Abstract
Background
FANCA mutations have been detected in a variety of cancers and found to be pro-carcinogenic. However, no functional studies have been identified regarding the involvement of FANCA in the occurrence and the immune response of LUAD.
Methods
The mRNA expression and overall survival rates of FANCA were evaluated by the TIMER, PrognoScan and TCGA database in LUAD tissues, and FANCA expression was further validated by clinical serum samples using ELISA. The correlation between FANCA and immune infiltration level was investigated via TISIDB database and CIBERSORT algorithm. The Kaplan–Meier plotter was used to further evaluate the prognostic value based on the expression levels of FANCA in related immune cells. Then, the influence of FANCA knockout on the proliferation, migration, and invasion of A549 and H1299 cells was validated using CCK8, cloning formation, and Transwell assays. Subsequently, HLA-A2-restricted FANCA antigenic peptides were predicted and synthesized by NetMHC4.0 and SYFPEITHI, and DCs were induced and cultured in vitro. Finally, DCs loaded with HLA-A2-restricted FANCA antigenic peptides were co-cultured with autologous peripheral blood lymphocyte to generate specific CTLs. The killing effects of different CTLs on LUAD cells were studied.
Results
The results showed that high levels of FANCA in patients with LUAD were significantly correlated with worse OS survival, which was correlated with age, clinical stage, pathological T stage, M stage, and N stage in LUAD. Knockdown of FANCA in A549 and H1299 cells significantly inhibited proliferation, metastasis, and invasion in vitro. In addition, FANCA was significantly related to immune infiltrate, genomic alterations and TMB. FANCA expression infuenced the prognosis of LUAD patients by directly affecting immune cell infltration. Finally, HLA-A2-restricted FANCA antigenic peptides were synthesized. And FANCA 146–154 (SLLEFAQYL) antigenic peptide exhibit a stronger affinity for DCs, and induce CTLs to produce stronger targeted killing ability for LUAD cells at an effector-to-target ratio of 40:1.
Conclusion
These results demonstrated that the elevation of FANCA promotes malignant phenotype of LUAD, and the potential peptide P2 (SLLEFAQYL) derived from FANCA may be used as an epitope vaccine for the treatment of LUAD.
Introduction
Lung cancer is one of the most common malignancies in the world with high incidence (2.206 million cases, 11.4%) and high mortality (1.796 million cases, 18%) [1]. Distinguished by distinct pathological categories, lung cancer delineates into small cell lung cancer (15%) and non-small cell lung cancer (85%), with Lung Adenocarcinoma (LUAD) constituting the primary pathological subtype within the non-small cell lung cancer spectrum[2, 3]. The insidious nature of LUAD often leads to late diagnoses, poor prognosis and absence of effective treatments[4]. Recent years have witnessed significant strides in the realm of LUAD treatment, notably in various immunotherapy, such as targeted immune checkpoint inhibitor, tumor vaccines, and adoptive immune cell therapy[5]. Nevertheless, the emergence of drug resistance and off-target effects, stemming from tumor heterogeneity, poses significant hurdles to the clinical efficacy of immunotherapy[6]. Consequently, enhancing the sensitivity and precision targeting of LUAD patients to immunotherapy is an imperative challenge demanding resolution.
Fanconi Anemia Complementation Group A (FANCA) stands as a pivotal member within the core complex gene associated with Fanconi anemia, a condition characterized by pronounced bone marrow failure, dysplasia, and heightened susceptibility to cancer[7, 8]. Extensive research had underscored the crucial roles of FANCA in orchestrating responses to DNA damage, facilitating DNA repair, and upholding genome stability[9]. Mutations in the FANCA gene can lead to deficiencies in DNA damage repair mechanisms, thereby promoting chromosome instability and increasing the risk of developing a number of cancers, such as metastatic ovarian cancer, prostate cancer, and chronic lymphocytic leukemia [9–12]. Recent investigations using genome-wide gene expression data have revealed that high expression of FANCA is not only associated with lung cancer risk, but also strongly associated with poor prognosis in patients with LUAD[13–15]. These studies suggested FANCA may play an important role in lung cancer and can be used as a target for cancer immunotherapy. Nevertheless, the effects of FANCA on the occurrence and development of LUAD, as well as the association with immune mechanisms, remain elusive and warrant further exploration.
In this study, we conducted an analysis of FANCA expression levels in LUAD tissues and explored its associations with clinicopathological features, prognosis, and immune cell infiltration utilizing bio-informatics technology. Furthermore, we investigated FANCA expression in the serum of LUAD patients, and evaluated its effect on the proliferation, migration and invasion abilities of lung adenocarcinoma cells in vitro. Subsequently, mature DCs loaded with HLA-A2-restricted FANCA peptides were co-cultured with autologous peripheral blood lymphocyte to generate specific CTLs to eradicate of LUAD cells. These findings furnish both theoretical insights and experimental evidence for the exploration of novel immunotherapy targets in the context of LUAD.
Methods
Clinical serum samples
Serum samples were collected from 91 LUAD patients and 54 healthy subjects in Fujian Provincial Hospital from January 2023 to January 2024. The inclusion criteria of LUAD patients were: (1) histopathologically confirmed lung adenocarcinoma; (2) no history of other malignant tumors; (3) no prior radiotherapy, chemotherapy, or other treatments. The exclusion criteria of LUAD patients were: (1) other pathological types of lung malignancies; (2) a previous history of surgery for malignant lung tumor; (3) presence of other malignant tumors. The inclusion criteria of healthy volunteer were: (1) adult (≥ 18 years); (2) no history of tumor-related diseases or serious illnesses; (3) no unexplained lung nodules or chronic inflammation. All specimens were obtained with the consent of the patients, in compliance with the Declaration of Helsinki and approved by the Ethics Committee of Fujian Provincial Hospital (K2023-03-010).
TCGA databases
We downloaded LUAD expression profile data from The Cancer Genome Atlas (TCGA) database (https://tcga-data.nci.nih.gov/tcga/). After excluding samples with missing sample expression values, survival information and clinicopathological parameters, a total of 597 cases of RNASeq data (59 cases of adjacent normal tissues and 538 cases of LUAD tissues) were included to normalize the gene matrix. The expression levels of FANCA mRNA in adjacent normal tissues and LUAD tissues were compared by rank sum test. Subsequently, FANCA mRNA expression levels were compared between 59 paired adjacent normal tissues and LUAD tissues. Analyses were performed with R statistical software (version 3.6.4). The correlation between FANCA expression and tumor mutational burden (TMB) was established by using the Spearman’s rank correlation coefficient. Mutation data were analyzed and visualized using the R package ‘maftools’.
TIMER databases
The TIMER database (http://timer.comp-genomics.org/timer/) is a web-based resource that analyzes the interactions between various cancers and immune cells. The database included 10,897 samples from 32 cancer types in the TCGA database. We used the “Gene_DE” module to analyze the expression of FANCA in pan-cancer.
TISIDB databases
The TISIDB database (http://cis.hku.hk/TISIDB/) is an integrated repository web, including data from PubMed literature, molecular profiling, para-cancerous multiomics data, high-throughput screening techniques, and various resources for immunological data retrieved from seven public databases. In our study, the correlation of FANCA with Overall Survival (OS) in LUAD was analyzed using the log-rank test by the TISIDB database.
PrognoScan databases
The PrognoScan database (http://dna00.bio.kyutech.ac.jp/PrognoScan/) is a comprehensive database designed for meta-analysis of the prognostic value of genes, particularly in the context of cancer research. It serves as a powerful tool for linking gene expression data to clinical outcomes such as OS and Disease-Free Survival (DFS). In our research, the GSE31210 cohort from the PrognoScan database was used to investigate the prognostic relevance of FANCA expression alterations in LUAD patients. By employing this database, we were able to categorize FANCA expression levels into “high” or “low” based on the median gene expression values. And blue and red curves correspond to low and high FANCA expressions, respectively.
Kaplan–Meier plotter analysis
The prognostic values of FANCA, based on different levels of immune cell infiltration, was estimated using the Kaplan–Meier plotter (http://kmplot.com). The analysis provided the hazard ratio (HR) along with 95% confidence intervals and the log-rank P-value.
Enzyme-linked immuno sorbent assay (ELISA)
The assay was conducted using the Human FANCA ELISA Kit (Biorbyt, Britain). A standard curve was prepared using concentrations of 2000 pg/mL, 1000 pg/mL, 500 pg/mL, 250 pg/mL, 125 pg/mL, 62.5 pg/mL, and 31.25 pg/mL. Standard material and 100µL of serum samples diluted at 1:2 were added and incubated at 37℃ for 1 h and 30 min. The liquid in the plate was discarded, 1 × washing buffer was added, and the 96-well plate was patted dry without soaking. Then, 100µL Biotin-labeled antibody was added to each well and incubated at 37℃ for 60 min. After washing the plate five times, 100µL HRP-Streptavidin Conjugate was added to each well with incubation in a 37℃ for 30 min. Then, 90µL tetramethylbenzidine(TMB) solution was added to each well and incubated in a 37 ℃ water bath for 10 min. A 50µL termination solution was added to each well, and the absorbance value was detected by a 450 nm microplate reader (Bio-Rad, USA). The standard curve was plotted based on the absorbance values corresponding to the concentrations of the standard substances, and the serum sample concentrations were calculated accordingly.
Cell culture and transfection
Human Non-small cell lung cancer (NSCLC) cell A549 was kindly provided by professor Zhaolei Cui from Fujian Medical University Cancer Hospital. Normal human lung epithelial BEAS-2B cells, NSCLC cell H1299, PC9 and human lymphoblastoid cell T2 (HLA-A2 +) were stored in a liquid nitrogen tank. These cells were cultured in RPMI 1640 medium containing 10% fetal bovine serum (FBS, Thermo Fisher Scientific, USA) and 1% Penicillin–Streptomycin Solution (P/S, BasalMedia, Shanghai, China) at 37 °C and 5% CO2. To investigate the influence of FANCA expression on the progression of LUAD, A549 and H1299 cells were separately transfected with siRNA. The experiment was designed with two groups: the knockdown group, which was treated with FANCA-specific siRNA (si-FANCA group), and the negative control group, which was transfected with non-targeting control siRNA (NC group). Both the FANCA siRNA sequence (5′-GCTCTGCTTTGCAGGATCA-3′) and the control siRNA were obtained from RiboBio (Guangzhou, China). For the 2 mL transfection system, 5 µL of siRNA, 12 µL of transfection reagent, and 120 µL of 1 × loading buffer were added to 1863 µL of RPMI 1640 medium containing 10% FBS.
RNA preparation and quantitative RT-PCR (qRT-PCR) assay
Total RNA was extracted using TRIZOL® reagent (Thermo Fisher Scientific, America) according to the manufacturer’s instructions. And cDNA synthesis was performed using a TaKaRa PrimeScrip™ RT reagent Kit (TaKaRa Biotechnology, Japan). The real-time PCR was conducted using a TB Green®Premix Ex Taq™ Green II (TaKaRa Biotechnology, Japan) on automatic fluorescent PCR amplification instrument (Cobas z480, Roche Diagnostics, USA). GAPDH was used as an internal control. Primers were listed as follows:
FANCA: forward, 5′- TTTGCTTGAGGTAGAAGGTCCA -3′
reverse, 5′-CCCGGCTGAGAGAATACCCA-3′;
GAPDH: forward,5′-TGACTTCAACAGCGACACCCA-3′
reverse, 5′-CACCCTGTTGCTGTAGCCAAA-3′.
Western blot analysis
To analyze the protein level of FANCA in cell lines (BEAS-2B, PC9, A549 and H1299), western blot analysis was conducted as described previously [16]. In summary, total protein was extracted from cells using RIPA lysis buffer (Beyotime Biotechnology, China). The protein extracts was separated by 12.5% SDS-PAGE gel, and then transferred onto PVDF membranes (Millipore, Bedford, MA, USA). The membranes were blocked with 5% non-fatted milk for 1 h at room temperature and incubated with the primary antibodies overnight at 4 °C: an anti-FANCA rabbit polyclonal antibody (1:2500, Proteintech, USA) and an anti-GAPDH rabbit antibody (1:1000, Bioss, USA). After washing the membranes four times for 5 min each with 1 × TBST, they were incubated for 1 h at room temperature with a horseradish peroxidase-conjugated (HRP) secondary antibody. Specific protein bands were detected with ECL immunoblotting and visualized with X-ray films (Kodak, Japan) according to the manufacturer’s instructions. The bands were scanned and further analyzed using ImageJ software to determine densitometry values.
CCK‑8 cell proliferation assay
To determine the effect of FANCA on the growth of A549 and H1299 cells, cell viability was measured using the Cell Counting Kit-8 (CCK8, MedChem Express, China) following the manufacturer's instructions. Briefly, A549-SiFANCA, A549-NC, H1299-SiFANCA, and H1299-NC cells were digested, centrifuged, resuspended, counted, and then seeded in 96-well cell culture plates at a density of 5 × 10^3 cells per well. Each group contained three replicate wells. They were cultured in an incubator at 37 °C with 5% CO2 for 24, 48, 72, and 96 h, respectively.
Subsequently, 10 µL of CCK-8 reagent was added to each well at the corresponding time points and incubated at 37 °C for 2 h. The optical density (OD) of the plates was then measured at 450 nm using a microplate reader (Bio-Rad, USA). Based on these measurements, the tumor cell proliferation curve was plotted.
Wound healing assay
A 2 mL suspension of 3 × 10^5 cells transfected with siRNA was seeded into 6-well plates (Saining Life Science, Suzhou, China). Once the cells reached 80% confluence, a straight wound was artificially made by utilizing a 100 µL pipette tip. Images of the wound area were captured at 0 and 24 h.
Migration assay
A 200 µL suspension of cells, containing 4 × 10^4 cells, was seeded into the upper Transwell chamber (8.0 µm pore size, Corning, Japan). To the lower chamber, 600 µL of RPMI 1640 medium supplemented with 10% FBS was added. After a 48-h incubation at 37 °C and 5% CO2, the upper chamber was washed twice with PBS. The upper chamber was then fixed with 4% formaldehyde (Solarbio® Life Sciences, Beijing, China) for 30 min. After three additional washes with PBS, the cells in the upper chamber were stained with 1% crystal violet (Solarbio® Life Sciences, Beijing, China) for 30 min and subsequently photographed.
Prediction, synthesis and purification of FANCA peptides
FANCA peptides were predicted, designed, synthesized, and purified by Shanghai Guixiong Biotech Company (Shanghai, China). Initially, the full-length amino acid sequence of FANCA was obtained using the NCBI database (https://www.ncbi.nlm.nih.gov/protein/NP_000126.2?report=fasta). Subsequently, the online databases such as NetMHC4.0 (https://services.healthtech.dtu.dk/services /NetMHC-4.0/) and SYFPEITHI (http://www.syfpeithi.de/0-Home.htm) were utilized to predict HLA-A2-restricted FANCA epitopes. The binding affinity was predicted using the NetMHCcons server. A peptide was identified as a strong binder if its %Rank (rank of the predicted affinity when compared to a set of 400,000 random natural peptides) was below 0.5%, or the binding affinity (IC50) was below 50 nM [17]. SYFPEITHI scores are based on binding motifs common to known naturally occurring epitopes. According to the SYFPEITHI scoring guidelines, a naturally expressed epitope should be rank in the top 2% of all peptide scores in least 80% of cases [18]. The top five HLA-A2-restricted 9 amino acid of FANCA epitope was predicted and designed (Table 2). The five FANCA peptides and the HLA-A2-restricted positive control peptide (CMV peptide: NLVPMVATV) were synthesized based on Fmoc solid-phase principle using the polypeptide synthesizer [19]. High-performance liquid chromatography (HPLC) was employed to purify the crude peptides obtained after synthesis. Subsequent identification and analysis of these peptides were conducted using mass spectrometry. A peptide was confirmed as the target if its measured molecular weight matched the theoretical value. The identified peptide solution was lyophilized and then was dissolved in either diethyl pyrocarbonate (DEPC) water ordimethyl sulfoxide (DMSO) before being stored at − 80 °C.
Table 2.
Information of five peptide candidates
| No | Amino acid sequences | At position | SYFPEITHI score | NetMHC4.0 Affinity(nM) | purity% | Molecular mass |
|---|---|---|---|---|---|---|
| P1 | LLLEAVWHL | 176 | 30 | 2.72 | 96.4 | 1093.32 |
| P2 | SLLEFAQYL | 146 | 27 | 4.18 | 99.7 | 1083.24 |
| P3 | GLAALAVHL | 782 | 26 | 22.13 | 95.5 | 864.5 |
| P4 | RLADLKVSI | 516 | 26 | 27.83 | 99.4 | 1014.23 |
| P5 | ALHFAIQQV | 1228 | 25 | 32.31 | 96.1 | 1026.19 |
T2 cells binding experiments
The HLA-A2 positive T2 cells, which have antigen-processing defects that facilitate the efficient loading of exogenous peptides, were utilized as an assay to evaluate the binding efficiency of candidate HLA-A2 peptides. T2 cells, at a density of 5 × 10^5 cells per well, were incubated with individual FANCA peptides (P1 to P5) or the CMV peptide across a range of concentrations: 2 µg/mL, 10 µg/mL, 50 µg/mL, 100 µg/mL, and 250 µg/mL. The incubation was performed in 48 well plates for 18 h at a temperature of 37 °C and 5% CO2, with a negative control group lacking peptide also included. Post-incubation, the mean fluorescence intensity (MFI) of HLA-A2 on the surface of T2 cells was measured after the addition of human HLA-A2-PerCP Cy5.5 antibodies (Becton Dickinson, USA). The measurements were conducted using a FACSCalibur flow cytometer (Becton Dickinson, USA). The binding affinity of the peptides to HLA-A2 molecules was quantified by the Fluorescence Index (FI), calculated as follows: FI = (MFI of each experimental group—MFI of the negative control) / MFI of the negative control. The FI values were interpreted as follows: FI > 1.5 indicated that the peptide had a high affinity for HLA-A2 molecules, 1.0 < FI ≤ 1.5indicated that the peptide had moderate affinity for the HLA-A2 molecule, and 0.5 < FI ≤ 1.0 indicated that the peptide had low affinity to the HLA-A2 molecule.
Isolation of human peripheral blood mononuclear cells
The peripheral blood samples of healthy volunteers were collected with their consent and approval from the ethics committee of the Fujian Provincial Hospital approval (K2023-03–010). Heparin-anticoagulant peripheral blood was diluted with an equal volume of PBS, and then carefully layered on top of an equal volume of Ficoll lymphocyte separation medium (Tianjing haoyang, China). The mononuclear cell layer was gently aspirated. The cells were then washed with PBS by centrifugation at 1400 rpm for 10 min at room temperature. The supernatant was discarded, and red blood cell lysis buffer (Solarbio, Beijing, China) was added to lyse red blood cells on ice for 15 min. Subsequently, the cells were resuspended in PBS, centrifuged at 1400 rpm for 10 min, and the supernatant was discarded. The cells were washed again with PBS, centrifuged at 1200 rpm for 10 min at room temperature, and the supernatant was discarded. Ultimately, the cells were resuspended in complete RPMI 1640 medium to yield a suspension of peripheral blood mononuclear cells (PBMCs).
Induction of mature DCs pulsed with peptides in vitro
In vitro DC induction was carried out following the established procedures as previously outlined [20]. PBMCs from HLA-A2 healthy donor were incubated at 37 °C overnight with 5% CO2. The non-adherent cells, which are predominantly lymphocytes, were cryopreserved for later use. In contrast, the Dendritic Cells (DCs), which had adhered to the culture surface, were harvested and seeded into 6-well plates at a concentration of 1 × 10^6 cells/mL. These DCs were then supplemented with a cytokine cocktail consisting of 100 ng/mL of recombinant human Granulocyte–Macrophage Colony-Stimulating Factor (rhGM-CSF) and 50 ng/mL of recombinant human Interleukin-4 (rhIL-4) (both from PeproTech, the United States). The DCs were observed, and the medium was changed at half volume every 2 days and supplemented with cytokines. On the 5th day, the cells were divided into control and experimental groups, with the following categorization: the P(1–3)-DC groups, where P1-P3 candidate peptides were co-incubated with the DCs respectively; the P + -DC group, where a CMV peptide was co-incubated with the DCs; and the NP-DC group, where no peptide was co-incubated with the DCs. The concentration of peptides in the experimental groups was 50 µg/mL. Additionally, all groups were treated with 50 ng/mL of recombinant human tumor necrosis factor-α (rhTNF-α) (PeproTech, the United States). After loading with peptides, the maturation of the DCs was assessed. The daily morphological changes of DCs were monitored using a microscope and photographed throughout the process.
FANCA peptides-induced cytotoxic T lymphocytes in vitro
Cytotoxic T lymphocytes (CTLs) were generated by co-culturing mature DCs loaded with specific peptides and autologous peripheral blood lymphocytes from HLA-A2-positive healthy donors. These lymphocytes were restimulated by mature DCs loaded with the relevant peptide at 7-day intervals, for a total of two rounds. Briefly, on day 7, autologous lymphocytes were first stimulated by autologous mature DC freshly loaded with FANCA peptides, and were cultured in complete RPMI 1640 medium supplemented with 50 ng/mL of recombinant human interleukin-2 (rhIL-2) (PeproTech, the United States). The DCs and lymphocytes were co-cultured at a ratio of 1:10. The culture medium was refreshed by replacing half of its volume with fresh medium every 48 h, along with the addition of essential cytokines. On day 14, the lymphocytes were re-stimulated with newly mature DCs, following the same protocol as the initial stimulation. On the 17 day, the CTLs were harvested for the subsequent experiments.
Enzyme-linked immunospot assay
The secretion of IFN-γ from CTLs against target cells was detected using an enzyme-linked immunospot (ELISPOT) assay with the IFN-γ ELISPOT kit (Dakewe, Shenzhen, China). CTLs, induced by peptide-loaded DCs, were added to ELISPOT plate per-coated with anti-human IFN-γantibody at a density of 1 × 10^6 cells/mL. Positive control wells (with PHA) and background control wells were included according to the kit instructions. After incubation at 37 °C with 5% CO2 for 36 h, cells were lysed by adding ice-cold water. Biotinylated anti-IFN-γ antibodies were then added to bind specifically to IFN-γ, followed by the addition of streptavidin-HRP to form a secondary antibody complex. The color development was induced using the ACE solution. The number of spot-forming cells in each well was quantified using Mabtech Apex software, Version 1.1.45.114, in a blinded manner.
Lactate dehydrogenase release assay
The cytolytic activity of CTLs against target cell was measured using an LDH Cytotoxicity Assay Kit (Beyotime, Shanghai, China). Firstly, the expression of HLA-A2 on H1299 and PC9 cells was confirmed through by Cellosaurus database (https://www.cellosaurus.org/CVCL_B260). The presence of the FANCA protein in LUAD cell lines was also detected by Western blot analysis. Isolation of CD8 + T cells from CTLs, induced by peptide-loaded DCs, was performed using the EasySep™ Human CD8 Positive Selection Kit II (StemCell Technologies). LUAD cells, PC9 (HLA-A2 + , FANCA +), H1299 (HLA-A2-, FANCA +), served as target cells and were plated in 96-well plates at a density of 2 × 10^4 cells/mL. CD8 + T lymphocytes were collected as effector cells. Effector and target cells were co-cultured at ratios of 10:1, 20:1, and 40:1 in experimental wells. Concurrently, target cell spontaneous LDH release wells and target cell maximum LDH release wells were established. After co-culture at 37 °C with 5% CO2 for 3 h, 10 μL of LDH releasing reagent was added to the target cell maximum LDH release wells. All the cells were further cultured for an additional hour. Subsequently, 80 μL supernatant from each well was taken and transferred to a new 96-well plate, and 40 μL of LDH detection solution was added. The plate was then incubated at room temperature in the dark for 30 min. Finally, OD was immediately measured at 490 nm using a microplate reader. The cytotoxicity rate was calculated as: (OD of experimental wells—OD of target cell spontaneous LDH release wells) / (OD of target cell maximum LDH release wells—OD of target cell spontaneous LDH release wells) × 100%.
T cell antigen epitope prediction
The X-ray crystal structures of HLA-A2(PDB code: 3v5k) were obtained from the Protein Data Bank. The protonation state of all the compounds was standardized to pH = 7.4, and their 3D structures were generated using Open Babel[21]. AutoDock Tools (ADT3) were applied to prepare and parametrize both the receptor protein and ligands. The docking grid documents were generated by AutoGrid of sitemap, and AutoDock Vina (1.2.0) was used for docking simulation[22, 23]. The optimal pose was selected to analysis interaction. Finally, the protein–ligand interaction figure was generated by PyMOL.
Flow cytometry analysis
The presence of HLA-A2 in peripheral blood of healthy volunteers and the expression of DC-related markers were assessed by flow cytometry. CD8 + T cells were isolated from PBMC of healthy volunteers using the EasySep™ Human CD8 Positive Selection Kit II (StemCell Technologies), following the manufacturer's protocol. The isolated cells were then resuspended in PBS at a concentration of 10^6 cells per tube. After that, 2 μL of HLA-A2-PerCP CY5.5 antibody (BD, the United States) was added to the CD8 + T cell samples. Dendritic cells were collected on the 3rd and 7th days of the DC induction process were centrifuged at 1000 rpm for 5 min at room temperature. The cells were washed twice with PBS and then resuspended in PBS at 10^6 cells per tube. Subsequently, the DCs were incubated with 5 μL each of CD80-PE antibody, CD83-APC antibody, and HLA-DR-PerCP Cy5.5 antibody (all from BD, the United States).
All treated cells above were incubated at 4 °C in the dark for 30 min, washed twice with PBS, and resuspended in 500 μL of PBS. Finally, the samples were analyzed using a FACSCalibur flow cytometer (Becton Dickinson, USA).
Statistical analysis
Except for T2 binding test, all experiments were performed three times independently. Statistical analyses were performed using SPSS 25.0 software package (SPSS Inc. Chicago, USA) and GraphPad Prism 9.0 (GraphPad Software, USA). Data are represented as mean ± standard deviation unless stated otherwise. The comparison between the two groups was performed by t test. The receiver operating characteristic (ROC) curves were used to assess the diagnostic value of the biological marker. P < 0.05 was statistically significant.
Results
The mRNA expression and prognosis of FANCA in LUAD
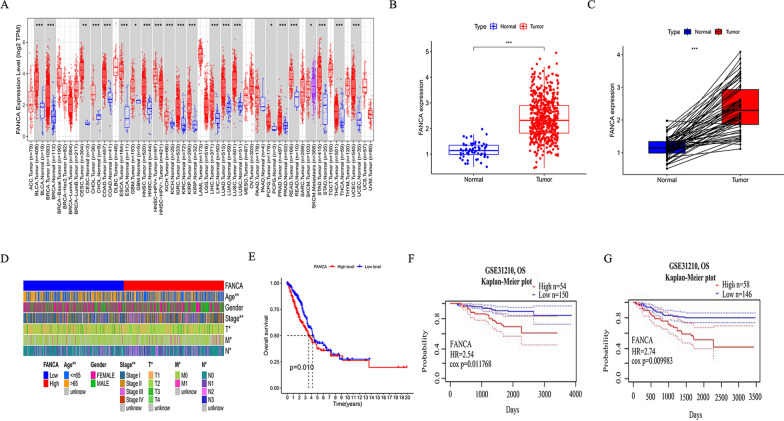
To determine the differential expression of FANCA between tumor and normal tissues, an analysis was performed on FANCA mRNA levels across various cancerous and normal tissues using the TIMER database. Relative to normal tissues, elevated FANCA expression was observed in a range of cancers, including LUAD (Fig. 1A). Further exploration of FANCA mRNA expression levels in LUAD tissues was conducted using the TCGA database. As shown in Fig. 1B, FANCA mRNA expression was significantly higher in 538 LUAD tissues compared to 59 paracancerous tissues (P < 0.001). This increase was also evident in 59 pairs of LUAD tissues and their adjacent non-cancerous counterparts (P < 0.001, Fig. 1C). Subsequently, the potential role of FANCA in LUAD progression was investigated by examining the correlation between FANCA expression and clinicopathological parameters. The mRNA levels of FANCA were significantly associated with age (P < 0.01), pathologic stage (P < 0.01), T stage (P < 0.01), M stage (P < 0.05), and N stage (P < 0.05) (Fig. 1D). Additionly, higher FANCA expression was correlated with poorer overall survival (OS) in LUAD, as indicated by the TISIDB database (Fig. 1E). This correlation was further supported by the PrognoScan database, which demonstrated that high FANCA expression was significantly linked to both poor OS and relapse-free survival (RFS) with hazard ratios (HR) of 2.54 for OS (Cox P-value = 0.011768, Fig. 1F) and 2.74 for RFS (Cox P-value = 0.009983, Fig. 1G).
Fig. 1.
Analysis the mRNA level and prognosis of FANCA in lung adenocarcinoma. A The mRNA expression levels of FANCA in pan-cancer using TIMER database. Comparison of FANCA mRNA expression in lung adenocarcinom with adjacent normal lung tissues (B) and matched adjacent normal lung tissues C using TCGA database. D The correlation between FANCA expression and clinicopathological parameters was analyzed in TGGA database using R language. E The correlation between FANCA expression and OS was analyzed by TISIDB in LUAD. GSE31210 cohort was used to analyze the correlation between FANCA expression with OS (F) and RFS (G) in patients with LUAD by PrognoScan database (*P < 0.05, **P < 0.01, ***P < 0.001)
FANCA protein levels in LUAD patient serum and LUAD cells
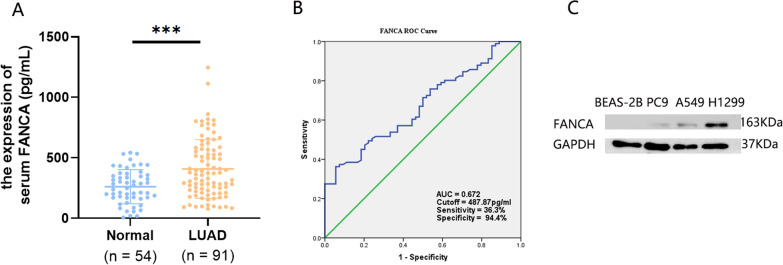
The protein levels of FANCA in LUAD patient serum were measured using ELISA. Compared with healthy individuals, an increase in FANCA protein levels was observed in LUAD patients (P < 0.05, Fig. 2A). Clinicopathological analysis revealed that FANCA expression was higher in male patients than in female patients and was positively correlated with tumor diameter, stage, and M stage (P < 0.05, Table 1). The receiver operating characteristic (ROC) curves revealed that FANCA demonstrated good performance, achieving a specificity of 94.4% and a sensitivity of 36.3% at a cut-off of 487.87 pg/ml (Fig. 2B). The protein levels of FANCA in LUAD cells were measured by Western blot analysis. As shown in Fig. 2C, FANCA protein expression levels were higher in LUAD cell lines compared to the normal human lung epithelial BEAS-2B cell line. Notably, the H1299 cell line exhibited particularly high expression of FANCA.
Fig. 2.
FANCA protein levels in LUAD patient serum and LUAD cells. A The protein level of FANCA in LUAD serum and healthy person’s serum. B ROC curve of serum FANCA. C The protein level of FANCA in LUAD cells
Table 1.
Correlation of FANCA protein expression and clinicopathological parameters
| Clinicopathological parameters | FANCA | ||
|---|---|---|---|
| n | Mean ± SEM | P value | |
| Age(yr) | 0.9904 | ||
| < 60 | 50 | 407.5 ± 32.45 | |
| ≥ 60 | 41 | 406.9 ± 40.71 | |
| Gender | 0.0176 | ||
| Male | 42 | 471.9 ± 39.71 | |
| Female | 49 | 351.8 ± 30.97 | |
| Diameter | 0.0092 | ||
| < 2 | 60 | 357.1 ± 27.79 | |
| ≥ 2 | 13 | 540.9 ± 73.47 | |
| Stage | 0.0240 | ||
| I | 64 | 370.1 ± 27.12 | |
| II + III + IV | 27 | 495.1 ± 53.84 | |
| T | 0.1618 | ||
| < 2 | 67 | 386.9 ± 28.23 | |
| ≥ 2 | 22 | 471.3 ± 59.38 | |
| N | 0.1451 | ||
| 0 | 63 | 383.4 ± 27.06 | |
| ≥ 1 | 26 | 466.7 ± 59.27 | |
| M | 0.0276 | ||
| 0 | 69 | 375.8 ± 26.53 | |
| ≥ 1 | 22 | 505.9 ± 60.88 | |
| CEA (ng/mL) | 0.1236 | ||
| < 5 | 68 | 367.0 ± 25.79 | |
| ≥ 5 | 17 | 461.8 ± 65.39 |
FANCA promotes the proliferation, migration and invasion of LUAD cells
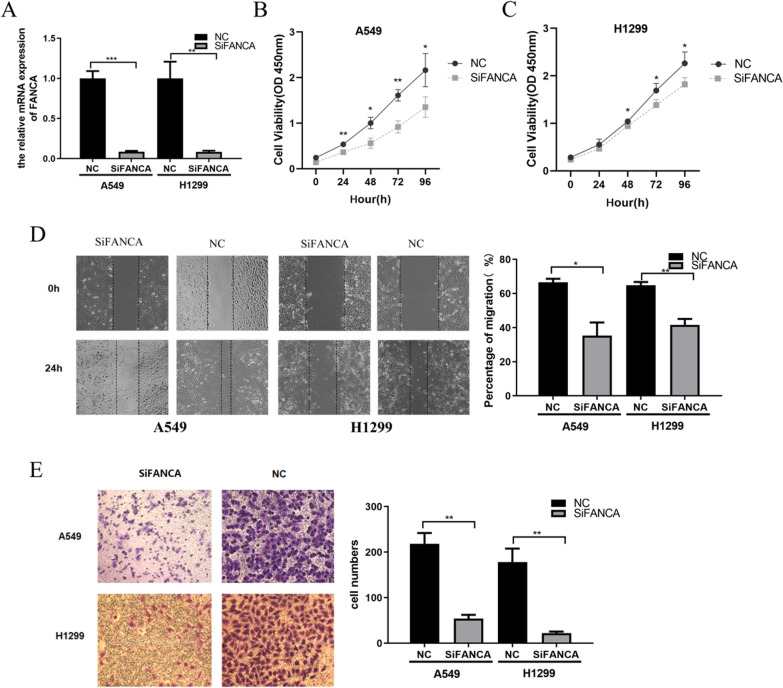
Elevated FANCA levels in LUAD tissues and serum suggest a potential role for FANCA as an oncogenic factor in LUAD. To test this hypothesis, CCK8, wound healing and transwell assays were performed following the knockdown of FANCA in A549 and H1299 cell lines. The results showed that the downregulation of FANCA significantly reduced the proliferative, migratory, and invasive capabilities of both cell lines (Figs. 3A–E).
Fig. 3.
FANCA promotes LUAD cell proliferation, migration and invasion. A Relative expression level of FANCA after siRNA transient knockdown in A549 and H1299 cells. B-C The low expression of FANCA suppressed A549 and H1299 cells proliferation by CCK8 viability analysis. D The low expression of FANCA significantly suppressed A549 and H1299 cells migration by wound healing experiment. EThe low expression of FANCA suppressed cell invasion by transwell assay. (*P < 0.05, **P < 0.01, ***P < 0.001)
FANCA expression is associated with the landscape of gene mutations
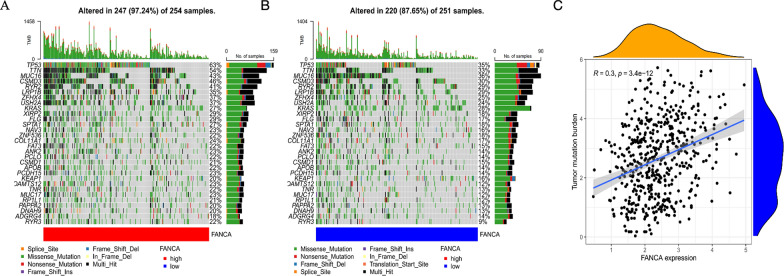
To evaluate the relationship between FANCA expression and somatic variants as well as tumor mutational burden (TMB) in LUAD patients, cases were classified into two subgroups based on the medain value of FANCA expression levels: a high-FANCA group with 245 cases and a low-FANCA group with 251 cases. The waterfall diagram, which visualized the integration status of somatic mutations in TCGA LUAD dataset, revealed that TP53 mutation was enriched in both the high-FANCA (63%) and the low-FANCA group (35%). In the high-FANCA group, the next four most frequently mutated genes were TTN (54%), MUC16 (43%), CSMD3 (46%), and RYP2 (35%). Whereas in the low-FANCA group, they were TTN (33%), MUC16 (36%), CSMD3 (30%), and RYR2 (29%) (Fig. 4A, B). Furthermore, as shown in Fig. 4C, our analysis showed a positive correlation between FANCA expression and TMB (R = 0.3, P < 3.4e-12).
Fig. 4.
The relationship between FANCA expression and somatic variants and TMBscore of LUAD patients in TCGA database. The oncoPrint plots of gene mutant frequency in high (A) and low (B) FANCA expression group. Correlation between FANCA expression and Tumor Mutuation Burden (C)
FANCA correlates with infiltration of immune cells
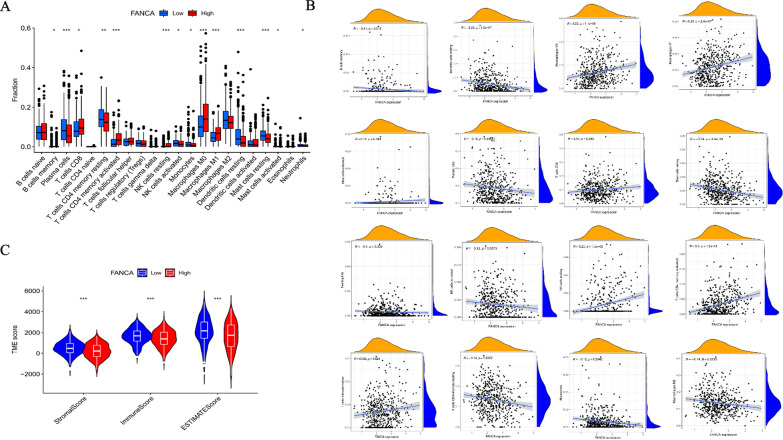
The differences in immune cell infiltration ratios between the low-FANCA and high-FANCA groups were investigated through the CIBERSORT tool. Our findings indicated that the high-expression group exhibited significantly higher levels of T cell CD8 (P < 0.005), T cells CD4 memory activated (P < 0.001), Macrophages M0 (P < 0.001), Macrophages M1(P < 0.001) and NK cells resting (P < 0.05) compared to low-expression group. Conversely, B cells memory (P < 0.05), Plasma cells (P < 0.001), T cells CD4 memory resting (P < 0.01), NK cells activated (P < 0.05), Monocytes(P < 0.05), Dendritic cells resting (P < 0.001), resting dendritic cell (P < 0.001), resting mast cell (P < 0.001), Mast cells resting (P < 0.001), Neutrophils (P < 0.05) were significantly decreased in the high-expression group (Figs. 5A). Additionally, Spearman's rank correlation analysis was employed to assess the potential correlation between FANCA and immune cell infiltration. The results demonstrated that FANCA expression was positively correlated with the levels of T cells follicular helper, T cells CD8, T cells CD4 memory activated, NK cells resting, Mast cells activated, Macrophages M1 and Macrophages M0. In contrast, FANCA expression was negatively associated with the levels of T cells CD4 memory resting, Mast cells resting, Dendritic cells resting, B cells memory, NK cells activated, Plasma cells, Monocytes, Macrophages M2 and neutrophils (Figs. 5B). Furthermore, the ESTIMATE method was applied to calculate the stromal score, immune score, and ESTIMATE score for both the high-expression and low-expression groups. The immune score, the stromal score, and ESTIMATE score were found to be lower in the high expression group than in the low expression group (Figs. 5C).
Fig. 5.
FANCA correlates with infiltration of immune cells. A Box plots of 22 immune cell infiltration levels between the high and low FANCA groups; B Correlation between FANCA expression and the immune cell infiltration levels; C Correlation between FANCA expression and the Stromal score, immune score and ESTIMATE score in the high and low expression groups
The prognostic value of FANCA according to immune cells in LUAD
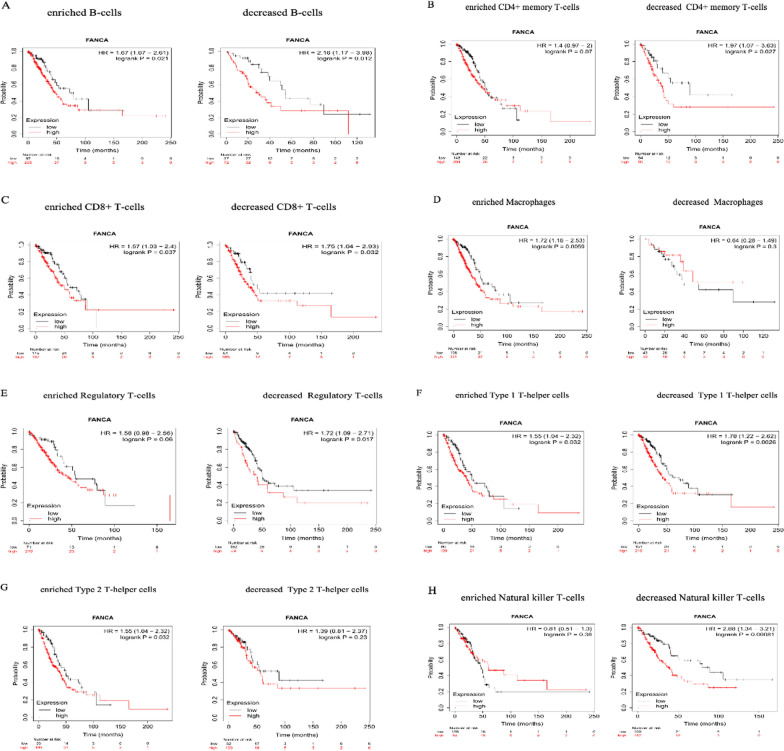
To investigated whether FANCA expression could directly affect the survival time of LUAD patients by altering the level of immune cell infiltration, the survival analysis based on FANCA expression levels within various immune cell subgroups in LUAD were conducted. The high expression of FANCA in enriched/decreased B cell, decreased CD4 + T cell, enriched/decreased CD8 + T cell, enriched macrophage, decreased natural killer T cell, decreased regulatory T cell (Treg), enriched/decreased type 1 T helper cell, and enriched type 2 T helper cell groups was correlated with the poor prognosis. Nevertheless, the relationship between FANCA and LUAD patient prognosis in the enriched CD4 + T cell, decreased macrophage, enriched regulatory T cell, enriched natural killer T cell and decreased type 2 T helper cell groups had no statistically significant difference (Figs. 6A–H).
Fig. 6.
Comparison of Kaplan–Meier survival curves of the high and low expression of FANCA in LUAD based on immune cell subgroups. A-H Relationships between FANCA of different immune cell subgroup and prognoses in in LUAD patients
Synthesis, prediction and screening of FANCA peptides
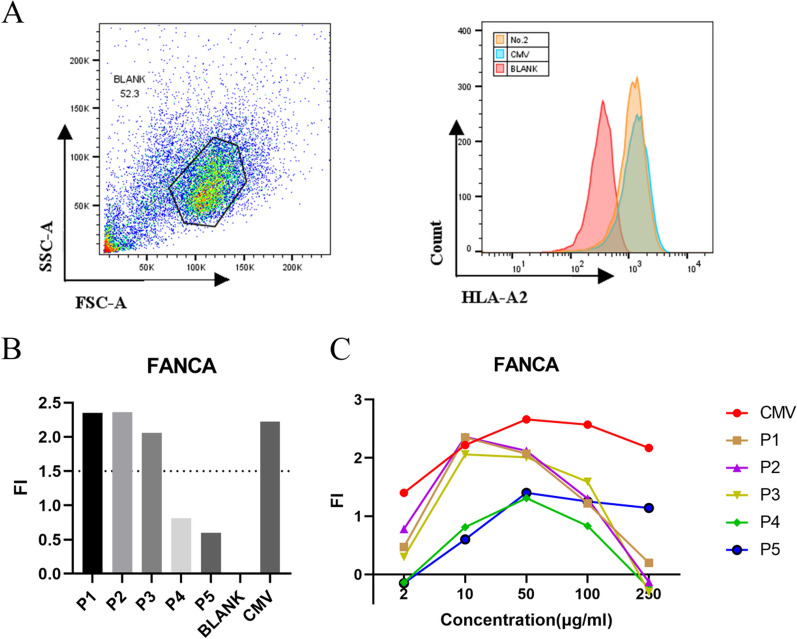
The full-length amino acid sequence of FANCA protein was retrieved from NCBI database. The prediction of HLA-A2-restricted FANCA peptides was conducted using SYFPEITHI and NetMHC4.0. The top five peptides, selected based on the highest composite scores, were designated as P1, P2, P3, P4, and P5 and were subsequently synthesized (Table 2). Following this, the HLA-A2 affinity of FANCA peptides (P1-P5) were evaluated by a T2 cell binding assay. As shown in Fig. 7A, the MFI of T2 cells increased significantly after the addition of peptides, indicating that the HLA-A2 binding peptides enhanced the stable expression of HLA-A2 molecules on the T2 cells. Among the five candidate peptides, three peptides (P1, P2, P3) exhibited high binding affinity (FI > 1.5). In contrast, peptides P4 and P5 demonstrated a lower binding affinity (Fig. 7B). At a concentration of 10 µg/mL, peptides P1, P2, and P3 demonstrated the highest FI. The optimal concentration for CMV was 50 µg/mL, where the FI was maximized (Fig. 7C). Therefore, peptides P1, P2, and P3 at a concentration of 10 µg/mL, as well as CMV at 50 µg/mL, were selected for subsequent experiments.
Fig. 7.
Screening of FANCA peptides by the T2 binding assay. A Representative multiparameter flow cytometry scatter diagram for T2 cell affinity detection. B Fluorescence intensity (FI) value of the five peptides at a concentration of 10 µg/mL. C FI values of these peptides across a range of concentrations
Identification of HLA-A2 positive healthy volunteers and induction of DCs in vitro
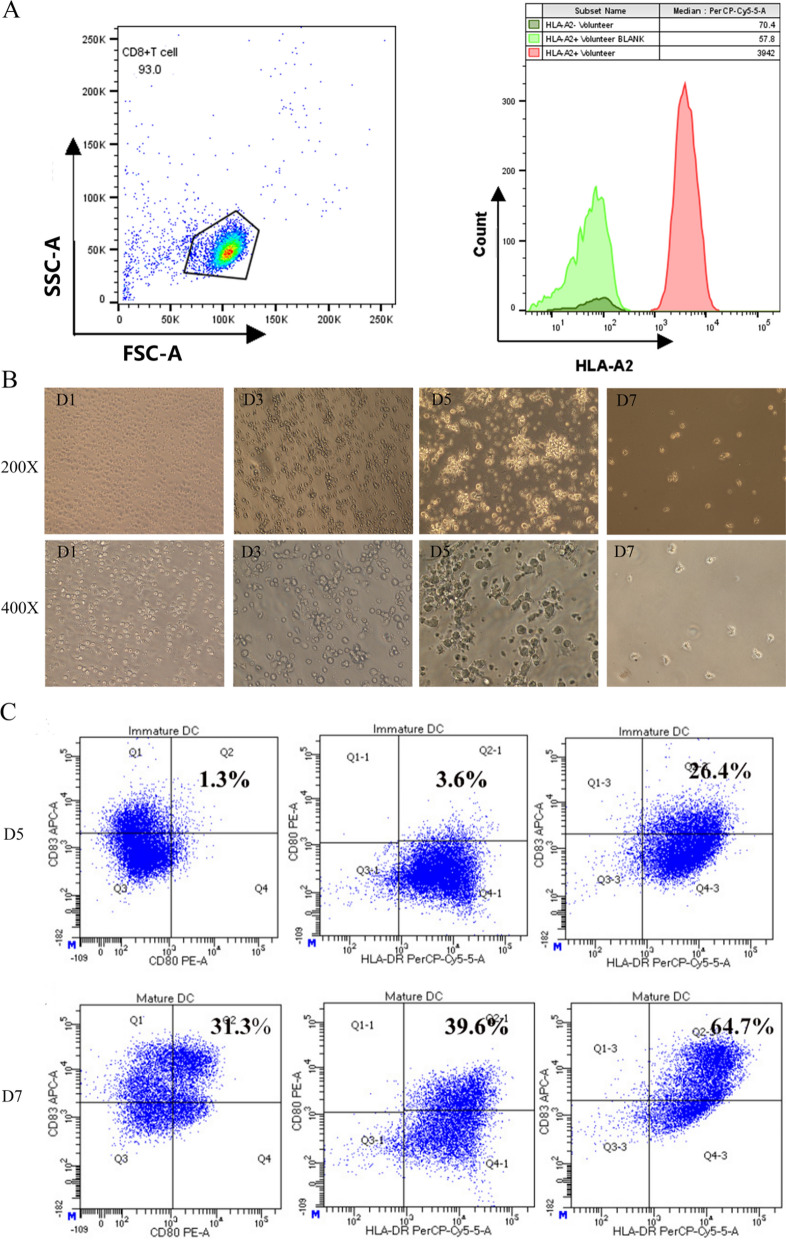
Healthy volunteers expressing HLA-A2 were identified and their peripheral blood was collected for further research (Fig. 8A). As shown in Fig. 8B, the morphological progression of DCs was monitored microscopically. On day 1, the cells appeared round or oval with a small volume. On day 3, some cells began to aggregate, exhibiting irregular morphology and an increase in volume. On day 5, a significant aggregation of DCs was observed, with the cells extending dendritic pseudopodia, creating an appearance reminiscent of a grape string in a semi-suspension state, indicative of immature DCs. On day 7, these cells had further matured, forming single suspended DCs. To assess the maturation of DCs, the expression levels of surface markers CD80, CD83 and HLA-DR were measured using flow cytometry on both day 5 and day 7. As illustrated in Fig. 8C, the maturation of DCs from day 5 to day 7 was characterized by a significant increase in the expression of costimulatory molecules. Specifically, there was a substantial increase in the percentage of CD80 + and CD83 + cells, from 1.3% to 31.3%. Additionally, the proportion of cells expressing both HLA-DR and CD80 saw a marked increase, from 3.6% to 39.6%. Similarly, the co-expression of HLA-DR and CD83 rose significantly, from 26.4% to 64.7%. Collectively, these observations indicate a functional maturation of DCs, thereby conferring upon them an augmented capacity for antigen presentation.
Fig. 8.
Identification of HLA-A2 positive healthy volunteers and induction of DCs in vitro. A Expression of HLA-A2 on the surface of CD8 + T cells in peripheral blood of healthy volunteers. B Morphological of dendritic cells which cultured in vitro. C Flow cytometry was utilized to analyze the expression levels of CD80, CD83, and HLA-DR as dendritic cells progress from an immature to a mature state
HLA-A2-restricted FANCA peptide P2-induced CTLs secrete IFN-γ and specifically lyse LUAD cell line PC9 in vitro
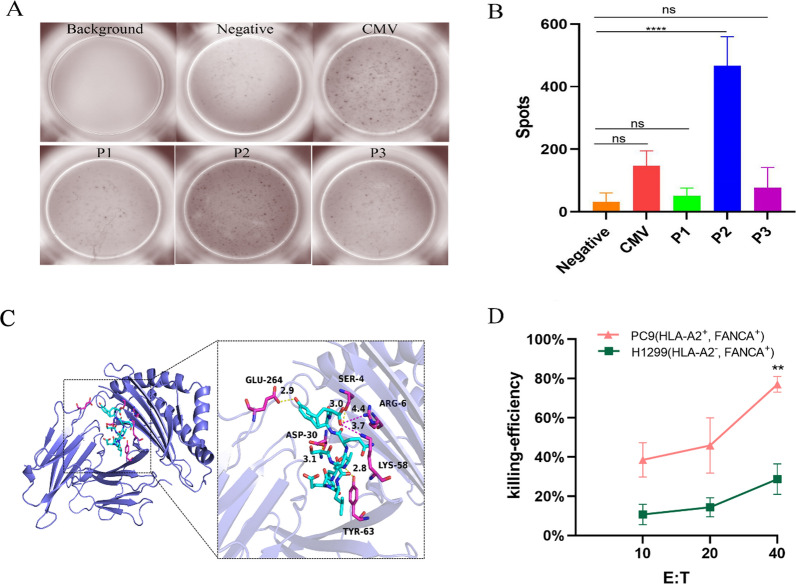
Considering the varying immunogenicity of the peptides, we assessed the immunostimulatory effects of three FANCA-derived peptides, P1, P2, and P3, using an ELISPOT assay. After 17 days of continuous culture, the formation of expanded lymphocyte clusters was observed microscopically. The results of ELISPOT assay, as shown in Fig. 9A, B, demonstrated that the positive control peptide CMV elicited lymphocyte secretion of over 150 IFN-γ spots, whereas the P2 peptide triggered lymphocyte secretion exceeding 400 IFN-γ spots. These findings indicate that P2 possesses the highest immunogenicity among the tested peptides.
Fig. 9.
HLA-A2-Restricted FANCA peptide P2-induced CTLs secrete IFN-γ and specifically lyse LUAD cell line PC9 in vitro. FANCA peptide-induced CTLs secrete IFN-γ in vitro. A Images result of the ELISPOT assay (B) Spot counts of the ELISPOT assay. Cytotoxic activity of FANCA peptide-induced CTLs to target cells detected by LDH release. C The interactions between FANCA protein and P2 ligand, all functional residues were identified and classified according to their interactions. D The killing efficiency of P2-induced specific CTLs against H1299 and PC9. (**P < 0.01, ****P < 0.0001)
To ensure the interaction between HLA molecules and our predicted potential epitopes, we performed molecular docking using autodock 4.0. To investigate the specific interaction between peptide P2 and HLA-A*0201 molecules, we performed molecular docking using AutoDock Tools. The electron analysis revealed that six binding bonds allow TL to exhibit considerable affinity for HLA-A201, and 9 amino acid residues of HLA-A201 are involved (SER4, ASP30, TYR63, GLU264, ARG6 and LYS58) (Fig. 9C). To further evaluate whether peptide P2 could induce CTL-specific killing to LUAD cell lines, we conducted a lactate dehydrogenase release assay. As shown in Fig. 9D, these CTLs induced by the HLA-A2-restricted FANCA peptide P2 demonstrated significant cytotoxicity against PC9 cell line, which increased with as the effector-to-target cell ratio increased. Notably, at an effector-to-PC9 cell (HLA-A2 + , FANCA +) ratio of 40:1, cytotoxicity reached nearly 80%. In comparison, the maximum cytotoxicity against H1299 cells (HLA-A2-, FANCA +) was less than 40%.
Discussion
Lung cancer is the leading type of malignant tumour among cancer-caused death worldwide, and the 5-year survival rate of lung cancer patients is only 18%[4]. The primary factors contributing to the unfavorable prognosis of lung cancer include the absence of a highly precise and sensitive biomarker for timely disease detection, as well as the effective drugs for lung cancer therapy[24]. Research on FANCA and its relationship with cancer has been an area of significant interest due to the protein's role in DNA repair and its implications in genomic stability[25]. FANCA mutations have been detected in a variety of cancers and found to be pro-carcinogenic[26–28]. In addition, FANCA has been reported to be involved in the occurrence and development of hepatocellular carcinoma and chronic lymphocytic leukemia[12, 29]. However, systematic research on the function and expression of FANCA in LUAD remains insufficient.
In the present study, our results underscored that high levels of FANCA in patients with LUAD were significantly correlated with worse OS survival, which was correlated with age, clinical stage, pathological T stage, M stage, and N stage in LUAD. Then, serum FANCA protein expression were evaluated by ELISA in total 54 healthy volunteers’ and 91 LUAD patients’ serum samples. The diagnostic value of FANCA in patients with LUAD was evaluated using the ROC curve, and with a specificity of 94.4% at a sensitivity of 36.3% and a cut-off value of 487.87 pg/ml, and were closely related to stage. In addition, knockdown of FANCA in A549 and H1299 cells significantly inhibited proliferation, metastasis, and invasion of LUAD cells in vitro. Recent studies showed that FANCA was associated with tumor progression and poor prognosis in patients with LIHC and this was consistent with our results [29]. All data revealed that FANCA might perform a critical function in the development and prediction of LUAD.
According to the FANCA expression in LUAD from TCGA database, we investigated the landscape of differential gene mutations and observed that more mutations were found in high FANCA group. Genomic changes may help to change the tumor microenvironment (TME), promote tumor progression and resistance to treatment[30]. The finding suggested that FANCA expression may be related to the tumor biological process. Another significant finding in this study was that FANCA expression was significantly associated with immunoreaction and diverse immune infiltration levels in LUAD. TME representsthe cellular environment of tumors and consists of the extracellular matrix, immune cells, and stromal cells[31]. Studies have suggested that the immune cell infiltration in the TME was closely related to LUAD progression and immunotherapeutic response in LUAD[32, 33]. Our findings suggested that FANCA expression was strongly positively associated with the degree of infiltration B cell, CD8 + T cell, CD4 + T cell, macrophage, neutrophil, and DC infiltration in LUAD. Furthermore, FANCA expression influenced the prognosis of LUAD patients by directly affecting immune cell infiltration by the Kaplan–Meier plotter analysis Thus, high expression of FANCA in LUAD may affect the prognosis of LUAD patients in part due to immune infiltration. Together these results demonstrated that FANCA expression was closely associated with immune infiltration and positively correlated with TMB, indicating that patients with FANCA were more suitable for immunotherapy.
DC vaccines induce T-cell activation, that is, CTL production, which has anti-tumor effects. As the principal effector cells within the cellular immune response, CTLs, often known as CD8 + T cells, can efficiently present antigens, thereby augmenting their capacity to eliminate tumor cells[34]. We have screened five potential antigenic peptides, and identified three high-affinity peptides that can bind HLA-A2 molecules in the T2 binding assay. The three peptides were able to induce the specific CTLs to secrete IFN-γ when encountering the cognate peptides. FANCA 146–154 (SLLEFAQYL), which showed the strongest immunogenicity, could induce specific CTLs that showed significant antitumor activity in vitro. The electron analysis revealed that six binding bonds allow TL to exhibit considerable affinity for HLA-A*0201, and 9 amino acid residues of HLA-A*0201 are involved (SER4, ASP30, TYR63, GLU264, ARG6 and LYS58). Finally, the SLLEFAQYL-specific CTLs can lyse the HLA-A2+ FANCA+ LUAD cell line PC9 cell in vitro. It can be inferred that FANCA 146–154 (SLLEFAQYL) antigenic peptide may exhibit a stronger affinity for DCs, and induce CTLs to produce stronger targeted killing ability for LUAD cells at an effector-to-target ratio of 40:1.
To sum up, our study firstly found the protein level of FANCA was elevated in LUAD tissue samples and was significantly associated with proliferation, metastasis, and invasion of LUAD cells. We uncovered that FANCA is involved in remolding LUAD immune infiltration and TME that is closely related to immunotherapy. We identified a dominant epitope P2 (SLLEFAQYL) that can effectively bind to HLA-A2. And the HLA-A2-restricted FANCA peptide P2 can induce CTLs to specifically lyse LUAD cell line. These results demonstrated that the elevation of FANCA promotes malignant phenotype of LUAD, and the potential peptide P2 (SLLEFAQYL) derived from FANCA may be used as an epitope vaccine for the treatment of LUAD.
Author contributions
The initial drafting of the manuscript was undertaken by CL and KY. Experimental procedures were conducted by KY, GY and ZR. YJ and GX contributed to providing algorithms and performing data analysis. Project administration was undertaken by FC. The preparation, writing, and fnal approval process before publication were all enriched by the valuable insights and comments provided by the authors.
Funding
This research was funded by Joint Funds for the innovation of science and Technology, Fujian province (Grant number: 2023Y9279); and Startup Fund for scientific research, Fujian Medical University (Grant number: 2020QH1177) and Sponsored by Fujian provincial health technology project (Grant number: 2022QNA004).
Availability of data and materials
The article and supplemental material encompass the dataset employed in our research. For any further inquiries, kindly reach out to the corresponding author directly.
Declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
All authors have agreed to publish the manuscript in this version.
Competing interests
The authors declared no competing interests that could potentially infuence or bias the outcomes of this research.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Yanli Kang and Ruifang Zhong have contributed equally to this work and share first authorship.
Contributor Information
Falin Chen, Email: Chenfalin008@126.com.
Liangyuan Chen, Email: liangyuan039083@163.com.
References
- 1.Thai AA, Solomon BJ, Sequist LV, Gainor JF, Heist RS. Lung cancer. Lancet. 2021;398(10299):535–54. 10.1016/S0140-6736(21)00312-3. [DOI] [PubMed] [Google Scholar]
- 2.Blandin Knight S, Crosbie PA, Balata H, Chudziak J, Hussell T, Dive C. Progress and prospects of early detection in lung cancer. Open Biol. 2017;7(9): 170070. 10.1098/rsob.170070. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Sorin M, Rezanejad M, Karimi E, Fiset B, Desharnais L, Perus LJM, Milette S, Yu MW, Maritan SM, Doré S, Pichette É, Enlow W, Gagné A, Wei Y, Orain M, Manem VSK, Rayes R, Siegel PM, Camilleri-Broët S, Fiset PO, Desmeules P, Spicer JD, Quail DF, Joubert P, Walsh LA. Single-cell spatial landscapes of the lung tumour immune microenvironment. Nature. 2023;614(7948):548–54. 10.1038/s41586-022-05672-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Denisenko TV, Budkevich IN, Zhivotovsky B. Cell death-based treatment of lung adenocarcinoma. Cell Death Dis. 2018;9(2):117. 10.1038/s41419-017-0063-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Chen D, Sha H, Hu T, Dong S, Zhang J, Liu S, Cao H, Ma R, Wu Y, Jing C, Wang Z, Wu J, Feng J. Cytokine-induced killer cells as a feasible adoptive immunotherapy for the treatment of lung cancer. Cell Death Dis. 2018;9(3):366. 10.1038/s41419-018-0404-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Liu J, Shen H, Gu W, Zheng H, Wang Y, Ma G, Du J. Prediction of prognosis, immunogenicity and efficacy of immunotherapy based on glutamine metabolism in lung adenocarcinoma. Front Immunol. 2022;11(13): 960738. 10.3389/fimmu.2022.960738. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Yuan F, Qian L, Zhao X, Liu JY, Song L, D’Urso G, Jain C, Zhang Y. Fanconi anemia complementation group A (FANCA) protein has intrinsic affinity for nucleic acids with preference for single-stranded forms. J Biol Chem. 2012;287(7):4800–7. 10.1074/jbc.M111.315366. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Kimble DC, Lach FP, Gregg SQ, Donovan FX, Flynn EK, Kamat A, Young A, Vemulapalli M, Thomas JW, Mullikin JC, Auerbach AD, Smogorzewska A, Chandrasekharappa SC. A comprehensive approach to identification of pathogenic FANCA variants in Fanconi anemia patients and their families. Hum Mutat. 2018;39(2):237–54. 10.1002/humu.23366. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Huang JP, Lin J, Tzen CY, Huang WY, Tsai CC, Chen CJ, Lu YJ, Chou KF, Su YW. FANCA D1359Y mutation in a patient with gastric polyposis and cancer susceptibility: a case report and review of literature. World J Gastroenterol. 2018;24(38):4412–8. 10.3748/wjg.v24.i38.4412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Qian B, Leng W, Yan Z, Lu J, Chen S, Yi H, Jiang Z. Clinical benefit With PARP inhibitor for pathogenic germline FANCA-mutated relapsed epithelial ovarian cancer: a case report. Front Oncol. 2022;25(12): 778545. 10.3389/fonc.2022.778545. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Gheybi K, Jiang J, Mutambirwa SBA, Soh PXY, Kote-Jarai Z, Jaratlerdsiri W, Eeles RA, Bornman MSR, Hayes VM. Evaluating germline testing panels in southern african males with advanced prostate cancer. J Natl Compr Canc Netw. 2023;21(3):289-296.e3. 10.6004/jnccn.2022.7097. [DOI] [PubMed] [Google Scholar]
- 12.Bravo-Navas S, Yáñez L, Romón Í, Pipaón C. Elevated FANCA expression determines a worse prognosis in chronic lymphocytic leukemia and interferes with p53 function. FASEB J. 2019;33(9):10477–89. 10.1096/fj.201802439RR. [DOI] [PubMed] [Google Scholar]
- 13.Rifkin AS, Less EM, Wei J, Shi Z, Zheng SL, Helfand BT, Hulick PJ, Krantz SB, Xu J. Association of reported candidate monogenic genes with lung cancer risk. Clin Lung Cancer. 2023;24(4):313–21. 10.1016/j.cllc.2023.01.005. [DOI] [PubMed] [Google Scholar]
- 14.Wu X, Zhao J, Yang L, Nie X, Wang Z, Zhang P, Li C, Hu X, Tang M, Yi Y, Du X, Xia X, Guan Y, Yu Z, Gu W, Quan X, Li L, Shi H. Next-generation sequencing reveals age-dependent genetic underpinnings in lung adenocarcinoma. J Cancer. 2022;13(5):1565–72. 10.7150/jca.65370. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Sanada H, Seki N, Mizuno K, Misono S, Uchida A, Yamada Y, Moriya S, Kikkawa N, Machida K, Kumamoto T, Suetsugu T, Inoue H. Involvement of Dual Strands of miR-143 (miR-143-5p and miR-143-3p) and their target oncogenes in the molecular pathogenesis of lung adenocarcinoma. Int J Mol Sci. 2019;20(18):4482. 10.3390/ijms20184482. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Kang Y, You J, Gan Y, Chen Q, Huang C, Chen F, Xu X, Chen L. Serum and serum exosomal CircRNAs hsa_circ_0001492, hsa_circ_0001439, and hsa_circ_0000896 as diagnostic biomarkers for lung adenocarcinoma. Front Oncol. 2022;7(12): 912246. 10.3389/fonc.2022.912246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Karosiene E, Lundegaard C, Lund O, Nielsen M. NetMHCcons: a consensus method for the major histocompatibility complex class I predictions. Immunogenetics. 2012;64(3):177–86. 10.1007/s00251-011-0579-8. [DOI] [PubMed] [Google Scholar]
- 18.Dong HL, Sui YF. Prediction of HLA-A2-restricted CTL epitope specific to HCC by SYFPEITHI combined with polynomial method. World J Gastroenterol. 2005;11(2):208–11. 10.3748/wjg.v11.i2.208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Guzmán F, Gauna A, Roman T, Luna O, Álvarez C, Pareja-Barrueto C, Mercado L, Albericio F, Cárdenas C. Tea bags for Fmoc solid-phase peptide synthesis: an example of circular economy. Molecules. 2021;26(16):5035. 10.3390/molecules26165035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Thomas R, Shaath H, Naik A, Toor SM, Elkord E, Decock J. Identification of two HLA-A*0201 immunogenic epitopes of lactate dehydrogenase C (LDHC): potential novel targets for cancer immunotherapy. Cancer Immunol Immunother. 2020;69(3):449–63. 10.1007/s00262-020-02480-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.O’Boyle NM, Banck M, James CA, Morley C, Vandermeersch T, Hutchison GR. Open babel: an open chemical toolbox. J Cheminform. 2011;7(3):33. 10.1186/1758-2946-3-33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Eberhardt J, Santos-Martins D, Tillack AF, Forli S. AutoDock Vina 1.2.0: new docking methods, expanded force field, and python bindings. J Chem Inf Model. 2021;61(8):3891–8. 10.1021/acs.jcim.1c00203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Trott O, Olson AJ. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem. 2010;31(2):455–61. 10.1002/jcc.21334. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Huang S, Yang J, Shen N, Xu Q, Zhao Q. Artificial intelligence in lung cancer diagnosis and prognosis: current application and future perspective. Semin Cancer Biol. 2023;89:30–7. 10.1016/j.semcancer.2023.01.006. [DOI] [PubMed] [Google Scholar]
- 25.Peake JD, Noguchi E. Fanconi anemia: current insights regarding epidemiology, cancer, and DNA repair. Hum Genet. 2022;141(12):1811–36. 10.1007/s00439-022-02462-9. [DOI] [PubMed] [Google Scholar]
- 26.Pawlikowska P, Delestré L, Gregoricchio S, Oppezzo A, Esposito M, Diop MB, Rosselli F, Guillouf C. FANCA deficiency promotes leukaemic progression by allowing the emergence of cells carrying oncogenic driver mutations. Oncogene. 2023;42(37):2764–75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Bertola N, Degan P, Cappelli E, Ravera S. Mutated FANCA gene role in the modulation of energy metabolism and mitochondrial dynamics in head and neck squamous cell carcinoma. Cells. 2022;11(15):2353. 10.3390/cells11152353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Errazquin R, Sieiro E, Moreno P, Ramirez MJ, Lorz C, Peral J, Ortiz J, Casado JA, Roman-Rodriguez FJ, Hanenberg H, Río P, Surralles J, Segrelles C, Garcia-Escudero R. Generating new FANCA-deficient hnscc cell lines by genomic editing recapitulates the cellular phenotypes of fanconi anemia. Genes. 2021;12(4):548. 10.3390/genes12040548. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Huang FD, Zhong YP, Sun GY, Xu QJ, Xing ZY, Chen KH, Liao LS, Dong MY. Fanconi anemia complementary group A (FANCA) facilitates the occurrence and progression of liver hepatocellular carcinoma. Dig Dis Sci. 2024;69(3):1035–54. 10.1007/s10620-024-08282-3. [DOI] [PubMed] [Google Scholar]
- 30.Robinson DR, Wu YM, Lonigro RJ, Vats P, Cobain E, Everett J, Cao X, Rabban E, Kumar-Sinha C, Raymond V, Schuetze S, Alva A, Siddiqui J, Chugh R, Worden F, Zalupski MM, Innis J, Mody RJ, Tomlins SA, Lucas D, Baker LH, Ramnath N, Schott AF, Hayes DF, Vijai J, Offit K, Stoffel EM, Roberts JS, Smith DC, Kunju LP, Talpaz M, Cieślik M, Chinnaiyan AM. Integrative clinical genomics of metastatic cancer. Nature. 2017;548(7667):297–303. 10.1038/nature23306. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Bejarano L, Jordāo MJC, Joyce JA. Therapeutic targeting of the tumor microenvironment. Cancer Discov. 2021;11(4):933–59. 10.1158/2159-8290.CD-20-1808. [DOI] [PubMed] [Google Scholar]
- 32.Li Y, Jiang M, Aye L, Luo L, Zhang Y, Xu F, Wei Y, Peng D, He X, Gu J, Yu X, Li G, Ge D, Lu C. UPP1 promotes lung adenocarcinoma progression through the induction of an immunosuppressive microenvironment. Nat Commun. 2024;15(1):1200. 10.1038/s41467-024-45340-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Tang L, Chen Z, Yang J, Li Q, Wang S, Mo T, Zeng W, Ding H, Pan S. Single-cell and Bulk RNA-Seq reveal angiogenic heterogeneity and microenvironmental features to evaluate prognosis and therapeutic response in lung adenocarcinoma. Front Immunol. 2024;8(15):1352893. 10.3389/fimmu.2024.1352893. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Raskov H, Orhan A, Christensen JP, Gögenur I. Cytotoxic CD8+ T cells in cancer and cancer immunotherapy. Br J Cancer. 2021;124(2):359–67. 10.1038/s41416-020-01048-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The article and supplemental material encompass the dataset employed in our research. For any further inquiries, kindly reach out to the corresponding author directly.