Abstract
Proper anther dehiscence is essential for successful pollination and reproduction in angiosperms, and jasmonic acid (JA) is crucial for the process. However, the mechanisms underlying the tight regulation of JA biosynthesis during anther development remain largely unknown. Here, we demonstrate that the rice (Oryza sativa L.) ethylene-response factor-associated amphiphilic repression (EAR) motif-containing protein TEOSINTE BRANCHED1/CYCLOIDEA/PROLIFERATING CELL FACTORS (TCP) INTERACTOR CONTAINING EAR MOTIF PROTEIN1 (OsTIE1) tightly regulates JA biosynthesis by repressing TCP transcription factor OsTCP1/PCF5 during anther development. The loss of OsTIE1 function in Ostie1 mutants causes male sterility. The Ostie1 mutants display inviable pollen, early stamen filament elongation, and precocious anther dehiscence. In addition, JA biosynthesis is activated earlier and JA abundance is precociously increased in Ostie1 anthers. OsTIE1 is expressed during anther development, and OsTIE1 is localized in nuclei and has transcriptional repression activity. OsTIE1 directly interacts with OsTCP1, and overexpression of OsTCP1 caused early anther dehiscence resembling that of Ostie1. JA biosynthesis genes including rice LIPOXYGENASE are regulated by the OsTIE1-OsTCP1 complex. Our findings reveal that the OsTIE1-OsTCP1 module plays a critical role in anther development by finely tuning JA biosynthesis and provide a foundation for the generation of male sterile plants for hybrid seed production.
A transcriptional repressor interacts with a transcription factor to modulate rice anther dehiscence and male sterility by finely regulating jasmonic acid biosynthesis during anther development.
IN A NUTSHELL.
Background: Anthers are crucial for pollen development and storage. The success of plant reproduction depends on anthers opening and pollen being released at the appropriate time. Jasmonic acid (JA) is a key plant hormone that promote anther dehiscence. In the staple food crop rice, the JA concentration is low at the early anther developmental stage but increases at the late developmental stage to promote anther opening.
Question: How is JA concentration finely regulated during rice anther development?
Findings: This study identifies a transcriptional regulatory module consisting of the transcriptional repressor OsTIE1 and transcription factor OsTCP1 that tightly regulates JA biosynthesis during anther development. The disruption of OsTIE1 function in the Ostie1 mutant led to anthers opening prematurely, before pollen maturation. OsTIE1 physically interacted with the transcription factor OsTCP1, which then bound directly to the promoter of JA biosynthesis genes to regulate their expression. Consistently, the JA concentration in the early anthers of the Ostie1 mutant was elevated, and inhibiting JA biosynthesis partially rescued the early anther opening of Ostie1 mutant. Overexpression of OsTCP1 resulted in the early anther opening phenotype, similar to that observed in the Ostie1 mutant.
Next steps: The OsTIE1-OsTCP1 module could be further investigated to create phytohormone genic male-sterile lines for hybrid seed production.
Introduction
In angiosperms, the stamen is the male reproductive organ, consisting of a basal filament and an apical anther (Goldberg et al. 1993; Marchant and Walbot 2022). The anther contains 2 thecae, each of which is composed of 2 locules connected by the septum and stomium (Matsui et al. 1999; Wilson et al. 2011). The stamen begins development from the initiation of the third whorl organ primordium in the flower meristem, after which it goes through 14 developmental stages to form a functional anther with mature pollen and a filament length comparative to that of the pistil (Zhang et al. 2011; Guo and Liu 2012). Proper coordination of the final steps in late stamen developmental stages, including pollen maturation, appropriate filament elongation, and anther dehiscence, is vital for successful pollination and plant reproduction, and disruption of these steps causes male sterility (Wilson et al. 2011). Consequently, a better understanding of the mechanisms controlling these processes should facilitate the generation of male sterility lines that could be used for the heterosis of hybrid plants (Pak et al. 2020; Abbas et al. 2021).
The phytohormone jasmonic acid (JA) plays a central role in coordinating pollen maturation, filament elongation, and anther dehiscence (Wilson et al. 2011; Ghorbel et al. 2021). JA biosynthesis is initiated in plastids by the lipase DEFECTIVE IN ANTHER DEHISCENCE1 (DAD1), which produces α-linolenic acid (Ishiguro et al. 2001). α-Linolenic acid is oxygenated into 13(S)-hydroperoxy-octadecatrienoic acid (13-HPOD) by the 13-lipoxygenases (LOXs) (Ruan et al. 2019). ALLENE OXIDE SYNTHASE (AOS) then converts 13-HPOD into (13S)-12,13-epoxy-octadecatrienoic acid, which is subsequently catalyzed into 12-oxo-phytodienoic acid (OPDA) by ALLENE OXIDE CYCLASE (AOC) (Ruan et al. 2019). OPDA is transported into peroxisomes, in which an OPDA reductase (OPR3) and 3 sequential β-oxidations finally convert it into JA (Schaller et al. 2000). JA is conjugated with isoleucine (Ile) to form bioactive jasmonoyl-L-isoleucine (JA-Ile) by the JA-amido synthetase JASMONATE RESISTANT1 in the cytosol (Riemann et al. 2008; Xiao et al. 2014). In JA signaling, JA-Ile binds to the receptor CORONATINE INSENSITIVE1 (COI1), an F-box protein, which is part of the Skp1-Cullin1-F-box (SCF) protein ligase complex. The binding of JA to COI1 causes SCFCOI1 E3 ligase to target the transcriptional repressor JASMONATE-ZIM-DOMAIN (JAZ) for degradation (Xie et al. 1998; Thines et al. 2007). Finally, MYC transcription factors repressed by JAZ are released, allowing them to regulate gene expression, leading to transcriptome changes and the resulting biological effects of JA (Lorenzo et al. 2004).
In the past few decades, genetic analyses have identified several components of JA biosynthesis and signaling that are essential for stamen development in Arabidopsis (Arabidopsis thaliana). For example, in Arabidopsis, disruption of JA biosynthesis in the mutants dad1 and opr3, as well as the double mutant lox3 lox4, causes male sterility due to inviable pollen, defective filament elongation, and indehiscent anthers (Stintzi and Browse 2000; Ishiguro et al. 2001; Caldelari et al. 2011). Compromised JA signaling in the coi1 mutant leads to defective stamen development resembling that of dad1, opr3, or lox3 lox4, except that male sterility is rescued in the mutants with disrupted JA biosynthesis by JA treatment, but not in coi1 (Thines et al. 2007; Jewell and Browse 2016). MYC transcription factors play a redundant role in controlling stamen development. Single and double myc mutants have no obvious phenotype, while triple myc mutants and the quadruple mutant myc2 myc3 myc4 myc5 display defects in pollen viability, filament elongation, and anther dehiscence (Qi et al. 2015). MYC5 interacts with R2R3 MYB transcription factors MYB21 and MYB24 to form basic helix-loop-helix (bHLH)-MYB transcriptional complexes to regulate stamen development, and disruption of MYBs leads to defects in late stamen development similar to those observed in myc multiple mutants (Figueroa and Browse 2015; Qi et al. 2015). These findings demonstrate that JA is critical for male fertility and stamen development. However, the mechanisms underlying the regulation of JA biosynthesis and signaling during stamen development remain to be clarified.
Several streams of evidence indicate that the phytohormone auxin acts as a negative regulator of JA biosynthesis at late stages of stamen development (Cecchetti et al. 2008; Wilson et al. 2011; Acosta and Przybyl 2019). First, in Arabidopsis, disruption of auxin receptor genes TRANSPORT INHIBITOR RESPONSE 1 (TIR1), AUXIN SIGNALLING F-BOX 1 (AFB1), AFB2, and AFB3 in various triple mutants leads to earlier pollen maturation and precocious anther dehiscence before full elongation of filaments, similar to phenotype of the tir1 afb1 afb2 afb3 quadruple mutant (Cecchetti et al. 2013). Expression of JA biosynthesis genes such as OPR3 and DAD1 is repressed by auxin treatment of flowers, while it is promoted in auxin receptor mutants (Cecchetti et al. 2013). Second, mutations in auxin signal components, including INDOLE-3-ACETIC ACID INDUCIBLE 19 (IAA19), IAA8, AUXIN RESPONSE FACTOR 6 (ARF6), and ARF8, produce defects in stamen maturation (Tashiro et al. 2009; Tabata et al. 2010; Wang et al. 2013). For example, the arf6 arf8 double mutant produces unelongated filaments and unopened flower buds with protruding pistils (Tabata et al. 2010). The expression of DAD1 is downregulated in arf6 arf8, and induction of the splice variant ARF8.2, but not ARF8.4, increases the expression level of DAD1, indicating that ARF8.2 is required for the regulation of JA biosynthesis during stamen maturation (Ghelli et al. 2018). Third, in rice, increased abundance of auxin during stamen development leads to anther indehiscence by decreasing the JA level. For example, DIOXYGENASE FOR AUXIN OXIDATION (DAO) encodes an enzyme that converts indole-3-acetic acid (IAA) into 2-oxoindole-3-acetic acid (OxIAA), inactivating IAA. Disruption of DAO in the dao mutant, overexpression of auxin biosynthesis gene OsYUCCA1 (OsYUC1), and application of IAA all cause anther indehiscence and inviable pollen in rice (Zhao et al. 2013). The level of auxin during anther development is tightly controlled by the FT-INTERACTING PROTEIN 7 (OsFTIP7)-O. sativa homeobox 1 (OSH1) module. In this module, the expression of OsFTIP7 in anthers facilitates translocation of Class I KNOTTED1-LIKE HOMEOBOX transcription factor OSH1 into nuclei, and nucleus-localized OSH1 in turn represses transcription of OsYUC4 to decrease the auxin level for anther dehiscence by increasing JA biosynthesis (Song et al. 2018). These findings indicate that negative regulation of JA biosynthesis by auxin in anthers is conserved in monocots and dicots. Until now, few transcription factors have been shown to regulate JA biosynthesis. By inducing the function of the homeotic gene AGAMOUS (AG) in the ag mutant over a time course, AG was found to play an essential role in early stamen initiation and late stamen maturation in Arabidopsis (Ito et al. 2007). AG is expressed at the late stage of stamen development, and AG promotes JA biosynthesis by directly binding to the promoter region of DAD1. During late stages of leaf development, the TEOSINTE BRANCHED1/CYCLOIDEA/PROLIFERATING CELL FACTORS (TCP) transcription factor family TCP4 directly upregulates expression of LOX2 to enhance JA biosynthesis during leaf senescence in Arabidopsis (Schommer et al. 2008; Danisman et al. 2012). However, the roles of TCP transcription factors in control of JA biosynthesis during anther development remain unclear.
The ethylene-response factor-associated amphiphilic repression (EAR) motif consists of a conserved short sequence: LXLXL (L indicates leucine and X represents any amino acid; Ohta et al. 2001; Kagale and Rozwadowski 2011). The EAR motif is a well-known plant-specific repression domain identified in more than 200 Arabidopsis transcriptional regulators (Hiratsu et al. 2003; Kagale and Rozwadowski 2011). Many putative orthologs of these Arabidopsis EAR motif-containing proteins are found in the rice genome (Kagale et al. 2010). EAR motif-containing proteins play pivotal roles in diverse biological processes, including the signaling pathways of auxin, JA and other hormones, leaf and floral organ development, and biotic and abiotic resistance (Szemenyei et al. 2008; Tao et al. 2013; Plant et al. 2021; Chow et al. 2023). We previously demonstrated that Arabidopsis EAR motif-containing protein TCP INTERACTOR CONTAINING EAR MOTIF PROTEIN1 (TIE1) regulates leaf development and shoot branching by connecting TCP transcription factors with TOPLESS (TPL)/TPL-RELATED PROTEIN corepressors (Tao et al. 2013, Yang et al. 2018). TIE1 has a close putative ortholog known as OsTIE1 (LOC_Os01g11430) in rice. However, the function of OsTIE1 remains unknown.
In this study, we generated rice Ostie1 mutants using CRISPR/Cas9 technology and found that OsTIE1 plays critical roles in controlling stamen development. The Ostie1 mutants display male infertility and produce inviable pollen. The anthers of Ostie1 mutants exhibit precocious dehiscence due to elevated JA abundance. OsTIE1 is a transcriptional repressor that interacts with the TCP transcription factor OsTCP1/PCF5. Overexpression of OsTCP1/PCF5 in stamens causes male fertility and defective anther dehiscence similar to that observed in Ostie1 mutants. We further demonstrate that OsTCP1/PCF5 activates the expression of OsLOX genes and OsTIE1 represses them. Therefore, our findings reveal the important roles of OsTIE1 in rice stamen development, as well as the existence of an OsTIE1-OsTCP1 transcriptional regulation module that finely tunes JA biosynthesis during rice stamen development.
Results
Disruption of OsTIE1 causes male sterility in rice
We previously identified Arabidopsis TIE1, which encodes an EAR motif-containing transcriptional repressor that plays pivotal roles in controlling leaf development (Tao et al. 2013), shoot branching (Yang et al. 2018), and root elongation (He et al. 2022). The rice LOC_Os01g11430 is a close putative ortholog to Arabidopsis TIE1, and we named it OsTIE1 (Supplementary Fig. S1A and Files S1 and S2). OsTIE1 encodes a protein containing 345 amino acid residues (Supplementary Fig. S1B). OsTIE1 contains an N-terminal nuclear localization signal with a helix region and a C-terminal EAR motif (Supplementary Fig. S1, B and C; Fig. 1A). To elucidate the function of OsTIE1, we first generated 35Spro:OsTIE1, in which OsTIE1 was driven by the cauliflower mosaic virus (CaMV) 35S promoter and transformed 35Spro:OsTIE1 into Arabidopsis. The resulting plants with overexpression of OsTIE1 had increased numbers of shoot branches and small leaves, as displayed by the Arabidopsis TIE1 overexpression lines (Yang et al. 2018), suggesting that OsTIE1 may have a biological function similar to that of Arabidopsis TIE1 (Fig. 1B). We then knocked out OsTIE1 by generating Ostie1-1 and Ostie1-2 in rice cultivar Nipponbare using CRISPR/Cas9 technology (Miao et al. 2013, Ma et al. 2015). The Ostie1-1 and Ostie1-2 mutants each carried a different 1-bp insert in the first exon of OsTIE1 (Supplementary Fig. S2). The two 1-bp inserts each caused a shift of the ORF and thus disrupted the function of OsTIE1 in Ostie1-1 or Ostie1-2 (Supplementary Fig. S2). Although Ostie1-1 and Ostie1-2 displayed no obvious phenotypes at the vegetative stage, both mutants showed defective fertility at the reproductive stage and very few seeds were harvested (Supplementary Fig. S3A; Fig. 1, C and D). To determine whether male or female fertility was affected in the Ostie mutants, we performed a reciprocal cross between wild-type (WT) Nipponbare and Ostie1-1 or Ostie1-2. When WT pollen was used to pollinate the pistils of Ostie1-1 or Ostie1-2, the seed-setting rate was comparable to that observed following self-crossing of WT Nipponbare (Fig. 1, E and F). However, when the pollen of Ostie1-1 or Ostie1-2 was used to pollinate WT pistils, extremely few seeds were harvested (Fig. 1, E and F), indicating that male fertility was compromised in Ostie1, but female fertility was not. These results indicate that OsTIE1 is critical for male fertility in rice.
Figure 1.
OsTIE1 is a key factor in maintaining rice male fertility. A) Conserved sequence alignment between OsTIE1 and TIE1. The same amino acid residues are marked with a star sign, while amino acid residues of the same type are marked with a colon. B) Heterologous expression of OsTIE1 in Arabidopsis. Scale bar: 5 cm. C) WT, Ostie1-1, and Ostie1-2 plants at the pustulation period. The mutant plants produced few seeds. Scale bar: 10 cm. D) Statistical analysis of the seed-setting rate of the WT, Ostie1-1, and Ostie1-2. Data are expressed as mean (±Sd) (n ≥ 4). The chi-square test was used to compare the seed-setting rate of different phenotypes. ***P < 0.001. E) Reciprocal cross between WT and mutant alleles. Scale bar: 2 cm. F) Statistical analysis of crossing combination. WT♀×WT♂ was used as the control. The seed-setting rate exhibited no significant difference from that of the control when the mutant was used as the female, while plants were completely sterile when the mutant was used as the pollen donor. Data are expressed as mean (±Sd) (n ≥ 3). The chi-square test was used to compare the seed-setting rate of different cross groups. ***P < 0.001.
Anther dehiscence, filament elongation, and pollen viability are defective in Ostie1 mutants
To reveal the mechanism underlying male sterility in Ostie1 mutants, we investigated the process of anther development. The morphology of Ostie1-1 florets had no obvious differences from that of the WT control (Supplementary Fig. S3B). However, some of the anthers were found to be dehiscent at developmental Stage 12, when the florets were still unopened, in Ostie1-1 and Ostie1-2 mutants (Fig. 2, A to H). Pollen grains were clearly observed on the pistils in the closed florets of Ostie1 mutants (Fig. 2, B and H). No anthers were dehiscent and no released pollen grains were observed in the unopened florets of the WT control at developmental Stage 12 (Fig. 2, A, E, and G). In the Ostie1 mutants, approximately 20% to 30% of anthers were precociously dehiscent (Fig. 2I). The filaments of the stamens from Ostie1 mutants were significantly longer, while the anthers of Ostie1 mutants were shorter than those of the WT control at Stage 12 (Fig. 2, J and K). These results clearly showed that the stamens of Ostie1 mutants underwent precocious filament elongation and anther dehiscence before maturation.
Figure 2.
The Ostie1-1 and Ostie1-2 mutants displayed premature anther dehiscence. A to D) Florets were observed at Stage 12 with a stereomicroscope. Ostie1-1 and Ostie1-2 showed premature anther dehiscence compared to the WT. Scale bar: 1 mm. E, F) SEM observation of the anthers of the WT and Ostie1-1 at Stage 12. Scale bar: 100 μm. G, H) SEM observation of the pistils of the WT and Ostie1-1 at Stage 12. Pollen grains marked with arrows were prematurely released and adhered to pistils at Stage 12 in Ostie1-1. Scale bar: 200 μm. I) Statistical analysis of the anther dehiscence rate at Stage 12 and Stage 13. Data are expressed as mean (±Se) (n ≥ 5). Two-tailed t-test was applied for comparisons between groups. ***P < 0.001. J) Comparison of WT, Ostie1-1, and Ostie1-2 anthers at Stage 12. Scale bar: 1 mm. K) Statistical analysis of the length of filaments and anthers in the WT, Ostie1-1, and Ostie1-2 at Stage 12. Data are expressed as the mean (±Sd) (n ≥ 100). Fisher's protected least significant difference was applied for multiple comparisons. Means were judged significantly different at P < 0.05. L, M) Cross-section of anthers at Stage 12. Anthers were prematurely dehisced in Ostie1-1. Scale bar: 100 μm. N, O) I2-KI staining of pollen grains at Stage 12. Pollen grains were inviable in Ostie1-1. Scale bar: 100 μm.
To examine the effects of the prematuration of anthers on pollen development in Ostie1 mutants, we performed a series of transverse sections of anthers at different developmental stages. The pollen development of WT and Ostie1-1 plants displayed no obvious differences before anther developmental Stage 12 (Supplementary Fig. S4, A to H). However, WT microspores became spherical at Stage 12 (from the falcate shape observed at Stage 11), while Ostie1-1 microspores maintained their falcate shape at Stage 12 (Supplementary Fig. S4, I to K). Transverse sections also showed that Ostie1-1 anthers were dehiscent at Stage 12 (Fig. 2, L and M). I2-KI staining showed that pollen grains from Ostie1-1 were stained less intensely in comparison with WT pollen grains (Fig. 2, N and O). Transmission electron microscopy (TEM) of pollen grains showed that WT grains were round and accumulated many starch granules, while Ostie1-1 pollen grains had a falcate shape with a large vacuole and lacked starch granules (Supplementary Fig. S4K). These observations suggest that pollen maturation and viability were disrupted in Ostie1 mutants, indicating that OsTIE1 plays essential roles in coordinating pollen maturation, filament elongation, and anther dehiscence.
OsTIE1 is predominantly expressed in anthers and encodes a nucleus-localized transcriptional repressor
To investigate the spatial and temporal expression pattern of OsTIE1, we first performed reverse transcription quantitative PCR (RT-qPCR) to measure the expression level of OsTIE1 in the anthers at different developmental stages. OsTIE1 began to be detected at Stage 8 (Fig. 3A), and OsTIE1 transcript abundance increased as anthers developed. The expression level of OsTIE1 was similar from Stage 9 to Stage 11 and increased markedly at Stage 12 (Fig. 3A). We then generated an OsTIE1pro:GUS construct by cloning a 3,000-bp-long promoter upstream of the OsTIE1 coding region and using it to drive GUS reporter gene expression. We transformed OsTIE1pro:GUS into the rice cultivar Nipponbare. Five randomly selected OsTIE1pro:GUS transgenic lines showed a similar GUS staining pattern. Although GUS staining was not observed in anther primordia and early anthers, GUS expression began to be detected in the anthers at Stage 8 (Fig. 3, B and C), and GUS staining was clearly observed in the anthers after Stage 9 (Fig. 3D). The GUS signal was similar from Stage 9 to Stage 11 and increased markedly in the anthers at Stage 12 (Fig. 3, E to G). The GUS staining patterns of the OsTIE1pro:GUS transgenic lines were consistent with the RT-qPCR results for OsTIE1. These results suggest that OsTIE1 is regulated during anther development, and the expression pattern is consistent with the male sterility observed in Ostie1 mutants, providing further evidence that OsTIE1 is critical for anther development.
Figure 3.
OsTIE1 is a nucleus-localized transcriptional repressor mainly expressed in anthers after Stage 9. A) RT-qPCR analysis of OsTIE1 expression during anther development. The ordinate represents the ratio of the gene expression level to that of the internal control OsUBQ. Data are expressed as the mean (±Sd) of 3 biological replicates. B to G) Tissue-specific staining with GUS driven by the OsTIE1 promoter. The signal was consistent with the results of the real-time assays. Scale bar: 1 mm. H, I) OsTIE1 was colocalized with FIB2, which was used as a nuclear marker in N. benthamiana and rice protoplasts. Scale bar: 50 μm. J) OsTIE1 is a transcriptional repressor. REN LUC was used as the internal control. The ratio of the luminescent signals from firefly LUC and REN LUC was calculated. OsTIE1△EAR indicates OsTIE1 without the EAR-motif. Data are expressed as the mean (±Sd) of 3 biological replicates. Two-tailed t-test was used for comparisons between groups. **P < 0.01.
OsTIE1 contains an EAR repression domain at its C-terminal end, and overexpression of OsTIE1 in Arabidopsis led to increased shoot branches, similar to the effect of overexpressing Arabidopsis TIE1 (Fig. 1B) (Yang et al. 2018). To determine whether OsTIE1 could act as a transcriptional repressor in a manner similar to Arabidopsis TIE1, we began by determining the subcellular localization of OsTIE1. We fused the GFP reporter gene to OsTIE1 and drove the fusion product with the CaMV 35S promoter to generate 35Spro:GFP-OsTIE1. 35Spro:RFP-FIBRILLARIN 2 (FIB2) was generated as a nuclear marker by using the CaMV 35S promoter to drive RFP fusion with a sequence encoding nuclear protein FIB2 (Degenhardt and Bonham-Smith 2008). We first cotransformed 35Spro:GFP-OsTIE1 with the marker 35Spro:RFP-FIB2 in Nicotiana benthamiana leaves, which produced clearly overlapping GFP and RFP fluorescence in the nuclei (Fig. 3H). When we coexpressed 35Spro:GFP-OsTIE1 and 35Spro:RFP-FIB2 in rice protoplasts, the GFP signal was colocalized with that of RFP, consistent with the results in N. benthamiana leaves (Fig. 3I). These results suggest that OsTIE1 was localized to nuclei and may have acted as a transcriptional regulator. To test the transcriptional repression activity of OsTIE1, we generated the reporter construct 35Spro-UAS:LUC-35Spro:REN, in which the firefly luciferase (LUC) gene was driven by the CaMV 35S promoter fused with 6 GAL4-binding sites (upstream activating sequence [UAS]), and the renilla (REN) LUC gene was driven by the CaMV 35S promoter as an internal control (Fig. 3J). The effectors, including the control 35Spro:DNA-binding domain (DBD), 35Spro:DBD-OsTIE1 and 35Spro:DBD-OsTIE1ΔEAR, were generated using the CaMV 35S promoter to drive a sequence encoding a GAL4 DBD, DBD fused with OsTIE1, or DBD fused with OsTIE1ΔEAR, in which the EAR motif was deleted. We coexpressed 35Spro:DBD, 35Spro:DBD-OsTIE1 or 35Spro:DBD-OsTIE1ΔEAR with the reporter 35Spro-UAS:LUC-35Spro:REN in rice protoplasts. The results showed that 35Spro:DBD-OsTIE1 significantly repressed the expression of the LUC reporter gene in comparison with 35Spro:DBD (Fig. 3J). In contrast, when the EAR motif was deleted, the repression activity of OsTIE1 was significantly decreased, as shown by the results obtained with 35Spro:DBD-OsTIE1ΔEAR (Fig. 3J). These results suggest that OsTIE1 is a transcriptional repressor, and its C-terminal EAR motif is very important for its transcriptional repression activity.
The anther transcriptome regulated by OsTIE1
To determine the molecular mechanisms through which OsTIE1 regulates anther development, we collected anthers at Stage 10 and Stage 12 from Ostie1-1 and WT control plants for RNA-sequencing (RNA-seq) analysis. The good repeatability of the data was determined by Principal Component Analysis (PCA) (Supplementary Fig. S5A). In the comparison of WT control and Ostie1-1 anthers (Ostie1-1 vs WT), a total of 5,208 differentially expressed genes (DEGs; log2 fold change ≥ 1.5 or ≤−1.5; q ≤ 0.05) were identified at Stage 10 and only 2,262 DEGs were identified at Stage 12 (Fig. 4A; Supplementary Data Sets S1 to S4), suggesting that TIE1 regulated more genes at the earlier anther developmental stage. Among the DEGs, 3,467 genes were upregulated and 1,741 genes were downregulated in Ostie1-1 at Stage 10, while 1,751 genes were upregulated and 511 genes were downregulated at Stage 12 in Ostie1-1 (Supplementary Fig. S5, B and C). The number of upregulated genes was about 2-fold or 3-fold the number of downregulated genes at Stage 10 or at Stage 12, suggesting that OsTIE1 acted as a transcriptional repressor. Gene ontology (GO) analysis showed that terms related to hormones, especially JA, were the most enriched terms at Stage 10 and Stage 12, including GO: 0032870 (cellular response to hormone stimulus), GO: 0009725 (response to hormone), GO: 0009753 (response to jasmonic acid), and GO: 0009694 (jasmonic acid metabolic process) (Fig. 4, B and C; Supplementary Tables S1 and S2).
Figure 4.
OsTIE1 regulates the JA synthesis pathway. A) Bar plot of DEGs at Stage 10 and Stage 12. The criteria for DEGs were log2 fold change of ≥1.5 or ≤−1.5 and q ≤ 0.05. B, C) GO enrichment analysis of DEGs between Ostie1-1 and WT at Stage 10 and Stage 12, respectively. Terms under the biological process category were enriched. Several terms related to JA pathway or cell wall biogenesis appeared among DEGs. D) Heat map of JA synthesis genes with log2 fold change revealed by RNA-seq. E, F)OsLOX6 and OsLOX11 were upregulated in Ostie1-1. Data are expressed as the mean (±Sd) of 3 biological replicates. Two-tailed t-test was used for comparisons between groups. *P < 0.05. ***P < 0.001. G) The concentration of JA in anthers at Stage 10 and Stage 12. Data are expressed as the mean (±Sd) of 3 biological replicates. Two-tailed t-test was used for comparisons between groups. ***P < 0.001.
It has been proposed that the JA pathway plays an important role in anther dehiscence (Wilson et al. 2011). Our transcriptome analysis suggests that OsTIE1 could control anther dehiscence by regulating the JA pathway. To provide more evidence to support the findings from our transcriptome analysis, we first determined changes in the expression levels of all JA biosynthesis genes expressed in the anthers using our RNA-seq data. Heatmap analysis showed that nearly all genes related to JA biosynthesis in the anthers of Ostie1-1 were upregulated at Stage 10 and/or at Stage 12. GAOYAO1 (GY1), which encodes a PLA1-type phospholipase that initiates the first step of JA biosynthesis, was significantly upregulated in the anthers of Ostie1-1 (Cai et al. 2014; Xiong et al. 2017). Other JA biosynthesis genes, including OsLOX2, OsLOX6, OsLOX7, OsLOX11, OsAOS1, OsOPR10, and OsAOC, were also upregulated more than 2-fold in Ostie1-1 (Fig. 4D) (Liu et al. 2017; Huang et al. 2023). Next, we used RT-qPCR to confirm that the expression levels of OsLOX6 and OsLOX11 were indeed significantly higher in the anthers of Ostie1-1 in comparison with those of the WT control at Stage 10 and Stage 12 (Fig. 4, E and F). Finally, we measured the JA concentration in anthers collected from Ostie1-1 and WT control plants at Stage 10 and Stage 12. The JA concentration increased as anthers developed in Ostie1-1 and the WT control (Fig. 4G). In the WT control anthers, the JA concentration was relatively low at Stage 10, while it increased by about 2-fold at Stage 12 (Fig. 4G), consistent with the role of JA in promoting anther dehiscence. In the anthers of Ostie1-1, the JA concentration at Stage 10 was higher than that in the WT control at Stage 10 and Stage 12 (Fig. 4G). At Stage 12, the JA concentration in the anthers of Ostie1-1 was significantly higher than that of the WT control anthers (Fig. 4G). The earlier accumulation of JA in the anthers of Ostie1-1 corresponded to the increased level of OsLOX6 and OsLOX11 and earlier anther dehiscence, suggesting that OsTIE1 plays a critical role in tightly controlling JA biosynthesis during anther development.
High JA abundance caused early anther dehiscence in Ostie1-1
To determine whether the high JA concentration in Ostie1-1 could be the reason for precocious anther dehiscence, we treated the florets of Ostie1-1 and the WT control with the JA biosynthesis inhibitor sodium diethyldithiocarbamate (DIECA) (Farmer et al. 1994). The DIECA treatment began at Stage 10 when the florets were still wrapped in the flag leaves. DIECA treatment significantly repressed precocious anther dehiscence, long filament formation, and shorter anther formation in Ostie1-1 at Stage 12 and Stage 13 (Fig. 5, A to E), and the defective pollen grains were partially rescued correspondingly (Fig. 5, F and G), suggesting that overproduction of JA was the primary cause of early anther dehiscence in Ostie1-1. We then treated the florets of Ostie1-1 and WT plants with JA. JA treatment significantly increased the ratio of dehiscent anthers in both WT and Ostie1-1 florets (Fig. 5, H to J; Supplementary Fig. S6, A and B). In addition, in WT florets, JA treatment led to shorter anthers and reduced the viability of pollen grains (Supplementary Fig. S6, C and D). These results demonstrate that OsTIE1 is a key regulator that plays an important role in determining the appropriate timing of JA accumulation during anther development and thus the proper timing of anther dehiscence for plant reproduction.
Figure 5.
Abnormal JA accumulation in Ostie1-1 is the main reason for premature anther dehiscence. A, B) DIECA treatment of Ostie1-1 rescued the phenotype of anther dehiscence. Dehisced anthers are marked with red arrows. Scale bar: 1 mm. C, I) Statistical analysis of the anther dehiscence rate of Ostie1-1 and WT plants treated with the mock treatment, 1 mm DIECA or 100 μm JA. A 1‰ ethanol solution was used as the mock treatment. Data are expressed as the mean (±Se) (n ≥ 5). Fisher’s protected least significant difference was used for multiple comparisons. Means were judged significantly different at P < 0.05. D, H) Effects of DIECA or JA treatment on WT and Ostie1-1 anthers at Stage 12. Scale bar: 1 mm. E, J) Statistical analysis of the length of anthers and filaments under mock, JA, or DIECA treatment. Data are expressed as the mean (±Sd) (n ≥ 100). Fisher’s protected least significant difference was used for multiple comparisons. Means were judged significantly different at P < 0.05. F, G) I2-KI staining of Ostie1-1 pollen grains under mock or DIECA treatment at Stage 12. Scale bar: 100 μm.
OsTIE1 interacts with the TCP transcription factor OsTCP1/PCF5 in controlling JA biosynthesis
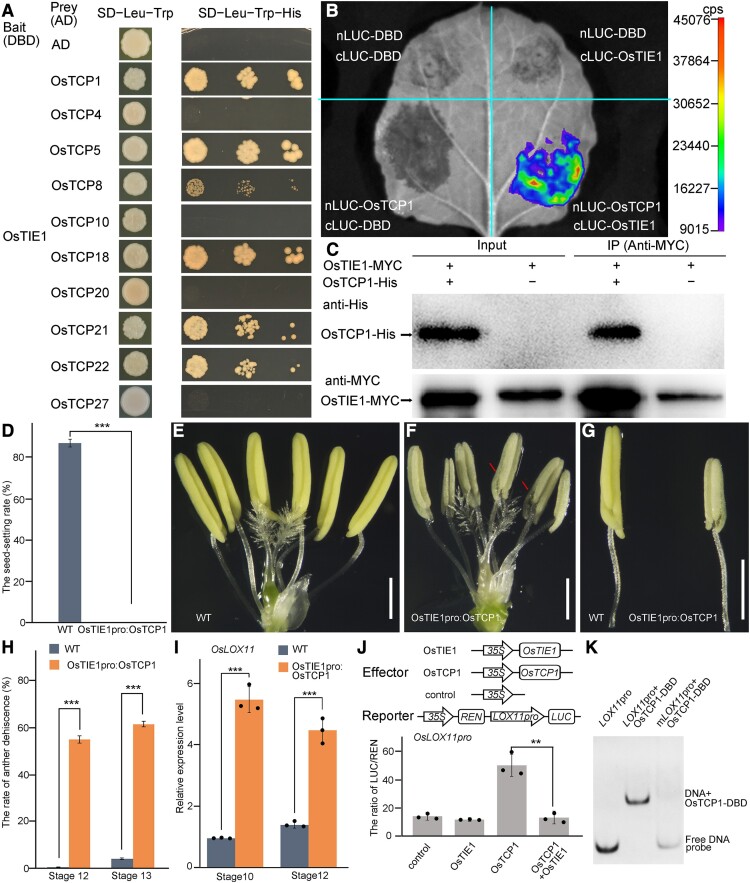
Like Arabidopsis TIE1, OsTIE1 has no apparent DBD (Tao et al. 2013). Arabidopsis TIE1 interacts with Class II TCP transcription factors in the control of leaf development and shoot branching (Tao et al. 2013; Yang et al. 2018). Considering that LOX2 is a direct target of Arabidopsis TCP transcription factors during leaf senescence (Schommer et al. 2008; Danisman et al. 2012), OsTCP21 regulates OsLOX genes in plant disease resistance (Zhang et al. 2016), and OsLOX6 and OsLOX11 were upregulated in Ostie1-1, we hypothesized that OsTIE1 could interact with rice TCP transcription factors in the control of JA biosynthesis during anther development. To test this hypothesis, we first used yeast 2-hybrid assays to examine the interaction between OsTIE1 and each of the 10 rice Class II TCP transcription factors (Martin-Trillo and Cubas 2010). OsTIE1 strongly interacted with OsTCP1/PCF5 (LOC_Os01g11550), OsTCP22/PCF8 (LOC_Os12g42190), OsTCP5 (LOC_Os01g55750), OsTCP18 (LOC_Os05g43760), and OsTCP21 (LOC_Os07g05720) (Fig. 6A). OsTCP8/PCF6 (LOC_Os03g57190) interacted weakly with OsTIE1, while no interaction was detected between OsTIE1 and OsTCP4/PCF7 (LOC_Os01g55100), OsTCP10 (LOC_Os02g51310), OsTCP20 (LOC_Os07g04510), and OsTCP27 (LOC_Os12g02090) (Fig. 6A). The RNA-seq results revealed that genes encoding rice TCPs that were found to interact with OsTIE1 (except OsTCP5) were all expressed at Stage 10 and Stage 12 (Supplementary Fig. S7A). Firefly LUC complementation imaging assays showed clear fluorescence in N. benthamiana leaves with coexpression of nLUC-OsTCP1 and cLUC-OsTIE1, while no fluorescence was detected in the control combinations (Fig. 6B). Coimmunoprecipitation experiments confirmed that OsTIE1 interacted with OsTCP1 in planta (Fig. 6C). We then generated the OsTIE1pro:OsTCP1 construct, in which OsTCP1 was driven by the OsTIE1 promoter. We transformed OsTIE1pro:OsTCP1 into rice cultivar Nipponbare to increase the expression level of OsTCP1 in the anthers. We obtained 5 independent OsTIE1pro:OsTCP1 transgenic lines, all of which were infertile (Fig. 6D). RT-qPCR analysis showed that the expression level of OsTCP1 was significantly increased in the anthers of the OsTIE1pro:OsTCP1 lines (Supplementary Fig. S7B). The anthers of the OsTIE1pro:OsTCP1 lines were precociously dehiscent (Fig. 6, E to H), and pollen was aborted (Supplementary Fig. S7C). These phenotypes caused by increased expression of OsTCP1 resembled that of the Ostie1 mutants, suggesting that the loss of OsTIE1 function may have released the activity of OsTCP transcription factors during anther development. Consistent with this possibility, the expression level of OsLOX11 was significantly elevated in the anthers of the OsTIE1pro:OsTCP1 transgenic lines (Fig. 6I). We also generated the reporter 35S:REN-OsLOX11pro:LUC construct, in which the LUC gene was driven by the OsLOX11 promoter, and the REN LUC gene was driven by the CaMV 35S promoter, as an internal expression control (Fig. 6J). Transient expression in protoplasts showed that OsTCP1 activated the promoter of OsLOX11, while OsTIE1 did not (Fig. 6J). When OsTIE1 was coexpressed with OsTCP1, activation of OsLOX11 by OsTCP1 was significantly attenuated by OsTIE1 (Fig. 6J), suggesting that OsTIE1 repressed the regulatory influence of OsTCP1 activity on the expression of OsLOX11. To determine whether OsTCP1 could directly bind to the promoter region of OsLOX11 and OsLOX6 that were both upregulated in Ostie1-1, we firstly analyzed the promoter sequence of OsLOX11 and OsLOX6 using the binding site prediction function in the plantTFDB database and found possible TCP-binding cis-elements in their promoter regions (Supplementary Fig. S8, A and B) (Jin et al. 2017). We next used electrophoretic mobility shift assay (EMSA) to test the binding of OsTCP1 to the promoter region of OsLOX11 or OsLOX6. The results showed that the TCP domain of OsTCP1 bound to the promoter of OsLOX11 and OsLOX6, while the mutations in cis-elements disrupted the binding of OsTCP1 (Fig. 6K; Supplementary Fig. S8C). To further test whether OsTCP1 could regulate the other genes in JA biosynthesis pathway, we cloned the promoters of OsLOX2, OsPLDα4 (LOC_Os06g40170), and OsOPR10 and used each of them to drive the LUC gene to generate 35S:REN-OsLOX2pro:LUC, 35S:REN-OsPLDα4pro:LUC, and 35S:REN-OsOPR10pro:LUC reporters. Transient expression analysis in protoplasts showed that the promoter of OsPLDα4 or OsOPR10, but not OsLOX2, was activated by OsTCP1 (Supplementary Fig. S8D), consistent with the upregulation of OsPLDα4 and OsOPR10 in Ostie1-1 (Fig. 4D). We also cloned OsTCP22 to generate 35Spro:OsTCP22. Coexpression of 35Spro:OsTCP22 with 35S:REN-OsLOX2pro:LUC, 35S:REN-OsPLDα4pro:LUC, or 35S:REN-OsOPR10pro:LUC in protoplasts showed that OsTCP22 also activated the reporters of OsPLDα4 and OsOPR10 (Supplementary Fig. S8D). These data suggested that TCP family members could redundantly regulate several genes in the JA biosynthesis pathway, finely tuning anther dehiscence.
Figure 6.
OsTIE1 functions by interacting with OsTCP1. A) The interactions between OsTIE1 and CIN-like TCPs in rice were tested using yeast 2-hybrid assays. B) The LUC signal was detected only when nLUC-OsTIE1 and cLUC-OsTCP1 were coexpressed in N. benthamiana leaves. DBD indicates the DBD of yeast GAL4, which was used as the control. C) OsTCP1 fused with His was detected when OsTIE1-MYC was enriched. D) Statistical analysis of the seed-setting rate of WT and OsTIE1pro:OsTCP1 transgenic plants. Data are expressed as the mean (±Sd) (n ≥ 5). The chi-square test was used to compare the seed-setting rate. ***P < 0.001. E to G) Comparison of WT and OsTIE1pro:OsTCP1 transgenic plants at Stage 12. Premature dehisced anthers are marked with red arrows. Scale bar: 1 mm. H) Statistical analysis of the anther dehiscence rate. Data are expressed as the mean (±Se) (n ≥ 5). Two-tailed t-test was used for comparisons between groups. ***P < 0.001. I)OsLOX11 was upregulated in OsTIE1pro:OsTCP1 anther. Data are expressed as the mean (±Sd) of 3 biological replicates. Two-tailed t-test was used for comparisons between groups. ***P < 0.001. J) The dual-LUC assay was employed to test the transcriptional regulation activity of the OsTIE1-OsTCP1 module. Firefly LUC was driven by the promoter of LOX11, which was 2,100-bp upstream of LOX11 coding region. The expression level of firefly LUC was increased when OsTCP1 was transformed and declined when OsTIE1 and OsTCP1 were cotransformed. Data are expressed as the mean (±Sd) of 3 biological replicates. Two-tailed t-test was used for comparisons between groups. **P < 0.01. K) EMSA revealed that OsTCP1 directly bound to the promoter region of OsLOX11.
The aberration TIE-TCP module in Arabidopsis causes defects in anther dehiscence
To evaluate the level of conservation of the function of OsTIE1 and the roles of its homologs in regulating anther dehiscence, we first examined the stamens of transgenic Arabidopsis plants overexpressing OsTIE1. In comparison with the WT control, the 35Spro:OsTIE1 transgenic Arabidopsis plants produced more branches (Fig. 1B). In addition, at the reproductive stage, the filaments of the stamens of the 35Spro:OsTIE1 transgenic Arabidopsis line did not elongate normally, and the filaments were obviously shorter than those of the WT control (Fig. 7, A and G). Interestingly, the anthers of the 35Spro:OsTIE1 transgenic Arabidopsis line were not dehiscent in the mature flowers (Fig. 7, B, C, and F). We next observed the anthers of the tie1-D mutant line, in which Arabidopsis TIE1 was overexpressed by activation tagging (Tao et al. 2013). The stamens and pistils of tie1-D and 35Spro:OsTIE1 transgenic lines were smaller than those of the WT control, and the filaments of stamens were shorter (Fig. 7, D, E, and G). In addition, anther dehiscence was delayed in the opened flowers of tie1-D plants (Fig. 7, D to F). The late dehiscence of anthers exhibited by the 35Spro:OsTIE1 transgenic lines and tie1-D suggests that TIE1 is very important for anther dehiscence. We further analyzed the anther dehiscence phenotype of the Arabidopsis tie1 tie2 tie3 tie4 quadruple mutant in which 4 TIEs were disrupted (He et al. 2022). The results showed that the anthers were frequently dehiscent before flower opening (Fig. 7, H to L), resembling that observed in Ostie1 (Fig. 2, A to F). We also observed the expression pattern of TIE1 protein in anthers using TIE1pro:TIE1-GUS transgenic line in which TIE1 gene fusion with GUS reporter was driven by a 2,790-bp-long TIE1 promoter. The GUS staining was clearly observed in anthers. Interestingly, the predominant expression of TIE1 protein matched well with the anther dehiscent areas. These results indicated that TIE1 plays an important role in the control of the proper anther dehiscence.
Figure 7.
TIE-TCP modules regulate anther dehiscence in Arabidopsis. A to C) Increased expression of OsTIE1 in Arabidopsis produced a phenotype of anther indehiscence and shortened filaments, opposite to that of Ostie1-1. D, E) The TIE1 gain-of-function mutant tie1-D displayed a phenotype similar to that of 35Spro:OsTIE1. Scale bar: 1 mm in A) and D) and 100 μm in B), C), and E). F) Statistical analysis of anther dehiscence rate at flowering stage. Data are means ± Se (n ≥ 5). Two-tailed t-test was applied for comparison between groups. ***P < 0.001. G) Statistical analysis of the length of filament or anther in WT, tie1-D, and 35S:OsTIE1 at flowering stage. Data are means ± Sd (n ≥ 100). Two-tailed t-test was applied for comparison between groups. ***P < 0.001. H to K) Phenotype observation of WT and tie1 tie2 tie3 tie4 1 d before flowering. tie1 tie2 tie3 tie4 exhibited premature dehiscence that phenocopied Ostie1. Scale bar: 1 mm in H) and I) and 200 μm in J) and K). L) Statistical analysis of anther dehiscence rate in H) to K). Data are means ± Se (n ≥ 5). Two-tailed t-test was used. ns, P > 0.05, *P < 0.05, and ***P < 0.001. M to P) GUS staining assay revealed that TIE1 protein was expressed in anthers. Scale bar: 1 mm in M) and N), 50 μm in O), and 20 μm in P). Q to U) Anther dehiscence of WT, tcpSEP, tcpOCT, and tcpUND at flowering stage. tcpOCT, tcpUND exhibited indehiscent anthers that were opposite to that of Ostie1. Scale bar: 1 mm in Q) and 200 μm in R) to U). V) Statistical analysis of anther dehiscence rate at flowering stage in Q) to U). Data are means ± Se (n ≥ 5). Two-tailed t-test was used. ns, P > 0.05, *P < 0.05, and ***P < 0.001. W to Z) GUS staining assay revealed that TCP4 protein was expressed in anthers. Scale bar: 1 mm in W), 500 μm in X), 50 μm in Y), and 20 μm in Z).
TCP transcription factors have high functional redundancy (Lan et al. 2023). To overcome the functional redundancy, we previously generated a series of multiple tcp mutants in Arabidopsis including the septuple mutant tcp2 tcp3 tcp4 tcp5 tcp10 tcp13 tcp17 (tcpSEP), the octuple mutant tcp2 tcp3 tcp4 tcp5 tcp10 tcp13 tcp17 tcp24 (tcpOCT), and the undecuple mutant tcp1 tcp2 tcp3 tcp4 tcp5 tcp10 tcp12 tcp13 tcp17 tcp18 tcp24 (tcpUND). To evaluate the possible redundant roles of TCPs in anther dehiscence, we observed the anther dehiscence phenotype of tcpSEP, tcpOCT, and tcpUND. The results showed that the anther dehiscence of tcpSEP was not significantly different from that of WT (Fig. 7, Q to S and V). However, a significant number of mature anthers were not dehiscent in tcpOCT, and even more mature anthers from tcpUND were indehiscent, when compared to that in WT control (Fig. 7, Q, R, and T to V). We then examined the expression pattern of TCP4 protein in anthers using TCP4pro:TCP4-GUS, which we previously generated (Zheng et al. 2022). The results showed that TCP4 protein was clearly expressed in anthers (Fig. 7, W to Z), and the predominant expression of TCP4 was found in the dehiscent areas of anthers (Fig. 7, Y and Z). These results indicate that TCP4 and TIE1 had overlapping expression pattern during anther development, and the loss of TCP function in tcpOCT and tcpUND leads to an anther dehiscence phenotype that was opposite to that observed in the tie1 tie2 tie3 tie4 mutant. These data suggest that the function and mechanism of the TIE-TCP module might be conserved in finely tuning the anther dehiscence in monocots and dicots.
Discussion
In this study, we demonstrated that OsTIE1 encodes an EAR motif-containing transcriptional repressor critical for anther dehiscence. The loss-of-function Ostie1 mutants generated by CRISPR/Cas9 technology display male sterility due to their prematurely dehiscent anthers, early elongation of stamen filaments, and abortive pollen. Transcriptome analysis and RT-qPCR showed that disruption of OsTIE1 caused increased expression of genes encoding key enzymes in JA biosynthesis during anther development. Moreover, the JA concentration was correspondingly increased in the early anthers of Ostie1, and the treatment with an inhibitor of JA biosynthesis partially rescued the defective anthers and pollen in Ostie1. We further showed that OsTIE1 has transcriptional repression activity and interacts with rice Class II TCP transcription factors, including OsTCP1/PCF5. In addition, we found that increased expression of OsTCP1 in rice anthers phenocopies the early anther dehiscence of Ostie1. Based on our findings, we propose a working model for the role of OsTIE1 in the control of anther dehiscence. OsTIE1 is developmentally regulated during anther development. The precise spatial and temporal expression patterns of OsTIE1 finely tune JA biosynthesis to avoid premature anther dehiscence and mature anther indehiscence. Tight regulation of JA biosynthesis by OsTIE1 is realized by the direct interaction between OsTIE1 and rice TCP transcription factors. OsTCP1 promotes JA biosynthesis by positively regulating the expression of key genes involved in JA biosynthesis, and OsTIE1 acts as a transcriptional repressor to suppress the activity of OsTCP1 (Fig. 8). In the absence of OsTIE1, the repression of OsTCP1 by OsTIE1 was released, increasing the expression of JA biosynthesis genes and leading to early accumulation of JA in anthers and finally inducing precocious anther dehiscence and defective pollen in Ostie1 (Fig. 8).
Figure 8.
Working model of OsTIE1 in the suppression of rice anther dehiscence. JA synthesis is suppressed by the OsTIE1-OsTCP module before Stage 12 in rice to ensure normal timing of anther dehiscence. OsTCP1 acts as a positive regulator of JA synthesis by activating the expression of JA synthesis genes, including LOXs, while OsTIE1 is a transcriptional repressor that functions by interacting with OsTCP1 and suppressing the expression of JA synthesis genes. In Ostie1-1, hyperactive JA synthesis causes premature accumulation of JA and anther dehiscence, while JA synthesis is suppressed until the normal dehiscence time in WT plants expressing OsTIE1.
JA is an important plant hormone that is essential for plant reproductive organ development and male fertility (Huang et al. 2023). In rice, disruption of critical enzymes, including EXTRA GLUME1 and COLEOPTILE PHOTOMORPHOGENESIS 2/HEBIBA, which are involved in JA biosynthesis, causes severe disorder in spikelet and anther development (Agrawal et al. 2003; Li et al. 2009; Riemann et al. 2013; Cai et al. 2014). In this paper, the precocious anther dehiscence in Ostie1 was partially rescued by treatment with the JA biosynthesis inhibitor DIECA, while JA treatment led to early anther dehiscence in the WT, further demonstrating the importance of JA in controlling anther dehiscence. Proper JA biosynthesis during anther development is critical for timely anther dehiscence. Early or late JA peaks cause improper anther dehiscence before or after pollen maturation, failing plant reproduction. Our data indicate that the fine-tuning of JA biosynthesis or peaks during anther development is tightly and elegantly governed by the TIE-TCP modules. Plants might use the TIE-TCP modules to flexibly control anther dehiscence in response to various internal and external signals.
In Arabidopsis, several factors have also been shown to tightly regulate JA biosynthesis in the control of stamen development. The plant hormones auxin and gibberellins (GAs) regulate JA biosynthesis to control stamen development (Huang et al. 2023). In auxin signaling, the transcription factors ARF6 and ARF8 positively regulate the expression of DAD1, LOX2, AOS, and OPR3 to facilitate JA biosynthesis (Tabata et al. 2010), whereas IAA8 negatively regulates JA biosynthesis by directly repressing the transactivation activity of ARF6 and ARF8 during stamen development (Wang et al. 2013). In GA signaling, DELLA proteins, including REPRESSOR-of-ga1-3 (RGA), RGA-LIKE 1 (RGL1), and RGL2, repress stamen development, and RGL2 negatively regulates JA biosynthesis by suppressing the expression of DAD1 (Cheng et al. 2009). Consequently, GAs promote JA biosynthesis by mediating DELLA degradation to control stamen development (Cheng et al. 2009). In addition, nutrients such as copper (Cu) also affect JA biosynthesis. Cu deficiency induces the expression of Cu-DEFICIENCY INDUCED TRANSCRIPTION FACTOR1 (CITF1) (Yan et al. 2017). CITF1 interacts with SQUAMOSA PROMOTER BINDING PROTEIN LIKE7, a master regulator of copper homeostasis, to promote Cu delivery to anthers and protect pollen by positively regulating JA biosynthesis (Yan et al. 2017). Some transcription factors have also been reported to regulate JA biosynthesis. The MADS-box transcription factor AG plays essential roles in early stamen initiation and late stamen maturation by directly binding to the promoter region of DAD1 to promote JA biosynthesis (Ito et al. 2007), while NAC-like protein ANTHER INDEHISCENCE FACTOR acts as a transcriptional repressor to negatively regulate JA biosynthesis and anther dehiscence (Shih et al. 2014). In addition, the chromatin remodeling factors of the Imitation of Switch complex promote stamen filament elongation by positively regulating JA biosynthesis (Zhao et al. 2021). Another recent study showed that JINGUBANG (JGB), a protein containing 7 WD40 repeats, inhibits pollen germination by interacting with TCP4 transcription factors to negatively regulate JA biosynthesis (Ju et al. 2016). However, until now, few factors have been found to tightly regulate JA biosynthesis in rice floral organs. In this study, we identified a transcriptional repressor, OsTIE1, which plays a crucial role in stamen development by repressing JA biosynthesis. We also showed that OsTIE1 directly interacts with Class II TCP transcription factor OsTCP1/PCF5. Several JA biosynthetic genes, including OsLOX genes, are greatly upregulated in the loss of function mutant Ostie1 and OsTCP1 overexpression transgenic lines. These findings indicate that OsTIE1 is a key regulator that finely tunes JA biosynthesis and the JA concentration by controlling the transactivation activity of TCPs during stamen development. Tight regulation of JA biosynthesis by the OsTIE1-OsTCP1 module guarantees proper and timely dehiscence of mature anthers, thus avoiding premature anther dehiscence.
TCP transcription factors are conserved in planta and act as master regulators in diverse plant developmental processes by regulating the biosynthesis of different hormones, including brassinosteroids, auxin, salicylic acid, and JA (Martin-Trillo and Cubas 2010; Nicolas and Cubas 2016). In Arabidopsis, Class II TCPs, including TCP4, promote JA biosynthesis by directly binding to the promoter region of LOX2, whereas TCP20, a Class I TCP, maintains the balance of JA biosynthesis by repressing LOX2 and Class I TCP TCP9 during leaf senescence (Schommer et al. 2008; Danisman et al. 2012). TCPs also promote JA biosynthesis in pollen germination and plant resistance to biotic stresses. The activity of TCPs is controlled to tightly regulate JA biosynthesis in different processes. For example, the microRNA (miRNA) miRNA319 negatively regulates the expression of some Class II TCPs at the posttranscriptional level in both Arabidopsis and rice (Danisman et al. 2012; Zhang et al. 2016). Rice ragged stunt virus inhibits JA biosynthesis to facilitate infection by inducing miRNA319, which represses the expression of OsTCPs, including OsTCP21 (Zhang et al. 2016). The aster yellows phytoplasma witches' broom (AY-WB) pathogen secretes AY-WB protein 11 (SAP11), which mediates the degradation of TCP proteins by directly interacting with TCPs to inhibit JA biosynthesis and facilitate infection (Sugio et al. 2011). The Arabidopsis WD40 protein JGB interacts with TCP4 to repress JA biosynthesis during pollen germination (Ju et al. 2016). We previously showed that the transcriptional repressor TIE1 suppresses the activity of TCPs by directly interacting with TCPs during leaf development in Arabidopsis (Tao et al. 2013). The activity of some Class II TCPs is also regulated by miRNA319 in Arabidopsis during leaf development (Danisman et al. 2012; Zhang et al. 2016). In this study, we found that the increased expression of OsTCP1/PCF5 via the OsTIE1 promoter phenocopies the early anther dehiscence of Ostie1 mutants in rice. We further demonstrated that OsTIE1 directly interacts with OsTCP1/PCF5 in regulating JA biosynthesis during stamen development, suggesting that the TIE-TCP regulatory module may be conserved in dicots and monocots in the control of JA biosynthesis in various biological processes. Furthermore, overexpression of OsTIE1 or overexpression of TIE1 in the activation tagging mutant tie1-D produced short filaments and indehiscent anthers, which were opposite to the phenotypes displayed by the Ostie1 and tie1 tie2 tie3 tie4 mutant, suggesting that the role of TIE transcriptional repressors in regulating stamen development may be conserved. We found that TCPs play redundant roles and are regulated by TIEs during anther dehiscence in both Arabidopsis and rice, suggesting that the regulation of TCPs by TIEs might also be conserved in dicots and monocots. Whether miRNA could play roles in the tight control of JA biosynthesis by regulating TCP activity during rice anther development is still an open question.
It has been proposed that phytohormone genic male-sterile (PHGMS) lines with disrupted JA biosynthesis may be useful in the production of rice hybrid seeds via 2-line systems because JA can be applied to maintain male sterility (Pak et al. 2020). Our findings reveal the molecular mechanism by which JA accumulation is finely regulated by the OsTIE1-OsTCP1/PCF5 module during stamen development and also provide targets that could be exploited to create PHGMS lines for the production of hybrid seeds by 2-line system strategies.
Methods
Plant materials and growth conditions
Rice (O. sativa L. ssp. japonica) cultivar Nipponbare and Arabidopsis (A. thaliana) ecotype Col-0 were used in this study. Rice plants were grown in a field during the rice growing season in Nanjing (N 32.02°, E 118.51°). Seeds of Arabidopsis plants were surface sterilized in 70% (v/v) ethanol for 10 min and then grown on 1/2 MS medium. The plates with seeds were stratified at 4 °C for 3 d and then placed in a 22 °C chamber with a long-day condition (16-h light and 8-h dark) and a light intensity of 120 µmol m−2 s−1 (Philips F17T8/TL841 17-W bulbs). The seedlings were planted in soil and grown in a greenhouse with the conditions described above. The seeds of N. benthamiana were germinated in soil and grown in the same greenhouse.
Generation of binary constructs, mutants, and transgenic plants
The site-specific mutant line Ostie1-1 was created using CRISPR/Cas9 technology as described previously (Miao et al. 2013). Briefly, the primers of gene-specific spacers (OsTIE-spc1-F and OsTIE-spc1-R) targeting OsTIE1 were annealed and cloned into the Bsa I site of the pOs-sgRNA vector to generate entry construct pOs-sgRNA-OsTIE1 by T4 ligation. pH-Ubi-Cas9-OsTIE1 was generated by an LR reaction between pOs-sgRNA-OsTIE1 and pH-Ubi-Cas9-7. pH-Ubi-Cas9-OsTIE1 was transformed into rice by Agrobacterium tumefaciens (EHA105) using embryo-derived calli of Nipponbare. The transgenic lines were screened with 50-mg/L hygromycin. The mutations were determined by sequencing the genomic region of the OsTIE1 gene in Ostie1-1.
The site-specific mutant line Ostie1-2 was created using CRISPR/Cas9 technology as described previously (Ma et al. 2015). Briefly, the primers for gene-specific spacers (OsTIE-spc2-F and OsTIE-spc2-R) targeting OsTIE1 were annealed and finally cloned into the Bsa I sites of the pYLCRISPR/Cas9Pubi-H vector by Golden Gate ligation to form pYLCRISPR/Cas9Pubi-H-OsTIE1 using pYLsgRNA-OsU3b as the donor of OsU3b and sgRNA. pYLCRISPR/Cas9Pubi-H-OsTIE1 was transformed into rice by A. tumefaciens (EHA105) using embryo-derived calli of Nipponbare. The transgenic lines were screened with 50-mg/L hygromycin. The mutations were determined by sequencing the genomic region of the OsTIE1 gene in Ostie1-2.
To investigate the spatial and temporal expression patterns of OsTIE1, a 3,000-bp-long promoter upstream of the coding region of OsTIE1 was amplified from the genomic DNA of Nipponbare. The promoter region was fused to the GUS coding sequence and NOS terminator from the vector pBI121 and cloned into pCAMBIA1300 to generate OsTIE1pro:GUS. OsTIE1pro:GUS transgenic lines were generated by Agrobacterium-mediated rice transformation.
To investigate the function of OsTCP1, the coding region of OsTCP1 driven by 3,000-bp-long OsTIE1 promoter as described above was cloned into pCAMBIA1300 backbone to generate OsTIE1pro:OsTCP1. OsTIE1pro:OsTCP1 transgenic lines were generated by Agrobacterium-mediated rice transformation.
To investigate the expression patterns of TIE1 and TCP4 during anther development, a 2,790-bp promoter with the full-length genomic coding region sequence of TIE1 without stop codon was amplified from Arabidopsis genomic DNA using the primer pair topo-TIE1promoter-FCF/topo-TIE1cds-FCR. The fragment was cloned into the pENTR/D-TOPO vector to generate pENTR-TIE1pg. TIE1pro:TIE1-GUS was finally generated by Gateway Cloning via the LR reaction between pENTR-TIE1pg and pB7GUSWG0 (Invitrogen). TCP4pro:TCP4-GUS construct and transgenic line were generated as described previously (Zheng et al. 2022).
To generate the construct 35Spro:OsTIE1, the coding region of OsTIE1 was amplified from genomic DNA extracted from mature rice leaves with primer pair OsTIE1-F/OsTIE1-R. 35Spro:OsTIE1 was first introduced into A. tumefaciens (GV3101) and then transformed into Arabidopsis by the floral dip method.
Primers used in this study are listed in Supplementary Data Set S7.
PCR reaction and gene expression assays
To perform PCR analysis, genomic DNA was extracted from WT plants, mutants, and transgenic plants using the CTAB method. The PCR reaction conditions were as follows: 94 °C for 30 s, 55 °C to 58 °C for 30 s, and 72 °C for 1 to 2 min for 35 cycles.
To evaluate gene expression using RT-qPCR, total RNA was extracted from anthers and other tissues collected from WT control plants, Ostie1 mutants, and OsTIE1pro:OsTCP1 transgenic lines. Genomic DNA was digested using DNase I (TaKaRa, Japan), after which total RNA was reverse-transcribed using the M-MLV kit (TIANGEN, China). The cDNA was diluted and used as the template to perform RT-qPCR with SYBR Green Mix (TOYOBO, Japan). The RT-qPCR condition was as follows: 95 °C for 2 min, 40 cycles of 94 °C for 10 s, 55 °C for 20 s, and 72 °C for 20 s. The ΔΔCT (cycle threshold) method was employed to quantify the relative expression levels of genes. The OsUBQ (LOC_Os03g13170) gene was used as an internal control.
Staining and microscopy
Histochemical GUS staining was performed as described previously (Tao et al. 2013). Briefly, the rice or Arabidopsis florets were treated with 90% acetone for 30 min on ice. The tissues were transferred into phosphate buffer (0.34 m Na2HPO4·12H2O, 0.4 mm K4[Fe(CN)6]·3H2O, 0.01 m NaH2PO4·2H2O, and 0.5 mm K3[Fe(CN)6]) for 2 washes. The florets were then placed into GUS staining solution (100 mm Na3PO4 [pH 7.0], 1 mm EDTA, 1 mm potassium ferrocyanide, 1% Triton X-100 [v/v], and 1-mg/mL 5-bromo-4-chloro-3-indolyl β-D-glucuronide [X-Gluc]) overnight at 37 °C in darkness. The tissues were finally decolorized with 75% ethanol (v/v) and photographed with a Leica M205 FCA stereoscope.
To perform SEM analysis, rice stamens and pistils from Ostie1 mutants and WT plants were dissected and fixed in 50% FAA buffer (v/v) overnight at 4 °C. Serial ethanol dehydration was performed before critical point drying in liquid CO2. Samples were observed using a scanning electron microscope (JEOL JSM-6610LV) following the procedures described in the user manual.
To observe the anther phenotype, semithin sections were made using spurr resin (SPI) according to a previously described procedure (Zhang et al. 2016). Images were photographed with an Olympus BX51 microscope using an Olympus DP 70 digital camera.
To perform the TEM analysis, rice tissue was fixed in 4% (v/v) glutaraldehyde diluted in 0.1 m PBS buffer (pH = 7.4) overnight at 4 °C. Samples were then washed 3 times with 0.1 m PBS buffer (pH = 7.4) and stained with 2% osmium tetroxide (w/v) along with 1.5% potassium ferricyanide (w/v) dissolved in the same buffer at 4 °C for 2 h, followed by 3 washes in water. Next, en bloc staining was performed with 2% uranyl acetate (w/v) overnight at 4 °C. A graded ethanol series was used for sample dehydration, and fresh resin was used for subsequent embedding. Ultrathin (70 nm) sections were taken after polymerization at 65 °C using a Leica UC7 ultramicrotome equipped with a Ditome diamond knife. Poststaining was implemented using uranyl acetate and lead citrate. Grids with samples were observed at 80 kV in a JEOL Jem-1400 TEM and imaged by a CMOS camera (XAROSA, EMSIS).
To carry out I2-KI staining of pollen grains, anthers from blooming florets were picked and immersed in 1% (w/v, 1-g I2 and 8-g KI dissolved in 100-mL water) I2-KI staining solution for 5 min. The stained anthers were then extruded with a cover slide and imaged by a ZEISS Axio Imager M2 microscope.
Yeast 2-hybrid assays
To test the interaction between OsTIE1 and CIN-like TCP members, the bait construct OsTIE1-BD was generated through LR reaction between TOPO-OsTIE1 and pDEST32 (Invitrogen) as described previously (Han et al. 2019). The coding sequences of OsTCP1, OsTCP4, OsTCP5, OsTCP8, OsTCP10, OsTCP18, OsTCP20, OsTCP21, OsTCP22, and OsTCP27 were amplified from cDNA as described above. These sequences were then cloned into the pDEST22 (Invitrogen) vector by LR reaction as prey. The prey and bait constructs were cotransformed into yeast strain AH109 (Clontech).
The yeasts were grown on SD-Leu-Trp selection medium in a 30 °C incubator for 3 d, after which they were transferred into SD-Leu-Trp-His with 5 mm of 3-amino-1,2,4-triazole for recording and imaging.
Firefly LUC complementation imaging assays
To confirm the interaction between OsTIE1 and OsTCP1, 35Spro:cLUC-OsTIE1 and 35Spro:nLUC-OsTCP1 were constructed through LR reaction with the pCAMBIA1300 backbone as described previously (Lan et al. 2021). Each of the resulting plasmids, as well as a blank plasmid with the yeast GAL4-binding domain, was transformed separately into A. tumefaciens strain GV3101. Different combinations of plasmids were transformed into N. benthamiana leaves through infiltration. After 72-h incubation, the N. benthamiana leaves were collected and injected with 100 mm luciferin and kept in darkness for 5 min. Luminescence was imaged using a NightOWL818 II LB983 imaging system with indiGO software.
Coimmunoprecipitation
To confirm the interaction between OsTIE1 and OsTCP1 in vivo, the coding sequences of OsTIE1 or OsTCP1 were fused with MYC or HIS-tag and cloned into the pK7WG2 backbone. The resulting plasmids were cotransformed into Arabidopsis protoplasts. After incubation in darkness for 12 h, total protein was extracted from protoplasts with IP buffer as described (Lan et al. 2021). The samples were incubated with anti-MYC agarose beads (Sigma, E6654) for 3 h, washed 5 times in 6× SDS solution, and heated to boiling at 100 °C for 10 min. The samples then underwent electrophoresis on a 12% PAGE gel and transferred onto a PVDF membrane (Millipore, ISEQ00010). The membrane was cut apart and hybridized with anti-His antibody (Abcam, ab18184, 1:5,000), or anti-Myc antibody (CWbio, CW0299, 1:5,000), respectively, followed by goat-anti-mouse HRP (CWbio, CW0102S, 1:10,000) for Myc or His detection, and imaged with a Tanon 5200 Multi Automatic Chemiluminescence Fluorescence Imaging Analysis System.
Transcriptional activity analysis using dual-LUC reporter assays
To test the transcriptional repression activity of OsTIE1, the EAR-motif (DLELKLGNC) was removed from OsTIE1 to generate OsTIE1△EAR. The coding sequence of OsTIE1 and OsTIE1△EAR were cloned in frame with the yeast GAL4-binding domain into pGreen II 62-SK. A pGreen II 0800-LUC vector fused with the CaMV 35S promoter and cis-element UAS was generated as a reporter plasmid (Hellens et al. 2005). The plasmids were cotransformed into fresh rice protoplasts and incubated as described above. The ratio of LUC/REN intensity was determined with a Glomax 20/20 Luminometer (Promega).
To test the transcriptional activity of the OsTIE1-OsTCP1 module, OsTIE1 and OsTCP1 were separately cloned into pGreen II 62-SK. A 2,100-bp-long promoter upstream of OsLOX11 was cloned into pGreen II 0800-LUC vector to drive the firefly LUC gene. Different combinations of plasmids were cotransformed into fresh Arabidopsis protoplasts and the ratio of LUC/REN intensity was measured.
To test the transcription activation activity of OsTCP1 and OsTCP22, the 2,000-bp-long promoter upstream of OsPLDα4 and OsOPR10 was cloned into the Green II 0800-LUC vector to drive the firefly LUC gene. The ratio of LUC/REN intensity was measured.
EMSA
The 30-bp sequences within the promoter regions of OsLOX6 and OsLOX11 were synthesized. The predicted TCP-binding motifs are “CTGCGGGCCCAC” in the OsLOX6 promoter and “AAGGGACCGC” in the OsLOX11 promoter. The sequence encoding the TCP domain from 88 to 159 amino acid residues of OsTCP1 was cloned into the pET-21b vector with a N-terminal His-tag. The construct was transformed into Escherichia coli strain BL21. The protein purification and electrophoresis of incubated protein–DNA mixture were conducted as described previously (Zheng et al. 2022).
RNA-seq analysis
Rice cultivar Nipponbare and Ostie1-1 were grown in a field until the booting stage. Anthers were collected at Stage 10 (1.5 to 1.7 mm) and Stage 12 (2.0 mm) and immediately frozen in liquid nitrogen. RNA-seq was performed by Annoroad Gene Technology Corporation (Beijing, China). DEGs were identified through DESeq2 software as genes with log2 fold change ≥ 1.5 and q ≤ 0.05 (Love et al. 2014). DEGs were visualized with bar plots and volcano plots, and common DEGs were visualized with Venn diagrams. GO enrichment analysis was implemented with the ClusterProfiler package to identify enriched terms related to biological processes (Yu et al. 2012). The raw RNA-seq data were submitted to https://www.ncbi.nlm.nih.gov/geo with accession number: GSE248536.
Phylogenetic analysis
To perform phylogenetic analysis of 4 TIE proteins from Arabidopsis and 6 OsTIE proteins from rice, full-length amino acid sequences of TIEs were obtained from The Arabidopsis Information Resource (TAIR) database (https://www.arabidopsis.org), and full-length amino acid sequences of OsTIEs were obtained from Rice Genome Annotation Project (http://rice.uga.edu/). Alignment of the sequences was conducted through Multiple Protein Sequence Alignment (Muscle) in MEGA11 with default settings. The best model of the maximum likelihood method was calculated in MEGA11. The phylogenetic tree was further generated using the maximum likelihood method with JTT + G model and 1,000 bootstrap replications. The alignment data of TIEs and OsTIEs were provided in Supplementary File S1. The phylogenetic tree file was provided in Supplementary File S2.
Accession numbers
Accession number of rice genes from this article can be found in Rice Genome Annotation Project (RGAP) under the following accession numbers: OsTIE1 (LOC_Os01g11430), OsTCP1/PCF5 (LOC_Os01g11550), OsTCP4/PCF7 (LOC_Os01g55100), OsTCP5 (LOC_Os01g55750), OsTCP10 (LOC_Os02g51310), OsTCP8/PCF6 (LOC_Os03g57190), OsTCP18 (LOC_Os05g43760), OsTCP20 (LOC_Os07g04510), OsTCP21 (LOC_Os07g05720), OsTCP27 (LOC_Os12g02090), OsTCP22/PCF8 (LOC_Os12g42190), GY1 (LOC_Os01g67430), OsPLDα4 (LOC_Os06g40170), OsLOX1 (LOC_Os02g10120), OsLOX2 (LOC_Os03g08220), OsLOX4 (LOC_Os03g49350), OsLOX6 (LOC_Os04g37430), OsLOX7 (LOC_Os05g23880), OsLOX11 (LOC_Os12g37260), OsAOS1 (LOC_Os03g55880), OsOPR6 (LOC_Os06g11200), OsOPR8 (LOC_Os02g35310), OsOPR10 (LOC_Os01g27230), OsAOC (LOC_Os03g32314), OsACX1 (LOC_Os06g01390), and OsACX3 (LOC_Os06g24704). Accession number of Arabidopsis genes from this article can be found in The Arabidopsis Information Resource (TAIR) under the following accession numbers: TIE1 (AT4G28840) and TCP4 (AT3G15030).
Supplementary Material
Acknowledgments
We thank the Core Facilities of Life Sciences, Peking University, and the National Center for Protein Sciences at Peking University for assistance with the SEM and TEM assays. We are grateful to Dr. Yiqun Liu and Dr. Feng Liu for technical support for the TEM analysis.
Contributor Information
Yuxing Fang, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Beijing 100871, China.
Dongshu Guo, Provincial Key Laboratory of Agrobiology, Institute of Germplasm Resources and Biotechnology, Jiangsu Academy of Agricultural Sciences, Nanjing 210014, China; Zhongshan Biological Breeding Laboratory, Nanjing, 210014, China.
Yi Wang, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Beijing 100871, China.
Ning Wang, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Beijing 100871, China.
Xianwen Fang, Provincial Key Laboratory of Agrobiology, Institute of Germplasm Resources and Biotechnology, Jiangsu Academy of Agricultural Sciences, Nanjing 210014, China.
Yunhui Zhang, Provincial Key Laboratory of Agrobiology, Institute of Germplasm Resources and Biotechnology, Jiangsu Academy of Agricultural Sciences, Nanjing 210014, China.
Xiao Li, Provincial Key Laboratory of Agrobiology, Institute of Germplasm Resources and Biotechnology, Jiangsu Academy of Agricultural Sciences, Nanjing 210014, China; Zhongshan Biological Breeding Laboratory, Nanjing, 210014, China.
Letian Chen, State Key Laboratory for Conservation and Utilization of Subtropical Agro-Bioresources, College of Life Sciences, South China Agricultural University, Guangzhou 510642, China.
Diqiu Yu, State Key Laboratory for Conservation and Utilization of Bio-Resources in Yunnan, School of Life Sciences, Yunnan University, Kunming 650091, China; Southwest United Graduate School, Kunming 650092, China.
Baolong Zhang, Provincial Key Laboratory of Agrobiology, Institute of Germplasm Resources and Biotechnology, Jiangsu Academy of Agricultural Sciences, Nanjing 210014, China; Zhongshan Biological Breeding Laboratory, Nanjing, 210014, China.
Genji Qin, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Beijing 100871, China; Southwest United Graduate School, Kunming 650092, China.
Author contributions
G.Q. and B.Z. conceived the project. G.Q., Y.F., B.Z., Y.W., and D.G. designed the experiments. Y.F., D.G., Y.W., N.W., X.F., Y.Z., and X.L. performed the experiments. G.Q., B.Z., Y.F., Y.W., D.G., D.Y., and L.C. analyzed the data. G.Q., Y.F., and D.G. wrote the paper.
Supplementary data
The following materials are available in the online version of this article.
Supplementary Figure S1. OsTIE1 is a close putative ortholog of Arabidopsis TIE1.
Supplementary Figure S2. The genotypes of Ostie1-1 and Ostie1-2.
Supplementary Figure S3. The defective phenotype of Ostie1 alleles appears after Stage 12.
Supplementary Figure S4. Cross-section of anthers and TEM observation of pollen in Ostie1-1 and WT.
Supplementary Figure S5. Transcriptional profiling of WT and Ostie1-1.
Supplementary Figure S6. Exogenous JA treatment of WT plants induced premature anther dehiscence and inviable pollen.
Supplementary Figure S7. The expression of OsTCPs in anthers and the upregulation of OsTCP1 in OsTIE1pro:OsTCP1 caused male sterility.
Supplementary Figure S8. The genes in JA biosynthesis pathway were regulated by OsTCP1 and OsTCP22.
Supplementary Table S1. The enriched GO terms of DEGs at Stage 10.
Supplementary Table S2. The enriched GO terms of DEGs at Stage 12.
Supplementary Data Set S1. Genes upregulated in Ostie1-1 at Stage 10.
Supplementary Data Set S2. Genes downregulated in Ostie1-1 at Stage 10.
Supplementary Data Set S3. Genes upregulated in Ostie1-1 at Stage 12.
Supplementary Data Set S4. Genes downregulated in Ostie1-1 at Stage 12.
Supplementary Data Set S5. Primers used in this study.
Supplementary Data Set S6. Statistical tests for figures.
Supplementary File S1. Multiple alignments of amino acid sequences of TIEs and OsTIEs
Supplementary File S2. Phylogenic tree of TIEs and OsTIEs
Funding
This research was supported by the National Science Fund for Distinguished Young Scholars of China (grant no. 31725005), the National Natural Science Foundation of China (grant no. 31970194), funding from the Jiangsu Provincial Key Laboratory of Agrobiology (JKLA2022-ZD01), and funding from Zhongshan Biological Breeding Laboratory (BM2022008-02 and ZSBBL-KY2023-06).
Dive Curated Terms
The following phenotypic, genotypic, and functional terms are of significance to the work described in this paper:
References
- Abbas A, Yu P, Sun L, Yang Z, Chen D, Cheng S, Cao L. Exploiting genic male sterility in rice: from molecular dissection to breeding applications. Front. Plant Sci. 2021:12:629314. 10.3389/fpls.2021.629314 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Acosta IF, Przybyl M. Jasmonate signaling during Arabidopsis stamen maturation. Plant Cell Physiol. 2019:60(12):2648–2659. 10.1093/pcp/pcz201 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Agrawal GK, Jwa N-S, Agrawal SK, Tamogami S, Iwahashi H, Rakwal R. Cloning of novel rice allene oxide cyclase (OsAOC): mRNA expression and comparative analysis with allene oxide synthase (OsAOS) gene provides insight into the transcriptional regulation of octadecanoid pathway biosynthetic genes in rice. Plant Sci. 2003:164(6):979–992. 10.1016/S0168-9452(03)00082-7 [DOI] [Google Scholar]
- Cai Q, Yuan Z, Chen M, Yin C, Luo Z, Zhao X, Liang W, Hu J, Zhang DB. Jasmonic acid regulates spikelet development in rice. Nat Commun. 2014:5(1):3476. 10.1038/ncomms4476 [DOI] [PubMed] [Google Scholar]
- Caldelari D, Wang G, Farmer EE, Dong X. Arabidopsis lox3 lox4 double mutants are male sterile and defective in global proliferative arrest. Plant Mol. Biol. 2011:75(1-2):25–33. 10.1007/s11103-010-9701-9 [DOI] [PubMed] [Google Scholar]
- Cecchetti V, Altamura MM, Brunetti P, Petrocelli V, Falasca G, Ljung K, Costantino P, Cardarelli M. Auxin controls Arabidopsis anther dehiscence by regulating endothecium lignification and jasmonic acid biosynthesis. Plant J. 2013:74(3):411–422. 10.1111/tpj.12130 [DOI] [PubMed] [Google Scholar]
- Cecchetti V, Altamura MM, Falasca G, Costantino P, Cardarelli M. Auxin regulates Arabidopsis anther dehiscence, pollen maturation, and filament elongation. Plant Cell. 2008:20(7):1760–1774. 10.1105/tpc.107.057570 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cheng H, Song S, Xiao L, Soo HM, Cheng Z, Xie D, Peng J. Gibberellin acts through jasmonate to control the expression of MYB21, MYB24, and MYB57 to promote stamen filament growth in Arabidopsis. PLoS Genet. 2009:5(3):e1000440. 10.1371/journal.pgen.1000440 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chow V, Kirzinger MW, Kagale S. Lend me your ears: a systematic review of the broad functions of ear motif-containing transcriptional repressors in plants. Genes (Basel). 2023:14(2):270. 10.3390/genes14020270 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Danisman S, van der Wal F, Dhondt S, Waites R, de Folter S, Bimbo A, van Dijk ADJ, Muino JM, Cutri L, Dornelas MC, et al. Arabidopsis Class I and Class II TCP transcription factors regulate jasmonic acid metabolism and leaf development antagonistically. Plant Physiol. 2012:159(4):1511–1523. 10.1104/pp.112.200303 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Degenhardt RF, Bonham-Smith PC. Arabidopsis ribosomal proteins RPL23aA and RPL23aB are differentially targeted to the nucleolus and are disparately required for normal development. Plant Physiol. 2008:147(1):128–142. 10.1104/pp.107.111799 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Farmer EE, Caldelari D, Pearce G, Walker-Simmons MK, Ryan CA. Diethyldithiocarbamic acid inhibits the octadecanoid signaling pathway for the wound induction of proteinase inhibitors in tomato leaves. Plant Physiol. 1994:106(1):337–342. 10.1104/pp.106.1.337 [DOI] [Google Scholar]
- Figueroa P, Browse J. Male sterility in Arabidopsis induced by overexpression of a MYC5-SRDX chimeric repressor. Plant J. 2015:81(6):849–860. 10.1111/tpj.12776 [DOI] [PubMed] [Google Scholar]
- Ghelli R, Brunetti P, Napoli N, De Paolis A, Cecchetti V, Tsuge T, Serino G, Matsui M, Mele G, Rinaldi G, et al. A newly identified flower-specific splice variant of AUXIN RESPONSE FACTOR8 regulates stamen elongation and endothecium lignification in Arabidopsis. Plant Cell. 2018:30(3):620–637. 10.1105/tpc.17.00840 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ghorbel M, Brini F, Sharma A, Landi M. Role of jasmonic acid in plants: the molecular point of view. Plant Cell Rep. 2021:40(8):1471–1494. 10.1007/s00299-021-02687-4 [DOI] [PubMed] [Google Scholar]
- Goldberg RB, Beals TP, Sanders PM. Anther development: basic principles and practical applications. Plant Cell. 1993:5(10):1217–1229. 10.1105/tpc.5.10.1217 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guo J-X, Liu Y-G. Molecular control of male reproductive development and pollen fertility in rice. J Integr Plant Biol. 2012:54(12):967–978. 10.1111/j.1744-7909.2012.01172.x [DOI] [PubMed] [Google Scholar]
- Han X, Yu H, Yuan R, Yang Y, An F, Qin G. Arabidopsis transcription factor TCP5 controls plant thermomorphogenesis by positively regulating PIF4 activity. iScience. 2019:15:611–622. 10.1016/j.isci.2019.04.005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- He Q, Yuan R, Zhang T, An F, Wang N, Lan J, Wang X, Zhang Z, Pan Y, Wang X, et al. Arabidopsis TIE1 and TIE2 transcriptional repressors dampen cytokinin response during root development. Sci Adv. 2022:8(36):eabn5057. 10.1126/sciadv.abn5057 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hellens RP, Allan AC, Friel EN, Bolitho K, Grafton K, Templeton MD, Karunairetnam S, Gleave AP, Laing WA. Transient expression vectors for functional genomics, quantification of promoter activity and RNA silencing in plants. Plant Methods. 2005:1(1):1–14. 10.1186/1746-4811-1-13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hiratsu K, Matsui K, Koyama T, Ohme-Takagi M. Dominant repression of target genes by chimeric repressors that include the EAR motif, a repression domain, in Arabidopsis. Plant J. 2003:34(5):733–739. 10.1046/j.1365-313X.2003.01759.x [DOI] [PubMed] [Google Scholar]
- Huang H, Chen Y, Wang S, Qi T, Song S. Jasmonate action and crosstalk in flower development and fertility. J Exp Bot. 2023:74(4):1186–1197. 10.1093/jxb/erac251 [DOI] [PubMed] [Google Scholar]
- Ishiguro S, Kawai-Oda A, Ueda J, Nishida I, Okada K. The DEFECTIVE IN ANTHER DEHISCENCE1 gene encodes a novel phospholipase A1 catalyzing the initial step of jasmonic acid biosynthesis, which synchronizes pollen maturation, anther dehiscence, and flower opening in Arabidopsis. Plant Cell. 2001:13(10):2191–2209. 10.1105/tpc.010192 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ito T, Ng K-H, Lim T-S, Yu H, Meyerowitz EM. The homeotic protein AGAMOUS controls late stamen development by regulating a jasmonate biosynthetic gene in Arabidopsis. Plant Cell. 2007:19(11):3516–3529. 10.1105/tpc.107.055467 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jewell JB, Browse J. Epidermal jasmonate perception is sufficient for all aspects of jasmonate-mediated male fertility in Arabidopsis. Plant J. 2016:85(5):634–647. 10.1111/tpj.13131 [DOI] [PubMed] [Google Scholar]
- Jin JP, Tian F, Yang DC, Meng YQ, Kong L, Luo JC, Gao G. PlantTFDB 4.0: toward a central hub for transcription factors and regulatory interactions in plants. Nucleic Acids Res. 2017:45(D1):D1040–D1045. 10.1093/nar/gkw982 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ju Y, Guo L, Cai Q, Ma F, Zhu Q-Y, Zhang Q, Sodmergen. Arabidopsis JINGUBANG is a negative regulator of pollen germination that prevents pollination in moist environments. Plant Cell. 2016:28(9):2131–2146. 10.1105/tpc.16.00401 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kagale S, Links MG, Rozwadowski K. Genome-wide analysis of ethylene-responsive element binding factor-associated amphiphilic repression motif-containing transcriptional regulators in Arabidopsis. Plant Physiol. 2010:152(3):1109–1134. 10.1104/pp.109.151704 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kagale S, Rozwadowski K. EAR motif-mediated transcriptional repression in plants: an underlying mechanism for epigenetic regulation of gene expression. Epigenetics. 2011:6(2):141–146. 10.4161/epi.6.2.13627 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lan J, Wang N, Wang Y, Jiang Y, Yu H, Cao X, Qin G. Arabidopsis TCP4 transcription factor inhibits high temperature-induced homeotic conversion of ovules. Nat Commun. 2023:14(1):5673. 10.1038/s41467-023-41416-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lan J, Zhang J, Yuan R, Yu H, An F, Sun L, Chen H, Zhou Y, Qian W, He H, et al. TCP transcription factors suppress cotyledon trichomes by impeding a cell differentiation-regulating complex. Plant Physiol. 2021:186(1):434–451. 10.1093/plphys/kiab053 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li H, Xue D, Gao Z, Yan M, Xu W, Xing Z, Huang D, Qian Q, Xue Y. A putative lipase gene EXTRA GLUME1 regulates both empty-glume fate and spikelet development in rice. Plant J. 2009:57(4):593–605. 10.1111/j.1365-313X.2008.03710.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu L, Zou Z, Qian K, Xia C, He Y, Zeng H, Zhou X, Riemann M, Yin C. Jasmonic acid deficiency leads to scattered floret opening time in cytoplasmic male sterile rice Zhenshan 97A. J Exp Bot. 2017:68(16):4613–4625. 10.1093/jxb/erx251 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lorenzo O, Chico JM, Saénchez-Serrano JJ, Solano R. JASMONATE-INSENSITIVE1 encodes a MYC transcription factor essential to discriminate between different jasmonate-regulated defense responses in Arabidopsis. Plant Cell. 2004:16(7):1938–1950. 10.1105/tpc.022319 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014:15(12):1–21. 10.1186/s13059-014-0550-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ma X, Zhang Q, Zhu Q, Liu W, Chen Y, Qiu R, Wang B, Yang Z, Li H, Lin Y, et al. A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol Plant. 2015:8(8):1274–1284. 10.1016/j.molp.2015.04.007 [DOI] [PubMed] [Google Scholar]
- Marchant DB, Walbot V. Anther development—the long road to making pollen. Plant Cell. 2022:34(12):4677–4695. 10.1093/plcell/koac287 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martin-Trillo M, Cubas P. TCP genes: a family snapshot ten years later. Trends Plant Sci. 2010:15(1):31–39. 10.1016/j.tplants.2009.11.003 [DOI] [PubMed] [Google Scholar]
- Matsui T, Omasa K, Horie T. Mechanism of anther dehiscence in rice (Oryza sativa L). Ann Bot. 1999:84(4):501–506. 10.1006/anbo.1999.0943 [DOI] [Google Scholar]
- Miao J, Guo D, Zhang J, Huang Q, Qin G, Zhang X, Wan J, Gu H, Qu L-J. Targeted mutagenesis in rice using CRISPR-Cas system. Cell Res. 2013:23(10):1233–1236. 10.1038/cr.2013.123 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nicolas M, Cubas P. TCP factors: new kids on the signaling block. Curr. Opin. Plant Biol. 2016:33:33–41. 10.1016/j.pbi.2016.05.006 [DOI] [PubMed] [Google Scholar]
- Ohta M, Matsui K, Hiratsu K, Shinshi H, Ohme-Takagi M. Repression domains of Class II ERF transcriptional repressors share an essential motif for active repression. Plant cell. 2001:13(8):1959–1968. 10.1105/TPC.010127 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pak H, Wang H, Kim Y, Song U, Tu M, Wu D, Jiang L. Creation of male-sterile lines that can be restored to fertility by exogenous methyl jasmonate for the establishment of a two-line system for the hybrid production of rice (Oryza sativa L). Plant Biotechnol J. 2020:19(2):365–374. 10.1111/pbi.13471 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Plant AR, Larrieu A, Causier B. Repressor for hire! The vital roles of TOPLESS-mediated transcriptional repression in plants. New Phytol. 2021:231(3):963–973. 10.1111/nph.17428 [DOI] [PubMed] [Google Scholar]
- Qi T, Huang H, Song S, Xie D. Regulation of jasmonate-mediated stamen development and seed production by a bHLH-MYB complex in Arabidopsis. Plant Cell. 2015:27(6):1620–1633. 10.1105/tpc.15.00116 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Riemann M, Haga K, Shimizu T, Okada K, Ando S, Mochizuki S, Nishizawa Y, Yamanouchi U, Nick P, Yano M, et al. Identification of rice Allene Oxide Cyclase mutants and the function of jasmonate for defence against Magnaporthe oryzae. Plant J. 2013:74(2):226–238. 10.1111/tpj.12115 [DOI] [PubMed] [Google Scholar]
- Riemann M, Riemann M, Takano M. Rice JASMONATE RESISTANT 1 is involved in phytochrome and jasmonate signalling. Plant Cell Environ. 2008:31(6):783–792. 10.1111/j.1365-3040.2008.01790.x [DOI] [PubMed] [Google Scholar]
- Ruan J, Zhou Y, Zhou M, Yan J, Khurshid M, Weng W, Cheng J, Zhang K. Jasmonic acid signaling pathway in plants. Int J Mol Sci. 2019:20(10):2479. 10.3390/ijms20102479 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schaller F, Biesgen C, Müssig C, Altmann T, Weiler EW. 12-Oxophytodienoate reductase 3 (OPR3) is the isoenzyme involved in jasmonate biosynthesis. Planta. 2000:210(6):979–984. 10.1007/s004250050706 [DOI] [PubMed] [Google Scholar]
- Schommer C, Palatnik JF, Aggarwal P, Chételat A, Cubas P, Farmer EE, Nath U, Weigel D. Control of jasmonate biosynthesis and senescence by miR319 targets. PLoS Biol. 2008:6(9):e230. 10.1371/journal.pbio.0060230 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shih C-F, Hsu W-H, Peng Y-J, Yang C-H. The NAC-like gene ANTHER INDEHISCENCE FACTOR acts as a repressor that controls anther dehiscence by regulating genes in the jasmonate biosynthesis pathway in Arabidopsis. J Exp Bot. 2014:65(2):621–639. 10.1093/jxb/ert412 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Song S, Chen Y, Liu L, See YHB, Mao C, Gan Y, Yu H. OsFTIP7 determines auxin-mediated anther dehiscence in rice. Nat Plants. 2018:4(7):495–504. 10.1038/s41477-018-0175-0 [DOI] [PubMed] [Google Scholar]
- Stintzi A, Browse J. The Arabidopsis male-sterile mutant, opr3, lacks the 12-oxophytodienoic acid reductase required for jasmonate synthesis. Proc Natl Acad Sci U S A. 2000:97(19):10625–10630. 10.1073/pnas.190264497 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sugio A, Kingdom HN, MacLean AM, Grieve VM, Hogenhout SA. Phytoplasma protein effector SAP11 enhances insect vector reproduction by manipulating plant development and defense hormone biosynthesis. Proc Natl Acad Sci U S A. 2011:108(48):E1254–E1263. 10.1073/pnas.1105664108 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Szemenyei H, Hannon M, Long JA. TOPLESS mediates auxin-dependent transcriptional repression during Arabidopsis embryogenesis. Science. 2008:319(5868):1384–1386. 10.1126/science.1151461 [DOI] [PubMed] [Google Scholar]
- Tabata R, Ikezaki M, Fujibe T, Aida M, Tian C-E, Ueno Y, Yamamoto KT, Machida Y, Nakamura K, Ishiguro S. Arabidopsis AUXIN RESPONSE FACTOR6 and 8 regulate jasmonic acid biosynthesis and floral organ development via repression of Class 1 KNOX genes. Plant Cell Physiol. 2010:51(1):164–175. 10.1093/pcp/pcp176 [DOI] [PubMed] [Google Scholar]
- Tao Q, Guo D, Wei B, Zhang F, Pang C, Jiang H, Zhang J, Wei T, Gu H, Qu L-J, et al. The TIE1 transcriptional repressor links TCP transcription factors with TOPLESS/TOPLESS-RELATED corepressors and modulates leaf development in Arabidopsis. Plant Cell. 2013:25(2):421–437. 10.1105/tpc.113.109223 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tashiro S, Tian C-E, Watahiki MK, Yamamoto KT. Changes in growth kinetics of stamen filaments cause inefficient pollination in massugu2, an auxin insensitive, dominant mutant of Arabidopsis thaliana. Physiol Plant. 2009:137(2):175–187. 10.1111/j.1399-3054.2009.01271.x [DOI] [PubMed] [Google Scholar]
- Thines B, Katsir L, Melotto M, Niu Y, Mandaokar A, Liu G, Nomura K, He SY, Howe GA, Browse J. JAZ repressor proteins are targets of the SCF(COI1) complex during jasmonate signalling. Nature. 2007:448(7154):661–665. 10.1038/nature05960 [DOI] [PubMed] [Google Scholar]
- Wang J, Yan D-W, Yuan T-T, Gao X, Lu Y-T. A gain-of-function mutation in IAA8 alters Arabidopsis floral organ development by change of jasmonic acid level. Plant Mol. Biol. 2013:82(1-2):71–83. 10.1007/s11103-013-0039-y [DOI] [PubMed] [Google Scholar]
- Wilson ZA, Song J, Taylor B, Yang C. The final split: the regulation of anther dehiscence. J Exp Bot. 2011:62(5):1633–1649. 10.1093/jxb/err014 [DOI] [PubMed] [Google Scholar]
- Xiao Y, Chen Y, Charnikhova T, Mulder PPJ, Heijmans J, Hoogenboom A, Agalou A, Michel C, Morel J-B, Dreni L, et al. OsJAR1 is required for JA-regulated floret opening and anther dehiscence in rice. Plant Mol. Biol. 2014:86(1-2):19–33. 10.1007/s11103-014-0212-y [DOI] [PubMed] [Google Scholar]
- Xie DX, Feys BF, James S, Nieto-Rostro M, Turner JG. COI1: an Arabidopsis gene required for jasmonate-regulated defense and fertility. Science. 1998:280(5366):1091–1094. 10.1126/science.280.5366.1091 [DOI] [PubMed] [Google Scholar]
- Xiong Q, Ma B, Lu X, Huang Y-H, He S-J, Yang C, Yin C-C, Zhao H, Zhou Y, Zhang W-K, et al. Ethylene-inhibited jasmonic acid biosynthesis promotes mesocotyl/coleoptile elongation of etiolated rice seedlings. Plant Cell. 2017:29(5):1053–1072. 10.1105/tpc.16.00981 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yan J, Chia J-C, Sheng H, Jung H-I, Zavodna T-O, Zhang L, Huang R, Jiao C, Craft EJ, Fei Z, et al. Arabidopsis pollen fertility requires the transcription factors CITF1 and SPL7 that regulate copper delivery to anthers and jasmonic acid synthesis. Plant Cell. 2017:29(12):3012–3029. 10.1105/tpc.17.00363 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang Y, Nicolas M, Zhang J, Yu H, Guo D, Yuan R, Zhang T, Yang J, Cubas P, Qin G. The TIE1 transcriptional repressor controls shoot branching by directly repressing BRANCHED1 in Arabidopsis. PLoS Genet. 2018:14(3):e1007296. 10.1371/journal.pgen.1007296 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu G, Wang L-G, Han Y, He Q-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 2012:16(5):284–287. 10.1089/omi.2011.0118 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang C, Ding Z, Wu K, Yang L, Li Y, Yang Z, Shi S, Liu X, Zhao S, Yang Z, et al. Suppression of jasmonic acid-mediated defense by viral-inducible microRNA319 facilitates virus infection in rice. Mol Plant. 2016:9(9):1302–1314. 10.1016/j.molp.2016.06.014 [DOI] [PubMed] [Google Scholar]
- Zhang D, Luo X, Zhu L. Cytological analysis and genetic control of rice anther development. J Genet Genomics. 2011:38(9):379–390. 10.1016/j.jgg.2011.08.001 [DOI] [PubMed] [Google Scholar]
- Zhao Y, Jiang T, Li L, Zhang X, Yang T, Liu C, Chu J, Zheng B. The chromatin remodeling complex imitation of switch controls stamen filament elongation by promoting jasmonic acid biosynthesis in Arabidopsis. J Genet Genomics. 2021:48(2):123–133. 10.1016/j.jgg.2021.02.003 [DOI] [PubMed] [Google Scholar]
- Zhao Z, Zhang Y, Liu X, Zhang X, Liu S, Yu X, Ren Y, Zheng X, Zhou K, Jiang L, et al. A role for a dioxygenase in auxin metabolism and reproductive development in rice. Dev Cell. 2013:27(1):113–122. 10.1016/j.devcel.2013.09.005 [DOI] [PubMed] [Google Scholar]
- Zheng X, Lan J, Yu H, Zhang J, Zhang Y, Qin Y, Su X-D, Qin G. Arabidopsis transcription factor TCP4 represses chlorophyll biosynthesis to prevent petal greening. Plant Comm. 2022:3(4):100309. 10.1016/j.xplc.2022.100309 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.