ABSTRACT
Staphylococcus aureus is usually regarded as a bacterial pathogen due to its ability to cause multiple types of invasive infections. Nevertheless, S. aureus colonizes about 30% of the human population asymptomatically in the nares, either transiently or persistently, and can therefore be regarded a human commensal as well, although carriage increases the risk of infection. Whereas many facets of the infection processes have been studied intensively, little is known about the commensal lifestyle of S. aureus. Recent studies highlight the major role of the composition of the highly variable nasal microbiota in promoting or inhibiting S. aureus colonization. Competition for limited nutrients, trace elements, and epithelial attachment sites, different susceptibilities to host defense molecules and the production of antimicrobial molecules by bacterial competitors may determine whether nasal bacteria outcompete each other. This chapter summarizes our knowledge about mechanisms that are used by S. aureus for efficient nasal colonization and strategies used by other nasal bacteria to interfere with its colonization. An improved understanding of naturally evolved mechanisms might enable us to develop new strategies for pathogen eradication.
INTRODUCTION
The human body offers several distinct niches for specific microbiomes, variable consortia of bacterial communities. The skin, for instance, is primarily colonized by members of the genera Propionibacterium (now Cutibacterium), Corynebacterium, and Staphylococcus (1). A similar pattern of genera is found in the human nose, which is regarded as a transition zone from the dry skin to the moist, mucoid airways (2). This rather confined area, more specifically, the region from the anterior nasal vestibule to the posterior nasopharyngeal cavity, is the favored colonization site of Staphylococcus aureus. While S. aureus is a member of the normal nasal microbiome in ca. 30% of the human population, it can also become an aggressive, life-threatening pathogen (3). Nasal carriage is a major risk factor for S. aureus infections. Accordingly, S. aureus is eradicated from the nose in at-risk patients by treatment with the antibiotic mupirocin. Interestingly, a significant percentage of humans seem never to be colonized by S. aureus, for reasons that are currently unknown. Increasing evidence suggests that the composition of the nasal microbiome is an important factor for the exclusion of S. aureus from the nose (2).
This article summarizes our knowledge of S. aureus nasal colonization and its interaction with other bacterial species that share the same habitat. Understanding naturally occurring S. aureus elimination strategies might enable the development of new eradication approaches in the future.
S. AUREUS AS A HUMAN COMMENSAL AND PATHOGEN
The preferred habitat of S. aureus is the anterior nares. Approximately 20% of the human population is permanently colonized by S. aureus, and another 20% are considered to be noncarriers. The remaining 60% of the population belong to the group of intermittent carriers (4). Although several factors that affect the S. aureus carrier status have been described, why only certain people are colonized remains elusive. In general, S. aureus occurs particularly frequently in men (5) and in women using hormonal contraception (6). Moreover, diabetic (7), hospitalized, and dialysis patients (4) are more often colonized than healthy humans, and there is a higher carriage rate during childhood compared to adulthood (8). A negative correlation exists between smoking and S. aureus carriage (9). Seasonal differences, such as the transition from winter to spring, and temperature and pollen or dust levels, could also affect the nasal microbiome composition (10). Host genetics appear to have only a moderate impact on the shape of the microbiome (11).
S. aureus possesses an extensive arsenal of virulence factors and, therefore, the clinical manifestations of S. aureus diseases are diverse and range from rather mild soft skin and tissue infections to severe and life-threatening infections, such as pneumonia, osteomyelitis, endocarditis, and sepsis (12). Although nasal colonization by S. aureus is asymptomatic, it increases the risk of invasive infections, especially in immunocompromised and hospitalized patients (4, 13). Analysis of infecting strains revealed that it is usually the nasal strain of a patient that is responsible for subsequent infections (13, 14). Therefore, decolonization of nasal S. aureus serves as a very efficacious measure for prevention of invasive infections (13). Since S. aureus transmission from the nose to other parts of the body occurs mainly via hand contact, S. aureus usually disappears also from other body sites when it is eliminated from the nose (15, 16).
The high risk of S. aureus carriers to infection justifies widely applied mupirocin treatment to eradicate S. aureus from the nares of patients. Mupirocin interferes with the synthesis of bacterial proteins by reversibly binding to bacterial isoleucyl-tRNA (17). Application of a mupirocin ointment twice a day for 5 consecutive days typically leads to eradication of S. aureus from the nose and to a reduced risk of invasive S. aureus infections (14, 17). The success rate of mupirocin treatment reaches approximately 90%, yet resistance is increasing, reaching up to 30% in some parts of the United States. Hence, new antimicrobial compounds for S. aureus decolonization are urgently needed (17, 18). Recolonization by S. aureus frequently occurs a few weeks after the end of mupirocin treatment, probably as a result of S. aureus cells using the posterior vestibula as a hidden reservoir, which can hardly be reached by mupirocin treatment (19–21).
Resistance to several common antibiotics has increased in S. aureus in the past decades. Resistance to the entire class of β-lactam antibiotics, including methicillin, characterizes the epidemic methicillin-resistant S. aureus (MRSA) strains. However, multidrug-resistant MRSA strains, which have acquired resistance to additional antibiotics, such as erythromycin, clindamycin, ciprofloxacin, tetracycline, or mupirocin, have also been observed (22). MRSA infections used to be acquired in hospitals by patients and are called hospital-associated MRSA infections. However, in the past decade, increasing numbers of MRSA infections have been acquired outside the health care system by emerging MRSA clones referred to as community-associated MRSA. Many community-associated MRSA clones are much more virulent and transmissible than hospital-associated MRSA clones (23).
DIVERSITY OF THE HUMAN NASAL MICROBIOME
In addition to the genus Staphylococcus, which belongs to the phylum Firmicutes, the nasal microbiome also contains members of other phyla, in particular, Actinobacteria and Proteobacteria. Within these phyla, the genera Corynebacterium, Propionibacterium, Staphylococcus, and Moraxella are most common in the human nose (24). Other genera are less frequently found (11). Interestingly, the nasal cavities are also colonized by various anaerobic species (19).
To classify different microbiome compositions, seven community state types (CSTs) were defined according to the dominance of specific nasal species, genera, or families. CST1 is dominated by S. aureus, and CST2 is defined by the major occurrence of Enterobacteriaceae, such as Escherichia spp., Proteus. spp., and Klebsiella spp. Staphylococcus epidermidis dominates CST3, whereas Propionibacterium spp. are major colonizers in CST4. Corynebacterium is the dominating genus of CST5, and Moraxella spp. are most prevalent in CST6, whereas CST7 is defined by the predominance of Dolosigranulum pigrum. The most common nasal CST is CST4, followed by CST3 and CST1, whereas CST6 is the least prevalent (Table 1). Although the conspicuous presence of various species or genera defines the CSTs, these species can also be identified at lower abundances in other CSTs (11). Comparing the microbiome compositions at various body sites, it is obvious that the nasal bacterial composition overlaps with those of the oral cavity and skin and can therefore be regarded as a bridge between the microbiomes of these two body sites (24, 25).
TABLE 1.
Community state types of the human nosea
| Community state type | Dominated by | Prevalence (%) |
|---|---|---|
| CST1 | S. aureus | 12.4 |
| CST2 | Enterobacteriaceae, e.g., Escherichia spp. (11), Proteus spp. (11), Klebsiella spp. (11) | 9.0 |
| CST3 | S. epidermidis | 22.5 |
| CST4 | Propionibacterium spp., e.g., P. acnes (88) | 28.7 |
| CST5 | Corynebacterium spp., e.g., C. accolens (88), C. pseudodiptheriticum (88), C. propinquum (88) | 11.2 |
| CST6 | Moraxella spp., e.g., M. lacunata (88), M. nonliquefaciens (88) | 5.6 |
| CST7 | Dolosigranulum pigrum | 10.7 |
Seven major community state types were defined according to the abundance of dominant nasal bacterial species, genera, or families (11).
Human skin areas with mostly dry, sebaceous, or moist properties differ in the abundance of bacterial microbiome members, while the core species are largely the same (1). It remains unclear if the individual differences in the skin microbiomes can also be attributed to specific CSTs. S. epidermidis, Staphylococcus capitis, Staphylococcus hominis, and Staphylococcus warneri are the most abundant coagulase-negative staphylococci (CoNS) on the skin (1). S. aureus also occurs transiently on the skin of healthy humans, especially in skin areas of the axillae and perineum (26, 27). Notably, the skin of atopic dermatitis patients is often permanently colonized by S. aureus on inflamed and noninflamed skin parts (28).
CoNS not only colonize the human skin; they can also occasionally be detected in the nose (29, 30). Only S. epidermidis is a core microbiome member of both the skin and nose (29). In general, the nasal microbiome is less dense and less diverse than, for instance, the gut microbiome (31, 32). S. epidermidis is particularly prevalent on moist areas of the body, such as the axillae, inguinal and perineal areas, toe tissue, and conjunctiva, and is also a common resident of the anterior nares (29). In addition, a variety of other CoNS occupy specific regions of the skin. S. hominis and Staphylococcus haemolyticus are often found in the axillae and pubic areas (29). S. capitis can be preferentially isolated from the sebaceous glands on the forehead and scalp (29) and is also found on the skin and in the nares (33). Whereas Staphylococcus auricularis is part of the external ear microbiome, Staphylococcus saprophyticus often colonizes the rectum and genitourinary tract (29). Staphylococcus lugdunensis is also part of the human skin flora and is particularly prevalent in the pelvic and perineal regions, the groin, the axillae, and in the nail bed of the first toe. S. lugdunensis has been isolated from the human nose at an incidence rate of 10 to 26% (19, 29, 34, 35). S. warneri is typically found at a lower percentage than other CoNS in the nares and on the skin, preferably on the head, arms, and legs (33). Compared to the other CoNS, S. lugdunensis and S. saprophyticus are considered species with slightly higher pathogenicity (29).
Compared to species such as S. epidermidis, which can be present in the nose with multiple strains simultaneously, nasal S. aureus isolates typically belong only to one specific clone (36, 37). Notably, the number of S. aureus cells, which can be cultivated from carriers by nasal swabbing, varies from only a few to more than 107 per swab (38). Attempts to distinguish persistent from intermediate carriers by a single nasal swab indicated that the detection of more than 103 S. aureus organisms CFU/swab corresponds to a high probability of persistent carriage (39).
The anterior nares are regarded as the primary habitat of S. aureus (4), but a recent study showed that this species can be isolated with an even higher incidence from deeper (posterior) areas of the nasal cavity, suggesting that the posterior vestibule or the entire nasal vestibule may be the principle habitat of S. aureus (19). Further studies indicated that S. aureus can penetrate into the nasal tissue of healthy humans, and individual cells could be visualized even at the stratum basale of the nasal epithelium, which differs in structure between the anterior and posterior areas (40, 41). While the anterior nares are characterized by a keratinized, stratified squamous epithelium, the posterior nares are defined by pseudostratified, columnar ciliated epithelial cells (2, 41). In the anterior vestibule, the moist squamous epithelium on the septum adjacent to the nasal ostium harbors the highest number of S. aureus cells. By contrast, CoNS prefer the skin of the nasal septum and the anterior, hair-covered epidermal portion of the lateral wall (5).
METABOLISM IN THE HUMAN NOSE
The nose is an environment with a very low nutrient supply, which might be the reason for the low species diversity of this habitat. Human nasal secretions contain sodium chloride at concentrations found also in other body fluids and small amounts of potassium, magnesium, and phosphate, but they are low in potential nutrients such as sugars, amino acids, and other major building blocks (30). Based on the composition of nasal secretions, a synthetic nasal medium (SNM3) was established that enabled the efficient growth of S. aureus, whereas CoNS did not steadily grow in SNM3 (30). This finding indicates that S. aureus is better adapted to life in the human nose than many CoNS, which may use the human nose only as a temporary, but not preferred, habitat. S. aureus and many other nasal bacteria, such as S. epidermidis, Finegoldia magna, Propionibacterium acnes, and Streptococcus pyogenes, secrete proteases that degrade human proteins, such as albumin, lactoferrin, mucin, cytokeratin 10, and hemoglobin. These proteins are present in considerable amounts in human nasal secretions and might serve as nutrient sources for various bacteria (42–44). It is likely that secretory proteases produced by a specific bacterial strain generate peptides and amino acids that can be utilized by many other microbiome members and may have a broad impact on microbiome metabolism.
MECHANISMS OF NASAL EPITHELIAL ATTACHMENT
Efficient attachment mechanisms are a prerequisite for bacterial nasal colonizers to remain in tight contact with epithelial cells and resist clearance by mucociliar movement. S. aureus binds to fully keratinized, dead desquamated cells of the anterior nasal cavity and adheres also to live ciliated cells in the posterior nasal cavity. S. aureus uses different adhesion mechanisms depending on the specific epithelial characteristics at different parts of the nares (45–48).
The cell wall glycopolymer wall teichoic acid (WTA) mediates the initial attachment of S. aureus to epithelial cells, and WTA is crucial for S. aureus nasal colonization (41, 47, 49). WTA-deficient S. aureus mutants are limited in their ability to bind to nasal epithelial cells. In addition, mutations in the dltABCD operon, leading to a loss of d-alanine modification of WTA, also exhibit diminished epithelial binding capacities (47, 49). Similarly, the modification of WTA with N-acetylglucosamine (GlcNAc) in α- or β-configuration is essential for efficient binding to nasal epithelial cells, and a lack of WTA glycosylation significantly abolished the ability of S. aureus to colonize cotton rat nares in vivo (50).
The scavenger receptor class-F member 1 (SREC1) on nasal epithelia in the posterior nasal cavity is a target for the WTA of S. aureus and allows S. aureus binding to epithelial cells (46). The charged zwitterionic properties of the ribitol-phosphate repeating units of WTA are essential in SREC1 binding (51). The loss of the d-alanine modification of the WTA significantly reduces the SREC1 binding.
The adherence of S. aureus to the nasal epithelium is also mediated by cell wall-anchored proteins, which are involved in long-term persistence within the anterior nasal cavity. S. aureus expresses a variety of cell wall proteins, which belong to the group of MSCRAMMs (microbial surface components recognizing adhesive matrix molecules) (41). MSCRAMMs such as clumping factor B (ClfB), iron-regulated surface determinant A (IsdA), and serine-aspartate repeat-containing protein D (SdrD) play an important role in nasal colonization by S. aureus.
ClfB is a fibrinogen-binding protein produced by S. aureus that binds to the matrix proteins cytokeratin 10 and loricrin. These proteins are exposed on human squamous epithelial cells (48, 52, 53). IsdA, which is involved in heme uptake and iron acquisition (54), is expressed during human infections and is responsible for nasal colonization and survival of S. aureus on the skin. Similar to ClfB, IsdA interacts with cytokeratin 10 and loricrin, as well as with the extracellular matrix protein involucrin (48, 55). SdrD mediates adhesion to the human squamous epithelium by binding to desmoglein 1, a desmosomal cadherin that is mainly expressed by the epidermis and mucosa (56, 57) (Fig. 1). The connective function of desmoglein 1 is required for the epidermis to maintain its integrity and structure (57). In addition, the surface proteins SdrC and SasG promote adhesion to squamous epithelium cells, but their binding partners are still unknown (45, 58). Interfering with glycopolymer-receptor interaction might become a new strategy for controlling S. aureus colonization in the nose (46). Furthermore, ClfB could represent an ideal target molecule for new decolonization strategies (53). In this context, ClfB can be considered a promising component for the development of a vaccine that would also reduce nasal colonization by S. aureus (59).
FIGURE 1.
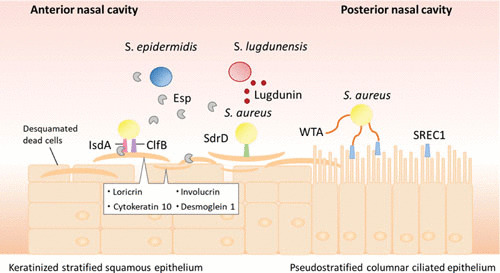
Attachment mechanisms of S. aureus in the human nasal cavity. The anterior and posterior parts of the human nose are lined by different types of epithelial cell, which require alternative bacterial adhesion mechanisms. For the keratinized stratified squamous epithelium in the anterior nasal cavity, S. aureus predominantly uses cell wall-attached surface proteins (MSCRAMMs) (2, 41). In contrast, the primary attachment in the posterior area, composed of a pseudostratified columnar ciliated epithelium, is mediated by specific interaction of the cell-wall linked wall teichoic acid (WTA) with the scavenger receptor class F member 1 (SREC1) (41, 46). The corneocytes (desquamated epithelial cells) in the anterior nasal cavity contain high levels of the proteins loricrin, cytokeratin 10, and involucrin (55). S. aureus can express a variety of cell wall proteins, which bind to these matrix proteins. The S. aureus adhesin clumping factor B (ClfB) binds to cytokeratin 10 and loricrin (48, 52), whereas the iron-regulated surface determinant A (IsdA) can also interact with involucrin (55). In addition, the S. aureus serine-aspartate repeat-containing protein D (SdrD) mediates adhesion to human squamous epithelial cells by binding to desmoglein 1 (56). Some S. epidermidis isolates secrete an extracellular serine protease (Esp), which inhibits S. aureus colonization by degradation of the surface proteins IsdA and SdrD and host receptors (77, 78). In addition, S. lugdunensis can prevent nasal colonization of S. aureus by producing the cyclic thiazolidine-containing peptide antibiotic lugdunin (35).
S. epidermidis colonizes the human nose at a higher frequency than S. aureus. In contrast to S. aureus, S. epidermidis has a different WTA structure, and adhesion proteins related to ClfB, IsdA, SdrD, and SdrC, are absent. Therefore, it remains unclear how S. epidermidis accomplishes adhesion in the nose (2, 60).
BACTERIAL COMPETITION
Bacteria from habitats with a limited nutrient supply such as the skin and nose have developed mechanisms to increase their fitness in competition with other microbiome members. Competition between bacterial species can be direct or indirect. Direct inhibition can be achieved, for instance, through production of antimicrobials, whereas indirect inhibition may occur via competition for nutrients or modification of living conditions (61).
Many nasal Staphylococcus isolates produce antimicrobial substances against bacterial competitors at an unexpectedly high frequency (84%), with S. epidermidis as the most frequent producer of antimicrobial activity (62). Importantly, production of many of the antibacterial activities is strongly enhanced or exclusively detectable under specific environmental stress conditions which are present in the human nose, such as hydrogen peroxide release and iron limitation (62). Antimicrobial substances, also called bacteriocins, are categorized into various groups and subgroups (63). Many Staphylococcus isolates are producers of lantibiotics, ribosomally synthesized antimicrobial peptides characterized by the presence of the thioether amino acids lanthionine and methyllanthionine. A variety of lantibiotics have been described for Staphylococcus strains (64). Epidermin (S. epidermidis) (65), Pep5 (S. epidermidis) (66), epilancin K7 and 15X (S. epidermidis) (67, 68), epicidin 280 (S. epidermidis) (69), staphylococcin C55 (S. aureus) (70), various nukacins (S. warneri [71], S. epidermidis [62], and S. hominis [72]), and lantibiotic-α and -β (S. hominis) (73), were documented. In addition, putative lantibiotic-biosynthetic gene clusters were found in the genome of S. capitis, which share homology with the biosynthetic systems of epidermin/gallidermin and the nonlantibiotic bacteriocin epidermicin (74). Lantibiotics are usually exclusively active against Gram-positive bacteria but often show no activity against S. aureus (62, 71). It is rather uncommon that nasal S. aureus is severely affected by lantibiotic-producing CoNS. Interestingly, a systematic analysis of P. acnes isolates revealed a frequent capacity to inhibit S. epidermidis (75). No compound has been identified that could explain this inhibitory activity; however, the genomes of various P. acnes strains encode a putative thiopeptide-biosynthetic gene cluster that is similar to those for antimicrobially active siomycin and berninamycin, which might explain the inhibitory effect (76).
A recent study showed that secretion of the extracellular serine protease Esp by S. epidermidis can efficiently inhibit S. aureus colonization. Artificial inoculation of Esp-secreting S. epidermidis to the nasal cavities of human volunteers was sufficient to eradicate S. aureus (77). Nasal colonization of S. aureus is probably abolished by Esp via the degradation of both bacterial adhesive surface proteins IsdA and SdrD and host receptor proteins (78). Although most S. epidermidis isolates produce Esp and some are producers of lantibiotics, there is no clear correlation of the absence of S. aureus with the presence of S. epidermidis (20, 64, 77).
Nasal S. lugdunensis can prevent colonization by S. aureus by producing an unusual antimicrobial compound termed lugdunin, which is a novel, cyclic thiazolidine-containing peptide antibiotic (35). The lugdunin gene cluster is encoded on the S. lugdunensis chromosome and is present in almost all S. lugdunensis strains. Lugdunin is generated by nonribosomal peptide synthetases. It has bactericidal activity against S. aureus and other Gram-positive bacteria, and it was shown to be active in vivo in animal models. Analysis of nasal microbiomes of hospitalized patients revealed that S. lugdunensis colonization is associated with a 6-fold reduced risk of S. aureus carriage in the nose, suggesting that lugdunin or lugdunin-producing commensals could become useful for the prevention of S. aureus colonization and infection.
Streptococcus spp. mainly occur in the oropharynx but can also be found at low frequency in the nostrils (79). Importantly, S. pneumoniae can modify the nasal habitat by releasing hydrogen peroxide, which induces the SOS response in S. aureus and activates DNA repair mechanisms as well as resident prophages. Subsequently, phage-produced lytic enzymes destroy S. aureus cells (80, 81). Hence, S. pneumonia is negatively correlated with nasal S. aureus carriage (82, 83).
An ambivalent correlation between Corynebacterium spp. and S. aureus was reported. While Corynebacterium accolens often occurs together with S. aureus, Corynebacterium pseudodiptheriticum is associated with the absence of S. aureus. A mutualistic relationship between C. accolens and S. aureus might rely on the joint mobilization of nutrients promoting the growth of these strains. In contrast, the competitive interaction between C. pseudodiptheriticum and S. aureus interferes with S. aureus colonization. Thus, the presence of C. accolens or C. pseudodiptheriticum might become a useful predictor of the propensity for nasal S. aureus carriage (20). A bacterial strain replacement study highlighted that nasal inoculation of persistent S. aureus carriers with a Corynebacterium sp. can lead to complete eradication of the pathogen in more than 70% of the probands (84).
Nostril- and skin-associated Propionibacterium spp. release coproporphyrin III, a porphyrin metabolite that promotes the aggregation and nasal colonization of S. aureus (85) (Fig. 2). A negative association with S. aureus carriage was reported for Simonsiella spp., D. pigrum, and F. magna, which are also regular members of the nasal microbiome (11, 31). The particular sensitivity of D. pigrum to a multitude of Staphylococcus isolates suggests that D. pigrum can only be present when no bacteriocin-producing Staphylococcus is in the same habitat (62).
FIGURE 2.
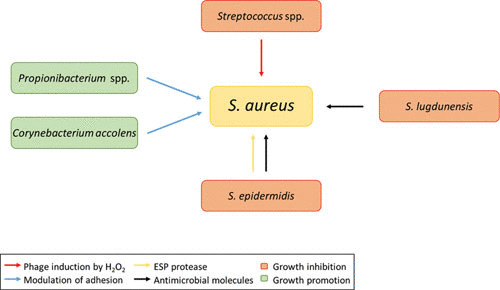
Established interactions between nasal bacteria. C. accolens and Propionibacterium spp. can promote the colonization by S. aureus (green boxes) by modulation of its adhesive capacities (20, 85) (blue arrows), while specific clones of S. epidermidis, S. lugdunensis, and Streptococcus spp. can lead to S. aureus growth inhibition (orange boxes). Some S. epidermidis isolates secrete high levels of extracellular serine protease (Esp), which inhibits S. aureus nasal colonization (77) (yellow arrow). In addition, S. epidermidis and S. lugdunensis can impede S. aureus colonization by producing antimicrobial molecules (35, 64) (black arrows). S. pneumoniae can release hydrogen peroxide, which leads to prophage activation in S. aureus along with phage-mediated lysis of S. aureus cells (80, 81) (red arrow).
COMPETITION BY INDUCTION OF HOST DEFENSE
In addition to the above-described competition scenarios, the human host can also impact the composition of the nasal microbiome. S. aureus expresses surfactant-like phenol-soluble modulin peptides, which mobilize proinflammatory lipoproteins from the staphylococcal cytoplasmic membrane. These lipoproteins activate the Toll-like receptor 2 and, consequently, lead to inflammation (86). The resulting inflammatory response in the nasal epithelium provokes the production of antimicrobial peptides. Although these hardly affect S. aureus, with its intrinsic immune evasion factors, they impair growth of other nasal commensal bacteria (5, 87).
CONCLUSION AND OUTLOOK
The mechanisms of microbiome-mediated exclusion of S. aureus from the human nose are probably multifactorial. It is likely that in addition to the processes described above, other factors are involved. It will be important to further investigate which other antimicrobial substances are produced by nasal commensals and which other strategies are used in their competition with S. aureus for nutrients and adhesion sites. New findings on nasal colonization may clarify why 20% of the human population is permanently colonized by S. aureus, while a similar percentage is never colonized. A better understanding could be helpful for developing new S. aureus eradication approaches that protect against recurrent S. aureus nasal colonization.
ACKNOWLEDGMENTS
Our research is supported by grants from the Deutsche Forschungsgemeinschaft (TRR34, TRR156, SFB766, SFB685, GRK1708, PE805/5-1), the Deutsches Zentrum für Infektionsforschung (TTU HAARBI), and the European Innovative Medicines Initiative (COMBACTE).
REFERENCES
- 1.Byrd AL, Belkaid Y, Segre JA. 2018. The human skin microbiome. Nat Rev Microbiol 16:143–155 10.1038/nrmicro.2017.157. [PubMed] 10.1038/nrmicro.2017.157 [DOI] [PubMed] [Google Scholar]
- 2.Krismer B, Weidenmaier C, Zipperer A, Peschel A. 2017. The commensal lifestyle of Staphylococcus aureus and its interactions with the nasal microbiota. Nat Rev Microbiol 15:675–687 10.1038/nrmicro.2017.104. [PubMed] 10.1038/nrmicro.2017.104 [DOI] [PubMed] [Google Scholar]
- 3.Tong SY, Davis JS, Eichenberger E, Holland TL, Fowler VG Jr. 2015. Staphylococcus aureus infections: epidemiology, pathophysiology, clinical manifestations, and management. Clin Microbiol Rev 28:603–661 10.1128/CMR.00134-14. [PubMed] 10.1128/CMR.00134-14 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Kluytmans J, van Belkum A, Verbrugh H. 1997. Nasal carriage of Staphylococcus aureus: epidemiology, underlying mechanisms, and associated risks. Clin Microbiol Rev 10:505–520. [PubMed] 10.1128/CMR.10.3.505 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Cole AM, Tahk S, Oren A, Yoshioka D, Kim YH, Park A, Ganz T. 2001. Determinants of Staphylococcus aureus nasal carriage. Clin Diagn Lab Immunol 8:1064–1069. [PubMed] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Zanger P, Nurjadi D, Gaile M, Gabrysch S, Kremsner PG. 2012. Hormonal contraceptive use and persistent Staphylococcus aureus nasal carriage. Clin Infect Dis 55:1625–1632 10.1093/cid/cis778. [PubMed] 10.1093/cid/cis778 [DOI] [PubMed] [Google Scholar]
- 7.Lipsky BA, Pecoraro RE, Chen MS, Koepsell TD. 1987. Factors affecting staphylococcal colonization among NIDDM outpatients. Diabetes Care 10:483–486 10.2337/diacare.10.4.483. [PubMed] 10.2337/diacare.10.4.483 [DOI] [PubMed] [Google Scholar]
- 8.Armstrong-Esther CA, Smith JE. 1976. Carriage patterns of Staphylococcus aureus in a healthy non-hospital population of adults and children. Ann Hum Biol 3:221–227 10.1080/03014467600001381. [PubMed] 10.1080/03014467600001381 [DOI] [PubMed] [Google Scholar]
- 9.Olsen K, Falch BM, Danielsen K, Johannessen M, Ericson Sollid JU, Thune I, Grimnes G, Jorde R, Simonsen GS, Furberg AS. 2012. Staphylococcus aureus nasal carriage is associated with serum 25-hydroxyvitamin D levels, gender and smoking status. The Tromsø Staph and Skin Study. Eur J Clin Microbiol Infect Dis 31:465–473 10.1007/s10096-011-1331-x. [PubMed] 10.1007/s10096-011-1331-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Camarinha-Silva A, Jáuregui R, Pieper DH, Wos-Oxley ML. 2012. The temporal dynamics of bacterial communities across human anterior nares. Environ Microbiol Rep 4:126–132 10.1111/j.1758-2229.2011.00313.x. [PubMed] 10.1111/j.1758-2229.2011.00313.x [DOI] [PubMed] [Google Scholar]
- 11.Liu CM, Price LB, Hungate BA, Abraham AG, Larsen LA, Christensen K, Stegger M, Skov R, Andersen PS. 2015. Staphylococcus aureus and the ecology of the nasal microbiome. Sci Adv 1:e1400216 10.1126/sciadv.1400216. [PubMed] 10.1126/sciadv.1400216 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Lowy FD. 1998. Staphylococcus aureus infections. N Engl J Med 339:520–532 10.1056/NEJM199808203390806. [PubMed] 10.1056/NEJM199808203390806 [DOI] [PubMed] [Google Scholar]
- 13.von Eiff C, Becker K, Machka K, Stammer H, Peters G, Study Group. 2001. Nasal carriage as a source of Staphylococcus aureus bacteremia. N Engl J Med 344:11–16 10.1056/NEJM200101043440102. [PubMed] 10.1056/NEJM200101043440102 [DOI] [PubMed] [Google Scholar]
- 14.Bode LG, Kluytmans JA, Wertheim HF, Bogaers D, Vandenbroucke-Grauls CM, Roosendaal R, Troelstra A, Box AT, Voss A, van der Tweel I, van Belkum A, Verbrugh HA, Vos MC. 2010. Preventing surgical-site infections in nasal carriers of Staphylococcus aureus. N Engl J Med 362:9–17 10.1056/NEJMoa0808939. [PubMed] 10.1056/NEJMoa0808939 [DOI] [PubMed] [Google Scholar]
- 15.Reagan DR, Doebbeling BN, Pfaller MA, Sheetz CT, Houston AK, Hollis RJ, Wenzel RP. 1991. Elimination of coincident Staphylococcus aureus nasal and hand carriage with intranasal application of mupirocin calcium ointment. Ann Intern Med 114:101–106 10.7326/0003-4819-114-2-101. [PubMed] 10.7326/0003-4819-114-2-101 [DOI] [PubMed] [Google Scholar]
- 16.Parras F, Guerrero MC, Bouza E, Blázquez MJ, Moreno S, Menarguez MC, Cercenado E. 1995. Comparative study of mupirocin and oral co-trimoxazole plus topical fusidic acid in eradication of nasal carriage of methicillin-resistant Staphylococcus aureus. Antimicrob Agents Chemother 39:175–179 10.1128/AAC.39.1.175. [PubMed] 10.1128/AAC.39.1.175 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Septimus EJ, Schweizer ML. 2016. Decolonization in prevention of health care-associated infections. Clin Microbiol Rev 29:201–222 10.1128/CMR.00049-15. [PubMed] 10.1128/CMR.00049-15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Antonov NK, Garzon MC, Morel KD, Whittier S, Planet PJ, Lauren CT. 2015. High prevalence of mupirocin resistance in Staphylococcus aureus isolates from a pediatric population. Antimicrob Agents Chemother 59:3350–3356 10.1128/AAC.00079-15. [PubMed] 10.1128/AAC.00079-15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Kaspar U, Kriegeskorte A, Schubert T, Peters G, Rudack C, Pieper DH, Wos-Oxley M, Becker K. 2016. The culturome of the human nose habitats reveals individual bacterial fingerprint patterns. Environ Microbiol 18:2130–2142 10.1111/1462-2920.12891. [PubMed] 10.1111/1462-2920.12891 [DOI] [PubMed] [Google Scholar]
- 20.Yan M, Pamp SJ, Fukuyama J, Hwang PH, Cho DY, Holmes S, Relman DA. 2013. Nasal microenvironments and interspecific interactions influence nasal microbiota complexity and S. aureus carriage. Cell Host Microbe 14:631–640 10.1016/j.chom.2013.11.005. [PubMed] 10.1016/j.chom.2013.11.005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Wertheim HFL, Verveer J, Boelens HAM, van Belkum A, Verbrugh HA, Vos MC. 2005. Effect of mupirocin treatment on nasal, pharyngeal, and perineal carriage of Staphylococcus aureus in healthy adults. Antimicrob Agents Chemother 49:1465–1467 10.1128/AAC.49.4.1465-1467.2005. [PubMed] 10.1128/AAC.49.4.1465-1467.2005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Otto M. 2012. MRSA virulence and spread. Cell Microbiol 14:1513–1521 10.1111/j.1462-5822.2012.01832.x. [PubMed] 10.1111/j.1462-5822.2012.01832.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.DeLeo FR, Otto M, Kreiswirth BN, Chambers HF. 2010. Community-associated meticillin-resistant Staphylococcus aureus. Lancet 375:1557–1568 10.1016/S0140-6736(09)61999-1. 10.1016/S0140-6736(09)61999-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Human Microbiome Project Consortium. 2012. Structure, function and diversity of the healthy human microbiome. Nature 486:207–214 10.1038/nature11234. [PubMed] 10.1038/nature11234 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Faust K, Sathirapongsasuti JF, Izard J, Segata N, Gevers D, Raes J, Huttenhower C. 2012. Microbial co-occurrence relationships in the human microbiome. PLOS Comput Biol 8:e1002606 10.1371/journal.pcbi.1002606. [PubMed] 10.1371/journal.pcbi.1002606 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Casewell MW, Hill RL. 1986. The carrier state: methicillin-resistant Staphylococcus aureus. J Antimicrob Chemother 18(Suppl A):1–12. [PubMed] 10.1093/jac/18.Supplement_A.1 [DOI] [PubMed] [Google Scholar]
- 27.Peacock SJ, de Silva I, Lowy FD. 2001. What determines nasal carriage of Staphylococcus aureus? Trends Microbiol 9:605–610 10.1016/S0966-842X(01)02254-5. 10.1016/S0966-842X(01)02254-5 [DOI] [PubMed] [Google Scholar]
- 28.Geoghegan JA, Irvine AD, Foster TJ. 2017. Staphylococcus aureus and atopic dermatitis: a complex and evolving relationship. Trends Microbiol 26:484–497. [PubMed] 10.1016/j.tim.2017.11.008 [DOI] [PubMed] [Google Scholar]
- 29.Becker K, Heilmann C, Peters G. 2014. Coagulase-negative staphylococci. Clin Microbiol Rev 27:870–926 10.1128/CMR.00109-13. [PubMed] 10.1128/CMR.00109-13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Krismer B, Liebeke M, Janek D, Nega M, Rautenberg M, Hornig G, Unger C, Weidenmaier C, Lalk M, Peschel A. 2014. Nutrient limitation governs Staphylococcus aureus metabolism and niche adaptation inthe human nose. PLoS Pathog 10:e1003862 10.1371/journal.ppat.1003862. [PubMed] 10.1371/journal.ppat.1003862 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Wos-Oxley ML, Plumeier I, von Eiff C, Taudien S, Platzer M, Vilchez-Vargas R, Becker K, Pieper DH. 2010. A poke into the diversity and associations within human anterior nare microbial communities. ISME J 4:839–851 10.1038/ismej.2010.15. [PubMed] 10.1038/ismej.2010.15 [DOI] [PubMed] [Google Scholar]
- 32.Quercia S, Candela M, Giuliani C, Turroni S, Luiselli D, Rampelli S, Brigidi P, Franceschi C, Bacalini MG, Garagnani P, Pirazzini C. 2014. From lifetime to evolution: timescales of human gut microbiota adaptation. Front Microbiol 5:587 10.3389/fmicb.2014.00587. [PubMed] 10.3389/fmicb.2014.00587 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Kloos WE, Schleifer KH. 1975. Isolation and characterization of staphylococci from human skin. II. Descriptions of four new species: Staphylococcus warneri, Staphylococcus capitis, Staphylococcus hominis, and Staphylococcus simulans. Int J Syst Bacteriol 25:62–79 10.1099/00207713-25-1-62. 10.1099/00207713-25-1-62 [DOI] [Google Scholar]
- 34.Bieber L, Kahlmeter G. 2010. Staphylococcus lugdunensis in several niches of the normal skin flora. Clin Microbiol Infect 16:385–388 10.1111/j.1469-0691.2009.02813.x. [PubMed] 10.1111/j.1469-0691.2009.02813.x [DOI] [PubMed] [Google Scholar]
- 35.Zipperer A, Konnerth MC, Laux C, Berscheid A, Janek D, Weidenmaier C, Burian M, Schilling NA, Slavetinsky C, Marschal M, Willmann M, Kalbacher H, Schittek B, Brötz-Oesterhelt H, Grond S, Peschel A, Krismer B. 2016. Human commensals producing a novel antibiotic impair pathogen colonization. Nature 535:511–516 10.1038/nature18634. [PubMed] 10.1038/nature18634 [DOI] [PubMed] [Google Scholar]
- 36.Hu L, Umeda A, Kondo S, Amako K. 1995. Typing of Staphylococcus aureus colonising human nasal carriers by pulsed-field gel electrophoresis. J Med Microbiol 42:127–132 10.1099/00222615-42-2-127. [PubMed] 10.1099/00222615-42-2-127 [DOI] [PubMed] [Google Scholar]
- 37.Hu L, Umeda A, Amako K. 1995. Typing of Staphylococcus epidermidis colonizing in human nares by pulsed-field gel electrophoresis. Microbiol Immunol 39:315–319 10.1111/j.1348-0421.1995.tb02207.x. [PubMed] 10.1111/j.1348-0421.1995.tb02207.x [DOI] [PubMed] [Google Scholar]
- 38.Warnke P, Devide A, Weise M, Frickmann H, Schwarz NG, Schäffler H, Ottl P, Podbielski A. 2016. Utilizing moist or dry swabs for the sampling of nasal MRSA carriers? An in vivo and in vitro study. PLoS One 11:e0163073 10.1371/journal.pone.0163073. [PubMed] 10.1371/journal.pone.0163073 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Verhoeven PO, Grattard F, Carricajo A, Lucht F, Cazorla C, Garraud O, Pozzetto B, Berthelot P. 2012. An algorithm based on one or two nasal samples is accurate to identify persistent nasal carriers of Staphylococcus aureus. Clin Microbiol Infect 18:551–557 10.1111/j.1469-0691.2011.03611.x. [PubMed] 10.1111/j.1469-0691.2011.03611.x [DOI] [PubMed] [Google Scholar]
- 40.Hanssen AM, Kindlund B, Stenklev NC, Furberg AS, Fismen S, Olsen RS, Johannessen M, Sollid JUE. 2017. Localization of Staphylococcus aureus in tissue from the nasal vestibule in healthy carriers. BMC Microbiol 17:89 10.1186/s12866-017-0997-3. [PubMed] 10.1186/s12866-017-0997-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Weidenmaier C, Goerke C, Wolz C. 2012. Staphylococcus aureus determinants for nasal colonization. Trends Microbiol 20:243–250 10.1016/j.tim.2012.03.004. [PubMed] 10.1016/j.tim.2012.03.004 [DOI] [PubMed] [Google Scholar]
- 42.Casado B, Pannell LK, Iadarola P, Baraniuk JN. 2005. Identification of human nasal mucous proteins using proteomics. Proteomics 5:2949–2959 10.1002/pmic.200401172. [PubMed] 10.1002/pmic.200401172 [DOI] [PubMed] [Google Scholar]
- 43.Tomazic PV, Birner-Gruenberger R, Leitner A, Obrist B, Spoerk S, Lang-Loidolt D. 2014. Nasal mucus proteomic changes reflect altered immune responses and epithelial permeability in patients with allergic rhinitis. J Allergy Clin Immunol 133:741–750 10.1016/j.jaci.2013.09.040. [PubMed] 10.1016/j.jaci.2013.09.040 [DOI] [PubMed] [Google Scholar]
- 44.Koziel J, Potempa J. 2013. Protease-armed bacteria in the skin. Cell Tissue Res 351:325–337 10.1007/s00441-012-1355-2. [PubMed] 10.1007/s00441-012-1355-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Corrigan RM, Miajlovic H, Foster TJ. 2009. Surface proteins that promote adherence of Staphylococcus aureus to human desquamated nasal epithelial cells. BMC Microbiol 9:22 10.1186/1471-2180-9-22. [PubMed] 10.1186/1471-2180-9-22 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Baur S, Rautenberg M, Faulstich M, Grau T, Severin Y, Unger C, Hoffmann WH, Rudel T, Autenrieth IB, Weidenmaier C. 2014. A nasal epithelial receptor for Staphylococcus aureus WTA governs adhesion to epithelial cells and modulates nasal colonization. PLoS Pathog 10:e1004089 10.1371/journal.ppat.1004089. [PubMed] 10.1371/journal.ppat.1004089 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Weidenmaier C, Kokai-Kun JF, Kristian SA, Chanturiya T, Kalbacher H, Gross M, Nicholson G, Neumeister B, Mond JJ, Peschel A. 2004. Role of teichoic acids in Staphylococcus aureus nasal colonization, a major risk factor in nosocomial infections. Nat Med 10:243–245 10.1038/nm991. [PubMed] 10.1038/nm991 [DOI] [PubMed] [Google Scholar]
- 48.Mulcahy ME, Geoghegan JA, Monk IR, O’Keeffe KM, Walsh EJ, Foster TJ, McLoughlin RM. 2012. Nasal colonisation by Staphylococcus aureus depends upon clumping factor B binding to the squamous epithelial cell envelope protein loricrin. PLoS Pathog 8:e1003092 10.1371/journal.ppat.1003092. [PubMed] 10.1371/journal.ppat.1003092 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Weidenmaier C, Peschel A. 2008. Teichoic acids and related cell-wall glycopolymers in Gram-positive physiology and host interactions. Nat Rev Microbiol 6:276–287 10.1038/nrmicro1861. [PubMed] 10.1038/nrmicro1861 [DOI] [PubMed] [Google Scholar]
- 50.Winstel V, Kühner P, Salomon F, Larsen J, Skov R, Hoffmann W, Peschel A, Weidenmaier C. 2015. Wall teichoic acid glycosylation governs Staphylococcus aureus nasal colonization. MBio 6:e00632-15 10.1128/mBio.00632-15. [PubMed] 10.1128/mBio.00632-15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Schade J, Weidenmaier C. 2016. Cell wall glycopolymers of Firmicutes and their role as nonprotein adhesins. FEBS Lett 590:3758–3771 10.1002/1873-3468.12288. [PubMed] 10.1002/1873-3468.12288 [DOI] [PubMed] [Google Scholar]
- 52.O’Brien LM, Walsh EJ, Massey RC, Peacock SJ, Foster TJ. 2002. Staphylococcus aureus clumping factor B (ClfB) promotes adherence to human type I cytokeratin 10: implications for nasal colonization. Cell Microbiol 4:759–770 10.1046/j.1462-5822.2002.00231.x. [PubMed] 10.1046/j.1462-5822.2002.00231.x [DOI] [PubMed] [Google Scholar]
- 53.Wertheim HF, Walsh E, Choudhurry R, Melles DC, Boelens HA, Miajlovic H, Verbrugh HA, Foster T, van Belkum A. 2008. Key role for clumping factor B in Staphylococcus aureus nasal colonization of humans. PLoS Med 5:e17 10.1371/journal.pmed.0050017. [PubMed] 10.1371/journal.pmed.0050017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Foster TJ, Geoghegan JA, Ganesh VK, Höök M. 2014. Adhesion, invasion and evasion: the many functions of the surface proteins of Staphylococcus aureus. Nat Rev Microbiol 12:49–62 10.1038/nrmicro3161. [PubMed] 10.1038/nrmicro3161 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Clarke SR, Andre G, Walsh EJ, Dufrêne YF, Foster TJ, Foster SJ. 2009. Iron-regulated surface determinant protein A mediates adhesion of Staphylococcus aureus to human corneocyte envelope proteins. Infect Immun 77:2408–2416 10.1128/IAI.01304-08. [PubMed] 10.1128/IAI.01304-08 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Askarian F, Ajayi C, Hanssen AM, van Sorge NM, Pettersen I, Diep DB, Sollid JU, Johannessen M. 2016. The interaction between Staphylococcus aureus SdrD and desmoglein 1 is important for adhesion to host cells. Sci Rep 6:22134 10.1038/srep22134. [PubMed] 10.1038/srep22134 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Hammers CM, Stanley JR. 2013. Desmoglein-1, differentiation, and disease. J Clin Invest 123:1419–1422 10.1172/JCI69071. [PubMed] 10.1172/JCI69071 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Roche FM, Meehan M, Foster TJ. 2003. The Staphylococcus aureus surface protein SasG and its homologues promote bacterial adherence to human desquamated nasal epithelial cells. Microbiology 149:2759–2767 10.1099/mic.0.26412-0. [PubMed] 10.1099/mic.0.26412-0 [DOI] [PubMed] [Google Scholar]
- 59.Schaffer AC, Solinga RM, Cocchiaro J, Portoles M, Kiser KB, Risley A, Randall SM, Valtulina V, Speziale P, Walsh E, Foster T, Lee JC. 2006. Immunization with Staphylococcus aureus clumping factor B, a major determinant in nasal carriage, reduces nasal colonization in a murine model. Infect Immun 74:2145–2153 10.1128/IAI.74.4.2145-2153.2006. [PubMed] 10.1128/IAI.74.4.2145-2153.2006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Winstel V, Liang C, Sanchez-Carballo P, Steglich M, Munar M, Bröker BM, Penadés JR, Nübel U, Holst O, Dandekar T, Peschel A, Xia G. 2013. Wall teichoic acid structure governs horizontal gene transfer between major bacterial pathogens. Nat Commun 4:2345 10.1038/ncomms3345. [PubMed] 10.1038/ncomms3345 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Brugger SD, Bomar L, Lemon KP. 2016. Commensal-pathogen interactions along the human nasal passages. PLoS Pathog 12:e1005633 10.1371/journal.ppat.1005633. [PubMed] 10.1371/journal.ppat.1005633 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Janek D, Zipperer A, Kulik A, Krismer B, Peschel A. 2016. High frequency and diversity of antimicrobial activities produced by nasal Staphylococcus strains against bacterial competitors. PLoS Pathog 12:e1005812 10.1371/journal.ppat.1005812. [PubMed] 10.1371/journal.ppat.1005812 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Lee H, Kim HY. 2011. Lantibiotics, class I bacteriocins from the genus Bacillus. J Microbiol Biotechnol 21:229–235. [PubMed] [PubMed] [Google Scholar]
- 64.Bierbaum G, Sahl HG. 2009. Lantibiotics: mode of action, biosynthesis and bioengineering. Curr Pharm Biotechnol 10:2–18 10.2174/138920109787048616. [PubMed] 10.2174/138920109787048616 [DOI] [PubMed] [Google Scholar]
- 65.Allgaier H, Jung G, Werner RG, Schneider U, Zähner H. 1986. Epidermin: sequencing of a heterodetic tetracyclic 21-peptide amide antibiotic. Eur J Biochem 160:9–22 10.1111/j.1432-1033.1986.tb09933.x. [PubMed] 10.1111/j.1432-1033.1986.tb09933.x [DOI] [PubMed] [Google Scholar]
- 66.Kaletta C, Entian KD, Kellner R, Jung G, Reis M, Sahl HG. 1989. Pep5, a new lantibiotic: structural gene isolation and prepeptide sequence. Arch Microbiol 152:16–19 10.1007/BF00447005. [PubMed] 10.1007/BF00447005 [DOI] [PubMed] [Google Scholar]
- 67.van de Kamp M, van den Hooven HW, Konings RN, Bierbaum G, Sahl HG, Kuipers OP, Siezen RJ, de Vos WM, Hilbers CW, van de Ven FJ. 1995. Elucidation of the primary structure of the lantibiotic epilancin K7 from Staphylococcus epidermidis K7. Cloning and characterisation of the epilancin-K7-encoding gene and NMR analysis of mature epilancin K7. Eur J Biochem 230:587–600 10.1111/j.1432-1033.1995.tb20600.x. [PubMed] 10.1111/j.1432-1033.1995.tb20600.x [DOI] [PubMed] [Google Scholar]
- 68.Ekkelenkamp MB, Hanssen M, Danny Hsu ST, de Jong A, Milatovic D, Verhoef J, van Nuland NA. 2005. Isolation and structural characterization of epilancin 15X, a novel lantibiotic from a clinical strain of Staphylococcus epidermidis. FEBS Lett 579:1917–1922 10.1016/j.febslet.2005.01.083. [PubMed] 10.1016/j.febslet.2005.01.083 [DOI] [PubMed] [Google Scholar]
- 69.Heidrich C, Pag U, Josten M, Metzger J, Jack RW, Bierbaum G, Jung G, Sahl HG. 1998. Isolation, characterization, and heterologous expression of the novel lantibiotic epicidin 280 and analysis of its biosynthetic gene cluster. Appl Environ Microbiol 64:3140–3146. [PubMed] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Navaratna MA, Sahl HG, Tagg JR. 1998. Two-component anti-Staphylococcus aureus lantibiotic activity produced by Staphylococcus aureus C55. Appl Environ Microbiol 64:4803–4808. [PubMed] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Sashihara T, Kimura H, Higuchi T, Adachi A, Matsusaki H, Sonomoto K, Ishizaki A. 2000. A novel lantibiotic, nukacin ISK-1, of Staphylococcus warneri ISK-1: cloning of the structural gene and identification of the structure. Biosci Biotechnol Biochem 64:2420–2428 10.1271/bbb.64.2420. [PubMed] 10.1271/bbb.64.2420 [DOI] [PubMed] [Google Scholar]
- 72.Wilaipun P, Zendo T, Okuda K, Nakayama J, Sonomoto K. 2008. Identification of the nukacin KQU-131, a new type-A(II) lantibiotic produced by Staphylococcus hominis KQU-131 isolated from Thai fermented fish product (Pla-ra). Biosci Biotechnol Biochem 72:2232–2235 10.1271/bbb.80239. [PubMed] 10.1271/bbb.80239 [DOI] [PubMed] [Google Scholar]
- 73.Nakatsuji T, Chen TH, Narala S, Chun KA, Two AM, Yun T, Shafiq F, Kotol PF, Bouslimani A, Melnik AV, Latif H, Kim JN, Lockhart A, Artis K, David G, Taylor P, Streib J, Dorrestein PC, Grier A, Gill SR, Zengler K, Hata TR, Leung DY, Gallo RL. 2017. Antimicrobials from human skin commensal bacteria protect against Staphylococcus aureus and are deficient in atopic dermatitis. Sci Transl Med 9:9 10.1126/scitranslmed.aah4680. [PubMed] 10.1126/scitranslmed.aah4680 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Kumar R, Jangir PK, Das J, Taneja B, Sharma R. 2017. Genome analysis of Staphylococcus capitis TE8 reveals repertoire of antimicrobial peptides and adaptation strategies for growth on human skin. Sci Rep 7:10447 10.1038/s41598-017-11020-7. [PubMed] 10.1038/s41598-017-11020-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Christensen GJM, Scholz CFP, Enghild J, Rohde H, Kilian M, Thürmer A, Brzuszkiewicz E, Lomholt HB, Brüggemann H. 2016. Antagonism between Staphylococcus epidermidis and Propionibacterium acnes and its genomic basis. BMC Genomics 17:152 10.1186/s12864-016-2489-5. [PubMed] 10.1186/s12864-016-2489-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Brzuszkiewicz E, Weiner J, Wollherr A, Thürmer A, Hüpeden J, Lomholt HB, Kilian M, Gottschalk G, Daniel R, Mollenkopf HJ, Meyer TF, Brüggemann H. 2011. Comparative genomics and transcriptomics of Propionibacterium acnes. PLoS One 6:e21581 10.1371/journal.pone.0021581. [PubMed] 10.1371/journal.pone.0021581 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Iwase T, Uehara Y, Shinji H, Tajima A, Seo H, Takada K, Agata T, Mizunoe Y. 2010. Staphylococcus epidermidis Esp inhibits Staphylococcus aureus biofilm formation and nasal colonization. Nature 465:346–349 10.1038/nature09074. [PubMed] 10.1038/nature09074 [DOI] [PubMed] [Google Scholar]
- 78.Sugimoto S, Iwamoto T, Takada K, Okuda K, Tajima A, Iwase T, Mizunoe Y. 2013. Staphylococcus epidermidis Esp degrades specific proteins associated with Staphylococcus aureus biofilm formation and host-pathogen interaction. J Bacteriol 195:1645–1655 10.1128/JB.01672-12. [PubMed] 10.1128/JB.01672-12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Lemon KP, Klepac-Ceraj V, Schiffer HK, Brodie EL, Lynch SV, Kolter R. 2010. Comparative analyses of the bacterial microbiota of the human nostril and oropharynx. MBio 1:e00129-10. [PubMed] 10.1128/mBio.00129-10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Selva L, Viana D, Regev-Yochay G, Trzcinski K, Corpa JM, Lasa I, Novick RP, Penadés JR. 2009. Killing niche competitors by remote-control bacteriophage induction. Proc Natl Acad Sci USA 106:1234–1238 10.1073/pnas.0809600106. [PubMed] 10.1073/pnas.0809600106 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Uehara Y, Kikuchi K, Nakamura T, Nakama H, Agematsu K, Kawakami Y, Maruchi N, Totsuka K. 2001. H(2)O(2) produced by viridans group streptococci may contribute to inhibition of methicillin-resistant Staphylococcus aureus colonization of oral cavities in newborns. Clin Infect Dis 32:1408–1413 10.1086/320179. [PubMed] 10.1086/320179 [DOI] [PubMed] [Google Scholar]
- 82.Bogaert D, van Belkum A, Sluijter M, Luijendijk A, de Groot R, Rümke HC, Verbrugh HA, Hermans PW. 2004. Colonisation by Streptococcus pneumoniae and Staphylococcus aureus in healthy children. Lancet 363:1871–1872 10.1016/S0140-6736(04)16357-5. 10.1016/S0140-6736(04)16357-5 [DOI] [PubMed] [Google Scholar]
- 83.Regev-Yochay G, Dagan R, Raz M, Carmeli Y, Shainberg B, Derazne E, Rahav G, Rubinstein E. 2004. Association between carriage of Streptococcus pneumoniae and Staphylococcus aureus in children. JAMA 292:716–720 10.1001/jama.292.6.716. [PubMed] 10.1001/jama.292.6.716 [DOI] [PubMed] [Google Scholar]
- 84.Uehara Y, Nakama H, Agematsu K, Uchida M, Kawakami Y, Abdul Fattah AS, Maruchi N. 2000. Bacterial interference among nasal inhabitants: eradication of Staphylococcus aureus from nasal cavities by artificial implantation of Corynebacterium sp. J Hosp Infect 44:127–133 10.1053/jhin.1999.0680. [PubMed] 10.1053/jhin.1999.0680 [DOI] [PubMed] [Google Scholar]
- 85.Wollenberg MS, Claesen J, Escapa IF, Aldridge KL, Fischbach MA, Lemon KP. 2014. Propionibacterium-produced coproporphyrin III induces Staphylococcus aureus aggregation and biofilm formation. MBio 5:e01286-14 10.1128/mBio.01286-14. [PubMed] 10.1128/mBio.01286-14 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Hanzelmann D, Joo HS, Franz-Wachtel M, Hertlein T, Stevanovic S, Macek B, Wolz C, Götz F, Otto M, Kretschmer D, Peschel A. 2016. Toll-like receptor 2 activation depends on lipopeptide shedding by bacterial surfactants. Nat Commun 7:12304 10.1038/ncomms12304. [PubMed] 10.1038/ncomms12304 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Riechelmann H, Essig A, Deutschle T, Rau A, Rothermel B, Weschta M. 2005. Nasal carriage of Staphylococcus aureus in house dust mite allergic patients and healthy controls. Allergy 60:1418–1423 10.1111/j.1398-9995.2005.00902.x. [PubMed] 10.1111/j.1398-9995.2005.00902.x [DOI] [PubMed] [Google Scholar]
- 88.Wos-Oxley ML, Chaves-Moreno D, Jauregui R, Oxley AP, Kaspar U, Plumeier I, Kahl S, Rudack C, Becker K, Pieper DH. 2016. Exploring the bacterial assemblages along the human nasal passage. Environ Microbiol 18:2259–2271. [PubMed] 10.1111/1462-2920.13378 [DOI] [PubMed] [Google Scholar]


