Abstract
Purpose
Retinoblastoma (RB) is most often diagnosed with clinical features and not diagnosed with tumor biopsy. This study describes tumor-derived analyte concentrations from aqueous humor (AH) liquid biopsy and its use in clinical assays.
Design
Case series study.
Participants
Sixty-two RB eyes from 55 children and 14 control eyes from 12 children from 4 medical centers.
Methods
This study included 128 RB AH samples including: diagnostic (DX) samples, samples from eyes undergoing treatment (TX), samples after completing treatment (END), and during bevacizumab injection for radiation therapy after completing RB treatment (BEV). Fourteen-control AH were analyzed for unprocessed analytes (double-stranded DNA [dsDNA], single-stranded DNA [ssDNA], micro-RNA [miRNA], RNA, and protein) with Qubit fluorescence assays. Double-stranded DNA from 2 RB AH samples underwent low-pass whole-genome sequencing to detect somatic copy number alterations. Logistic regression was used to predict disease burden given analyte concentrations.
Main Outcome Measures
Unprocessed analyte (dsDNA, ssDNA, miRNA, RNA and protein) concentrations.
Results
Results revealed dsDNA, ssDNA, miRNA, and proteins, but not RNA, were quantifiable in most samples (up to 98%) with Qubit fluorescence assays. Median dsDNA concentration was significantly higher in DX (3.08 ng/μl) compared to TX (0.18 ng/μl; P < 0.0001) at an order of 17 times greater and 20 times greater than END samples (0.15 ng/μl; P = 0.001). Using logistic regression, nucleic acid concentrations were useful in predicting higher versus lower RB disease burden. Retinoblastoma somatic copy number alterations were identified in a TX, but not in a BEV sample, indicating the correlation with RB activity.
Conclusions
Aqueous humor liquid biopsy in RB is a high-yield source of dsDNA, ssDNA, miRNA, and protein. Diagnostic samples are most useful for RB 1 gene mutational analyses. Genomic analysis may be more informative of tumor activity status than quantification alone and can be performed even with smaller analyte concentrations obtained from TX samples.
Financial Disclosure(s)
Proprietary or commercial disclosure may be found after the references.
Keywords: Aqueous humor, Liquid biopsy, Circulating analytes, Retinoblastoma, Multicenter
Retinoblastoma (RB) is rare among cancers in that it is diagnosed without a tumor biopsy due to concern for seeding tumor outside of the eye.1 Thus, until recently, there have been no means to identify or quantify tumor-specific biomarkers in RB at diagnosis or during treatment. However, in 2017 we demonstrated that aqueous humor (AH) liquid biopsy is a robust source of tumor derived cell-free DNA (cfDNA) attained via paracentesis.2 Aqueous humor liquid biopsy has enabled the identification of multiple tumor-derived analytes. Our group and others have demonstrated that tumor DNA is present in the AH of eyes with RB and can be utilized to identify retinoblastoma 1 gene (RB1) single-nucleotide variants (SNVs), somatic copy number alterations (SCNAs), tumor methylation status, and to estimate tumor fraction.2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20 Other groups have evaluated the AH as a liquid biopsy in RB for other analytes including survivin,21 metabolomic signatures,19 expression of secreted peptides,22 and nucleic acids,23 and in other intraocular tumor types and proteins.24
Given the potential clinical application of AH liquid biopsy in RB patients, there is a need for quantification of analytes that may guide the type of genomic analysis testing and biomarker identification that can be done at various clinical time points. Further, quantifying analytes from RB eyes that are treatment-naive versus actively undergoing treatment may facilitate more accurate and objective ways to determine intraocular disease activity than clinical examination alone.
The purpose of our report is to describe:
-
(1)
The expected concentrations of AH nucleic acids and proteins based on RB International Intraocular Retinoblastoma Classification (IIRC) Groups and at various therapeutic time points ranging from diagnosis to the end of therapy.
-
(2)
The potential clinical implications for AH liquid biopsy analyses based on quantification of AH analytes and the input needed for various assays.
-
(3)
A stratification of disease burden through logistic modeling based on AH nucleic acids and protein concentration alone.
Methods
This investigation is a multicenter case series study at tertiary care hospitals with RB treatment centers. Sites that actively contribute samples to the AH biorepository include: Children’s Hospital Los Angeles, Emory Eye Center, Oregon Health & Science University Hospital Casey Eye Institute, Wills Eye Hospital, and University of Washington Medical Center. All samples were processed in the same laboratory at Children’s Hospital Los Angeles.
Samples were taken between August 2018 and June 2022. Institutional Review Board approval was obtained at each institution for this study. The study was conducted in accordance with the Declaration of Helsinki. Written informed consent was obtained from the parents of all participants and that included permission for publication.
Specimen Collection and Storage
The detailed methods have been previously published.25 In brief, a paracentesis with extraction of 0.1 ml of AH with a 32-gauge needle was performed. There was no contact between the needle and the retinal tumor or the vitreous cavity. Aqueous humor was extracted at various time points throughout therapy, all while the child was under general anesthesia for routine clinical care for their ocular condition. Aqueous humor samples were immediately stored on dry ice and then transferred to a −80°C refrigerator within hours of extraction.
Quantification of Nucleic Acid and Protein Content in AH
Nucleic acids (double-stranded DNA [dsDNA], single-stranded DNA [ssDNA], RNA, and micro-RNA [miRNA]) were assayed using high-sensitivity Qubit Assays for dsDNA and RNA and Qubit Assay Kits for ssDNA and miRNA (Thermo Fisher), with the Qubit Fluorometer following the manufacturer’s manual. Protein concentrations were assayed using the Qubit Protein broad range (BR) assay. The lower limit of detection of these kits are:
dsDNA = 0.005 ng/μl
ssDNA = 0.05 ng/μl
RNA = 0.2 ng/μl
miRNA = 0.025 ng/μl
protein = 100 ng/μl
Genomic Analysis of Samples
Aqueous humor samples underwent DNA isolation, sequencing, and analysis within 1 month of collection at Children's Hospital Los Angeles. The methods for this have been previously published.4,26 While this was done for each sample the results of genomic sequencing have been (or will be) published separately4,5,7, 8, 9,27 and are not the focus of this report, aside from an illustrative case study (see Discussion section).
Statistical Analysis
Analyte concentration samples (dsDNA, ssDNA, miRNA, RNA, and protein) were categorized based on IIRC classification (Groups A–E) of the source eye, as well as disease burden at the time of sample collection. Samples collected at diagnosis prior to subsequent eye-salvage therapy or at primary enucleation were considered diagnostic samples (DX), showing the highest disease burden. Those collected at secondary enucleation, at the time of systemic chemotherapy, intra-arterial chemotherapy, or intravitreal chemotherapy injection were categorized as treatment (TX) samples, reflecting moderate disease burden. Eyes that received bevacizumab injection for radiation retinopathy after completing RB treatment are categorized as sample taken at bevacizumab injection (BEV) and those collected up to 6 months after completing RB treatment were categorized as END samples. Bevacizumab injection and END samples were from RB patients with no clinical evidence of active intraocular RB tumors or seeding. Samples from eyes diagnosed with conditions other than RB (cataracts, glaucoma, pediatric retinal disease [PRD]) were used as controls.
Differences in analyte values were evaluated overall using Kruskal-Wallis rank tests and tested pairwise using Dunn’s test with Benjamini-Hochberg correction for multiple tests. Analyte concentration values were log-transformed for further evaluation in logistic regression models, which were used to determine cut-off values that distinguish the highest disease burden (DX) from samples with lower burden (TX, BEV, and END) and also examine whether moderate disease burden (TX) could be distinguished from the lowest (BEV and END) based on analyte concentrations. For each analyte, a concentration value was selected that optimized sensitivity and specificity of a given model; these values were back-transformed for use by clinicians. All analyses were conducted using Stata/SE 14.2.
Results
Patient Demographics and AH Sample Characteristics
A total of 128 RB AH samples from 62 RB eyes in 55 children and 14 AH samples from 14 non-RB eyes (3 pediatric cataracts, 2 pediatric glaucoma, and 9 PRD) in 12 children were obtained. No patients had complications secondary to extraction of AH. Thirty-one RB diagnostic AH samples were taken prior to subsequent eye-salvage therapy or at primary enucleation (DX), 82 samples were taken during treatment (TX), 10 samples were taken during bevacizumab injection (BEV), and 5 samples 6 months after completing RB treatment (END). Among DX samples, 2 Group A, 2 Group B, 3 Group C, 20 Group D, and 4 Group E eyes samples were evaluated.
Retinoblastoma tumor IIRC Group, The American Joint Committee on Cancer stage, clinical characteristics, and AH sample characteristics for each case are summarized in Tables 1 and 2. The median age for RB patients at diagnosis was 14.5 months (range 11 days–46 months) and median age of control patients was 87.25 months (range 1–208 months).
Table 1.
RB Patient Demographic and Clinical Information
| Case ID | Disease | Age at Diagnosis (Months) | Sex | Disease Laterality | Eye Included in Analysis | IIRC at Diagnosis | AJCC | Vitreous Seeding at Diagnosis | Blood RB1 Mutation | Initial Treatment | Required IVM/IVC? | Required Enucleation? | Time from Diagnosis to Enucleation (Months) | Samples Included in Analysis |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | RB | 10 | M | U | OS | D | CT2B | None | Negative | IAC | Yes | Yes | 22 | 1 |
| 2_OD | RB | 2 | F | B | OD | E | CT2B | None | c.1421+12_1421+32del21bp | CEV | Yes | Yes | 50 | 1 |
| 2_OS | RB | 2 | F | B | OS | B | CT1B | None | c.1421+12_1421+32del21bp | CEV | Yes | No | N/A | 1 |
| 3 | RB | 3 | F | B | OD | D | CT2B | None | Familial RB, mutation unknown | CEV | Yes | Yes | 8 | 1 |
| 4 | RB | 38 | M | B | OD | D | CT2B | Dust and Sphere | c.1961-2A>G | CEV | No | Yes | 9 | 1 |
| 5 | RB | 10 | F | U | OS | D | CT3D | Dust | Negative | PE | N/A | Yes | 0 | 1 |
| 6 | RB | 22 | M | U | OS | D | CT2B | Sphere | Negative | IAC | Yes | Yes | 5 | 5 |
| 7 | RB | 16 | F | B | OS | D | CT2B | None | c.1389+2T>C | PE | N/A | Yes | 0 | 1 |
| 8 | RB | 6 | M | U | OD | C | CT1B | Dust | Negative | CEV | Yes | No | N/A | 2 |
| 9 | RB | 13 | M | B | OS | E | CT3C | Dust | c.1494T>G | CEV | Yes | No | N/A | 3 |
| 10 | RB | 4 | F | U | OD | D | CT2B | Dust | Negative | CEV | Yes | No | N/A | 4 |
| 11 | RB | 18 | F | U | OD | D | CT2B | Dust | c.958C>T | CEV | Yes | No | N/A | 3 |
| 12 | RB | 19 | F | B | OS | D | CT2B | Cloud | c.1875delT | CEV | Yes | No | N/A | 5 |
| 13 | RB | 28 | M | U | OS | D | CT2B | Dust | c.1847_1848delAA | IAC | Yes | No | N/A | 6 |
| 14 | RB | 13 | F | U | OS | D | CT2B | Sphere | Negative | IAC | Yes | No | N/A | 6 |
| 15_OD | RB | 4 | F | B | OD | B | CT1B | None | c.1666C>T | CEV, IAC | No | No | N/A | 2 |
| 15_OS | RB | 4 | F | B | OS | D | CT2B | Dust | c.1666C>T | CEV, IAC | Yes | No | N/A | 9 |
| 16 | RB | 8 | F | U | OD | D | CT2B | None | Negative | CEV | No | No | N/A | 1 |
| 17 | RB | 5 | M | U | OS | C | CT2A | None | Negative | CEV | No | No | N/A | 2 |
| 18 | RB | 15 | F | U | OD | D | CT2B | Sphere and Dust | Negative | CEV | Yes | No | N/A | 4 |
| 19 | RB | 18 | M | U | OD | D | CT2B | Predominantly cloud | Negative | IAC | No | Yes | 1 | 1 |
| 20 | RB | 35 | M | U | OD | D | CT1B | Predominantly cloud | Negative | PE | N/A | Yes | 0 | 1 |
| 21 | RB | 24 | F | U | OD | D | CT2B | None | Negative | PE | N/A | Yes | 0 | 1 |
| 22 | RB | 30 | M | U | OD | D | CT2B | Dust | Negative | IAC | No | No | N/A | 1 |
| 23 | RB | 15 | F | U | OS | D | CT2B | Sphere | Negative | PE | N/A | Yes | 0 | 1 |
| 24 | RB | 24 | F | U | OD | E | CT3C | Sphere | Negative | PE | N/A | Yes | 0 | 1 |
| 25 | RB | 25 | M | U | OD | E | CT3C | Sphere | Negative | PE | N/A | Yes | 0 | 1 |
| 26 | RB | 24 | M | U | OS | D | CT2B | Sphere | Negative | IAC | Yes | Yes | 5 | 2 |
| 27 | RB | 18 | F | B | OS | D | CT2B | Unknown, presented post treatment | Positive, mutation unknown | CEV | Yes | No | N/A | 6 |
| 28 | RB | 24 | F | U | OD | E | CT3C | Unknown, filled globe | Negative | PE | N/A | Yes | 0 | 1 |
| 29 | RB | 13 | M | U | OS | D | CT2B | Dust | c.607+1G>T | IAC | No | No | N/A | 1 |
| 30 | RB | 24 | F | U | OD | D | CT2B | Dust | Negative | CEV | Yes | No | N/A | 5 |
| 31_OD | RB | 9 | M | B | OD | E | CT2B | Sphere | c.1732delG | CEV | Yes | Yes | 19 | 3 |
| 31_OS | RB | 9 | M | B | OS | D | CT3B | Sphere | c.1732delG | CEV | Yes | Yes | 9 | 2 |
| 32_OD | RB | 6 | M | B | OD | C | CT2B | Dust and Sphere | c.2027T>G | CEV | No | No | N/A | 1 |
| 32_OS | RB | 6 | M | B | OS | D | CT2B | Dust | c.2027T>G | CEV | No | No | N/A | 1 |
| 33 | RB | 19 | F | U | OD | D | CT2B | Sphere | Negative | IAC | Yes | Yes | 11 | 5 |
| 34 | RB | 15 | F | B | OS | E | CT3C | Unknown, filled globe | Deletion of exons 24–27 | CEV | No | Yes | 7 | 1 |
| 35 | RB | 24 | M | U | OS | D | CT2B | Filled globe (Cloud, Dust, Sphere) | c.3920T>A | PE | N/A | Yes | 0 | 2 |
| 36 | RB | 24 | F | B | OS | D | CT2B | Cloud | c.1362 C>G | IAC | Yes | Yes | 9 | 5 |
| 37 | RB | 24 | M | U | OS | D | CT2B | Dust | Negative | CEV | No | Yes | 36 | 2 |
| 38 | RB | 1 | F | U | OS | D | CT2B | Dust | Negative | CEV | Yes | No | N/A | 4 |
| 39 | RB | 24 | F | U | OD | D | CT2B | Cloud | Negative | PE | N/A | Yes | 0 | 1 |
| 40_OD | RB | 2 | M | B | OD | A | CT1A | None | c.607+1G>C | Laser | No | No | N/A | 1 |
| 40_OS | RB | 2 | M | B | OS | A | CT1A | None | c.607+1G>C | Laser | No | No | N/A | 1 |
| 41 | RB | 8 | F | U | OD | D | CT2B | Sphere | c.1362C>G | CEV | Yes | No | N/A | 2 |
| 42_OD | RB | 27 | F | B | OD | A | CT1A | None | c.1399C>T | Laser | No | No | N/A | 1 |
| 42_OS | RB | 27 | F | B | OS | D | CT2B | Cloud | c.1399C>T | CEV | No | Yes | 7 | 1 |
| 43 | RB | 35 | M | U | OS | D | CT2B | Cloud | Negative | CEV | Yes | No | N/A | 1 |
| 44_OD | RB | 4 | F | B | OD | B | CT1B | None | c.1215+1G>A | CEV, Laser | No | No | N/A | 1 |
| 44_OS | RB | 4 | F | B | OS | C | CT2B | None | c.1215+1G>A | CEV | No | No | N/A | 1 |
| 45 | RB | 46 | M | U | OS | D | CT2B | None | Negative | CEV | No | No | N/A | 1 |
| 46 | RB | 4 | M | U | OS | C | CT2B | None | c.958C>T | CEV | No | No | N/A | 1 |
| 47 | RB | 8 | F | U | OS | D | CT2B | Sphere | Negative | CEV | No | No | N/A | 1 |
| 48 | RB | 6 | M | U | OD | D | CT2B | Unknown, filled globe | Negative | CEV | No | No | N/A | 1 |
| 49 | RB | 20 | F | U | OD | D | CT2B | Cloud | Positive, mutation unknown | IAC | Yes | No | N/A | 1 |
| 50 | RB | 36 | M | B | OS | E | CT2B | Cloud | Positive, mutation unknown | IAC | Yes | No | N/A | 1 |
| 51 | RB | < 1 (11 days) | F | B | OD | A | CT1A | None | N/A | CEV | Yes | No | N/A | 1 |
| 52 | RB | 10 | M | U | OD | D | CT2B | Cloud | Negative | IAC | Yes | No | N/A | 1 |
| 53 | RB | 4 | F | U | OD | E | CT3C | None | Negative | PE | N/A | Yes | 0 | 1 |
| 54 | RB | 14 | F | U | OS | C | CT1B | None | Positive, mutation unknown | IAC | Yes | Yes | Unknown | 1 |
| 55 | RB | 22 | F | U | OS | D | CT2B | Sphere | Negative | IAC | Yes | Yes | Unknown | 1 |
AJCC = American Joint Committee on Cancer; B = bilateral disease; CEV = systemic carboplatin [C]/etoposide[E]/vincristine[V]; F = female; IAC = intra-arterial chemotherapy; Case ID = case identification; IIRC = International Intraocular Retinoblastoma Classification; IVC = intravenous chemotherapy; IVM = intravitreal melphalan; M= male; N/A = not applicable; OD = right eye; OS = left eye; PE = primary enucleation; RB = retinoblastoma; RB1 = retinoblastoma 1 gene; U = unilateral disease.
Table 2.
Control Sample Demographic Information
| Case ID | Diagnosis | Age at Diagnosis (Months) | Sex | Disease Laterality | AH Sample Eye |
|---|---|---|---|---|---|
| Ret09 | PRD | 92 | M | U | OD |
| Ret011 | PRD | 181.9 | F | U | OD |
| Ret017 | PRD | 36 | M | U | OS |
| Ret018 | PRD | 181.7 | M | U | OS |
| Ret013 | PRD | 146 | M | U | OD |
| Ret014 | PRD | 204 | M | B | OD |
| Ret014 | PRD | 204 | M | B | OS |
| Ret019 | PRD | 208 | M | U | OD |
| Ret015 | PRD | 82.5 | M | U | OS |
| Glc05 | Pediatric Glaucoma | 4 | M | B | OD |
| Glc09 | Pediatric Glaucoma | 50 | F | U | OD |
| Cat07 | Pediatric Cataract | 55 | M | B | OD |
| Cat07 | Pediatric Cataract | 55 | M | B | OS |
| Cat08 | Pediatric Cataract | 1 | M | U | OD |
AH = aqueous humor; B = bilateral disease; F = female; Case ID = case identification; M = male; OD = right eye; OS = left eye; PRD = pediatric retinal disease; U = unilateral disease.
AH Nucleic Acid (dsDNA, ssDNA, RNA, and miRNA) and Protein Concentrations
All AH samples were stratified by time of sample collection and IIRC classification. Mean (with standard deviation) and median (with range) concentration values for these groupings are shown in Table 3. Most samples demonstrated measurable concentrations of dsDNA (84.4% measurable), ssDNA (95.2%), miRNA (36.7%), and protein (97.7%). Samples that had levels of dsDNA, ssDNA, RNA, miRNA, and protein below detection thresholds are counted in Table 3 as “ND” (not detected).
Table 3.
Sample Analyte Ranges Including Mean, Median, and Number of AH Samples for dsDNA, ssDNA, RNA, miRNA, and Protein by Time of AH Sample, Disease Type, and Burden
| Diagnosis | dsDNA (ng/μl) |
ssDNA (ng/μl) |
RNA (ng/μl) |
miRNA (ng/μl) |
Protein (ng/μl) |
|||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample Size |
Mean ± SD | Median (Range) | Sample Size |
Mean ± SD | Median (Range) | Sample Size |
Mean ± SD | Median (Range) | Sample Size |
Mean ± SD | Median (Range) | Sample Size |
Mean ± SD | Median (Range) | ||||||
| #Read | ND | #Read | ND | #Read | ND | #Read | ND | #Read | ND | |||||||||||
| RB AH taken at Diagnosis∗ (n = 31) |
27 | 4 | 13.65 ± 18.63 | 3.08 (0.41–68) | 28 | 1 | 43.62 ± 69.06 | 8.03 (0.28–256) | 3 | 22 | 5.56 ± 1.05 | 5.84 (4.40–6.44) | 25 | 4 | 43.51 ± 56.37 | 11.9 (0.80–232) | 29 | 0 | 2853 ± 3635 | 2020 (118–19 200) |
| All Samples (n = 128) | 108 | 20 | 3.74 ± 10.93 | 0.22 (0.02–68) | 120 | 6 | 11.04 ± 37.67 | 0.68 (0.10–256) | 3 | 117 | 5.56 ± 1.05 | 5.84 (4.40–6.44) | 47 | 79 | 24.3 ± 45.89 | 2.5 (0.06–232) | 125 | 1 | 1868 ± 2086 | 1304 (118–19200) |
| By Eye Classification AH taken at Diagnosis∗: |
||||||||||||||||||||
| A (n = 2) | 0 | 2 | – | 1 | 1 | 0.28 | 0.28 | 0 | 2 | – | 0 | 2 | – | 2 | 0 | 264 ± 206 | 264 (118–410) | |||
| B (n = 2) | 2 | 0 | 1.15 ± 0.05 | 1.18 (1.11–1.18) | 1 | 0 | 3.12 | 3.12 | 0 | 1 | – | 1 | 0 | 2.78 | 2.78 | 1 | 0 | 220 | 220 | |
| C (n = 3) | 3 | 0 | 0.78 ± 0.59 | 0.47 (0.41–1.46) | 3 | 0 | 2.32 ± 1.36 | 1.7 (1.39–3.88) | 0 | 3 | – | 3 | 0 | 2.14 ± 0.97 | 1.86 (1.35–3.22) | 3 | 0 | 589 ± 137 | 618 (440–710) | |
| D (n = 20) | 18 | 2 | 14.9 ± 16.86 | 7.29 (0.51–56.6) | 19 | 0 | 43.87 ± 61.19 | 11.9 (0.64–239) | 1 | 14 | 6.44 | 6.44 | 17 | 2 | 42.25 ± 39.78 | 26.3 (0.80–120) | 19 | 0 | 2527 ± 1785 | 2040 (460–6680) |
| E (n = 4) | 4 | 0 | 23.99 ± 31.27 | 13.35 (1.27–68) | 4 | 0 | 94.34 ± 118.94 | 58.6 (4.14–256) | 2 | 2 | 5.12 ± 1.02 | 5.12 (4.4–5.84) | 4 | 0 | 90.06 ± 108.71 | 61.94 (4.34–232) | 4 | 0 | 8055 ± 7449 | 4720 (3580–19 200) |
| Diagnosis | dsDNA (ng/μl) |
ssDNA (ng/μl) |
RNA (ng/μl) |
miRNA (ng/μl) |
Protein (ng/μl) |
|||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample Size | Mean ± SD | Median (Range) | Sample Size | Mean ± SD | Median (Range) | Sample Size | Mean ± SD | Median (Range) | Sample Size | Mean ± SD | Median (Range) | Sample Size | Mean ± SD | Median (Range) | ||||||
| #Read | ND | #Read | ND | #Read | ND | #Read | ND | #Read | ND | |||||||||||
| By Disease Burden | ||||||||||||||||||||
| Highest∗ (n = 31) | 27 | 4 | 13.65 ± 18.63 | 3.08 (0.41–68) | 28 | 1 | 43.62 ± 69.06 | 8.03 (0.28–256) | 3 | 22 | 5.56 ± 1.05 | 5.84 (4.40–6.44) | 25 | 4 | 43.51 ± 56.37 | 11.9 (0.80–232) | 29 | 0 | 2853 ± 3635 | 2020 (118–19 200) |
| Moderate∗∗ (n = 82) | 68 | 14 | 0.25 ± 0.26 | 0.18 (0.02–1.57) | 77 | 5 | 0.75 ± 0.79 | 0.56 (0.10–5.6) | 0 | 81 | – | 19 | 63 | 1.12 ± 1.46 | 0.64 (0.06–5.22) | 81 | 1 | 1514 ± 1106 | 1264 (200–5040) | |
| Bevacizumab at the End of RB Treatment∗∗∗ (n = 10) | 10 | 0 | 1.74 ± 4.7 | 0.193 (0.114–15.1) | 10 | 0 | 4.42 ± 11.5 | 0.72 (0.38–37) | 0 | 10 | – | 3 | 7 | 11 ± 17.7 | 0.8 (0.68–31.4) | 10 | 0 | 2461 ± 1570 | 2225 (710–5000) | |
| End of RB Treatment∗∗∗∗ (n = 5) | 3 | 2 | 0.114 ± 0.075 | 0.152 (0.028–0.162) | 5 | 0 | 0.352 ± 0.198 | 0.24 (0.22–0.68) | 0 | 5 | – | 0 | 5 | – | 5 | 0 | 697 ± 516 | 380 (276–1474) | ||
| Other Conditions | 9 | 5 | 1.2 ± 2.06 | 0.23 (0.07–6.50) | 12 | 2 | 1.46 ± 1.73 | 0.54 (0.11–4.98) | 4 | 10 | 3.5 ± 2.34 | 3.75 (0.7–5.8) | 7 | 7 | 3.99 ± 5.97 | 1.91 (0.16–17) | 14 | 0 | 268 ± 605 | 11.48 (0.30–2080) |
| Cataracts | 0 | 3 | – | – | 1 | 2 | 0.24 | 0.24 | 0 | 3 | 0 | 3 | – | – | 3 | 0 | 169 ± 208 | 80 (20–406) | ||
| Glaucoma | 2 | 0 | 0.09 ± 0.03 | 0.09 (0.07–0.11) | 2 | 0 | 0.51 ± 0.01 | 0.51 (0.50–0.52) | 0 | 2 | – | – | 0 | 2 | – | 2 | 0 | 1602 ± 676 | 1602 (1124–2080) | |
| PRD | 7 | 2 | 1.52 ± 2.27 | 0.74 (0.13–6.50) | 9 | 0 | 1.8 ± 1.89 | 1.03 (0.11–4.98) | 4 | 5 | 3.5 ± 2.34 | 3.75 (0.7–5.8) | 7 | 2 | 3.99 ± 5.97 | 1.91 (0.16-17) | 9 | 0 | 5.18 ± 5.99 | 1.75 (0.30–14.96) |
AH = aqueous humor; dsDNA = double-stranded DNA; miRNA = micro-RNA; ND = not detected; PRD = pediatric retinal disease; RB = retinoblastoma; SD = standard deviation; ssDNA = single-stranded DNA.
Highest disease burden: ∗Includes samples taken at diagnosis prior to any subsequent treatment and at primary enucleation. Moderate disease burden: ∗∗During treatment, during intravitreal melphalan, intra-arterial chemotherapy, systemic chemotherapy or at secondary enucleation. Lowest disease burden: ∗∗∗End of RB treatment during bevacizumab injection and ∗∗∗∗End of RB treatment.
#Read indicates the number of samples with detectable levels of each analyte. ND, not detected, indicates the number of samples with a concentration too low for detection.
Evaluated together, RB AH samples had 113.6 times higher median protein concentrations than Controls, a significant difference (1304 ng/μl [118–19 200] vs. 11.48 ng/μl [0.30–2080], P < 0.0001), but no significant differences were found between RB and control samples for median concentrations of dsDNA, ssDNA, RNA, or miRNA (Fig 1A–E and Table 3). It should be noted that AH from eyes with PRD have higher concentrations than cataract or glaucoma samples, but all control AH samples were evaluated collectively due to the limited number available.
Figure 1.
Quantification of aqueous humor analytes in all retinoblastoma (RB) samples and controls: (A) double-stranded DNA (dsDNA), (B) single-stranded DNA (ssDNA), (C) RNA (D) micro-RNA (miRNA), and (E) protein concentration of RB eyes and control eyes were analyzed. SEM = standard error of the mean.
Analyte Concentration by Disease Burden
When RB AH samples were further stratified by treatment time point, DX samples had significantly higher analyte concentrations than TX for all analyte types: DX versus TX dsDNA: 3.08 ng/μl (0.41–68) versus 0.18 ng/μl (0.02–1.57), P < 0.0001; ssDNA: 8.03 ng/μl (0.28–256) versus 0.56 ng/μl (0.10–5.60), P < 0.0001; miRNA: 11.90 ng/μl (0.80–232) versus 0.64 ng/μl (0.06–5.22), P = 0.009; protein: 2020 ng/μl (118–19 200) versus 1264 ng/μl (200–5040), P = 0.01; see Fig 2A–E and Table 3.
Figure 2.
Quantification of retinoblastoma (RB) aqueous humor analytes from: diagnostic (DX), during RB treatment (TX), during bevacizumab injection at the end of RB treatment (BEV), and end of RB treatment (END) samples. A, double-stranded DNA (dsDNA),(B) single-stranded DNA (ssDNA), (C) RNA, (D) micro-RNA (miRNA), and (E) protein concentration. NA = not available; SEM = standard error of the mean.
DNA concentrations were significantly higher in DX samples compared to BEV and END samples, in which there was no active intraocular disease clinically, for both dsDNA and ssDNA: dsDNA (DX 3.08 ng/μl [0.41–68]) versus (BEV 0.19 ng/μl [0.11–15.10]; P = 0.005) and (END 0.15 ng/μl [0.03–0.16]; P = 0.001); ssDNA (DX 8.03 ng/μl [0.28–256]) versus (BEV 0.72 ng/μl [0.38–37]; P = 0.01); and (END 0.24 ng/μl [0.22–0.68]; P = 0.04). Of note, only 3 out of 5 END samples had detectable dsDNA concentrations, but all 5 samples had detectable ssDNA and protein [380 ng/μl (276–1474)]. Micro-RNA and RNA concentrations were not detectable in END samples.
Analyte Concentrations at Diagnosis by IIRC Group
Diagnostic samples were evaluated by IIRC Groups and are shown in Table 3. Out of only 2 Group A DX samples, proteins were detectable in both (264 ng/μl [118–410]), but only 1 sample yielded detectable ssDNA concentration (0.28 ng/μl) and other nucleic acids were undetectable. Samples from eyes with moderate disease (Groups B and C) were compared to those with more extensive disease (Groups D and E). As shown in Figure 3A–E, analyte concentrations in Group D/E eyes were significantly higher than concentrations in Group B/C eyes: dsDNA: 7.29 ng/μl (2.10–24.60) versus 1.11 ng/μl (0.47–1.18), P = 0.005; ssDNA: 11.90 ng/μl (2.60–80.60) versus 2.41 ng/μl (1.55–3.5), P = 0.049; miRNA: 26.30 ng/μl (5.48–77) versus 2.32 ng/μl (1.61–3), P = 0.01; protein: 2920 ng/μl (1008–4660) versus 529 ng/μl (330–664), P = 0.001; see Figure 3A–E.
Figure 3.
Quantification of retinoblastoma (RB) aqueous humor (AH) analytes from diagnostic AH samples grouped by International Intraocular Retinoblastoma Classification. A, double-stranded DNA (dsDNA), (B) single-stranded DNA (ssDNA), (C) RNA, (D) micro-RNA (miRNA), and (E) protein concentration. NA = not available; SEM = standard error of the mean.
Analyte Concentrations as Predictors of RB Disease Burden
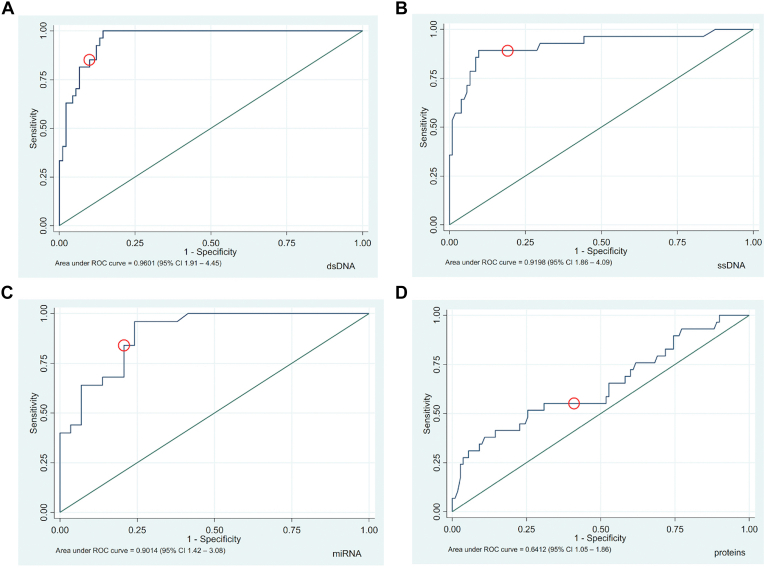
Analyte concentrations were used to predict RB disease burden (e.g., DX vs. TX vs. END) in logistic regression models. As illustrated in Figure 4A–D, sample concentrations were useful predictors of whether a sample was obtained from an eye with the highest disease burden. Aqueous humor concentration cut-offs of dsDNA 0.61 ng/μl (89% accuracy), ssDNA 1.38 ng/μl (90% accuracy) and miRNA 2.4 ng/μl (81% accuracy) were able to distinguish higher and lower disease burden, with higher analyte concentrations correlating to higher disease burden. Protein concentrations were less useful in identifying highest disease burden, where concentration of 1310 ng/μl could be used to identify a sample with highest disease burden eye with only 56% accuracy. Logistic model calculations were not performed on RNA concentrations due to insufficient quantifiable RNA samples. Analyte concentrations were not useful in distinguishing moderate disease burden (TX) from low-disease burden samples (END or BEV).
Figure 4.
Receiver operating curves (ROC) curve analysis of the utility of aqueous humor analyte concentrations to determine retinoblastoma disease burden. Cut-off values for analyte concentration that maximizes sensitivity and specificity are circled in red. Disease burden was defined as: Highest disease burden (Diagnostic: diagnostic or primarily enucleated samples), Moderate disease burden (During treatment: at secondary enucleation, systemic chemotherapy, intravitreal melphalan injection, or intra-arterial chemotherapy), Lowest disease burden (End of treatment and during bevacizumab injection after the end of treatment). A, double-stranded DNA (dsDNA), (B) single-stranded DNA (ssDNA), (C) micro-RNA (miRNA), and (D) protein concentration models. CI = confidence interval.
Clinical Correlates of RB Somatic Copy Number Alteration Analysis
As previously reported, common highly recurrent RB SCNAs include 1q, 2p and 6p gain, and 16q and 13q loss are identifiable in AH samples.4,7,8,28 To demonstrate the impact of SCNA analysis versus analyte quantification alone, 2 representative AH samples from the same eye at 2 different clinical time points involving intravitreal injections were examined; one obtained during RB therapy using intravitreal chemotherapy for treating seeds and 1 at the end of therapy during bevacizumab injection to treat radiation retinopathy (Fig 5A). Both TX and BEV samples had similar dsDNA, ssDNA, and protein analyte concentrations, while miRNA and RNA levels were undetectable. The SCNA profile of a sample taken during intravitreal chemotherapy displayed RB SCNAs of 6p gain, as well as 16q loss (Fig 5B). The genomic profile of an RB case undergoing additional bevacizumab injections had no discernable SCNAs on whole genome sequencing (Fig 5C).
Figure 5.
Analyte concentrations and somatic copy number alteration (SCNA) profiles from the same eye at different clinical times. A, AH analyte (double-stranded DNA [dsDNA], single-stranded DNA [ssDNA], micro-RNA [miRNA], RNA, and protein) concentrations from Case 14 undergoing intravitreal melphalan (IVM) and the same eye at the end of treatment receiving bevacizumab (BEV) injections. B, Case 14 IVM, retinoblastoma SCNAs of 6p gain and 16q loss are indicated with an ∗. C, Case 14 BEV, a flat SCNA profile.
Discussion
This is a multicenter study analyzing the use of AH liquid biopsy with 128 RB AH and 14 Control AH samples to investigate the quantification of AH nucleic acid and protein analytes. Herein we demonstrate that (1) the AH is a high-yield source of multiple circulating analytes (dsDNA, ssDNA, miRNA, and protein) in RB with concentrations that can be measured at diagnosis and through therapy, (2) AH analyte (dsDNA, ssDNA, miRNA, and protein) concentrations were highest in samples taken before treatment (DX), either at diagnosis or primary enucleation, and (3) analyte concentrations alone during treatment may not be sufficient in determining whether or not there is active intraocular disease, but the addition of genomic analysis may be more useful. To the authors’ knowledge, this is the first multicenter report that investigates quantifiable analytes in AH liquid biopsy of RB and has important implications for future use of the AH in clinical assays of various analytes as a diagnostic and prognostic liquid biopsy.
Herein we report mean and median nucleic acid (dsDNA, ssDNA, miRNA) and protein concentrations from diagnosis to the end of treatment (Table 3). Most analytes were readily detectable from diagnosis through therapy. However, RNA was only detectable in DX samples with Qubit high sensitivity RNA assay, likely due to high degree of degradation. This RNA assay is designed to detect RNA fragments longer than 100 nucleotides. Aqueous humor RNA may be better detected using other methods. For dsDNA, median concentrations from DX AH samples were on the order of 17 times higher than TX, 16 times higher than BEV, and 20 times higher than END samples. Diagnostic AH sample concentrations were also significantly higher than TX samples for dsDNA, ssDNA, miRNA, and protein (Fig 2A–E). Diagnostic samples had higher concentrations than BEV (Fig 2A) and END (Fig 2B) samples for dsDNA and ssDNA. As hypothesized, DX concentrations had the highest concentration of all analytes, which implicates DX AH liquid biopsy as a promising liquid biopsy source of tumor-derived analytes for various RB research and clinical assays.
International Intraocular Retinoblastoma Classification Groups also correlated with analyte concentrations, with more severe Groups D and E having higher analyte concentrations. When comparing DX samples from different IIRC Groups, Groups E dsDNA median concentrations were on the order of 1.8 times higher than Group D, 28.4 times greater than Group C, and 11.3 times greater than Group B; almost directly correlating to the increase in tumor size between Groups. Group A samples had mostly undetectable nucleic acid concentrations. Group D and E eyes had significantly higher concentrations than Group B and C for dsDNA, ssDNA, miRNA, and protein, but not RNA (Fig 3A–E). This demonstrates that DX AH samples from all except Group A eyes with the smallest tumor (< 3 mm tumors) are expected to have detectible analyte concentrations.
We then investigated AH analyte concentrations’ relationship to disease burden, defined by DX and primary enucleation samples as highest disease burden. With a logistic regression model using analyte concentrations, we found dsDNA, ssDNA, and miRNA levels were useful in predicting disease burden, with higher values clearly defining greater disease burden. All nucleic acid (dsDNA, ssDNA, and miRNA) models were strongly correlated with one another, while protein levels were less useful in distinguishing disease burden (Fig 4A–D). Diagnostic samples could be identified by AH concentration cut-offs of dsDNA 0.61 ng/μl, ssDNA 1.38 ng/μl, and miRNA 2.4 ng/μl with 81% to 90% accuracy, where concentrations greater than these cut-offs correlated with higher disease burden. Thus, at diagnosis, in the setting of a diagnostic dilemma, analyte concentration alone may be useful, along with facilitating specific genetic and genomic assays. However, eyes that were undergoing therapy and eyes at the end of RB therapy were not distinguishable with nucleic acid concentrations alone in this sample.
What are the implications of this data? Our group and others have already demonstrated that the AH cfDNA can be utilized to identify RB1 SNVs, SCNAs, tumor methylation status, and to estimate tumor fraction.2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20 However, as the AH liquid biopsy moves from an area of bench research to one of standard-of-care clinical assays, the expected ranges of analytes at different clinical time points will impact the kind of analyses that can be done. For example, next-generation sequencing for SNV analysis of genetic mutations requires target enrichment and a higher DNA input in any sample (whether AH or another liquid biopsy source). Thus, only treatment naive AH samples, with a median 3.08 ng/μl, may routinely provide enough DNA for next-generation sequencing RB1 gene mutation analysis. Le Gall et al16 also reported that AH DNA concentration was too low to perform gene analysis in a patient after several intravitreal melphalan treatments. While this technology continues to improve, the current clinical industry standard for SNV analysis requires > 20 ng of dsDNA input6 which can be found in as little as 10 μl from a DX sample or require > 100 μl during therapy. However, current laboratory developed tests for research require lower concentrations, demonstrating that reliable results are possible with lower inputs. During treatment, the concentration of DNA in the AH may or may not be adequate for RB1 mutation analysis, but at diagnosis, it is adequate in nearly all cases.
However, that is not to say there is no utility of AH analysis at other clinical time points. While a lower concentration of AH cfDNA is found in samples taken during RB treatment (e.g., dsDNA, with a median concentration of 0.18 ng/μl), this amount is sufficient to perform genomic analysis via low-pass whole-genome sequencing,15 as seen in Figure 5B. The advantage of SCNA identification is multifold. First, SCNA analysis requires a lower DNA concentration than SNV analysis, so it can be done even when the tumor burden is low25. Furthermore, SCNAs can also be utilized to determine tumor fraction, which correlates to intraocular disease activity.29
Aside from establishing expected concentrations of analytes at various clinical time points, we also wanted to evaluate whether evaluation of AH concentration alone could help indicate that a tumor is active. While ocular oncologists currently do this based on clinical examination, this is significantly hampered if there is a loss of view to the back of the eye. This can happen due to various reasons including vitreous hemorrhage, and anti-VEGF agents are often given intravitreally 30. Although BEV AH samples were thought to not have clinically active RB, we predicted that the presence of vitreous hemorrhage secondary to radiation retinopathy would affect analyte concentrations. As hypothesized, TX and BEV AH samples had similar analyte concentrations (dsDNA: TX, 0.104 ng/μl; BEV 0.192 ng/μl), but these values are not significantly different from the median dsDNA concentration observed in AH samples from patients with PRD (0.74 ng/μl; see Table 3). Thus, in the setting of loss of view due to vitreous hemorrhage, AH analyte concentrations alone cannot indicate whether the RB tumor is active in the eye, except for a large tumor at diagnosis. However, genomic analysis of the AH may provide additional objective evidence of the resolution of the RB tumor. For example, in the same eye at different clinical time points and with similar analyte concentrations between the samples, there were no SCNAs identified in the BEV AH, while the TX AH had positive RB SCNAs 6p gain, and 16q loss4,7,8,28 (Fig 5A–C). We suggest that the positive RB SCNAs reflect the presence of active RB tumor, as we expected in an eye undergoing treatment (Fig 5B). Thus, identification of SCNA via low-pass whole-genome sequencing analysis may provide additional disease status information beyond analyte concentration, especially in the presence of vitreous hemorrhage.
A limitation in this report is our small sample size from each disease burden group, since a larger cohort would produce a better powered model. Further investigations with larger sample sizes from multiple centers should be pursued for further validation of our results.
In summary, AH liquid biopsy at the time of diagnosis, prior to any therapy, provides the highest nucleic acid and protein concentrations, and may be most useful for RB1 mutational testing. Aqueous humor biopsy during therapy can also yield high enough dsDNA concentration to perform low-pass whole-genome sequencing and SCNA detection to facilitate prognosis and trends in tumor fraction which correlate with ocular outcomes.7, 8, 9 Nucleic acid quantifications can stratify high- versus low-disease burden eyes and higher analyte concentrations correlated with increasing IIRC group and disease burden; however, it was not possible to reliably distinguish low-disease burden from eyes without active disease (TX vs. BEV/END), which may be complicated by vitreous hemorrhage. Aqueous humor liquid biopsy is a high-yield source of nucleic acids and proteins in RB specifically at diagnosis. Evaluating AH analyte quantifications will facilitate potential applications of clinical assays.
Acknowledgments
The authors would like to acknowledge Brianne Brown for the coordination of this study.
Manuscript no. XOPS-D-22-00267.
Footnotes
Disclosures:
All authors have completed and submitted the ICMJE disclosures form.
The authors made the following disclosures: J.LB. and L.X.: Patent application entitled: Aqueous humor cell free DNA for diagnostic and prognostic evaluation of ophthalmic disease.
J.L.B.: Research support – National Cancer Institute of the National Institute of Health Award Number K08CA232344, The Wright Foundation, Children’s Oncology Group/ St. Baldrick’s Foundation, Danhakl Family Foundation, Hyundai Hope on Wheels, Childhood Eye Cancer Trust, Children’s Cancer Research Fund, A. Linn Murphree, MD, Chair in Ocular Oncology, The Berle & Lucy Adams Chair in Cancer Research, The Larry and Celia Moh Foundation, The Institute for Families, Inc, Children's Hospital Los Angeles.
L.X.: Research support – The Knights Templar Eye Foundation.
A.N.: Research support – NIH K08EY030924, the Las Madrinas Endowment in Experimental Therapeutics for Ophthalmology, Research to Prevent Blindness Career Development Award, Knights Templar Eye Foundation Endowment.
H.E.G.: Research support – NIH R01 EY028450.
G.B.H.: Research support – the National Eye Institute as Protocol Co-Chair for studies on retinopathy of prematurity (ROP3 and ROP4 Trials).
The sponsors or funding organizations had no role in the design or conduct of this research.
Drs Alison Skalet and Kellyn Bellsmith are supported by grant P30 EY010572, from the National Institutes of Health (Bethesda, MD), and by unrestricted departmental funding from Research to Prevent Blindness (New York, NY).
HUMAN SUBJECTS: Human Subjects were used in this study. Institutional Review Board approval was obtained at Children's Hospital Los Angeles, University of Southern California (IRB CHLA-17-00248). The study was conducted in accordance with the Declaration of Helsinki. Written informed consent was obtained from the parents of all participants and that included permission for publication.
No animal subjects were used in this study.
Author Contributions:
Conception and design: Xu, Berry
Analysis and interpretation: Im, Pike, Reid, Peng, Xu, Berry
Data Collection: Im, Pike, Peng, Grossniklaus, Hubbard, Skalet, Bellsmith, Shields, Lally, Stacey, Reiser, Nagiel, Shah, Xu, Berry
Obtained funding: N/A
Overall responsibility: Im, Pike, Reid, Peng, Sirivolu, Grossniklaus, Hubbard, Skalet, Bellsmith, Shields, Lally, Stacey, Reiser, Nagiel, Shah, Xu, Berry
References
- 1.Karcioglu Z.A., Gordon R.A., Karcioglu G.L. Tumor seeding in ocular fine needle aspiration biopsy. Ophthalmology. 1985;92:1763–1767. doi: 10.1016/s0161-6420(85)34105-2. [DOI] [PubMed] [Google Scholar]
- 2.Berry J.L., Xu L., Murphree A.L., et al. Potential of aqueous humor as a surrogate tumor biopsy for retinoblastoma. JAMA Ophthalmol. 2017;135:1221–1230. doi: 10.1001/jamaophthalmol.2017.4097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Berry J.L., Cobrinik D., Hicks J. Potential of aqueous humor as a surrogate tumor biopsy for retinoblastoma-reply. JAMA Ophthalmol. 2018;136:598. doi: 10.1001/jamaophthalmol.2018.0395. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Berry J.L., Xu L., Kooi I., et al. Genomic cfDNA analysis of aqueous humor in retinoblastoma predicts eye salvage: the surrogate tumor biopsy for retinoblastoma. Mol Cancer Res. 2018;16:1701–1712. doi: 10.1158/1541-7786.MCR-18-0369. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Kim M.E., Polski A., Xu L., et al. Comprehensive somatic copy number analysis using aqueous humor liquid biopsy for retinoblastoma. Cancers (Basel) 2021;13:3340. doi: 10.3390/cancers13133340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Xu L., Shen L., Polski A., et al. Simultaneous identification of clinically relevant RB1 mutations and copy number alterations in aqueous humor of retinoblastoma eyes. Ophthalmic Genet. 2020;41:526–532. doi: 10.1080/13816810.2020.1799417. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Xu L., Polski A., Prabakar R.K., et al. Chromosome 6p amplification in aqueous humor cell-free DNA is a prognostic biomarker for retinoblastoma ocular survival. Mol Cancer Res. 2020;18:1166–1175. doi: 10.1158/1541-7786.MCR-19-1262. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Xu L., Kim M.E., Polski A., et al. Establishing the clinical utility of ctDNA analysis for diagnosis, prognosis, and treatment monitoring of retinoblastoma: the aqueous humor liquid biopsy. Cancers (Basel) 2021;13:1282. doi: 10.3390/cancers13061282. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Polski A., Xu L., Prabakar R.K., et al. Cell-free DNA tumor fraction in the aqueous humor is associated with therapeutic response in retinoblastoma patients. Transl Vis Sci Technol. 2020;9:30. doi: 10.1167/tvst.9.10.30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Li H.T., Xu L., Weisenberger D.J., et al. Characterizing DNA methylation signatures of retinoblastoma using aqueous humor liquid biopsy. Nat Commun. 2022;13:5523. doi: 10.1038/s41467-022-33248-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Liu J., Ottaviani D., Sefta M., et al. A high-risk retinoblastoma subtype with stemness features, dedifferentiated cone states and neuronal/ganglion cell gene expression. Nat Commun. 2021;12:5578. doi: 10.1038/s41467-021-25792-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Wong E.Y., Xu L., Shen L., et al. Inter-eye genomic heterogeneity in bilateral retinoblastoma via aqueous humor liquid biopsy. NPJ Precis Oncol. 2021;5:73. doi: 10.1038/s41698-021-00212-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Berry J.L., Xu L., Polski A., et al. Aqueous humor is superior to blood as a liquid biopsy for retinoblastoma. Ophthalmology. 2020;127:552–554. doi: 10.1016/j.ophtha.2019.10.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Ghose N., Kaliki S. Liquid biopsy in retinoblastoma: a review. Semin Ophthalmol. 2022;37:813–819. doi: 10.1080/08820538.2022.2078165. [DOI] [PubMed] [Google Scholar]
- 15.Gerrish A., Stone E., Clokie S., et al. Non-invasive diagnosis of retinoblastoma using cell-free DNA from aqueous humour [published correction appears in Br J Ophthalmol. 2020 Mar;104(3):415-416] Br J Ophthalmol. 2019;103:721–724. doi: 10.1136/bjophthalmol-2018-313005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Le Gall J., Dehainault C., Benoist C., et al. Highly sensitive detection method of retinoblastoma genetic predisposition and biomarkers. J Mol Diagn. 2021;23:1714–1721. doi: 10.1016/j.jmoldx.2021.08.014. [DOI] [PubMed] [Google Scholar]
- 17.Raval V., Racher H., Wrenn J., Singh A.D. Aqueous humor as a surrogate biomarker for retinoblastoma tumor tissue. J AAPOS. 2022;26:137.e1–137.e5. doi: 10.1016/j.jaapos.2022.03.005. [DOI] [PubMed] [Google Scholar]
- 18.Gerrish A., Jenkinson H., Cole T. The impact of cell-free DNA analysis on the management of retinoblastoma. Cancers (Basel) 2021;13:1570. doi: 10.3390/cancers13071570. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Liu W., Luo Y., Dai J., et al. Monitoring retinoblastoma by machine learning of aqueous humor metabolic fingerprinting. Small Methods. 2022;6 doi: 10.1002/smtd.202101220. [DOI] [PubMed] [Google Scholar]
- 20.Cancellieri F., Peter V.G., Quinodoz M., et al. Genetic bases of retinoblastoma from liquid biopsies. Invest Ophthalmol Vis Sci. 2022;63:503. A0080. [Google Scholar]
- 21.Alfaar A.S., Chantada G., Qaddoumi I. Survivin is high in retinoblastoma, but what lies beneath? J AAPOS. 2018;22:482. doi: 10.1016/j.jaapos.2017.02.021. [DOI] [PubMed] [Google Scholar]
- 22.Busch M.A., Haase A., Miroschnikov N., et al. TFF1 in aqueous humor-a potential new biomarker for retinoblastoma. Cancers (Basel) 2022;14:677. doi: 10.3390/cancers14030677. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Ghiam B.K., Xu L., Berry J.L. Aqueous humor markers in retinoblastoma, a review. Transl Vis Sci Technol. 2019;8:13. doi: 10.1167/tvst.8.2.13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Wierenga A.P.A., Cao J., Mouthaan H., et al. Aqueous humor biomarkers identify three prognostic groups in uveal melanoma. Invest Ophthalmol Vis Sci. 2019;60:4740–4747. doi: 10.1167/iovs.19-28309. [DOI] [PubMed] [Google Scholar]
- 25.Kim M.E., Xu L., Prabakar R.K., et al. Aqueous humor as a liquid biopsy for retinoblastoma: clear corneal paracentesis and genomic analysis. J Vis Exp. 2021 doi: 10.3791/62939. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Baslan T., Kendall J., Rodgers L., et al. Genome-wide copy number analysis of single cells [published correction appears in Nat Protoc. 2016 Mar;11(3):616] Nat Protoc. 2012;7:1024–1041. doi: 10.1038/nprot.2012.039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Sirivolu S., Xu L., Warren M., et al. Chromosome 6p amplification detected in blood cell-free DNA in advanced intraocular retinoblastoma. Ophthalmic Genet. 2022;43:866–870. doi: 10.1080/13816810.2022.2142246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Kooi I.E., Mol B.M., Massink M.P., et al. A meta-analysis of retinoblastoma copy numbers refines the list of possible driver genes involved in tumor progression. PLoS One. 2016;11 doi: 10.1371/journal.pone.0153323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Polski A., Xu L., Prabakar R., et al. Longitudinal aqueous humor sampling reflects treatment response in retinoblastoma patients. Invest Ophthalmol Vis Sci. 2020;61:1394. [Google Scholar]
- 30.Stathopoulos C., Gaillard M.C., Moulin A., et al. Intravitreal anti-vascular endothelial growth factor for the management of neovascularization in retinoblastoma after intravenous and/or intraarterial chemotherapy: long-term outcomes in a series of 35 eyes. Retina. 2019;39:2273–2282. doi: 10.1097/IAE.0000000000002339. [DOI] [PubMed] [Google Scholar]