Abstract
The accurate characterization of triplet repeats, especially the overrepresented CAG repeats, is increasingly relevant for several reasons. First, germline expansion of CAG repeats above a gene-specific threshold causes multiple neurodegenerative disorders; for instance, Huntington’s disease (HD) is triggered by >36 CAG repeats in the huntingtin (HTT) gene. Second, extreme expansions up to 800 CAG repeats have been found in specific cell types affected by the disease. Third, synonymous single nucleotide variants within the CAG repeat stretch influence the age of disease onset. Thus, new sequencing-based protocols that profile both the length and the exact nucleotide sequence of triplet repeats are crucial. Various strategies to enrich the target gene over the background, along with sequencing platforms and bioinformatic pipelines, are under development. This review discusses the concepts, challenges, and methodological opportunities for analyzing triplet repeats, using HD as a case study. Starting with traditional approaches, we will explore how sequencing-based methods have evolved to meet increasing scientific demands. We will also highlight experimental and bioinformatic challenges, aiming to provide a guide for accurate triplet repeat characterization for diagnostic and therapeutic purposes.
Graphical Abstract
Graphical Abstract.
Introduction
Clinical relevance of C-A-G triplet repeats in the nervous system
Triplet repeats are genomic sequences composed of tandem repetitions of three nucleotide motifs, which are abundant in the genomes of higher organisms (1). Many triplet repeats are overrepresented in the human genome, with the C-A-G (cytosine-adenine-guanine, CAG) being the most represented one in exons (2). Their main supposed purpose is to allow for evolutionary plasticity, given their highly unstable nature (3,4).
Importantly, expansions of certain triplet repeats above a gene-specific threshold cause multiple human neurological and neuromuscular diseases, such as Huntington’s disease (HD), spinocerebellar ataxias (SCAs), spinal and bulbar muscular atrophy (SBMA), myotonic dystrophy (DM), fragile X syndrome type A, Friedreich’s ataxia and many others (5,6). The fact that the majority of diseases caused by triplet repeat expansions involve neurological dysfunctions is suggestive of a specific need to tightly regulate trinucleotide repeats in the nervous system (7).
In 1993, the Huntington Disease Study Group identified Huntingtin (HTT) as the disease-causing gene (8). The CAG repeat, encoding for glutamine (Q), is located in the first of 67 exons of the HTT transcript. In patients, the CAG is repeated above the threshold of 36 copies, driving the clinical symptoms and neurodegeneration typical of the disease (9). Longer CAG repeats are associated with earlier onset (8), highlighting the inverse correlation between CAG repeat length and age of onset (AOO). The most frequent pathogenic repeat size reported is around 40–42 CAG repeats, with a mean AOO typically at 45 years of age (10), with a variability of 10–20 years (11). Moreover, the CAG repeat is followed by a CCG repeat, encoding for proline (P), which may stabilize the glutamine-rich tract, by keeping it soluble (12). For many years, efforts focused on the elucidation of the toxic effects of the mutant Huntingtin protein (mHTT). These research endeavours shed light on its deleterious effect in a variety of molecular and cellular processes, leading to dysfunction and selective degeneration of the neurons in the brain cerebral cortex and striatum regions (13). The inherited CAG repeat can further expand from generation to generation, particularly through paternal transmission (14,15). This evidence underscores the unstable nature of the CAG repeat in the gene, a phenomenon that has come under scrutiny, especially regarding female and male germline transmission (8,14). This instability is likely linked to the high number of mitotic events occurring over time in the male germline, compared to the female germline (14,15). Consequently, an increase in CAG size during parental transmission results in earlier AOO and leads to a clinical feature known as anticipation, where the longer CAG repeat inherited by the progeny causes the disease phenotype to manifest much earlier in the offspring than in the gene-positive father (16).
In the early era of measuring inherited pathological CAG sizes in the HTT gene for HD diagnosis, methods based on polymerase chain reaction (PCR) amplification and electrophoresis were first developed. These techniques facilitated the straightforward measurement of the CAG repeat length in HTT, but did not provide information on its nucleotide composition. For a long time, these approaches represented the gold-standard in diagnostics and underwent only minor optimizations.
Characterizing CAG repeat dynamics: meeting the challenges of scientific progress
In the last few years, the field has been transformed by evidence that the pathological CAG triplets carried in genes such as HTT vary in size among different cells within affected tissues (17–20). A summary of these results and their implications are provided in the accompanying review (157). This variability also suggests that the diseased brain is a mosaic of genomes (21), each carrying a genome with varying CAG sizes in the responsible gene. This mosaicism results from the instability of the CAG tract, which occurs not only in germline transmission but also in somatic tissues, accumulating over the patients’ lifetimes. Notably, in 2003, Kennedy and colleagues (22) first reported extreme somatic expansion events in the striatum of HD patients, though these findings were initially overlooked due to technological limitations. Since then, data on somatic instability (SI) have been documented for multiple disease genes, including those associated with HD (14,20,23), SCA1 (24–26) and DM (27–30).
Interest in SI was renewed when Genome Wide Association Studies (GWAS) identified polymorphisms in genes involved in the mismatch repair (MMR) pathway as modifiers of AOO in HD (11,31,33). Specifically, MMR protein complexes recognize large extra-helical extrusions (i.e. mismatches between the two strands) at the HTT CAG repeat, and repair them, potentially altering the CAG length (32). Recent GWAS using larger cohorts, confirmed previously identified genes as trans modifiers of AOO, such as FAN1, MSH3, PMS1, PMS2, MLH1 and LIG1, and also identified new modifiers, including POLD1 and MED15 (34).
More recently, two studies have shown that not only do different tissues exhibit variable CAG instability, but individual cell types within those tissues do as well. Importantly, within the striatum, instability is cell-type specific, with medium spiny neurons (MSNs) being the most affected neuronal type in HD and exhibiting the highest instability levels in both HD and SCA3 (17,18). Mätlik and colleagues studied postmortem HD brain tissue and used fluorescence-activated nuclear sorting to isolate brain cells based on marker genes expression. They performed CAG sizing via bulk PCR and Illumina MiSeq sequencing, supporting a model where cell-type-specific SI is necessary but not sufficient for cell death. Notably, similar levels of somatic mosaicism were observed in both MSNs and cholinergic neurons, despite different vulnerability (17). Handsaker and colleagues developed a single-cell protocol using long-read sequencing to characterize CAG repeats in the HTT gene and the transcriptional profiles of the same individual cells from postmortem HD brains (18). Using a PCR protocol with unique molecular identifiers (UMIs) and Pacific Biosciences (PacBio) high fidelity (HiFi) sequencing – discussed in subsequent sections – they detected extreme somatic expansions exceeding 800 CAG repeats in a subset of human MSNs, linking these expansions to transcriptional dysregulation and neuron death (18).
GWAS have also identified elements within the HTT locus acting as (cis) modifiers of CAG instability. The HTT gene contains a CAA–CAG sequence at the end of the CAG stretch, where CAA – that also encodes for glutamine – interrupts the otherwise ‘pure’ stretch of CAG repeats. Variants like loss of the CAA interruption have been described, in which patients carry a CAG codon instead of CAA at this position, or with a CAACAG duplication (DUP) experience significant changes in AOO – with loss of interruption (LOI) leading to a 25-year earlier onset and DUP delaying onset by 4.2-year (33). These variations are linked to CAG instability in blood cells, though it remains unclear if they affect brain cells similarly (11,33,34).
This evidence supports a model of HD pathogenesis where an initially harmless CAG stretch expands progressively, eventually crossing a toxic threshold that triggers transcriptional abnormalities and cell dysfunction (35). Therapeutic strategies are shifting toward reducing SI, alongside clinical trials focussed on lowering mHTT levels through RNA interference or allele-specific mHTT reduction (23,32,33,36).
In summary, the growing importance of accurately characterizing CAG repeats for diagnostic, prognostic and therapeutic purposes is evident (18), but the repetitive nature of these sequences presents technical challenges (37). The following sections will review traditional triplet repeat characterization methods (38,39), introduce high-throughput sequencing-based techniques and discuss bioinformatic approaches for analyzing sequencing data. Finally, we will guide the selection of methods tailored to specific applications, with a focus on HD.
Traditional methods for triplet repeats characterization
Since the 1990s, several methods have been developed to size triplet repeats (26,40–42). One class includes amplification-free methods like southern blotting, where genomic DNA is digested, separated by gel electrophoresis and probed with a labeled DNA fragment that hybridizes to the repeat-containing region (43) (Figure 1A). Southern blotting avoids PCR amplification steps – therefore bypassing amplification-related artifacts – but requires large amounts of DNA. It is commonly used for detecting long triplet repeat expansions that may not amplify well by PCR, especially when PCR produces a homozygous band for the wild-type allele (39,44–47). Southern blotting was pivotal in 2003 for identifying large expansions in HD patients’ striatal cells early in the disease progression (22).
Figure 1.
Traditional methods for triplet repeats characterization. (A) Southern blotting requires genomic DNA digestion with restriction enzymes, followed by blotting and probing with a labeled DNA fragment that specifically hybridizes to the repeat containing region. (B) Fluorescence-PCR uses at least one fluorescent primer and performs fragment analysis using a capillary electrophoresis system. (C) Small-pool PCR relies on serial dilutions and multiple independent PCRs across the repeat, followed by electrophoresis and blotting. (D) Sanger sequencing of PCR amplicons, after allelic separation by electrophoresis, detects fluorescence emitted by chain-terminating nucleotides.
Another set of methods employs capillary electrophoresis to analyze bulk PCR products (Figure 1B). These methods employ either one or two fluorescent primers (Fluorescence PCR) (48,49). Another method based on three primers has been developed, where one primer lies outside the repeat, the second is within the repeat with a sequence-tail complementary to a third universal and fluorescently labeled primer (Repeat-primed PCR) (40,50–53). Fluorescence PCR accurately sizes repeats but struggles to detect low-frequency alleles from SI, due to preferential PCR amplification of shorter alleles and unclear boundary between signal and noise in electrophoresis (43,49). In contrast, repeat-primed PCR produces ladder-shaped amplification products, minimizing misses of mutant alleles, and offering a cost-effective alternative to southern blotting for determining true homozygosity (43,51,52). However, it is less effective for accurate sizing of repeats longer than 100 CAGs (52).
For HD, PCR amplification of the target locus followed by gel or capillary electrophoresis has been the traditional method for measuring CAG repeats, with single-repeat resolution (41,42,54). To detect low-frequency expanded alleles resulting from somatic mosaicism, methods like small-pool or single-molecule PCR, which use serial dilutions of genomic DNA and multiple independent PCR reactions, have been developed (small-pool or single-molecule PCR) (28,55) (Figure 1C). Serial dilution sampling enables the detection of very rare repeat variants within a sample, which has been challenging for traditional bulk PCR approaches. Notably, these methods use less than a few hundred molecules of input DNA per reaction, allowing for the detection of rare genomic variants. Though effective, these methods are labour-intensive and susceptible to DNA contamination, requiring separation of all PCR products by gel or electrophoresis, with detection by southern blotting.
While these methods offer reliable estimates of triplet repeats lengths and insights into SI, they do not provide nucleotide sequence information. As a result, they cannot detect clinically relevant cis modifiers within the disease-causing haplotype (11,33,38,56).
Sanger sequencing, developed in the 1970s, has been used to characterize the HTT gene’s repetitive region (8) and remains widely employed in clinical practice, mainly as a confirmatory orthogonal assay (39,57–61) (Figure 1D). This method uses PCR amplicons followed by a chain termination reaction, with fragments separated by capillary electrophoresis and sequenced by dye fluorescence (57,58,62). While accurate up to 1000 bp (63), Sanger sequencing reads from a bulk population, which results in the loss of sequence phase coherence, limiting its utility for studying somatic mosaicism – a common feature in triplet repeats disorders (39,56,64,65). Therefore, high-throughput sequencing-based methods are being developed to provide accurate sizing, single-nucleotide composition and somatic mosaicism assessment for triplet repeats.
Advancing high-throughput sequencing methods for triplet repeats characterization
The discovery of nucleotide variants within the CAG tract of the HTT gene, which correlate with varying AOO of HD, and the evidence of significant somatic expansion of CAG repeats up to hundreds of units, have driven a shift in sequencing strategies. This shift focuses on accurately sequencing long sequences and determining their nucleotide composition at this specific locus. Advanced sequencing techniques that offer precise readings of these repeats and their variants have emerged (11,33,38,60,66,67) (Figure 2). These advancements are driven by high-throughput sequencing platforms, categorized into PCR-based and PCR-free enrichment methods and short-reads versus long-reads sequencing platforms. While optical genome mapping systems, like those offered by Bionano Genomics, can detect short tandem repeats, they do not provide detailed sequence information for the repeats, and their current resolution is limited to ∼200–500 bp (68–70). However, these methods may become a high-throughput, genome-wide alternative to southern blotting for identifying the most likely disease-related genes in the future.
Figure 2.
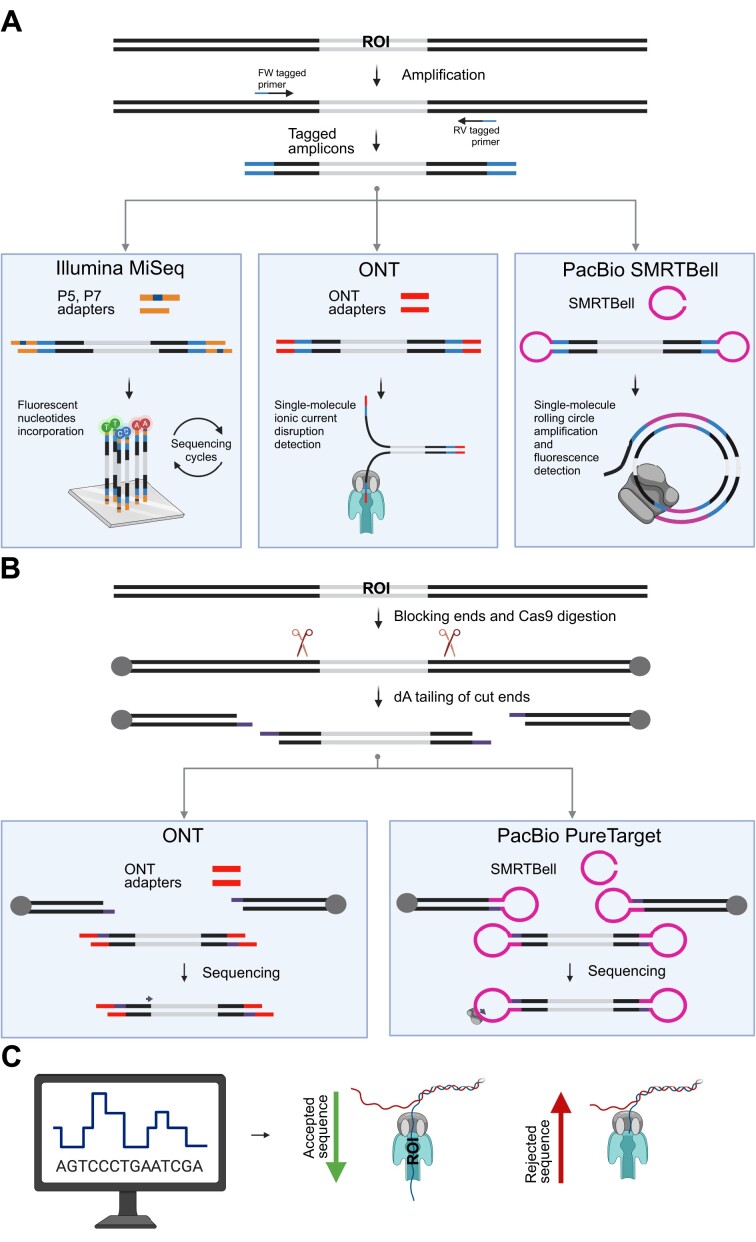
High-throughput sequencing methods for triplet repeat characterization. (A) PCR-based methods begin with PCR amplification of the region of interest; the resulting amplicons then undergo platform-specific library preparation for high-throughput sequencing. (B) CRISPR/Cas9-based enrichment methods involve cutting DNA using the Cas9-CRISPR RNAs (crRNAs) complex, followed by ligation of sequencing adapters to the free DNA ends. (C) In-silico-based enrichment methods (adaptive sampling or ‘Read Until’) are used with Oxford Nanopore Technologies (ONT) devices to selectively sequence DNA molecules. Based on the first sequenced bases, the voltage across the nanopore can be reversed to eject the molecule if it does not match an on-target region.
PCR-based approaches for triplet repeats characterization using short-read sequencing
A primary class of sequencing approaches involves PCR amplification of the target region, followed by sequencing on short-read platforms, primarily led by Illumina. Emerging competitors such as MGI, Element Biosciences and Ultima Genomics are expected to gain prominence in the near future (71). Illumina’s sequencing-by-synthesis method utilizes ‘bridge amplification’, where DNA molecules with appropriate adapters serve as templates for repeated cycles of synthesis on a solid support (Figure 2A). This process generates millions of clonal clusters, with each cluster containing about a thousand copies of the same oligonucleotide fragment. During each synthesis cycle, a terminating, fluorescently labeled nucleotide is incorporated, and the corresponding fluorescence signal is detected (72).
Sequencing accuracy is typically expressed as the probability of correct base-calling, and it is often represented by Phred Quality scores (Q) and defined as Q = −10*log10 (P) (73). As such, a Q score of 10 (Q10) reflects a 90% base-calling accuracy, Q20 equates to 99%, Q30 to 99.9% and so on. Illumina short-reads typically achieve accuracy levels exceeding Q30 (>99.9%). However, they are known to experience phasing issues, which progressively degrade sequencing quality, dependent on the sequencing cycle (74). This is due to the incomplete removal of blocker nucleotides, causing some molecules from each cluster to lag one cycle behind, reducing the signal-to-noise ratio (74). In this context, Illumina MiSeq is a frequently used sequencer for these applications, supporting sequencing runs of up to 600 cycles and generating up to 15 Gb of data in 56 h (60,66,67) (Table 1). In particular, MiSeq can sequence DNA fragments up to 600–1000 bp in paired-end mode, where each read corresponds to one end of the fragment (75,76). Asymmetric cycle distribution between paired reads allows flexibility, with one read being longer (e.g. 400 bp) and the other shorter (17,38). This approach supports sequencing of repeats up to ∼350 bp long, assuming PCR primers are positioned near the repeat flanking region (17,38). Despite its strengths, MiSeq faces challenges with phasing issues, especially towards the end of each read (38,74).
Table 1.
High-throughput sequencing platforms for triplet repeats characterization
| Sequencing platform | Num. flow-cells | Maximum throughput | Modal quality Score | Mean read length | Run time | Platform cost ($) | Cost ($)/Gba |
|---|---|---|---|---|---|---|---|
Illumina MiSeq
|
1 | Up to 15 Gb | Q40 = 99.99% | Up to 400 bp | 56 hrs | 125 000 | 111, 6 |
PacBio Sequel IIe
|
1 | Up to 30 Gb per SMRT Cell | Q30 = 99.9% | Up to 20 kb | 30 hrs | 495 000 | 41, 7 |
PacBio Revio
|
4 | Up to 90 Gb per SMRT Cell | Q33 = 99.95% | Up to 20 kb | 30 hrs | 779 000 | 11, 05 |
ONT MinION
|
1 | Up to 48 Gb | Simplex: Q20 = 99% Duplex: Q30 = 99.9% |
Up to 300 kb | Up to 72 hrs | 1080 | 9, 9 |
ONT GridION
|
5 | Up to 48 Gb per MinION flow-cell | Simplex: Q20 = 99% Duplex: Q30 = 99.9% |
Up to 300 kb | Up to 72 hrs | 68 830 | 9, 9 |
ONT PromethION (2 Solo, 24, 48)
|
2/24/48 | Up to 200 Gb per PromethION flow-cell | Simplex: Q20 = 99% Duplex: Q30 = 99.9% |
Up to 300 kb | Up to 72 hrs | P2 Solo: 9800 PromethION 24: 461 300 PromethION 48: 692 800 |
3 |
For each sequencing platform, the maximum number of flow-cells that can be run in parallel, the maximum theoretical throughput, the modal quality score, the mean read length, the sequencing run time and the expected platform cost are reported.
aCost ($)/Gb is estimated based on the cost per single sequencing run and the maximum yield per flow cell, considering the costs for a standard library preparation. The reported costs do not include target enrichment strategies, such as CRISPR/Cas9 enrichment. Cost per sequencing run is estimated from manufacturer’s reports.
While the maximum read length of MiSeq is generally suitable for characterizing most mutant alleles in the germline for pathologies with shorter expansions (e.g. HD) (17), or for ruling out expanded alleles when sequencing two alleles with below-threshold repeat counts (66), it may fall short for fully characterizing expanded alleles in disorders like DM2 (39). Table 2 highlights HD studies that utilized MiSeq for HTT CAG sizing, discussing its utility and limitations. Recent investigations by Ciosi et al. (38) and Mätlik et al. (17) used MiSeq to size expanded HTT CAG repeats, achieving resolutions up to ∼110 CAG repeats with as little as 20 ng of genomic DNA. However, concerns have been raised about MiSeq’s suitability for studying extremely long repeats arising from SI. Ciosi etal. highlighted these concerns in their comparison to orthogonal methods, while Handsaker et al. reported extreme somatic expansions linked to transcriptional dysregulation and tissue degeneration in HD (18,38). Similarly, MiSeq is not suitable to study SI in HD mouse models, where the CAG repeat in the germline frequently exceeds 100 CAG, such as in Q140, R6/1, R6/2 and zQ175 models (36,38,77,78). These limitations highlight the need for sequencing technologies capable of handling longer read lengths and addressing phasing issues intrinsic to short-read platforms, in studies involving dynamic repeat expansions.
Table 2.
Studies using sequencing-based approaches to characterize CAG repeats in HD
| Study | Enrichment method | Sequencing platform | Bioinformatic workflow | Max. num. CAG repeats detected | Required amount of DNA (μg) | On-target coverage |
|---|---|---|---|---|---|---|
| Miyatake et al. (123) |
Read Until | ONT GridION R9.4.1 | Tandem-genotypes + lamassemble + RepeatAnalysisTools | 44 | 2–3 μg | 15–37x |
| McAllister et al. (126) | PCR | Illumina MiSeq | ScaleHD | 55 | 20 ng | NA (ultra-high) |
| Höijer et al. (113) | Crispr/Cas9 (no-Amp) | PacBio RSII HiFi | HTT-repeat-analysis | 57 | Variable (5–20 μg) | 75–301x |
| Tsai et al.(107) | Crispr/Cas9 (no-Amp) | PacBio RSII HiFi | Alignment to custom references (Blasr) | 60 | 5–20 μg | 400–800x |
| Fang et al. (127) | PCR | ONT GridION R9.4.1, Sanger | DeepRepeat | 72 | NA | 2000–4000x |
| Stevanovski et al.(122) | Read Until (ReadFish) | ONT MinION R9.4.1, | Flye + Racon + TRF | 74 | 1.5–5 μg | 9–40x |
| Fang et al.(96) | PCR | ONT MinION R9.4.1 | NanoRepeat | 82 | 1 μg | >100x |
| Mätlik et al. (17) | PCR | Illumina MiSeq | Alignment to custom references (Burrow-Wheeler Aligner, BWA) | 113 | Up to 10 ng | NA (high-depth) |
| Ciosi et al.(38) | PCR | PacBio RSII HiFi | Alignment to custom references (BWA) | 550 | 20 ng | 577–1715 |
| Ciosi et al.(38) | PCR | Illumina MiSeq | Alignment to custom references (BWA) | 123 | 20 ng | 5262–111 873 |
| Tayloret al.(128) | PCR | PacBio Sequel IIe HiFi | RepeatDetector | 550 | NA | 623–60 466 |
| Taylor et al.(128) | PCR | Illumina MiSeq | RepeatDetector | 55 | 20 ng | 1886–147 924 |
| Handsaker et al.(18) | PCR + UMI | PacBio Sequel IIe HiFi | HTT-CAG | 842 | NA [4 ul complementary DNA (cDNA)] | NA |
For each study, the adopted enrichment method, sequencing platform, and bioinformatic workflow are reported, along with the maximum number of CAG repeats detected.
PCR-based approaches for triplet repeats characterization using long-read sequencing
The limitations of traditional methods have driven the development of single-molecule long-read sequencing platforms, mainly represented by Pacific Biosciences (PacBio) and ONT (84). These platforms, with their increased read lengths, have enabled researchers to address previously intractable biological problems, such as differential isoform usage and haplotype reconstruction, while also significantly improving the detection of structural variants and the assembly of genomes (63,79–83).
PacBio sequencing requires each target DNA molecule to be circularized into a structure known as SMRTbell, which uses hairpin adaptors. The technology utilizes optical nanostructures fabricated in a thin metallic film (zero-mode waveguide), with a polymerase immobilized at the bottom. As this polymerase incorporates single nucleotides, it emits nucleotide-specific fluorescence (84). This process produces Continuous Long Reads (CLR). In this protocol, both strands of the SMRTbell can be sequenced multiple times. The resulting CLR are then subdivided into multiple subreads, which are collapsed to generate a Circular Consensus Sequence (CCS). Each subread is reported to have ∼85% accuracy, but accuracy increases logarithmically with the number of passes the polymerase makes over the template, i.e. of subreads collapsed into a single CCS, reaching on average 99% with 10 passes and 99.8% with 20 passes (85,86). Since PacBio read lengths are constrained by the lifespan of the DNA polymerase, CCS reads are used to improve sequencing accuracy, although this comes at the cost of read length. For a short SMRTbell, the polymerase can synthesize the DNA template doing multiple passes, generating many subreads, that can then be collapsed into a single CCS read (65,84). Recent advancements have significantly improved PacBio CCS reads, with accuracy levels exceeding 99.8% and average read lengths of 15–20 kbp, now termed HiFi reads (86,87) (Figure 2A).
ONT sequencing platforms, on the other hand, rely on an array of nanopores embedded in an insulating membrane, submerged in an electrolytic solution. A constant voltage is applied across the membrane, causing ions to flow through each nanopore, generating an electric current that is measured in real-time. Driven by the voltage, single-stranded electrically charged DNA or RNA molecules translocate through the pores, with the speed controlled by a motor protein. As each nucleotide passes through the pore, it causes a specific disruption in the electric current, which varies depending on the nucleotide. In ONT’s R9.4.1 sequencing chemistry, typically 5–6 nucleotides can occupy a nanopore simultaneously. This collective effect of nucleotides on the electric signal, and possibly their methylation state (if PCR amplification is not used), is decoded by computational algorithms in real-time, enabling single-molecule sequencing (88–90).
However, homopolymer runs (long stretches of identical nucleotides, longer than 5–6) pose a challenge, as they produce a flat electric signal due to the lack of nucleotide diversity. The latest R10.4.1 sequencing chemistry has greatly improved performance also in these homopolymeric regions, allowing the generation of ultra-long reads with single-pass (a.k.a. ‘simplex’) single-molecule accuracy of over 99% (81,91). Furthermore, by pairing reads from both strands of a single DNA molecule to create ‘duplex’ reads, ONT claims to achieve accuracy levels of 99.9% (Q30) (Figure 2A).
Both PacBio and ONT sequencing technologies have been successfully coupled to PCR amplification to characterize triplet repeats up to 4 kbp long (38,56,65,92–95). However, PCR artifacts like stuttering, caused by polymerase slippage, and the preferential amplification of smaller repeats, can complicate the accurate determination of exact CAG repeat lengths, particularly when detecting expansions due to SI (37,38,65,94,95). Fang et al.(96) utilized ONT sequencing on 10 kbp amplicons from the HTT locus, in a study involving 983 individuals with HD. They developed a workflow that enabled the joint sizing of CAG and CCG repeats and the diploid genotyping of alleles containing up to 82 CAG repeats. However, their focus did not extend to detecting rare expansions. Conversely, Ciosi et al.(38) successfully characterized expanded HD alleles with up to ∼255 CAG repeats, using PacBio HiFi amplicon sequencing. Nevertheless, they encountered challenges in accurately characterizing repeats with ∼470 CAG, likely due to loading and/or sequencing bias toward smaller fragments on the PacBio RSII sequencing platform. This was evidenced by comparisons with Small pool-PCR and bulk-PCR capillary electrophoresis on the same samples. The comparisons revealed that longer triplet repeats were largely missed by PacBio HiFi sequencing of bulk PCR products, despite some PacBio CCS reads with over 450 CAG repeats (38) (Table 2).
Incorporating UMIs into PCR-based approaches for triplet repeats characterization
PCR-based approaches, while effective for amplifying specific DNA regions, have the drawback of preferentially amplifying shorter alleles. This bias can lead to the under-representation or even omission of sequencing reads from longer alleles in samples containing a mix of molecules with variable lengths. To mitigate this issue, PCR primers can be tagged with UMIs – random molecular barcodes ligated to template DNA molecules before amplification (97) (Figure 2A). UMIs enable bioinformatic correction of PCR artifacts after sequencing. By identifying sequencing reads sharing the same UMI, duplicates originating from PCR amplification of the same molecule can be identified and removed from the dataset. This process involves assigning reduced weight to each read carrying the same UMI, thereby improving the accuracy of allele quantification and reducing the impact of PCR bias (97).
Identifying UMI sequences in long-read datasets can be more challenging and typically involves building a reference database of high-quality UMIs using bioinformatic strategies. However, the benefits of obtaining highly accurate consensus sequences from each set of reads grouped by UMI are even higher. UMIs not only facilitate the deduplication of reads but also enable the generation of consensus sequences for each UMI group, combining reads with the same UMI. Due to the random distribution of sequencing errors across reads within each UMI group, the consensus sequences obtained exhibit significantly higher accuracy than individual reads alone, often exceeding 99.99% accuracy (Q40) (98–102).
The decision on which UMIs to retain can influence both sequencing accuracy and throughput. UMIs supported by fewer reads typically exhibit lower accuracy, prompting considerations to discard such UMIs. However, raising the threshold also results in a lower number of UMIs and molecules characterized (98). Handsaker and colleagues (18) developed a protocol based on PCR amplification of UMI-tagged molecules from HTT exon 1, followed by sequencing on the PacBio HiFi platform. Incorporating UMIs during first-strand cDNA synthesis was crucial in counteracting PCR bias toward shorter sequences. By leveraging UMI sequences, allelic proportions were preserved, ensuring accurate representation of each molecule amplified by PCR.
However, very long triplet repeats, on the order of tens of kbp, may be difficult to amplify using PCR, especially due to their extreme GC content. This high GC content is known to hamper PCR amplification by forming self-dimers and secondary structures in the DNA template (65,103). For these reasons, unless whole-genome, PCR-free sequencing data have previously been generated from the same sample for genome-wide variant calling (104–106), and there is no need to study somatically expanded alleles, PCR-free enrichment methods are recommended (38). A critical test of the ability of UMI-tagged amplicons to perform as accurately as PCR-free methods would involve multi-platform characterization of samples with known CAG lengths. Specifically, mixing DNA from multiple samples with variable CAG lengths in known proportions would enable a thorough assessment of each approach in terms of accuracy and sensitivity for detecting rare alleles (i.e. those present in a minority of cells). This experiment has not yet been conducted but is urgently needed.
PCR-free enrichment methods based on CRISPR/Cas9 for triplet repeats characterization using long-read sequencing
Recent advancements in molecular research have enabled the development of PCR-free enrichment methods, including those based on the CRISPR/Cas9 system. These methods allow for the enrichment of target regions over the genomic background, avoiding an amplification step and the associated biases (39,107–110).
This is the most common class of PCR-free enrichment methods, and can be coupled with either PacBio or ONT sequencing. PacBio amplification-free protocol, called No-Amp, involves cutting the SMRTbell templates containing the target region using Cas9 and a crRNA designed to be complementary to a sequence adjacent to the region of interest. A capture adapter is then ligated to the digested templates, specifically enriching for the region of interest (107). This protocol, when combined with PacBio HiFi reads, has successfully characterized short tandem repeats up to 20 kbp in length (108,111–114) (Figure 2B). Recently, PacBio introduced a new protocol named PureTarget, offering amplification-free HiFi reads across a panel of 20 genes, including those prone to repeat expansion and associated with neurological disorders. Thanks to the ultra-high-throughput Revio sequencing machine, which can produce up to 360 Gb in 24 h (Table 1), each of the 20 genes can be sequenced with high-coverage.
While ONT does not yet have an officially supported CRISPR/Cas9 protocol compatible with the latest R10.4.1 chemistry, an official protocol existed for the R9.4.1 chemistry, and many were proposed by the scientific community, following seminal work by Giesselmann and colleagues (39,115–118). Most of these protocols are based on dephosphorylation of DNA ends, followed by Cas9 cuts guided by a pair of crRNAs, and ligation of ONT sequencing adapters to phosphorylated ends near the region of interest (116) (Figure 2B). Overall, CRISPR/Cas9 enrichment combined with ONT sequencing has enabled the characterization of short tandem repeats up to 50 kbp in length, corresponding to ∼12 500 repeats of the CCTG motif (39).
Regarding the detection of CAG repeats in the HTT gene, no-one has yet applied CRISPR/Cas9 enrichment in combination with ONT sequencing. Only Tsai et al. (107) and Höijer et al.(113) have used the PacBio no-Amp protocol, starting with blood samples from HD patients, detecting up to 60 CAG repeats (Table 2). These protocols have not yet been thoroughly tested on HD brain samples; however, they may represent the most unbiased strategy for detecting rare extreme expansions in affected tissues. Moreover, their accuracy and sensitivity may be crucial for capturing subtle contraction events over short time scales, potentially enabling the monitoring of pharmacological treatments’ impact on SI.
PCR-free in-silico enrichment methods for triplet repeats characterization using long-read ONT sequencing
A second class of PCR-free enrichment methods, compatible only with ONT devices, exploits adaptive sampling – also known as ‘Read Until’ – to in-silico enrich for genomic regions of interest (119–123) (Figure 2C). Adaptive sampling refers to selectively sampling or ‘fishing out’ DNA fragments of interest from a pool, dynamically changing the sampling process based on observed data (124). This method allows Nanopore sequencers to reject individual molecules while they are being sequenced. Real-time analysis of the first portion of a molecule, as it passes through the pore, is performed. This initial read portion is computationally compared to a target sequence, and a quick decision is made to either fully sequence the molecule (if it matches the target) or reverse the voltage across the pore to eject the read, replacing it with a new one (121). Adaptive sampling, now available in MinKNOW ONT proprietary sequencing software, has been tested on MinION (coupled to an external High-Performance Computing cluster), GridION and PromethION devices, providing sufficient sequencing coverage for characterizing triplet repeats (122,123,125). These devices span the full range of ONT sequencers in terms of throughput and computational power. PromethION, ONT’s highest throughput sequencer, can run up to 48 independent flow-cells in parallel, generating up to 13.3 Tb in 72 h, roughly corresponding to 130 human genomes sequenced at 30× average coverage. Further details on these devices are provided in Table 1.
Stevanovski et al. (122) and Miyatake et al. (123) both tested ‘Read Until’ to enrich for multiple loci associated with triplet repeat disorders, including HTT, starting from patient-derived blood samples, and identifying up to 74 CAG (Table 2). This system represents a significant advantage for CAG sizing in HD and other triplet repeat diseases because it does not require any laborious ad hoc enrichment protocols. It only requires 1.5–5 μg of genomic DNA and a powerful computer with a suitable GPU for rapid base-calling.
However, the current enrichment levels achieved, about 5–10 fold over the genomic background (122), correspond to ∼100–200× coverage depth on a PromethION flow-cell for each target locus (including the HTT locus). While sufficient for germline genotyping, where the inherited number of CAG repeats is measured, this coverage may not allow the detection of rare alleles (<1% frequency within a sample) generated by extreme somatic mosaicism in neurons, which correspond to the right-tail of the distribution.
The bioinformatic challenge: deciphering CAG repeats from sequencing data
While the characterization of triplet repeats by means of sequencing-based approaches is becoming more and more popular, there is no general consensus or gold-standard strategy for the bioinformatic data analysis yet (129). This may be due to multiple reasons, including the highly variable quality of sequencing data that requires tailored analysis approaches – despite both PacBio and ONT having drastically improved sequencing accuracy in recent years (79) – the platform-specific raw data types that can be interrogated to retrieve additional information (e.g. raw current signal produced by ONT sequencers), and the plethora of biological questions the users may want to address (i.e. diploid or germline genotyping, interruptions detection and mosaicism quantification).
Although each bioinformatic tool for triplet repeat characterization has its own peculiarities regarding input data format, underlying algorithms, and implementation, these tools can be grouped into classes based on various criteria. These criteria include: (i) the type of input data utilized; (ii) the distinction between de-novoand reference-based methods; and (iii) the level of information provided, whether at the single-read versus allele-level. Table 3 reports available tools for triplet repeats characterization, along with the suitable type of sequencing data, their need for a reference sequence, their capability to detect mosaicism and interruptions in the canonical repeated motif and their capability to perform diploid genotyping.
Table 3.
Available bioinformatic tools for triplet repeat characterization from high-throughput sequencing data
| Tool | Input data type | Reference-based versus de novo | Mosaicism detection | Diploid genotyping | Interruptions detection |
|---|---|---|---|---|---|
| LobSTR (131) | Illumina whole-genome sequencing (WGS) | Reference-based | No | Yes | No |
| HipSTR (132) | Illumina WGS | Reference-based | No | Yes | No |
| TredPARSE (133) | Illumina WGS | Reference-based | No | Yes | No |
| STRetch (134) | Illumina WGS | Reference-based | No | Yes | No |
| exSTRA (135) | Illumina WGS | Reference-based | No | Yes | No |
| ExpansionHunter (106) | Illumina WGS | Reference-based | No | Yes | No |
| GangSTR (130) | Illumina WGS | Reference-based | No | Yes | No |
| STRling (136) | Illumina WGS | Reference-based | No | Yes | No |
| ScaleHD (126) | Illumina MiSeq | Reference-based | Yes | Yes | Yes |
| Repeat Detector (RD) (128) | Illumina MiSeq/PacBio | De novo | Yes | No | Only indirect |
| PacmonSTR (80) | PacBio | Reference-based | Yes | Yes | Yes |
| RepeatHMM (137) | PacBio/ONT | Reference-based | Yes | Yes | No |
| Tandem-genotypes (138) | PacBio/ONT | Reference-based | Yes | Yes | No |
| TRICoLOR (140) | PacBio/ONT | Reference-based | No | Yes | No |
| Straglr (141) | PacBio/ONT | Reference-based | Yes | Yes | No |
| NanoRepeat (96) | PacBio/ONT | Reference-based | Yes | Yes | No |
| NanoSTR (143) | ONT | Reference-based | No | Yes | No |
| NanoSatellite (105) | ONT raw R9.4 | Reference-based | Yes | Yes | Only indirect |
| STRique (115) | ONT raw R9.4 | Reference-based | Yes | No | No |
| DeepRepeat (127) | ONT raw R9.4 | Reference-based | No | Yes | No |
| WarpSTR (148) | ONT raw R9.4 | Reference-based | Yes | Yes | No |
| Tandem Repeat Genotyping Tool (TRGT) (146) | PacBio HiFi | Reference-based | Yes | Yes | Yes |
| Noise Cancelling Repeat Finder (NCRF) (139) | PacBio/ONT | De novo | Yes | No | No |
| HMMSTR (144) | PacBio/ONT | De novo | Yes | Yes | No |
| Lamassemble (142) | PacBio/ONT | De novo | No | No | Yes |
| MosaicViewer (56) | ONT | De novo | Yes | No | Only indirect |
| CharONT2 (39) | ONT | De novo | Only qualitative |
Yes | Yes |
For each tool, we report the input data, the distinction between reference-based and de novo approaches, and the capabilities of detecting mosaicism, performing diploid genotyping and identifying interruptions in the repeated motif.
Choosing the appropriate CAG sizing tool according to the available input data
Regarding input data, tools may be designed to work with Illumina short paired-end reads, PacBio and ONT long-reads, PacBio HiFi long reads or ONT raw electric signals.
Illumina short-reads: A first group of tools is designed for Illumina short-reads. Although these reads may not be long enough to cover expanded triplet repeats – reaching up to 115 CAG repeats with MiSeq amplicon sequencing, algorithms have been developed to estimate repeat length from Illumina WGS data, which may potentially be able genotyping triplet repeats longer than the read length (130). While early tools like LobSTR (131) and HipSTR (132) mainly exploited sequencing reads enclosing the full repeat, more recently developed tools such as TredPARSE (133), STRetch (134), exSTRA (135), ExpansionHunter (106), GangSTR (130) and STRling (136) also utilize the average fragment length of the sequencing library, the expected sequencing coverage in the target locus, and reads partially enclosing the repeat. This allows performing diploid genotyping, even when the triplet repeats are longer than the sequenced DNA fragment.
PacBio and ONT long-reads: A second group of tools is based on PacBio or ONT long-read sequencing data, which exploit direct information for triplet repeat characterization. However, the higher error rate typical of long-read sequencing platforms requires the adoption of sophisticated probabilistic methods to reconstruct the most likely alleles of the target gene (129). Following PacmonSTR (80), which was the first probabilistic approach developed to characterize short tandem repeats sequenced with PacBio, several tools have been created to handle noisy long-reads from PacBio and ONT platforms. These include RepeatHMM (137), Tandem-genotypes (138), NCRF (139), TRICoLOR (140), Straglr (141), lamassemble (142), MosaicViewer (56), CharONT2 (39), NanoRepeat (96), NanoSTR (143) and HMMSTR (144). Notably, many of these tools adapt Tandem Repeats Finder (TRF) (145) – a tool developed 25 years ago for identifying tandem repeats in genome assemblies – to accommodate the higher error rate (56,137,139,141). With the increasing accuracy of PacBio HiFi reads, new tools have been developed, such as RD (128), TRGT (146) and RepeatAnalysisTools (provided by PacBio). These tools enable precise sizing of triplet repeats and accurate assessment of somatic mosaicism, thanks to the high sequencing accuracy of each read.
ONT long-reads (raw signal):Although ONT platforms have similarly reduced their error rate, very long repetitive regions remain challenging to base-call, i.e. to accurately convert from the raw electric signal into the sequence of nucleotides (105,127,147). For this reason, several bioinformatic tools have been developed that exploit the raw electric signal produced by ONT sequencers, such as NanoSatellite (105), STRique (115), DeepRepeat (127) and WarpSTR (148). These tools use the base-called sequence only to localize the triplet repeat in the raw electric signal from each read, and rely on raw signal (a.k.a. squiggle) analysis for inferring the number of repeats. These tools provided accurate quantification of triplet repeat length from ONT data generated with R9.4.1 sequencing chemistry, but they have quickly become obsolete, as they are not compatible with the latest R10.4.1 sequencing chemistry, which fully replaced the previous version in March 2024. Unfortunately, no tools based on the analysis of raw signals generated with R10.4.1 sequencing chemistry are available yet.
CAG sizing tools exploiting a reference sequence: pros and cons compared to de novo tools
Aside from the required input data, tools can be classified based on their need for a reference sequence for aligning the reads (alignment-based tools versus reference-free or de-novo tools) (129). Alignment-based tools represent the majority of the available triplet repeat characterization tools. These tools require aligning the sequencing reads to the reference genome, allowing them to genotype multiple triplet repeats at once, enabling genome-wide scans (141). Additionally, users can adopt custom reference sequences to distinguish reads from the two alleles, by leveraging preferential mapping based on expected triplet repeats length. An example of this is ScaleHD, a tool developed by McAllister and colleagues, which maps reads to a reference set composed of 4000 possible HTT allele structures with variable CAG/CCG length, predicting the most likely genotypes for a sample (126). Conversely, reference-free tools may be the best solution to save computational time and resources required for reads alignment, especially in the case of extremely long triplet repeats that may cause alignment issues, particularly with short high-complexity sequences flanking the repeat (115,138,141). However, these tools often require higher sequencing coverage to obtain an accurate representation of the two alleles (129). To minimize alignment issues, which may be frequent in case of long repeat expansions, Taylor and colleagues (128) developed RD, an alignment-free tool flexible enough to work with datasets from multiple repeated motifs and sequencing platforms, such as Illumina MiSeq and PacBio HiFi. Notably, RD showed high agreement with available tools on 609 PCR-amplified blood-derived samples from HD, predicting the same modal allele length for 98.3% of the samples compared to ScaleHD (128) (Table 2).
CAG sizing tools providing allele-level versus single-read information
A third possible classification of bioinformatic tools for deciphering CAG repeats is based on the provided output: allele-level versus single-read information. The distinction separates tools that provide an aggregate genotype for the two alleles from those that analyze each single read and then may combine them into the genotype of the two alleles.
Allele-level genotype: The allele-level genotype may be achieved either through de novoassembly/consensus sequence generation or through reference-based variant-calling approaches. In de novoassembly, all on-target reads are assembled to obtain one consensus sequence for each allele, which then undergoes triplet repeat counting. In variant-calling approaches, all reads are aligned to a reference sequence, and differences in the reads compared to the reference in the target locus are identified and represented as heterozygous or homozygous variants. This aggregated allele-level information is crucial for detecting single nucleotide repeat interruptions. Multiple reads provide the evidence necessary for reliable detection of repeat interruptions, ruling out sequencing errors at specific genomic coordinate in single reads (56,146). In fact, a single long-read may contain a spurious single nucleotide variant with biological significance (such as the LOI, associated with earlier HD onset), but only due to a random sequencing error, rather than a true biological polymorphism.
Single-read information: Some other tools provide triplet repeat length and motif for each sequencing read. These tools may also include a genotyping step that aggregates read-level information into the genotype for the two alleles. In this case, the software first provides triplet repeats length for each on-target read, then combines single-read results into up to two clusters, corresponding to the two alleles, providing the diploid genotype. For example, this aggregation can be achieved with k-means clustering or by exploiting Gaussian Mixture Models (GMM) (39,96). Fang and colleagues (96) developed NanoRepeat for HTT alleles genotyping, enabling the joint quantification of the prone-to-expansion CAG repeat and the adjacent CCG repeat. NanoRepeat uses GMMs to group reads into the two alleles based on their CAG and CCG size, offering additional precision in genotyping samples with similar CAG repeat sizes for the two alleles.
Importance of single-read genotyping: Single-read genotyping is crucial in cases of repeat instability, as somatic expansions may affect a small portion of cells to varying degrees (17,38,39,56). For instance, the ‘armadillo shape’ repeat length distribution observed in postmortem HD brains by Handsaker and colleagues (18) shows that the majority of cells have a CAG size similar to the germline, with moderate expansions, while very few cells exhibit extreme expansions. These extreme expansions, though potentially key in HD pathogenesis, do not increase the modal triplet repeat length, as they are a minority (18). Several tools enable qualitative assessment of SI (38,56,117,146), through visual inspection of aligned reads in a genome browser or by showing the CAG size distributions of all reads. However, this qualitative information is insufficient for accurately quantifying the response to drug candidates in terms of modulation of SI. Quantitative instability indexes originally developed for capillary electrophoresis data should be adapted to long-read sequencing, considering the reduced noise and suitability for time-course experimental designs (49,149,150).
Choosing the right tool: In short, when selecting a tool, the type of available data is the primary consideration. Tools providing single-read genotyping are preferred over those offering only the aggregated diploid genotype, if sequencing coverage allows for studying somatic mosaicism. De novo tools may be preferred for studying extreme somatic expansions, as corresponding sequencing reads may align poorly to the reference, and be discarded by alignment-based tools. In case TRGT is used, Tandem Repeat Visualization represents a companion tool for visually inspecting the reads aligned to the repeat alleles, thus determining the accuracy of the genotype results (144). Regardless of the chosen tool, visual assessment of sequencing reads in a genome browser, such as Integrative Genomics Viewer, is recommended to exclude any unintended or unexpected software behaviour (151). Although an independent benchmarking study exploiting samples with known genotype is still missing, some of the tools were benchmarked on both simulated and real long-read sequencing data. In particular, in the manuscript presenting Straglr, the tool was shown to outperform both RepeatHMM and Tandem-genotypes (141). Similarly, a recent paper reported higher accuracy for HMMSTR compared to Straglr across multiple datasets (144).
Achieving the most accurate HTT CAG repeat characterization
As previously discussed, a range of target enrichment methods, sequencing platforms and analysis pipelines have been developed for triplet repeats characterization. While each method can address specific biological questions in particular use cases, several key factors must be considered to select the most appropriate approach. These factors include the expected maximum triplet repeat length in the disease gene, the need to assess somatic mosaicism in brain tissue, the detection of repeat interruptions and contractions, the amount of available DNA and budget constraints. In Figure 3, we propose a strategy for designing the optimal HTT CAG repeat characterization experiment, based on the specific biological questions the users need to address.
Figure 3.
Proposed experimental strategies to characterize CAG repeats in HD. Based on the biological questions users may wish to address, a workflow outlining the optimal experimental setup is proposed. An estimate of the costs for each approach is also provided: $ represents 10$; $$ represents 100$; $$$ represents 1000$.
Retrieving DNA sequence composition of triplet repeats with PCR-based sequencing approaches
Of the >40 documented disorders caused by triplet repeat expansions, the majority have germline pathogenic allele size thresholds below 50 repeats, corresponding to sequences shorter than 150 bp (16,129). Therefore, for these disorders, triplet repeat characterization can be effectively achieved using PCR enrichment coupled with short-read sequencing, such as Illumina MiSeq (Figure 3). However, users should note that longer somatic expansions may go undetected due to the limited read length of this platform.
Cost and efficiency of PCR-based sequencing protocols
PCR-based amplification protocols (i.e. amplicon sequencing) are reported to cost around $10 per sample, provided that a sufficient number of samples can be multiplexed in a single sequencing run to fully utilize the capacity of the latest Illumina MiSeq and PacBio Sequel IIe or Revio sequencing platforms (38). ONT also offers the more affordable MinION sequencer with lower throughput, where each flow-cell can be used for a smaller number of samples (e.g. 24) (Table 1).
Multiplexing involves attaching a sample-specific barcode to each molecule before pooling the samples for a single sequencing run. After sequencing, reads are assigned to their respective samples using these unique barcodes, a process called demultiplexing. Illumina and PacBio generate a fixed number of reads for run, with Illumina MiSeq producing up to 15 Gb and the PacBio Revio up to 90 Gb per flow-cell.
To optimize costs, multiple samples are multiplexed to fully exploit the flow-cell capacity. This can sometimes lead to delays while waiting to collect enough samples, particularly for protocols with specific requirements, such as HTT samples sequenced on Illumina MiSeq that need an extended read length of 400 bp for read 1.
Retaining allelic proportions for somatic mosaicism assessment
For diseases involving long triplet repeats or where it is critical to maintain allelic proportions for accurate somatic mosaicism, a PCR-free long-read sequencing approach represents the ideal choice (Figure 3). However, these protocols typically require larger quantities of genomic DNA (5–10 μg) (39,104,112,114,116,117,138) and are reported to cost several hundred dollars per sample (38,39). PacBio HiFi reads offer higher single-molecule accuracy (ranging from 99.5% to 99.9%, or Q30) (152) compared to the latest chemistry of ONT long-reads (>99% accuracy for single-pass ‘simplex’ reads, or Q20+) (152), and are less prone to systematic errors in repetitive regions (79,110,147). However, ONT reads provide significant advantages.
First, ONT’s portable MinION sequencer, which does not require large capital investments, allows easy setup of sequencing experiments in various environments, such as hospital bedsides or limited-resource settings like rainforests (63,89,152–154) (Table 1). Second, ONT sequencing does not require a predetermined number of reads, allowing users to stop sequencing once sufficient data have been obtained (153).
Third, ONT has introduced adaptive sampling (the ‘Read Until’ mentioned above), an in-silico enrichment strategy for targeted sequencing. Read Until has been reported by both Stevanovski et al.(122) and Miyatake et al.(123) to provide sufficient coverage to genotype 59 and 37 loci associated with triplet repeat expansions, respectively. However, current coverage levels (∼5–10 fold enrichment) generated for each locus may not be sufficient to assess rare alleles generated by somatic mosaicism (122,123).
A possible solution to the PCR versus PCR-free dilemma
A potential compromise between PCR-based and PCR-free methods is the use of PCR-based approaches with UMI-tagged primers (38) (Figure 3). This technique requires as little DNA as standard PCR protocols (∼20 ng), but produces a consensus sequence for each molecule (i.e. for each group of reads tagged with the same UMI), reducing PCR bias (38). This approach has been coupled with ONT and PacBio sequencing platforms to generate up to 30 000 consensus sequences with Q40 sequencing quality from a single flow-cell (98). However, the application of UMI-based long-read sequencing has only been minimally explored for triplet repeats characterization, and only in the context of single-cell cDNA sequencing (18).
Choosing the most suitable CAG sizing pipeline
In bioinformatics, a comprehensive benchmarking of tools for triplet repeat characterization is still lacking (129). However, several long-read alignment-based tools, such as Tandem-genotypes (138), Straglr (141), HMMSTR (144) and TRGT (146) have demonstrated accurate triplet repeat length estimation in both simulated and real datasets. These tools can also assess somatic mosaicism quantitatively, through instability indexes. For detecting interruptions within repeats, reference-free tools like CharONT2 (39) and lamassemble (142), can be used for de novo assembly of expanded repeats. General-purpose long-read assemblers like Flye (155) and Canu (156) are also effective (129). Often, the best approach involves running multiple tools on the same dataset, and cross-checking results to obtain a clearer picture of the triplet repeat sequences and their features.
Conclusions
Accurate characterization of triplet repeats requires careful selection of the target enrichment strategy, sequencing platform, and bioinformatic analysis pipeline. We recommend prioritizing long-read sequencing platforms over short-read ones, particularly when extreme expansion events associated with somatic mosaicism are expected. Additionally, to retain true allelic proportions, it is crucial to use protocols that avoid known PCR biases. Bioinformatic tools should be selected based on their compatibility with the data type and their ability to provide comprehensive information, including diploid genotype, interruption detection and mosaicism identification. Visual inspection of aligned reads in a genome browser is always recommended as a quality check.
Overall, the variability in triplet repeat lengths, enrichment protocols, sequencing platforms and data analysis tools make comparisons across studies extremely challenging. Moreover, there is growing interest in combining accurate triplet repeat characterization with transcriptional profiling from the same cells, as demonstrated in pioneering work by Handsaker and colleagues (18). Therefore, a benchmarking study using DNA standards with known genotypes is urgently needed. Such a study would help dissect the impact of each methodological choice on the accuracy of the protocol, paving the way for the integration of sequencing-based protocols for short tandem repeat characterization into studies of disease mechanisms, clinical guidelines and the monitoring of drug candidates for HD treatment. In fact, a thorough benchmarking study is essential to ensure the standardization and reproducibility across approaches, clarifying how different techniques and algorithms perform. By defining a set of rigorous quality checks throughout the workflow, it will be possible to assess the technical soundness of the results, aiding in the transition from research to clinical care.
Contributor Information
Simone Maestri, Department of Biosciences, University of Milan, Street Giovanni Celoria, 26, 20133, Milan, Italy; INGM, Istituto Nazionale Genetica Molecolare ‘Romeo ed Enrica Invernizzi’, Street Francesco Sforza, 35, 20122, Milan, Italy.
Davide Scalzo, Department of Biosciences, University of Milan, Street Giovanni Celoria, 26, 20133, Milan, Italy; INGM, Istituto Nazionale Genetica Molecolare ‘Romeo ed Enrica Invernizzi’, Street Francesco Sforza, 35, 20122, Milan, Italy.
Gianluca Damaggio, Department of Biosciences, University of Milan, Street Giovanni Celoria, 26, 20133, Milan, Italy; INGM, Istituto Nazionale Genetica Molecolare ‘Romeo ed Enrica Invernizzi’, Street Francesco Sforza, 35, 20122, Milan, Italy.
Martina Zobel, Department of Biosciences, University of Milan, Street Giovanni Celoria, 26, 20133, Milan, Italy; INGM, Istituto Nazionale Genetica Molecolare ‘Romeo ed Enrica Invernizzi’, Street Francesco Sforza, 35, 20122, Milan, Italy.
Dario Besusso, Department of Biosciences, University of Milan, Street Giovanni Celoria, 26, 20133, Milan, Italy; INGM, Istituto Nazionale Genetica Molecolare ‘Romeo ed Enrica Invernizzi’, Street Francesco Sforza, 35, 20122, Milan, Italy.
Elena Cattaneo, Department of Biosciences, University of Milan, Street Giovanni Celoria, 26, 20133, Milan, Italy; INGM, Istituto Nazionale Genetica Molecolare ‘Romeo ed Enrica Invernizzi’, Street Francesco Sforza, 35, 20122, Milan, Italy.
Data availability
No new data were generated or analyzed in support of this research.
Funding
European Research Council, Advanced Grant [742436]; NSC-Reconstruct Consortium, European Union's Horizon 2020 Research and Innovation Program [874758]; C.H.D.I. Foundation, New York, U.S.A. [JSC A11103]; Leslie Gehry Prize for Innovation in Science from the Hereditary Disease Foundation (New York, U.S.A.); Fondazione Telethon [GMR23T1059 and GMR23T1216]; Ministero dell'Istruzione, dell'Università e della Ricerca [2022LBENTH]. Funding for open access charge: H2020 European Research Council Grant [742436].
Conflict of interest statement. None declared.
References
- 1. Ellegren H. Microsatellites: simple sequences with complex evolution. Nat. Rev. Genet. 2004; 5:435–445. [DOI] [PubMed] [Google Scholar]
- 2. Kozlowski P., Sobczak K., Krzyzosiak W.J.. Trinucleotide repeats: triggers for genomic disorders?. Genome Medicine. 2010; 2:29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Kashi Y., King D.G.. Simple sequence repeats as advantageous mutators in evolution. Trends Genet. 2006; 22:253–259. [DOI] [PubMed] [Google Scholar]
- 4. Iennaco R., Formenti G., Trovesi C., Rossi R.L., Zuccato C., Lischetti T., Bocchi V.D., Scolz A., Martínez-Labarga C., Rickards O.et al.. The evolutionary history of the polyQ tract in huntingtin sheds light on its functional pro-neural activities. Cell Death Differ. 2022; 29:293–305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. López Castel A., Cleary J.D., Pearson C.E.. Repeat instability as the basis for human diseases and as a potential target for therapy. Nat. Rev. Mol. Cell Biol. 2010; 11:165–170. [DOI] [PubMed] [Google Scholar]
- 6. La Spada A.R., Taylor J.P.. Repeat expansion disease: progress and puzzles in disease pathogenesis. Nat. Rev. Genet. 2010; 11:247–258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Orr H.T., Zoghbi H.Y.. Trinucleotide repeat disorders. Annu. Rev. Neurosci. 2007; 30:575–621. [DOI] [PubMed] [Google Scholar]
- 8. MacDonald M.E., Ambrose C.M., Duyao M.P., Myers R.H., Lin C., Srinidhi L., Barnes G., Taylor S.A., James M., Groot N.et al.. A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington’s disease chromosomes. Cell. 1993; 72:971–983. [DOI] [PubMed] [Google Scholar]
- 9. MacDonald M.E., Gines S., Gusella J.F., Wheeler V.C.. Huntington’s disease. Neuromolecular Med. 2003; 4:7–20. [DOI] [PubMed] [Google Scholar]
- 10. Bakels H.S., Roos R.A.C., van Roon-Mom W.M.C., de Bot S.T.. Juvenile-onset Huntington disease pathophysiology and neurodevelopment: a review. Mov. Disord. 2022; 37:16–24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Genetic Modifiers of Huntington’s Disease (GeM-HD) Consortium. Electronic address: gusella@helix.mgh.harvard.edu, Genetic Modifiers of Huntington’s Disease (GeM-HD) Consortium CAG repeat not polyglutamine length determines timing of Huntington’s disease onset. Cell. 2019; 178:887–900. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Zuccato C., Valenza M., Cattaneo E.. Molecular mechanisms and potential therapeutical targets in Huntington’s disease. Physiol. Rev. 2010; 90:905–981. [DOI] [PubMed] [Google Scholar]
- 13. Sari Y. Huntington’s disease: from mutant Huntingtin protein to neurotrophic factor therapy. Int. J. Biomed. Sci. 2011; 7:89–100. [PMC free article] [PubMed] [Google Scholar]
- 14. Telenius H., Almqvist E., Kremer B., Spence N., Squitieri F., Nichol K., Grandell U., Starr E., Benjamin C., Castaldo I.. Somatic mosaicism in sperm is associated with intergenerational (CAG)n changes in Huntington disease. Hum. Mol. Genet. 1995; 4:189–195. [DOI] [PubMed] [Google Scholar]
- 15. Ridley R.M., Frith C.D., Crow T.J., Conneally P.M.. Anticipation in Huntington’s disease is inherited through the male line but may originate in the female. J. Med. Genet. 1988; 25:589–595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Budworth H., McMurray C.T.. A brief history of triplet repeat diseases. Methods Mol. Biol. 2013; 1010:3–17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Mätlik K., Baffuto M., Kus L., Deshmukh A.L., Davis D.A., Paul M.R., Carroll T.S., Caron M.-C., Masson J.-Y., Pearson C.E.et al.. Cell-type-specific CAG repeat expansions and toxicity of mutant Huntingtin in human striatum and cerebellum. Nat. Genet. 2024; 56:383–394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Handsaker R.E., Kashin S., Reed N.M., Tan S., Lee W.-S., McDonald T.M., Morris K., Kamitaki N., Mullally C.D., Morakabati N.et al.. Long somatic DNA-repeat expansion drives neurodegeneration in Huntington disease. 2024; bioRxiv doi:20 May 2024, preprint: not peer reviewed 10.1101/2024.05.17.592722 [DOI]
- 19. Roy J.C.L., Vitalo A., Andrew M.A., Mota-Silva E., Kovalenko M., Burch Z., Nhu A.M., Cohen P.E., Grabczyk E., Wheeler V.C.et al.. Somatic CAG expansion in Huntington’s disease is dependent on the MLH3 endonuclease domain, which can be excluded via splice redirection. Nucleic Acids Res. 2021; 49:3907–3918. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Mouro Pinto R., Arning L., Giordano J.V., Razghandi P., Andrew M.A., Gillis T., Correia K., Mysore J.S., Grote Urtubey D.-M., Parwez C.R.et al.. Patterns of CAG repeat instability in the central nervous system and periphery in Huntington’s disease and in spinocerebellar ataxia type 1. Hum. Mol. Genet. 2020; 29:2551–2567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Bizzotto S., Walsh C.A.. Genetic mosaicism in the human brain: from lineage tracing to neuropsychiatric disorders. Nat. Rev. Neurosci. 2022; 23:275–286. [DOI] [PubMed] [Google Scholar]
- 22. Kennedy L., Evans E., Chen C.-M., Craven L., Detloff P.J., Ennis M., Shelbourne P.F.. Dramatic tissue-specific mutation length increases are an early molecular event in Huntington disease pathogenesis. Hum. Mol. Genet. 2003; 12:3359–3367. [DOI] [PubMed] [Google Scholar]
- 23. Swami M., Hendricks A.E., Gillis T., Massood T., Mysore J., Myers R.H., Wheeler V.C.. Somatic expansion of the Huntington’s disease CAG repeat in the brain is associated with an earlier age of disease onset. Hum. Mol. Genet. 2009; 18:3039–3047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Chong S.S., McCall A.E., Cota J., Subramony S.H., Orr H.T., Hughes M.R., Zoghbi H.Y.. Gametic and somatic tissue–specific heterogeneity of the expanded SCA1 CAG repeat in spinocerebellar ataxia type 1. Nat. Genet. 1995; 10:344–350. [DOI] [PubMed] [Google Scholar]
- 25. Lopes-Cendes I., Maciel P., Kish S., Gaspar C., Silveira I., Robitaille Y., Clark H.B., Koeppen A.H., Nance M., Schut L.et al.. Somatic mosaicism in the central nervous system in spinocerebellar ataxia type 1 and machado-joseph disease. Ann. Neurol. 1996; 40:199–206. [DOI] [PubMed] [Google Scholar]
- 26. Hashida H., Goto J., Kurisaki H., Mizusawa H., Kanazawa I.. Brain regional differences in the expansion of a CAG repeat in the spinocerebellar ataxias: dentatorubral-pallidoluysian atrophy, machado-joseph disease, and spinocerebellar ataxia type 1. Ann. Neurol. 1997; 41:505–511. [DOI] [PubMed] [Google Scholar]
- 27. Thornton C.A., Johnson K., Moxley R.T. III. Myotonic dystrophy patients have larger CTG expansions in skeletal muscle than in leukocytes. Ann. Neurol. 1994; 35:104–107. [DOI] [PubMed] [Google Scholar]
- 28. Monckton D.G., Wong L.-J.C., Ashizawa T., Caskey C.T.. Somatic mosaicism, germline expansions, germline reversions and intergenerational reductions in myotonic dystrophy males: small pool PCR analyses. Hum. Mol. Genet. 1995; 4:1–8. [DOI] [PubMed] [Google Scholar]
- 29. Morales F., Couto J.M., Higham C.F., Hogg G., Cuenca P., Braida C., Wilson R.H., Adam B., del Valle G., Brian R.et al.. Somatic instability of the expanded CTG triplet repeat in myotonic dystrophy type 1 is a heritable quantitative trait and modifier of disease severity. Hum. Mol. Genet. 2012; 21:3558–3567. [DOI] [PubMed] [Google Scholar]
- 30. Morales F., Vásquez M., Corrales E., Vindas-Smith R., Santamaría-Ulloa C., Zhang B., Sirito M., Estecio M.R., Krahe R., Monckton D.G.. Longitudinal increases in somatic mosaicism of the expanded CTG repeat in myotonic dystrophy type 1 are associated with variation in age-at-onset. Hum. Mol. Genet. 2020; 29:2496–2507. [DOI] [PubMed] [Google Scholar]
- 31. Genetic Modifiers of Huntington’s Disease (GeM-HD) Consortium Identification of genetic factors that modify clinical onset of Huntington’s disease. Cell. 2015; 162:516–526. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Ferguson R., Goold R., Coupland L., Flower M., Tabrizi S.J.. Therapeutic validation of MMR-associated genetic modifiers in a human ex vivo model of Huntington disease. Am. J. Hum. Genet. 2024; 111:1165–1183. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Wright G.E.B., Collins J.A., Kay C., McDonald C., Dolzhenko E., Xia Q., Bečanović K., Drögemöller B.I., Semaka A., Nguyen C.M.et al.. Length of uninterrupted CAG, independent of polyglutamine size, results in increased somatic instability, hastening onset of Huntington disease. Am. Hum. Genet. 2019; 104:1116–1126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Genetic Modifiers of Huntington's Disease (GeM-HD) Consortium Lee J.-M., McLean Z.L., Correia K., Shin J.W., Lee S., Jang J.-H., Lee Y., Kim K.-H., Choi D.E.. Genetic modifiers of somatic expansion and clinical phenotypes in Huntington's disease reveal shared and tissue-specific effects. 2024; bioRxiv doi:18 June 2024, preprint: not peer reviewed 10.1101/2024.06.10.597797 [DOI]
- 35. Hong E.P., MacDonald M.E., Wheeler V.C., Jones L., Holmans P., Orth M., Monckton D.G., Long J.D., Kwak S., Gusella J.F.et al.. Huntington’s disease pathogenesis: two sequential components. J. Huntingtons Dis. 2021; 10:35–51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Wang N., Zhang S., Langfelder P., Ramanathan L., Plascencia M., Gao F., Vaca R., Gu X., Deng L., Dionisio L.E.et al.. Msh3 and Pms1 set neuronal CAG-repeat migration rate to drive selective striatal and cortical pathogenesis in HD mice. 2024; bioRxiv doi:15 July 2024, preprint: not peer reviewed 10.1101/2024.07.09.602815 [DOI]
- 37. Chintalaphani S.R., Pineda S.S., Deveson I.W., Kumar K.R.. An update on the neurological short tandem repeat expansion disorders and the emergence of long-read sequencing diagnostics. Acta Neuropathol. Commun. 2021; 9:98. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Ciosi M., Cumming S.A., Chatzi A., Larson E., Tottey W., Lomeikaite V., Hamilton G., Wheeler V.C., Pinto R.M., Kwak S.et al.. Approaches to sequence the HTT CAG repeat expansion and quantify repeat length variation. J. Huntingtons Dis. 2021; 10:53–74. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Alfano M., De Antoni L., Centofanti F., Visconti V.V., Maestri S., Degli Esposti C., Massa R., D’Apice M.R., Novelli G., Delledonne M.et al.. Characterization of full-length CNBP expanded alleles in myotonic dystrophy type 2 patients by Cas9-mediated enrichment and nanopore sequencing. eLife. 2022; 11:e80229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Warner J.P., Barron L.H., Goudie D., Kelly K., Dow D., Fitzpatrick D.R., Brock D.J.. A general method for the detection of large CAG repeat expansions by fluorescent PCR. J. Med. Genet. 1996; 33:1022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Warner J.P., Barron L.H., Brock D.J.. A new polymerase chain reaction (PCR) assay for the trinucleotide repeat that is unstable and expanded on Huntington’s disease chromosomes. Mol. Cell. Probes. 1993; 7:235–239. [DOI] [PubMed] [Google Scholar]
- 42. Andrew S.E., Goldberg Y.P., Theilmann J., Zeisler J., Hayden M.R.. A CCG repeat polymorphism adjacent to the CAG repeat in the Huntington disease gene: implications for diagnostic accuracy and predictive testing. Hum. Mol. Genet. 1994; 3:65–67. [DOI] [PubMed] [Google Scholar]
- 43. Massey T., McAllister B., Jones L.. Precious S.V., Rosser A.E., Dunnett S.B.. Methods for assessing DNA repair and repeat expansion in Huntington’s disease. Methods in Molecular Biology. 2018; 1780:Humana Press, Clifton, N.J; 483–495. [DOI] [PubMed] [Google Scholar]
- 44. Guida M., Fenwick R.G., Papp A.C., Snyder P.J., Sedra M., Prior T.W.. Southern transfer protocol for confirmation of Huntington disease. Clin. Chem. 1996; 42:1711–1712. [PubMed] [Google Scholar]
- 45. Day J.W., Ricker K., Jacobsen J.F., Rasmussen L.J., Dick K.A., Kress W., Schneider C., Koch M.C., Beilman G.J., Harrison A.R.et al.. Myotonic dystrophy type 2. Neurology. 2003; 60:657–664. [DOI] [PubMed] [Google Scholar]
- 46. Filipovic-Sadic S., Sah S., Chen L., Krosting J., Sekinger E., Zhang W., Hagerman P.J., Stenzel T.T., Hadd A., Latham G.J.et al.. A novel FMR1 PCR method that reproducibly amplifies fragile X full mutations in concordance with southern blotting and reliably detects low abundance expanded alleles. Clin. Chem. 2010; 56:399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Spector E., Behlmann A., Kronquist K., Rose N.C., Lyon E., Reddi H.V.. Laboratory testing for fragile X, 2021 revision: a technical standard of the American College of Medical Genetics and Genomics (ACMG). Genet. Med. 2021; 23:799–812. [DOI] [PubMed] [Google Scholar]
- 48. Vnencak-Jones C.L. Potter N.T. Fluorescence PCR and GeneScan® analysis for the detection of CAG repeat expansions associated with Huntington’s disease. Neurogenetics: Methods and Protocols. 2003; Totowa, NJ: Springer New York; 101–108. [DOI] [PubMed] [Google Scholar]
- 49. Lee J.-M., Zhang J., Su A.I., Walker J.R., Wiltshire T., Kang K., Dragileva E., Gillis T., Lopez E.T., Boily M.-J.et al.. A novel approach to investigate tissue-specific trinucleotide repeat instability. BMC Syst. Biol. 2010; 4:29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Saluto A., Brussino A., Tassone F., Arduino C., Cagnoli C., Pappi P., Hagerman P., Migone N., Brusco A.. An enhanced polymerase chain reaction assay to detect pre- and full mutation alleles of the fragile X mental retardation 1 gene. J. Mol. Diagn. 2005; 7:605–612. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Kamsteeg E.-J., Kress W., Catalli C., Hertz J.M., Witsch-Baumgartner M., Buckley M.F., van Engelen B.G.M., Schwartz M., Scheffer H.. Best practice guidelines and recommendations on the molecular diagnosis of myotonic dystrophy types 1 and 2. Eur. J. Hum. Genet. 2012; 20:1203–1208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52. Jama M., Millson A., Miller C.E., Lyon E.. Triplet repeat primed PCR simplifies testing for Huntington disease. J. Mol. Diagn. 2013; 15:255–262. [DOI] [PubMed] [Google Scholar]
- 53. Loureiro J.R., Oliveira C.L., Sequeiros J., Silveira I.. A repeat-primed PCR assay for pentanucleotide repeat alleles in spinocerebellar ataxia type 37. J. Hum. Genet. 2018; 63:981–987. [DOI] [PubMed] [Google Scholar]
- 54. Losekoot M., van Belzen M.J., Seneca S., Bauer P., Stenhouse S.A.R., Barton D.E.. EMQN/CMGS best practice guidelines for the molecular genetic testing of Huntington disease. Eur. J. Hum. Genet. 2013; 21:480–486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55. Gomes-Pereira M., Bidichandani S.I., Monckton D.G.. Kohwi Y. Analysis of unstable triplet repeats using small-pool polymerase chain reaction. Trinucleotide Repeat Protocols. 2004; Totowa, NJ: Humana Press; 61–76. [DOI] [PubMed] [Google Scholar]
- 56. Grosso V., Marcolungo L., Maestri S., Alfano M., Lavezzari D., Iadarola B., Salviati A., Mariotti B., Botta A., D’Apice M.R.et al.. Characterization of FMR1 repeat expansion and intragenic variants by indirect sequence capture. Front. Genet. 2021; 12:743230. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Sanger F., Coulson A.R.. A rapid method for determining sequences in DNA by primed synthesis with DNA polymerase. J. Mol. Biol. 1975; 94:441–448. [DOI] [PubMed] [Google Scholar]
- 58. Keogh M.J., Chinnery P.F.. Next generation sequencing for neurological diseases: new hope or new hype?. Clin. Neurol. Neurosurg. 2013; 115:948–953. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59. Lee J.-M., Kim K.-H., Shin A., Chao M.J., Abu Elneel K., Gillis T., Mysore J.S., Kaye J.A., Zahed H., Kratter I.H.et al.. Sequence-level analysis of the major European Huntington disease haplotype. Am. J. Hum. Genet. 2015; 97:435–444. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60. de Leeuw R.H., Garnier D., Kroon R.M.J.M., Horlings C.G.C., de Meijer E., Buermans H., van Engelen B.G.M., de Knijff P., Raz V.. Diagnostics of short tandem repeat expansion variants using massively parallel sequencing and componential tools. Eur. J. Hum. Genet. 2019; 27:400–407. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61. De Cario R, Kura A., Suraci S., Magi A., Volta A., Marcucci R., Gori A.M., Pepe G., Giusti B., Sticchi E.. Sanger validation of high-throughput sequencing in genetic diagnosis: still the best practice?. 2020; 11:592588. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62. Sanger F., Nicklen S., Coulson A.R.. DNA sequencing with chain-terminating inhibitors. Proc. Natl Acad. Sci. U.S.A. 1977; 74:5463–5467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63. Maestri S., Maturo M.G., Cosentino E., Marcolungo L., Iadarola B., Fortunati E., Rossato M., Delledonne M.. A long-read sequencing approach for direct haplotype phasing in clinical settings. Int. J. Mol. Sci. 2020; 21:9177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64. Monckton D.G., Caskey C.T.. Unstable triplet repeat diseases. Circulation. 1995; 91:513–520. [DOI] [PubMed] [Google Scholar]
- 65. Loomis E.W., Eid J.S., Peluso P., Yin J., Hickey L., Rank D., McCalmon S., Hagerman R.J., Tassone F., Hagerman P.J.. Sequencing the unsequenceable: expanded CGG-repeat alleles of the fragile X gene. Genome Res. 2013; 23:121–128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66. Gettings K.B., Kiesler K.M., Faith S.A., Montano E., Baker C.H., Young B.A., Guerrieri R.A., Vallone P.M.. Sequence variation of 22 autosomal STR loci detected by next generation sequencing. Forensic Sci. Int. Genet. 2016; 21:15–21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67. Riman S., Iyer H., Borsuk L., Gettings K., Vallone P.. Investigating the effects of different library preparation protocols on STR sequencing. Forensic Sci. Int. Genet. Suppl. Ser. 2017; 6:e418–e420. [Google Scholar]
- 68. Facchini S., Dominik N., Manini A., Efthymiou S., Currò R., Rugginini B., Vegezzi E., Quartesan I., Perrone B., Kutty S.K.et al.. Optical genome mapping enables detection and accurate sizing of RFC1 repeat expansions. Biomolecules. 2023; 13:1546. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69. van der Sanden B., Neveling K., Pang A.W.C., Shukor S., Gallagher M.D., Burke S.L., Kamsteeg E.-J., Hastie A., Hoischen A.. Optical genome mapping for applications in repeat expansion disorders. Curr. Protoc. 2024; 4:e1094. [DOI] [PubMed] [Google Scholar]
- 70. Zarouchlioti C., Efthymiou S., Facchini S., Dominik N., Bhattacharyya N., Liu S., Costa M.A., Szabo A., Sadan A.N., Jun A.S.et al.. Tissue-specific TCF4 triplet repeat instability revealed by optical genome mapping. Ebiomedicine. 2024; 108:105328. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71. Eisenstein M. Innovative technologies crowd the short-read sequencing market. Nature. 2023; 614:798–800. [DOI] [PubMed] [Google Scholar]
- 72. Slatko B.E., Gardner A.F., Ausubel F.M.. Overview of next generation sequencing technologies. Curr. Protoc. Mol. Biol. 2018; 122:e59. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73. Ewing B., Hillier L., Wendl M.C., Green P.. Base-calling of automated sequencer traces UsingPhred. I. Accuracy assessment. Genome Res. 1998; 8:175–185. [DOI] [PubMed] [Google Scholar]
- 74. Stoler N., Nekrutenko A.. Sequencing error profiles of Illumina sequencing instruments. NAR Genom. Bioinform. 2021; 3:lqab019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75. Bronner I.F., Quail M.A., Turner D.J., Swerdlow H.. Improved protocols for Illumina sequencing. Curr. Protoc. Hum. Genet. 2009; 18: 10.1002/0471142905.hg1802s62 [DOI] [PubMed] [Google Scholar]
- 76. Iadarola B., Xumerle L., Lavezzari D., Paterno M., Marcolungo L., Beltrami C., Fortunati E., Mei D., Vetro A., Guerrini R.et al.. Shedding light on dark genes: enhanced targeted resequencing by optimizing the combination of enrichment technology and DNA fragment length. Sci. Rep. 2020; 10:9424. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77. Li J.Y., Popovic N., Brundin P.. The use of the R6 transgenic mouse models of Huntington’s disease in attempts to develop novel therapeutic strategies. NeuroRx. 2005; 2:447–464. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78. Menalled L.B., Kudwa A.E., Miller S., Fitzpatrick J., Watson-Johnson J., Keating N., Ruiz M., Mushlin R., Alosio W., McConnell K.et al.. Comprehensive behavioral and molecular characterization of a new knock-in mouse model of Huntington’s disease: zQ175. PLoS One. 2012; 7:e49838. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79. Marx V. Method of the year: long-read sequencing. Nat. Methods. 2023; 20:6–11. [DOI] [PubMed] [Google Scholar]
- 80. Ummat A., Bashir A.. Resolving complex tandem repeats with long reads. Bioinformatics. 2014; 30:3491–3498. [DOI] [PubMed] [Google Scholar]
- 81. Sereika M., Kirkegaard R.H., Karst S.M., Michaelsen T.Y., Sørensen E.A., Wollenberg R.D., Albertsen M.. Oxford Nanopore R10.4 long-read sequencing enables the generation of near-finished bacterial genomes from pure cultures and metagenomes without short-read or reference polishing. Nat. Methods. 2022; 19:823–826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82. Maestri S., Gambino G., Lopatriello G., Minio A., Perrone I., Cosentino E., Giovannone B., Marcolungo L., Alfano M., Rombauts S.et al.. Nebbiolo’ genome assembly allows surveying the occurrence and functional implications of genomic structural variations in grapevines (Vitis viniferaL.). BMC Genomics. 2022; 23:159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83. Darian J.C., Kundu R., Rajaby R., Sung W.-K.. Constructing telomere-to-telomere diploid genome by polishing haploid nanopore-based assembly. Nat. Methods. 2024; 21:574–583. [DOI] [PubMed] [Google Scholar]
- 84. Rhoads A., Au K.F.. PacBio sequencing and its applications. Genomics Proteomics Bioinformatics. 2015; 13:278–289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85. Pourmohammadi R., Abouei J., Anpalagan A.. Error analysis of the PacBio sequencing CCS reads. Int. J. Biostat. 2023; 19:439–453. [DOI] [PubMed] [Google Scholar]
- 86. Wenger A.M., Peluso P., Rowell W.J., Chang P.-C., Hall R.J., Concepcion G.T., Ebler J., Fungtammasan A., Kolesnikov A., Olson N.D.et al.. Accurate circular consensus long-read sequencing improves variant detection and assembly of a human genome. Nat. Biotechnol. 2019; 37:1155–1162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87. Hon T., Mars K., Young G., Tsai Y.-C., Karalius J.W., Landolin J.M., Maurer N., Kudrna D., Hardigan M.A., Steiner C.C.et al.. Highly accurate long-read HiFi sequencing data for five complex genomes. Sci. Data. 2020; 7:399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88. Kovaka S., Fan Y., Ni B., Timp W., Schatz M.C.. Targeted nanopore sequencing by real-time mapping of raw electrical signal with UNCALLED. Nat. Biotechnol. 2021; 39:431–441. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89. Wang Y., Zhao Y., Bollas A., Wang Y., Au K.F.. Nanopore sequencing technology, bioinformatics and applications. Nat. Biotechnol. 2021; 39:1348–1365. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90. Zhang H., Li H., Jain C., Cheng H., Au K.F., Li H., Aluru S.. Real-time mapping of nanopore raw signals. Bioinformatics. 2021; 37:i477–i483. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91. Ni Y., Liu X., Simeneh Z.M., Yang M., Li R.. Benchmarking of Nanopore R10.4 and R9.4.1 flow cells in single-cell whole-genome amplification and whole-genome shotgun sequencing. Comput. Struct. Biotechnol. J. 2023; 21:2352–2364. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92. Doi K., Monjo T., Hoang P.H., Yoshimura J., Yurino H., Mitsui J., Ishiura H., Takahashi Y., Ichikawa Y., Goto J.et al.. Rapid detection of expanded short tandem repeats in personal genomics using hybrid sequencing. Bioinformatics. 2014; 30:815–822. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93. Landrian I., McFarland K.N., Liu J., Mulligan C.J., Rasmussen A., Ashizawa T.. Inheritance patterns of ATCCT repeat interruptions in spinocerebellar ataxia type 10 (SCA10) expansions. PLoS One. 2017; 12:e0175958. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94. Cumming S.A., Hamilton M.J., Robb Y., Gregory H., McWilliam C., Cooper A., Adam B., McGhie J., Hamilton G., Herzyk P.et al.. De novorepeat interruptions are associated with reduced somatic instability and mild or absent clinical features in myotonic dystrophy type 1. Eur. J. Hum. Genet. 2018; 26:1635–1647. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95. Mangin A., de Pontual L., Tsai Y.-C., Monteil L., Nizon M., Boisseau P., Mercier S., Ziegle J., Harting J., Heiner C.et al.. Robust detection of somatic mosaicism and repeat interruptions by long-read targeted sequencing in myotonic dystrophy type 1. Int. J. Mol. Sci. 2021; 22:2616. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96. Fang L., Monteys A.M., Dürr A., Keiser M., Cheng C., Harapanahalli A., Gonzalez-Alegre P., Davidson B.L., Wang K.. Haplotyping SNPs for allele-specific gene editing of the expanded huntingtin allele using long-read sequencing. HGG Adv. 2022; 4:100146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97. Woerner A.E., Mandape S., King J.L., Muenzler M., Crysup B., Budowle B.. Reducing noise and stutter in short tandem repeat loci with unique molecular identifiers. Forensic Sci. Int. Genet. 2021; 51:102459. [DOI] [PubMed] [Google Scholar]
- 98. Karst S.M., Ziels R.M., Kirkegaard R.H., Sørensen E.A., McDonald D., Zhu Q., Knight R., Albertsen M.. High-accuracy long-read amplicon sequences using unique molecular identifiers with Nanopore or PacBio sequencing. Nat. Methods. 2021; 18:165–169. [DOI] [PubMed] [Google Scholar]
- 99. Lebrigand K., Magnone V., Barbry P., Waldmann R.. High throughput error corrected Nanopore single cell transcriptome sequencing. Nat. Commun. 2020; 11:4025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100. Amstler S., Forer L., Streiter G., Maio S.D., Paulweber B., Kronenberg F., Schönherr S., Coassin S.. Nanopore sequencing with unique molecular identifiers preserves SNP haplotypes of the LPA KIV-2 copy number variation. Atherosclerosis. 2023; 379:S47 [Google Scholar]
- 101. Ivančić D., Mir-Pedrol J., Jaraba-Wallace J., Rafel N., Sanchez-Mejias A., Güell M.. INSERT-seq enables high-resolution mapping of genomically integrated DNA using Nanopore sequencing. Genome Biol. 2022; 23:227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102. Zurek P.J., Knyphausen P., Neufeld K., Pushpanath A., Hollfelder F.. UMI-linked consensus sequencing enables phylogenetic analysis of directed evolution. Nat. Commun. 2020; 11:6023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103. Bastepe M., Xin W.. Huntington disease: molecular diagnostics approach. Curr. Protoc. Hum. Genet. 2015; 87:9.26.1–9.26.23. [DOI] [PubMed] [Google Scholar]
- 104. Sone J., Mitsuhashi S., Fujita A., Mizuguchi T., Hamanaka K., Mori K., Koike H., Hashiguchi A., Takashima H., Sugiyama H.et al.. Long-read sequencing identifies GGC repeat expansions in NOTCH2NLC associated with neuronal intranuclear inclusion disease. Nat. Genet. 2019; 51:1215–1221. [DOI] [PubMed] [Google Scholar]
- 105. De Roeck A., De Coster W., Bossaerts L., Cacace R., De Pooter T., Van Dongen J., D’Hert S., De Rijk P., Strazisar M., Van Broeckhoven C.et al.. NanoSatellite: accurate characterization of expanded tandem repeat length and sequence through whole genome long-read sequencing on PromethION. Genome Biol. 2019; 20:239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106. Dolzhenko E., Deshpande V., Schlesinger F., Krusche P., Petrovski R., Chen S., Emig-Agius D., Gross A., Narzisi G., Bowman B.et al.. ExpansionHunter: a sequence-graph-based tool to analyze variation in short tandem repeat regions. Bioinformatics. 2019; 35:4754–4756. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107. Tsai Y.-C., Greenberg D., Powell J., Höijer I., Ameur A., Strahl M., Ellis E., Jonasson I., Pinto R.M., Wheeler V.C.et al.. Amplification-free, CRISPR-Cas9 targeted enrichment and SMRT sequencing of repeat-expansion disease causative genomic regions. 2017; bioRxiv doi:16 October 2017,preprint: not peer reviewed 10.1101/203919 [DOI]
- 108. Hafford-Tear N.J., Tsai Y.-C., Sadan A.N., Sanchez-Pintado B., Zarouchlioti C., Maher G.J., Liskova P., Tuft S.J., Hardcastle A.J., Clark T.A.et al.. CRISPR/Cas9-targeted enrichment and long-read sequencing of the Fuchs endothelial corneal dystrophy–associated TCF4 triplet repeat. Genet. Med. 2019; 21:2092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109. López-Girona E., Davy M.W., Albert N.W., Hilario E., Smart M.E.M., Kirk C., Thomson S.J., Chagné D.. CRISPR-Cas9 enrichment and long read sequencing for fine mapping in plants. Plant Methods. 2020; 16:121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110. Lopatriello G., Maestri S., Alfano M., Papa R., Di Vittori V., De Antoni L., Bellucci E., Pieri A., Bitocchi E., Delledonne M.et al.. CRISPR/Cas9-mediated enrichment coupled to nanopore sequencing provides a valuable tool for the precise reconstruction of large genomic target regions. Int. J. Mol. Sci. 2023; 24:1076. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111. Schüle B., McFarland K.N., Lee K., Tsai Y.-C., Nguyen K.-D., Sun C., Liu M., Byrne C., Gopi R., Huang N.et al.. Parkinson’s disease associated with pure ATXN10 repeat expansion. NPJ Parkinsons Dis. 2017; 3:27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112. Ebbert M.T.W., Farrugia S.L., Sens J.P., Jansen-West K., Gendron T.F., Prudencio M., McLaughlin I.J., Bowman B., Seetin M., DeJesus-Hernandez M.et al.. Long-read sequencing across the C9orf72 ‘GGGGCC’ repeat expansion: implications for clinical use and genetic discovery efforts in human disease. Mol. Neurodegener. 2018; 13:46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113. Höijer I., Tsai Y., Clark T.A., Kotturi P., Dahl N., Stattin E., Bondeson M., Feuk L., Gyllensten U., Ameur A.. Detailed analysis of HTT repeat elements in human blood using targeted amplification-free long-read sequencing. Hum. Mutat. 2018; 39:1262–1272. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114. DeJesus-Hernandez M., Aleff R.A., Jackson J.L., Finch N.A., Baker M.C., Gendron T.F., Murray M.E., McLaughlin I.J., Harting J.R., Graff-Radford N.R.et al.. Long-read targeted sequencing uncovers clinicopathological associations for C9orf72-linked diseases. Brain. 2021; 144:1082–1088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115. Giesselmann P., Brändl B., Raimondeau E., Bowen R., Rohrandt C., Tandon R., Kretzmer H., Assum G., Galonska C., Siebert R.et al.. Analysis of short tandem repeat expansions and their methylation state with nanopore sequencing. Nat. Biotechnol. 2019; 37:1478–1481. [DOI] [PubMed] [Google Scholar]
- 116. Gilpatrick T., Lee I., Graham J.E., Raimondeau E., Bowen R., Heron A., Downs B., Sukmar S., Sedlazeck F.J., Timp W.. Targeted nanopore sequencing with Cas9-guided adaptor ligation. Nat. Biotechnol. 2020; 38:433–438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117. Mizuguchi T., Toyota T., Miyatake S., Mitsuhashi S., Doi H., Kudo Y., Kishida H., Hayashi N., Tsuburaya R.S., Kinoshita M.et al.. Complete sequencing of expanded SAMD12 repeats by long-read sequencing and Cas9-mediated enrichment. Brain. 2021; 144:1103–1117. [DOI] [PubMed] [Google Scholar]
- 118. Wallace A.D., Sasani T.A., Swanier J., Gates B.L., Greenland J., Pedersen B.S., Varley K.E., Quinlan A.R.. CaBagE: a Cas9-based background elimination strategy for targeted, long-read DNA sequencing. PLoS One. 2021; 16:e0241253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119. Loose M., Malla S., Stout M.. Real-time selective sequencing using nanopore technology. Nat. Methods. 2016; 13:751–754. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120. Edwards H.S., Krishnakumar R., Sinha A., Bird S.W., Patel K.D., Bartsch M.S.. Real-time selective sequencing with RUBRIC: read Until with basecall and reference-informed criteria. Sci. Rep. 2019; 9:11475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121. Payne A., Holmes N., Clarke T., Munro R., Debebe B.J., Loose M.. Readfish enables targeted nanopore sequencing of gigabase-sized genomes. Nat. Biotechnol. 2021; 39:442–450. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122. Stevanovski I., Chintalaphani S.R., Gamaarachchi H., Ferguson J.M., Pineda S.S., Scriba C.K., Tchan M., Fung V., Ng K., Cortese A.et al.. Comprehensive genetic diagnosis of tandem repeat expansion disorders with programmable targeted nanopore sequencing. Sci. Adv. 2022; 8:eabm5386. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123. Miyatake S., Koshimizu E., Fujita A., Doi H., Okubo M., Wada T., Hamanaka K., Ueda N., Kishida H., Minase G.et al.. Rapid and comprehensive diagnostic method for repeat expansion diseases using nanopore sequencing. NPJ Genom. Med. 2022; 7:62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124. Weilguny L., De Maio N., Munro R., Manser C., Birney E., Loose M., Goldman N.. Dynamic, adaptive sampling during nanopore sequencing using Bayesian experimental design. Nat. Biotechnol. 2023; 41:1018–1025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125. Chen Z., Gustavsson E.K., Macpherson H., Anderson C., Clarkson C., Rocca C., Self E., Alvarez Jerez P., Scardamaglia A., Pellerin D.et al.. Adaptive long-read sequencing reveals GGC repeat expansion in ZFHX3 associated with spinocerebellar ataxia type 4. Mov. Disord. 2024; 39:486–497. [DOI] [PubMed] [Google Scholar]
- 126. McAllister B., Donaldson J., Binda C.S., Powell S., Chughtai U., Edwards G., Stone J., Lobanov S., Elliston L., Schuhmacher L.-N.et al.. Exome sequencing of individuals with Huntington’s disease implicates FAN1 nuclease activity in slowing CAG expansion and disease onset. Nat. Neurosci. 2022; 25:446–457. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127. Fang L., Liu Q., Monteys A.M., Gonzalez-Alegre P., Davidson B.L., Wang K.. DeepRepeat: direct quantification of short tandem repeats on signal data from nanopore sequencing. Genome Biol. 2022; 23:108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128. Taylor A.S., Barros D., Gobet N., Schuepbach T., McAllister B., Aeschbach L., Randall E.L., Trofimenko E., Heuchan E.R., Barszcz P.et al.. Repeat Detector: versatile sizing of expanded tandem repeats and identification of interrupted alleles from targeted DNA sequencing. NAR Genom. Bioinform. 2022; 4:lqac089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129. Chiara M., Zambelli F., Picardi E., Horner D.S., Pesole G.. Critical assessment of bioinformatics methods for the characterization of pathological repeat expansions with single-molecule sequencing data. Brief. Bioinform. 2020; 21:1971–1986. [DOI] [PubMed] [Google Scholar]
- 130. Mousavi N., Shleizer-Burko S., Yanicky R., Gymrek M.. Profiling the genome-wide landscape of tandem repeat expansions. Nucleic Acids Res. 2019; 47:e90. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131. Gymrek M., Golan D., Rosset S., Erlich Y.. lobSTR: a short tandem repeat profiler for personal genomes. Genome Res. 2012; 22:1154–1162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132. Willems T., Zielinski D., Yuan J., Gordon A., Gymrek M., Erlich Y.. Genome-wide profiling of heritable and de novo STR variations. Nat. Methods. 2017; 14:590–592. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133. Tang H., Kirkness E.F., Lippert C., Biggs W.H., Fabani M., Guzman E., Ramakrishnan S., Lavrenko V., Kakaradov B., Hou C.et al.. Profiling of short-tandem-repeat disease alleles in 12,632 human whole genomes. Am. J. Hum. Genet. 2017; 101:700–715. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134. Dashnow H., Lek M., Phipson B., Halman A., Sadedin S., Lonsdale A., Davis M., Lamont P., Clayton J.S., Laing N.G.et al.. STRetch: detecting and discovering pathogenic short tandem repeat expansions. Genome Biol. 2018; 19:121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135. Tankard R.M., Bennett M.F., Degorski P., Delatycki M.B., Lockhart P.J., Bahlo M.. Detecting expansions of tandem repeats in cohorts sequenced with short-read sequencing data. Am. J. Hum. Genet. 2018; 103:858–873. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136. Dashnow H., Pedersen B.S., Hiatt L., Brown J., Beecroft S.J., Ravenscroft G., LaCroix A.J., Lamont P., Roxburgh R.H., Rodrigues M.J.et al.. STRling: a k-mer counting approach that detects short tandem repeat expansions at known and novel loci. Genome Biol. 2022; 23:257. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 137. Liu Q., Zhang P., Wang D., Gu W., Wang K.. Interrogating the “unsequenceable” genomic trinucleotide repeat disorders by long-read sequencing. Genome Med. 2017; 9:65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138. Mitsuhashi S., Frith M.C., Mizuguchi T., Miyatake S., Toyota T., Adachi H., Oma Y., Kino Y., Mitsuhashi H., Matsumoto N.. Tandem-genotypes: robust detection of tandem repeat expansions from long DNA reads. Genome Biol. 2019; 20:58. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139. Harris R.S., Cechova M., Makova K.D.. Noise-cancelling repeat finder: uncovering tandem repeats in error-prone long-read sequencing data. Bioinformatics. 2019; 35:4809–4811. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140. Bolognini D., Magi A., Benes V., Korbel J.O., Rausch T.. TRiCoLOR: tandem repeat profiling using whole-genome long-read sequencing data. GigaScience. 2020; 9:giaa101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141. Chiu R., Rajan-Babu I.-S., Friedman J.M., Birol I.. Straglr: discovering and genotyping tandem repeat expansions using whole genome long-read sequences. Genome Biol. 2021; 22:224. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 142. Frith M.C., Mitsuhashi S., Katoh K.. lamassemble: multiple alignment and consensus sequence of long reads. Methods Mol. Biol. 2021; 2231:135–145. [DOI] [PubMed] [Google Scholar]
- 143. Lang J., Xu Z., Wang Y., Sun J., Yang Z.. NanoSTR: a method for detection of target short tandem repeats based on nanopore sequencing data. Front. Mol. Biosci. 2023; 10:1093519. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 144. Deynze K.V., Mumm C., Maltby C.J., Switzenberg J.A., Todd P.K., Boyle A.P.. Enhanced detection and genotyping of disease-associated tandem repeats using HMMSTR and targeted long-read sequencing. 2024; medRxiv doi:03 May 2024, preprint: not peer reviewed 10.1101/2024.05.01.24306681 [DOI] [PMC free article] [PubMed]
- 145. Benson G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res. 1999; 27:573–580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146. Dolzhenko E., English A., Dashnow H., De Sena Brandine G., Mokveld T., Rowell W.J., Karniski C., Kronenberg Z., Danzi M.C., Cheung W.A.et al.. Characterization and visualization of tandem repeats at genome scale. Nat. Biotechnol. 2024; 42:1606–1614. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 147. Tan K.-T., Slevin M.K., Meyerson M., Li H.. Identifying and correcting repeat-calling errors in nanopore sequencing of telomeres. Genome Biol. 2022; 23:180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148. Sitarčík J., Vinař T., Brejová B., Krampl W., Budiš J., Radvánszky J., Lucká M.. WarpSTR: determining tandem repeat lengths using raw nanopore signals. Bioinformatics. 2023; 39:btad388. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149. Nakamori M., Panigrahi G.B., Lanni S., Gall-Duncan T., Hayakawa H., Tanaka H., Luo J., Otabe T., Li J., Sakata A.et al.. A slipped-CAG DNA-binding small molecule induces trinucleotide-repeat contractions in vivo. Nat. Genet. 2020; 52:146–159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 150. Sanchez-Flores M., Corral-Juan M., Gasch-Navalón E., Cirillo D., Sanchez I., Matilla-Dueñas A.. Novel genotype–phenotype correlations, differential cerebellar allele-specific methylation, and a common origin of the (ATTTC)n insertion in spinocerebellar ataxia type 37. Hum. Genet. 2024; 143:211–232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 151. Robinson J.T., Thorvaldsdóttir H., Winckler W., Guttman M., Lander E.S., Getz G., Mesirov J.P.. Integrative genomics viewer. Nat. Biotechnol. 2011; 29:24–26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 152. Oehler J.B., Wright H., Stark Z., Mallett A.J., Schmitz U.. The application of long-read sequencing in clinical settings. Hum. Genomics. 2023; 17:73. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 153. Maestri S., Cosentino E., Paterno M., Freitag H., Garces J.M., Marcolungo L., Alfano M., Njunjić I., Schilthuizen M., Slik F.et al.. A rapid and accurate MinION-based workflow for tracking species biodiversity in the field. Genes (Basel). 2019; 10:468. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 154. Tarquini G., Martini M., Maestri S., Firrao G., Ermacora P.. The virome of ‘Lamon Bean’: application of MinION sequencing to investigate the virus population associated with symptomatic beans in the Lamon area, Italy. Plants. 2022; 11:779. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 155. Kolmogorov M., Yuan J., Lin Y., Pevzner P.A.. Assembly of long, error-prone reads using repeat graphs. Nat. Biotechnol. 2019; 37:540–546. [DOI] [PubMed] [Google Scholar]
- 156. Koren S., Walenz B.P., Berlin K., Miller J.R., Bergman N.H., Phillippy A.M.. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 2017; 27:722–736. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 157. Elena C., Scalzo D., Iennaco R., Camilla M., Zobel M., Besusso D., Maestri S.. When repetita no-longer iuvant: somatic instability of the CAG triplet in Huntington's disease. Nucleic Acids Res. 2024; 10.1093/nar/gkae1204 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
No new data were generated or analyzed in support of this research.