Abstract
Aging is characterized by a decline in various biological functions that is associated with changes in gene expression programs. Recent transcriptome-wide integrative studies in diverse organisms and tissues have revealed a gradual uncoupling between RNA and protein levels with aging, which highlights the importance of post-transcriptional regulatory processes. Here, we provide an overview of multi-omics analyses that show the progressive uncorrelation of transcriptomes and proteomes during the course of healthy aging. We then describe the molecular changes leading to global downregulation of protein synthesis with age and review recent work dissecting the mechanisms involved in gene-specific translational regulation in complementary model organisms. These mechanisms include the recognition of regulated mRNAs by trans-acting factors such as miRNA and RNA-binding proteins, the condensation of mRNAs into repressive cytoplasmic RNP granules, and the pausing of ribosomes at specific residues. Lastly, we mention future challenges of this emerging field, possible buffering functions as well as potential links with disease.
Keywords: Aging, Post-transcriptional Regulation, Tanslation, RNA, RNA-Binding Proteins
Subject terms: Translation & Protein Quality, RNA Biology
This review discusses the progressive uncoupling of transcriptomes and proteomes during healthy aging, and the underlying mechanisms of translation, mRNA and RNP regulation in various model organisms.

Introduction
Aging is associated with physiological changes that affect most biological functions and increase susceptibility to diseases. Distinguishing between “driver and passenger mechanisms of aging” (de Magalhaes, 2024) is a difficult task, but a number of cellular and molecular processes that are functionally contributing to the aging process have been defined (Lopez-Otin et al, 2023). Among the primary hallmarks of aging are epigenetic alterations that affect nuclear DNA methylation patterns, post-translational modifications of histones and chromatin remodeling and thus profoundly modify gene expression by altering transcription programs (Sen et al, 2016). While regulation of nuclear transcription is undoubtedly a critical step in the control of gene expression, other post-transcriptional mechanisms affecting pre-mRNA fate and outputs through control of isoform selection (Ule and Blencowe, 2019), RNA stability (Houseley and Tollervey, 2009), RNA localization (Das et al, 2021) or RNA translation (Kong and Lasko, 2012) have been described (Medioni and Besse, 2018). These regulatory steps are mediated by specialized cellular machineries (spliceosomes, ribosomes, exosomes, transport machineries and so on) recruited in trans to RNA molecules, but also involve cis-regulatory marks found in RNA molecules that modulate the recruitment and/or the efficiency of these machineries (Fig. 1). These marks can be RNA modifications (e.g., m6A methylation, RNA editing) (Roundtree et al, 2017), regulatory sequences located in the non-translated 5′ or 3’UTR region of mRNAs (e.g., binding sites for RNA-binding proteins or complementary noncoding RNAs) (Mayr, 2019) or specific coding sequence features (e.g., codon composition) (Hanson and Coller, 2018).
Figure 1. Overview of post-transcriptional regulatory mechanisms.
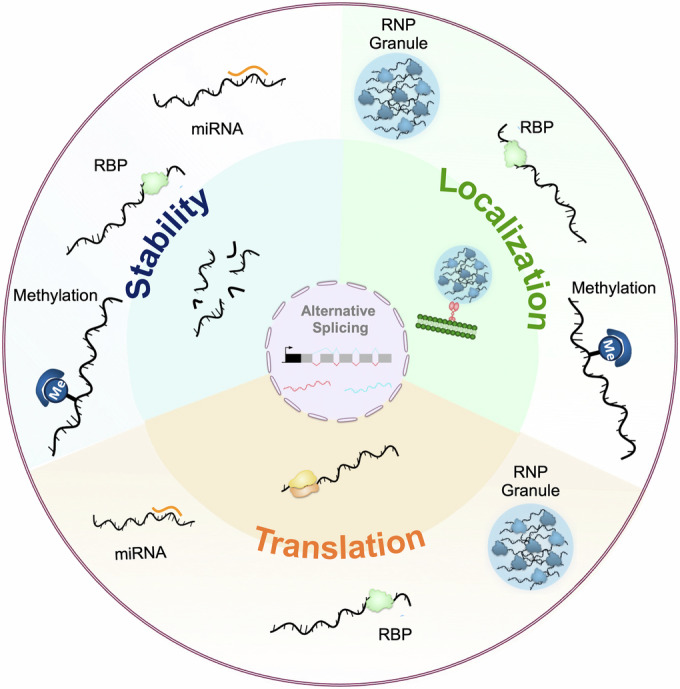
Once exported from the nucleus to the cytoplasm, mRNAs undergo post-transcriptional regulation at three different levels: stability, localization, and translation. These processes, represented in the inner circle, are modulated by trans-acting factors such as RNA-Binding Proteins (RBPs) or miRNAs, but also by epigenetic processes such as methylation or condensation into so-called ribonucleoproteic (RNP) granules (outer circle). RBPs regulate mRNA fate at different levels: translation, localization and stability. RBPs can have an opposite impact on translation by either promoting the recruitment of the translation machinery or preventing it through competition with translation initiation factors. Similarly, RBPs play a dual role on mRNA stability: they can protect mRNAs through capping and polyadenylation modulation or induce their cleavage and decay. Lastly, RBPs can recruit molecular motors to promote mRNA transport and remote translation. miRNAs hybridize to target mRNAs via complementary base pairing, leading to the recruitment of the RNA-induced silencing complex (RISC). This interaction can result in either translational repression or mRNA degradation. Methylation modifies the structure of RNA molecules and their binding affinity for RBPs, which can impact their stability or localization. RNP granules mediate the clustering of mRNA subsets into RBP-rich condensates that are depleted of the translation machinery and can be transported over long distances.
Despite expanding knowledge on the mechanisms controlling RNA fate, the role and regulation of post-transcriptional processes in the context of aging have so far been poorly investigated. However, recent large-scale integrative studies combining transcriptome, proteome and translatome profiling have revealed that aging is characterized by a significant loss of correlation between RNA and protein levels, indicating the importance of post-transcriptional regulatory processes in shaping the proteomes of aging organisms (Cellerino and Ori, 2017). Functional studies performed in a number of complementary model organisms (Box 1) have also underscored the contribution of these processes to age-related physiological changes. In this review, we describe recent work about the uncoupling between RNA and protein expression during aging and focus on the mechanisms contributing to age-dependent changes in RNA translation patterns. These mechanisms include both general modulation of the protein synthesis machinery and more specific processes that target selected subsets of transcripts.
Box 1 Common model organisms in aging studies.
The yeast Saccharomyces cerevisae is a unicellular eukaryotic organism whose genome size is ~12 Mb, with around 6000 protein-coding genes. In this model, aging is mostly defined as replicative aging and measured by the number of times a mother cell has divided. It can also be defined as chronological aging and measured by how long cells stays alive under nutrient deprivation (typically 1 week).
The nematode Caenorhabditis elegans has a genome size of ~97 Mb, with around 19,800 protein-coding genes. In this invertebrate model, age is defined as the number of days post-eclosion and the average lifespan is 2–3 weeks.
The fly Drosophila melanogaster has a genome size of ~180 Mb, with around 13,600 protein-coding genes. In this invertebrate model, age is defined as the number of days post-hatching and the average lifespan is 2–3 months.
S. cerevisiae, C. elegans, and Drosophila all are genetically tractable model systems in which gene function can easily be manipulated via diverse tools enabling tissue-specific gene overexpression or inactivation. They are also amenable to systematic genetics and drug screenings. Providing evolutionary support to the molecular mechanisms underlying aging, major genetic pathways involved in vertebrate aging, such as the Insulin or the mTor pathways, have been initially discovered and dissected in these models.
The turquoise killifish Nothobranchius furzeri has a genome size of ~1.2 Gb, with around 28,500 protein-coding genes. This vertebrate model attains sexual maturity within 3–4 weeks of hatching and has an extremely short lifespan of 4–6 months. It genetically tractable, although its generation time is 40 days.
The mice Mus musculus has a genome size of ~2.5 Gb, with around 30,000 protein-coding genes, 85% of them having orthologues in the human genome. This mammalian model attains sexual maturity within 4–7 weeks and lives for 2.5–3 years. This model is amenable to gene manipulations and editing. It possesses a developed tool kit for generating transgenics, knockins and knockouts, although generation time is 12 weeks. This model is used for screening of anti-aging drugs.
Beyond differences in lifespan, amenability to genetic manipulation, and transcriptome sizes, other parameters should be considered when studying post-transcriptional regulation in model organisms. Unicellular and invertebrate model organisms are particularly adapted to bulk transcriptome-wide studies where each datapoint represents an average population value, thus minimizing age-related inter-individual variabilities. Tissue-specific profiling cannot be done in unicellulars, and is easier to perform in vertebrate model organisms, where individual organs can easily be dissected and provide enough material for transcriptomics or proteomics.
Aging is characterized by a progressive uncoupling of RNA and protein levels
Recent multi-omics studies combining RNA-seq and Mass-Spectrometry on a range of model organisms (human, mouse, killifish, nematode, and so on) and tissues such as kidneys, heart, brain, etc, have uncovered that age-dependent changes in protein levels overall poorly correlate with changes in corresponding RNA levels (Di Fraia et al, 2024; Gerdes Gyuricza et al, 2022; Keele et al, 2023; Khatir et al, 2023; Ori et al, 2015; Takemon et al, 2021; Waldera-Lupa et al, 2014; Walther et al, 2015; Wei et al, 2015; Winsky-Sommerer et al, 2023). As demonstrated by comparing multiple time points, uncoupling of protein and RNA levels is much higher during aging than during development (Wei et al, 2015) and occurs progressively in the course of aging (Janssens et al, 2015; Kelmer Sacramento et al, 2020). It manifests as both positive uncoupling (increase in protein abundance relative to transcript level) and negative uncoupling (decrease in protein abundance relative to transcript level), the latter explaining up to 30–40% of the decoupling variance (Di Fraia et al, 2024; Wei et al, 2015).
Remarkably, combination of hierarchical clustering and definition of concordant and discordant classes of genes after analysis of RNA and protein expression levels in primate prefrontal cortex samples indicated that discordant gene classes tend to be enriched in regulatory and signaling functions. In addition, a strong conservation of mRNA and protein co-expression profiles was observed by comparing human and rhesus macaque samples, further suggesting functional importance (Wei et al, 2015). At the gene-specific level, uncoupling coefficients appeared to be highly heterogenous, leading to a decreased overall coordination between functionally related mRNAs. In the case of multi-molecular complexes, this translates into age-related alterations in the relative stoichiometry of constituent subunits (Gerdes Gyuricza et al, 2022; Janssens et al, 2015; Kelmer Sacramento et al, 2020). Together, a comparison of RNA and protein levels has highlighted the importance of post-transcriptional regulatory mechanisms in age-dependent alterations in gene expression.
Dysregulated proteostasis is a well-established hallmark of aging and could explain in part the observed RNA-protein uncoupling (Hipp et al, 2019; Kaushik and Cuervo, 2015; Lopez-Otin et al, 2023). The contribution of additional upstream mechanisms regulating translation efficiency has however emerged through two complementary transcriptome-wide assays providing comprehensive snapshots of tissue translatomes: polysome profiling, in which polysome-associated mRNAs are isolated through sucrose density fractionation before RNA-sequencing; and ribosome-sequencing (Ribo-seq), in which ribosome-associated mRNA fragments (or ribosome footprints) are specifically purified and sequenced (Brar and Weissman, 2015; Iwasaki and Ingolia, 2017). Comparison of Ribo-seq and proteomic data from young and aged rat brain and liver, for example, revealed the existence of hundreds of transcripts exhibiting significant changes in translation efficiency during aging and an overall good correlation between changes in protein levels and translation efficiency (Ori et al, 2015). An independent study performed in killifish brain also observed that variations in translational efficiency are better suited to explain changes in protein abundance than variations in RNA levels (Di Fraia et al, 2024), further suggesting the importance of translational regulation in aging. Of note, strong tissue-specific differences were seen when comparing gene expression profiles, both in terms of the identity of genes undergoing age-dependent changes and in terms of the extent of protein synthesis-dependent control, as up to 15% of the transcripts analyzed exhibited alteration in translation efficiency in the aging rat brain compared to only 2% in the liver (Ori et al, 2015). As further described below, it has become clear that age-dependent changes in translation efficiency result both from a general modification of the activity of the protein synthesis machinery and from translational regulation of selected subsets of transcripts (Fig. 2).
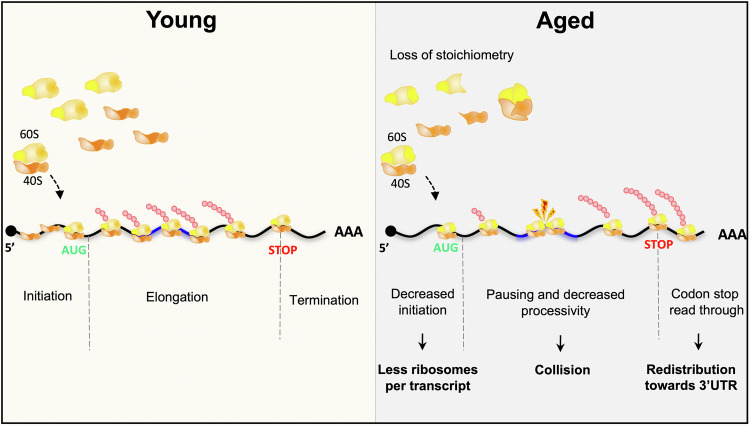
Figure 2. Age-dependent alterations in the efficiency of the translation machinery.
Schematic representation of the three main steps of elongation (left panel) and their alteration in aged individuals (right panel). During the initiation phase, the pre-initiation complex containing the small ribosomal subunit 40S scans the mRNA 5′ untranslated region (UTR) to find the start codon AUG. At this position, the large ribosomal subunit 60S is recruited to form a competent 80S ribosome. The loss of ribosomal protein stoichiometry observed with aging leads to a decreased initiation rate and a reduced number of 80S ribosome per transcript. During the elongation step, the polypeptide chain is synthesized through ribosome-mediated incorporation of amino acids via aminoacyl-tRNA codon recognition. In aged individuals, the rate of elongation, which reflects the processivity of the ribosome along the coding sequence, is decreased, thereby increasing the frequency of collisions. At the end of the coding sequence, the stop codon triggers the termination of translation and the dissociation of the ribosome from the mRNA. An increased frequency of STOP codon readthrough is observed in aging cells, inducing a partial relocalization of ribosomes to the 3’ UTR.
Global decline in protein synthesis with aging
Pioneer experiments performed in the 1970s and 80s via metabolic incorporation of radiolabeled amino acids in a wide range of cell-free systems and tissues from vertebrates (rat, mouse) or invertebrates (Drosophila) have shown a significant decrease in the rate of global protein synthesis with aging (Blazejowski and Webster, 1983; Dwyer et al, 1980; Fando et al, 1980; Kim and Pickering, 2023; Layman et al, 1976; Ward and Richardson, 1991; Webster and Webster, 1979). More recent ribosome profiling experiments, in which samples from replicatively aged yeast, old C. elegans or rat brains were sedimented on a sucrose gradient to separate free RNAs from ribosome-engaged RNAs, have confirmed these initial discoveries and demonstrated a marked decrease in the global amount of RNA associated with ribosomes in old compared to young cells (Fando et al, 1980; Hu et al, 2018; Motizuki and Tsurugi, 1992; Stein et al, 2022).
The number of ribosomes loaded per mRNA molecule is controlled by regulation of two main regulatory steps of translation (Hershey et al, 2012; Kim and Pickering, 2023; Yuan et al, 2024): (i) the rate-limiting initiation step, in which the small 40S ribosomal subunit is recruited to the mRNA and then engages into 5′UTR scanning until reaching an initiation codon whereby the large 60S subunit is recruited to assemble the translation-competent 80S initiation complex; and (ii) the elongation step in which codon-specific aminoacyl-tRNAs are sequentially recruited to the ribosome and their associated amino acids are covalently linked to the growing nascent peptide chain before translocation to the next codon. Thus, regulation of initiation and elongation rates can shift the balance between ribosome-free or monosome-associated RNAs (not or poorly translated RNAs) and polysome-associated RNAs (efficiently translated RNAs). Studies performed in different models have shown that both translation initiation and translation elongation are affected by aging (Anisimova et al, 2018; Kim and Pickering, 2023).
Decrease in translation initiation
Historical experiments performed in cell-free extracts from young and old mouse livers have pointed to a decreased activity of the 40S ribosomal subunit in forming the initiation complex with aging (Nakazawa et al, 1984). More recent estimation of the amount of monosome-engaged versus polysome-engaged RNAs, combined with global profiling of ribosome occupancy along transcripts, further demonstrated a strong reduction of translation initiation in vivo in aged animals, whether replicatively aged yeasts or >10-day-old C. elegans (Hu et al, 2018; Stein et al, 2022).
Decrease in the pool of functional ribosome subunits upon aging
Such a general decrease in translation initiation can be explained to a great extent by a loss of functional ribosomes, as the biogenesis of the protein synthesis machinery is altered in different ways during aging. First, levels of ribosomal proteins were found to be strongly downregulated in aging tissues of vertebrate and invertebrate models (Keele et al, 2023; Khatir et al, 2023; Ubaida-Mohien et al, 2019; Walther et al, 2015), a phenomenon that was identified as upstream and causal in a high-resolution systems-level model of aging, modeling the interdependence of gene expression changes occurring throughout replicative aging in yeast (Janssens et al, 2015). Second, decrease in protein levels is seen for many, but not all ribosomal proteins, leading to a progressive imbalance in the relative levels of the different ribosomal machinery constituents from yeast to mammals (Gerdes Gyuricza et al, 2022; Janssens et al, 2015; Keele et al, 2023; Kelmer Sacramento et al, 2020). Such a loss of ribosomal protein stoichiometry is associated with defects in ribosome assembly, as seen in size-exclusion chromatography experiments in which components of the ribosome co-eluted at lower-than-expected molecular weight in old killifish brain lysates (Kelmer Sacramento et al, 2020). Third, age-induced ribosomal protein imbalance creates a pool of orphan subunits at risk of aggregation. Consistent with this, various studies have shown that ribosomal proteins tend to accumulate in the insoluble fractions collected from different tissues and different organisms (Chen et al, 2024; David et al, 2010; Di Fraia et al, 2024; Harel et al, 2024; Kelmer Sacramento et al, 2020; Reis-Rodrigues et al, 2012), with a particularly high propensity to aggregate in organs such as the gut, the liver and the brain (Chen et al, 2024). Notably, the decrease in the pool of functional ribosomes may not only be explained by defective ribosome assembly, but also by defective ribosome recycling. Accumulation of isolated 3’UTR fragments loaded with ribosomes was indeed described in the aging rat brain in response to oxidative stress and altered translation termination (Sudmant et al, 2018). Furthermore, an increased frequency of stop codon readthrough was observed in aged Drosophila (Chen et al, 2020; Martinez-Miguel et al, 2021).
Decreased amount and/or activity of translation initiation factors
The decrease in translation efficiency observed during aging has been explained in parallel by changes in the activity and levels of translation initiation factors. For example, the amount of eIF2, a factor essential for recruiting the initiator Met-tRNAi to the 40S subunit and further assembly of the 43S pre-initiation complex, was shown to decline with age and to linearly scale with protein synthesis in rat liver, kidney, lung and brain (Kimball et al, 1992). Age-dependent changes in the phosphorylation patterns of distinct translation initiation factors were also observed in rat brains (Ori et al, 2015) and yeast (Hu et al, 2018). Phosphorylation of eIF2α, a process known to prevent regeneration of active eIF2 and thus reduce protein synthesis (Rowlands et al, 1988; Sonenberg and Hinnebusch, 2009), was in particular shown to be induced during yeast replicative aging in response to the activity of the Gcn2 kinase (Hu et al, 2018). eIF2α phosphorylation has in fact been observed in various biological aging models, including C. elegans (Derisbourg et al, 2021b) and mouse liver and kidney (Ladiges et al, 2000). It represents the main target of the Integrated Stress Response (ISR) pathway, whose age-related increase in activity was proposed to fine-tune translation in the context of aging (Derisbourg et al, 2021a).
eIF4E, a factor recognizing the 7-methyl-guanosine 5′ cap structure of mRNAs and required for assembly of the eIF4F complex and further recruitment of the 43S pre-initiation complex, is also under tight control (Hershey et al, 2012; Sonenberg and Hinnebusch, 2009). Work performed in C. elegans, for example, has shown that not only the levels, but also the availability of IFE-2, the worm somatic eIF4E, decrease with aging (Rieckher et al, 2018). Specifically, the observed accumulation of IFE-2 into P-bodies, cytoplasmic condensates concentrating RNA molecules and RNA-binding proteins involved in RNA decay and translational repression (Ripin and Parker, 2023), was proposed to sequester IFE-2, thus contributing to the general decrease in protein synthesis with aging (Rieckher et al, 2018).
Changes in translation elongation
Decreased efficiency of translation elongation has also been implicated in the overall reduction of protein synthesis during aging (Connors et al, 2008; Kim and Pickering, 2023). As revealed by comparing ribosome half transit time in cell extracts from young and aged rats, a significant decrease in the rate of peptide chain elongation was indeed observed in liver cells (Coniglio et al, 1979). Pulse-chase experiments performed in mice to estimate the in vivo rate of amino acid incorporation further confirmed this finding (Gerashchenko et al, 2021). In this work, the sequential application of harringtonine, an inhibitor that specifically blocks translation initiation and cycloheximide, an inhibitor that blocks translation elongation, was followed by Ribo-seq and reconstitution of ribosome coverage tracks, which revealed a near 20% decrease in the rate of elongation in the liver. Age-dependent decrease in the efficiency of peptide chain elongation was additionally observed in cell-free extracts prepared from Drosophila whole bodies (Webster and Webster, 1979), and attributed to a large extent to alteration in the binding of aminoacyl-tRNA to ribosomes (Webster and Webster, 1982).
As recruitment of aminoacyl-tRNAs to the translating ribosomes is mediated by the eukaryotic elongation factor 1 (eEF-1), age-dependent changes in the levels and activity of eEF-1 were analyzed. In Drosophila cell extracts, a marked decrease in the synthesis of eEF-1 was apparent as early as day 7, that is, prior to the main drop in cellular protein synthesis (Webster and Webster, 1983). In rat, eEF-1 activity was found to be 30–40% higher in young compared to old brain and liver (Moldave et al, 1979). In addition, the activity of eEF-2, another elongation factor involved in the translocation of the peptidyl-tRNA from the A- to the P-site of the ribosome, was found to undergo age-related changes in rat liver cell-free extracts. Specifically, the activity of the eEF-2 kinase and the phosphorylation level of its target eEF-2, both associated with a decreased activity of eEF-2 and a reduction in peptide elongation, were shown to be more than 2-fold higher in extracts from aged rats compared to young ones (Riis et al, 1993). Together, these changes participate in the general reduction in the efficiency of the protein synthesis machinery with age.
Mechanisms regulating the translation of specific subsets of transcripts with aging
In addition to the global decrease in translation efficiency observed with aging, specific signatures of mRNA-protein uncoupling were identified in transcriptome-wide sequencing studies (Di Fraia et al, 2024; Ori et al, 2015; Wei et al, 2015), pointing to gene-specific post-transcriptional regulatory processes. Although the mechanisms underlying such specific changes largely remain to be explored, emerging studies have highlighted the role of distinct molecular pathways, the regulation of which impacts on the expression of classes of downstream transcripts in the context of aging. For example, longitudinal Ribo-seq analyses performed on livers of up to 32-month-old mice indicated that mRNAs containing 5′-terminal oligopyrimidine (5′-TOP) regulatory sequences responsive to changes in the activity of the mTor pathway (Thoreen et al, 2012) exhibit particularly strong translational repression in aging (Anisimova et al, 2020). As described below, several other mechanisms involving noncoding RNAs (3.1), RNA-binding proteins (RBPs) and their condensation (3.2 and 3.3), or coding sequence properties (3.4) have been dissected in further detail (Fig. 3 and Table 1).
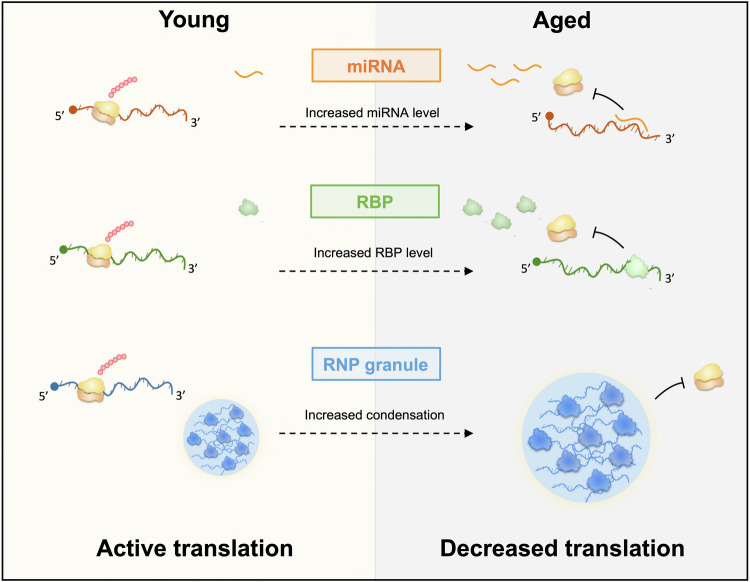
Figure 3. Trans-acting factors contributing to translational repression with aging.
Age-dependent increase in the level of trans-acting factors such as miRNA or RBPs (upper and middle parts, respectively) were shown to induce the translational repression of their target mRNAs. Condensation of RBPs and their associated mRNAs into bigger granules devoid of ribosomes (lower part) can induce their translational repression.
Table 1.
Table listing the cited trans-acting factors regulating translation, together with their associated targets and age-related phenotypes.
| Molecule | Regulation upon aging | Targets | Tissue | Age-associated phenotype | Species | References | |
|---|---|---|---|---|---|---|---|
| miRNA | mir-34 | Up | Eip74EF, Su(z)12, Pcl | Brain | Vacuolarization, polyglutamine aggregation, and neurodegeneration | Drosophila | Kennerdell et al, 2018; Liu et al, 2012 |
| Up | Atg9A | NA | Decreased autophagic flux | Nematode HEK293 Rat | Yang et al, 2013 | ||
| Up | PNUTS | Heart | Increased cardiomyocyte cell death | Mouse | Boon et al, 2013 | ||
| mir-188-3p | Up | ITGβ3 | endothelial cells | Decreased bone-type H capillary vessel number | Mouse | He et al, 2022 | |
| mir-29 | Up | IRP2 | Brain | Buffered intracellular iron content and oxidative stress | Killifish | Ripa et al, 2017 | |
| RBP | Pumilio2 | Down | Mff | Muscle | Altered mitochondria homeostasis | Nematode Mouse | D’Amico et al, 2019 |
| CPEB1 | Down | Ccnb1 | Oocyte | Decreased oocytes function | Mouse | Takahashi et al, 2023 | |
| RNP | Me31B/DDX-6, Imp/IGF2BP | Increased condensation | profilin | Brain | NA | Drosophila | Pushpalatha et al, 2022 |
| C1q | Increased incorporation in neuronal RNP | 28 targets | Brain | NA | Mouse | Scott-Hewitt et al, 2024 | |
NA not available.
Age-dependent changes in levels of miRNAs
miRNAs are small noncoding RNAs of 19–24 nucleotides in length that recognize complementary sequences in the 3’UTR regions of cellular mRNAs and repress their expression by inducing their degradation or translational repression (Shang et al, 2023). Each miRNA can simultaneously regulate multiple targets, thereby potentially co-targeting different components of functional networks involved in specific biological processes. A number of transcriptomics studies, performed over the years in complementary model organisms and tissues, have indicated that dozens of miRNAs are differentially expressed (either up- or downregulated) during aging (Ibanez-Ventoso et al, 2006; Inukai et al, 2012; Kato et al, 2011; Liu et al, 2012; Somel et al, 2010; Wood et al, 2015); reviewed in (Kinser and Pincus, 2020; Smith-Vikos and Slack, 2012). In these studies, algorithms predicting miRNA targets were used and combined with expression-profiling data to identify numerous transcripts potentially regulated by miRNAs during aging.
As computational-based predictions can generate false positives, it is essential to validate both the miRNA-dependent regulation of potential targets and its functional relevance. So far, this has been done for a few miRNA-target mRNA combinations in both invertebrate and vertebrate model organisms (Kinser and Pincus, 2020). mir-34, for example, was shown in Drosophila to undergo age-dependent up-regulation in the brain (Liu et al, 2012), inducing an opposite downregulation of its targets Eip74EF (a component of the steroid hormone signaling pathway) and Su(z)12 and Pcl (two components of the PRC2 polycomb repressive complex) (Kennerdell et al, 2018; Liu et al, 2012). Remarkably, mir-34 mutants exhibit accelerated brain aging characterized on the one hand by early onset, Eip74EF-dependent, vacuolarization and on the other hand by a PRC2-dependent premature sensitivity to polyglutamine aggregation and neurodegeneration (Liu et al, 2012).
As another example, mir-188-3p was shown to be upregulated in aged mouse endothelial cells and to negatively regulate the formation of bone-type H capillary vessels (He et al, 2022). While the decline of H-vessel number got alleviated in aged mir-188-3p knock-out mice, over-expressing mir-188-3p in endothelial cells of young mice decreased H vessels, a process mimicked in vitro by downregulation of the mir-188-3p mRNA target ITGβ3 (He et al, 2022).
Together, these and other functional studies have pointed to strong tissue-specificities of miRNAs undergoing age-dependent changes in expression, their target mRNAs and their physiological impacts (Cellerino and Ori, 2017; Kinser and Pincus, 2020; Smith-Vikos and Slack, 2012). Few miRNAs were, however, found to exhibit consistent and evolutionary-conserved regulation during aging. This includes mir-34, the expression of which not only increases with age in Drosophila, but also in C. elegans (de Lencastre et al, 2010), mammalian heart (Boon et al, 2013) and primate brain (Somel et al, 2010). This also includes mir-29, which undergoes age-dependent up-regulation in the nervous system of different vertebrates (Baumgart et al, 2012; Ripa et al, 2017; Somel et al, 2010; Takahashi et al, 2012), and was shown in the killifish to downregulate the expression of its target IRP2, thereby limiting intracellular iron concentration upon aging (Ripa et al, 2017).
Although clear examples of miRNAs with conserved functions in regulating the expression of target genes upon aging have been identified, the extent to which miRNAs explain age-dependent RNA-protein level uncoupling remains to be clarified. In C. elegans, 30% of the proteins, the levels of which increase with aging, were shown to be sensitive to the loss of function of Dicer, an enzyme required for miRNA processing (Walther et al, 2015). Bioinformatics analyses focusing on gene classes showing discordant RNA-protein profiles in primate brains however revealed that miRNA-binding sites are overall less predictive than RBP-binding sites to separate discordant from concordant classes (Wei et al, 2015). Furthermore, integration of RNA-seq, microRNA-seq and Ribo-seq data from aging killifish brains estimated that changes in miRNA-mediated repression explain less than 7% of the changes in protein abundance over time (Kelmer Sacramento et al, 2020). Although miRNAs each can, in theory, target up to hundreds of mRNAs, they may, in the end, be involved only in specific, yet physiologically important, regulatory processes during aging. In the future, it will be important to better understand how that specificity is achieved and how aging influences the expression of miRNAs, as our knowledge on the upstream regulatory factors is so far limited.
Age-dependent changes in levels of RNA-binding proteins
RBPs play a fundamental role in the post-transcriptional regulation of RNA fate. Through binding of selective sets of mRNAs and either masking or recruitment of cellular machineries involved in RNA degradation, translation and/or transport, they regulate the spatio-temporal expression of their target mRNAs (Glisovic et al, 2008). In their bioinformatics analysis of RNA-protein level uncoupling during primate brain aging, Wei et al uncovered a specific and significant enrichment in RBP-binding site number and density—as defined by CLiP experiments—in gene classes showing discordant RNA-protein expression profiles with aging when compared to concordant classes (Wei et al, 2015). Remarkably, different gene classes were enriched for different RBP-binding sites, but gene-class expression patterns during aging were consistent both with the expression profiles of the corresponding RBPs and with the known functions of these RBPs. For example, 85% of the transcripts of one of the defined discordant gene classes, in which protein levels but not RNA levels decrease with aging, contained one or more binding sites for the TIAL1 RBP. Moreover, TIAL1 expression was shown to negatively correlate with target protein levels, consistent with its described function in translational repression (Wei et al, 2015).
Both gene-specific and transcriptome-wide studies have further identified regulatory RBPs whose expression levels change with aging (Chaturvedi et al, 2015; D’Amico et al, 2019; Winsky-Sommerer et al, 2023). In a longitudinal study combining polysome profiling and RNA-seq of aging mouse hippocampus samples, Winsky-Sommerer et al, for example, identified a cluster enriched in RBPs and characterized by a strong past mid-age decrease in expression (Winsky-Sommerer et al, 2023). Upon integration of their multi-omics analyses performed on aging rat brain samples, Ori et al further identified functional protein-interaction networks affected by brain aging and highlighted several networks composed of RBPs involved in RNA degradation, translation or splicing (Ori et al, 2015).
Despite emerging transcriptome-wide sequencing studies predicting the importance of RBP-mediated processes in aging, very few studies have so far functionally explored the role of specific RBPs in driving age-dependent changes in gene expression. The role of Pumilio2 (Pum2), a translational repressor the levels of which were found to be consistently upregulated in various muscle and brain samples from aging mice and C. elegans (D’Amico et al, 2019), has however been dissected. In this study, cross-comparison of multi-omics experiments identified the mRNA encoding the mitochondrial fission factor MFF as a candidate Pum2 target. Mff, indeed, was found to be bound by Pum2 through a 3’UTR-located Pumilio-binding element (PBE) conserved from mammals to nematodes and to exhibit age-dependent downregulation at the protein but not RNA level. Remarkably, both RNAi-induced depletion of Pum2 ortholog in nematode and Cas9-induced silencing of Pum2 in old mouse muscles led to increased accumulation of the MFF protein and improvement in age-related alteration of mitochondrial homeostasis, providing strong in vivo evidence of the importance of Pum2-mediated translational repression during aging (D’Amico et al, 2019).
CPEB1 is another RBP whose age-related decline in level was shown in mouse to be causally involved in the altered translation activation of mRNAs essential for the developmental competence of meiosis II-arrested oocytes and fertility (Takahashi et al, 2023). Prematurely reducing the levels of CPEB1 in young oocytes decreased the translation of CPE (cytoplasmic polyadenylation element)-containing mRNAs normally activated during egg maturation, and led to infertility. Conversely, increasing the levels of CPEB1 in aged early-stage oocytes restored both the altered translation pattern of the CPEB1 mRNA target Ccnb1 and the timing of oocyte maturation (Takahashi et al, 2023). Together, these examples showcase the contribution of RBPs to normal physiological aging. More functional studies are however needed to further illustrate their role and identify their relevant targets.
Increased condensation of RNA molecules in cytoplasmic RNP granules
Extensive recent work has described and studied the recruitment of tightly regulated mRNAs and their associated proteins into large cytoplasmic assemblies termed RNA condensates or RNP (ribonucleoprotein) granules (Ripin and Parker, 2023). Different types of granules, with overlapping yet distinct RNA and protein content have been defined: some are constitutive (e.g., P-bodies, neuronal RNP granules), others are induced by stress (e.g., Stress Granules) (An et al, 2021; Bauer et al, 2023; Ripin and Parker, 2023). Condensation of RNA molecules into cytoplasmic granules has been implicated in RNA buffering, translational repression or RNA compartmentalization (Adekunle and Hubstenberger, 2020; Putnam et al, 2023). Under normal conditions, cytoplasmic RNP granules undergo highly dynamic molecular turnover. In the context of age-related diseases such as neurodegenerative diseases, static pathological inclusions enriched in RNP granule components, including ALS disease-causing RBPs such as TDP-43, FUS or hnRNPA1, have repeatedly been observed (Ling et al, 2013). These observations, combined with in vitro studies showing that mutant RBPs assemble into less dynamic assemblies, led to a model in which abnormal aggregation of granule components may occur over time, favored by disease mutations and the local concentration of interaction-prone molecules (Alberti and Hyman, 2021; Bauer et al, 2023; Kiebler and Bauer, 2024). As these studies were performed in disease contexts, it had remained unclear until recently if and how aging impacts on RNP granules in healthy contexts.
Proteomic analysis of the insoluble proteome has however revealed a propensity of RBPs, and particularly RBPs with disordered prion-like domains such as the helicase DDX-5, to aggregate in the aging killifish brains (Harel et al, 2024). Similarly, work performed in gonad-less aging C.elegans has identified proteins whose insolubility increase with age in control individuals, but not in long-lived daf-2 individuals with reduced insulin signaling, highlighting a high over-representation of RNP granule components (Lechler et al, 2017). Age-dependent decrease in RNP granule component solubility was however not systematically observed (Kelmer Sacramento et al, 2020; Molzahn et al, 2023), which may reflect differences of biochemical purifications, time windows and/or tissue-specificity. Indeed, systematic comparison of aggregation patterns across various killifish organs revealed very little overlap in the aggregating proteomes isolated from brain, gut, liver, heart, muscle, skin, and testis (Chen et al, 2024).
At the cellular level, different studies have described an increased recruitment of RBPs into cytoplasmic RNP granules with aging. In yeast, the number of cells with visible P-bodies was shown to increase in the course of replicative aging (Hu et al, 2018; Rieckher et al, 2018). In mouse neocortex tissues, an increased accumulation of Pum2 into cytoplasmic granules was observed in aged individuals (D’Amico et al, 2019). In C. elegans, the Stress Granule components PAB-1 and TIAR-2, which are diffusely localizing in pharyngial muscles of young animals, were found to accumulate in Stress Granule-like solid aggregates from mid-age (7 days) onwards, with nearly half of the end-stage individuals showing significant aggregate accumulation (Lechler et al, 2017). As revealed by a detailed analysis performed in Drosophila brain, the accumulation of Stress Granule-like punctae may represent an end-of-live phenomenon, as no such accumulation was observed in mid-aged individuals (De Graeve et al, 2022; Pushpalatha et al, 2022). Rather, conserved components of constitutive RNP granules (e.g., Me31B/DDX-6 and Imp/IGF2BP) were found to gradually accumulate into larger, yet dynamic, granules distinct from solid aggregates with aging (Pushpalatha et al, 2022). Age-related condensation was induced by increased levels of the granule nucleator Me31B, and required the activity of the PKA signaling pathway. Furthermore, RBP-bound mRNAs also exhibited increased, 3’UTR-mediated, condensation into RNP granules, which triggers their translational repression with aging.
These results suggest that the recruitment of selected mRNAs to repressive granules of increased size and number may represent a mechanism underlying transcript-specific post-transcriptional regulation in the context of aging. Consistent with a regulatory function of RNA condensation, recent work discovered that the innate immune complement protein C1q is upregulated in microglial cells of the aged mouse brain and internalized by neuronal cells to integrate into neuronal RNP granules rich in ribosomes and to regulate the translation of specific sets of proteins (Scott-Hewitt et al, 2024). This function appears to be independent of the complement pathway, but how it is mediated remains to be investigated.
Increased translation pausing at specific positions
The above-mentioned examples point to the importance of regulatory elements in the non-translated regulatory 5′ and 3’UTR regions of mRNAs in driving age-dependent post-transcriptional regulation. Features present in the coding sequences of specific mRNAs were also recently shown to alter the kinetics of translation elongation and the production of associated proteins. Stein et al, uncovered through Ribo-seq analyses in yeast and C. elegans a specific age-related increase in ribosome pausing at positions coding for Proline as well as basic Arginine and Lysine residues (Stein et al, 2022). Pausing was particularly exacerbated at polybasic stretches, where it associated with both increased ribosome collision and increased aggregation of the elongating peptide chain, a process triggered by a deficient ribosome-associated quality-control (RQC) pathway likely overwhelmed in the context of aged cells. Remarkably, a similar conclusion was reached in a multi-omics study performed in killifish brains that observed increased ribosome pausing and collision at codons encoding basic residues (Di Fraia et al, 2024). In this study, modeling also suggested that elongation pausing may be an important process driving the uncoupling of RNA and protein levels, particularly affecting a proteome subset enriched in basic residues. It will now be interesting to investigate why ribosome pausing increases specifically at these positions in aging organisms and tissues.
Concluding remarks
The various physiological changes characterizing the aging process are accompanied by alterations in both tissue-specific and systemic gene expression programs. Although most of the observed changes in gene expression correspond to small variations in absolute level, a significant fraction of the genome undergoes strong age-dependent variations in expression levels that cannot simply be explained by a decreased efficiency of the different cellular machineries involved in gene expression (Cellerino and Ori, 2017).
Control of RNA translation has emerged as an important post-transcriptional mechanism explaining both global and gene-specific expression changes with age. Molecular players contributing to translation regulation in the context of aging have been identified, notably translation initiation and elongation factors, miRNAs or RNA-binding proteins. The role of other molecular pathways, the activity of which varies with age, now remains to be investigated. m6A RNA methylation patterns were shown in different tissues and organisms to be modified with aging (Castro-Hernandez et al, 2023; Jiang et al, 2021; Perlegos et al, 2024; Wu et al, 2023a; Wu et al, 2023b) which suggests their contribution to changes in RNA stability and translation (Perlegos et al, 2024; Tassinari et al, 2023). More recent work has however challenged the view that m6A marks influence RNA translation (Zaccara and Jaffrey, 2024), thus questioning the importance of this process in the control of age-dependent changes in translation profiles. Long noncoding RNAs have for years been identified as important players of aging (Grammatikakis et al, 2014), but have only recently been suggested to regulate this process through translation regulation (Anver et al, 2024), opening new perspectives related to their contribution to age-dependent translation changes. Lastly, alternative splicing site selection has also been shown to vary with age in various tissues and organisms (Bhadra et al, 2020; Ham et al, 2022; Kumar et al, 2024; Mazin et al, 2013; Rodriguez et al, 2016; Tollervey et al, 2011; Ubaida-Mohien et al, 2019), causing isoforms with different UTR length, cis-regulatory elements and post-transcriptional dynamics (Kumar et al, 2024). How such changes functionally contribute to age-dependent translational regulation remains to be investigated.
More generally, a number of challenges remain to be addressed to better characterize the post-transcriptional mechanisms driving changes in gene expression relevant to aging physiology (Box 2). First, one needs to go beyond correlative analyses and establish functional causality links. This can be achieved using modeling approaches (Di Fraia et al, 2024; Janssens et al, 2015), but requires transcriptome-wide data sets of high temporal resolution and replicate numbers, as increased variability in RNA and protein levels is observed in aging tissues and organisms (Cellerino and Ori, 2017). This can also be achieved through functional studies which inactivate trans-acting machineries or cis-regulatory elements, and assess the direct impact on gene expression in vertebrate and/or invertebrate model organisms (Box 1).
Second, one needs to understand if the observed post-transcriptional changes are protective or if they rather participate to the decline in biological functions occurring with age. Although both scenarios are likely to be at play in a tissue-, organism- and gene-specific manner, it is interesting to consider that post-transcriptional regulation, and in particular translation regulation, may have buffering functions (Kusnadi et al, 2022). Translational compensation mechanisms have indeed been shown in different species to maintain proteome composition in contexts with initial differences in mRNA abundance (Cenik et al, 2015; Kusnadi et al, 2022; McManus et al, 2014). Lastly, one needs to better understand if and how age-dependent post-transcriptional changes contribute to disease susceptibility. Consistent with shared common mechanisms, genetic mutations in translation regulatory factors and ribosomal components were causally implicated in a number of age-related diseases (Tahmasebi et al, 2018), including cancer (Robichaud et al, 2019) and neurodegenerative diseases (Skariah and Todd, 2021).
Box 2 In need of answers.
-
i.
Are there sex-specific differences in age-related post-transcriptional regulatory processes?
-
ii.
Is the age-dependent uncoupling between protein and RNA levels progressive or are there critical periods?
-
iii.
What are the key RNA-binding proteins mediating post-transcriptional changes in gene expression upon aging? Do they control conserved “regulons” contributing to physiological aging?
-
iv.
Are the changes in gene expression regulated post-transcriptionally detrimental or protective in the context of aging? Are these changes involved in translational buffering?
-
v.
Does the age-dependent condensation of RNAs and associated proteins represent a seed for the assembly of pathological inclusions observed in late-onset degenerative diseases? More generally, how independent are the age-related changes in post-transcriptional regulatory mechanisms from the disease-causing mechanisms?
Supplementary information
Acknowledgements
The authors apologize to those whose work could not be cited owing to space constraints. This study was supported by the CNRS, as well as grants from the INCA (2021-162), the ANR (ANR22-CE12-0024), and the CEFIPRA (IFC/6503-E/2021/193). MS received fellowships from the French Ministry of Research and the ARC Foundation (ARCDOC42024010007708). The authors thank Hiba Laghrissi for the critical reading of the manuscript.
Author contributions
Mathilde Solyga: Writing—original draft; Writing—review and editing. Amitabha Majumdar: Writing—original draft; Writing—review and editing. Florence Besse: Conceptualization; Funding acquisition; Writing—original draft; Project administration; Writing—review and editing.
Disclosure and competing interests statement
The authors declare no competing interests.
Peer review information
A peer review file is available at 10.1038/s44319-024-00315-2
References
- Adekunle DA, Hubstenberger A (2020) The multiscale and multiphase organization of the transcriptome. Emerg Top Life Sci 4:265–280 [DOI] [PubMed] [Google Scholar]
- Alberti S, Hyman AA (2021) Biomolecular condensates at the nexus of cellular stress, protein aggregation disease and ageing. Nat Rev Mol Cell Biol 22:196–213 [DOI] [PubMed] [Google Scholar]
- An H, de Meritens CR, Shelkovnikova TA (2021) Connecting the “dots”: RNP granule network in health and disease. Biochim Biophys Acta Mol Cell Res 1868:119058 [DOI] [PubMed] [Google Scholar]
- Anisimova AS, Alexandrov AI, Makarova NE, Gladyshev VN, Dmitriev SE (2018) Protein synthesis and quality control in aging. Aging 10:4269–4288 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Anisimova AS, Meerson MB, Gerashchenko MV, Kulakovskiy IV, Dmitriev SE, Gladyshev VN (2020) Multifaceted deregulation of gene expression and protein synthesis with age. Proc Natl Acad Sci USA 117:15581–15590 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Anver S, Sumit AF, Sun XM, Hatimy A, Thalassinos K, Marguerat S, Alic N, Bähler J (2024) Ageing-associated long non-coding RNA extends lifespan and reduces translation in non-dividing cells. EMBO Rep 25(11):4921–4949 [DOI] [PMC free article] [PubMed]
- Bauer KE, de Queiroz BR, Kiebler MA, Besse F (2023) RNA granules in neuronal plasticity and disease. Trends Neurosci 46:525–538 [DOI] [PubMed] [Google Scholar]
- Baumgart M, Groth M, Priebe S, Appelt J, Guthke R, Platzer M, Cellerino A (2012) Age-dependent regulation of tumor-related microRNAs in the brain of the annual fish Nothobranchius furzeri. Mech Ageing Dev 133:226–233 [DOI] [PubMed] [Google Scholar]
- Bhadra M, Howell P, Dutta S, Heintz C, Mair WB (2020) Alternative splicing in aging and longevity. Hum Genet 139:357–369 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blazejowski CA, Webster GC (1983) Decreased rates of protein synthesis by cell-free preparations from different organs of aging mice. Mech Ageing Dev 21:345–356 [DOI] [PubMed] [Google Scholar]
- Boon RA, Iekushi K, Lechner S, Seeger T, Fischer A, Heydt S, Kaluza D, Treguer K, Carmona G, Bonauer A et al (2013) MicroRNA-34a regulates cardiac ageing and function. Nature 495:107–110 [DOI] [PubMed] [Google Scholar]
- Brar GA, Weissman JS (2015) Ribosome profiling reveals the what, when, where and how of protein synthesis. Nat Rev Mol Cell Biol 16:651–664 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Castro-Hernandez R, Berulava T, Metelova M, Epple R, Pena Centeno T, Richter J, Kaurani L, Pradhan R, Sakib MS, Burkhardt S et al (2023) Conserved reduction of m(6)A RNA modifications during aging and neurodegeneration is linked to changes in synaptic transcripts. Proc Natl Acad Sci USA 120:e2204933120 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cellerino A, Ori A (2017) What have we learned on aging from omics studies? Semin Cell Dev Biol 70:177–189 [DOI] [PubMed] [Google Scholar]
- Cenik C, Cenik ES, Byeon GW, Grubert F, Candille SI, Spacek D, Alsallakh B, Tilgner H, Araya CL, Tang H et al (2015) Integrative analysis of RNA, translation, and protein levels reveals distinct regulatory variation across humans. Genome Res 25:1610–1621 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chaturvedi P, Neelamraju Y, Arif W, Kalsotra A, Janga SC (2015) Uncovering RNA binding proteins associated with age and gender during liver maturation. Sci Rep 5:9512 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen Y, Sun T, Bi Z, Ni JQ, Pastor-Pareja JC, Javid B (2020) Premature termination codon readthrough in Drosophila varies in a developmental and tissue-specific manner. Sci Rep 10:8485 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen YR, Harel I, Singh PP, Ziv I, Moses E, Goshtchevsky U, Machado BE, Brunet A, Jarosz DF (2024) Tissue-specific landscape of protein aggregation and quality control in an aging vertebrate. Dev Cell 59:1892–1911 [DOI] [PMC free article] [PubMed]
- Coniglio JJ, Liu DS, Richardson A (1979) A comparison of protein synthesis by liver parenchymal cells isolated from Fischer F344 rats of various ages. Mech Ageing Dev 11:77–90 [DOI] [PubMed] [Google Scholar]
- Connors MT, Poppi DP, Cant JP (2008) Protein elongation rates in tissues of growing and adult sheep. J Anim Sci 86:2288–2295 [DOI] [PubMed] [Google Scholar]
- D’Amico D, Mottis A, Potenza F, Sorrentino V, Li H, Romani M, Lemos V, Schoonjans K, Zamboni N, Knott G et al (2019) The RNA-binding protein PUM2 impairs mitochondrial dynamics and mitophagy during aging. Mol Cell 73:775–787.e710 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Das S, Vera M, Gandin V, Singer RH, Tutucci E (2021) Intracellular mRNA transport and localized translation. Nat Rev Mol Cell Biol 22:483–504 [DOI] [PMC free article] [PubMed] [Google Scholar]
- David DC, Ollikainen N, Trinidad JC, Cary MP, Burlingame AL, Kenyon C (2010) Widespread protein aggregation as an inherent part of aging in C. elegans. PLoS Biol 8:e1000450 [DOI] [PMC free article] [PubMed] [Google Scholar]
- De Graeve F, Formicola N, Pushpalatha KV, Nakamura A, Debreuve E, Descombes X, Besse F (2022) Detecting stress granules in Drosophila neurons. Methods Mol Biol 2428:229–242 [DOI] [PubMed] [Google Scholar]
- de Lencastre A, Pincus Z, Zhou K, Kato M, Lee SS, Slack FJ (2010) MicroRNAs both promote and antagonize longevity in C. elegans. Curr Biol 20:2159–2168 [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Magalhaes JP (2024) Distinguishing between driver and passenger mechanisms of aging. Nat Genet 56:204–211 [DOI] [PubMed] [Google Scholar]
- Derisbourg MJ, Hartman MD, Denzel MS (2021a) Perspective: modulating the integrated stress response to slow aging and ameliorate age-related pathology. Nat Aging 1:760–768 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Derisbourg MJ, Wester LE, Baddi R, Denzel MS (2021b) Mutagenesis screen uncovers lifespan extension through integrated stress response inhibition without reduced mRNA translation. Nat Commun 12:1678 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Di Fraia D, Marino A, Lee JH, Kelmer Sacramento E, Baumgart M, Bagnoli S, Tomaz da Silva P, Kumar Sahu A, Siano G, Tiessen M et al (2024) Impaired biogenesis of basic proteins impacts multiple hallmarks of the aging brain. Preprint at bioRxiv 10.1101/2023.07.20.549210
- Dwyer BE, Fando JL, Wasterlain CG (1980) Rat brain protein synthesis declines during postdevelopmental aging. J Neurochem 35:746–749 [DOI] [PubMed] [Google Scholar]
- Fando JL, Salinas M, Wasterlain CG (1980) Age-dependent changes in brain protein synthesis in the rat. Neurochem Res 5:373–383 [DOI] [PubMed] [Google Scholar]
- Gerashchenko MV, Peterfi Z, Yim SH, Gladyshev VN (2021) Translation elongation rate varies among organs and decreases with age. Nucleic Acids Res 49:e9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gerdes Gyuricza I, Chick JM, Keele GR, Deighan AG, Munger SC, Korstanje R, Gygi SP, Churchill GA (2022) Genome-wide transcript and protein analysis highlights the role of protein homeostasis in the aging mouse heart. Genome Res 32:838–852 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Glisovic T, Bachorik JL, Yong J, Dreyfuss G (2008) RNA-binding proteins and post-transcriptional gene regulation. FEBS Lett 582:1977–1986 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grammatikakis I, Panda AC, Abdelmohsen K, Gorospe M (2014) Long noncoding RNAs(lncRNAs) and the molecular hallmarks of aging. Aging 6:992–1009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ham S, Kim SS, Park S, Kim EJE, Kwon S, Park HH, Jung Y, Lee SV (2022) Systematic transcriptome analysis associated with physiological and chronological aging in Caenorhabditis elegans. Genome Res 32:2003–2014 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hanson G, Coller J (2018) Codon optimality, bias and usage in translation and mRNA decay. Nat Rev Mol Cell Biol 19:20–30 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harel I, Chen YR, Ziv I, Singh PP, Heinzer D, Navarro Negredo P, Goshtchevsky U, Wang W, Astre G, Moses E et al (2024) Identification of protein aggregates in the aging vertebrate brain with prion-like and phase-separation properties. Cell Rep 43:112787 [DOI] [PMC free article] [PubMed] [Google Scholar]
- He WZ, Yang M, Jiang Y, He C, Sun YC, Liu L, Huang M, Jiao YR, Chen KX, Hou J et al (2022) miR-188-3p targets skeletal endothelium coupling of angiogenesis and osteogenesis during ageing. Cell Death Dis 13:494 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hershey JW, Sonenberg N, Mathews MB (2012) Principles of translational control: an overview. Cold Spring Harb Perspect Biol 4:a011528 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hipp MS, Kasturi P, Hartl FU (2019) The proteostasis network and its decline in ageing. Nat Rev Mol Cell Biol 20:421–435 [DOI] [PubMed] [Google Scholar]
- Houseley J, Tollervey D (2009) The many pathways of RNA degradation. Cell 136:763–776 [DOI] [PubMed] [Google Scholar]
- Hu Z, Xia B, Postnikoff SD, Shen ZJ, Tomoiaga AS, Harkness TA, Seol JH, Li W, Chen K, Tyler JK (2018) Ssd1 and Gcn2 suppress global translation efficiency in replicatively aged yeast while their activation extends lifespan. eLife 7:e35551 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ibanez-Ventoso C, Yang M, Guo S, Robins H, Padgett RW, Driscoll M (2006) Modulated microRNA expression during adult lifespan in Caenorhabditis elegans. Aging Cell 5:235–246 [DOI] [PubMed] [Google Scholar]
- Inukai S, de Lencastre A, Turner M, Slack F (2012) Novel microRNAs differentially expressed during aging in the mouse brain. PLoS ONE 7:e40028 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iwasaki S, Ingolia NT (2017) The growing toolbox for protein synthesis studies. Trends Biochem Sci 42:612–624 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Janssens GE, Meinema AC, Gonzalez J, Wolters JC, Schmidt A, Guryev V, Bischoff R, Wit EC, Veenhoff LM, Heinemann M (2015) Protein biogenesis machinery is a driver of replicative aging in yeast. eLife 4:e08527 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jiang ZX, Wang YN, Li ZY, Dai ZH, He Y, Chu K, Gu JY, Ji YX, Sun NX, Yang F et al (2021) The m6A mRNA demethylase FTO in granulosa cells retards FOS-dependent ovarian aging. Cell Death Dis 12:744 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kato M, Chen X, Inukai S, Zhao H, Slack FJ (2011) Age-associated changes in expression of small, noncoding RNAs, including microRNAs, in C. elegans. RNA 17:1804–1820 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaushik S, Cuervo AM (2015) Proteostasis and aging. Nat Med 21:1406–1415 [DOI] [PubMed] [Google Scholar]
- Keele GR, Zhang JG, Szpyt J, Korstanje R, Gygi SP, Churchill GA, Schweppe DK (2023) Global and tissue-specific aging effects on murine proteomes. Cell Rep 42:112715 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kelmer Sacramento E, Kirkpatrick JM, Mazzetto M, Baumgart M, Bartolome A, Di Sanzo S, Caterino C, Sanguanini M, Papaevgeniou N, Lefaki M et al (2020) Reduced proteasome activity in the aging brain results in ribosome stoichiometry loss and aggregation. Mol Syst Biol 16:e9596 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kennerdell JR, Liu N, Bonini NM (2018) MiR-34 inhibits polycomb repressive complex 2 to modulate chaperone expression and promote healthy brain aging. Nat Commun 9:4188 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khatir I, Brunet MA, Meller A, Amiot F, Patel T, Lapointe X, Avila Lopez J, Guilloy N, Castonguay A, Husain MA et al (2023) Decoupling of mRNA and protein expression in aging brains reveals the age-dependent adaptation of specific gene subsets. Cells 12:615 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kiebler MA, Bauer KE (2024) RNA granules in flux: dynamics to balance physiology and pathology. Nat Rev Neurosci 25:711–725 [DOI] [PubMed] [Google Scholar]
- Kim HS, Pickering AM (2023) Protein translation paradox: Implications in translational regulation of aging. Front Cell Dev Biol 11:1129281 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kimball SR, Vary TC, Jefferson LS (1992) Age-dependent decrease in the amount of eukaryotic initiation factor 2 in various rat tissues. Biochem J 286:263–268 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kinser HE, Pincus Z (2020) MicroRNAs as modulators of longevity and the aging process. Hum Genet 139:291–308 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kong J, Lasko P (2012) Translational control in cellular and developmental processes. Nat Rev Genet 13:383–394 [DOI] [PubMed] [Google Scholar]
- Kumar NH, Kluever V, Barth E, Krautwurst S, Furlan M, Pelizzola M, Marz M, Fornasiero EF (2024) Comprehensive transcriptome analysis reveals altered mRNA splicing and post-transcriptional changes in the aged mouse brain. Nucleic Acids Res 52:2865–2885 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kusnadi EP, Timpone C, Topisirovic I, Larsson O, Furic L (2022) Regulation of gene expression via translational buffering. Biochim Biophys Acta Mol Cell Res 1869:119140 [DOI] [PubMed] [Google Scholar]
- Ladiges W, Morton J, Blakely C, Gale M (2000) Tissue specific expression of PKR protein kinase in aging B6D2F1 mice. Mech Ageing Dev 114:123–132 [DOI] [PubMed] [Google Scholar]
- Layman DK, Ricca GA, Richardson A (1976) The effect of age on protein synthesis and ribosome aggregation to messenger RNA in rat liver. Arch Biochem Biophys 173:246–254 [DOI] [PubMed] [Google Scholar]
- Lechler MC, Crawford ED, Groh N, Widmaier K, Jung R, Kirstein J, Trinidad JC, Burlingame AL, David DC (2017) Reduced insulin/IGF-1 signaling restores the dynamic properties of key stress granule proteins during aging. Cell Rep 18:454–467 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ling SC, Polymenidou M, Cleveland DW (2013) Converging mechanisms in ALS and FTD: disrupted RNA and protein homeostasis. Neuron 79:416–438 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu N, Landreh M, Cao K, Abe M, Hendriks GJ, Kennerdell JR, Zhu Y, Wang LS, Bonini NM (2012) The microRNA miR-34 modulates ageing and neurodegeneration in Drosophila. Nature 482:519–523 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lopez-Otin C, Blasco MA, Partridge L, Serrano M, Kroemer G (2023) Hallmarks of aging: An expanding universe. Cell 186:243–278 [DOI] [PubMed] [Google Scholar]
- Martinez-Miguel VE, Lujan C, Espie-Caullet T, Martinez-Martinez D, Moore S, Backes C, Gonzalez S, Galimov ER, Brown AEX, Halic M et al (2021) Increased fidelity of protein synthesis extends lifespan. Cell Metab 33:2288–2300 e2212 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mayr C (2019) What are 3’ UTRs doing? Cold Spring Harb Perspect Biol 11:a034728 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mazin P, Xiong J, Liu X, Yan Z, Zhang X, Li M, He L, Somel M, Yuan Y, Phoebe Chen YP et al (2013) Widespread splicing changes in human brain development and aging. Mol Syst Biol 9:633 [DOI] [PMC free article] [PubMed] [Google Scholar]
- McManus CJ, May GE, Spealman P, Shteyman A (2014) Ribosome profiling reveals post-transcriptional buffering of divergent gene expression in yeast. Genome Res 24:422–430 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Medioni C, Besse F (2018) The secret life of RNA: lessons from emerging methodologies. Methods Mol Biol 1649:1–28 [DOI] [PubMed] [Google Scholar]
- Moldave K, Harris J, Sabo W, Sadnik I (1979) Protein synthesis and aging: studies with cell-free mammalian systems. Fed Proc 38:1979–1983 [PubMed] [Google Scholar]
- Molzahn C, Kuechler ER, Zemlyankina I, Nierves L, Ali T, Cole G, Wang J, Albu RF, Zhu M, Cashman NR et al (2023) Shift of the insoluble content of the proteome in the aging mouse brain. Proc Natl Acad Sci USA 120:e2310057120 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Motizuki M, Tsurugi K (1992) The effect of aging on protein synthesis in the yeast Saccharomyces cerevisiae. Mech Ageing Dev 64:235–245 [DOI] [PubMed] [Google Scholar]
- Nakazawa T, Mori N, Goto S (1984) Functional deterioration of mouse liver ribosomes during aging: translational activity and activity for formation of the 47 S initiation complex. Mech Ageing Dev 26:241–251 [DOI] [PubMed] [Google Scholar]
- Ori A, Toyama BH, Harris MS, Bock T, Iskar M, Bork P, Ingolia NT, Hetzer MW, Beck M (2015) Integrated transcriptome and proteome analyses reveal organ-specific proteome deterioration in old rats. Cell Syst 1:224–237 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perlegos AE, Byrns CN, Bonini NM (2024) Cell type-specific regulation of m(6) A modified RNAs in the aging Drosophila brain. Aging Cell 23:e14076 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pushpalatha KV, Solyga M, Nakamura A, Besse F (2022) RNP components condense into repressive RNP granules in the aging brain. Nat Commun 13:2782 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Putnam A, Thomas L, Seydoux G (2023) RNA granules: functional compartments or incidental condensates? Genes Dev 37:354–376 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reis-Rodrigues P, Czerwieniec G, Peters TW, Evani US, Alavez S, Gaman EA, Vantipalli M, Mooney SD, Gibson BW, Lithgow GJ et al (2012) Proteomic analysis of age-dependent changes in protein solubility identifies genes that modulate lifespan. Aging Cell 11:120–127 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rieckher M, Markaki M, Princz A, Schumacher B, Tavernarakis N (2018) Maintenance of proteostasis by P body-mediated regulation of eIF4E availability during aging in Caenorhabditis elegans. Cell Rep 25:199–211 e196 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Riis B, Rattan SI, Palmquist K, Nilsson A, Nygard O, Clark BF (1993) Elongation factor 2-specific calcium and calmodulin dependent protein kinase III activity in rat livers varies with age and calorie restriction. Biochem Biophys Res Commun 192:1210–1216 [DOI] [PubMed] [Google Scholar]
- Ripa R, Dolfi L, Terrigno M, Pandolfini L, Savino A, Arcucci V, Groth M, Terzibasi Tozzini E, Baumgart M, Cellerino A (2017) MicroRNA miR-29 controls a compensatory response to limit neuronal iron accumulation during adult life and aging. BMC Biol 15:9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ripin N, Parker R (2023) Formation, function, and pathology of RNP granules. Cell 186:4737–4756 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robichaud N, Sonenberg N, Ruggero D, Schneider RJ (2019) Translational control in cancer. Cold Spring Harb Perspect Biol 11:a032896 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rodriguez SA, Grochova D, McKenna T, Borate B, Trivedi NS, Erdos MR, Eriksson M (2016) Global genome splicing analysis reveals an increased number of alternatively spliced genes with aging. Aging Cell 15:267–278 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roundtree IA, Evans ME, Pan T, He C (2017) Dynamic RNA modifications in gene expression regulation. Cell 169:1187–1200 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rowlands AG, Panniers R, Henshaw EC (1988) The catalytic mechanism of guanine nucleotide exchange factor action and competitive inhibition by phosphorylated eukaryotic initiation factor 2. J Biol Chem 263:5526–5533 [PubMed] [Google Scholar]
- Scott-Hewitt N, Mahoney M, Huang Y, Korte N, Yvanka de Soysa T, Wilton DK, Knorr E, Mastro K, Chang A, Zhang A et al (2024) Microglial-derived C1q integrates into neuronal ribonucleoprotein complexes and impacts protein homeostasis in the aging brain. Cell 187:4193–4212.e24 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sen P, Shah PP, Nativio R, Berger SL (2016) Epigenetic mechanisms of longevity and aging. Cell 166:822–839 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shang R, Lee S, Senavirathne G, Lai EC (2023) microRNAs in action: biogenesis, function and regulation. Nat Rev Genet 24:816–833 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Skariah G, Todd PK (2021) Translational control in aging and neurodegeneration. Wiley Interdiscip Rev RNA 12:e1628 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith-Vikos T, Slack FJ (2012) MicroRNAs and their roles in aging. J Cell Sci 125:7–17 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Somel M, Guo S, Fu N, Yan Z, Hu HY, Xu Y, Yuan Y, Ning Z, Hu Y, Menzel C et al (2010) MicroRNA, mRNA, and protein expression link development and aging in human and macaque brain. Genome Res 20:1207–1218 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sonenberg N, Hinnebusch AG (2009) Regulation of translation initiation in eukaryotes: mechanisms and biological targets. Cell 136:731–745 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stein KC, Morales-Polanco F, van der Lienden J, Rainbolt TK, Frydman J (2022) Ageing exacerbates ribosome pausing to disrupt cotranslational proteostasis. Nature 601:637–642 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sudmant PH, Lee H, Dominguez D, Heiman M, Burge CB (2018) Widespread accumulation of ribosome-associated isolated 3’ UTRs in neuronal cell populations of the aging brain. Cell Rep. 25:2447–2456 e2444 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tahmasebi S, Khoutorsky A, Mathews MB, Sonenberg N (2018) Translation deregulation in human disease. Nat Rev Mol Cell Biol 19:791–807 [DOI] [PubMed] [Google Scholar]
- Takahashi M, Eda A, Fukushima T, Hohjoh H (2012) Reduction of type IV collagen by upregulated miR-29 in normal elderly mouse and klotho-deficient, senescence-model mouse. PLoS ONE 7:e48974 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Takahashi N, Franciosi F, Daldello EM, Luong XG, Althoff P, Wang X, Conti M (2023) CPEB1-dependent disruption of the mRNA translation program in oocytes during maternal aging. Nat Commun 14:416 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Takemon Y, Chick JM, Gerdes Gyuricza I, Skelly DA, Devuyst O, Gygi SP, Churchill GA, Korstanje R (2021) Proteomic and transcriptomic profiling reveal different aspects of aging in the kidney. eLife 10:e62585 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tassinari V, La Rosa P, Guida E, Colopi A, Caratelli S, De Paolis F, Gallo A, Cenciarelli C, Sconocchia G, Dolci S et al (2023) Contribution of A-to-I RNA editing, M6A RNA Methylation, and Alternative Splicing to physiological brain aging and neurodegenerative diseases. Mech Ageing Dev 212:111807 [DOI] [PubMed] [Google Scholar]
- Thoreen CC, Chantranupong L, Keys HR, Wang T, Gray NS, Sabatini DM (2012) A unifying model for mTORC1-mediated regulation of mRNA translation. Nature 485:109–113 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tollervey JR, Wang Z, Hortobagyi T, Witten JT, Zarnack K, Kayikci M, Clark TA, Schweitzer AC, Rot G, Curk T et al (2011) Analysis of alternative splicing associated with aging and neurodegeneration in the human brain. Genome Res 21:1572–1582 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ubaida-Mohien C, Lyashkov A, Gonzalez-Freire M, Tharakan R, Shardell M, Moaddel R, Semba RD, Chia CW, Gorospe M, Sen R et al (2019) Discovery proteomics in aging human skeletal muscle finds change in spliceosome, immunity, proteostasis and mitochondria. eLife 8:e49874 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ule J, Blencowe BJ (2019) Alternative splicing regulatory networks: functions, mechanisms, and evolution. Mol Cell 76:329–345 [DOI] [PubMed] [Google Scholar]
- Waldera-Lupa DM, Kalfalah F, Florea AM, Sass S, Kruse F, Rieder V, Tigges J, Fritsche E, Krutmann J, Busch H et al (2014) Proteome-wide analysis reveals an age-associated cellular phenotype of in situ aged human fibroblasts. Aging 6:856–878 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walther DM, Kasturi P, Zheng M, Pinkert S, Vecchi G, Ciryam P, Morimoto RI, Dobson CM, Vendruscolo M, Mann M et al (2015) Widespread Proteome remodeling and aggregation in aging C. elegans. Cell 161:919–932 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ward W, Richardson A (1991) Effect of age on liver protein synthesis and degradation. Hepatology 14:935–948 [DOI] [PubMed] [Google Scholar]
- Webster GC, Webster SL (1979) Decreased protein synthesis by microsomes from aging Drosophila melanogaster. Exp Gerontol 14:343–348 [DOI] [PubMed] [Google Scholar]
- Webster GC, Webster SL (1982) Effects of age on the post-initiation stages of protein synthesis. Mech Ageing Dev 18:369–378 [DOI] [PubMed] [Google Scholar]
- Webster GC, Webster SL (1983) Decline in synthesis of elongation factor one (EF-1) precedes the decreased synthesis of total protein in aging Drosophila melanogaster. Mech Ageing Dev 22:121–128 [DOI] [PubMed] [Google Scholar]
- Wei YN, Hu HY, Xie GC, Fu N, Ning ZB, Zeng R, Khaitovich P (2015) Transcript and protein expression decoupling reveals RNA binding proteins and miRNAs as potential modulators of human aging. Genome Biol 16:41 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Winsky-Sommerer R, King HA, Iadevaia V, Moller-Levet C, Gerber AP (2023) A post-transcriptional regulatory landscape of aging in the female mouse hippocampus. Front Aging Neurosci 15:1119873 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wood SH, van Dam S, Craig T, Tacutu R, O’Toole A, Merry BJ, de Magalhaes JP (2015) Transcriptome analysis in calorie-restricted rats implicates epigenetic and post-translational mechanisms in neuroprotection and aging. Genome Biol 16:285 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu Z, Lu M, Liu D, Shi Y, Ren J, Wang S, Jing Y, Zhang S, Zhao Q, Li H et al (2023a) m(6)A epitranscriptomic regulation of tissue homeostasis during primate aging. Nat Aging 3:705–721 [DOI] [PubMed] [Google Scholar]
- Wu Z, Ren J, Liu GH (2023b) Deciphering RNA m(6) A regulation in aging: perspectives on current advances and future directions. Aging Cell 22:e13972 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang J, Chen D, He Y, Melendez A, Feng Z, Hong Q, Bai X, Li Q, Cai G, Wang J et al (2013) MiR-34 modulates Caenorhabditis elegans lifespan via repressing the autophagy gene atg9. Age 35:11–22 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yuan S, Zhou G, Xu G (2024) Translation machinery: the basis of translational control. J Genet Genomics 51:367–378 [DOI] [PubMed] [Google Scholar]
- Zaccara S, Jaffrey SR (2024) Understanding the redundant functions of the m(6)A-binding YTHDF proteins. RNA 30:468–481 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.