Abstract
Cellular senescence is an aging mechanism characterized by cell cycle arrest and a senescence-associated secretory phenotype (SASP). Preclinical studies demonstrate that senolytic drugs, which target survival pathways in senescent cells, can counteract age-associated conditions that span several organs. The comparative efficacy of distinct senolytic drugs for modifying aging and senescence biomarkers in vivo has not been demonstrated. Here, we established aging- and senescence-related plasma proteins and tissue transcripts that changed in old versus young female and male mice. We investigated responsivity to acute treatment with venetoclax, navitoclax, fisetin or luteolin versus transgenic senescent cell clearance in aged p16-InkAttac mice. We discovered that age-dependent changes in plasma proteins, including IL-23R, CCL5 and CA13, were reversed by senotherapeutics, which corresponded to expression differences in tissues, particularly in the kidney. In plasma from humans across the lifespan, IL-23R increased with age. Our results reveal circulating factors as candidate mediators of senescence-associated interorgan signal transduction and translationally impactful biomarkers of systemic senescent cell burden.
Subject terms: Ageing, Transcription, Biomarkers
Using mouse and human plasma, Carver et al. identify factors that are altered with age and test which are reverted by a panel of genetic and pharmacological senolytic interventions in aged mice. They identify IL-23R as a senescence-associated, age-increased circulating biomarker.
Main
Cellular senescence is an age-associated mechanism that contributes to pathogenesis in diverse organ systems, including immune dysfunction1,2, cardiovascular and metabolic diseases3–5, pulmonary fibrosis6, osteoporosis7, kidney disease8, hepatic damage9, sarcopenia10 and cognitive decline11,12. Factors in aged blood can drive senescent and inflammatory activation in numerous tissues13,14. The tissue- and cell-specific origins of progeronic circulating factors are not well characterized. Circulating proteins that originate from senescent cells may reflect systemic senescent cell burden, may be exploited as translational diagnostic or prognostic biomarkers, and are possible mediators of progeronic interorgan communication.
Characteristic properties of senescent cells include upregulation of cell cycle regulatory proteins, including p16ink4a (encoded by Cdkn2a)15, production and secretion of a SASP16,17 and activation of senescent cell anti-apoptotic pathways (SCAPs)18. The SASP is cell-type- and context-specific19,20 and can confer adverse changes to local tissue environments3,6,21 and systemic organs22,23. Use of the p16-InkAttac transgenic model, which permits systemic clearance of p16-positive cells through a temporally controlled suicide gene, has demonstrated that senescent cell deletion alleviates features of age-related pathology in several organs, including kidney, adipose, skeletal muscle, eye, heart and brain4,11,12,24–27. A recent comparison of cell-type-specific versus whole-body transgenic targeting of p16-postitive cells demonstrated greater benefits in aged bone composition following systemic clearance, which supports the notion that the adverse influence of senescent cells and the SASP results from both local and distant signaling28.
Senolytic compounds selectively target and deactivate SCAPs to enable senescent cell apoptosis18,29. Two classes of SCAP-directed senolytics include BCL-family inhibitors and natural-product flavonoids. Venetoclax (ABT-199; VEN) and navitoclax (ABT-263; NAV) are BH3 mimetic/BCL-family inhibitors with differential selectivity and specificity to BCL2, BCL-XL and/or BCL-W30,31. Fisetin (FIS) and luteolin (LUT) are structurally similar flavonoids that reduce markers of senescence in aging mice, confer anti-inflammatory properties and mitigate age-related pathologies32–34. Despite considerable senolytic testing underway in preclinical models and humans, understanding of the comparative effects of senolytic drugs and senescent cell targeting efficiency across tissues is limited.
The central goal of this study was to identify age- and senescence-related plasma and tissue biomarkers that are responsive to senotherapeutic intervention. We assessed system-wide profiles of senescence, SASP and inflammatory biomarkers in aging and their alteration by clinically relevant senolytic compounds versus transgenic p16-InkAttac senescent cell targeting in mice. We discovered that the abundance of IL-23R, CCL5 and other proteins showing age-dependent increases in circulation were reduced by senotherapeutic agents. CA13 decreased in aged plasma, and senolytics restored this factor toward youthful levels. In secretory tissues, gene expression of Il23r and Ccl5 coincided with expression of senescence markers and aged plasma protein abundance in vivo, and these factors were significantly upregulated and secreted by senescent cells in vitro. Our results suggest that senescent cells in aged kidney, liver and spleen are viable sources of these aging biomarkers in blood circulation. Among the drugs tested, VEN suppressed age-related changes in the greatest number of circulating and tissue biomarkers in aged mice. In human plasma, we discovered that IL-23R abundance increased with age in both women and men. Overall, this study identifies candidate progeronic and rejuvenative aging biomarkers as indicators of systemic senescence burden and possible mediators of senescence-regulated interorgan signaling that can be controlled by senolytic intervention in vivo.
Results
Circulating plasma proteins change in abundance with aging
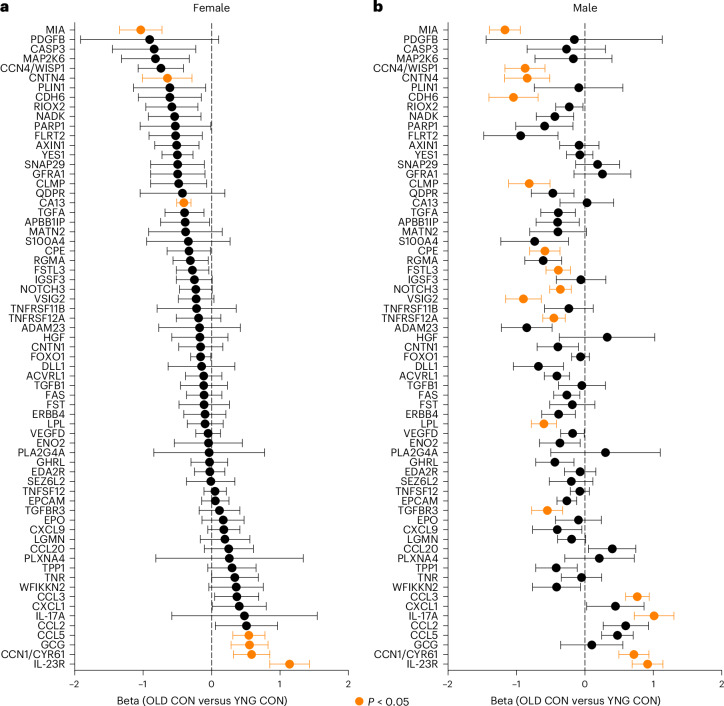
We first sought to identify circulating biomarkers that were altered by aging between young (3–4 months) and old (16–20 months) mice using Olink proximity extension assays (PEAs). The protein panel included SASP, inflammatory, metabolic, developmental, cell scaffolding and signaling factors (Fig. 1). The utility of this approach was to probe both senescence-related and generalized factors of systemic health in aging. Relative to young females, four proteins were higher (CCL5, CCN1, IL-23R, GCG) and three proteins were lower (CA13, CNTN4, MIA) in old female plasma (Fig. 1a). Relative to young males, 4 proteins were higher (CCL3, CCN1, IL-17A, IL-23R) and 12 proteins were lower (CCN4, CDH6, CNTN4, CLMP, CPE, FSTL3, LPL, MIA, NOTCH3, TGFBR3, TNFRSF12A, VSIG2) in old male plasma (Fig. 1b). Across both sexes, IL-23R and CCN1 were higher and CNTN4 and MIA were lower in aged plasma. We next sought to measure the transcriptional expression of these and other aging biomarkers in tissues.
Fig. 1. Conserved and distinct plasma proteins are altered by age in female and male mice.
a,b, Beta estimates corresponding to comparison of old control (OLD CON) versus young control (YNG CON) protein abundance determined by Olink PEAs in female (a) and male (b) plasma samples. Statistically significant differences between OLD CON and YNG CON are highlighted in orange (P < 0.05, two-sided Kruskal–Wallis rank sum test), with negative values indicating a decrease in protein levels in aging, and positive values indicating an increase in protein abundance in aging samples. Datapoints summarize the mean beta estimate and bars summarize the ± standard error. Sample sizes: YNG CON female, n = 6; YNG CON male, n = 5; OLD CON female, n = 6; OLD CON male, n = 8.
Senescence and aged plasma genes increase in aged tissues
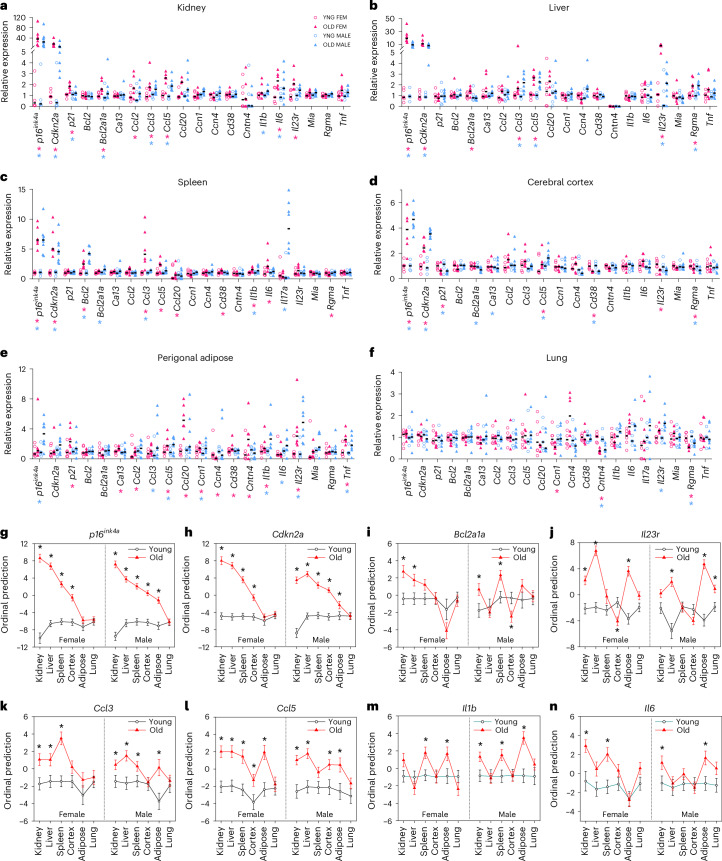
We hypothesized that proteins that increase in aged plasma may originate from senescent cells. Therefore, we measured transcript expression of genes encoding age-increased plasma factors alongside canonical senescence markers across tissues in old versus young p16-InkAttac mice. To support dynamic interpretation across tissues, we demonstrate the real-time PCR (RT-PCR) gene expression data in individual sample cycle threshold (CT) (Extended Data Fig. 1), relative expression (Fig. 2a–f), ordinal regression (Fig. 2g–n) and normalized relative expression heatmap (Supplementary Fig. 2) plots. Compared with young mice, we detected greater p16ink4a expression in old mouse kidney (Fig. 2a), liver (Fig. 2b), spleen (Fig. 2c) and brain cerebral cortex (Fig. 2d) of both sexes. p16ink4a was higher in perigonadal adipose of old male mice (Fig. 2e). Expression of the broader Cdkn2a locus detecting both p19arf and p16ink4a closely tracked with expression of p16ink4a per organ (Fig. 2a–h). In examination of relative CT values, we observed that aged spleen harbored the greatest abundance of p16 transcript across young and old tissues (Extended Data Fig. 1). However, ordinal prediction models demonstrate the largest age-associated change in p16ink4a and Cdkn2a relative expression in kidney, followed by liver and then spleen (Fig. 2g,h). p21 expression was higher in aged kidney in both sexes and in female adipose but lower in aged cortex (Fig. 2a,d,e). The SCAP gene Bcl2 increased in old male and female spleen (Fig. 2c). Moreover, Bcl2a1a, a downstream effector and cytokine-stimulated mediator of the BCL2 cell survival pathway35,36, was higher in aged kidney in both sexes, liver in females and spleen in males (Fig. 2i). We observed higher expression of Il23r in old female kidney, liver and adipose, and in old male liver, adipose and lung (Fig. 2a,b,e,f,j). Expression of SASP chemokines Ccl3 (Fig. 2k) and Ccl5 (Fig. 2l), Il1b (Fig. 2m) and Il6 (Fig. 2n) were systemically increased across several aged tissues, including kidney, liver, spleen, cortex and adipose, with Ccl5 increased more consistently relative to other analyzed chemokines.
Extended Data Fig. 1. Cycle threshold values from RT-PCR gene expression in young (Y) and old (O) tissues from female (pink) and male (blue) mice.
The maximum number of cycles for each experiment was 40. n = 11 Y KID, 14 O KID, 11 Y LIV, 14 O LIV, 11 Y SPLN, 14 O SPLN, 11 Y LNG, 14 O LNG, 11 Y FAT, 14 O FAT, 11 Y CTX, 14 O CTX. Bars summarize the mean ± s.e.m.
Fig. 2. Senescence- and age-associated genes are expressed across tissues in old female and male p16-InkAttac mice.
a–f, RT-PCR gene expression values from young female, old female, young male and old male tissues for kidney (a), liver (b), spleen (c), cerebral cortex (d), perigonadal adipose (e) and lung (f). Values are normalized relative to the expression of young control mice of each sex per tissue; black bars indicate median values for each group. g–n, Shown are ordinal logistic regression models comparing gene expression from young and old kidney, liver, spleen, cortex, adipose and lung for p16ink4a (g), Cdkn2a (h), Bcl2a1a (i), Il23r (j), Ccl3 (k), Ccl5 (l), Il1b (m) and Il6 (n). Symbols denote the mean ordinal prediction beta estimate ± standard error. *P < 0.05 significance between old and young mice of same sex by two-sided ordinal logistic regression. Samples sizes: young female, n = 6; young male, n = 5; old female, n = 6; old male, n = 8.
We next applied Spearman’s correlations to determine whether plasma protein levels were associated with tissue-specific transcript abundance in young and old mice (Extended Data Fig. 2). We discovered several age-altered plasma proteins that correlated positively with their corresponding transcripts. In females, Il23r expression in several tissues correlated positively with protein abundance in plasma (kidney r = 0.78, P = 0.004; liver r = 0.87, P < 0.001; adipose r = 0.70, P = 0.014; lung r = 0.74, P = 0.008). Male liver Ccl5 correlated positively with CCL5 plasma protein (r = 0.56, P = 0.047). We also assessed associations between age-altered plasma proteins and expression of canonical senescence markers. The plasma IL-23R protein correlated with Cdkn2a and p16ink4a transcript abundance across tissues (Extended Data Fig. 2), with the greatest positive association observed between plasma IL-23R and kidney Cdkn2a expression in female mice (r = 0.83, P = 0.002; Extended Data Fig. 2a). The kidney was characterized by consistent age-elevated senescence gene signatures that correlated with circulating proteins (Fig. 2); p16ink4a, Cdkn2a, Bcl2a1a, Ccl2, Ccl3, Ccl5, Il1b, Il6 and Il23r, and/or Tnf gene expression associated positively with plasma IL-23R, CCL2, CCL3, CCL5 and/or IL-17A and negatively with MIA (Extended Data Fig. 2). We discovered that IL-23R and/or CCL5 plasma protein also increased concomitant with expression of p16, SASP and SCAP factors in kidney, liver, spleen and adipose (Extended Data Fig. 2).
Extended Data Fig. 2. Correlation between tissue gene expression and plasma protein senescence in young and old mice.
Spearman correlations between tissue gene expression of senescence factors in young and old (A, B) kidney, (C, D) liver, (E, F) spleen, and (G, H) perigonadal adipose, and the plasma proteins CA13, CCL2, CCL3, CCL5, CCN1, IL17A, IL23R, and MIA for (A, C, E, G) female and (B, D, F, H) male mice. R-values are provided, and p values can be found in the source data. Samples sizes of per group: young female n = 6; old female n = 6; young male n = 5; old male n = 8.
Since Cdkn2a/p16 and Il23r (established and emerging aging biomarkers, respectively) both increased in aged kidney, we performed fluorescent in situ hybridization to localize and validate their expression in aged kidney (Extended Data Fig. 3). p16ink4a expression was abundant throughout the nephrons and interstitium. Moreover, Il23r was highly concentrated in cells found in the renal sinus cavity and perivascular tissue, in which p16ink4a was colocalized. Furthermore, we observed that Il23r and p16ink4a colocalized with Cd3e+ cells in immune nodes throughout the aged kidney, present in glomeruli and pervasive in the mesenchymal tissue, whereas young kidney lacked the presence of Il23r (Extended Data Fig. 3b–j).
Extended Data Fig. 3. In situ hybridization (RNAscope) of Il23r+ cells co-localized to p16ink4a and Cd3e in the renal sinus and perivascular areas of aged mouse kidney.
(A) Representative images of p16ink4a (green) and Il23r (red) in C57BL/6 aged kidney perivascular tissue (21 mo. old). Blue color is an open channel for contrast of tissue architecture. (B) RNAscope of p16ink4a (green), Il23r (red), and Cd3e (cyan) from young (top, 2 mo.) and old (bottom, 24 mo.) kidney tissue from mice on p16-InkAttac background. DAPI nuclear stain is depicted in dark blue. Old kidney exhibits dense and perivascular Cd3e + Il23r+ immune nodes. (C–F) Young and (G–J) old kidney tissue from RNAscope, similar to that shown in B. (C, D, G, H) Interstitial and perivascular tissue exhibit co-localization of Cd3e, Il23r, and p16ink4a in old but not young kidney. (E, I) Aged glomeruli show the presence of Cd3e + Il23r+ cells. (F, J) Connective tissue shows dense accumulation of Cd3e+ cells in both young and old kidney, but only old tissue exhibits co-localization with Il23r and p16ink4a. All scale bars = 100 μm. Experiments were repeated in 2 biological replicates of young and 4 replicates of old mice.
Senescent cells express Il23r and Ccl5
To definitively determine whether IL-23R or CCL5 can originate from senescent cells, we implemented an established cell culture model of irradiation (IR)-induced senescence in mouse embryonic fibroblasts (MEFs) (Extended Data Fig. 4). We confirmed IR-exposed MEFs were senescent through senescence-associated β-galactosidase (SA-β-gal) staining and nuclear morphology assessments, RT-PCR gene expression profiling, and multiplexed immunoassay analysis of the SASP in conditioned medium. IR induced a greater percentage (Extended Data Fig. 4b) and total number (Extended Data Fig. 4c) of SA-β-gal+ cells compared with sham controls. Cells from IR cultures were characterized by larger nucleus area (Extended Data Fig. 4d). For gene expression profiling, we examined established senescence and SASP genes, as well as genes encoding proteins that displayed age-associated changes in plasma (Extended Data Fig. 4e). Relative to sham, p16ink4a, Cdkn2a, p21, Gdf15, Il6 and Mmp3 expression increased in IR MEFs. Coinciding with this senescence signature, Il23r expression was 2.4 ± 0.3-fold greater, and Ccl5 expression was 6.3 ± 0.5-fold greater in IR MEFs, compared with sham cultures. Mia also increased in IR MEFs, but Ccn1 was lower than sham. We next measured the abundance of canonical SASP proteins and emerging biomarkers secreted from IR senescent and control cells in conditioned media using multiplexed Luminex and Olink immunoassays (Extended Data Fig. 4f,g). Across the two assays, CCL2, CCL3, CCL5, CCL11, CCL20, CXCL1, CXCL9, GDF15, IL-23R, MMP2, MMP12, OPN, TNFSF12 and TNFRSF1A proteins increased significantly in IR MEF conditioned media. Collectively, Ccl5/CCL5 and Il23r/IL-23R expression are induced in and secreted from senescent cells, which suggests that elevated CCL5 and IL-23R in aged blood circulation could originate from senescent cells.
Extended Data Fig. 4. Senescence- and age-associated genes, including Il23r, are expressed in senescent fibroblasts in vitro.
(A) Representative 10x images of senescence-associated β-galactosidase (SA-β-gal) staining in sham and irradiated (IR) mouse embryonic fibroblasts (MEFs). Left panels are bright-field images, right panels are DAPI fluorescent images of the same field of view. Experiments were repeated in (B, C) Shown are quantifications for (B) percentage and (C) total number of SA-β-gal cells per image field. (D) Quantification of nucleus area in sham and irradiated cells as depicted in A. For B-D, bars denote mean ± s.e.m., p values provided vs. sham by two-tailed Mann-Whitney U test, with Holm-Sidak multiple-comparison correction. n = 12 sham, n = 12 IR. (E) RT-PCR gene expression of senescence- and age-associated markers in sham (open blue circles) and irradiated (solid orange triangle) MEFs. Values are expression relative to sham control. (F) Concentrations of senescence-related proteins in conditioned media (MEF CM) measured in pg/mL with Luminex multiplex assay. (G) Normalized protein expression (NPX) of senescence-related proteins in conditioned media measured with Olink PEA assay. Inset shows IL-23R values re-scaled. For E-G, bars denote mean ± s.e.m., * p < 0.05 vs. sham by two-tailed Mann-Whitney U test, with Holm-Sidak multiple-comparison correction for data. n = 6 samples per group.
Senotherapeutics suppress age-associated plasma proteins
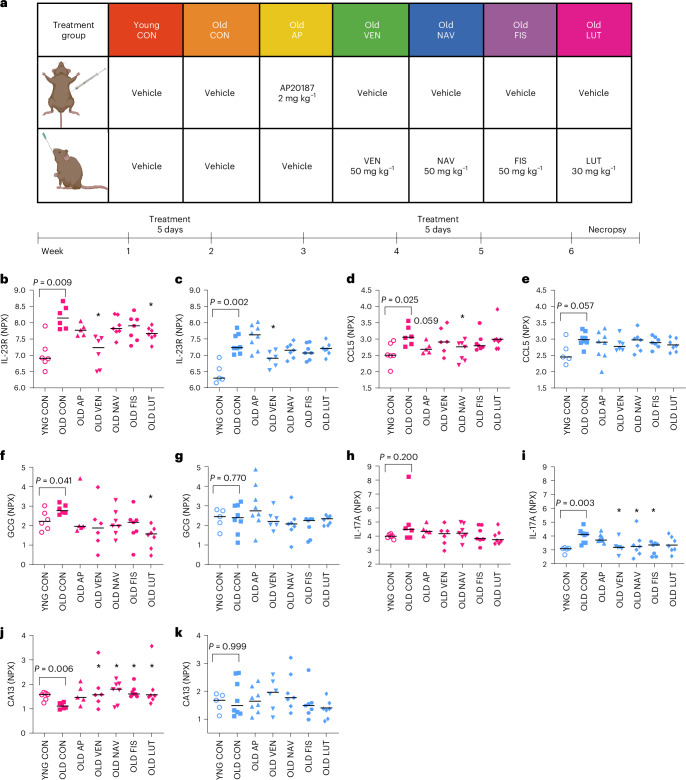
We next tested whether putative age-associated plasma proteins are modulated by drugs that target senescent cells in vivo. In old p16-InkAttac mice, we conducted head-to-head comparison of AP20187 (AP)-mediated transgenic versus senolytic interventions through a short-term, intermittent delivery strategy (Fig. 3a). All four compounds (VEN, NAV, FIS and LUT) demonstrated selective ability to induce apoptosis via cleaved caspase-3 activity in etoposide-induced senescent lung fibroblast in vitro (Extended Data Fig. 5). We hypothesized that senotherapeutic agents would reverse the direction of age-related changes of plasma proteins in vivo. Therefore, we analyzed drug responses for proteins demonstrating an age-dependent change using nonparametric, two-tailed statistical tests with correction for multiple comparisons (Fig. 3b–k). In old female mice, VEN and LUT reduced plasma abundance of IL-23R (Fig. 3b). In old male mice, VEN reduced plasma abundance of IL-23R (Fig. 3c). NAV reduced plasma abundance of CCL5 in old females and AP displayed a trending reduction of CCL5 (Fig. 3d). LUT reduced plasma abundance of GCG in old females (Fig. 3f). In old males, VEN, NAV and FIS reduced plasma abundance of IL-17A (Fig. 3i). CA13 was lower in old female plasma compared with young plasma and increased in old mice following treatment with VEN, NAV, FIS or LUT (Fig. 3j). Thus, senotherapeutic compounds revert the abundance of age-associated circulating proteins to more youthful levels, including modulation of factors that are both increased and decreased with aging. Among these data, IL-23R was elevated in aged mice across sexes and showed significant modulation by senotherapeutic VEN. We also tested whether any drugs ‘exacerbated’ the age-dependent change in plasma protein levels. In male plasma, FIS further increased the age-dependent change in CDH6, FSTL3, MIA and CCN4 (Source data).
Fig. 3. Age-related changes in plasma proteins are reverted by short-term senotherapeutic interventions.
a, Treatment groups (top) and senotherapeutic treatment timeline (bottom) of young and old mice that received either vehicle or AP20187 (AP) by i.p. injection and either vehicle or one of four senolytic drugs (VEN, NAV, FIS or LUT) by oral gavage. Beginning on week 1, mice were treated once daily for 5 days, followed by 2 weeks without treatment, followed by 5 days of further treatment, followed by 1 week without treatment before necropsy. b–k, Plasma protein levels are demonstrated for all biomarkers that differed by age and at least one senolytic intervention significantly reversed the age-related change in at least one sex. This criterion was met for IL-23R (b,c), CCL5 (d,e), GCG (f,g), IL-17A (h,i) and CA13 (j,k). Pink, female groups; blue, male groups. Protein values were measured by Olink PEAs and are reported as NPX. Bars summarize median values. Samples sizes: YNG CON female, n = 6; OLD CON female, n = 6; OLD AP female, n = 5; OLD VEN female, n = 5; OLD NAV female, n = 7; OLD FIS female, n = 7; OLD LUT female, n = 7; YNG CON male, n = 5; OLD CON male, n = 8; OLD AP male, n = 8; OLD VEN male, n = 6; OLD NAV male, n = 7; OLD FIS male, n = 7; OLD LUT male, n = 7. YNG CON versus OLD CON reflect statistics provided in Fig. 1 with exact P values provided. Kruskal–Wallis tests with multiple comparison correction were used for comparison of old treatment groups with OLD CON. We report *P < 0.05. See Methods for additional description of the two-step statistical approach. a, Created with BioRender.com.
Extended Data Fig. 5. Senolytic compounds have differential selectivity, potency, and efficacy in apoptosis of senescent cells in vitro.
Cleaved caspase-3 activity was measured in proliferating or etoposide-induced senescent primary human lung fibroblasts. (A–D) Concentration-response curves for percent change in cleaved caspase-3 intensity within senescent (red diamonds) or proliferating (white circles) cells and after incubation (3 days) with senolytic compounds (A) VEN, (B) NAV, (C) FIS, or (D) LUT. (E) Etoposide-induced senescent fibroblast or (F) proliferating fibroblast concentration-response curves of normalized cleaved caspase intensity by application of VEN (green circles), NAV (purple squares), FIS (orange triangles), or LUT (blue downward triangles) compounds. Symbols denote mean ± s.e.m., n = 4 samples per group.
Senescence-targeting drugs concentrate in secretory organs
Based on our observation that senolytics reversed age-associated changes in plasma biomarker abundance, we performed pharmacokinetic analysis in plasma and tissues using mass spectrometry at 20 min, 2 h, 6 h and 3 days following 4 consecutive days of treatment with AP or VEN (Supplementary Table 1). AP exhibited a low volume of distribution with relatively high plasma concentration compared with tissues. The steady-state concentrations of AP achieved in plasma, kidney and liver decreased gradually with an approximate half-life of 3–4 h, but detectable levels remained at 72 h (plasma, 2.1–6.6 µM; kidney, 0.2–0.8 µM; liver, 0.8–2.2 µM). In mice treated with VEN, maximum concentrations in plasma (2.9–6.0 µM), kidney (2.8–4.1 µM) and liver (18.8–29.5 µM) were detected 2 h after the final dose, and most of the drug was eliminated by 72 h. We note that these measurements were derived from 18-month-old mice that were drug-treated for only 4 days; older or diseased mice treated longer-term are expected to display distinct drug kinetics. Nevertheless, these results demonstrate that AP and VEN, drugs targeting senescent cells, concentrate and persist in kidney and liver, which harbor a substantial burden of senescent cell markers (Fig. 2) and, thus, are likely to induce on-target effects in these tissues.
Il23r and Ccl5 increase with age and are reduced by senotherapeutics
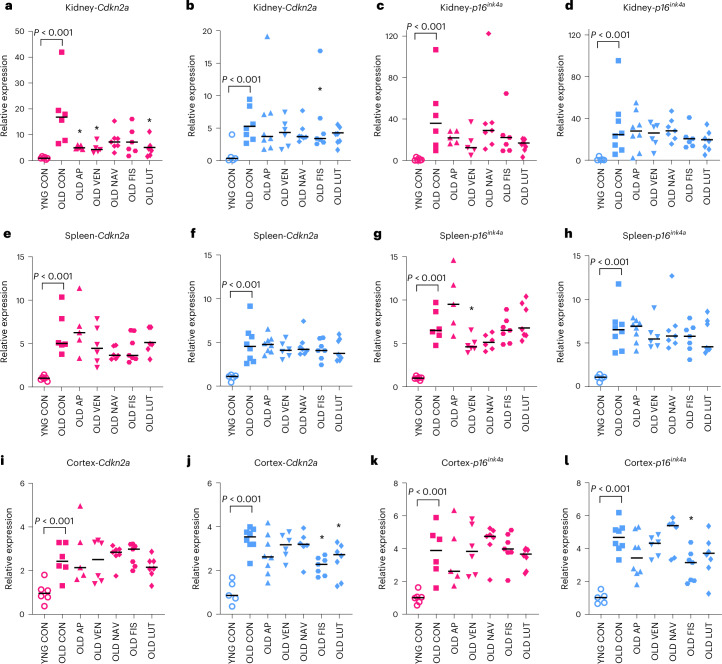
We next examined whether senotherapeutic compounds reduced the abundance of transcripts encoding age- and senolytic-altered plasma proteins in register with transcripts encoding canonical senescence factors. We hypothesized that senolytic treatments would reduce age-increased transcript levels, relative to old vehicle-treated mice and conducted hypothesis-directed two-tailed, nonparametric comparisons with multiple comparison corrections for females and males per tissue (Fig. 4). Previous p16-InkAttac targeting of p16-positive cells through long-term treatment with AP denoted clearance in a variable and tissue-specific manner, with substantial effects in kidney26. Consistent with those findings, we discovered that AP treatment reduced Cdkn2a expression in the aged female kidney (Fig. 4a). VEN and LUT also reduced Cdkn2a expression in female kidney (Fig. 4a) and FIS reduced Cdkn2a in male kidney (Fig. 4b). In female spleen, senolytics did not significantly alter Cdkn2a (Fig. 4e) but VEN reduced p16ink4a (Fig. 4g). FIS and LUT reduced Cdkn2a in cortex of male mice (Fig. 4j) and FIS reduced p16ink4a (Fig. 4l). We did not observe senolytic-mediated reversal of age-increased p16 expression in liver, lung or perigonadal adipose (Extended Data Fig. 6). The trend toward age-dependent increases in p21 expression in the spleen was reduced by NAV (Extended Data Fig. 7). The lower age-associated p21 expression in male cortex was reverted toward youthful levels by VEN (Extended Data Fig. 7). Additional tissue-specific senotherapeutic modulation of gene expression can be found in the Source Data. To further discern senolytic-dependent shifts in age-related factors, we performed principal component analysis on the combined kidney gene expression and plasma protein variables from young, old and treated old mice. We qualitatively observed that aged mice treated with AP or senotherapeutic drugs tended to cluster in between aged or young mice treated with vehicle (Supplementary Fig. 3 and Supplementary Table 2).
Fig. 4. p16 expression in old p16-InkAttac mice is reduced in a subset of aged tissues in response to short-term transgenic or pharmacological senescent cell targeting.
a–l, RT-PCR was used to assess Cdk2na (a,b,e,f,i,j) and p16ink4a (c,d,g,h,k,l) gene expression between old control (OLD CON) versus senolytic drug-treated mice, in kidney (a–d), spleen (e–h), and cortex (i–l). Values are expression relative to YNG CON per group. Pink, female groups; blue, male groups. Cdkn2a detects both transcript variants 1 (p19arf) and 2 (p16ink4a) (a,b,e,f,i,j) and p16ink4a detects only transcript variant 2 (c,d,g,h,k,l). YNG CON versus OLD CON reflect statistics provided in Fig. 2 with exact P values provided. Kruskal–Wallis tests with multiple comparison correction were used for comparison of old treatment groups with OLD CON. We report two-tailed *P < 0.05. Bars summarize the median group values. Samples sizes: YNG CON female, n = 6; OLD CON female, n = 6; OLD AP female, n = 5; OLD VEN female, n = 5; OLD NAV female, n = 7; OLD FIS female, n = 7; OLD LUT female, n = 7; YNG CON male, n = 5; OLD CON male, n = 8; OLD AP male, n = 8; OLD VEN male, n = 6; OLD NAV male, n = 7; OLD FIS male, n = 7; OLD LUT male, n = 7.
Extended Data Fig. 6. p16 gene expression in aged liver, lung or perigonadal adipose in response to short-term transgenic or pharmacological senescent cell targeting.
RT-PCR was used to assess p16 gene expression between old control (OLD CON) and senolytic drug treated mice, in (A–D) liver, (E–H) lung, and (I–L) perigonadal adipose. (A, C, E, G, I, K) Female groups are shown in pink, and (B, D, F, H, J, L) males are shown in blue. YNG CON versus OLD CON reflect statistics provided in Fig. 2 with exact p-values provided. Kruskal-Wallis tests with multiple comparison correction were used for comparison of old treatment groups with OLD CON. Bars summarize the median group values. Samples sizes per group, YNG CON female n = 6; OLD CON female n = 6; OLD AP female n = 5; OLD VEN female n = 5; OLD NAV female n = 7; OLD FIS female n = 7; OLD LUT female n = 7; YNG CON male n = 5; OLD CON male n = 8; OLD AP male n = 8; OLD VEN male n = 6; OLD NAV male n = 7; OLD FIS male n = 7; OLD LUT male n = 7.
Extended Data Fig. 7. p21 gene expression in aged tissues in response to short-term transgenic or pharmacological senescent cell targeting.
RT-PCR was used to assess p21 gene expression between old control (OLD CON) and senolytic drug treated mice, in (A, B) kidney, (C, D) liver, (E, F) spleen, (G, H) lung, (I, J) perigonadal adipose, and (K, L) cortex. (A, C, E, G, I, K). Female groups are shown in pink, and (B, D, F, H, J, L) males are shown in blue. YNG CON versus OLD CON reflect statistics provided in Fig. 2 with exact p-values provided. Kruskal-Wallis tests with multiple comparison correction were used for comparison of old treatment groups with OLD CON. We report two-tailed * p < 0.05, based on our hypothesis that each treatment would revert the levels of each age-associated factor closer to youthful levels. Bars summarize the median group values. Samples sizes per group, YNG CON female n = 6; OLD CON female n = 6; OLD AP female n = 5; OLD VEN female n = 5; OLD NAV female n = 7; OLD FIS female n = 7; OLD LUT female n = 7; YNG CON male n = 5; OLD CON male n = 8; OLD AP male n = 8; OLD VEN male n = 6; OLD NAV male n = 7; OLD FIS male n = 7; OLD LUT male n = 7.
We next determined whether Ccl5 expression in tissues decreased in response to each intervention (Fig. 5). Ccl5 exhibited widespread differential expression among aged organs and was expressed most abundantly in the aged spleen (Extended Data Fig. 1). VEN treatment decreased Ccl5 transcript abundance in female kidney (Fig. 5a) and spleen (Fig. 5e). FIS treatment also decreased Ccl5 expression in female spleen (Fig. 5e).
Fig. 5. Ccl5 gene expression decreases in a subset of aged tissues in response to short-term transgenic or pharmacological senescent cell targeting.
a–l, Comparison of Ccl5 gene expression measured by RT-PCR between senolytic drug intervention and OLD CON in kidney (a,b), liver (c,d), spleen (e,f), lung (g,h), perigonadal adipose (i,j) and cortex (k,l). Pink, female groups; blue, male groups. YNG CON versus OLD CON reflect statistics provided in Fig. 2 with exact P values provided. Kruskal–Wallis tests with multiple comparison correction were used for comparison of old treatment groups with OLD CON. We report two-tailed *P < 0.05. Bars summarize the median group values. Samples sizes: YNG CON female, n = 6; OLD CON female, n = 6; OLD AP female, n = 5; OLD VEN female, n = 5; OLD NAV female, n = 7; OLD FIS female, n = 7; OLD LUT female, n = 7; YNG CON male, n = 5; OLD CON male, n = 8; OLD AP male, n = 8; OLD VEN male, n = 6; OLD NAV male, n = 7; OLD FIS male, n = 7; OLD LUT male, n = 7.
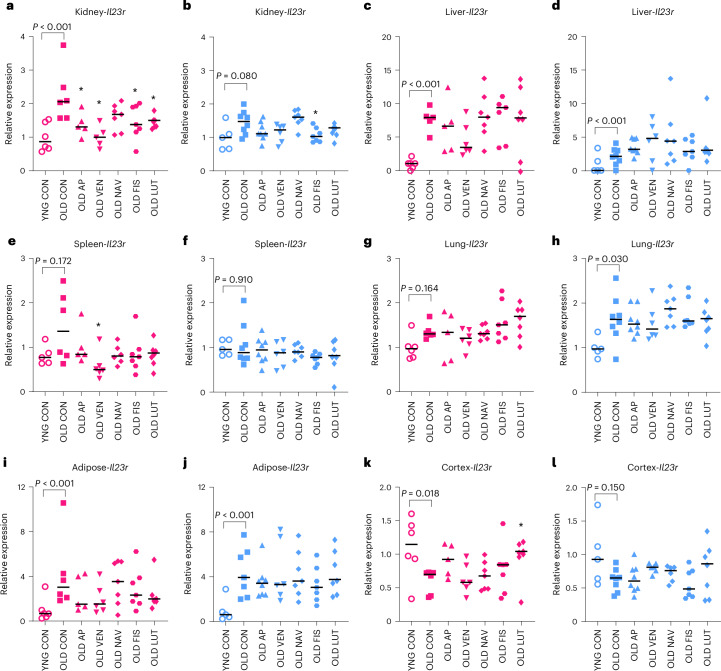
We next tested whether the senotherapeutics mitigated age-associated Il23r transcriptional changes (Fig. 6). In aged female kidney, AP treatment reduced Il23r expression (Fig. 6a). VEN reversed the age-dependent increases in Il23r in female kidney (Fig. 6a) and spleen (Fig. 6e). FIS and LUT also reduced Il23r expression in old female kidney compared with controls (Fig. 6a). In males, FIS reversed the age-dependent trend increase in Il23r in kidney (Fig. 6b). Antithetical to the peripheral tissues, LUT increased Il23r in old female cortex, toward the higher level of cortical expression in young mice (Fig. 6k). VEN exhibited consistent efficacy in mitigating senescence-related gene expression in the acute treatment paradigm, with partial phenocopy of AP-mediated senescent cell clearance in kidney. Altogether, the age- and senotherapeutic-mediated changes in plasma protein and gene expression suggest that IL-23R and CCL5 are important aging and senescence biomarkers.
Fig. 6. Il23r gene expression is altered in a subset of aged tissues in response to short-term transgenic or pharmacological senescent cell targeting.
a–l, Comparison of Il23r gene expression measured by RT-PCR between senolytic drug intervention and OLD CON in kidney (a,b), liver (c,d), spleen (e,f), lung (g,h), perigonadal adipose (i,j) and cortex (k,l). Pink, female groups; blue, male groups. YNG CON versus OLD CON reflect statistics provided in Fig. 2 with exact P values provided. Kruskal–Wallis tests with multiple comparison correction were used for comparison of old treatment groups with OLD CON. We report two-tailed *P < 0.05. Bars summarize the median group values. Samples sizes: YNG CON female, n = 6; OLD CON female, n = 6; OLD AP female, n = 5; OLD VEN female, n = 5; OLD NAV female, n = 7; OLD FIS female, n = 7; OLD LUT female, n = 7; YNG CON male, n = 5; OLD CON male, n = 8; OLD AP male, n = 8; OLD VEN male, n = 6; OLD NAV male, n = 7; OLD FIS male, n = 7; OLD LUT male, n = 7.
IL-23R protein increases in aged human plasma
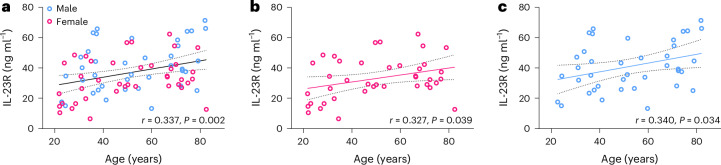
Our findings suggest that IL-23R is an age-increased plasma protein sensitive to senotherapeutic intervention. To begin to assess the translational relevance of this discovery, we measured IL-23R using enzyme-linked immunosorbent assay (ELISA) in human plasma from a cross-sectional sample of 79 Mayo Clinic Biobank participants. IL-23R increased with age in the combined sample (r = 0.337, P = 0.002) and separately in female (r = 0.327, P = 0.039) and male (r = 0.340, p = 0.034) participants (Fig. 7). We next examined IL-23R protein abundance in young and aged mouse and human samples. In mouse samples, we performed Olink assays that included plasma standards spiked with IL-23R protein. We found that average IL-23R plasma concentration for 22- to 24-month-old female mice was 78 ± 5 pg ml−1 (n = 17), whereas young plasma samples (n = 7) were below the lower limit of detection of the standard curve (Supplementary Table 3). Using ELISAs with IL-23R protein standards, we determined that average IL-23R in human plasma was 37 ± 3 ng ml−1 for aged females 66–82 years old and 46 ± 4 ng ml−1 for males 67–82 years old. Therefore, IL-23R increases with age in both mice and humans.
Fig. 7. Plasma IL-23R protein increases with age in humans.
a–c, Measurement of IL-23R in human EDTA plasma in combined (a), female (pink) (b) and male (blue) (c) Mayo Clinic Biobank participants aged 20–82 years. Best fit line and 95% confidence interval (dashed lines) were determined by linear regression. Female participants (F = 4.545, d.f. = 38, P = 0.0395). Male participants (F = 4.837, d.f. = 37, P = 0.0342). Two-tailed test of Pearson correlation values (r) and P values are provided for each plot.
Discussion
Our study was developed under the hypothesis that some age-associated proteins in circulation originate from, or are influenced by, senescent cells. We further hypothesized that senescent cell clearance would reduce age-increased (putative progeronic) plasma proteins and increase age-reduced (putative rejuvenative) plasma proteins, ultimately restoring the circulating milieu to a more youthful status. These hypotheses are linked to the logic that benefits derived from systemic deletion of senescent cells may result from both tissue-localized and circulating effects. We provide evidence for both modalities in response to senescent cell targeting. Our study reveals age-associated plasma proteins and tissue-expressed genes that are responsive to short-term senotherapeutic intervention. We demonstrate circulating biomarkers that are reflective of systemic senescent cell burden and senescent cell targeting.
IL-23R, CCL5, CA13 and additional factors have emerged from this work as critical aging biomarkers that are linked to abundance of systemic senescence gene expression, are responsive to senotherapeutics and are possible mediators of interorgan signaling mediated by senescent cells. According to gene ontology terms, the proteins that were lower in aged plasma were potentially cell membrane or cytoplasmic in origin and possibly related to DNA damage and transcriptional defects37. For example, CA13 is a cytosolic enzyme that catalyzes hydration of carbon dioxide and participates in bicarbonate regulation and pH homeostasis38. CA13 decreased in aged female plasma and the protein levels were elevated by the four pharmacological senotherapeutic strategies, suggestive of an intriguing candidate rejuvenative factor under senotherapeutic control. Many of the proteins that were higher in aged plasma were secreted factors16,39. For example, CCL5 is a T cell regulatory chemokine and an established inflammatory SASP factor20,40. We also identified IL-23R as a notable age-increased circulating protein that is suppressed by senotherapeutics in both sexes.
The IL-23–IL-23R signaling pathway is critical in the maturation of T helper 17 (TH17) cells and the orchestration of inflammatory signals41–44. Mice with genomic deficit of IL-23R do not elicit TH17 responses and are resistant to experimental autoimmune encephalomyelitis45, and IL-23R dysfunction is linked to susceptibility to several autoimmune diseases46. As the body ages, proinflammatory TH17 cells increase that are IL-23R positive47,48. Furthermore, age-associated upregulation of IL-23 may facilitate shifts of IL-23R in circulation in concert with other inflammatory mechanisms49. Although most widely studied in immune cells, IL-23R is also expressed in nonlymphoid cells, including renal tubular epithelial cells, wherein tubular epithelial cell-specific IL-23R deletion suppresses pathogenic kidney inflammation in vivo50. In the described contexts, IL-23R is a surface-expressed receptor; however, the cellular origins and mechanistic influence of circulating soluble IL-23R are not well established. Detected in humans but not yet confirmed in mice, a splice variant of IL-23R is manufactured as incapable of transmembrane anchoring and therefore, escapes the cell as soluble IL-23R, which may have an entirely different function to its membrane-bound counterpart51,52. Furthermore, ectodomain shedding of both human and murine IL-23R can occur by interaction with ADAM enzymes, which could be another source of soluble IL-23R53. We speculate that age-related circulating IL-23R may correspond to a soluble form of the receptor and/or vesicle-associated protein, which could originate from membrane-bound IL-23R. Overexpression of soluble IL-23R in vivo interferes with the IL-23–IL-23R interaction that promotes inflammation52. Importantly, the Elisseeff laboratory recently demonstrated reciprocal control between senescent cells and TH17 T cells in aging and injury models, wherein targeting either population can attenuate pathology and/or promote regeneration54–56. Related to these findings, we discovered that plasma IL-23R concentration increases in both aged mouse (using Olink assays) and human (using ELISA, since Olink assays are not available) plasma samples. Through senotherapeutic control of systemic IL-23R, we provide foundational evidence that circulating IL-23R is linked to cell senescence, but much remains to be understood concerning the modulatory roles of IL-23R in organ-distributed function in aging. Our data further the conclusion that the IL-23R axis is a central regulatory node in aged cell identities throughout the body, and we establish circulating IL-23R as a translationally relevant aging biomarker for ongoing clinical and preclinical research.
We implemented system-wide gene expression profiling with a goal of tracing circulating biomarkers to their origin tissues. Il23r expression increased in the aged kidney, liver, perigonadal adipose and lung, which may reflect increased abundance of Il23r-expressing senescent cells and/or tissue-infiltrating immune cells. In situ hybridization confirmed that Il23r and Cdkn2a were expressed abundantly and colocalized in the renal sinus and interstitium of the aged kidney. We conclude that senescent targeting was most effective in reducing Il23r in kidney, based on the observation that several drugs were effective in both sexes. Il23r transcript reduction through senolytic intervention was also achieved in spleen. This suggests that several tissues are viable sources of increased soluble IL-23R in aged plasma, and senotherapies may act in these tissues to control its abundance in blood circulation. We highlight the potential for senescent cell clearance in a primary tissue to have benefits in a secondary tissue through circulatory control.
A previous report showed that VEN can target BCL2-expressing, senescent beta cells in the pancreas, preventing type I diabetic pathology in mice57. Although we did not study the pancreas, we were intrigued to discover that senotherapeutic LUT intervention restored GCG plasma protein concentrations to youthful levels in aged females, which may coincide with metabolic benefits originating in the pancreas. Furthermore, we observed BCL2-inhibition through VEN as efficacious in reducing aging and senescence markers in the kidney and spleen. We detected qualitatively higher Bcl2 and/or Bcl2a1a expression in aged spleen and kidney, relative to other tissues analyzed. Therefore, BCL2-targeted efficacy may require a threshold of target induction before modulation can occur and/or be detected.
There are at least three scenarios by which age-increased plasma factors discovered here may be altered by senotherapeutics. First, these factors could originate from senescent cells, and eliminating senescent cells could reduce their production and secretion. Senomorphic effects may also occur, in which senescent cells are modified but not killed by the drug treatments. Second, these factors could be secreted from immune or other cells that are under regulatory control of senescent cells, wherein targeting of senescent cells could reduce the plasma factors through cell nonautonomous mechanisms. Third, these factors could be secreted from immune or other cells that are not associated with senescent cells but are modulated by the drugs. Related to points two and three, BCL2 inhibitors and flavanoid compounds have immunomodulatory effects that may or may not involve senescence58–60. We observed that Il23r/IL-23R is expressed in, and secreted from, senescent fibroblasts. Parallel senotherapeutic testing revealed that senolytic modulation mirrored and, in several cases, performed better than the p16-InkAttac transgenic approach. These results collectively suggest that eliminating p16-positive senescent cells underlies a portion, but not all, of the reductions in IL-23R and other age-related plasma factors following senolytic intervention. Senolytic drugs may act more broadly on other types of senescent and potentially nonsenescent cells, relative to exclusive elimination of p16ink4a-expressing cells, and senotherapeutics may also confer immunomodulatory properties that do not involve senescent cells. Disentangling senescence-dependent and independent mechanisms altered by senotherapeutics and interactions between senescent cells and immune cells are important directives in the field of translational aging research.
Beyond the key factors highlighted above, the breadth of our results reveals important findings that build on previous foundational discoveries. For example, Ccn1/Cyr61 was recently identified as a top upregulated gene in the ‘SenMayo’ senescent cell geneset61. CCN1 is a proinflammatory, matricellular protein that induces cytokine/chemokine synthesis in cooperation of tissue repair by activated macrophages62,63. We confirmed that CCN1 is increased significantly in aged mouse plasma and is also transcriptionally increased in aged adipose. Plasma CCN1 correlated positively with other senescence and inflammatory factors but was not altered significantly by short-term senolytic modulation. We note that the age of old transgenic mice used in this study (16–20 months) may reflect a heterogenous onset of old age, and we anticipate that longer-term treatment and larger sample sizes may be necessary for confirming senotherapeutic modulation of many emerging senescence biomarkers, including CCN1. Furthermore, we observed inverse aging patterns for select transcripts in the brain, relative to peripheral organs. This was true for Il23r and p21. p21 is expressed in quiescent progenitor cells, wherein it plays a role in maintenance of stemness, likely independent of a senescence phenotype64,65. While the source of the antithetical expression pattern for Il23r is undetermined, we speculate that reduced p21 transcript abundance in aging may reflect a reduction in progenitor cells in the aged brain. VEN increased p21 expression in aged male cortex closer to youthful levels; whether this reflects a rejuvenation effect of brain progenitor cells is yet unknown. However, this finding raises caution against use of p21 as a single biomarker for identification of senescent cells in the aged brain.
Select study design choices incorporated in our study warrant commentary. A strength of our design is the high number of head-to-head comparisons. Comparing five experimental drugs across biomarkers in plasma and six tissues for both sexes provides valuable quantitative and qualitative resource data applicable to a multitude of questions. However, limited sample sizes required for study feasibility challenged the statistical power. Nevertheless, we used hypothesis-directed nonparametric statistical tests with multiple comparison correction to examine the senotherapeutic modulation of aging. From this approach, identification of consistent tissue-specific and drug-response patterns fostered our discovery of aging- and senescence-related biomarkers in context with well-established molecular signatures. The p16-InkAttac transgenic mouse, maintained on C57BL/6 background, may display variable traits to that of other C57BL/6 aged mice. We also selected doses of drugs informed by our experience and the literature11,33,66, but additional senolytic drug and dose comparison experiments are warranted for lifespan and healthspan studies. Findings provided herein will be very useful for single-agent and/or tissue-specific senescence investigations, which may include larger sample sizes and altered treatment paradigms (for example, longer-term or variable dosing strategies) for interrogation of functional changes resultant of specific signaling factors. Sex is increasingly and appropriately recognized as a critical variable in aging biology. In the present study, female and male data were analyzed separately, which revealed sex-conserved and sex-specific results. IL-17A levels were significantly higher in aged male mice. CA13 and GCG significantly changed in aged female plasma, and IL-23R was significantly greater in both aged female and male plasma. Interestingly, these age-regulated factors were reverted by select senolytic interventions.
Our results suggest that several tissues, and particularly the kidney, have the potential to become endocrine mediators of aging via expression and secretion of Il23r, IL-23R and other factors that are controlled by senescent cells. The accumulation kinetics, identity and effects of senescent cell and SASP profiles are variable and heterogenous67–69. However, key senescence-related genes, including p16 and Il23r, identified by accumulation in aged kidney, were subject to acute modulation by senolytics in aged mice. Mapping the cellular origins and targets of progeronic and rejuvenative factors in circulation, including those that are regulated by cell senescence, is an important area of investigation anticipated to advance through emerging methods, including high-dimensional profiling, system-wide multiomics and bioorthogonal labeling. Our discoveries build on these trajectories and provide foundational insight into how senotherapeutics differentially alter age-related circulating factors, molecular fates of secretory organs and acute control over senescence-related biomarkers in old age.
Methods
This research complies will all ethical regulations, research practices, and experimental protocols by the Mayo Clinic Institutional Animal Care and Committee and the Mayo Clinic Institutional Review Board.
Animals
Mouse experiments were performed under protocols approved by Mayo Clinic Institutional Animal Care and Use Committee. Male and female heterozygous p16-InkAttac mice (C57BL/6 background) were used in this study, and sex-specific features of age were analyzed independently. Mice were group-housed in ventilated cages with a constant temperature of 25 °C, 30–70% humidity and a 12-h light/dark cycle. Water and the 5053 PicoLab Rodent Diet 20 were provided ad libitum. At the time of tissue collection, young mice were 3–4 months of age and old mice were 16–20 months of age. Mice were euthanized with a lethal dose of pentobarbital. Blood was acquired from the inferior vena cava with EDTA-coated syringes and stored in EDTA-coated tubes. Blood samples were centrifuged (1,300g for 15 min, 4 °C), and the plasma fractions were separated and stored at −80 °C. Following blood collection, mice were perfused transcardially with ice-cold PBS. All mice were examined for gross pathology and tumor prevalence, with exclusion of mice harboring pronounced splenomegaly. Brain cortex, liver, kidney, spleen, lung and perigonadal adipose were removed immediately and placed in TRIzol for gene expression analyses.
Senotherapeutic treatment of mice
For AP20187 drug dilution, a 12.5 mg ml−1 stock solution in 100% ethanol was created and stored at −20 °C. AP20187 working solution (1 ml; 2 mg kg−1) was prepared by mixing 40 µl stock solution with 100 µl PEG-400 and 860 µl of 2% Tween-20 in ddH2O. The solution was vortexed and administered to mice by intraperitoneal (i.p.) injection within 30 min of diluting the stock. The corresponding vehicle was composed of ethanol, PEG-400 and Tween-20 and was administered to control animals via i.p. injection. For senolytic compounds, a stock oral gavage vehicle was created as a solution of 60% phosphatidylcholine in propylene glycol (Phosal 50 PG, Lipoid GmbH), 30% PEG-400 and 10% ethanol. Senolytic compounds were freshly vortexed into vehicle solution at room temperature and stored at 4 °C the week of injections. Old p16-InkAttac mice were randomized (based on body weight and age) to receive vehicle or AP20187 (2 mg kg−1, i.p. injection), NAV (50 mg kg−1), VEN (50 mg kg−1), FIS (50 mg kg−1) or LUT (30 mg kg−1) by oral gavage. All mice received both gavage and i.p. treatments, with AP mice receiving gavage vehicle and senolytic drug groups also receiving vehicle by i.p. Young and old p16-InkAttac control mice (YNG CON, OLD CON) received the same vehicle dosing strategy (Fig. 3a). NAV (ABT-263, cat. no. 11500) and LUT (3′,4′,5,7-tetrahydroxyflavone, cat. no. 10004161) were acquired from Cayman Chemical. VEN (ABT-199, cat. no. A8194) was acquired from APExBio Technology. FIS (3,7,3′,4′-tetrahydroxyflavone, cat. no. T0121) was acquired from TCI America. AP20187 was acquired from WuXi AppTec. All other reagents, unless otherwise denoted, were acquired from Sigma-Aldrich/Millipore.
For the gene expression and plasma profiling focused study, mice received five consecutive daily administrations for 1 week, 2 weeks off and then another 5-day treatment. Treatments were discontinued 1 week before necropsy (Fig. 3a). Samples sizes of each senotherapeutic group, YNG CON: n = 6 female/5 male; OLD CON: n = 6 female/8 male; OLD AP: n = 5 female/8 male; OLD VEN: n = 5 female/6 male; OLD NAV: n = 7 female/7 male; OLD FIS: n = 7 female/7 male; OLD LUT: n = 7 female/7 male. For the pharmacokinetic study of drug accumulation/elimination in tissue, 18-month-old mice were treated once daily with 2 mg kg−1 AP (i.p.), 50 mg kg−1 VEN (oral gavage), or control vehicle for 4 days. Mice were euthanized 20 min, 2 h, 6 h or 72 h following the fourth and final treatment, perfused with ice-cold PBS, and collected tissues were flash-frozen in liquid nitrogen. Sample sizes of each pharmacokinetic group according to time of collection post injection: vehicle (20 min): two female, two male; AP (20 min): two female, two male; AP (2 h): two female, two male; AP (6 h): four female; AP (3 days): two female, one male; VEN (20 min): two female, two male; VEN (2 h): two female, two male; VEN (6 h): one female, one male; VEN (3 days): two female, two male.
Real-time PCR
Tissue samples collected at necropsy were stored immediately in TRIzol reagent. RNA was isolated using TRIzol-based chloroform–isopropanol precipitation, followed by nanodrop concentration and purity analysis. For all tissues, 2 µg of total RNA was used for cDNA synthesis through M-MLV reverse transcription (Invitrogen, cat. no. 18091200). RT-PCR was performed on a QuantStudio5 RT-PCR system (ThermoFisher) with PerfeCTa FastMix II Low ROX (Quantabio) and Taqman PrimeTime qPCR Primer assays from Integrated DNA Technologies (Supplementary Table 4). Quantstudio 5 Design and Analysis Software v.1.5.2 was used for data collection of PCR CT values. Experimental genes were normalized to Hprt or Tbp housekeeper gene expression. Relative gene expression compared with the young control group was derived from the 2−ΔΔCT value for each tissue sample. In studies involving RT-PCR analyses in young versus old tissues, we report CT values in addition to relative expression results (Extended Data Fig. 1). In system-wide comparisons across several drug treatments, we found it very informative to examine both relative expression and total abundance of the transcripts (CT values) according to tissue of origin and age. The total abundance qualitative comparison is feasible since cDNA was synthesized from the same amount of input RNA per tissue sample (2 µg).
In situ hybridization
Kidneys were collected from 4-month- and 24-month-old mice, fixed in 4% paraformaldehyde for 24 h, embedded in paraffin and sectioned at 5 μm. The RNAscope fluorescent multiplex assay (ACD-Bio/Bio-Techne) was performed on FFPE (formalin fixed, paraffin embedded) tissue, according to the manufacturer’s specifications and protocols. Cdkn2a/p16ink4a probe (cat. no. 447491, Mm-Cdkn2a-tv2) was used in channel 1 and paired with TSA Vivid 520 fluorophore (cat. no. 323271). Il23r (cat. no. 403751-C2, Mm-Il23r-C2) was used in channel 2 and paired with TSA Vivid 570 fluorophore (cat. no. 323272). Cd3e (cat. no. 314721-C3, Mm-Cd3e-C3) was used in channel 3 and paired with TSA Vivid 650 (cat. no. 323273). Negative control probe DapB (cat. no. 320871) was used on parallel sections as control. TSA Vivid fluorophores were reconstituted in 100 μl dimethylsulfoxide and diluted with TSA Buffer at a 1:1,000 concentration. Tissues were quenched with TrueBlackPlus (1× in PBS, Biotium) for 10 min before adding Vectashield and coverslip. Sections were imaged with a Nikon Ti2 Eclipse Inverted microscope with ×10, ×20 and ×40 Plan Apo objectives, eight-channel Spectra III light engine and using the Orca Fusion BT sCMOS camera with Nikon Elements AR software; 405, 488, 561 and 647 laser lines were used with an image exposure time of 100–200 ms per channel in 16-bit readout mode.
Mouse plasma protein analysis
The Olink Target 96 Mouse Exploratory panel was used to analyze 92 protein biomarkers from mouse plasma by PEA. Target-specific antibody pairs are linked to DNA oligonucleotides and, upon binding to the target protein, the antibodies are brought into close proximity and the oligonucleotides hybridize, enabling amplification with signal quantification. Protein CT values were normalized by log2 scale, denoted as normalized protein expression (NPX). A total of 67 proteins were analyzed and reported. 25 proteins were excluded from age-associated and senolytic comparisons due to low or lack of detection. 1 µl plasma was used for PEA per sample.
Human plasma protein analysis
Human plasma analyses were approved by the Mayo Clinic Institutional Review Board. Archived Mayo Clinic Biobank human plasma samples were used for this study70. All study participants provided written informed consent at the time of sample collection. Samples corresponded to 40 women and 40 men between 20 and 90 years of age. Participants with a history of cancer, other than breast cancer and melanoma, before the age of 50 years, or autoimmune diseases (for example, rheumatoid arthritis, lupus), and women with body mass index (BMI) less than 18.5 kg m−2 or greater than 40.0 kg m−2 and men with BMI less than 18.5 kg m−2 or greater than 38.0 kg m−2 were excluded. One male participant was excluded as an outlier on the basis of BMI by Grubbs outlier test. EDTA blood was collected and processed into platelet-poor plasma immediately, and aliquots were frozen at −80 °C until used. EDTA platelet-poor plasma (1 ml) was thawed on ice and spun down at 15,000g, 4 °C for 5 min to remove insoluble particles. The supernatants were transferred to a fresh clean tube, homogenized and re-aliquoted into smaller aliquots to use on each day to avoid repeated freeze–thaw cycles, and these aliquots were kept at −80 °C. We used the Ancillary Reagent Kit for ELISA (cat. no. DY008B, R&D Systems/Bio-Techne) and the antibody pair from the Human IL-23R DuoSet ELISA (cat. no. DY1400-05, R&D Systems/Bio-Techne) for the development of an ELISA protocol with feasible performance to quantify the amount of soluble IL-23R in EDTA plasma. We accessed the optimal condition for linear range, accuracy, precision and parallelism. The experiments were run according to the manufacturer’s protocol, except for the blocking buffer (2% BSA, 4 mM EDTA in PBS) and sample/standard diluent (BSA 1%, EDTA 4 mM, in PBS). The linear range was 94-6000 pg ml−1, and the recovery ranged from 80% to 105%. The optical density was read using iD3 Spectramax (Molecular Devices) and five parameter nonlinear regression was used to generate the standard curve and interpolate the results using SoftMax Pro Software v.6.1 (Molecular Devices).
MEF cell culture and senescence confirmation
A fibroblast culture model was used to validate hallmark markers of senescence26. Embryonic day 13.5 fibroblasts were harvested from C57BL/6 mice and cultured in DMEM containing 10% fetal bovine serum (FBS) until passage 4 (P4). P4 cells were plated in six-well plates or T75 flasks. At 40% confluency, cells were subjected to sham conditions or irradiated at 10 Gy (RadSource Technologies RS 2000). Cells were harvested 24 h after sham treatment or 10 days postirradiation. At 24 h before harvest or fixing, the culture medium was replaced with FBS-free medium. Conditioned medium was collected from T75 flasks, 0.2 µm-filtered and snap-frozen at −80 °C. Cells were trypsinized, resuspended in 1 ml TRIzol and snap-frozen at −80 °C before processing for RT-PCR. For SA-β-gal staining, cells in six-well plates were fixed with 4% paraformaldehyde for 10 min, rinsed with PBS and stained with solution containing 40 mM citric acid/Na2HPO4 buffer at pH 6.0, 5 mM potassium ferrocyanide, 5 mM potassium ferricyanide, 150 mM NaCl2, 2 mM MgCl2 and 1 mg ml−1 (25 mM) 5-bromo-4-chloro-3-indolyl-β-d-galactopyranoside (VWR) in water. Cells were stained for 27.5 h at 37 °C with low shaking. Cells were rinsed with PBS, and counterstained with 4′,6-diamidino-2-phenylindole (DAPI) at 1 µg ml−1 for nuclear colocalization. Images were acquired with 405 nm fluorescence and brightfield on an Eclipse Ti inverted microscope (Nikon Instruments) for visualization of SA-β-gal-converted chromogenic compound. iLastik v.1.4.0 pixel classification workflow was used to create a binary mask of brightfield images71. Unstained regions and SA-β-Gal stained regions were added manually for training, with one image per well for both sham and IR included in the training set. The resulting classifier was then run on the entire set of images producing binary masks. CellProfiler v.4.2.5 was used to quantify and describe the cells and SA-β-Gal stained regions within each image72. Tiff files of DAPI staining were used for the cells and the tiff files produced by iLastik were used for SA-β-Gal. DAPI images were run through the ColorToGray module to generate a grayscale image, after which both the DAPI grayscale and the iLastik SA-β-Gal binary images were run through the following modules (with distinct settings): IdentifyPrimaryObjects, MeasureObjectSizeShape, OverlayOutlines, SaveImages and ExportToSpreadsheet. The objects identified in IdentifyPrimaryObjects were overlaid onto images and saved for review. All measurements were exported into *.csv files and used for quantification.
For cell preparation, conditioned medium was aspirated off from cells grown in T75 flasks and the cell monolayer was washed once with 5 ml of 1× PBS; 2 ml of TrypLE (Gibco) was added for 4 min at room temperature, after which the flasks were tapped against the bench three times to dislodge all cells, and the TrypLE was quenched with 5 ml of MEF medium. The cell suspension was pipetted over the flask three times, then transferred to a 15 ml tube, spun at 188g for 5 min at 22 °C, the supernatant was aspirated, the pellet resuspended in 1 ml of Trizol, solution was transferred to a 1.5-ml tube, and stored at −80 °C. RNA extraction was then performed as described above.
Conditioned medium from sham and IR cells were used for multiplexed sandwich ELISAs using the Luminex platform (Bio-Techne). Analyte-specific antibodies were precoated onto magnetic beads embedded with fluorophores. Microparticles, standards and medium samples were pipetted into wells and analytes of interest bound by immobilized antibodies. After washing off unbound material, biotinylated antibody cocktail was added to each well. Following another wash step, Streptavidin-PE conjugate was added. After a final wash, microparticles were resuspended in buffer and read using the Intelliflex xMAP to magnetically capture the beads and read the fluorophore signals unique to each analyte. Conditioned medium, with samples run in duplicate, was also tested with the Olink PEA assay Target 96 Mouse Exploratory panel. For PCR and conditioned medium analyte measurements, six sham and six irradiated experiment samples were used.
Senolytic drug testing in senescent lung fibroblasts
Primary human lung fibroblasts were purchased from ScienCell and cultured in EMEM containing 10% FBS. Senescence was induced by treating cells with 20 µM etoposide for 48 h, followed by 6 days in normal culture medium as we have performed previously6,73. Proliferating and senescent cells were then plated into 96-well plates and allowed to attach. After 6 h, the medium was changed to EMEM containing 0% FBS and the indicated concentration of compound was added. After 72 h the cells were fixed with 4% formalin, permeabilized with 0.25% Triton X-100, blocked with 1% BSA and immunostained with an antibody for cleaved caspase-3 (Cell Signaling, cat. no. 9661, RRID:AB_2341188, 1:500 dilution). Alexa Flour-488 secondary anti-rabbit (ThermoFisher, cat. no. A-11008, RRID:AB_143165, 1:1,000 dilution) was incubated along with DAPI. Images were collected using a Cytation5 automated microscope (Biotek) using a ×4 objective and cleaved caspase-3 intensity was quantified using onboard software (Gen5). To validate the detection of apoptosis, alternatively, etoposide-induced senescent primary human lung fibroblasts were cultured for 24 h with 1 µM staurosporine to induce apoptosis and determine the ceiling of cleaved caspase-3 activity (Supplementary Fig. 4). Cells were fixed, and immunostained for cleaved caspase-3 and DAPI, with four samples collected per group. Percent change in cleaved caspase-3 intensity was normalized to the greatest response, and concentration-response curves were fit by the Hill curve.
Pharmacokinetic analyses
Quantitation of AP and VEN in mouse plasma, kidney, liver, brain and muscle was performed on a Sciex 6500 MS/MS interfaced with a Sciex ExionLC 2.0 liquid chromatography system. Plasma samples were extracted, and proteins were precipitated by mixing with three times v:v acetonitrile containing carbamazepine as an internal standard. Samples were allowed to sit on ice for approximately 30 min with occasional agitation before filtration through a 0.2-µm filter plate. Standards were prepared by spiking AP and VEN standards into blank plasma and processed using the same procedure as the samples. Kidney, liver, brain and muscle samples were diluted threefold (w:v) with PBS and dissociated with a probe tip sonicator. The homogenates were processed similarly to the plasma samples. VEN was detected in positive ion mode following the parent ion 868.3 and fragment ion 636.2 and AP used parent ion 742 and fragment ion 209.2.
Statistics and reproducibility
We describe a stepwise statistical approach for assessing age- and treatment-dependent changes in plasma protein and tissue gene expression in the Supplementary Text and Supplementary Fig. 1. Data were not assumed to be normally distributed and nonparametric tests were used to be as rigorous as possible. Briefly, for initial plasma screening, two-tailed Kruskal–Wallis rank sum tests were used to determine group differences for any of the proteins, separately for each sex. Only those proteins in which there is an overall group level P value < 0.10 were selected for testing reversal by senotherapeutic treatment. We then performed a two-tailed test comparing each treated group of old mice (OLD AP, OLD VEN, OLD NAV, OLD FIS and OLD LUT) with OLD VEH, with significance set at P value < 0.05 (Source Data Figs. 1 and 3). RT-PCR genes with an age group effect at the P < 0.05 significance level were identified for further evaluation. For genes with an overall group difference of P < 0.05, two-tailed comparisons were made between old drug-treated and the old vehicle group for each tissue type. P values were adjusted using Dunnett correction to account for multiple comparisons compared with the OLD VEH group. Correlations between RT-PCR and plasma protein data were interrogated using Spearman correlations and significance was determined with the criterion P < 0.05. Sample sizes were determined according to current standards used for mice in experimental biology, based on the minimal number of mice required to detect significance with an alpha set at 0.05 in two-tailed test in a standard powered experiment6. Sample sizes were five to eight mice per group. Mouse data collection and analyses were performed by the experimenter blinded to conditions of the experiments. GraphPad Prism v.9 and R v.4.1.2 were used for statistical analyses.
For gene expression from cultured MEFs or protein measurement from their conditioned medium, comparisons were made between sham control and irradiated samples using Mann–Whitney test with significance at P < 0.05, adjusted for multiple comparisons with the Holm–Sidak method; n = 6 samples per group.
Human plasma IL-23R data were first tested for normality by Shapiro–Wilk and Kolmogorov–Smirnov tests. Both male and female data were found to be normally distributed and best fit by Pearson linear regression with correlation R values determined.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Supplementary information
Supplementary Information Text, Figs. 1–4 and Tables 1–4.
Pharmacokinetic quantification of AP20187 and venetoclax in mouse tissues.
Principal component analysis of kidney gene expression in response to senolytic drug treatment.
Quantification of Olink plasma IL-23R in mouse.
Taqman PrimeTime qPCR Primer assays from Integrated DNA Technologies used in gene expression analysis.
Source data
Olink mouse plasma protein data.
RT-PCR gene expression for young and old mouse tissues.
Olink mouse plasma protein data of in vivo senolytic drug treatment.
RT-PCR gene expression for in vivo senolytic drug treatment in aged mouse tissues.
IL-23R ELISA quantification in human plasma.
Spearman correlations of RT-PCR gene expression and plasma protein.
Cleaved caspase-3 quantification for in vitro cells in response to senolytics.
Acknowledgements
We thank M. Cameron (UF Scripps) for technical assistance with the mass spectrometer. We thank M. Romero (Mayo Clinic) for guidance with kidney tissue histology. We thank Bio-Techne R&D Systems (K. Nelson and N. Schrantz) for generously performing the Luminex experiments on MEF conditioned medium with Intelliflex xMAP equipment platform at the R&D Systems facility. We are grateful to the Mayo Clinic Biobank and support from the Mayo Clinic Center for Individualized Medicine for providing human plasma samples. We thank the Mayo Clinic Department of Comparative Medicine for support in mouse colony housing and care. We thank the University of Minnesota Genomics Center for assistance with Olink sample experiments. This study was funded by NIH K99/R00 AG058798, R01 AG071686 and UG3 CA275669 (M.J.S.). The pharmacokinetic data were acquired by a mass spectrometer funded by NIH grant 1 S10OD030332-01 (T.M.K.) and were supported by the Mayo Clinic Robert and Arlene Kogod Center NIH Program Project Grant P01 AG062413 (N.K.L.).
Extended data
Author contributions
M.J.S., C.M.C. and S.L.R. conceived and designed the study. A.J.H. and N.K.L. contributed to study design. D.J.B. and K.B.J. provided the mouse embryonic fibroblast protocol and advice in its application. M.J.S., C.M.C. and S.L.R. managed the aging mouse colonies, performed animal treatments, and collected and processed animal tissues. C.M.C., N.C.A., A.J.D. and M.J.S. analyzed data and generated figures. E.J.A. performed statistical modeling and data analysis. A.J.D. and N.C.A. performed mouse embryonic fibroblast in vitro experiments. P.T.G. and E.A.L. provided technical assistance in experiments. T.M.K. performed and provided funding for the pharmacokinetic experiments and analyses. A.J.H. and M.G.W. conducted comparative testing of senolytic drugs in vitro. J.M.E.-N. and C.M.C. analyzed human plasma samples. M.J.S. provided study funding. C.M.C. and M.J.S. drafted the paper. All authors reviewed and approved the paper.
Peer review
Peer review information
Nature Aging thanks Nathan Basisty and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Data availability
The original contributions presented in the study are included in the article and Supplementary Information. Pharmacokinetic data of AP20187 and venetoclax are included in Supplementary Table 1. Principal component analysis values for combined kidney gene expression and plasma protein are included in Supplementary Table 2 and also shown in Supplementary Fig. 3. All primer assays used for RT-PCR are included in Supplementary Table 4. All data supporting the findings of this study are also available from the corresponding author upon request. Source data are included with this paper.
Code availability
The R code used for statistical analysis of plasma protein and gene expression in Figs. 1–6 is publicly available via Code Ocean at 10.24433/CO.4129341.v1 (ref. 74) and details of its use are included with the Supplementary Information. iLastik and CellProfiler software are open source, well-documented and available to the public.
Competing interests
M.J.S., C.M.C., A.J.H., E.J.A., N.K.L. and Mayo Clinic have intellectual property related to this research. This research was reviewed by the Mayo Clinic Conflict of Interest Review Board and was conducted in compliance with Mayo Clinic Conflict of Interest policies. D.J.B. has a potential financial interest related to this research. He is a co-inventor on patents held by Mayo Clinic, patent applications licensed to or filed by Unity Biotechnology, and a Unity Biotechnology shareholder. Research in the Baker laboratory has been reviewed by the Mayo Clinic Conflict of Interest Review Board and is being conducted in compliance with Mayo Clinic Conflict of Interest policies. The other authors declare no competing interests.
Footnotes
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
is available for this paper at 10.1038/s43587-024-00752-7.
Supplementary information
The online version contains supplementary material available at 10.1038/s43587-024-00752-7.
References
- 1.Bianchi, T., Rufer, N., MacDonald, H. R. & Migliaccio, M. The tumor suppressor p16Ink4a regulates T lymphocyte survival. Oncogene25, 4110–4115 (2006). [DOI] [PubMed] [Google Scholar]
- 2.Yousefzadeh, M. J. et al. An aged immune system drives senescence and ageing of solid organs. Nature594, 100–105 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Childs, B. G. et al. Senescent intimal foam cells are deleterious at all stages of atherosclerosis. Science354, 472–477 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Roos, C. M. et al. Chronic senolytic treatment alleviates established vasomotor dysfunction in aged or atherosclerotic mice. Aging Cell15, 973–977 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Schafer, M. J., Miller, J. D. & LeBrasseur, N. K. Cellular senescence: implications for metabolic disease. Mol. Cell. Endocrinol.455, 93–102 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Schafer, M. J. et al. Cellular senescence mediates fibrotic pulmonary disease. Nat. Commun.8, 14532 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Khosla, S., Farr, J. N., Tchkonia, T. & Kirkland, J. L. The role of cellular senescence in ageing and endocrine disease. Nat. Rev. Endocrinol.16, 263–275 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Li, C., Shen, Y., Huang, L., Liu, C. & Wang, J. Senolytic therapy ameliorates renal fibrosis postacute kidney injury by alleviating renal senescence. FASEB J.35, e21229 (2021). [DOI] [PubMed] [Google Scholar]
- 9.Ogrodnik, M. et al. Cellular senescence drives age-dependent hepatic steatosis. Nat. Commun.8, 15691 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Zhang, X. et al. Characterization of cellular senescence in aging skeletal muscle. Nat. Aging2, 601–615 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Bussian, T. J. et al. Clearance of senescent glial cells prevents tau-dependent pathology and cognitive decline. Nature562, 578–582 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Zhang, X. et al. Rejuvenation of the aged brain immune cell landscape in mice through p16-positive senescent cell clearance. Nat. Commun.13, 5671 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Villeda, S. A. et al. The ageing systemic milieu negatively regulates neurogenesis and cognitive function. Nature477, 90–94 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Yousefzadeh, M. J. et al. Heterochronic parabiosis regulates the extent of cellular senescence in multiple tissues. Geroscience42, 951–961 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Krishnamurthy, J. et al. Ink4a/Arf expression is a biomarker of aging. J. Clin. Invest.114, 1299–1307 (2004). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Coppé, J. P. et al. Senescence-associated secretory phenotypes reveal cell-nonautonomous functions of oncogenic RAS and the p53 tumor suppressor. PLoS Biol.6, 2853–2868 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Coppé, J. P., Desprez, P. Y., Krtolica, A. & Campisi, J. The senescence-associated secretory phenotype: the dark side of tumor suppression. Annu. Rev. Pathol.5, 99–118 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Zhu, Y. et al. The Achilles’ heel of senescent cells: from transcriptome to senolytic drugs. Aging Cell14, 644–658 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Basisty, N. et al. A proteomic atlas of senescence-associated secretomes for aging biomarker development. PLoS Biol.18, e3000599 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Schafer, M. J. et al. The senescence-associated secretome as an indicator of age and medical risk. JCI Insight5, e133668 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Sturmlechner, I. et al. p21 produces a bioactive secretome that places stressed cells under immunosurveillance. Science374, eabb3420 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Jeon, O. H. et al. Systemic induction of senescence in young mice after single heterochronic blood exchange. Nat. Metab.4, 995–1006 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Pálovics, R. et al. Molecular hallmarks of heterochronic parabiosis at single-cell resolution. Nature603, 309–314 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Baker, D. J. et al. Clearance of p16Ink4a-positive senescent cells delays ageing-associated disorders. Nature479, 232–236 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Xu, M. et al. Targeting senescent cells enhances adipogenesis and metabolic function in old age. eLife4, e12997 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Baker, D. J. et al. Naturally occurring p16(Ink4a)-positive cells shorten healthy lifespan. Nature530, 184–189 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Carver, C. M. et al. Senescent and disease-associated microglia are modifiable features of aged brain white matter. Preprint at Research Square10.21203/rs.3.rs-3467812/v1 (2023).
- 28.Farr, J. N. et al. Local senolysis in aged mice only partially replicates the benefits of systemic senolysis. J. Clin. Invest.133, e162519 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Xu, M. et al. Senolytics improve physical function and increase lifespan in old age. Nat. Med.24, 1246–1256 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Park, C. M. et al. Discovery of an orally bioavailable small molecule inhibitor of prosurvival B-cell lymphoma 2 proteins. J. Med. Chem.51, 6902–6915 (2008). [DOI] [PubMed] [Google Scholar]
- 31.Souers, A. J. et al. ABT-199, a potent and selective BCL-2 inhibitor, achieves antitumor activity while sparing platelets. Nat. Med.19, 202–208 (2013). [DOI] [PubMed] [Google Scholar]
- 32.Kellenberger, E., Kuhn, I., Schuber, F. & Muller-Steffner, H. Flavonoids as inhibitors of human CD38. Bioorg. Med. Chem. Lett.21, 3939–3942 (2011). [DOI] [PubMed] [Google Scholar]
- 33.Yousefzadeh, M. J. et al. Fisetin is a senotherapeutic that extends health and lifespan. EBioMedicine36, 18–28 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Huang, X. F. et al. A network pharmacology strategy to investigate the anti-inflammatory mechanism of luteolin combined with in vitro transcriptomics and proteomics. Int. Immunopharmacol.86, 106727 (2020). [DOI] [PubMed] [Google Scholar]
- 35.Métais, J. Y. et al. BCL2A1a over-expression in murine hematopoietic stem and progenitor cells decreases apoptosis and results in hematopoietic transformation. PLoS ONE7, e48267 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Schenk, R. L., Strasser, A. & Dewson, G. BCL-2: long and winding path from discovery to therapeutic target. Biochem. Biophys. Res. Commun.482, 459–469 (2017). [DOI] [PubMed] [Google Scholar]
- 37.Stegeman, R. & Weake, V. M. Transcriptional signatures of aging. J. Mol. Biol.429, 2427–2437 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Lehtonen, J. et al. Characterization of CA XIII, a novel member of the carbonic anhydrase isozyme family. J. Biol. Chem.279, 2719–2727 (2004). [DOI] [PubMed] [Google Scholar]
- 39.Freund, A., Orjalo, A. V., Desprez, P. Y. & Campisi, J. Inflammatory networks during cellular senescence: causes and consequences. Trends Mol. Med.16, 238–246 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Zeng, Z., Lan, T., Wei, Y. & Wei, X. CCL5/CCR5 axis in human diseases and related treatments. Genes Dis.9, 12–27 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Cua, D. J. et al. Interleukin-23 rather than interleukin-12 is the critical cytokine for autoimmune inflammation of the brain. Nature421, 744–748 (2003). [DOI] [PubMed] [Google Scholar]
- 42.Duvallet, E., Semerano, L., Assier, E., Falgarone, G. & Boissier, M. C. Interleukin-23: a key cytokine in inflammatory diseases. Ann. Med.43, 503–511 (2011). [DOI] [PubMed] [Google Scholar]
- 43.Yago, T. et al. IL-23 and Th17 disease in inflammatory arthritis. J. Clin. Med.6, 81 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Łukasik, Z., Gracey, E., Venken, K., Ritchlin, C. & Elewaut, D. Crossing the boundaries: IL-23 and its role in linking inflammation of the skin, gut and joints. Rheumatology (Oxford)60, iv16–iv27 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Awasthi, A. & Kuchroo, V. K. Th17 cells: from precursors to players in inflammation and infection. Int. Immunol.21, 489–498 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Gaffen, S. L., Jain, R., Garg, A. V. & Cua, D. J. The IL-23–IL-17 immune axis: from mechanisms to therapeutic testing. Nat. Rev. Immunol.14, 585–600 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Schmitt, V., Rink, L. & Uciechowski, P. The Th17/Treg balance is disturbed during aging. Exp. Gerontol.48, 1379–1386 (2013). [DOI] [PubMed] [Google Scholar]
- 48.Uematsu, T., Fujita, T. & Kobayashi, N. Characterization of IL-17-producing T helper cells-like autoreactive T cells in aged mice. Exp. Anim.68, 483–490 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.El Mezayen, R., El Gazzar, M., Myer, R. & High, K. P. Aging-dependent upregulation of IL-23p19 gene expression in dendritic cells is associated with differential transcription factor binding and histone modifications. Aging Cell8, 553–565 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Li, H. et al. IL-23 reshapes kidney resident cell metabolism and promotes local kidney inflammation. J. Clin. Invest.131, e142428 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Yu, R. Y. & Gallagher, G. A naturally occurring, soluble antagonist of human IL-23 inhibits the development and in vitro function of human Th17 cells. J. Immunol.185, 7302–7308 (2010). [DOI] [PubMed] [Google Scholar]
- 52.Miralles, M. et al. Clinical and histopathological amelioration of experimental autoimmune encephalomyelitis by AAV vectors expressing a soluble interleukin-23 receptor. Neurotherapeutics14, 1095–1106 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Franke, M. et al. Human and murine interleukin 23 receptors are novel substrates for a disintegrin and metalloproteases ADAM10 and ADAM17. J. Biol. Chem.291, 10551–10561 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Chung, L. et al. Interleukin 17 and senescent cells regulate the foreign body response to synthetic material implants in mice and humans. Sci. Transl. Med.12, eaax3799 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Faust, H. J. et al. IL-17 and immunologically induced senescence regulate response to injury in osteoarthritis. J. Clin. Invest.130, 5493–5507 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Han, J. et al. Age-associated senescent—T cell signaling promotes type 3 immunity that inhibits the biomaterial regenerative response. Adv. Mater.36, e2310476 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Thompson, P. J. et al. Targeted elimination of senescent beta cells prevents type 1 diabetes. Cell Metab.29, 1045–1060.e10 (2019). [DOI] [PubMed] [Google Scholar]
- 58.Park, H. H. et al. Anti-inflammatory activity of fisetin in human mast cells (HMC-1). Pharmacol. Res.55, 31–37 (2007). [DOI] [PubMed] [Google Scholar]
- 59.Ludwig, L. M. et al. Venetoclax imparts distinct cell death sensitivity and adaptivity patterns in T cells. Cell Death Dis.12, 1005 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Hytti, M. et al. Two dietary polyphenols, fisetin and luteolin, reduce inflammation but augment DNA damage-induced toxicity in human RPE cells. J. Nutr. Biochem.42, 37–42 (2017). [DOI] [PubMed] [Google Scholar]
- 61.Saul, D. et al. A new gene set identifies senescent cells and predicts senescence-associated pathways across tissues. Nat. Commun.13, 4827 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Bai, T., Chen, C. C. & Lau, L. F. Matricellular protein CCN1 activates a proinflammatory genetic program in murine macrophages. J. Immunol.184, 3223–3232 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Jun, J. I. & Lau, L. F. The matricellular protein CCN1 induces fibroblast senescence and restricts fibrosis in cutaneous wound healing. Nat. Cell Biol.12, 676–685 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Marqués-Torrejón, M. et al. Cyclin-dependent kinase inhibitor p21 controls adult neural stem cell expansion by regulating Sox2 gene expression. Cell Stem Cell12, 88–100 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Porlan, E. et al. Transcriptional repression of Bmp2 by p21(Waf1/Cip1) links quiescence to neural stem cell maintenance. Nat. Neurosci.16, 1567–1575 (2013). [DOI] [PubMed] [Google Scholar]
- 66.Sawmiller, D. et al. Luteolin reduces Alzheimer’s disease pathologies induced by traumatic brain injury. Int. J. Mol. Sci.15, 895–904 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Nelson, G. et al. A senescent cell bystander effect: senescence-induced senescence. Aging Cell11, 345–349 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Acosta, J. C. et al. A complex secretory program orchestrated by the inflammasome controls paracrine senescence. Nat. Cell Biol.15, 978–990 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Teo, Y. V. et al. Notch signaling mediates secondary senescence. Cell Rep.27, 997–1007.e5 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Olson, J. E. et al. The Mayo Clinic Biobank: a building block for individualized medicine. Mayo Clin. Proc.88, 952–962 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Berg, S. et al. ilastik: interactive machine learning for (bio)image analysis. Nat. Methods16, 1226–1232 (2019). [DOI] [PubMed] [Google Scholar]
- 72.Stirling, D. R. et al. CellProfiler 4: improvements in speed, utility and usability. BMC Bioinf.22, 433 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Zhang, L. et al. Novel small molecule inhibition of IKK/NF-κB activation reduces markers of senescence and improves healthspan in mouse models of aging. Aging Cell20, e13486 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Atkinson, E. J. & Carver, C. M. IL23R is a senescence-linked circulating and tissue biomarker of aging. Code Ocean10.24433/CO.4129341.v1 (2024). [DOI] [PMC free article] [PubMed]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary Information Text, Figs. 1–4 and Tables 1–4.
Pharmacokinetic quantification of AP20187 and venetoclax in mouse tissues.
Principal component analysis of kidney gene expression in response to senolytic drug treatment.
Quantification of Olink plasma IL-23R in mouse.
Taqman PrimeTime qPCR Primer assays from Integrated DNA Technologies used in gene expression analysis.
Olink mouse plasma protein data.
RT-PCR gene expression for young and old mouse tissues.
Olink mouse plasma protein data of in vivo senolytic drug treatment.
RT-PCR gene expression for in vivo senolytic drug treatment in aged mouse tissues.
IL-23R ELISA quantification in human plasma.
Spearman correlations of RT-PCR gene expression and plasma protein.
Cleaved caspase-3 quantification for in vitro cells in response to senolytics.
Data Availability Statement
The original contributions presented in the study are included in the article and Supplementary Information. Pharmacokinetic data of AP20187 and venetoclax are included in Supplementary Table 1. Principal component analysis values for combined kidney gene expression and plasma protein are included in Supplementary Table 2 and also shown in Supplementary Fig. 3. All primer assays used for RT-PCR are included in Supplementary Table 4. All data supporting the findings of this study are also available from the corresponding author upon request. Source data are included with this paper.
The R code used for statistical analysis of plasma protein and gene expression in Figs. 1–6 is publicly available via Code Ocean at 10.24433/CO.4129341.v1 (ref. 74) and details of its use are included with the Supplementary Information. iLastik and CellProfiler software are open source, well-documented and available to the public.