystery.
Localization and Role in Autophagosome Formation
Atg18 is a peripheral membrane-associate protein that localizes to the PAS, but also endosomes and vacuoles (Barth et al., 2001; Krick et al., 2006; Obara et al., 2008). Like WIPI proteins, Atg18 is recruited onto autophagic membranes upon the induction of autophagy and this redistribution depends on its ability to bind PtdIns3P (Barth et al., 2001; Krick et al., 2006; Efe et al., 2007; Obara et al., 2008; Proikas-Cezanne et al., 2015). Consistently with its localization, Atg18 is involved in autophagy, endosomal trafficking, and regulation of vacuolar morphology (Barth et al., 2001; Courtellemont et al., 2022; Marquardt et al., 2023). Atg18 functions as a subunit of the PtdIns3P 5-kinase Fab1 complex to regulate vacuolar morphology at the vacuole via binding to PtdIns3,5P2 (Watanabe et al., 2012), whereas it interacts with the Vps5-Vps17-Vps35 retromer complex for endosomal traffic (Courtellemont et al., 2022; Marquardt et al., 2023). Atg18 is also essential for autophagy (Barth et al., 2001; Obara et al., 2008) and forms a complex with Atg2 (Obara et al., 2008; Watanabe et al., 2012; Rieter et al., 2013). Where and how this complex is assembled is controversial. On the one hand, it has been proposed that the Atg2-Atg18 complex is formed constitutively and recruited en bloc to the PAS in a PtdIns3P-dependent manner (Obara et al., 2008). On the other hand, it has been shown that Atg18 is recruited to the PAS by coincidence binding to PtdIns3P and Atg2 (Rieter et al., 2013), and that Atg18 interaction with Atg2 requires Atg2–Atg9 binding (Gomez-Sanchez et al., 2018). Nonetheless, Atg18 binding to Atg2 is essential for its function in autophagy (Rieter et al., 2013) and together with Atg2 and Atg9, it localizes to the extremities of the phagophore where it is part of the phagophore-ER MCSs (Graef et al., 2013; Suzuki et al., 2013; Gomez-Sanchez et al., 2018).
Atg18 could have a possible regulatory role in lipid transfer at the phagophore-ER MCSs, since phagophores fail to expand into an autophagosome in the absence of ATG18 (Gomez-Sanchez et al., 2018; Chumpen Ramirez et al., 2022). Additionally, Atg18 is involved in Atg9 retrieval from autophagosomal membranes (Reggiori et al., 2004), although this could be linked to its function together with the retromer complex (Courtellemont et al., 2022; Marquardt et al., 2023). Since deletion of ATG2 leads to the same Atg9 trafficking defect (Reggiori et al., 2004), however, it may be that the Atg9 retrograde transport from autophagosomal intermediates is a downstream event following phagophore expansion and/or closure.
All WIPI proteins function as PtdIns3P effectors in autophagy (Thumm et al., 2013; Proikas-Cezanne et al., 2015). While WIPI1 and WIPI2 have a role in the early stages of autophagosome biogenesis, in which WIPI2 coordinates the recruitment of the two ubiquitin-like conjugation systems (Proikas-Cezanne et al., 2004; Proikas-Cezanne et al., 2007; Itakura and Mizushima, 2010; Polson et al., 2010; Dooley et al., 2014; Strong et al., 2021), WIPI3 and WIPI4 act in later stages since their simultaneous knockdown leads to the accumulation of expanded phagophores (Bakula et al., 2017; Ji et al., 2021). Depletion of WIPI4 leads to a modest defect in autophagy (Bakula et al., 2017; Ji et al., 2021), indicating that it could have a redundant role with WIPI3 or eventually another factor.
WIPI4 is a stronger binding partner of ATG2A and ATG2B than WIPI1 or WIPI2 (Zheng et al., 2017). Alignments of multiple amino acid sequences have underlined that the residues in loop 2 of Atg18 are conserved in WIPI4, while the ones reported by Watanabe and co-workers are not (Rieter et al., 2013; Zheng et al., 2017). However, mutations in the loop 2 of WIPI4 do not alter its interaction with ATG2B; suggesting that ATG2B binding site is located elsewhere in WIPI4. Indeed, changes in loop 3 abolish WIPI4 binding to ATG2B (Zheng et al., 2017). Thus, additional studies are required to identify how WIPI4 interact with mammalian ATG2 proteins.
Conclusions and Perspectives
The current notion is that ATG9, ATG2 and Atg18/WIPI4 proteins play an essential role during the phagophore expansion until autophagosome completion by both establishing the phagophore-ER MCSs and regulating lipid transport from the ER to the phagophore (Figure 5). Although these three proteins distinctly localize to the phagophore-ER MCSs in yeast (Graef et al., 2013; Suzuki et al., 2013; Gomez-Sanchez et al., 2018), it remains to be shown whether they have a similar distribution in mammalian cells. In this context, ATG9A appears to associate with nascent autophagosomes only transitionally (Orsi et al., 2012; Olivas et al., 2023) and consequently it may have a less prominent role than its yeast counterpart as an organizational hub.
Figure 5.
Putative Molecular Function of ATG9, ATG2 and Atg18/WIPI4 Proteins at the Phagophore-ER MCSs. The ER is considered as a key lipids source during phagophore expansion. The current model is that ATG2 proteins localize to the extremities of the phagophore together with ATG9 proteins and Atg18/WIPI4. Atg18/WIPI4 and possibly ATG2 proteins bind to PtdIns3P (orange lipids). ATG2 proteins also bind the ER/ERES through a still unclear mechanism (question mark), and thus they have a key role in establishing the phagophore-ER MCSs. The oval structures on the ER represent the putative ATG2 protein binding partners, which experimental evidence has indicated to be ERES components although which one(s) of them is directly interacting with the ATG2 proteins remains to be determined. Although it remains unknown what is providing the force for the flow of lipids, those are transferred from the ER to the phagophore by ATG2 proteins (black arrow), a process that also requires the presence of Atg18/WIPI4 at the phagophore-ER MCSs. Lipids from ATG2 proteins may or may not be delivered to ATG9 scramblases, which translocate bidirectionally the lipids across the phagophore membrane bilayer, maintaining the symmetric distribution of lipids and ultimately allowing phagophore expansion.
In vitro data clearly show that ATG2 proteins are central for membrane tethering and lipid transfer activities (Figure 5) (Chowdhury et al., 2018; Kotani et al., 2018; Otomo et al., 2018; Maeda et al., 2019; Osawa et al., 2019; Valverde et al., 2019; Chumpen Ramirez et al., 2022). Additionally, a recent study has demonstrated that ATG2A mediates membrane tethering in vivo using the dual-color particle tracking technique (Broadbent et al., 2023). Lipids are very likely transferred from ER to the cytosolic leaflet of the phagophore membrane creating a lipid asymmetry and possibly a tension that could hamper phagophore elongation. Therefore, it is not surprising that ATG2 proteins would need the cooperation of lipid scramblases, like ATG9 proteins, to dissipate the lipid asymmetry that they create, by translocating lipids from the cytosolic leaflet to the inner one of the phagophore membrane and thus promoting its expansion (Figure 5) (Maeda et al., 2020; Matoba et al., 2020; Melia and Reinisch, 2022; Zhang et al., 2022). Since ATG2 proteins probably also generate a lipid asymmetry in the ER membrane because the lipid extraction from the outer leaflet of the ER limiting membrane, it is not unexpected that ER-localized lipid scramblases such as mammalian VMP1 and TMEM41B are also crucial for autophagy (Ghanbarpour et al., 2021; McEwan and Ryan, 2022). This idea is supported by in vitro observations showing that insertion of Atg9 in the acceptor or donor liposomes enhances the lipid transfer activity of Atg2 (Chumpen Ramirez et al., 2022). The force for lipids to enter and pass through the ATG2 proteins could be provided by the biosynthesis of lipids on the cytoplasmic leaflet of the ER. Within such mechanism, however, ER-localized scramblases would lower the tension between the ER membrane leaflets, blocking lipid transfer by ATG2 proteins. Thus, additional investigations are necessary to unveil how lipid are transferred at the phagophore-ER MCSs and the role of ER-localized scramblases in this.
The function of Atg18 and WIPI4 is more enigmatic, but the in vitro data revealed that the tethering and lipid transfer activity of ATG2 proteins is enhanced by these proteins, respectively (Otomo et al., 2018; Osawa and Noda, 2019; Chumpen Ramirez et al., 2022). A simple hypothesis is that Atg18 and WIPI4 are regulators that proof-read whether ATG2 proteins and eventually ATG9 scramblases are localized and/or assembled at the right place before starting lipid transfer (Figure 5). The ability of Atg18 to coincidentally associated with Atg9-bound Atg2 and PtdIns3P could allow this protein to distinguish the phagophore membrane from other ones (Figure 5) (Rieter et al., 2013; Gomez-Sanchez et al., 2018). However, other scenarios are also possible.
Although enormous advances on the molecular role of ATG9, ATG2 and Atg18/WIPI4 proteins in autophagosome formation have been achieved in recent years, numerous questions still wait for an answer. For instance, it is unknown how ATG2 proteins associate to the ER, even if studies in yeast have shown the interaction with a specialized ERES may mediate this anchoring (Graef et al., 2013; Suzuki et al., 2013). Moreover, it is still unclear whether the establishment of the phagophore-ER MCSs and lipid transfer involved additional players. In this respect, it is totally unknown which kind of force makes it possible that a massive amount of lipids rapidly passes from the ER to the phagophore, although lipid biosynthesis may play a role in this (Nishimura et al., 2017; Schutter et al., 2020). Understanding these aspects will not only be important to decipher the mechanism of autophagy, but it would also provide molecular insights into lipid droplet biogenesis, since both ATG2A and ATG9A have a role in lipid mobilization/size regulation of this organelle (Velikkakath et al., 2012; Mailler et al., 2021). Additionally, addressing these questions could help to uncover the mechanism of lysosomal repair, in which ATG2 proteins appear to have an important function (Tan and Finkel, 2022). Overall ATG9, ATG2 and/or Atg18/WIPI4 proteins may represent a modular system that eukaryotic cells may use to establish MCSs between specific organelles for lipid transfer. The relevance of this system is also underlined by the fact that mutations in WIPI4 cause ß-propeller protein associated neurodegeneration (BPAN), a lethal neurodevelopmental disorder (Cong et al., 2021; Shimizu et al., 2023). Henceforth, the elucidation of the molecular function of the phagophore-ER MCSs could aid to understand the generation of autophagosomes and possibly other organelles, in health and disease.
Footnotes
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding: The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Schweizerischer Nationalfonds zur Förderung der Wissenschaftlichen Forschung, H2020 Marie Skłodowska-Curie Actions, Novo Nordisk Fonden, ENW, (grant number CRSII5_189952, 754513, 0066384, OCENW.KLEIN.118).
ORCID iD: Fulvio Reggiori https://orcid.org/0000-0003-2652-2686
References
- Bakula D, Muller AJ, Zuleger T, Takacs Z, Franz-Wachtel M, Thost AK, Brigger D, Tschan MP, Frickey T, Robenek H, … (2017). WIPI3 And WIPI4 beta-propellers are scaffolds for LKB1-AMPK-TSC signalling circuits in the control of autophagy. Nature Communications 8, 15637. doi: 10.1038/ncomms15637 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barth H, Meiling-Wesse K, Epple UD, Thumm M. (2001). Autophagy and the cytoplasm to vacuole targeting pathway both require Aut10p. FEBS Letters 508, 23–28. doi: 10.1016/s0014-5793(01)03016-2 [DOI] [PubMed] [Google Scholar]
- Baskaran S, Ragusa MJ, Boura E, Hurley JH. (2012a). Two-site recognition of phosphatidylinositol 3-phosphate by PROPPINs in autophagy. Molecular Cell 47, 339–348. doi: 10.1016/j.molcel.2012.05.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baskaran S, Ragusa MJ, Hurley JH. (2012b). How Atg18 and the WIPIs sense phosphatidylinositol 3-phosphate. Autophagy 8, 1851–1852. doi: 10.4161/auto.22077. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Behrends C, Sowa ME, Gygi SP, Harper JW. (2010). Network organization of the human autophagy system. Nature 466, 68–76. doi: 10.1038/nature09204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bieber A, Capitanio C, Erdmann PS, Fiedler F, Beck F, Lee CW, Li D, Hummer G, Schulman BA, Baumeister W, Wilfling F. (2022). In situ structural analysis reveals membrane shape transitions during autophagosome formation. Proceedings of the National Academy of Sciences of the United States of America 119, e2209823119. doi: 10.1073/pnas.2209823119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Birgisdottir AB, Lamark T, Johansen T. (2013). The LIR motif - crucial for selective autophagy. Journal of Cell Science 126, 3237–3247. doi: 10.1242/jcs.126128. [DOI] [PubMed] [Google Scholar]
- Bohnert M. (2020). Tether me, tether me not-dynamic organelle contact sites in metabolic rewiring. Developmental Cell 54, 212–225. doi: 10.1016/j.devcel.2020.06.026. [DOI] [PubMed] [Google Scholar]
- Bozic M, van den Bekerom L, Milne BA, Goodman N, Roberston L, Prescott AR, Macartney TJ, Dawe N, McEwan DG. (2020). A conserved ATG2-GABARAP family interaction is critical for phagophore formation. EMBO Reports 21, e48412. doi: 10.15252/embr.201948412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Broadbent DG, Barnaba C, Perez GI, Schmidt JC. (2023). Quantitative analysis of autophagy reveals the role of ATG9 and ATG2 in autophagosome formation. Journal of Cell Biology 222. doi: 10.1083/jcb.202210078. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chowdhury S, Otomo C, Leitner A, Ohashi K, Aebersold R, Lander GC, Otomo T. (2018). Insights into autophagosome biogenesis from structural and biochemical analyses of the ATG2A-WIPI4 complex. Proceedings of the National Academy of Sciences of the United States of America 115, E9792–E9801. doi: 10.1073/pnas.1811874115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chumpen Ramirez S, Gomez-Sanchez R, Verlhac P, Hardenberg R, Margheritis E, Cosentino K, Reggiori F, Ungermann C. (2022). Atg9 interactions via its transmembrane domains are required for phagophore expansion during autophagy. Autophagy, 1–20. doi: 10.1080/15548627.2022.2136340 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cong Y, So V, Tijssen MAJ, Verbeek DS, Reggiori F, Mauthe M. (2021). WDR45, One gene associated with multiple neurodevelopmental disorders. Autophagy 17, 3908–3923. doi: 10.1080/15548627.2021.1899669. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Courtellemont T, De Leo MG, Gopaldass N, Mayer A. (2022). CROP: A retromer-PROPPIN complex mediating membrane fission in the endo-lysosomal system. EMBO Journal 41, e109646. doi: 10.15252/embj.2021109646. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dabrowski R, Tulli S, Graef M. (2023). Parallel phospholipid transfer by Vps13 and Atg2 determines autophagosome biogenesis dynamics. Journal of Cell Biology 222. doi: 10.1083/jcb.202211039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Daleke DL. (2003). Regulation of transbilayer plasma membrane phospholipid asymmetry. Journal of Lipid Research 44, 233–242. doi: 10.1194/jlr.R200019-JLR200. [DOI] [PubMed] [Google Scholar]
- Dooley HC, Razi M, Polson HE, Girardin SE, Wilson MI, Tooze SA. (2014). WIPI2 Links LC3 conjugation with PI3P, autophagosome formation, and pathogen clearance by recruiting Atg12-5-16L1. Molecular Cell 55, 238–252. doi: 10.1016/j.molcel.2014.05.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dove SK, Piper RC, McEwen RK, Yu JW, King MC, Hughes DC, Thuring J, Holmes AB, Cooke FT, Michell RH, … (2004). Svp1p defines a family of phosphatidylinositol 3,5-bisphosphate effectors. EMBO Journal 23, 1922–1933. doi: 10.1038/sj.emboj.7600203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Efe JA, Botelho RJ, Emr SD. (2007). Atg18 regulates organelle morphology and Fab1 kinase activity independent of its membrane recruitment by phosphatidylinositol 3,5-bisphosphate. Molecular Biology of the Cell 18, 4232–4244. doi: 10.1091/mbc.e07-04-0301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ghanbarpour A, Valverde DP, Melia TJ, Reinisch KM. (2021). A model for a partnership of lipid transfer proteins and scramblases in membrane expansion and organelle biogenesis. Proceedings of the National Academy of Sciences of the United States of America 118. doi: 10.1073/pnas.2101562118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gomez-Sanchez R, Rose J, Guimaraes R, Mari M, Papinski D, Rieter E, Geerts WJ, Hardenberg R, Kraft C, Ungermann C, Reggiori F. (2018). Atg9 establishes Atg2-dependent contact sites between the endoplasmic reticulum and phagophores. J. Cell Biol 217, 2743–2763. doi: 10.1083/jcb.201710116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gomez-Sanchez R, Tooze SA, Reggiori F. (2021). Membrane supply and remodeling during autophagosome biogenesis. Current Opinion in Cell Biology 71, 112–119. doi: 10.1016/j.ceb.2021.02.001. [DOI] [PubMed] [Google Scholar]
- Graef M, Friedman JR, Graham C, Babu M, Nunnari J. (2013). ER Exit sites are physical and functional core autophagosome biogenesis components. Molecular Biology of the Cell 24, 2918–2931. doi: 10.1091/mbc.E13-07-0381. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guardia CM, Tan XF, Lian T, Rana MS, Zhou W, Christenson ET, Lowry AJ, Faraldo-Gomez JD, Bonifacino JS, Jiang J, Banerjee A. (2020). Structure of human ATG9A, the only transmembrane protein of the core autophagy machinery. Cell Reports 31, 107837. doi: 10.1016/j.celrep.2020.107837. [DOI] [PMC free article] [PubMed] [Google Scholar]
- He C, Song H, Yorimitsu T, Monastyrska I, Yen WL, Legakis JE, Klionsky DJ. (2006). Recruitment of Atg9 to the preautophagosomal structure by Atg11 is essential for selective autophagy in budding yeast. Journal of Cell Biology 175, 925–935. doi: 10.1083/jcb.200606084. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Imai K, Hao F, Fujita N, Tsuji Y, Oe Y, Araki Y, Hamasaki M, Noda T, Yoshimori T. (2016). Atg9A trafficking through the recycling endosomes is required for autophagosome formation. Journal of Cell Science 129, 3781–3791. doi: 10.1242/jcs.196196. [DOI] [PubMed] [Google Scholar]
- Itakura E, Mizushima N. (2010). Characterization of autophagosome formation site by a hierarchical analysis of mammalian atg proteins. Autophagy 6, 764–776. doi: 10.4161/auto.6.6.12709. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jain A, Holthuis JCM. (2017). Membrane contact sites, ancient and central hubs of cellular lipid logistics. Biochimica et Biophysica Acta - Molecular Cell Research 1864, 1450–1458. doi: 10.1016/j.bbamcr.2017.05.017. [DOI] [PubMed] [Google Scholar]
- Ji C, Zhao H, Chen D, Zhang H,, & Zhao YG. (2021). beta-propeller proteins WDR45 and WDR45B regulate autophagosome maturation into autolysosomes in neural cells. Current Biology 31, 1666–1677e1666. doi: 10.1016/j.cub.2021.01.081 [DOI] [PubMed] [Google Scholar]
- Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, Tunyasuvunakool K, Bates R, Zidek A, Potapenko A, … (2021). Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589. doi: 10.1038/s41586-021-03819-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim J, Huang WP, Stromhaug PE, Klionsky DJ. (2002). Convergence of multiple autophagy and cytoplasm to vacuole targeting components to a perivacuolar membrane compartment prior to de novo vesicle formation. Journal of Biological Chemistry 277, 763–773. doi: 10.1074/jbc.M109134200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kolakowski D, Kaminska J, Zoladek T. (2020). The binding of the APT1 domains to phosphoinositides is regulated by metal ions in vitro. Biochimica et Biophysica Acta Biomembranes 1862, 183349. doi: 10.1016/j.bbamem.2020.183349. [DOI] [PubMed] [Google Scholar]
- Kotani T, Kirisako H, Koizumi M, Ohsumi Y, Nakatogawa H. (2018). The Atg2-Atg18 complex tethers pre-autophagosomal membranes to the endoplasmic reticulum for autophagosome formation. Proceedings of the National Academy of Sciences of the United States of America 115, 10363–10368. doi: 10.1073/pnas.1806727115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krick R, Tolstrup J, Appelles A, Henke S, Thumm M. (2006). The relevance of the phosphatidylinositolphosphat-binding motif FRRGT of Atg18 and Atg21 for the cvt pathway and autophagy. FEBS Letters 580, 4632–4638. doi: 10.1016/j.febslet.2006.07.041. [DOI] [PubMed] [Google Scholar]
- Lahiri V, Hawkins WD, Klionsky DJ. (2019). Watch what you (self-) eat: Autophagic mechanisms that modulate metabolism. Cell Metabolism 29, 803–826. doi: 10.1016/j.cmet.2019.03.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lai LTF, Yu C, Wong JSK, Lo HS, Benlekbir S, Jiang L, Lau WCY. (2020). Subnanometer resolution cryo-EM structure of Arabidopsis thaliana ATG9. Autophagy 16, 575–583. doi: 10.1080/15548627.2019.1639300. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lei Y, Tang D, Liao G, Xu L, Liu S, Chen Q, Li C, Duan J, Wang K, Wang J, … (2021). The crystal structure of Atg18 reveals a new binding site for Atg2 in Saccharomyces cerevisiae. Cellular and Molecular Life Sciences 78, 2131–2143. doi: 10.1007/s00018-020-03621-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Levine TP. (2022). Sequence analysis and structural predictions of lipid transfer bridges in the repeating Beta groove (RBG) superfamily reveal past and present domain variations affecting form, function and interactions of VPS13, ATG2, SHIP164, hobbit and tweek. Contact (Thousand Oaks) 5, 251525642211343. doi: 10.1177/25152564221134328. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li D, Roberts R. (2001). Human genome and diseases: WD-repeat proteins: Structure characteristics, biological function, and their involvement in human diseases. CMLS 58, 2085–2097. doi: 10.1007/PL00000838. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li M, Tripathi-Giesgen I, Schulman BA, Baumeister W,, & Wilfling F. (2023). In situ snapshots along a mammalian selective autophagy pathway. Proceedings of the National Academy of Sciences of the United States of America 120, e2221712120. 10.1073/pnas.2221712120 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Longatti A, Lamb CA, Razi M, Yoshimura S, Barr FA, Tooze SA. (2012). TBC1D14 Regulates autophagosome formation via Rab11- and ULK1-positive recycling endosomes. Journal of Cell Biology 197, 659–675. doi: 10.1083/jcb.201111079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maeda S, Otomo C, Otomo T. (2019). The autophagic membrane tether ATG2A transfers lipids between membranes. Elife 8. doi: 10.7554/eLife.45777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maeda S, Yamamoto H, Kinch LN, Garza CM, Takahashi S, Otomo C, Grishin NV, Forli S, Mizushima N, Otomo T. (2020). Structure, lipid scrambling activity and role in autophagosome formation of ATG9A. Nature Structural & Molecular Biology 27, 1194–1201. doi: 10.1038/s41594-020-00520-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mailler E, Guardia CM, Bai X, Jarnik M, Williamson CD, Li Y, Maio N, Golden A, Bonifacino JS. (2021). The autophagy protein ATG9A enables lipid mobilization from lipid droplets. Nature Communications 12, 6750. doi: 10.1038/s41467-021-26999-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mari M, Griffith J, Rieter E, Krishnappa L, Klionsky DJ, Reggiori F. (2010). An Atg9-containing compartment that functions in the early steps of autophagosome biogenesis. Journal of Cell Biology 190, 1005–1022. doi: 10.1083/jcb.200912089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marquardt L, Taylor M, Kramer F, Schmitt K, Braus GH, Valerius O, Thumm M. (2023). Vacuole fragmentation depends on a novel Atg18-containing retromer-complex. Autophagy 19, 278–295. doi: 10.1080/15548627.2022.2072656. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matoba K, Kotani T, Tsutsumi A, Tsuji T, Mori T, Noshiro D, Sugita Y, Nomura N, Iwata S, Ohsumi Y, … (2020). Atg9 is a lipid scramblase that mediates autophagosomal membrane expansion. Nature Structural & Molecular Biology 27, 1185–1193. doi: 10.1038/s41594-020-00518-w. [DOI] [PubMed] [Google Scholar]
- Matscheko N, Mayrhofer P, Rao Y, Beier V, Wollert T. (2019). Atg11 tethers Atg9 vesicles to initiate selective autophagy. PLoS Biology 17, e3000377. doi: 10.1371/journal.pbio.3000377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McEwan DG, Ryan KM. (2022). ATG2 and VPS13 proteins: Molecular highways transporting lipids to drive membrane expansion and organelle communication. FEBS Journal 289, 7113–7127. doi: 10.1111/febs.16280. [DOI] [PubMed] [Google Scholar]
- Melia TJ, Reinisch KM. (2022). A possible role for VPS13-family proteins in bulk lipid transfer, membrane expansion and organelle biogenesis. Journal of Cell Science 135. doi: 10.1242/jcs.259357. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nair U, Jotwani A, Geng J, Gammoh N, Richerson D, Yen WL, Griffith J, Nag S, Wang K, Moss T, … (2011). SNARE Proteins are required for macroautophagy. Cell 146, 290–302. doi: 10.1016/j.cell.2011.06.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakatogawa H. (2020). Mechanisms governing autophagosome biogenesis. Nature Reviews Molecular Cell Biology 21, 439–458. doi: 10.1038/s41580-020-0241-0. [DOI] [PubMed] [Google Scholar]
- Neuman SD, Levine TP, Bashirullah A. (2022). A novel superfamily of bridge-like lipid transfer proteins. Trends in Cell Biology 32, 962–974. doi: 10.1016/j.tcb.2022.03.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nguyen TN, Padman BS, Usher J, Oorschot V, Ramm G, Lazarou M. (2016). Atg8 family LC3/GABARAP proteins are crucial for autophagosome-lysosome fusion but not autophagosome formation during PINK1/parkin mitophagy and starvation. Journal of Cell Biology 215, 857–874. doi: 10.1083/jcb.201607039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nishimura T, Tamura N, Kono N, Shimanaka Y, Arai H, Yamamoto H, Mizushima N. (2017). Autophagosome formation is initiated at phosphatidylinositol synthase-enriched ER subdomains. EMBO Journal 36, 1719–1735. doi: 10.15252/embj.201695189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nishimura T, Tooze SA. (2020). Emerging roles of ATG proteins and membrane lipids in autophagosome formation. Cell Discovery 6, 32. doi: 10.1038/s41421-020-0161-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Noda T, Kim J, Huang W-P, Baba M, Tokunaga C, Ohsumi Y, Klionsky DJ. (2000). Apg9p/Cvt7p is an integral membrane protein required for transport vesicle formation in the CVT and autophagy pathways. Journal of Cell Biology 148, 465–480. doi: 10.1083/jcb.148.3.465. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Obara K, Sekito T, Niimi K, Ohsumi Y. (2008). The Atg18-Atg2 complex is recruited to autophagic membranes via phosphatidylinositol 3-phosphate and exerts an essential function. Journal of Biological Chemistry 283, 23972–23980. doi: 10.1074/jbc.M803180200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olivas TJ, Wu Y, Yu S, Luan L, Choi P, Guinn ED, Nag S, De Camilli PV, Gupta K, Melia TJ. (2023). ATG9 vesicles comprise the seed membrane of mammalian autophagosomes. Journal of Cell Biology 222. doi: 10.1083/jcb.202208088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Orii M, Tsuji T, Ogasawara Y, Fujimoto T. (2021). Transmembrane phospholipid translocation mediated by Atg9 is involved in autophagosome formation. Journal of Cell Biology 220. doi: 10.1083/jcb.202009194. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Orsi A, Razi M, Dooley HC, Robinson D, Weston AE, Collinson LM, Tooze SA. (2012). Dynamic and transient interactions of Atg9 with autophagosomes, but not membrane integration, are required for autophagy. Molecular Biology of the Cell 23, 1860–1873. doi: 10.1091/mbc.E11-09-0746. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Osawa T, Kotani T, Kawaoka T, Hirata E, Suzuki K, Nakatogawa H, Ohsumi Y, Noda NN. (2019). Atg2 mediates direct lipid transfer between membranes for autophagosome formation. Nature Structural & Molecular Biology 26, 281–288. doi: 10.1038/s41594-019-0203-4. [DOI] [PubMed] [Google Scholar]
- Osawa T, Noda NN. (2019). Atg2: A novel phospholipid transfer protein that mediates de novo autophagosome biogenesis. Protein Science 28, 1005–1012. doi: 10.1002/pro.3623. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Otomo T, Chowdhury S, Lander GC. (2018). The rod-shaped ATG2A-WIPI4 complex tethers membranes in vitro. Contact (Thousand Oaks 1. doi: 10.1177/2515256418819936. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paoli M. (2001). Protein folds propelled by diversity. Progress in Biophysics and Molecular Biology 76, 103–130. doi: 10.1016/S0079-6107(01)00007-4 [DOI] [PubMed] [Google Scholar]
- Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE. (2004). UCSF Chimera-a visualization system for exploratory research and analysis. Journal of Computational Chemistry 25, 1605–1612. doi: 10.1002/jcc.20084. [DOI] [PubMed] [Google Scholar]
- Pfisterer SG, Bakula D, Frickey T, Cezanne A, Brigger D, Tschan MP, Robenek H, Proikas-Cezanne T. (2014). Lipid droplet and early autophagosomal membrane targeting of Atg2A and Atg14L in human tumor cells. Journal of Lipid Research 55, 1267–1278. doi: 10.1194/jlr.M046359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Polson HE, de Lartigue J, Rigden DJ, Reedijk M, Urbe S, Clague MJ, Tooze SA. (2010). Mammalian Atg18 (WIPI2) localizes to omegasome-anchored phagophores and positively regulates LC3 lipidation. Autophagy 6, 506–522. doi: 10.4161/auto.6.4.11863. [DOI] [PubMed] [Google Scholar]
- Prinz WA, Toulmay A, Balla T. (2020). The functional universe of membrane contact sites. Nature Reviews Molecular Cell Biology 21, 7–24. doi: 10.1038/s41580-019-0180-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Proikas-Cezanne T, Ruckerbauer S, Stierhof YD, Berg C, Nordheim A. (2007). Human WIPI-1 puncta-formation: A novel assay to assess mammalian autophagy. FEBS Letters 581, 3396–3404. doi: 10.1016/j.febslet.2007.06.040. [DOI] [PubMed] [Google Scholar]
- Proikas-Cezanne T, Takacs Z, Donnes P, Kohlbacher O. (2015). WIPI Proteins: Essential PtdIns3P effectors at the nascent autophagosome. Journal of Cell Science 128, 207–217. doi: 10.1242/jcs.146258. [DOI] [PubMed] [Google Scholar]
- Proikas-Cezanne T, Waddell S, Gaugel A, Frickey T, Lupas A, Nordheim A. (2004). WIPI-1alpha (WIPI49), a member of the novel 7-bladed WIPI protein family, is aberrantly expressed in human cancer and is linked to starvation-induced autophagy. Oncogene 23, 9314–9325. doi: 10.1038/sj.onc.1208331. [DOI] [PubMed] [Google Scholar]
- Rao Y, Perna MG, Hofmann B, Beier V, Wollert T. (2016). The Atg1-kinase complex tethers Atg9-vesicles to initiate autophagy. Nature Communications 7, 10338. doi: 10.1038/ncomms10338. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reggiori F, Tucker KA, Stromhaug PE,, & Klionsky DJ. (2004). The Atg1-Atg13 complex regulates Atg9 and Atg23 retrieval transport from the pre-autophagosomal structure. Developmental Cell 6, 79–90. doi: 10.1016/S1534-5807(03)00402-7 [DOI] [PubMed] [Google Scholar]
- Ren J, Liang R, Wang W, Zhang D, Yu L, Feng W. (2020). Multi-site-mediated entwining of the linear WIR-motif around WIPI beta-propellers for autophagy. Nature Communications 11, 2702. doi: 10.1038/s41467-020-16523-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rieter E, Vinke F, Bakula D, Cebollero E, Ungermann C, Proikas-Cezanne T, Reggiori F. (2013). Atg18 function in autophagy is regulated by specific sites within its beta-propeller. Journal of Cell Science 126, 593–604. doi: 10.1242/jcs.115725. [DOI] [PubMed] [Google Scholar]
- Sakai Y, Koyama-Honda I, Tachikawa M, Knorr RL, Mizushima N. (2020). Modeling membrane morphological change during autophagosome formation. iScience 23, 101466. doi: 10.1016/j.isci.2020.101466. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sawa-Makarska J, Baumann V, Coudevylle N, von Bulow S, Nogellova V, Abert C, Schuschnig M, Graef M, Hummer G, Martens S. (2020). Reconstitution of autophagosome nucleation defines Atg9 vesicles as seeds for membrane formation. Science 369. doi: 10.1126/science.aaz7714. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scacioc A, Schmidt C, Hofmann T, Urlaub H, Kuhnel K, Perez-Lara A. (2017). Structure based biophysical characterization of the PROPPIN Atg18 shows Atg18 oligomerization upon membrane binding. Scientific Reports 7, 14008. doi: 10.1038/s41598-017-14337-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schutter M, Giavalisco P, Brodesser S,, & Graef M. (2020). Local fatty acid channeling into phospholipid synthesis drives phagophore expansion during autophagy. Cell 180, 135–149e114. doi: 10.1016/j.cell.2019.12.005 [DOI] [PubMed] [Google Scholar]
- Sekito T, Kawamata T, Ichikawa R, Suzuki K, Ohsumi Y. (2009). Atg17 recruits Atg9 to organize the pre-autophagosomal structure. Genes to Cells 14, 525–538. doi: 10.1111/j.1365-2443.2009.01299.x. [DOI] [PubMed] [Google Scholar]
- Shimizu T, Tamura N, Nishimura T, Saito C, Yamamoto H,, & Mizushima N. (2023). Comprehensive analysis of autophagic functions of WIPI family proteins and their implications for the pathogenesis of beta-propeller associated neurodegeneration. doi: 10.1101/2023.03.08.531694 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shintani T, Suzuki K, Kamada Y, Noda T, Ohsumi Y. (2001). Apg2p functions in autophagosome formation on the perivacuolar structure. Journal of Biological Chemistry 276, 30452–30460. doi: 10.1074/jbc.M102346200. [DOI] [PubMed] [Google Scholar]
- Soreng K, Munson MJ, Lamb CA, Bjorndal GT, Pankiv S, Carlsson SR, Tooze SA, Simonsen A. (2018). SNX18 regulates ATG9A trafficking from recycling endosomes by recruiting Dynamin-2. EMBO Reports 19. doi: 10.15252/embr.201744837. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strong LM, Chang C, Riley JF, Boecker CA, Flower TG, Buffalo CZ, Ren X, Stavoe AK, Holzbaur EL, Hurley JH. (2021). Structural basis for membrane recruitment of ATG16L1 by WIPI2 in autophagy. Elife 10. doi: 10.7554/eLife.70372. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Suzuki K, Akioka M, Kondo-Kakuta C, Yamamoto H, Ohsumi Y. (2013). Fine mapping of autophagy-related proteins during autophagosome formation in Saccharomyces cerevisiae. Journal of Cell Science 126, 2534–2544. doi: 10.1242/jcs.122960. [DOI] [PubMed] [Google Scholar]
- Suzuki K, Kirisako T, Kamada Y, Mizushima N, Noda T, Ohsumi Y. (2001). The pre-autophagosomal structure organized by concerted functions of APG genes is essential for autophagosome formation. EMBO Journal 20, 5971–5981. doi: 10.1093/emboj/20.21.5971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Suzuki K, Kubota Y, Sekito T, Ohsumi Y. (2007). Hierarchy of ATG proteins in pre-autophagosomal structure organization. Genes to Cells 12, 209–218. doi: 10.1111/j.1365-2443.2007.01050.x. [DOI] [PubMed] [Google Scholar]
- Suzuki SW, Yamamoto H, Oikawa Y, Kondo-Kakuta C, Kimura Y, Hirano H, Ohsumi Y. (2015). Atg13 HORMA domain recruits Atg9 vesicles during autophagosome formation. Proceedings of the National Academy of Sciences of the United States of America 112, 3350–3355. doi: 10.1073/pnas.1421092112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tamura N, Nishimura T, Sakamaki Y, Koyama-Honda I, Yamamoto H, Mizushima N. (2017). Differential requirement for ATG2A domains for localization to autophagic membranes and lipid droplets. FEBS Letters 591, 3819–3830. doi: 10.1002/1873-3468.12901. [DOI] [PubMed] [Google Scholar]
- Tan JX, Finkel T. (2022). A phosphoinositide signalling pathway mediates rapid lysosomal repair. Nature 609, 815–821. doi: 10.1038/s41586-022-05164-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tang Z, Takahashi Y, He H, Hattori T, Chen C, Liang X, Chen H, Young MM,, & Wang HG. (2019). TOM40 Targets Atg2 to mitochondria-associated ER membranes for phagophore expansion. Cell Reports 28, 1744–1757e1745. doi: 10.1016/j.celrep.2019.07.036 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thumm M, Busse RA, Scacioc A, Stephan M, Janshoff A, Kuhnel K, Krick R. (2013). It takes two to tango: PROPPINs use two phosphoinositide-binding sites. Autophagy 9, 106–107. doi: 10.4161/auto.22400. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ungermann C, Reggiori F. (2018). Atg9 proteins, not so different after all. Autophagy 14, 1456–1459. doi: 10.1080/15548627.2018.1477382. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Valverde DP, Yu S, Boggavarapu V, Kumar N, Lees JA, Walz T, Reinisch KM, Melia TJ. (2019). ATG2 Transports lipids to promote autophagosome biogenesis. Journal of Cell Biology 218, 1787–1798. doi: 10.1083/jcb.201811139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van Vliet AR, Chiduza GN, Maslen SL, Pye VE, Joshi D, De Tito S, Jefferies HBJ, Christodoulou E, Roustan C, Punch E, … (2022). ATG9A And ATG2A form a heteromeric complex essential for autophagosome formation. Molecular Cell 82, 4324–4339e4328. doi: 10.1016/j.molcel.2022.10.017 [DOI] [PubMed] [Google Scholar]
- Varadi M, Anyango S, Deshpande M, Nair S, Natassia C, Yordanova G, Yuan D, Stroe O, Wood G, Laydon A, … (2022). Alphafold protein structure database: Massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Research 50, D439–D444. doi: 10.1093/nar/gkab1061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Velikkakath AK, Nishimura T, Oita E, Ishihara N, Mizushima N. (2012). Mammalian Atg2 proteins are essential for autophagosome formation and important for regulation of size and distribution of lipid droplets. Molecular Biology of the Cell 23, 896–909. doi: 10.1091/mbc.E11-09-0785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang CW, Kim J, Huang WP, Abeliovich H, Stromhaug PE, Dunn WA, Jr.,, & Klionsky DJ. (2001). Apg2 is a novel protein required for the cytoplasm to vacuole targeting, autophagy, and pexophagy pathways. Journal of Biological Chemistry 276, 30442–30451. doi: 10.1074/jbc.M102342200 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Watanabe Y, Kobayashi T, Yamamoto H, Hoshida H, Akada R, Inagaki F, Ohsumi Y, Noda NN. (2012). Structure-based analyses reveal distinct binding sites for Atg2 and phosphoinositides in Atg18. Journal of Biological Chemistry 287, 31681–31690. doi: 10.1074/jbc.M112.397570. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weidberg H, Shvets E, Shpilka T, Shimron F, Shinder V, Elazar Z. (2010). LC3 And GATE-16/GABARAP subfamilies are both essential yet act differently in autophagosome biogenesis. The EMBO Journal 29, 1792–1802. doi: 10.1038/emboj.2010.74. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamamoto H, Kakuta S, Watanabe TM, Kitamura A, Sekito T, Kondo-Kakuta C, Ichikawa R, Kinjo M, Ohsumi Y. (2012). Atg9 vesicles are an important membrane source during early steps of autophagosome formation. Journal of Cell Biology 198, 219–233. doi: 10.1083/jcb.201202061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Young AR, Chan EY, Hu XW, Kochl R, Crawshaw SG, High S, Hailey DW, Lippincott-Schwartz J, Tooze SA. (2006). Starvation and ULK1-dependent cycling of mammalian Atg9 between the TGN and endosomes. Journal of Cell Science 119, 3888–3900. doi: 10.1242/jcs.03172. [DOI] [PubMed] [Google Scholar]
- Zhang Y, Ge J, Bian X, Kumar A. (2022). Quantitative models of lipid transfer and membrane contact formation. Contact (Thousand Oaks 5, 1–21. doi: 10.1177/25152564221096024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zheng JX, Li Y, Ding YH, Liu JJ, Zhang MJ, Dong MQ, Wang HW, Yu L. (2017). Architecture of the ATG2B-WDR45 complex and an aromatic Y/HF motif crucial for complex formation. Autophagy 13, 1870–1883. doi: 10.1080/15548627.2017.1359381. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zheng Q, Chen Y, Chen D, Zhao H, Feng Y, Meng Q, Zhao Y, Zhang H. (2022). Calcium transients on the ER surface trigger liquid-liquid phase separation of FIP200 to specify autophagosome initiation sites. Cell 185, 4082–4098. e4022. doi: 10.1016/j.cell.2022.09.001. [DOI] [PubMed] [Google Scholar]
- Zwilling E, Reggiori F. (2022). Membrane Contact Sites in autophagy. Cells 11. doi: 10.3390/cells11233813. [DOI] [PMC free article] [PubMed] [Google Scholar]