Abstract
The upper airway is an important site of infection, but immune memory in the human upper airway is poorly understood, with implications for COVID-19 and many other human diseases1–4. Here we demonstrate that nasal and nasopharyngeal swabs can be used to obtain insights into these challenging problems, and define distinct immune cell populations, including antigen-specific memory B cells and T cells, in two adjacent anatomical sites in the upper airway. Upper airway immune cell populations seemed stable over time in healthy adults undergoing monthly swabs for more than 1 year, and prominent tissue resident memory T (TRM) cell and B (BRM) cell populations were defined. Unexpectedly, germinal centre cells were identified consistently in many nasopharyngeal swabs. In subjects with SARS-CoV-2 breakthrough infections, local virus-specific BRM cells, plasma cells and germinal centre B cells were identified, with evidence of local priming and an enrichment of IgA+ memory B cells in upper airway compartments compared with blood. Local plasma cell populations were identified with transcriptional profiles of longevity. Local virus-specific memory CD4+ TRM cells and CD8+ TRM cells were identified, with diverse additional virus-specific T cells. Age-dependent upper airway immunological shifts were observed. These findings provide new understanding of immune memory at a principal mucosal barrier tissue in humans.
Although peripheral blood sampling has proven useful for answering myriad clinical and research questions, sampling blood inherently misses important information at sites of disease. The upper airway is the primary site of infection for many human pathogens but has proved to be a challenging site to study immunologic memory, which is central to the protective immunity provided by almost all vaccines and infections. Critical outstanding questions remain about human germinal centre (GC) biology and human upper respiratory tract (URT) B cell and T cell immune memory, including development, abundance, maintenance and residency. This is illustrated by two of the largest and most comprehensive human tissue immunological studies, which had no representation of URT tissues5,6. Tissue resident memory T (TRM) cells are distinct memory T cells that do not circulate and are detectable in numerous human tissues5–11. T cell analysis from organ donors provided evidence for human tissue-specific TRM cells9,12. Human tissue resident memory B (BRM) cells also exist in several tissues13, but their biology in the upper airway is unknown. Additionally, while GCs are important for the generation of immune memory, they are classically very difficult to study in humans due to their inaccessibility14–19. Here we show that nasal and nasopharyngeal (NP) swabs can be used to obtain insights into immune memory of the human upper airway in adults, including the presence and persistence of antigen-specific B cell and T cell memory.
Several swab types and swabbing techniques were initially explored before suitable swabs and protocols were established for collecting upper airway cells. To determine the feasibility of upper airway swab sampling for studying human URT immune cell populations, we enroled a cohort of approximately 30 healthy adults to undergo monthly NP swabs for more than 1 year, with samples analysed by spectral flow cytometry. Monthly NP swab collections were well tolerated with minimal attrition of volunteers from the cohort. Up to 5 million viable cells could be collected with a single NP swab (median of more than 100,000; Fig. 1a), with approximately one-third being CD45+ immune cells (median 42,190 and maximum 2.75 million; Extended Data Fig. 1a), including B cells, CD4+ T cells and CD8+ T cells (Fig. 1b,c and Extended Data Fig. 1b). Epithelial cells were the primary CD45− cell types (Extended Data Fig. 1c–e). Upper airway immune cell populations, including total CD45+ cells, B cells, T cells, CD4+ T cells and CD8+ T cells, seemed stable over time (Fig. 1c). T cells from NP swabs included prominent putative TRM cell populations (Fig. 1d). CD8+ TRM cells were defined by coexpression of CD69 and CD103 on non-naive CD8+ T cells20 (Fig. 1d). URT CD4+ TRM cells were defined as CD69+ non-naive cells, with subpopulations coexpressing other putative TRM cell markers such as CD103 (ref. 20) (Fig. 1d). Putative BRM cells were observed in the NP swabs as CD69+ B cells13 (Fig. 1d). On examining the longitudinal donor cohort for these immune memory cell types, CD8+ TRM cells, CD4+ TRM cells and putative BRM cell populations were each observed to be stable over time (Fig. 1d).
Fig. 1 |. Distinct human upper airway cell populations sampled by NP swabs.
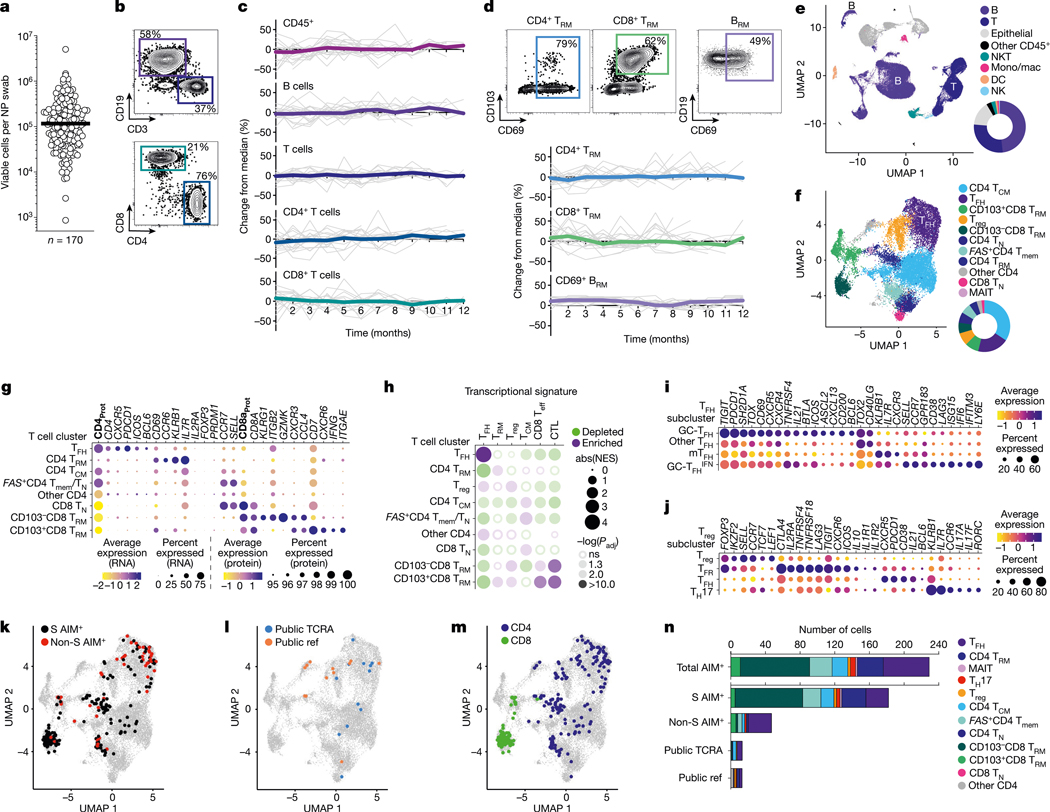
a, Number of viable cells per NP swab; each dot represents one NP swab per donor, black line is the median (n = 170 swabs). b, Representative flow cytometry plots of NP swab total B cells and T cells (top), and CD4+ and CD8+ T cells (bottom). c, URT CD45+, B cell and T cell stability over time, calculated as percentage change from median frequency for each donor; individual donor data in grey (n = 20 donors). d, Representative flow cytometry of URT CD4+ and CD8+ TRM cells, and putative BRM cell populations; stability of these populations over time as percentage change from median; individual donor data in grey (n = 20 donors). e, Combined Seurat UMAP for approximately 70,000 cells from 12 NP swabs (n = 12 donors); doughnut plot shows representative proportions of each cell type by scRNA-seq/CITE-seq. f, T cell subset UMAP generated from the approximately 19,000 T cells in e; T cell subset identities assigned by gene expression profiling and CITE-seq. g, Dotplot of select cluster-defining genes for T cell subsets in f. Protein markers (CITE-Seq) in bold with ‘Prot’. h, GSEA analysis of T cell subsets in f. i,j, Dotplots of select genes from subset analysis of TFH cell (i) and Treg cell (j) dominated clusters from f. k,l, SARS2-specific T cells in NP swabs determined experimentally by AIM assay (spike (S) or non-spike SARS2 antigen-specific) and personalized SARS2-specific TCR repertoire datasets for each subject (k), or identified bioinformatically using public datasets (l). m, SARS2-specific CD4+ versus CD8+ T cells, overlaid on UMAP from f. n, SARS2-specific T cell subtypes from k–m. See Extended Data Figs. 2d–g,i–k and 3 for further details and stratifications. abs(NES), absolute value of the normalized enrichment score; CTL, cytotoxic T lymphocyte; DC, dendritic cell; mono/mac, monocyte/macrophage; mTFH, mantle TFH; NK, natural killer; ns, not significant; Padj, adjusted P value; Teff, effector T.
To better characterize URT immune cell populations, we performed single-cell RNA sequencing (scRNA-seq) together with cellular indexing of transcriptomes and epitopes by sequencing (CITE-seq) using a nine antibody CITE-seq panel on single-cell suspensions of NP swabs from 12 adult donors (Supplementary Table 1). Numerous epithelial cell (Extended Data Fig. 1d,e) and immune cell (Fig. 1e) populations were identified by single-cell transcriptome profiling, and were conserved across donors (Extended Data Fig. 1f). Approximately one-half of all NP cells identified were B cells, 28% were T cells, 15% were epithelial cells and the remainder (less than 10%) other immune cells (Fig. 1e), with Seurat analysis confirming high concordance in cell identities by gene expression analysis and CITE-seq antibody binding (Extended Data Fig. 1g,h). B cells and T cells were further clustered independently, and distinct cell types were defined on the basis of differential gene expression, gene set enrichment analysis (GSEA)21–23 and B cell receptor (BCR) and T cell receptor (TCR) VDJ sequencing analysis. Several B cell populations were identified, including memory B (Bmem) cells, antibody-secreting cells (ASCs), BRM cells and others (Extended Data Fig. 2a,b), discussed in more detail later. Numerous distinct CD4+ and CD8+ T cell populations were identified and well represented, including CD4+ TRM cells, CD8+ TRM cells, regulatory T (Treg) cells, mucosal-associated invariant T (MAIT) cells and follicular helper T (TFH) cells (Fig. 1f–h and Extended Data Fig. 2c–g). Two CD8+ TRM cell populations were identified, with similar transcriptional patterns to CD103+ and CD103−CD8+ TRM cells observed in the human intestine24 (Fig. 1h). Compared with CD8+ TRM cells, CD4+ TRM cells are less well defined in human tissues; however, a distinct CD4+ TRM cell cluster was identified that was enriched for tissue residency markers and chemokine receptors, including CD69, ITGAE (CD103), CCR6 and CXCR6 (Fig. 1h). The CD4+ TRM cell cluster was also enriched for KLRB1 and IL7R (Fig. 1h). Higher resolution analysis of several clusters revealed more populations of interest. Closer examination of the ‘naive CD4+ T cell’ cluster (CD4 TN) showed an overlapping CD4+ memory T cell cluster distinguished by FAS expression (FAS+CD4 Tmem) (Fig. 1f and Extended Data Fig. 2e). Within the TFH cell cluster, it was possible to identify GC-TFH cells, GC-TFH cells with evidence of type I IFN signalling (GC-TFHIFN) and mantle TFH cells (Fig. 1i and Extended Data Fig. 2f)25. More granular examination of the Treg cell dominated cluster revealed follicular regu latory T (TFR) cells, TFH cells and T helper 17 (TH17) cells (expressing IL17a and IL17f but not FOXP3) in addition to Treg cells (Fig. 1j and Extended Data Fig. 2g). Overall, a rich diversity of CD4+ and CD8+ T cell populations were identified in the human upper airway.
It is important to better understand pathogen-specific immune memory in human tissues that are common sites of infection. Single-cell TCR sequencing (scTCR-seq) was completed on all 12 NP swab samples (Fig. 1e). Among these donors, all reported primary COVID-19 vaccinations and seven had evidence of a SARS-CoV-2 (SARS2) breakthrough infection (Supplementary Table 1). We used a combination of experimental and computational methods to query whether SARS2-specific CD4+ and CD8+ T cells were present in NP tissue1 (Fig. 1k–n). SARS2 spike-specific T cell clonotypes were defined experimentally from peripheral blood mononuclear cells (PBMCs) by an activation induced marker (AIM) assay antigen-specific CD4+ T cell and CD8+ T cell sorting and scRNA-seq pipeline (Extended Data Figs. 2h and 3), generating personalized SARS2-specific TCR repertoire reference datasets for each subject, specific for SARS2 spike and non-spike epitopes. Upper airway SARS2 spike- and non-spike-specific memory T cells were then identified by matching TCRs with 100% CDR3 amino acid sequence identity between NP swabs and the PBMC-derived personalized SARS2-specific TCR repertoire datasets for each subject (AIM+; Fig. 1k). Additional SARS2-specific CD4+ T cells were identified in swab samples by matches to a TCRα sequence motif known to recognize an immune-dominant HLA–DPB1*04 S167–180 epitope26 (public TCRA; Fig. 1l) or TCRs with 100% amino acid sequence identity to TCRs in select databases27,28 and papers29,30 (public ref; Fig. 1l). Altogether, 250 SARS2-specific CD4+ and CD8+ T cells were identified in the NP swabs of these 12 subjects (Fig. 1m,n). SARS2-specific CD4+ and CD8+ T cells were identified in the upper airways of all seven subjects who had recent SARS2 infections (Extended Data Fig. 2k and Supplementary Table 1). These included both SARS2 spike and non-spike epitope specificities. Upper airway SARS2-specific CD8+ TRM cells were found in most of the previously infected individuals and represented a plurality of the SARS2-specific T cells present (Fig. 1n and Extended Data Fig. 2k). SARS2-specific CD4+ TFH cells and TRM cells were also identified in upper airway tissue of most subjects (Fig. 1n and Extended Data Fig. 2k). Several other types of differentiated SARS2-specific CD4+ T cells were identified in upper airway samples from previously infected individuals, including central memory T (TCM) cells, Treg cells and TH17 cells (Fig. 1n and Extended Data Fig. 2g). A small number of spike-specific CD4+ or CD8+ T cells were identified in subjects who only had evidence of COVID-19 vaccination or had evidence of a remote subclinical SARS2 infection (Extended Data Fig. 2k and Supplementary Table 1). In sum, these experiments demonstrate both virus-specific memory CD4+ TRM cells and CD8+ TRM cells in the human upper airway after an infection. The experiments further demonstrate diverse differentiated T cell types in the upper airway in these same individuals to the same antigens. Last, these experiments demonstrate antigen-specific T cell clones with identical TCRs located in both peripheral blood and as TRM cells in the human upper airway.
Unexpectedly, B cells with surface phenotypes consistent with GC B (BGC) cells (CD19+CD20+CD38+) were present in NP swab samples from many individuals according to flow cytometry (Fig. 2a and Extended Data Fig. 4a). GC-TFH CD4+ T cells were identified in the same individuals by flow cytometry (CD4+CXCR5+PD-1hi) (Fig. 2a and Extended Data Fig. 4a). BGC cells and GC-TFH cells were also validated by intracellular staining for BCL6 (Extended Data Fig. 4b). Collection of BGC cells and GC-TFH cells by NP swabs was confirmed by scRNA-seq (Fig. 1i and Extended Data Fig. 2f for GC-TFH cells; Fig. 2b–d and Extended Data Fig. 2a for BGC cells), including SARS2-specific GC-TFH cells (Extended Data Fig. 2f). Among SARS2-specific TFH cells, several subpopulations were observed (Extended Data Fig. 2f). For subjects with recent SARS2 infection, TFH cells with spike and non-spike specificities were observed (Extended Data Fig. 2i), and were the most abundant SARS2-specific CD4+ T cell subset overall (Fig. 1n and Extended Data Fig. 2k). SARS2-specific TFH cells were found exclusively in subjects with breakthrough infections (Extended Data Fig. 2i–k and Supplementary Table 1). More granular BGC cell scRNA-seq analyses identified dark zone (‘DZ’; for example, AICDA, CXCR4, MKI67) and light zone/intermediate (‘LZ/Int’; for example, LMO2, CD83) differentiation states (Fig. 2b,c and Extended Data Figs. 2a and 4c,d; GSEA, Fig. 2d). Finally, cell cycle of the BGC cells was assessed and demonstrated to match canonical BGC cell-cycle states (Extended Data Fig. 4c). Reproducible sampling of GC lymphocytes by NP swabs indicated that swabbing may sample immune cells derived from both URT lymphoid tissue and mucosal epithelium.
Fig. 2 |. Upper airway mucosal BGC cells and GC-TFH cells consistently sampled by NP swabs.
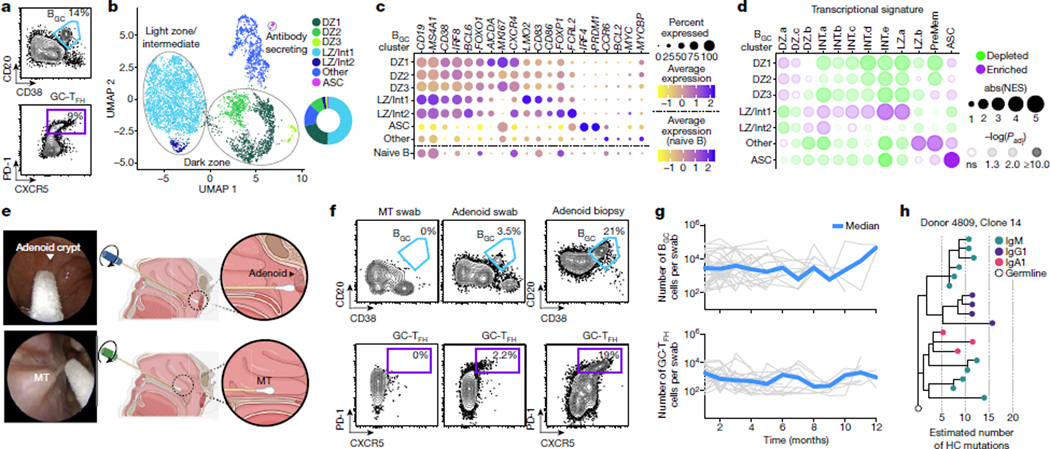
a, Representative flow cytometry plots of GC populations from NP swabs (BGC: CD20+CD38+ of CD19+ B cells and GC-TFH: CXCR5+PD-1hi of CD4+ T cells). b, BGC cell subset UMAP of approximately 5,500 cells from the BGC cell clusters shown in Extended Data Fig. 2a, demonstrating distinct dark zone (DZ) and light zone (LZ)/intermediate (Int) states. c,d, Characterization of BGC cells from NP swabs by scRNA-seq transcriptomic profiling (c) and GSEA (d). e, Schematics and representative images of endoscopic-guided MT and NP (adenoid) swab sampling from the same donor. f, Immunophenotyping from paired swabs collected by endoscopy compared with control adenoid tissue (from a different donor). g, Stability of URT GC populations (BGC cells and GC-TFH cells) from NP swabs over time by flow cytometry (data shown for NP swabs from original longitudinal cohort donors with BGC cell counts greater than 100; n = 15 donors). h, Clonal lineage of a BGC cell from an NP swab, represented as a phylogenic tree. Each tip of the tree represents a unique heavy chain (HC) sequence determined by its transcriptomic profile; coloured by heavy chain constant region gene (isotype). Grey dotted lines indicate intervals of five estimated heavy chain nucleotide mutations. Panel e was created in part using BioRender (https://Biorender.com).
Adenoids are a secondary lymphoid organ located at the posterior NP wall. Unlike encapsulated lymph nodes, adenoids have a single layer of fenestrated epithelium with channel-like invaginations known as crypts. On the basis of the swab results, we considered that adenoid tissue with functional GCs may be present in many adults and that NP swabs may directly sample GC lymphocytes from adenoid tissue through crypts. To test this suggestion, we pursued several experimental approaches. First, using nasal endoscopy to directly visualize the anatomic site being sampled, we separately collected swab samples from the nasal cavity epithelium approximately at the midpoint of the inferior nasal turbinate (MT) and from the adenoids of healthy adult volunteers (Fig. 2e and Supplementary Videos 1–3). Spectral flow cytometry immunophenotyping of samples collected by endoscopic-guidance showed that BGC cells and GC-TFH cells were found exclusively in adenoid swabs (Fig. 2f). Cells from an adenoid tissue biopsy were examined as a comparator. Longitudinal NP swab sampling of numerous individuals with consistently detectable adenoids by flow cytometry demonstrated that GC cells were sampled reproducibly, and relatively stable over time (Fig. 2g). Furthermore, substantial somatic hypermutation (SHM) was observed in BGC cells in clonal lineages collected by swabs (Fig. 2h and Extended Data Fig. 4e).
Paired NP posterior wall and MT swab samples were next collected from 38 adult donors for high parameter spectral cytometry immunophenotyping exploring immune memory cell types. Highly divergent frequencies of immune cell types existed between the two URT anatomic sites (Fig. 3a–q,u and Extended Data Fig. 5a–h). Compared with adenoids, turbinate epithelium possessed a significantly lower CD4+:CD8+ T cell ratio (P < 0.0001, Fig. 3g), few naive B cells (P < 0.0001; Extended Data Fig. 5b), few naive T cells (P < 0.0001; Extended Data Fig. 5c,d) and a higher frequency of Treg cells (P < 0.001; Fig. 3u and Extended Data Fig. 5a,e). BGC cells and GC-TFH cells were found exclusively in NP tissue (P < 0.0001 for both; Fig. 3h,j). Among memory lymphocytes, turbinate epithelium possessed higher frequencies of CD8+ TRM cells (CD69+CD103+; 95% versus 78%; P < 0.001; Extended Data Fig. 5f), most of which were also CD49a+ (Fig. 3m and Extended Data Fig. 5a). CD4+ TRM cells (Fig. 3l) and putative BRM cells (Fig. 3k) were detected in turbinate epithelium in addition to NP tissue. Many of these cell types involved in immune memory were not detected in blood (Extended Data Fig. 5i). Notably, Bmem cells in turbinate epithelium (including BRM cells) were significantly more likely to be IgA+ than Bmem cells in NP tissue or blood (P < 0.001 and P < 0.0001, respectively; Fig. 3o–q). Differential class-switch usage was also seen between NP BGC cells and Bmem cells (Fig. 3r–t). In sum, immune memory populations were substantially different between blood and two URT sites, with samples from upper airway tissue sites separated by centimetres demonstrating distinct local immune memory in healthy adults.
Fig. 3 |. Nasal cavity and nasopharynx mucosa possess distinct adaptive immune cell populations.
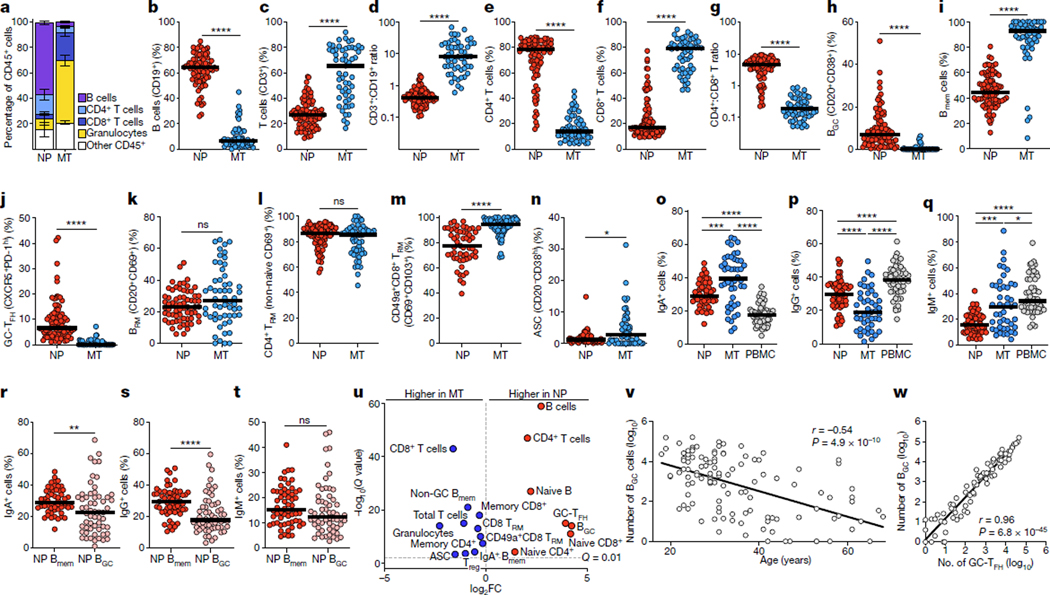
a, Parts of a whole graph demonstrating average percentages of principal immune cell populations in MT versus NP swabs (n = 38 donors). Error bars, s.e.m. b–n, Differences in immune cell frequencies, ratios and subsets in MT versus NP swabs by flow cytometry (n = 88 NP and 55 MT swabs; 38 donors); see Extended Data Fig. 4a for gating strategy: percentage of total CD19+ B cells and CD3+ T cells (of live CD45+) (b,c); ratio of B cells to T cells (b:c) (d); percentage of CD4+ (e) and CD8+ T cells (f); ratio of CD4+ to CD8+ T cells (e:f) (g); percentage of BGC cells (h), Bmem cells (CD20+CD38−IgD−) (i) and BRM cells (CD20+CD69+) (k) of total B cells; percentage GC-TFH cells of total T cells (j); percentage CD4+ TRM cells of non-naive CD4+ T cells (l); percentage CD49a+ of CD69+CD103+CD8+ TRM cells (m); percentage of antibody-secreting cells (CD20−CD38hi) of total B cells (n). o–q, Differences in B cell isotype profiles in MT and NP swabs and PBMCs (frequency of IgA+ (o), IgG+ (p) and IgM+ (q)), n = 58 NP and 46 MT swab, 56 PBMC samples; 22 donors. r–t, Differences in isotype profiles of NP Bmem cells versus BGC cells (IgA+ (r), IgG+ (s) and IgM+ (t)), n = 58 NP swabs; 22 donors. Bars in b–t, median (pairwise comparisons by Mann–Whitney test); *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001, ****P ≤ 0.0001. u, Volcano plot summarizing significant differences in NP versus MT populations. v, Correlation between number of BGC cells sampled per NP swab and age across healthy adults; Pearson r and P values (n = 117 donors; one swab per donor). w, Correlation between number of BGC cells and GC-TFH cells sampled per NP swab (n = 80 donors; one swab per donor); Pearson r and P values.
With the ability to non-invasively sample adenoid lymphoid tissue from healthy adult donors, it was possible to address the question of how long immunologically functional adenoid tissue lasts during adulthood, as this relates to the capacity for new generation of immune memory. NP swabs collected from more than 100 healthy adult subjects aged 18–68 years were used to determine adenoid GC activity across a large age spectrum. Seventy percent of the adult donors had detectable adenoid BGC cells. BGC cells per swab decreased with age, with approximately log-linear progression (Pearson r = −0.54, P = 4.9 × 10−10; Fig. 3v). Naive B cells per swab correlated with BGC cells (r = 0.71, P = 7.4 × 10−37; Extended Data Fig. 5k,n,o), indicating that BGC cell detection was both an indicator of the presence of mucosal lymphoid tissue and the functionality of B cell responses in that tissue. GC-TFH cell numbers decreased commensurately with age (r = −0.58, 6.1 × 10−12; Extended Data Fig. 5l–m). GC-TFH cell frequencies in NP swabs correlated tightly with BGC cells (r = 0.96, P = 6.8 × 10−45; Fig. 3w). BGC cells were observed in individuals as old as 68 years. Together, the data indicate that, whereas adenoid tissue atrophies with ageing, most healthy adults possess functional adenoid tissue for GC responses.
Adults with and without a history of recent SARS2 infection were enroled in a longitudinal study with monthly sample collections to better understand both functional attributes and durability of human URT B cell memory (Supplementary Table 1). Using fluorescently conjugated SARS2 antigen probes plus multiparametric spectral flow cytometry, upper airway SARS2-specific Bmem cells were detected in NP tissue in 100% of subjects and in MT tissue in 31% of subjects following breakthrough infection (Fig. 4a–c and Extended Data Fig. 6a,b). SARS2-specific Bmem cells in NP samples were preferentially IgA+ compared with blood (20% versus 5.9% for nucleocapsid, P = 0.0004; 9.1% versus 3.1% for spike, P < 0.0001; Fig. 4a,b). IgA class switching was even more pronounced in the turbinate epithelium, with around 60% of nucleocapsid-specific Bmem cells and around 30% of spike-specific Bmem cells being IgA+ in that tissue (Fig. 4a). Notably, SARS2 nucleocapsid-, receptor binding domain (RBD)-, and spike-specific Bmem cells were maintained in the upper airway (NP) for at least 6 months, and included both IgA and IgG isotypes (Fig. 4c and Extended Data Fig. 6c). Thus, both IgA+ and IgG+ virus-specific Bmem cells can be durable in the URT for more than 6 months.
Fig. 4 |. SARS2-specific upper airway mucosal B cells.
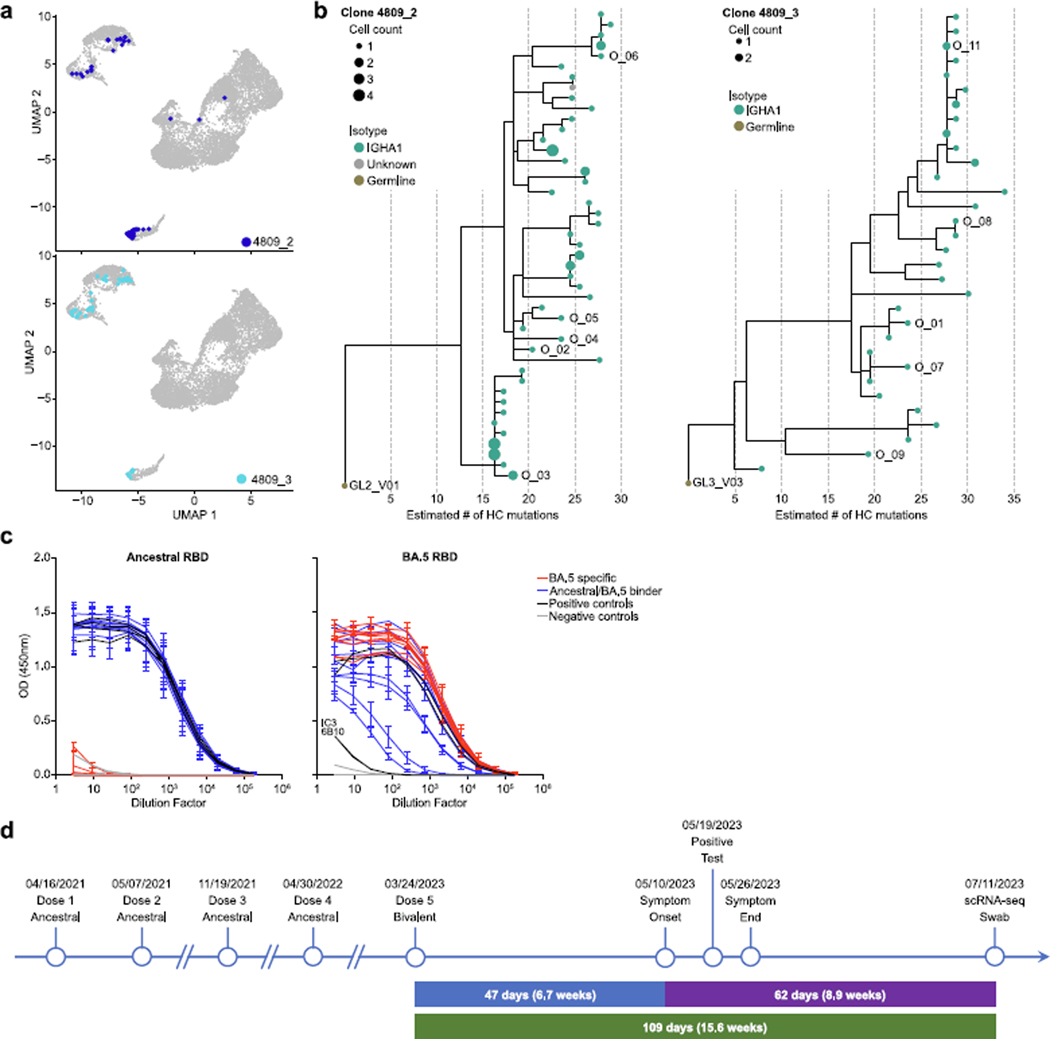
a,b, Frequencies of SARS2 nucleocapsid (N)-, RBD-, and spike-specific IgA+ (a) and IgG+ Bmem cells (b) in NP and MT swabs and PBMCs. Bars, median. Pairwise comparisons by Mann–Whitney; *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001, ****P ≤ 0.0001. c, Longitudinal probe staining of SARS2 nucleocapsid-, RBD- and spike-specific NP Bmem cells by flow cytometry (for gating strategy, see Extended Data Fig. 6a; n = 29–47 donors). Grey lines, data for individual donors. Dotted line, limit of detection. d, Seurat UMAP, 9,462 NP B cells from breakthrough donors 4202, 4809 and 4379 (one NP swab per donor) coloured by B cell subsets. e, Expression of genes of interest for cells in d. f,g, SHM (f) and class-switch recombination (CSR) (g) (isotype) analysis of cells in d. h, SARS2 RBD-, spike- and nucleocapsid-specific B cells overlaid on UMAP from d. i, Phenotypes of SARS2-specific B cells. Two cells that were class switched (IgG1, IgA1) with moderate-to-high SHM were reclassified from naive to class switched resting Bmem cells (Resting BCSR). j, Isotypes (left) and heavy chain (HC) SHM (right) for early and late BPC. k, SARS2 RBD-, spike-specific BPC. l, Phylogenic trees representing Omicron-specific B cell clonal lineages from donor 4809. Tip sizes, number of cells with identical features; colours, cell type; labels, validated antibodies and KD by SPR; *KD for BA.4/5 spike. m, Summary of ELISA, PSV neutralization and SPR data for ancestral (D614G), Omicron BA.4/5 spike, RBD for antibodies in l. B, binding of ancestral and Omicron; O, Omicron-specific. Grey, undetectable binding/neutralization. n, Comparison of heavy chain mutations for donor 4809 RBD- and nucleocapsid-specific BGC. Student’s t-test, ****P ≤ 0.0001.
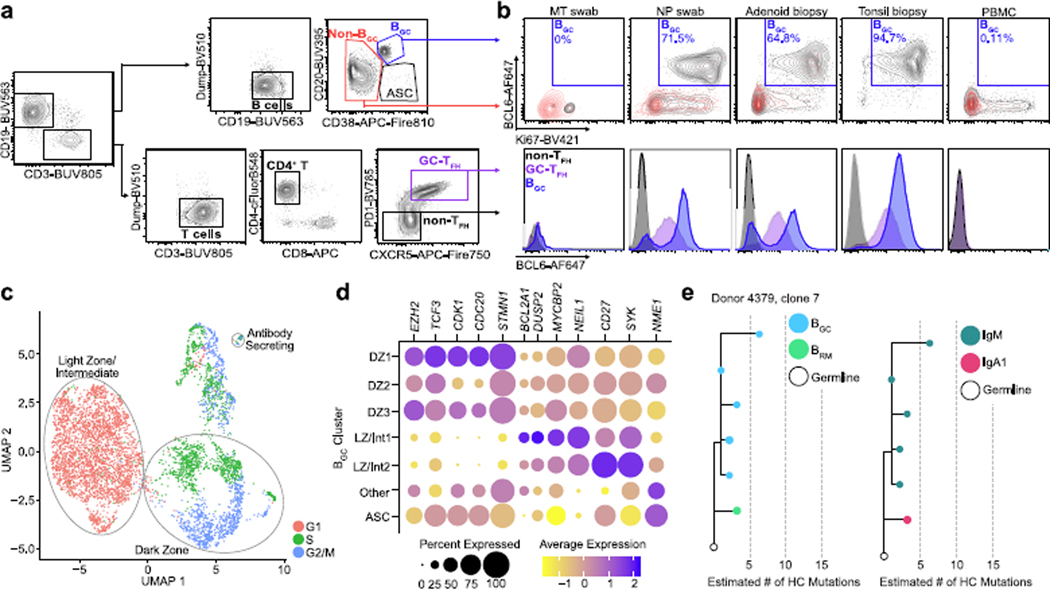
Paired PBMC samples and NP swabs were collected from three further subjects with recent breakthrough infection for scRNA-seq, CITE-seq and single-cell BCR sequencing (scBCR-seq), combined with oligonucleotide-barcoded, fluorescently conjugated SARS2 spike, RBD and nucleocapsid probes (Methods). Data were obtained for more than 9,400 total NP swab B cells. Twelve distinct B cell subtypes or states were identified among the URT B cells, including BGC cells, ASCs and BRM cells (Fig. 4d,e). Several URT Bmem cell clusters were delineated from each other by differences in class-switch recombination and SHM, following a continuum from low to high SHM and IgG class-switch recombination (Fig. 4f,g). Putative URT BRM cells had the highest proportion of IgA isotypes, with IgA1+ cells comprising nearly 50% of BRM cells (Fig. 4g).
Using an experimental and computational pipeline combining (1) SARS2 antigen probe B cell binding data (n = 102 cells), (2) personalized SARS2-specific BCR reference datasets from matched peripheral blood samples for each subject (n = 7 cells), and (3) BCR amino acid sequences identified computationally by matches in public SARS2 antibody datasets (n = 37 cells, with specificity calls revised on the basis of experimentally validated recombinant antibodies) (Extended Data Fig. 6e,f; Methods), we identified 146 SARS2-specific B cells in NP samples (Fig. 4h). RBD, spike non-RBD, and nucleocapsid protein specificities were all identified. Notably, SARS2-specific B cells identified in upper airway swabs included Bmem cell, BGC cell, BRM cell and ASC populations (Fig. 4i), demonstrating rich complexity of virus-specific B cells in the adult upper airway.
There is recent evidence in animal models of the importance of BRM cells in protective immunity at barrier tissues31–34, and thus analysis of antigen-specific putative human URT BRM cells was of particular interest, given the limited knowledge of this immune memory cell type. URT BRM cell SHM equalled the highest SHM amounts of any Bmem cell cluster (Fig. 4f). Among SARS2-specific Bmem cells, both putative BRM cells and conventional Bmem cells were observed (Fig. 4i). The putative URT BRM cell cluster had a distinct gene signature, including upregulation of FCRL4, FCRL5, CCR6, CCR1, ZEB2 and SOX5 (Fig. 4e), that is, genes previously reported in tissue-associated Bmem cells in palatine tonsils and atypical Bmem (aBmem) cells in other tissues19,31,35–37. Expression of TBX21 (T-bet)—a transcription factor seen in many circulating aBmem cells38,39—was absent in URT BRM cells (Extended Data Fig. 6d). In sum, URT BRM cells have distinctive features and both SARS2-specific IgA+ and IgA− BRM cells can be found in the URT of previously infected individuals.
ASCs comprised a distinct B cell cluster with high expression of XBP1, IRF4, PRDM1, CD27 and SDC1 (CD138) (Fig. 4e). To investigate whether this ASC cluster was composed entirely of short-lived plasmablasts, which would not constitute memory, we analysed the NP ASC cluster by comparison with published transcriptomic data of human bone marrow plasmablasts and plasma cells (BPC)40 (Fig. 4j and Extended Data Fig. 6g). There was minimal evidence of URT plasmablasts, on the basis of expression of MKI67 or other genes (Extended Data Fig. 6g). Instead, the ASCs exhibited gene expression profiles consistent with human bone marrow early and late BPC (Extended Data Fig. 6g). Compared with early BPC, the late BPC cluster had increased class switching and SHM (Fig. 4j). SARS2-specific cells were found as both early and late BPC (Fig. 4k and Extended Data Fig. 6h). In sum, mature SARS2-specific BPC were found locally in the URT, with transcriptional profiles of long-lived BPC.
We identified two extensive spike-specific B cell clonal lineages that included several BPC that were highly related to BGC cell clones (Fig. 4l), indicating both active local maturation of SARS2 Bmem cells and a potential reservoir for local long-lived antibody production in the human nasopharynx. These clonal families consisted primarily of BGC cells and BPC, with further differentiation into BRM cells and other Bmem cell subtypes (Fig. 4l and Extended Data Fig. 7a). These spike-specific B cell clonal family members were class switched ubiquitously to IgA1 (Extended Data Fig. 7b). Most cells had heavy chain SHM counts ranging from 15 to 30 mutations (Extended Data Fig. 7b). To further investigate these clonal families from donor 4809, we synthesized antibodies distributed across each phylogeny. Many antibodies did not bind ancestral proteins or neutralize ancestral virus appreciably but bound BA.4/5 proteins and neutralized BA.4/5 virus strongly, indicating an Omicron-primed B cell response (Fig. 4m and Extended Data Fig. 7c). Antibody affinities of clonal lineage 3 to BA.4/5 RBD ranged from 0.11 to 36 nM, with 3.3- to 8.5-fold higher affinity for local BPC (O_01) and Bmem cell (O_08) clones compared with the nearest neighbour to the inferred germline (O_10) (Fig. 4l,m), indicating local affinity maturation. Antibodies based on inferred germline BCRs for both clonal lineage 2 and 3 (GL2_V01 and GL3_V03) bound weakly to BA.4/5 RBD or spike, providing further evidence of local affinity maturation (Fig. 4l,m). This subject had received a bivalent COVID vaccine (containing both ancestral and BA.4/5 spike) before Omicron breakthrough infection and sample collection (Extended Data Fig. 7d). Analysis of BGC cell SHM across SARS2 antigen-specificities in donor 4809 revealed a striking dichotomy in the number of heavy chain mutations between nucleocapsid-specific and RBD-specific BGC cells (median 7.5 versus 20, P < 0.0001; Fig. 4n). The significantly lower number of heavy chain mutations for nucleocapsid-specific BGC cells provides evidence for local priming of B cells in response to SARS2 breakthrough infection in the adult URT. Together, a parsimonious explanation of the RBD-specific B cell data is priming and differentiation of Omicron RBD-specific Bmem cells after bivalent vaccine administration, followed by recall of Bmem cells to the URT upon breakthrough infection to initiate secondary GCs in the adenoid, whereas the nucleocapsid-specific B cell responses were primed locally in the URT. In summary, complex local B cell responses can occur in the URT in response to viral infection, including local priming, recall, class-switch recombination and affinity maturation, and develop complex populations of local immune memory, including BPC and BRM cells.
Discussion
The URT is the primary site of infection for many human pathogens, but there has been limited knowledge available on URT immune memory cellular content, durability, differentiation states or anatomical complexity in adult humans. Here, using more than 315 total swabs from over 100 donors, comprised of several cohorts of adults in a normal physiologic state or following recent acute viral upper airway infections, with sampling of NP posterior wall and nasal turbinate epithelium sites, combining diverse experimental techniques, we provide new insights into human upper airway immune memory and provide rich evidence of complexities of normal upper airway immunology. CD8+ TRM cells, CD4+ TRM cells and putative BRM cell populations were defined, and total proportions of these memory populations were stable over time for more than 1 year in healthy adults. Active adenoids with GCs were demonstrated in most healthy adults, with age-dependent declines in adenoid immunobiology observed over a 50-year span. Upper airway Bmem cells and BRM cells were shown to be preferentially IgA+, with highest amounts of SHM in BRM cells. Evidence of BPC longevity was demonstrated, which may represent a reservoir of long-lived mucosal immunity in the human URT. Complex populations of SARS2-specific T cells were demonstrated41, with numerous differences in T cell and B cell memory between adjacent lymphoid and mucosal epithelial tissue sites and many differences between the upper airway and blood. Shared CD4+ and CD8+ T cell clones were shown between the upper airway and blood, also demonstrating interactions between these compartments. Antigen-specific URT CD8+ TRM cells, CD4+ TRM cells and BRM cells were all demonstrated. SARS2-specific Bmem cells were demonstrated in both NP and MT sites, enriched for IgA+ cells, and durability of antigen-specific IgA+ and IgG+ Bmem cells for more than 6 months was shown. Local SARS2-specific GC responses were also shown. Finally, the evidence indicated local B cell priming, class switching, affinity maturation and differentiation into several immune cell types including Bmem cells, BRM cells and BPC with a transcriptional signature of longevity.
In sum, these findings highlight the presence and maintenance of diverse antigen-specific B cell and T cell subtypes that contribute to human upper airway immune memory and may contribute to protection against respiratory pathogens. Finally, these experimental approaches are applicable to diverse studies of human upper airway health and disease.
Online content
Any methods, additional references, Nature Portfolio reporting summaries, source data, extended data, supplementary information, acknowledgements, peer review information; details of author contributions and competing interests; and statements of data and code availability are available at https://doi.org/10.1038/s41586-024-07748-8.
Methods
Study populations
Healthy adults aged 18 years or older were eligible to participate in the study. Sample sizes were not predetermined by statistical methods. Randomization and blinding were not performed. The La Jolla Institute for Immunology (LJI) institutional review board provided oversight for all human subject research performed in this study, including all sample and clinical data collection. Further institutional review board oversight for tonsillar and adenoid tissue collection was provided by the University of California San Diego and Rady Children’s Hospital. All individuals provided written informed consent and assent for participation. Consent was provided to display and publish de-identified photographic images and videos. Please see Supplementary Table 1 for further details about the participants in the study cohorts.
Sample collection and processing
Swab (NP posterior wall/adenoid and MT) and blood samples were obtained using standardized collection protocols and processed the same day as collection. Participants were asked to sit in a chair designed for phlebotomy, to blow their nose and to close their eyes before swab sample collection to help with relaxation and ease of sampling. A registered nurse, doctor or other qualified and trained provider performed NP swab sampling. A flocked swab (Puritan) was inserted gently into one nostril, along the floor of the nasal cavity (parallel to the hard palate), until the posterior NP was reached (around 8–9 cm maximum insertion distance), followed by approximately five circular rotations of the swab while in the posterior NP. MT swabs were collected in a similar fashion, but by inserting the swab approximately half the depth as for an NP swab (around 3–4 cm maximum insertion distance) into the nasal cavity, then rotating the swab in a circular motion to sample the inferior turbinate mucosa. We performed swab sampling bilaterally for both NP and MT swabs, using the same flocked swab to sample both sides before placing the swab in a collection tube containing RPMI medium (Gibco) supplemented with 10% fetal bovine serum. One NP and/or MT swab was collected per donor per session. If NP and MT swab samples were paired, MT samples were collected first. In the case of endoscopic swab sampling, a licensed and board-certified rhinologist performed the endoscopic sample collection using standard technique to perform an endoscopic examination of the nasal cavity and NP with a rigid nasal endoscope (Stryker), followed by endoscopic swab sample collection and photo/video documentation of the sampling site. After collection, swabs were vortexed briefly in the capped collection tube to release cells from the swab. More medium was then used to rinse adherent cells from the swab by placing the swab in a 40 μm mesh filter over a 50 ml conical tube and applying medium on top of the swab. The filtered cell suspension was then centrifuged at 500g for 7 min at 4 °C. Swab cells were then processed for downstream applications.
PBMCs and plasma were isolated from blood using standard protocols, and cryopreserved until ready for use42,43. Surgically resected, deidentified human adenoid tissue was collected using standard protocols, and processed the same day as collection to yield single-cell suspensions, as previously described44. These adenoid cells were cryopreserved and stored in liquid nitrogen and thawed in the same fashion as cryopreserved PBMCs when ready for use.
Immunophenotyping of URT cell populations by flow cytometry
For flow cytometry, samples were sequentially stained with Fc block (BioLegend), viability dye, SARS2 probes (45–60 min at 4 °C) and/or antibody (30–60 min at 4 °C) before data acquisition with a 5-laser Cytek Aurora spectral analyser. For details about flow cytometry reagents/Ab panels, please see the Supplementary Information.
Detection of antigen-specific B cell subsets by flow cytometry/fluorescence-activated cell sorting
Biotinylated SARS-CoV-2 proteins were multimerized individually using fluorescently labelled streptavidin at 4 °C for 1 h. Full-length SARS-CoV-2 spike (AcroBiosystems, catalogue no. SPN-C82E9–200ug) and RBD (AcroBiosystems, catalogue no. SPD-C82E9–200ug) were used. Biotinylated spike was mixed with streptavidin BUV737 (BD, catalogue no. 612775) and streptavidin Alexa Fluor 647 (ThermoFisher Scientific, catalogue no. S21374) at a 20:1 ratio (around 4:1 molar ratio). Biotinylated RBD was mixed with streptavidin BV605 (BioLegend, catalogue no. 405229) and BV711 (BioLegend, catalogue no. 405241) at a 2.2:1 ratio (around 4:1 molar ratio). Biotinylated SARS-CoV-2 full-length nucleocapsid (Sino Biological, catalogue no. 40588-V27B-B) was multimerized using streptavidin BV421 (BioLegend, catalogue no. 405226) and streptavidin BB515 (BD, catalogue no. 564453) at a 5.5:1 ratio (around 6:1 molar ratio)42. Individual tubes of the antigen probes were prepared as described and were then mixed in Brilliant Buffer (BD Bioscience, catalogue no. 566349) containing 5 μM free d-biotin (Avidity, catalogue no. Bir500A). Samples were prestained using LIVE/DEAD Fixable Blue Stain Kit (ThermoFisher Scientific, catalogue no. L34962) and human Fc block (BioLegend catalogue no. 422302) in DPBS buffer at room temperature for 15 min. PBMCs and MT swabs were stained with 50 μl of antigen probe cocktail containing 100 ng of spike per probe (total 200 ng), 27.5 ng of RBD, 40 ng of nucleocapsid per probe (total 80 ng) and 5 μM free d-biotin at 4 °C for 1 h to ensure maximal staining quality before surface staining with antibodies as listed in Supplementary Table 2 in Brilliant Buffer at 4 °C for 30 min. NP swabs were stained with 100 μl of the same probe master mix and surface antibodies. PBMCs were counted and stained in an appropriate volume of probe master mix and surface antibodies for fluorescence-activated cell sorting (FACS) in preparation for scRNA-seq. For PBMC sorting by FACS for scRNA-seq, one of the streptavidins for each biotinylated target (RBD, spike, nucleocapsid) was replaced with a custom streptavidin dual conjugated with a fluorophore (typically BV421; BioLegend) and a unique oligonucleotide hashtag (BioLegend TotalSeq) such that RBD, spike and nucleocapsid were each assigned one unique fluorophore and one unique oligonucleotide hashtag and cells could be sorted as shown in Extended Data Fig. 5a. For flow cytometric analysis, stained samples were acquired on a 5-laser Cytek Aurora (with SpectroFlo software v.2 or higher) and analysed using FlowJo10 (BD Bioscience). For FACS, PBMCs were stained and sorted as described below.
The limit of detection was set at a minimum of three probe double-positive cells for calling a sample positive for SARS2 nucleocapsid-, RBD- or spike-specific Bmem cells and five probe double-positive cells for determining the isotype frequencies of SARS2 nucleocapsid-, RBD- or spike-specific Bmem cells. For Fig. 4a,b and Extended Data Fig. 5b, only data points above the limit of detection are shown. For any given donor’s samples to be included in longitudinal analyses (Fig. 4c and Extended Data Fig. 5c), they had to have detectable SARS2-specific Bmem cells for at least 2 consecutive months. Please see Supplementary Table 1 for further details of sample sizes.
AIM assay
PBMCs were isolated from peripheral blood samples drawn on the same day as swab collection. As previously described42,43,45, PBMCs were plated in 96-well plates with 1 × 106 PBMCs per well, and cultured for 24 h at 37 °C in an incubator with 5% CO2 in the presence of negative control (equimolar amount of DMSO vehicle), positive control (Staphylococcal enterotoxin B at 1 μg ml−1) or SARS-CoV-2 peptide megapools (1 μg ml−1 per megapool)43,46,47 containing spike or non-spike epitopes (CD4-RE or CD8-RE dominant and subdominant megapools). Before stimulation, PBMCs were incubated at 37 °C for 15 min with 0.5 μg ml−1 anti-human CD40 monoclonal blocking antibody (Miltenyi Biotec). After a 24-h incubation, the plates were centrifuged, cells were washed with PBS and then stained with LIVE/DEAD Fixable Blue (Invitrogen) 1:1,000 in PBS with Fc block (5 μl per sample; BioLegend) for 10–15 min at room temperature, washed with FACS buffer (3% FBS in DPBS without calcium or magnesium), then surface stained for 30 min at 4 °C (see Supplementary Information for details of antibody staining panels). Cells were then sorted according to surface marker expression. AIM+ gates were drawn on the basis of T cell responses to megapools relative to controls. AIM+ memory T cells that were reactive to SARS2 spike and/or non-spike megapools were isolated by FACS and subjected to scRNA-seq/scTCR-seq to generate SARS2-specific TCR repertoire reference libraries. Samples were labelled with TotalSeq-C anti-human hashtag antibody (BioLegend) for each donor and megapool stimulation condition. Samples from each donor and stimulation condition were sorted separately, with CD4+ and CD8+ T cells sorted into separate tubes. Cells were then washed, centrifuged, pooled (if appropriate) and subjected to downstream 10x processing (10x Genomics). CD4+ T cells were never pooled with CD8+ T cells.
Cell sorting
Cells were surface stained, as described in other sections, then subjected to FACS isolation using a 4-laser BD FACSAria Fusion (with FACSDiva software v.6 or higher) or 5-laser Cytek Aurora CS cell sorter (with SpectroFlo software v.2 or higher) in the LJI Flow Cytometry Core. For FACS antibody panels and other details, see the respective Methods sections and Supplementary Table 2.
Sample preparation for scRNA-seq
For scRNA-seq experiments using swabs, cell suspensions were subjected to dead cell removal (Miltenyi Dead Cell Removal Kit) before sequential staining with Fc block (10 min at room temperature; BioLegend), and then SARS2 probes and CITE-seq antibody. For CITE-seq, swab-derived cells were stained with the TotalSeq-C Human TBNK Cocktail (BioLegend) according to the manufacturer’s protocol. Most dead epithelial cells, as well as a portion of viable epithelial cells from swabs, were removed by the dead cell removal method, which also significantly enriched overall cell viability of swab samples. PBMCs were stained sequentially with Fc block and viability dye, SARS2 probes and then surface antibody for FACS. SARS2 probes for scRNA-seq were generated as described above for flow cytometry but using a custom streptavidin containing both a fluorophore and unique oligonucleotide hashtag (BioLegend TotalSeq-C) to allow for both FACS sorting of PBMC-derived probe-binding B cells using a combination of fluorophore-conjugated probes and bioinformatic identification of probe-binding B cells according to the unique oligonucleotide ‘hashtag’ present on one of the probes for each antigen. Swab cells were stained using only the hashtag-containing probe as these samples were not sorted and did not require dual probe staining for FACS. Cell suspensions from sorted PBMCs or processed swabs were prepared for Chromium Single Cell Immune Profiling using Chromium Next GEM Single Cell 5′ HT Reagent Kits (v.2), recommended supplies and equipment, according to the manufacturer’s protocol (10x Genomics). Samples were prepared for sequencing and sequenced on a NovaSeq 6000 sequencer (Illumina) in the LJI NGS Core. Please see Supplementary Information for further details.
Multimodal scRNA-seq analysis
Data were prepared for analysis in Cell Ranger (v.7.0.0 or newer; 10x Genomics) to process and align FASTQ reads from the gene expression (GEX), ADT/HTO and V(D)J sequencing libraries to the provided prebuilt GRCh38 human genome references. Analysis was then performed in R studio using R (v.4.0.0 or newer) and Seurat (v.4 for most figures; v.5 for Extended Data Fig. 5d)48. The data were filtered using the default parameters min.cells = 3, percent.mt <5, min.features = 200. The data were filtered to exclude cells with more than 4,500 unique feature counts. For Fig. 4, percent.mt was set to less than 10. Before analysis, gene expression was normalized using the ‘NormalizeData’ function in Seurat (default parameters). CITE-seq data were normalized in Seurat using centred log ratio transformation across features (margin 1). The 10,000 most highly variable features were selected and used for data scaling and dimension reduction through principal component analysis (PCA). Individual samples were then integrated using the reciprocal PCA method with default parameters except as follows: the top 50 dimensions were used to specify the neighbour search space; the top 10,000 variable features defined above were used to find anchors; two representative datasets (samples) were used as references. Following integration, data scaling and PCA dimension reduction were again performed on the complete, integrated dataset. Cells were clustered using the ‘FindNeighbours’ and ‘FindClusters’ functions. Further dimension reduction and visualization were performed with uniform manifold approximation and projection (UMAP). The ‘subset’ function was used to examine B cells (including all B cells and then B cells in the BGC cell and ASC clusters) and T cells (including all T cells, and then T cells in the TFH cell and Treg cell clusters) more closely independently. The resolution of each UMAP was adjusted to achieve optimal separation for cell cluster identification using a combination of gene expression and CITE-seq data. The resolution was set to 1.2 for the UMAP in Fig. 1e, 0.6–0.8 in Fig. 1f, 0.4 in Extended Data Fig. 2a, 0.4 in Extended Data Fig. 2e,f, 0.8 in Fig. 4d and 0.4 in Fig. 4k. We analysed differentially expressed genes (DEGs) with the ‘FindAllMarkers’ function in Seurat using a Wilcoxon Rank Sum test with a log2 fold-change threshold of 0.25, and including only genes expressed in at least 25% of cells. We performed GSEA using the Fast GSEA package v.1.24 with the fgseaMultilevel function and using default arguments49. MSigDB human gene sets were imported using the msigdbr package v.7.5.1. Several published, curated T cell21–23 and B cell37,40,50–52 gene sets were used for GSEA analysis (please see Supplementary Information for details). For Fig. 4, when curated gene sets contained more than 100 DEGs, only the top 100 upregulated and downregulated genes according to the adjusted P value and average log2 fold-change were included for GSEA. Cells mapping to naive B cell clusters that were class switched to IgA or IgG isotypes were reassigned as ‘Resting BCSR’. Similarly, a single cell in clone 4809_2 (Fig. 4l) mapped to a naive cell cluster and was reassigned as ‘Resting BCSR’ as it had 20 heavy chain mutations, no isotype call and the remaining 58 out of 59 cells in the clone all had an IgA1 isotype. Please see Supplementary Information for more details.
Single-cell TCR sequencing analysis
Cell Ranger V(D)J sequencing outputs were further processed and analysed using the Platypus R package53. The TCR data were processed initially using the ‘VDJ_GEX_matrix’ function with default parameters, and GEX data were processed separately in Seurat, as described above. We performed clonotyping using the ‘VDJ_clonotype’ function, and cells with 100% nucleotide sequence identity for paired CDR3α and CDR3β sequences were grouped as clones. MAIT were identified by a combination of GEX and scTCR-seq, and included 52 canonical (TRAV1–2, TRAJ33 expressing) MAIT cells.
Assignment of SARS2-specific T cells
A combination of computational and experimental methods was used to identify SARS2-specific T cells. We examined T cells in NP swabs by searching public databases27,28 and peer-reviewed publications29,30 for TCR sequences with 100% amino acid sequence identity to our paired TCRα and TCRβ scTCR-seq data (public reference). Putative SARS2-specific T cells were also identified by TCRs with CDR3α sequences matching an immunodominant CDR3α motif (CA[G/A/V] XNYGGSQGNLIF) that recognizes the HLA-DPB1*04 restricted spike (S167–180) epitope (spike public TCRA26). Separately, we defined SARS2 spike-specific T cells experimentally by matching TCRα and TCRβ CDR3 sequences with 100% amino acid sequence identity between NP swabs and PBMC-derived, sorted AIM+ T cells. Approximately 99% concordance between amino acid and nucleotide sequence identity was observed, demonstrating that nearly all the overlap identified between these sites represents true clonal relationships between NP and PBMC-derived TCR sequences rather than false assignment of TCR sequence overlap on the basis of convergent amino acid sequences.
Antibody-secreting cell subtype analysis
The ‘subset’ function was used to examine more closely the phenotypes of ASCs found in the B cell clusters. The resolution of the UMAP was adjusted to achieve optimal separation for cell cluster identification using GEX sequencing data. Published transcriptomic data from human ASC maturation states in bone marrow were used to define the upper airway ASCs into BPC subclusters40. DEGs identified by Duan et al.40 in the bone marrow ‘plasmablast’ and ‘early’ plasma cell clusters 1–2 were treated as a single gene set defined as ‘early’ in our analysis, whereas DEGs identified in ‘late’ cell clusters 1–4 were treated as a single gene set defined as ‘late’ in our analysis. DEGs from other cell clusters in ref. 40 were treated as discrete gene sets and retained cluster nomenclature from the original study. GSEA was used to compare our ASC data with the published DEGs, as described above.
Single-cell BCR sequencing analysis
Cell Ranger V(D)J sequencing outputs were further processed and analysed using R and Python packages from the Immcantation framework54–57. The National Center for Biotechnology Information IgBLAST v.1.21.0 (ref. 58) and the ‘AssignGenes.py’ function from Change-O v.1.3.0 were used to reannotate V(D)J genes by alignment to the ImMuno GeneTics reference database59. Cells with non-functional sequences, multiple heavy chain sequences or with unpaired heavy or light chain sequences were removed. For cells containing heavy chain paired with both κ and λ light chains, the κ chains were removed as λ chain rearrangement occurs only when rearrangement of both κ chain alleles is ineffective. We performed V(D)J clonotyping using scoper v.1.2.1 (refs. 56,57). For each donor, nucleotide sequences were first grouped by shared HC V genes, J genes and CDR3 sequence lengths. A normalized Hamming distance threshold for the CDR3 sequence repertoire was determined using the ‘findThreshold’ function in SHazaM v.1.1.2 (ref. 54) with default parameters. Nucleotide sequences differing from one another by this threshold were defined as distinct clones. Clones differing by light chain V gene, J gene or CDR3 sequence length were further subdivided to generate the final clone assignments. For each clone, consensus clonal germline V and J sequences were reconstructed using the ‘createGermlines’ function in dowser v.1.2.0 (refs. 60–62). The ‘observedMutations’ function in SHazaM was used to calculate SHM as the total number and frequency of non-ambiguous nucleotide mismatches from each cell to the V and J genes of its consensus clonal germline sequence. The SHazaM SHM outputs were plotted in R using geom_boxplot in the ggplot2 package (https://ggplot2.tidyverse.org/reference/geom_boxplot.html#summary-statistics).
B cell lineage tree construction
We constructed maximum-likelihood lineage trees for B cell clonal families using the IgPhyML model in the ‘GetTrees’ function of the R Package dowser (v.2.0)60–62. Branch lengths are indicative of the estimated cumulative number of mutations that have occurred in each heavy chain sequence since its divergence from the most recent common ancestor within the lineage. Tip sizes are indicative of the number of cells with identical features (heavy chain nucleotide sequence, isotype, cell type); tip colours are indicative of the cell type or isotype, as in the corresponding figures and figure legends.
Single-cell probe binding analysis of SARS2-specific B cells
Unnormalized probe unique molecular identifier (UMI) counts were extracted from the Seurat ADT assay for downstream analysis in R using proprietary scripts. UMI count thresholds for NP swab B cells and sorted Bmem cells were set for each probe to achieve an equivalent percentage of probe-positive cells as observed in the flow cytometry data from an equivalent timepoint from the same donors. In a minority of cells, UMI counts were above the probe-positive threshold for more than one antigen probe. In these cases, final antigen specificity was assigned according to the following criteria: (1) RBD-specific cells have RBD probe UMI counts more than the RBD probe-positive threshold and greater than twice the nucleocapsid probe UMI counts for the same cell; (2) spike (non-RBD)-specific cells have spike probe UMI counts greater than the spike probe-positive threshold, RBD probe UMI counts less than the RBD probe-positive threshold, and spike probe UMI counts more than twice the nucleocapsid probe UMI counts for the same cell; (3) nucleocapsid-specific cells have nucleocapsid probe UMI counts greater than nucleocapsid probe-positive threshold and more than twice the spike and RBD probe UMI counts for the same cell; (4) if none of the above criteria were met, the antigen specificity was assigned a ‘not applicable’ value. Antigen specificity was determined experimentally through a combination of direct probe binding to NP swab B cells and personalized references prepared as follows: (1) matched PBMCs from the same donors were stained with SARS2 antigen probes (RBD, spike, nucleocapsid) and sorted to isolate SARS2-specific Bmem cells before being subjected to scRNA-seq and analysis to generate BCR references for each donor; (2) NP swab B cells with BCRs that matched PBMC references on the basis of 70% or greater sequence identity and exact matches for heavy chain and light chain V and J genes as well as heavy chain CDR3 sequence length were considered antigen specific.
Computational identification of SARS2-specific BCRs
To identify putative SARS2-specific B cells from NP swabs, we developed a computational pipeline to match our paired heavy chain and light chain BCR sequencing data to publicly available SARS2-specific BCR and antibody sequences. We first looked for BCRs with exact matches to heavy chain V and J genes, CDRH3 amino acid sequence length and at least 70% CDRH3 amino acid sequence identity to human, SARS2-specific, reference sequences in the Coronavirus Antibody Database (CoV-AbDab)63. BCRs matching these criteria were further filtered by (1) exact matches to paired LC V and J genes in CoV-AbDab63, (2) heavy chain and light chain V gene pairs enriched in RBD-specific antibodies64,65, or (3) public heavy chain V genes and CDR3 sequence lengths enriched in RBD-specific antibodies64–66. We synthesized 20 antibodies corresponding to predicted RBD-specific BCRs and used these antibodies to experimentally validate RBD binding by enzyme-linked immunosorbent assays (ELISA; details below). Of the 20 antibodies, 7 were confirmed as RBD binders, and the computational pipeline was refined as follows: (1) method 3 above was removed from the pipeline as it had poor predictive value for determining SARS2-specificity and (2) the threshold for percent CDRH3 sequence identity to CoV-AbDab reference sequences was increased to 80%. Together, these changes resulted in a revised computational pipeline with a sensitivity of 35%, specificity of 87.5%, positive predictive value of 77.8% and negative predictive value of 51.9% (Supplementary Table 3). Predicted and validated RBD-specific BCRs were included in the final SARS2-specificity annotations, including three cells from the breakthrough infection donors shown in Fig. 4.
Validation of expanded SARS2-specific clones
Clone 4809_2 comprised 59 cells, including 1 cell that was validated experimentally as an RBD probe binder and 1 cell that was identified computationally as RBD-specific using the above computational methods. Similarly, clone 4809_3 comprised 36 cells, including 1 cell that was identified computationally as RBD-specific. As 56 out of 59 cells in clone 4809_2 and 35 out of 36 cells in clone 4809_3 were BGC cells and BPC, which have decreased BCR surface expression, we considered that the antigen probe assay may not have been sensitive enough to detect most RBD-specific cells within these clones. We thus synthesized recombinant human IgG monoclonal antibody representing BCRs distributed across distinct clades within these clonal lineages and validated them by ELISA, pseudovirus (PSV) neutralization and surface plasmon resonance (SPR) affinity measurements, as described in detail below; 100% of the antibodies tested from these clonal lineages (n = 6 for 4809_2 and n = 6 for 4809_3), including antibodies based on inferred clonal germline sequences, were validated as Omicron RBD-specific by all three methods. As such, all BCRs/B cells belonging to these clones were assigned as RBD-specific.
Antibody validation of SARS2-specific B cells
An initial batch of 20 bioinformatically identified putative SARS2-specific BCRs from the SARS2 breakthrough infection donors whose sequencing data are shown in Figs. 1 and 4 (and related Extended Data figures) were cloned as recombinant human IgG monoclonal antibodies, expressed in CHO cells (GenScript) and purified for validation. These 20 monoclonal antibodies were confirmed to be human IgG by ELISA. The antibodies were then tested for binding and neutralization of ancestral (D614G) and Omicron BA.4/5 SARS2 by ELISA and PSV neutralization assays, respectively. In total, 8 out of 20 antibodies recognized variants of SARS2 RBD as determined by ELISA: 6 bound strongly to both ancestral and BA.4/5 RBD and 2 bound strongly only to BA.4/5 RBD (the index antibodies from clones 4809_2 and 4809_3). Antigen-specificity was confirmed by performing ELISAs and PSV neutralization assays. The validated B cells that corresponded to the three breakthrough infection donors shown in Fig. 4 (three of the eight validated RBD binding antibodies) were added to Fig. 4h–i. A second batch of 25 antibodies, including 4 more antibodies from clone 4809_2 and 5 more antibodies from clone 4809_3 (11 in total, including the index antibodies), was synthesized for further validation. These also included eight antibodies (four per clone) based on inferred clonal germline sequences from clones 4809_2 and 4809_3, which were determined as follows: (1) for each clone, the nucleotide sequences for the germline junction regions predicted by both dowser and IgPhyML R packages were aligned with the junction regions of seven distinct sequenced BCRs with the fewest computed mutations; (2) ambiguous nucleotides were assigned by inferring the most likely nucleotide on the basis of the sequence alignments; (3) the final inferred germline nucleotide sequence for the entire heavy chain was compared with the nearest neighbour in the lineage tree using the National Center for Biotechnology Information BLASTn function to verify that the computed mutation number matched the number of nucleotide differences between the two sequences. Owing to the inherent ambiguity in predicting germline antibody sequences, four inferred germline iterations per clone were synthesized, representing the antibody sequences most likely to match the true germline. Only one iteration per clone showed evidence of binding to BA.4/5 RBD/spike, and these antibodies are indicated as GL2_V01 and GL3_V03 in the main text. Full variable region sequences for all validated SARS2-specific antibodies can be found in Supplementary Table 3. The antibody data are summarized in Fig. 4m.
SARS2 spike and RBD IgG ELISA
ELISAs were performed as described previously67. Positive control plasma pooled from convalescent COVID-19 donors was used to standardize each assay and normalize across experiments. Previously published and validated SARS2-specific monoclonal antibody were also used as positive controls68, and a previously published HIV envelope broadly neutralizing antibody was used as a negative control69. For the assay, 96-well half area plates were coated with 1 μg ml−1 of SARS2 Ancestral (D614G) spike or RBD (Saphire laboratory), Omicron BA.4/5 RBD (AcroBiosystems) or a negative control (EBV Nuclear antigen 1, EBNA1; Abcam) diluted in PBS overnight at 4 °C. Plates were then blocked with 3% milk in PBS containing 0.05% Tween-20 for 1.5 h at room temperature. Monoclonal antibodies were diluted in 1% milk containing 0.05% Tween-20 in PBS and added to the plate at a starting concentration of approximately 16,667 ng ml−1 (1 μg) of monoclonal antibodies, followed by serial threefold dilutions across the plate. Similarly, heat-inactivated, pooled human plasma controls were added to the appropriate wells and serially diluted across the plate. Plates were incubated for 1.5 h at room temperature, then washed. Plates were then incubated with conjugated IgG secondary antibody for 1 h. Plates were then washed and developed with 3,3′,5,5′- tetramethylbenzidine ELISA substrate solution (ThermoFisher). The optical density of the wells was then read using a SpectraMax Plate Reader at 450 nm using the SoftMax Pro software (v.7.1).
SARS2 PSV neutralization
PSV neutralization assays were performed as described previously67. In brief, a monolayer of Vero cells (ATCC) was generated by seeding 2.5 × 104 cells in black flat-bottom 96-well plate (Corning) and cells were incubated at 37 °C for 5–6 h. Pretitrated recombinant virus was incubated with serially diluted monoclonal antibody (threefold or sixfold serial dilutions from a starting concentration of 25,000 or 2,000 ng ml−1, respectively, depending on the activity of each specific antibody) for 1 h at 37 °C, added to confluent Vero cell monolayers, and incubated for 16–20 h at 37 °C in 5% CO2. Previously published and validated SARS2-specific monoclonal antibodies were used as positive controls, and a previously published HIV envelope trimer-binding broadly neutralizing antibody was used as a negative control (as above for ELISAs). Cells were then fixed in 4% paraformaldehyde in PBS with 10 μg ml−1 Hoechst (ThermoFisher) in the dark for 30 min at room temperature. Cells were imaged using a Cell Insight CX5 imager to quantify the total number of cells and infected GFP expressing cells. Neutralization titre (inhibition dose 50; ID50) were calculated from complete dose-response curves using the One-Site Fit Log IC50 model in GraphPad Prism. Recombinant SARS2 spike-pseudotyped VSV-ΔG-GFP were generated as described previously70 by transfecting HEK293T cells (ATCC) with phCMV-3-SARS-CoV-2 spike and collecting supernatants containing rVSV-SARS2 spike for virus variants with the specific amino acid mutations, as listed in the GISAID database: ancestral (D614G) or Omicron BA.4/5 (BA.5; T19I, L24S del25/27, del69/70, G142D, V213G, G339D, S371F, S373P, S375F, T376A, D405N, R408S, K417N, N440K, L452R, S477N, T478K, E484A, F486V, Q498R, N501Y, Y505H, D614G, H655Y, H655Y, N679K, P681H, N764K, D796Y, Q954H, N969K). Cells obtained from ATCC were not tested to authenticate cell line identity or for Mycoplasma contamination.
SARS2 spike protein production and purification for affinity testing
Recombinant SARS2 spike proteins were generated in the HexaPro background (containing residues 14–1,208 (GenBank: MN908947) of the ectodomain, six proline substitutions (F817P, A892P, A899P, A942P, K986P, V987P)), as well as the D614G mutation and replacement of cleavage site residues 682–685 (RRAR to GSAS)68. Each spike was cloned into a phCMV mammalian expression vector containing an N-terminal Gaussia luciferase signal sequence, a C-terminal HRV-3C cleavage site and a Twin-StrepII-Tag purification. HexaPro constructs also contained a foldon trimerization domain between the C terminus of S2 and the HRV-3C site. Plasmids were transformed into Stellar competent cells and isolated using a Plasmid Plus Midi kit (Qiagen). All clones were fully sequenced to ensure that no further mutations were introduced.
ExpiCHO-S cells (ThermoFisher) were transiently transfected with SARS2 spike constructs. CHO cells (ATCC) were maintained and transfected according to the manufacturer’s protocols. For all ExpiCHO cultures, the manufacturer’s High Titer protocol was used with a 7-day culture incubation to assess relative expression. In brief, plasmid DNA and Expifectamine were mixed in Opti-PRO SFM (Gibco) according to the manufacturer’s instructions and added to the cells. On day 1, cells were fed with manufacturer-supplied feed and enhancer as specified in the manufacturer’s protocol, and cultures were moved to a shaker incubator set to 32 °C, 5% CO2 and 115 rpm. On day 7, cultures were clarified by centrifugation, followed by addition of BioLock (IBA Life Sciences), passage through a 0.22 μM sterile filter and purification on an ÄKTA Pure system (Cytiva) using a 5 ml Strep-Tactin XT column equilibrated with TBS buffer (25 mM Tris pH 7.6, 200 mM NaCl, 0.02% NaN3), and eluted in TBS buffer supplemented with 10 mM d-desthiobiotin (Sigma Aldrich). Proteins were then purified by size-exclusion-chromatography on a Superdex 6 Increase 10/300 column (Cytiva) in the same TBS buffer.
High-throughput SPR binding kinetics and data analysis
Binding kinetics measurements were performed on the Carterra LSA platform using a CDMP sensor chip (Carterra). The chip was activated with a freshly prepared solution of 130 mM 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide hydrochloride (catalogue no. PG82079, Pierce) and 33 mM N-hydroxysulfosuccinimide (catalogue no. 24510, ThermoFisher Scientific) in 0.1 M MES pH 5.5 using a single flow cell. A surface capture lawn was prepared with 50 μg ml−1 of goat anti-human IgG Fc secondary antibody (catalogue no. 103255–066, VWR) in 10 mM sodium acetate (pH 4.5)/0.01% Tween. Unreactive esters were quenched with a 7-min injection of 1 M ethanolamine-HCl (pH 8.5). Purified monoclonal antibodies were captured using the 96-print head with 1× HBSTE buffer (10 mM HEPES pH 7.4, 150 mM NaCl, 3 mM EDTA and 0.01% Tween-20) as running buffer and antibody diluent. Three independent dilutions, at two concentrations (5 μg ml−1 and 0.5 μg ml−1 for spike analytes and 10 μg ml−1 and 20 μg ml−1 for RBD analytes), were prepared for each monoclonal antibody, with each dilution captured on two independent array spots.
A twofold dilution series of spike or RBD was prepared in 1× HBSTE-BSA buffer (10 mM HEPES pH 7.4,150 mM NaCl, 3 mM EDTA, 0.05% Tween-20, supplemented with 0.5 mg ml−1 bovine serum albumin (BSA)). Protein was injected onto the chip surface (at 25 °C) using the single flow cell, from the lowest to the highest concentration, without regeneration in between. Five buffer injections of the buffer before the lowest non-zero concentration were used for signal stabilization. For each concentration, baseline data were collected for 120 s, association data for 300 s and dissociation data for 900 s. After the titration of each analyte, the huFc capture lawn surface was regenerated with two pulses (17 s per pulse) of 10 mM glycine, pH 2.0 and monoclonal antibodies recaptured in between each analyte. The running buffer for all kinetic steps was 1× HBSTE-BSA. Titration data were processed with the Kinetics software package (Carterra v.1.9.1.4215 or higher), including reference subtraction, buffer subtraction and data smoothing. Analyte binding timecourses for each antibody were fitted to a 1:1 Langmuir model to derive ka, kd and KD values.
Statistics
Statistical analyses were performed in R (v.4.0.0 or newer), GraphPad Prism 10 (GraphPad Software) and FlowJo (v.9 or higher, BD). All flow cytometry and FACS data were analysed using FlowJo (v.9 or higher). For all analyses, a two-sided 5% type I error rate was used unless otherwise noted. Further details can be found in relevant Methods sections, the results, figures and corresponding figure legends.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Extended Data
Extended Data Fig. 1 |. NP swab sampling reveals diverse URT cell populations.
a. Total viable CD45+ immune cells collected per NP swab (n = 315 swabs). b. Example flow gating of CD45+ immune cell populations upstream of the TRM and putative BRM population gates shown in Fig. 1d c. Left: total viable CD45−EpCAM+ (CD326) epithelial cells collected per NP swab (n = 164 swabs); Right: example CD45 × EpCAM gating with CD45−EpCAM+ epithelial cell populations in the lower right quadrant. d. Epithelial cell UMAP from scRNA-seq of the approximately 8,000 epithelial cells derived from the same 12 NP swabs/donors as Fig. 1e. e. Dot plot demonstrating expression patterns of epithelial cell cluster defining genes. f. UMAP from Fig. 1e split by donor to show the contribution of NP swabs from each donor. g. Feature plots of gene (top panels) vs surface protein (bottom panels) expression for CD45, CD19, CD3E RNA and their corresponding proteins for the combined immune cell UMAP shown in Fig. 1e. h. Feature plots of gene (top panels) vs surface protein (bottom panels) expression for CD4 and CD8A RNA and their corresponding proteins for the T cell subset UMAP shown in Fig. 1f. Related to Fig. 1.
Extended Data Fig. 2 |. Characterization of adaptive immune populations of the human URT.
a. B cell subset UMAP generated from the approximately 35,000 B cells from Fig. 1e, with subset identities assigned by gene expression. b-c. (b) B cell and (c) T cell profiles (percentage of each lymphocyte subset for each donor) for each of the 12 swab donors whose B and T cell data are shown in Extended Data Fig. 2a and Fig. 1f, respectively. d. GSEA of the T cell subsets in Fig. 1f for the MAIT gene signature: NES = normalized enrichment score; -log(padj) = negative log10 Benjamini-Hochberg adjusted p-values (see Methods for details). MAIT were identified by a combination of gene expression and scTCRseq analysis (see Methods for details). e. Feature plot of FAS expression with the CD4+ TMem cluster from Fig. 1f circled. f-g. The clusters labelled TFH and Treg in Fig. 1f were named based on their most prominent cell type. Subset UMAPs generated from each of the (f) ~3,700 cell TFH and (g) ~1,500 Treg cluster cells from Fig. 1f showing additional cell types identified by subset analysis. Within the TFH cluster: mantle TFH (mTFH), GC-TFH, and GC-TFH with a type 1 IFN signaling signature (GC-TFHIFN). Within the Treg cluster: Treg, TH17, TFH, and TFR. See Fig. 1j for gene expression profiles. Virus-specific T cells were identified in these clusters and are indicated by colored halos based on the key shown in panels f,g. h. Example gating for T cell sorting for scRNA-seq/scTCR-seq based on AIM marker expression. i. SARS2 spike- and non-spike—specific TFH counts identified by AIM assay for each donor. j. SARS2-specific TFH counts (number per breakthrough infection donor swab) plotted by days post-COVID symptom onset. k. SARS2-specific T cell subsets identified in NP swabs split by donor. Related to Fig. 1.
Extended Data Fig. 3 |. Experimental workflow for defining personalized SARS2-specific T cell TCR references to identify upper airway antigen-specific T cells.
a. Workflow diagram for the 12 participants who underwent paired sample collection of peripheral blood (to isolate PBMCs) and NP swabs for CITE-seq plus scRNA-seq/scTCR-seq. SARS2-specific NP T cells were identified based on TCR overlap between NP swabs and blood. b-d. Representative flow plots for stimulated PBMC-derived SARS2-specific (b) CD4 T cells as OX40+CD40L+, (c) CD4 T cells as OX40+41BB+, and (d) CD8 T cells as CD69+41BB+ by AIM assay from a convalescent COVID-19 control donor with robust CD4 and CD8 S and non-S responses. e-g. Frequency of SARS2-specific (e) OX40+41BB+ CD4 T cells, (f) OX40+CD40L+ CD4 T cells, and (g) CD69+41BB+ CD8 T cells by AIM assay. For e-g the dotted line = limit of quantitation. h-j Stimulation index (SI) = fold-change above negative control for SARS2-specific CD4 (h) OX40+41BB+ and (i) OX40+CD40L+ and CD8 (j) CD69+41BB+ T cell responses by AIM assay. For h-j the dotted line = SI of 2. For e-j, data is shown from the 12 donors in panel a; positive responses are greater than the dotted line and negative responses are set to baseline (0.5 × LOQ or SI = 1); bar = geomean; dots are color-coded based on COVID-19 status as per the legend. Related to Fig. 1. Panel a was created using BioRender (https://Biorender.com).
Extended Data Fig. 4 |. Adult adenoids are URT secondary lymphoid organs with functional GCs.
a. Example gating for NP swab (adenoid) BGC and GC-TFH. ASC = antibody secreting cells. b. Top panels: flow plots showing intracellular Ki67 and BCL6 staining for BGC compared with non-BGC; Bottom panels: histograms showing intracellular BCL6 expression in GC-TFH, non-TFH CD4+ T cells, and BGC. c. UMAP showing the cell cycle phases of the BGC in Fig. 2b. d. Dot plot showing expression of additional BGC genes (related to Fig. 2c). e. A B cell clonal lineage from the NP swab of donor 4379, represented as linear phylogenic trees by B cell type (left) and by isotype (right). Related to Fig. 2.
Extended Data Fig. 5 |. Adenoids and nasal turbinate mucosa have distinct immune cell profiles.
a. Example gating for B and T cell subsets and B cell isotypes from NP and MT swabs shown in Fig. 3 and in b-h. b-h. Additional differences between NP (adenoid) and MT swab populations: (b) % naive B of total B cells, (c) % naive CD4+ of total T cells, (d) % naive CD8+ of total CD8+ T cells, (e) %Treg of CD4+ T cells, (f) %CD8+ TRM (CD69+CD103+) of CD8+ T cells (g) % non-naive CD4+ of total CD4+ T cells, and (h) % non-naive CD8+ of total CD8+ T cells. For b-h: Pairwise testing by Mann-Whitney; bars = median, two-tailed p-values: *** = p < 0.001, **** = p < 0.0001 (n = 38 donors; 88 NP and 55 MT swabs). i. Example flow plots for putative BRM (CD69+ B cells), CD4+ TRM (CD69+CD103+/−), CD8+ TRM (CD69+CD103+), BGC (CD19+CD20+CD38+), and GC-TFH (CXCR5+PD-1hi) in blood (PBMC). j-o. Additional correlations related to adenoid atrophy with aging: (j) frequency of BGC (% of total B cells) per NP swab vs age (n = 138 donors; 1 swab/donor), (k) BGC per NP swab in log10 vs % naive B (of total B cells) (n = 233 swabs), (l) frequency of GC-TFH (% of CD4+ T cells) per NP swab vs age (n = 138 donors; 1 swab/donor), (m) GC-TFH in log10 per NP swab vs age (n = 119 donors; 1 swab/donor), (n) frequency of naive B (% of total B cells) per NP swab vs age (n = 138 donors; 1 swab/donor), (o) naive B per NP swab in log10 vs age (n = 138 donors; 1 swab/donor); For j, k, n, and o: Spearman r and two-tailed p-values. For l-m: Pearson r and two-tailed p-values. Related to Fig. 3.
Extended Data Fig. 6 |. Multimodal validation of SARS2-specific URT mucosal B cells.
a. Example flow gating related to SARS2 probe staining for SARS2 S-, RBD-, and N-specific B cells. b. Proportion of IgG+ SARS2-specific Bmem in NP (n = 58 subjects) and MT swabs (n = 11 subjects) versus PBMC (n = 24 subjects). Bars = median. Pairwise comparisons by Mann-Whitney; two-tailed p-values: ns = not significant, * = p ≤ 0.05, **** = p ≤ 0.0001. Please see Methods for additional details. c. % SARS2 N-, RBD-, and spike (S)-specific IgA+ or IgG+ Bmem of total Bmem over time post-breakthrough infection by flow cytometry with fluorescent probe staining (n = 32–41 subjects). Dotted line = limit of detection. d. Select differences in gene expression patterns between NP swab putative BRM (orange) and PBMC-derived atypical Bmem (teal). PBMC-derived atypical Bmem often express TBX21. e. Gating strategy for sorting SARS2 S-, RBD-, and N-specific Bmem in PBMC for scRNA-seq/scBCR-seq. f. SARS2-specific B cells from the 3 breakthrough infection donors in Fig. 4h color-coded by method used for assigning Ag-specificity. g. Heat map of human bone marrow PB and early vs late PC gene expression by NP ASC cells in Fig. 4k based on published gene signatures. h. UMAP related to Fig. 4k showing BPC belonging to donor 4809 that are from the SARS2-specific clones 2 and 3 shown in Fig. 4l, and BPC belonging to a non-SARS2-specific clone (4809_10). Related to Fig. 4.
Extended Data Fig. 7 |. Expansion of SARS2 Omicron BA.4/5 variant-specific Bmem in the upper airway.
a. UMAP plots demonstrating the distribution of Omicron-specific clones 2 and 3 from donor 4809 across the B cell clusters from Fig. 4d. b. Two SARS2 Omicron-specific B cell clonal lineages from donor 4809 represented as linear phylogenic trees as in Fig. 4l. Tip sizes represent the number of cells with identical features. Tip colors represent the isotype. Tip labels represent validated antibodies. c. ELISA IgG dilution curves for ancestral and Omicron BA.4/5 RBD for the antibodies generated for SARS2-specific B cell validation (related to Fig. 4m summary data). Mean and standard deviation (error bars) for three independent experiments are shown for controls and the 23 mAb in Fig. 4m. d. Timeline of vaccination, breakthrough infection, and NP swab collection for single-cell experiments for donor 4809. Related to Fig. 4.
Supplementary Material
Acknowledgements
We would like to thank our donors for providing the samples that made this study possible. We gratefully acknowledge provision of epitope peptide pools from A. Sette and A. Grifoni (LJI), and sharing of early, unpublished results by R. Da Silva Antunes (LJI). We also thank P. Vijayanand (Vijay) (LJI) for advice and discussion. We would also like to thank the LJI core facilities for providing their expertise, services and support to aid in the completion of this study: the Clinical Studies Core staff, in particular the Clinical Studies Nurse and Supervisor J. Cardenas, the LJI Flow Cytometry Core facility director C. Kim and staff, including M. Shil, the Next Generation Sequencing Core facility director S. Alarcon and staff. Fig. 2e and Extended Data Fig. 3a were created with BioRender.com. This work was supported in part by the National Institute of Allergy and Infectious diseases (NIAID) of the National Institutes of Health (NIH), Department of Health and Human Services, under award U19 AI142742 Collaborative Center for Human Immunology. Further support was provided in part by the NIH T32 AI007036 (S.I.R.) and an A.P. Giannini Foundation fellowship award (S.I.R.). The content presented is the sole responsibility of the authors and may not represent the opinions of the NIH.
Footnotes
Competing interests S.C. has consulted for GSK, JP Morgan, Citi, Morgan Stanley, Avalia NZ, Nutcracker Therapeutics, University of California, California State Universities, United Airlines, Adagio, Sanofi and Roche. LJI has filed for patent protection for various aspects of T cell epitope and vaccine design work. The other authors declare no competing interests.
Additional information
Supplementary information The online version contains supplementary material available at https://doi.org/10.1038/s41586-024-07748-8.
Data availability
The authors confirm that the source data underlying the findings are fully available. Gene expression, BCR and TCR sequencing data generated in the current study are available in the Gene Expression Omnibus database (GEO) under the GEO Series record GSE270518. All data obtained from publicly available sources is cited in the text and the references include the URL for the data source.
References
- 1.Sette A & Crotty S Immunological memory to SARS-CoV-2 infection and COVID-19 vaccines. Immunol. Rev. 310, 27–46 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Allie SR & Randall TD Resident memory B cells. Viral Immunol. 33, 282–293 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Topol EJ & Iwasaki A Operation nasal vaccine-lightning speed to counter COVID-19. Sci. Immunol. 7, eadd9947 (2022). [DOI] [PubMed] [Google Scholar]
- 4.Zheng MZM & Wakim LM Tissue resident memory T cells in the respiratory tract. Mucosal Immunol. 15, 379–388 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Poon MML et al. Tissue adaptation and clonal segregation of human memory T cells in barrier sites. Nat. Immunol. 24, 309–319 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Domínguez Conde C et al. Cross-tissue immune cell analysis reveals tissue-specific features in humans. Science 376, eabl5197 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Clark RA Resident memory T cells in human health and disease. Sci. Transl. Med. 7, 269rv1 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Lange J., Rivera-Ballesteros O. & Buggert M. Human mucosal tissue-resident memory T cells in health and disease. Mucosal Immunol. 15, 389–397 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Thome JJC et al. Spatial map of human T cell compartmentalization and maintenance over decades of life. Cell 159, 814–828 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Gray JI & Farber DL Tissue-resident immune cells in humans. Annu. Rev. Immunol. 40, 195–220 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Masopust D & Soerens AG Tissue-resident T cells and other resident leukocytes. Annu. Rev. Immunol. 37, 521–546 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Snyder ME et al. Generation and persistence of human tissue-resident memory T cells in lung transplantation. Sci. Immunol. 4, eaav5581 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Weisel NM et al. Comprehensive analyses of B-cell compartments across the human body reveal novel subsets and a gut-resident memory phenotype. Blood 136, 2774–2785 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Havenar-Daughton C, Lee JH & Crotty S Tfh cells and HIV bnAbs, an immunodominance model of the HIV neutralizing antibody generation problem. Immunol. Rev. 275, 49–61 (2017). [DOI] [PubMed] [Google Scholar]
- 15.Havenar-Daughton C et al. Normal human lymph node T follicular helper cells and germinal center B cells accessed via fine needle aspirations. J. Immunol. Methods 479, 112746 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Alsoussi WB et al. SARS-CoV-2 Omicron boosting induces de novo B cell response in humans. Nature 617, 592–598 (2023). [DOI] [PubMed] [Google Scholar]
- 17.Leggat DJ et al. Vaccination induces HIV broadly neutralizing antibody precursors in humans. Science 378, eadd6502 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Victora GD & Nussenzweig MC Germinal Centers. Annu. Rev. Immunol. 40, 413–442 (2022). [DOI] [PubMed] [Google Scholar]
- 19.Xu Q et al. Adaptive immune responses to SARS-CoV-2 persist in the pharyngeal lymphoid tissue of children. Nat. Immunol. 24, 186–199 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Kumar BV et al. Human tissue-resident memory T cells are defined by core transcriptional and functional signatures in lymphoid and mucosal sites. Cell Rep. 20, 2921–2934 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Kusnadi A et al. Severely ill COVID-19 patients display impaired exhaustion features in SARS-CoV-2-reactive CD8+ T cells. Sci. Immunol. 6, eabe4782 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Meckiff BJ et al. Imbalance of regulatory and cytotoxic SARS-CoV-2-reactive CD4+ T cells in COVID-19. Cell 183, 1340–1353.e16 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Schmiedel BJ et al. Single-cell eQTL analysis of activated T cell subsets reveals activation and cell type-dependent effects of disease-risk variants. Sci. Immunol. 7, eabm2508 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.FitzPatrick MEB et al. Human intestinal tissue-resident memory T cells comprise transcriptionally and functionally distinct subsets. Cell Reports 34, 108661 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Crotty S T follicular helper cell biology: a decade of discovery and diseases. Immunity 50, 1132–1148 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Mudd PA et al. SARS-CoV-2 mRNA vaccination elicits a robust and persistent T follicular helper cell response in humans. Cell 185, 603–613.e15 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Nolan S et al. A large-scale database of T-cell receptor beta (TCRb) sequences and binding associations from natural and synthetic exposure to SARS-CoV-2. Preprint at Research Square 10.21203/rs.3.rs-51964/v1 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Goncharov M et al. VDJdb in the pandemic era: a compendium of T cell receptors specific for SARS-CoV-2. Nat. Methods 19, 1017–1019 (2022). [DOI] [PubMed] [Google Scholar]
- 29.Rowntree LC et al. SARS-CoV-2-specific T cell memory with common TCRαβ motifs is established in unvaccinated children who seroconvert after infection. Immunity 55, 1299–1315.e4 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Pogorelyy MV et al. Resolving SARS-CoV-2 CD4+ T cell specificity via reverse epitope discovery. Cell Rep. Med. 3, 100697 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Tan H-X et al. Lung-resident memory B cells established after pulmonary influenza infection display distinct transcriptional and phenotypic profiles. Sci. Immunol. 7, eabf5314 (2022). [DOI] [PubMed] [Google Scholar]
- 32.Barker KA et al. Lung-resident memory B cells protect against bacterial pneumonia. J. Clin. Invest. 131, e141810 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Gregoire C et al. Viral infection engenders bona fide and bystander subsets of lung-resident memory B cells through a permissive mechanism. Immunity 55, 1216–1233.e9 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Allie SR et al. The establishment of resident memory B cells in the lung requires local antigen encounter. Nat. Immunol. 20, 97–108 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Ehrhardt GRA et al. Expression of the immunoregulatory molecule FcRH4 defines a distinctive tissue-based population of memory B cells. J. Exp. Med. 202, 783–791 (2005). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Ehrhardt GRA et al. Discriminating gene expression profiles of memory B cell subpopulations. J. Exp. Med. 205, 1807–1817 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.King HW et al. Single-cell analysis of human B cell maturation predicts how antibody class switching shapes selection dynamics. Sci. Immunol. 6, eabe6291 (2021). [DOI] [PubMed] [Google Scholar]
- 38.Zumaquero E et al. IFNγ induces epigenetic programming of human T-bethi B cells and promotes TLR7/8 and IL-21 induced differentiation. eLife 8, e41641 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Knox JJ et al. T-bet+ B cells are induced by human viral infections and dominate the HIV gp140 response. JCI Insight 2, e92943 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Duan M et al. Understanding heterogeneity of human bone marrow plasma cell maturation and survival pathways by single-cell analyses. Cell Rep. 42, 112682 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Lim JME et al. SARS-CoV-2 breakthrough infection in vaccinees induces virus-specific nasal-resident CD8+ and CD4+ T cells of broad specificity. J. Exp. Med. 219, e20220780 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Dan JM et al. Immunological memory to SARS-CoV-2 assessed for up to 8 months after infection. Science 371, eabf4063 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Grifoni A et al. Targets of T cell responses to SARS-CoV-2 coronavirus in humans with COVID-19 disease and unexposed individuals. Cell 181, 1489–1501.e15 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Dan JM et al. Recurrent group A Streptococcus tonsillitis is an immunosusceptibility disease involving antibody deficiency and aberrant T FH cells. Sci. Transl. Med. 11, eaau3776 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Ramirez SI et al. Bamlanivimab therapy for acute COVID-19 does not blunt SARS-CoV-2–specific memory T cell responses. JCI Insight 7, e163471 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Yu ED et al. Development of a T cell-based immunodiagnostic system to effectively distinguish SARS-CoV-2 infection and COVID-19 vaccination status. Cell Host Microbe 30, 388–399.e3 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Grifoni A et al. SARS-CoV-2 human T cell epitopes: adaptive immune response against COVID-19. Cell Host Microbe 29, 1076–1092 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Hao Y et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e29 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Korotkevich G et al. Fast gene set enrichment analysis. Preprint at bioRXiv 10.1101/060012 (2016). [DOI] [Google Scholar]
- 50.Lee JH et al. Long-primed germinal centres with enduring affinity maturation and clonal migration. Nature 609, 998–1004 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Holla P et al. Shared transcriptional profiles of atypical B cells suggest common drivers of expansion and function in malaria, HIV, and autoimmunity. Sci. Adv. 7, eabg8384 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Lopes De Assis F et al. Tracking B cell responses to the SARS-CoV-2 mRNA-1273 vaccine. Cell Rep. 42, 112780 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Yermanos A et al. Platypus: an open-access software for integrating lymphocyte single-cell immune repertoires with transcriptomes. NAR Genom. Bioinform. 3, lqab023 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Gupta NT et al. Change-O: a toolkit for analyzing large-scale B cell immunoglobulin repertoire sequencing data. Bioinformatics 31, 3356–3358 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Vander Heiden JA et al. pRESTO: a toolkit for processing high-throughput sequencing raw reads of lymphocyte receptor repertoires. Bioinformatics 30, 1930–1932 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Gupta NT et al. Hierarchical clustering can identify B cell clones with high confidence in Ig repertoire sequencing data. J. Immunol. 198, 2489–2499 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Nouri N & Kleinstein SH Somatic hypermutation analysis for improved identification of B cell clonal families from next-generation sequencing data. PLoS Comput. Biol. 16, e1007977 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Ye J, Ma N, Madden TL & Ostell JM IgBLAST: an immunoglobulin variable domain sequence analysis tool. Nucleic Acids Res. 41, W34–W40 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Giudicelli V, Chaume D & Lefranc M-P IMGT/GENE-DB: a comprehensive database for human and mouse immunoglobulin and T cell receptor genes. Nucleic Acids Res. 33, D256–D261 (2005). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Hoehn KB et al. Repertoire-wide phylogenetic models of B cell molecular evolution reveal evolutionary signatures of aging and vaccination. Proc. Natl Acad. Sci. USA 116, 22664–22672 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Hoehn KB et al. Human B cell lineages associated with germinal centers following influenza vaccination are measurably evolving. eLife 10, e70873 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Hoehn KB, Pybus OG & Kleinstein SH Phylogenetic analysis of migration, differentiation, and class switching in B cells. PLoS Comput. Biol. 18, e1009885 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Raybould MIJ, Kovaltsuk A, Marks C & Deane CM CoV-AbDab: the coronavirus antibody database. Bioinformatics 37, 734–735 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Kaku CI et al. Evolution of antibody immunity following Omicron BA.1 breakthrough infection. Nat. Commun. 14, 2751 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Chen EC et al. Convergent antibody responses to the SARS-CoV-2 spike protein in convalescent and vaccinated individuals. Cell Rep. 36, 109604 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Barnes CO et al. Structures of human antibodies bound to SARS-CoV-2 spike reveal common epitopes and recurrent features of antibodies. Cell 182, 828–842.e16 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Tarke A et al. SARS-CoV-2 vaccination induces immunological T cell memory able to cross-recognize variants from Alpha to Omicron. Cell 185, 847–859.e11 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Hastie KM et al. Potent Omicron-neutralizing antibodies isolated from a patient vaccinated 6 months before Omicron emergence. Cell Rep. 42, 112421 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Lee JH et al. A broadly neutralizing antibody targets the dynamic HIV envelope trimer apex via a long, rigidified, and anionic β-hairpin structure. Immunity 46, 690–702 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Zhang Z et al. Humoral and cellular immune memory to four COVID-19 vaccines. Cell 185, 2434–2451.e17 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The authors confirm that the source data underlying the findings are fully available. Gene expression, BCR and TCR sequencing data generated in the current study are available in the Gene Expression Omnibus database (GEO) under the GEO Series record GSE270518. All data obtained from publicly available sources is cited in the text and the references include the URL for the data source.