Abstract
Background
Epitranscriptomics, the study of RNA modifications such as N6-methyladenosine (m6A), provides a novel layer of gene expression regulation with implications for numerous biological processes, including cellular adaptation to hypoxia. Hypoxia-inducible factor-1 (HIF-1), a master regulator of the cellular response to low oxygen, plays a critical role in adaptive and pathological processes, including cancer, ischemic heart disease, and metabolic disorders. Recent discoveries accent the dynamic interplay between m6A modifications and HIF-1 signaling, revealing a complex bidirectional regulatory network. While the roles of other RNA modifications in HIF-1 regulation remain largely unexplored, emerging evidence suggests their potential significance.
Main body
This review examines the reciprocal regulation between HIF-1 and epitranscriptomic machinery, including m6A writers, readers, and erasers. HIF-1 modulates the expression of key m6A components, while its own mRNA is regulated by m6A modifications, positioning HIF-1 as both a regulator and a target in this system. This interaction enhances our understanding of cellular hypoxic responses and opens avenues for clinical applications in treating conditions like cancer and ischemic heart disease. Promising progress has been made in developing selective inhibitors targeting the m6A-HIF-1 regulatory axis. However, challenges such as off-target effects and the complexity of RNA modification dynamics remain significant barriers to clinical translation.
Conclusion
The intricate interplay between m6A and HIF-1 highlights the critical role of epitranscriptomics in hypoxia-driven processes. Further research into these regulatory networks could drive therapeutic innovation in cancer, ischemic heart disease, and other hypoxia-related conditions. Overcoming challenges in specificity and off-target effects will be essential for realizing the potential of these emerging therapies.
Keywords: HIF-1, Hypoxia-inducible factor-1, Epitranscriptomics, m6A, Cancer, Heart
Background
Hypoxia-inducible factor (HIF) has been recognized as a key regulator of the cellular response to low oxygen levels, essential for maintaining oxygen homeostasis (Wang et al. 1995). Hypoxia can occur in both physiological and pathological conditions, positioning HIF at the core of processes such as development (Bohuslavova et al. 2019), metabolism (Kierans and Taylor 2021), angiogenesis (Monaci et al. 2024), and diseases like cancer (Pezzuto and Carico 2018) and ischemic heart disease (Semenza 2014).
HIF is a heterodimer composed of an oxygen-sensitive α-subunit and a constitutively expressed β-subunit (Wang and Semenza 1995). Three paralogues of the HIF-α subunit (HIF-1α, HIF-2α, and HIF-3α), and three paralogues of HIF-β have been identified in mammals (Zagórska and Dulak 2004). Among these, HIF-1α, expressed ubiquitously, plays a central role in regulating the hypoxic response. However, HIF-2α expressed in specific tissues also contributes significantly to this regulation (Wiesener et al. 2003). The function of HIF-3α is less understood, and it has traditionally been considered an inhibitor of HIF-1α activity (Makino et al. 2001). However, its role appears to be isoform-dependent, as some splicing variants have been shown to induce the expression of certain genes, including erythropoietin (EPO) (Tolonen et al. 2020). While HIF activity has traditionally been linked to protein-level regulation and oxygen-dependent mechanisms, recent studies have highlighted the crucial involvement of RNA modifications in its regulation (Nan et al. 2023).
Epitranscriptomics, also known as RNA epigenetics, is the study of chemical modifications on RNA molecules that regulate their function without altering the underlying RNA sequence. These modifications provide an additional layer of control over gene expression, much like epigenetic changes on DNA or histones (Peixoto et al. 2020). By dynamically regulating RNA fate, epitranscriptomic modifications enable cells to fine-tune gene expression in response to environmental cues and cellular signals. These mechanisms are especially critical under conditions of stress or altered metabolism, such as hypoxia, which directly affects HIF-1 signaling pathway (Hlavackova et al. 2018). First discovered in the 1960s, these modifications were initially thought to be adaptive markers that adjust the structure of mature RNA (Davis and Allen 1957; Cohn 1960). Today, over 170 different RNA modifications have been identified (Cappannini et al. 2024). Among these N6-methyladenosine (m6A) stands out as the most abundant one in mRNA (Desrosiers et al. 1974). m6A is implicated in nearly every stage of RNA metabolism, including mRNA translation, degradation, splicing, export, and folding (Longenecker et al. 2020). By modulating the activity of target transcripts, m6A subsequently influences various cellular processes and physiological functions. Importantly, it is a reversible and dynamic modification that responds to environmental conditions (Batista et al. 2014). Dysregulation of m6A signaling has been linked to many severe human diseases, including neurodegenerative, metabolic, and cardiovascular disorders (Meier et al. 2016).
Recent studies have uncovered a complex, bidirectional relationship between m6A modifications and HIF-1 signaling, in which m6A regulates HIF1A expression (Liang et al. 2022), while HIF-1, in turn, influences the m6A machinery (Lu et al. 2024). This review aims to explore the intricate crosstalk between m6A and HIF-1, focusing on their reciprocal regulation and its biological and clinical implications. By integrating these findings, we aim to shed light on the growing importance of epitranscriptomics in hypoxic signaling and its therapeutic potential.
HIF-1
HIF-1 is a heterodimeric transcription factor composed of an oxygen-sensitive HIF-1α subunit and a constitutively expressed HIF-1β subunit, also known as the aryl hydrocarbon receptor nuclear translocator (ARNT) (Wang and Semenza 1995). The activity of HIF-1 is tightly controlled by oxygen-dependent post-translational modifications that regulate the stability and activity of the HIF-1α subunit.
Under normoxic conditions, HIF-1α is rapidly degraded through the ubiquitin–proteasome pathway. This degradation is initiated by proline hydroxylation, which is catalyzed by the prolyl hydroxylase domain (PHD) family of enzymes (PHD1, PHD2, and PHD3) (Epstein et al. 2001). Proline hydroxylation promotes the binding of HIF-1α to the von Hippel-Lindau (pVHL) protein, an E3 ubiquitin ligase that targets HIF-1α for proteasomal degradation. Simultaneously, asparagine hydroxylation by the enzyme factor inhibiting HIF-1 (FIH-1) prevents the recruitment of transcriptional co-activators, further suppressing HIF-1 activity under normal oxygen conditions (Koivunen et al. 2004).
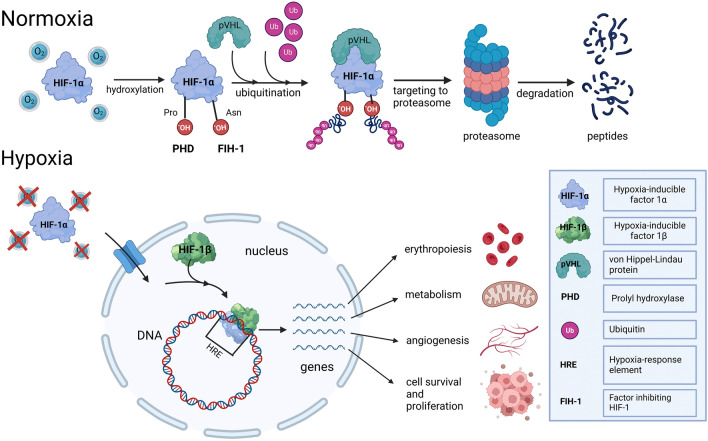
During hypoxia (Fig. 1), reduced oxygen availability inhibits the enzymatic activity of PHDs and FIH. This allows HIF-1α to escape degradation, accumulate in the cytoplasm, and translocate to the nucleus, where it dimerizes with HIF-1β. The HIF-1α/HIF-1β heterodimer binds to hypoxia-response elements (HREs) within the promoters and enhancers of target genes, initiating their transcription (Semenza 2001). The activation of HIF-1 occurs in a stepwise manner. Initially, the inhibition of PHD activity under mild hypoxia allows for the stabilization of HIF-1α. Under more severe hypoxic conditions, FIH-1 activity is also suppressed, enhancing the transactivation potential of HIF-1. This coordinated regulation enables HIF-1 to achieve maximal activity, driving the transcription of a diverse array of genes. These target genes, numbering over 1000, mediate critical adaptive responses. HIF-1 was initially believed to play a crucial role in erythropoiesis by stimulating EPO production (Semenza et al. 1991). However, more recent studies have identified HIF-2 as the primary transcription factor driving EPO expression (Warnecke et al. 2004; Haase 2010). Despite this, HIF-1 remains essential for hypoxic adaptation (Alanova et al. 2024), particularly by promoting angiogenesis through VEGF upregulation, which enhances blood vessel formation and oxygen delivery (Forsythe et al. 1996). Metabolic reprogramming is another key function, ensuring anaerobic ATP production by increasing glycolytic enzymes and glucose transporters (Fukuda et al. 2007). Finally, HIF-1 supports cell survival by regulating genes that mitigate oxidative stress and sustain viability in hypoxic conditions (Ong and Hausenloy 2012). Together, these adaptations ensure effective responses to oxygen deprivation, helping cells overcome this challenge.
Fig. 1.
Overview of HIF-1 regulation. Under normoxia, PHDs hydroxylate HIF-1α, triggering pVHL-mediated degradation, while FIH-1 inhibits its transcriptional activity. In hypoxia, HIF-1α is stabilized, translocates into the nucleus, dimerizes with HIF-1β, and binds to HREs to activate genes involved in erythropoiesis, metabolism, angiogenesis, and cell survival
While HIF-1 signaling plays a critical role in cellular adaptation to hypoxia, it can be a double-edged sword. For example, in cardiomyocytes, HIF-1 activation during ischemic states promotes cell survival and initiates protective pathways that increase the ability of the heart to cope with reduced oxygen availability (Ong et al. 2014). However, in the context of cancer, the same mechanisms can be detrimental. By enhancing tumor cell survival, angiogenesis, and metabolic reprogramming, HIF-1 contributes to tumor growth, invasion, and resistance to therapy (Semenza 2003). These contrasting effects point out the tissue-specific and disease-dependent nature of HIF-1 signaling. Its adaptability underscores its importance in both beneficial physiological processes, such as cardiac protection, and pathological conditions, such as cancer progression, making it a pivotal target for therapeutic interventions across diverse medical fields, including oncology and cardiovascular medicine.
In addition to oxygen-dependent regulation, HIF-1 signaling is influenced by mechanisms operating at multiple levels, ranging from transcriptional to post-translational regulation. For example, epigenetic modifications regulate the expression of HIF-1 transcripts, while post-translational modifications such as SUMOylation and deubiquitylation modulate HIF-1 stability and activity (Koyasu et al. 2018; Catrina and Zheng 2021; Semenza 2017). This review focuses on the post-transcriptional regulation of HIF-1 by the most prevalent epitranscriptomic modification, m6A, with an emphasis on the bidirectional interactions between HIF-1 signaling and this RNA modification.
N6-methyladenosine (m6A)
m6A is the most prevalent and well-studied epitranscriptomic modification in eukaryotic mRNAs, though it also occurs in non-coding RNAs such as long non-coding RNAs (lncRNAs), ribosomal RNAs (rRNAs), and small nuclear RNAs (snRNAs) (Zhang et al. 2021; Sendinc and Shi 2023). It is an evolutionary conserved modification found across a wide range of species, from plants to mammals, underscoring its essential role in regulating gene expression (Liang et al. 2020). Mechanistically, this modification alters RNA structure by forcing the methylamino group into an anti-conformation, destabilizing the thermodynamics of the RNA duplex (Sweaad et al. 2021). This structural change facilitates the interaction of RNA-binding proteins with their targets and subsequently influences various stages of RNA metabolism, including translation, stability, and decay (Niu et al. 2013).
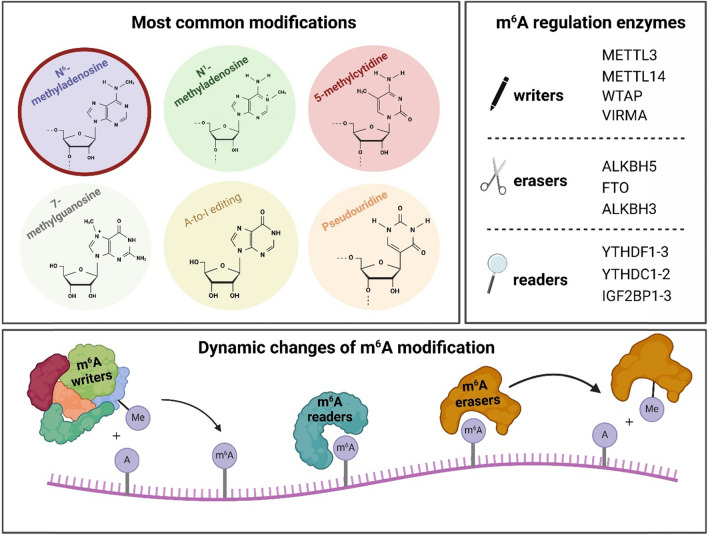
The regulation of m6A is mediated by a group of specialized RNA-modifying enzymes (overview in Fig. 2). Writers catalyze the addition of methyl groups to RNA, creating modifications that are recognized by RNA-binding proteins known as readers. While many irreversible RNA modifications are regulated through the degradation of modified RNA, the reversible nature of m6A allows for its removal by specialized enzymes called erasers, enabling dynamic regulation of RNA function without requiring its decay (Benak et al. 2024a).
Fig. 2.
Overview of RNA modifications and m6A regulation. The top left panel illustrates the most common RNA modifications, including N⁶-methyladenosine, N1-methyladenosine, 5-methylcytidine, 7-methylguanosine, A-to-I editing, and pseudouridine. The top right panel lists key m6A regulatory enzymes, categorized as writers (METTL3, METTL14, WTAP, VIRMA), erasers (ALKBH5, FTO, ALKBH3), and readers (YTHDF1-3, YTHDC1-2, IGF2BP1-3). The bottom panel represents the dynamic cycle of m6A modification, where m6A writers deposit methyl groups on adenosine, readers recognize and interpret the modification, and erasers remove m6A to regulate RNA fate and function. ALKBH5: AlkB homolog 5; FTO: fat mass and obesity-associated; IGF2BP1-3: insulin-like growth factor 2 mRNA-binding protein 1–3; METTL3/14: methyltransferase-like 3/14; m6A: N6-methyladenosine; VIRMA: vir-like m6A methyltransferase associated; WTAP: Wilms’ tumor 1-associating protein; YTHDC1-2: YTH domain-containing protein 2; YTHDF1-3: YTH domain-containing family protein 1–3
The primary protein complex responsible for adding the m6A modification to RNA consists of several key components. The most critical include methyltransferase-like 3 (METTL3), which serves as the catalytic subunit (Wang et al. 2016a, b); methyltransferase-like 14 (METTL14), which facilitates RNA binding (Wang et al. 2016a, b); and Wilms’ tumor 1-associating protein (WTAP), which aids in localizing the complex to nuclear speckles (Ping et al. 2014). Vir-like m6A methyltransferase associated (VIRMA, also known as KIAA1429) is another component of the methyltransferase complex, which mediates preferential methylation in specific RNA sites (Yue et al. 2018).
Several RNA-binding proteins have been identified as m6A readers, with the YTH domain-containing family proteins (YTHDF1-3) and YTH domain-containing proteins (YTHDC1-2) being among the most significant. These proteins play crucial roles in mRNA metabolism. While YTHDF1-3 orthologs are primarily involved in mRNA degradation (Zaccara and Jaffrey 2020; Lasman et al. 2020), both YTHDF1 and YTHDF3 have also been implicated in regulating translation (Wang et al. 2015; Shi et al. 2017). YTHDC1 is involved in mRNA splicing (Xiao et al. 2016), and YTHDC2 promotes translation (Hsu et al. 2017). IGF2BP1-3 proteins (insulin-like growth factor 2 mRNA-binding proteins 1–3) also bind to m6A and promote the stability and storage of their target mRNAs and therefore affect gene expression output (Huang et al. 2018).
AlkB homolog 5 (ALKBH5) and fat mass and obesity-associated protein (FTO) are the most well-known demethylases that remove the methyl group from m6A-modified transcripts. ALKBH5 is the primary m6A eraser, specifically targeting and demethylating m6A in mRNA and snRNA (Zheng et al. 2013; Wang et al. 2023). Although FTO is not exclusively specific to m6A, it predominantly acts on m6A in the nucleus, where it interacts with various RNA species, including mRNA, snRNA, and tRNA. In addition to m6A, FTO also demethylates N6,2ʹ-O-dimethyladenosine (m6Am) and N1-methyladenosine (m1A), further expanding its regulatory role in RNA metabolism (Benak et al. 2024b; Jia et al. 2011; Wei et al. 2018; Relier et al. 2021). Another eraser, AlkB homolog 3 (ALKBH3) has been reported to promote the m6A demethylation of mammalian tRNA (Ueda et al. 2017).
The reciprocal relationship: HIF-1 and m6A
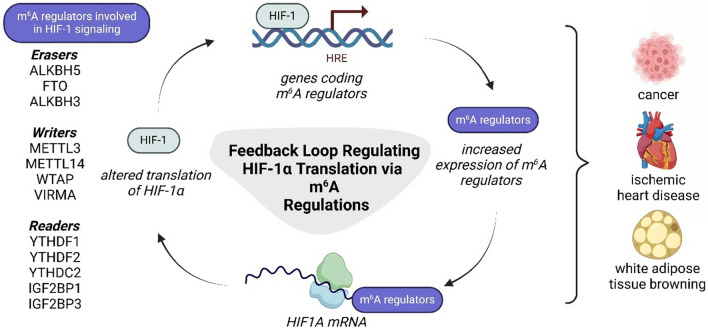
The dynamic interaction between HIF-1 and m6A plays a crucial role in gene expression regulation and had been described mainly in the cancer research (overview in Fig. 3).
Fig. 3.
The reciprocal relationship of HIF-1 and m6A. This figure illustrates the interaction between m6A regulators and HIF-1α translation in a feedback loop. HIF-1 binds to HREs in genes encoding m6A writers, erasers, and readers, leading to their increased expression. These m6A regulators modify HIF1A mRNA, affecting its stability and translation, which in turn alters HIF-1α protein levels. This regulatory mechanism plays a crucial role in various physiological and pathological conditions, including cancer, ischemic heart disease, and white adipose tissue browning. ALKBH5: AlkB homolog 5; FTO: fat mass and obesity-associated; HIF-1/HIF1A: hypoxia-inducible factor 1; HRE: hypoxia response element; IGF2BP1/3: insulin-like growth factor 2 mRNA-binding protein 1/3; METTL3/14: methyltransferase-like 3/14; m6A: N6-methyladenosine; VIRMA: vir-like m6A methyltransferase associated; WTAP: Wilms’ tumor 1-associating protein; YTHDC2: YTH domain-containing protein 2; YTHDF1-2: YTH domain-containing family protein 1–2
HIF-1 and m6A erasers
One of the earliest discoveries connecting m6A regulators and HIF-1 involves the m6A demethylase ALKBH5, which was identified as a direct transcriptional target of HIF-1α already in 2011 (Thalhammer et al. 2011), two years before its role as an m6A demethylase was revealed (Zheng et al. 2013). In this landmark study, Thalhammer et al. (2011) demonstrated that ALKBH5 is upregulated in response to hypoxia through HIF-1α signaling. This regulation was confirmed across multiple cell lines, where ALKBH5 expression increased significantly under hypoxic conditions. Following this discovery, further research revealed that the upregulation of ALKBH5 under hypoxia contributes to cancer progression. Specifically, in breast cancer cells, ALKBH5 induces pluripotency factors, leading to the formation of a breast cancer stem cell phenotype that is essential for tumor initiation and metastasis (Zhang et al. 2016a, b). More recently, another study demonstrated that in ovarian cancer, HIF-1α-driven upregulation of ALKBH5 stimulates downstream focal adhesion kinase (FAK)/Src proto-oncogene (Src) signaling and boosts integrin subunit beta 1 (ITGB1) expression by disrupting the YTHDF2 protein-mediated m6A degradation pathway, ultimately promoting lymph node metastasis and lymphangiogenesis (Sun et al. 2023).
Furthermore, recent studies indicated that ALKBH5-mediated m6A demethylation also plays a critical role in regulating HIF1A transcription, suggesting a reciprocal relationship between m6A and HIF-1. ALKBH5 was found to regulate m6A modifications on circular RNA circCCDC134 in cervical cancer. ALKBH5-mediated demethylation of circCCDC134 enhanced its stability, enabling it to recruit the transcription factor p65 (a subunit of NF-κB) and to act as a miR-503-5p sponge. These processes promoted the transcription of HIF1A, contributing to cancer progression and metastasis (Liang et al. 2022).
In addition to ALKBH5, also FTO plays a significant role in the regulation of HIF-1. FTO is upregulated in various cancers and contributes to tumor progression by influencing HIF-1 activity. One key study demonstrated that FTO demethylates and stabilizes the long intergenic non-coding RNA for kinase activation (LINK-A) in esophageal squamous cell carcinoma. LINK-A, in turn, disrupts the interaction between minichromosome maintenance complex component 3 (MCM3) and HIF-1α, abrogating MCM3-mediated HIF-1α transcriptional repression and promoting glycolysis and chemoresistance (Nan et al. 2023).
Beyond the direct regulation of HIF-1 by FTO, these two proteins also act synergistically. This was demonstrated with the hypoxia-responsive gene aldolase A (ALDOA), whose transcription is increased by HIF-1α under low oxygen conditions. At the same time, FTO demethylates ALDOA transcripts, stabilizing them and promoting their persistence, ultimately leading to the metabolic adaptation of cancer cells to hypoxia, enhancing glycolysis and hepatocellular carcinoma (HCC) progression (Niu et al. 2021).
These findings underscore the critical roles of m6A demethylases, ALKBH5 and FTO, in regulating HIF-1α activity. Both enzymes enhance the stability of key transcripts, like circCCDC134 and LINK-A, driving cancer progression. Moreover, these erasers can act synergistically with HIF-1α, as demonstrated with ALDOA, promoting hypoxia adaptation and tumor growth. These insights highlight the potential of targeting m6A demethylation in cancers dependent on HIF-1 signaling.
Moreover, influence of FTO on HIF-1α extends beyond cancer, playing a pivotal role in cardiovascular disorders, including ischemic heart disease and myocardial infarction (MI), both of which are associated with hypoxia. In MI, FTO expression was significantly downregulated in affected tissues, suggesting its potential protective role in cardiac injury (Shi et al. 2021; Wang et al. 2024). Dysregulation of the m6A machinery, including FTO, has been linked to key pathways such as PI3K-Akt and HIF-1, highlighting capacity of FTO to modulate cardiac function and hypoxic responses in ischemic conditions (Shi et al. 2021). In neonatal cardiac fibroblasts exposed to hypoxia (1% O₂ for 24 h), a decrease in FTO levels was associated with an increase in HIF-1α levels. Further experiments revealed a reciprocal regulatory mechanism wherein HIF-1α bound to HREs in the Fto promoter, suppressing its expression. In contrast, inhibition or knockdown of HIF-1α restored FTO expression, underlining their dynamic interplay (Wang et al. 2024). These findings are particularly intriguing, as HIF-1 is traditionally associated with transcriptional activation rather than suppression. Interestingly, in fasting induced cardioprotection, FTO levels were markedly increased, while HIF-1α levels were reduced at both transcript and protein levels in the heart (Benak et al. 2024c, d). Despite this downregulation, the m6A/m (m6A + m6Am) methylation levels on Hif1a mRNA remained stable, showing only a non-significant trend toward increased methylation (Benak et al. 2024c). These findings emphasize the intricate relationship between HIF-1 and FTO in cardiovascular contexts.
Another study showed that FTO deficiency in metabolic contexts, such as white adipose tissue browning, increases m6A methylation on Hif1a mRNA. This methylation, recognized by YTHDC2, promotes Hif1a translation and enhances HIF-1α protein levels, thereby activating thermogenic genes. This, in turn, promotes the white-to-beige fat transition and thermogenesis, leading to increased energy expenditure and protection against diet-induced obesity (Wu et al. 2021).
The association between HIF-1 and the third known demethylase, ALKBH3, remains poorly understood. However, one study reported that HIF-1α transcriptionally activates the expression of the lncRNA ALKBH3-AS1, which stabilizes ALKBH3 mRNA and promotes HCC cell proliferation and invasion (Lu et al. 2022).
Currently, all three known m6A erasers have been implicated in the regulation of HIF-1 signaling, drawing attention to their significant roles in cancer progression, energy homeostasis, and cardiovascular resistance to ischemia.
HIF-1 and m6A writers
Since the association of m6A erasers on HIF-1 signaling was established, it was only a matter of time to reveal the involvement of m6A writers as well. Similarly to erasers, these connections were mostly studied in cancer contexts.
METTL3, the most extensively studied m6A writer, has been shown to promote HCC progression by mediating m6A modifications on HIF1A mRNA, resulting in reprogramming of cellular metabolism, as well as enhanced proliferation, invasion and metastasis of liver cancer (Yang et al. 2021a; Zhang et al. 2024). In colorectal cancer, which is characterized by elevated METTL3 and HIF-1α levels, METTL3 knockdown reduced the m6A modification of HIF1A and lowered its translation efficiency, leading to the suppression of the Warburg effect. Moreover, this group also reported that HIF-1α binds to two main HREs in the promoter of METTL3, inducing its expression under hypoxia (Yang et al. 2021b). Similarly, in arecoline-induced oral squamous cell carcinoma, elevated METTL3 expression is stimulated by HIF-1α, forming a positive autoregulatory loop with MYC proto-oncogene that influences both carcinogenesis and cisplatin resistance (Wang et al. 2022). Beyond these direct interactions, METTL3 also promotes the m6A-dependent miR-21-5p maturation, which targets hypoxia-inducible factor 1 subunit alpha inhibitor (HIF1AN; gene name for FIH-1), a negative regulator of HIF-1α. This results in the activation of the HIF-1/VEGF signaling axis, promoting choriocarcinoma progression (Ye et al. 2022). A recent study revealed the role of the HIF-1α/METTL3/YTHDF2 axis in inhibiting the NF-κB/CCL3 pathway, contributing to benzene-induced hematotoxicity (Cong et al. 2024). This suggests that METTL3’s involvement in HIF-1 signaling extends beyond cancer, influencing immune and hematopoietic responses under toxic conditions.
Other subunits of the m6A methyltransferase complex are associated with HIF-1 signaling as well. METTL14-induced ferroptosis, mediated through m6A-YTHDF2-dependent degradation of solute carrier family 7 member 11 (SLC7A11), is dramatically abolished in hypoxic environment in a HIF-1α-dependent manner, contributing to HCC progression (Fan et al. 2021). WTAP, a key subunit of the m6A methyltransferase complex, is upregulated by HIF-1α in ovarian cancer, where it promotes the Warburg effect by stabilizing the mRNA of the glycolytic enzyme hexokinase 2 (HK2) through miR-200 in an m6A-dependent manner, thereby accelerating tumor progression under hypoxic conditions (Lyu et al. 2022). In acute myeloid leukemia (AML), WTAP is similarly up-regulated by HIF-1α and stabilizes lysine demethylase 4B (KDM4B) mRNA via m6A modification, driving cancer cell proliferation and survival (Shao et al. 2023). VIRMA is up-regulated in colon adenocarcinoma, and its silencing in colonic adenocarcinoma cell lines has been shown to inhibit tumor growth by blocking the HIF-1 signaling pathway. However, reactivation of the HIF-1 signaling pathway can counteract the antitumor effects of VIRMA silencing (Ouyang et al. 2024). Similarly, VIRMA is highly expressed in pancreatic ductal adenocarcinoma and is associated with poor prognosis. Mechanistically, VIRMA is promoting signaling through the STRA6/STAT3 axis, leading to elevated HIF-1α levels, which in turn enhances glycolysis and drives tumor progression (Yang et al. 2024).
Currently, multiple m6A writers, including METTL3, METTL14, WTAP, and VIRMA, have been shown to play critical roles in HIF-1 signaling, primarily in cancer progression. Their involvement influences metabolic reprogramming, tumor proliferation, and adaptation to hypoxia, underlining the significance of m6A methylation in regulating HIF-1 signaling and its potential as a therapeutic target in cancer.
HIF-1 and m6A readers
Compared to m6A writers and erasers, the relationship between m6A readers and HIF-1 signaling remains less explored. However, as mediators of RNA modification functions, m6A readers play a crucial role in shaping cellular responses to hypoxia and metabolic stress.
Among key m6A readers, YTHDF1 is directly regulated under hypoxic conditions. HIF-1α induces YTHDF1 expression, which in turn drives hypoxia-induced autophagy and malignancy of HCC by promoting translation of autophagy-related genes ATG2A and ATG14 in a m6A-dependent manner (Li et al. 2021). However, the regulation of YTHDF1 in hypoxia appears complex. In contrast to its upregulation in HCC, YTHDF1 levels are reduced in both the kidneys and liver of highland cattle compared to lowland cattle, suggesting context-dependent negative regulation in hypoxia (Shi et al. 2019). Another key reader, YTHDF2, cooperates with polybromo 1 (PBRM1) to regulate HIF-1α protein translation (Shmakova et al. 2021), which is also promoted by reader YTHDC2 (Wu et al. 2021).
Beyond the YTH domain family, HIF-1α-dependent upregulation of m6A reader IGF2BP1 has been also reported. The study revealed that this axis facilitates peripheral nerve injury recovery by enhancing SLC7A11 mRNA stabilization (An et al. 2023). Reader IGF2BP3 is highly expressed in stomach cancer tissues and hypoxia-treated stomach cancer cells alongside HIF-1α. Mechanistically, IGF2BP3 directly binds to an m6A site within the HIF1A mRNA coding region, positively regulating its expression. Knockdown of IGF2BP3 inhibits hypoxia-induced cell migration and angiogenesis by modulating HIF-1α in stomach cancer (Jiang et al. 2021). Additionally, IGF2BP3 stabilizes HIF1A mRNA promoting its expression during hepatocyte reprogramming in acute-on-chronic liver failure (Cheng et al. 2023).
These findings accentuate the dynamic and context-dependent interaction between HIF-1 signaling and m6A readers, highlighting their critical role in fine-tuning hypoxic adaptation and disease progression.
Collectively, current data bring attention to the intricate crosstalk between HIF-1 signaling and m6A erasers, writers, and readers. A summary of these bidirectional regulatory pathways is provided in Table 1.
Table 1.
Summary of bidirectional regulatory pathways between HIF-1 and m6A RNA modification regulators
| m6A regulator | Cell/tissue type | Regulation | Ref |
|---|---|---|---|
| HIF-1 → regulation of m6A pathway | |||
| ALKBH5 | MCF7, U2OS, and IMR32 cells | hypoxia → HIF-1 ↑ → ALKHBH5 ↑ | Thalhammer et al. (2011) |
| FTO | Neonatal cardiac fibroblasts | hypoxia → HIF-1 ↑ → Fto ↓ | Wang et al. (2024) |
| METTL3 | CRC cell lines | hypoxia → HIF-1 ↑ → METTL3 ↑ | Yang et al. (2021b) |
| Arecoline-induced oral squamous cell carcinoma | HIF-1 ↑ → METTL3 ↑ → tumorigenesis and cisplatin resistance | Wang et al. (2022) | |
| METTL14 | Hepatocellular carcinoma | hypoxia → HIF-1 ↑ → METTL14 ↓ → abrogated ferroptosis and cancer progression | Fan et al. (2021) |
| WTAP | Ovarian cancer | HIF-1 ↑ → WTAP ↑ → Warburg effect and cancer progression | Lyu et al. (2022) |
| Acute myeloid leukemia | HIF-1 ↑ → WTAP ↑ → cancer progression | Shao et al. (2023) | |
| YTHDF1 | Hepatocellular carcinoma | HIF-1 ↑ → YTHDF1 ↑ → autophagy and cancer progression | Li et al. (2021) |
| IGF2BP1 | Dorsal root ganglion neurons | hypoxia → HIF-1 ↑ → IGF2BP1 ↑ → peripheral nerve injury recovery | An et al. (2023) |
| regulation of m6A pathway → HIF-1 | |||
| FTO | Mouse adipose tissue | FTO deletion → m6A levels on Hif1a ↑ → YTHDC2 binding → HIF-1α ↑ → adipocyte browning | Wu et al. (2021) |
| METTL3 | Hepatocellular carcinoma | METTL3 ↑ → m6A levels on HIF1A ↑ → cancer progression | Yang et al. (2021a), Zhang et al. (2024) |
| Colorectal cancer | METTL3 knockdown → m6A levels on HIF1A ↓ → HIF-1α ↓ → Warburg effect ↑ | Yang et al. (2021b) | |
| VIRMA | Colon adenocarcinoma | VIRMA ↑ → VIRMA silencing → HIF-1 signaling ↓ → tumor growth inhibition | Ouyang et al. (2024) |
| Pancreatic ductal adenocarcinoma | VIRMA ↑ → HIF-1α ↑ → cancer progression | Yang et al. (2024) | |
| YTHDF2 | HeLa and H1299 cells | MG132 treatment + YTHDF2 silencing → HIF-1α ↓ | Shmakova et al. (2021) |
| YTHDC2 | Mouse adipose tissue | FTO deletion → m6A levels on Hif1a ↑ → YTHDC2 binding → HIF-1α ↑ → adipocyte browning | Wu et al. (2021) |
| IGF2BP3 | Stomach cancer | IGF2BP3 ↑→ HIF-1α ↑ → hypoxia-induced cell migration and angiogenesis | Jiang et al. (2021) |
| Hep3B and HepG2 cells | IGF2BP3 silencing → HIF-1α ↓ | Cheng et al. (2023) | |
ALKBH5: AlkB homolog 5; CRC: colorectal cancer; FTO: fat mass and obesity-associated; HIF-1/HIF1A: hypoxia-inducible factor 1; IGF2BP1/3: insulin-like growth factor 2 mRNA-binding protein; METTL3/14: methyltransferase-like 3/14; m6A: N6-methyladenosine; VIRMA: protein virilizer homolog; WTAP: Wilms’ tumor 1-associating protein; YTHDC2: YTH domain-containing protein 2; YTHDF1-2: YTH domain-containing family protein 1–2
HIF-1 and other RNA modifications
Besides m6A, the reciprocal relationship between RNA modifications and HIF-1 remains relatively unexplored. Notably, two negative regulators of HIF-1α—the natural antisense transcript HIF1A-AS2 and ubiquitin ligase scaffold LIMD1—have been identified as targets of adenosine-to-inosine (A-to-I) RNA editing, mediated by adenosine deaminase acting on RNA 1 (ADAR1). This ADAR1-dependent modification facilitates the robust and timely accumulation of HIF-1α upon oxygen depletion, thereby reinforcing downstream angiogenesis (Ma et al. 2019). In renal cell carcinoma cells, it has been shown that HIF-1 accumulation leads to downregulation of pseudouridine synthase 10 (PUS10) (Luo et al. 2023), an enzyme responsible for converting uridine to pseudouridine (Ψ) in tRNA (Gurha and Gupta 2008). More recently, a study on colorectal cancer cells reported that HIF-1 binds HRE in the promoter region of methyltransferase METTL1, ultimately leading to inhibition of its transcription and subsequent decrease in the levels of N7-methylguanosine (m7G) in tRNA (Mi et al. 2024).
The demethylase ALKBH1, known for its role in RNA modification such as N1-methyladenosine (m1A), 5-methylcytidine (m5C), or N3-methylcytidine (m3C), has also been implicated in HIF-1 signaling (Zhong et al. 2024; Wu et al. 2019; Guo et al. 2020; Liu et al. 2022). However, ALKBH1 can demethylate not only RNA but also DNA modifications, including N6-methyldeoxyadenosine (6mA) (Zhong et al. 2024). Notably, ALKBH1-mediated 6mA DNA demethylation within the promoter of HIF-1 target gene MIAT (lncRNA myocardial infarction-associated transcript) facilitates HIF-1 binding and enhances its transcriptional activation (Wu et al. 2019). Additionally, direct ALKBH1-mediated regulation of 6mA in the HIF1A gene has been observed (Guo et al. 2020; Liu et al. 2022). These findings suggest that ALKBH1 primarily regulates HIF-1 signaling through epigenetic (DNA) modifications rather than an epitranscriptomic (RNA) mechanism. However, given its broad activity across both RNA and DNA substrates, the potential for RNA modification-mediated effects on HIF-1 remains an open question.
A similar issue arises with ten-eleven translocation (TET) proteins, which function as erasers of m5C in RNA as well as 5-methyl-2′-deoxycytidine (5mdC) in DNA. All TET proteins (TET1-3) have been implicated in hypoxia-related responses (Cheng et al. 2018; Zhang et al. 2022; Cao et al. 2020). However, while their expression is regulated in a HIF-1-dependent manner (Cao et al. 2020; Ma et al. 2019; Hains et al. 2022), the reciprocal TET-HIF axis is mediated by DNA hydroxymethylation regulation rather than epitranscriptomic changes (Cheng et al. 2018; Cao et al. 2020; Ma et al. 2019). Thus, further research is needed to determine whether TET enzymes participate in hypoxia-driven RNA epitranscriptomic modifications, akin to their role in DNA demethylation.
While m6A remains the most studied RNA modification in the context of HIF-1 regulation, emerging evidence suggests that other epitranscriptomic marks, including A-to-I editing, Ψ, and m7G, may also contribute to hypoxia responses. At the same time, enzymes such as ALKBH1 and TET proteins, though primarily studied in DNA modification, may have underexplored roles in RNA demethylation under hypoxia. The reciprocal relationship between HIF-1 and these RNA modifications remains unresolved, highlighting the need for further research to uncover the full extent of epitranscriptomic regulation in hypoxic signaling.
Clinical implications
Hypoxia is a defining characteristic of the tumor microenvironment, arising as rapidly proliferating cancer cells outpace their blood supply. To survive and proliferate under these conditions, cancer cells rely on HIF-1α, a master regulator of the adaptive response to low oxygen. HIF-1α drives the transcription of genes involved in angiogenesis, glycolysis, and cell survival (Semenza 2003). Concurrently, epitranscriptomic modifications, particularly m6A methylation, modulate the HIF-1 signaling pathway, contributing to the malignant progression of tumors. Beyond cancer, epitranscriptomic regulation and the HIF-1 pathway are implicated in hypoxia-driven cardioprotection, such as the adaptive response to ischemia–reperfusion injury. As such, targeting these regulatory systems could present innovative therapeutic strategies to both inhibit cancer progression and address ischemic heart disease—two major global health challenges.
HIF-1α has been a focal point for drug development. Inhibitors such as topotecan (Parmakhtiar et al. 2019) or PX-478 (Jacoby et al. 2010) and activators of PHDs like KRH102053 (Choi et al. 2008) or KRH102140 (Nepal et al. 2011) have shown efficacy in tumor cells. Despite these advances, these compounds often suffer from off-target effects, toxicity to normal cells, low selectivity, and the formation of toxic metabolites. These limitations stress the need for innovative strategies to enhance the specificity and safety of such compounds.
Recent studies have also demonstrated the potential of small molecule inhibitors targeting m6A regulators in vitro and in animal studies (An and Duan 2022). Recently, STC-15, an inhibitor of the m6A writer METTL3, has become the first RNA-modifying enzyme inhibitor to enter clinical trials (NCT05584111) for cancer treatment (Medicine NLo, 2022). However, inhibitors of other m6A regulators have not yet reached this stage. Several hurdles remain before these inhibitors can be effectively translated into clinical practice. Epitranscriptomic regulators influence a broad array of cellular processes beyond their roles in cancer and ischemia, creating a significant risk of off-target effects. The widespread involvement of these enzymes in RNA metabolism makes selective targeting of disease-relevant pathways a critical but unmet need.
The substrate selectivity and dynamics of m6A regulators are still not well understood, raising questions whether their activity depends on sequence specificity (Li et al. 2019), subcellular localization (Relier et al. 2022), or regulation by specific proteins (Nabeel-Shah et al. 2024). Recent work by He et al. (He et al. 2023) elucidated the mechanisms by which m6A is selectively deposited and suppressed on mRNA, emphasizing the role of exon junction complexes (EJCs) in this regulation. Their findings demonstrate that EJCs act as suppressors of m6A methylation, particularly near exon-exon junctions, which limit methylation to certain regions of the mRNA transcript. This regulation ensures that only specific transcripts are targeted for m6A modifications, a process that is critical for maintaining stability and proper expression of mRNAs. Another current study demonstrated that the interaction between FTO and its target transcripts can be modulated by the telomeric zinc finger protein ZBTB48, uncovering a previously unrecognized mechanism that regulates FTO effects on RNA expression (Nabeel-Shah et al. 2024).
Moving forward, comprehensive basic research is paramount to address these challenges. Investigating the interaction between epitranscriptomics and HIF-1 signaling under pathological conditions will enable the identification of precise therapeutic windows. Furthermore, integrating cutting-edge technologies such as single-cell transcriptomics and CRISPR-based screens could accelerate the discovery of disease-specific regulatory networks. Such efforts will be instrumental in transforming promising preclinical findings into viable clinical interventions, offering new treatment for patients with cancer and ischemic heart disease.
Conclusions
The growing body of research on the interplay between RNA modifications, especially m6A, and HIF-1 signaling underscores the complexity of cellular responses to oxygen deprivation. While HIF-1 has long been recognized as a central regulator in hypoxia-driven processes, the recent discovery of its interaction with epitranscriptomic modulators offers new insights into the fine-tuning of gene expression. A particularly intriguing aspect is that many epitranscriptomic regulators not only influence HIF-1 signaling but may also be direct targets of HIF-1 itself. The bidirectional regulation between m6A modifications and HIF signaling not only enhances our understanding of cellular hypoxic responses but also points to potential therapeutic strategies for diseases such as cancer, ischemic heart disease, and other pathologies linked to dysregulated oxygen homeostasis.
Acknowledgements
Figures were created with BioRender.com.
Abbreviations
- 5mdC
5-methyl-2′-deoxycytidine
- 6mA
N6-methyldeoxyadenosine
- ADAR1
Adenosine deaminase acting on RNA 1
- ALDOA
Aldolase A
- ALKBH1/3/5
AlkB homolog 1/3/5
- AML
Acute myeloid leukemia
- ARNT
Aryl hydrocarbon receptor nuclear translocator
- ATG
Autophagy-related genes
- CCDC
Coiled-coil domain-containing
- CCL
Chemokine (C–C motif) ligand
- EJC
Exon junction complex
- EPO
Erythropoietin
- FIH
Factor inhibiting HIF
- FIH-1
Factor inhibiting HIF-1
- FTO
Fat mass and obesity-associated protein
- HCC
Hepatocellular carcinoma
- HIF-1
Hypoxia-inducible factor 1
- HK2
Hexokinase 2
- HRE
Hypoxia-responsive element
- IGF2BP
Insulin-like growth factor 2 mRNA-binding protein
- KDM
Lysine demethylase
- lncRNA
Long non-coding RNA
- MCM3
Minichromosome maintenance complex component 3
- METTL1/3/14
Methyltransferase-like 1/3/14
- MI
Myocardial infarction
- m1A
N1-methyladenosine
- m6A
N6-methyladenosine
- m6Am
N6, 2’-O-dimethyladenosine
- m3C
N3-methylcytidine
- m5C
5-methylcytidine
- m7G
N7-methylguanosine
- MIAT
Myocardial infarction-associated transcript
- NF-κB
Nuclear factor kappa B
- PI3K-Akt
Phosphoinositide 3-kinase–protein kinase B pathway
- PHD
Prolyl hydroxylase domain
- PBRM1
Protein polybromo 1
- pVHL
Von Hippel-Lindau protein
- SLC
Solute carrier family
- snRNA
Small nuclear RNA
- STRA6
Signaling receptor and transporter of retinol STRA6
- TET
Ten-eleven translocation proteins
- VEGF
Vascular endothelial growth factor
- VIRMA
Vir-like m6A methyltransferase associated
- WTAP
Wilms’ tumor 1-associating protein
- YTHDC1-2
YTH domain-containing protein 1–2
- YTHDF1-3
YTH domain-containing family protein 1–3
- ZBTB48
Zinc finger and BTB domain-containing protein 48
- Ψ
Pseudouridine
Author contributions
D.B., P.A., K.H., M.C., B.O., drafted the article and created the figures. F.K., G.P., and M.H. provided substantive revisions.
Funding
This work was supported by the Czech Science Foundation (Grant Number 24-10497S) to M.H.; the Charles University Grant Agency (Grant Numbers 80824 and 270623) to M.C. and B.O.; the Ministry of Education, Youth and Sports of the CR [Grant Inter-COST LUC24089], and the project National Institute for Research of Metabolic and Cardiovascular Diseases (Programme EXCELES, ID project No. LX22NPO5104)—Funded by the European Union—Next Generation EU.
Availability of data and materials
No datasets were generated or analysed during the current study.
Declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- Alanova P, et al. HIF-1α limits myocardial infarction by promoting mitophagy in mouse hearts adapted to chronic hypoxia. Acta Physiol (Oxf). 2024;240(9):e14202 [DOI] [PubMed]
- An Y, Duan H. The role of m6A RNA methylation in cancer metabolism. Mol Cancer. 2022;21(1):14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- An S, Shi J, Huang J, Li Z, Feng M, Cao G. HIF-1α-induced upregulation of m6A reader IGF2BP1 facilitates peripheral nerve injury recovery by enhancing SLC7A11 mRNA stabilization. In Vitro Cell Dev Biol Anim. 2023;59(8):596–605. [DOI] [PubMed] [Google Scholar]
- Batista PJ, Molinie B, Wang J, Qu K, Zhang J, Li L, et al. m(6)A RNA modification controls cell fate transition in mammalian embryonic stem cells. Cell Stem Cell. 2014;15(6):707–19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benak D, Kolar F, Hlavackova M. Epitranscriptomic regulations in the heart. Physiol Res. 2024a. 10.33549/physiolres.935265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benak D, Sevcikova A, Holzerova K, Hlavackova M. FTO in health and disease. Front Cell Dev Biol. 2024b;12:1500394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benak D, Holzerova K, Hrdlicka J, Kolar F, Olsen M, Karelson M, et al. Epitranscriptomic regulation in fasting hearts: implications for cardiac health. RNA Biol. 2024c;21(1):1–14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benak D, Holzerova K, Kolar F, Chalupova M, Hlavackova M. Unveiling the proteome of the fasting heart: insights into HIF-1 pathway regulation. Front Physiol. 2024d;15:1462014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bohuslavova R, Cerychova R, Papousek F, Olejnickova V, Bartos M, Görlach A, et al. HIF-1α is required for development of the sympathetic nervous system. Proc Natl Acad Sci USA. 2019;116(27):13414–23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cao JZ, Liu H, Wickrema A, Godley LA. HIF-1 directly induces TET3 expression to enhance 5-hmC density and induce erythroid gene expression in hypoxia. Blood Adv. 2020;4(13):3053–62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cappannini A, Ray A, Purta E, Mukherjee S, Boccaletto P, Moafinejad SN, et al. MODOMICS: a database of RNA modifications and related information: 2023 update. Nucleic Acids Res. 2024;52(D1):D239–44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Catrina SB, Zheng X. Hypoxia and hypoxia-inducible factors in diabetes and its complications. Diabetologia. 2021;64(4):709–16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cheng Y, Sun M, Chen L, Li Y, Lin L, Yao B, et al. Ten-eleven translocation proteins modulate the response to environmental stress in mice. Cell Rep. 2018;25(11):3194-203.e4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cheng K, Liu S, Li C, Zhao Y, Wang Q. IGF2BP3/HIF1A/YAP signaling plays a role in driving acute-on-chronic liver failure through activating hepatocyte reprogramming. Cell Signal. 2023;108:110727. [DOI] [PubMed] [Google Scholar]
- Choi HJ, Song BJ, Gong YD, Gwak WJ, Soh Y. Rapid degradation of hypoxia-inducible factor-1alpha by KRH102053, a new activator of prolyl hydroxylase 2. Br J Pharmacol. 2008;154(1):114–25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cohn WE. Pseudouridine, a carbon-carbon linked ribonucleoside in ribonucleic acids: isolation, structure, and chemical characteristics. J Biol Chem. 1960;235:1488–98. [PubMed] [Google Scholar]
- Cong X, Li X, Xu K, Yin L, Liang G, Sun R, et al. HIF-1α/m(6)A/NF-κB/CCL3 axis-mediated immunosurveillance participates in low level benzene-related erythrohematopoietic development toxicity. Environ Int. 2024;184:108493. [DOI] [PubMed] [Google Scholar]
- Davis FF, Allen FW. Ribonucleic acids from yeast which contain a fifth nucleotide. J Biol Chem. 1957;227(2):907–15. [PubMed] [Google Scholar]
- Desrosiers R, Friderici K, Rottman F. Identification of methylated nucleosides in messenger RNA from Novikoff hepatoma cells. Proc Natl Acad Sci USA. 1974;71(10):3971–5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Epstein AC, Gleadle JM, McNeill LA, Hewitson KS, O’Rourke J, Mole DR, et al. C. elegans EGL-9 and mammalian homologs define a family of dioxygenases that regulate HIF by prolyl hydroxylation. Cell. 2001;107(1):43–54. [DOI] [PubMed] [Google Scholar]
- Fan Z, Yang G, Zhang W, Liu Q, Liu G, Liu P, et al. Hypoxia blocks ferroptosis of hepatocellular carcinoma via suppression of METTL14 triggered YTHDF2-dependent silencing of SLC7A11. J Cell Mol Med. 2021;25(21):10197–212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Forsythe JA, Jiang BH, Iyer NV, Agani F, Leung SW, Koos RD, et al. Activation of vascular endothelial growth factor gene transcription by hypoxia-inducible factor 1. Mol Cell Biol. 1996;16(9):4604–13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fukuda R, Zhang H, Kim JW, Shimoda L, Dang CV, Semenza GL. HIF-1 regulates cytochrome oxidase subunits to optimize efficiency of respiration in hypoxic cells. Cell. 2007;129(1):111–22. [DOI] [PubMed] [Google Scholar]
- Guo Y, Pei Y, Li K, Cui W, Zhang D. DNA N(6)-methyladenine modification in hypertension. Aging. 2020;12(7):6276–91. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gurha P, Gupta R. Archaeal Pus10 proteins can produce both pseudouridine 54 and 55 in tRNA. RNA. 2008;14(12):2521–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haase VH. Hypoxic regulation of erythropoiesis and iron metabolism. Am J Physiol Renal Physiol. 2010;299(1):F1-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hains AE, Uppal S, Cao JZ, Salwen HR, Applebaum MA, Cohn SL, et al. MYCN and HIF-1 directly regulate TET1 expression to control 5-hmC gains and enhance neuroblastoma cell migration in hypoxia. Epigenetics. 2022;17(13):2056–74. [DOI] [PMC free article] [PubMed] [Google Scholar]
- He PC, Wei J, Dou X, Harada BT, Zhang Z, Ge R, et al. Exon architecture controls mRNA m(6)A suppression and gene expression. Science. 2023;379(6633):677–82. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hlavackova M, Benak D, Sotakova D, Cyprova M, Telensky P, Bendova Z, et al. 4007 Fat mass and obesity-associated protein in chronically hypoxic myocardium. High Alt Med Biol. 2018;19(4):A-443. [Google Scholar]
- Hsu PJ, Zhu Y, Ma H, Guo Y, Shi X, Liu Y, et al. Ythdc2 is an N(6)-methyladenosine binding protein that regulates mammalian spermatogenesis. Cell Res. 2017;27(9):1115–27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang H, Weng H, Sun W, Qin X, Shi H, Wu H, et al. Recognition of RNA N(6)-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nat Cell Biol. 2018;20(3):285–95. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jacoby JJ, Erez B, Korshunova MV, Williams RR, Furutani K, Takahashi O, et al. Treatment with HIF-1alpha antagonist PX-478 inhibits progression and spread of orthotopic human small cell lung cancer and lung adenocarcinoma in mice. J Thorac Oncol. 2010;5(7):940–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jia G, Fu Y, Zhao X, Dai Q, Zheng G, Yang Y, et al. N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat Chem Biol. 2011;7(12):885–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jiang L, Li Y, He Y, Wei D, Yan L, Wen H. Knockdown of m6A reader IGF2BP3 inhibited hypoxia-induced cell migration and angiogenesis by regulating hypoxia inducible factor-1α in stomach cancer. Front Oncol. 2021;11:711207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kierans SJ, Taylor CT. Regulation of glycolysis by the hypoxia-inducible factor (HIF): implications for cellular physiology. J Physiol. 2021;599(1):23–37. [DOI] [PubMed] [Google Scholar]
- Koivunen P, Hirsilä M, Günzler V, Kivirikko KI, Myllyharju J. Catalytic properties of the asparaginyl hydroxylase (FIH) in the oxygen sensing pathway are distinct from those of its prolyl 4-hydroxylases. J Biol Chem. 2004;279(11):9899–904. [DOI] [PubMed] [Google Scholar]
- Koyasu S, Kobayashi M, Goto Y, Hiraoka M, Harada H. Regulatory mechanisms of hypoxia-inducible factor 1 activity: two decades of knowledge. Cancer Sci. 2018;109(3):560–71. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lasman L, Krupalnik V, Viukov S, Mor N, Aguilera-Castrejon A, Schneir D, et al. Context-dependent functional compensation between Ythdf m(6)A reader proteins. Genes Dev. 2020;34(19–20):1373–91. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li Y, Wu K, Quan W, Yu L, Chen S, Cheng C, et al. The dynamics of FTO binding and demethylation from the m(6)A motifs. RNA Biol. 2019;16(9):1179–89. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li Q, Ni Y, Zhang L, Jiang R, Xu J, Yang H, et al. HIF-1α-induced expression of m6A reader YTHDF1 drives hypoxia-induced autophagy and malignancy of hepatocellular carcinoma by promoting ATG2A and ATG14 translation. Signal Transduct Target Ther. 2021;6(1):76. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liang Z, Riaz A, Chachar S, Ding Y, Du H, Gu X. Epigenetic modifications of mRNA and DNA in plants. Mol Plant. 2020;13(1):14–30. [DOI] [PubMed] [Google Scholar]
- Liang L, Zhu Y, Li J, Zeng J, Wu L. ALKBH5-mediated m6A modification of circCCDC134 facilitates cervical cancer metastasis by enhancing HIF1A transcription. J Exp Clin Cancer Res. 2022;41(1):261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Y, Chen Y, Wang Y, Jiang S, Lin W, Wu Y, et al. DNA demethylase ALKBH1 promotes adipogenic differentiation via regulation of HIF-1 signaling. J Biol Chem. 2022;298(1):101499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Longenecker JZ, Gilbert CJ, Golubeva VA, Martens CR, Accornero F. Epitranscriptomics in the heart: a focus on m(6)A. Curr Heart Fail Rep. 2020;17(5):205–12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu Q, Wang H, Lei X, Ma Q, Zhao J, Sun W, et al. LncRNA ALKBH3-AS1 enhances ALKBH3 mRNA stability to promote hepatocellular carcinoma cell proliferation and invasion. J Cell Mol Med. 2022;26(20):5292–302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu X, Li D, Lin Z, Gao T, Gong Z, Zhang Y, et al. HIF-1α-induced expression of the m6A reader YTHDF1 inhibits the ferroptosis of nucleus pulposus cells by promoting SLC7A11 translation. Aging Cell. 2024;23(9): e14210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luo W, Xu Z, Wang H, Lu Z, Ding L, Wang R, et al. HIF1A-repressed PUS10 regulates NUDC/Cofilin1 dependent renal cell carcinoma migration by promoting the maturation of miR-194-5p. Cell Biosci. 2023;13(1):153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lyu Y, Zhang Y, Wang Y, Luo Y, Ding H, Li P, et al. HIF-1α regulated WTAP overexpression promoting the Warburg effect of ovarian cancer by m6A-dependent manner. J Immunol Res. 2022;2022:6130806. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ma L, Qi T, Wang S, Hao M, Sakhawat A, Liang T, et al. Tet methylcytosine dioxygenase 1 promotes hypoxic gene induction and cell migration in colon cancer. J Cell Physiol. 2019;234(5):6286–97. [DOI] [PubMed] [Google Scholar]
- Ma CP, Liu H, Yi-Feng Chang I, Wang WC, Chen YT, Wu SM, et al. ADAR1 promotes robust hypoxia signaling via distinct regulation of multiple HIF-1α-inhibiting factors. EMBO Rep. 2019;20(5): e47107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Makino Y, Cao R, Svensson K, Bertilsson G, Asman M, Tanaka H, et al. Inhibitory PAS domain protein is a negative regulator of hypoxia-inducible gene expression. Nature. 2001;414(6863):550–4. [DOI] [PubMed] [Google Scholar]
- Medicine NLo. Oral administration of STC-15 in subjects with advanced malignancies (NCT05584111) 2022–2025. 2022. https://clinicaltrials.gov/study/NCT05584111?term=NCT05584111&rank=1.
- Meier JC, Kankowski S, Krestel H, Hetsch F. RNA editing-systemic relevance and clue to disease mechanisms? Front Mol Neurosci. 2016;9:124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mi S, Cai S, Xue M, Wu W. HIF-1α/METTL1/m(7)G axis is involved in CRC response to hypoxia. Biochem Biophys Res Commun. 2024;693:149385. [DOI] [PubMed] [Google Scholar]
- Monaci S, Coppola F, Filippi I, Falsini A, Carraro F, Naldini A. Targeting hypoxia signaling pathways in angiogenesis. Front Physiol. 2024;15:1408750. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nabeel-Shah S, Pu S, Burke GL, Ahmed N, Braunschweig U, Farhangmehr S, et al. Recruitment of the m(6)A/m6Am demethylase FTO to target RNAs by the telomeric zinc finger protein ZBTB48. Genome Biol. 2024;25(1):246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nan Y, Liu S, Luo Q, Wu X, Zhao P, Chang W, et al. m(6)A demethylase FTO stabilizes LINK-A to exert oncogenic roles via MCM3-mediated cell-cycle progression and HIF-1α activation. Cell Rep. 2023;42(10):113273. [DOI] [PubMed] [Google Scholar]
- Nepal M, Gong YD, Park YR, Soh Y. An activator of PHD2, KRH102140, decreases angiogenesis via inhibition of HIF-1α. Cell Biochem Funct. 2011;29(2):126–34. [DOI] [PubMed] [Google Scholar]
- Niu Y, Zhao X, Wu YS, Li MM, Wang XJ, Yang YG. N6-methyl-adenosine (m6A) in RNA: an old modification with a novel epigenetic function. Genomics Proteomics Bioinform. 2013;11(1):8–17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Niu Y, Lin Z, Wan A, Sun L, Yan S, Liang H, et al. Loss-of-function genetic screening identifies aldolase a as an essential driver for liver cancer cell growth under hypoxia. Hepatology. 2021;74(3):1461–79. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ong SG, Hausenloy DJ. Hypoxia-inducible factor as a therapeutic target for cardioprotection. Pharmacol Ther. 2012;136(1):69–81. [DOI] [PubMed] [Google Scholar]
- Ong SG, Lee WH, Theodorou L, Kodo K, Lim SY, Shukla DH, et al. HIF-1 reduces ischaemia-reperfusion injury in the heart by targeting the mitochondrial permeability transition pore. Cardiovasc Res. 2014;104(1):24–36. [DOI] [PubMed] [Google Scholar]
- Ouyang C, Xu G, Xie J, Xie Y, Zhou Y. Silencing of KIAA1429, a N6-methyladenine methyltransferase, inhibits the progression of colon adenocarcinoma via blocking the hypoxia-inducible factor 1 signalling pathway. J Biochem Mol Toxicol. 2024;38(9): e23829. [DOI] [PubMed] [Google Scholar]
- Parmakhtiar B, Burger RA, Kim JH, Fruehauf JP. HIF inactivation of p53 in ovarian cancer can be reversed by topotecan, restoring cisplatin and paclitaxel sensitivity. Mol Cancer Res. 2019;17(8):1675–86. [DOI] [PubMed] [Google Scholar]
- Peixoto P, Cartron PF, Serandour AA, Hervouet E. From 1957 to nowadays: a brief history of epigenetics. Int J Mol Sci. 2020;21(20):7571. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pezzuto A, Carico E. Role of HIF-1 in cancer progression: novel insights. A Review Curr Mol Med. 2018;18(6):343–51. [DOI] [PubMed] [Google Scholar]
- Ping XL, Sun BF, Wang L, Xiao W, Yang X, Wang WJ, et al. Mammalian WTAP is a regulatory subunit of the RNA N6-methyladenosine methyltransferase. Cell Res. 2014;24(2):177–89. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Relier S, Ripoll J, Guillorit H, Amalric A, Achour C, Boissière F, et al. FTO-mediated cytoplasmic m(6)A(m) demethylation adjusts stem-like properties in colorectal cancer cell. Nat Commun. 2021;12(1):1716. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Relier S, Rivals E, David A. The multifaceted functions of the Fat mass and Obesity-associated protein (FTO) in normal and cancer cells. RNA Biol. 2022;19(1):132–42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Semenza GL. Hypoxia-inducible factor 1: control of oxygen homeostasis in health and disease. Pediatr Res. 2001;49(5):614–7. [DOI] [PubMed] [Google Scholar]
- Semenza GL. Targeting HIF-1 for cancer therapy. Nat Rev Cancer. 2003;3(10):721–32. [DOI] [PubMed] [Google Scholar]
- Semenza GL. Hypoxia-inducible factor 1 and cardiovascular disease. Annu Rev Physiol. 2014;76:39–56. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Semenza GL. A compendium of proteins that interact with HIF-1α. Exp Cell Res. 2017;356(2):128–35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Semenza GL, Nejfelt MK, Chi SM, Antonarakis SE. Hypoxia-inducible nuclear factors bind to an enhancer element located 3’ to the human erythropoietin gene. Proc Natl Acad Sci USA. 1991;88(13):5680–4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sendinc E, Shi Y. RNA m6A methylation across the transcriptome. Mol Cell. 2023;83(3):428–41. [DOI] [PubMed] [Google Scholar]
- Shao YL, Li YQ, Li MY, Wang LL, Zhou HS, Liu DH, et al. HIF1α-mediated transactivation of WTAP promotes AML cell proliferation via m(6)A-dependent stabilization of KDM4B mRNA. Leukemia. 2023;37(6):1254–67. [DOI] [PubMed] [Google Scholar]
- Shi H, Wang X, Lu Z, Zhao BS, Ma H, Hsu PJ, et al. YTHDF3 facilitates translation and decay of N(6)-methyladenosine-modified RNA. Cell Res. 2017;27(3):315–28. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shi Y, Fan S, Wu M, Zuo Z, Li X, Jiang L, et al. YTHDF1 links hypoxia adaptation and non-small cell lung cancer progression. Nat Commun. 2019;10(1):4892. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shi X, Cao Y, Zhang X, Gu C, Liang F, Xue J, et al. Comprehensive analysis of N6-methyladenosine RNA methylation regulators expression identify distinct molecular subtypes of myocardial infarction. Front Cell Dev Biol. 2021;9:756483. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shmakova A, Frost M, Batie M, Kenneth NS, Rocha S. PBRM1 cooperates with YTHDF2 to control HIF-1α protein translation. Cells. 2021;10(6):1425. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun R, Yuan L, Jiang Y, Wan Y, Ma X, Yang J, et al. ALKBH5 activates FAK signaling through m6A demethylation in ITGB1 mRNA and enhances tumor-associated lymphangiogenesis and lymph node metastasis in ovarian cancer. Theranostics. 2023;13(2):833–48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sweaad WK, Stefanizzi FM, Chamorro-Jorganes A, Devaux Y, Emanueli C. Relevance of N6-methyladenosine regulators for transcriptome: implications for development and the cardiovascular system. J Mol Cell Cardiol. 2021;160:56–70. [DOI] [PubMed] [Google Scholar]
- Thalhammer A, Bencokova Z, Poole R, Loenarz C, Adam J, O’Flaherty L, et al. Human AlkB homologue 5 is a nuclear 2-oxoglutarate dependent oxygenase and a direct target of hypoxia-inducible factor 1α (HIF-1α). PLoS ONE. 2011;6(1): e16210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tolonen JP, Heikkilä M, Malinen M, Lee HM, Palvimo JJ, Wei GH, et al. A long hypoxia-inducible factor 3 isoform 2 is a transcription activator that regulates erythropoietin. Cell Mol Life Sci. 2020;77(18):3627–42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ueda Y, Ooshio I, Fusamae Y, Kitae K, Kawaguchi M, Jingushi K, et al. AlkB homolog 3-mediated tRNA demethylation promotes protein synthesis in cancer cells. Sci Rep. 2017;7:42271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang GL, Semenza GL. Purification and characterization of hypoxia-inducible factor 1. J Biol Chem. 1995;270(3):1230–7. [DOI] [PubMed] [Google Scholar]
- Wang GL, Jiang BH, Rue EA, Semenza GL. Hypoxia-inducible factor 1 is a basic-helix-loop-helix-PAS heterodimer regulated by cellular O2 tension. Proc Natl Acad Sci USA. 1995;92(12):5510–4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang X, Zhao BS, Roundtree IA, Lu Z, Han D, Ma H, et al. N(6)-methyladenosine modulates messenger RNA translation efficiency. Cell. 2015;161(6):1388–99. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang P, Doxtader KA, Nam Y. Structural basis for cooperative function of Mettl3 and Mettl14 methyltransferases. Mol Cell. 2016a;63(2):306–17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang X, Feng J, Xue Y, Guan Z, Zhang D, Liu Z, et al. Structural basis of N(6)-adenosine methylation by the METTL3-METTL14 complex. Nature. 2016b;534(7608):575–8. [DOI] [PubMed] [Google Scholar]
- Wang C, Kadigamuwa C, Wu S, Gao Y, Chen W, Gu Y, et al. RNA N6-methyladenosine (m6A) methyltransferase-like 3 facilitates tumorigenesis and cisplatin resistance of arecoline-exposed oral carcinoma. Cells. 2022;11(22):3605. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Y, Traugot CM, Bubenik JL, Li T, Sheng P, Hiers NM, et al. N(6)-methyladenosine in 7SK small nuclear RNA underlies RNA polymerase II transcription regulation. Mol Cell. 2023;83(21):3818-34.e7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang J, Li Y, Deng L, Zha Y, Zhang S. FTO suppresses cardiac fibrosis after myocardial infarction via m(6)A-mediated epigenetic modification of EPRS. Mol Med. 2024;30(1):213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Warnecke C, Zaborowska Z, Kurreck J, Erdmann VA, Frei U, Wiesener M, et al. Differentiating the functional role of hypoxia-inducible factor (HIF)-1alpha and HIF-2alpha (EPAS-1) by the use of RNA interference: erythropoietin is a HIF-2alpha target gene in Hep3B and Kelly cells. FASEB J. 2004;18(12):1462–4. [DOI] [PubMed] [Google Scholar]
- Wei J, Liu F, Lu Z, Fei Q, Ai Y, He PC, et al. Differential m(6)A, m(6)A(m), and m(1)A demethylation mediated by FTO in the cell nucleus and cytoplasm. Mol Cell. 2018;71(6):973-85.e5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wiesener MS, Jürgensen JS, Rosenberger C, Scholze CK, Hörstrup JH, Warnecke C, et al. Widespread hypoxia-inducible expression of HIF-2alpha in distinct cell populations of different organs. FASEB J. 2003;17(2):271–3. [DOI] [PubMed] [Google Scholar]
- Wu L, Pei Y, Zhu Y, Jiang M, Wang C, Cui W, et al. Association of N(6)-methyladenine DNA with plaque progression in atherosclerosis via myocardial infarction-associated transcripts. Cell Death Dis. 2019;10(12):909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu R, Chen Y, Liu Y, Zhuang L, Chen W, Zeng B, et al. m6A methylation promotes white-to-beige fat transition by facilitating Hif1a translation. EMBO Rep. 2021;22(11): e52348. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xiao W, Adhikari S, Dahal U, Chen YS, Hao YJ, Sun BF, et al. Nuclear m(6)A reader YTHDC1 regulates mRNA splicing. Mol Cell. 2016;61(4):507–19. [DOI] [PubMed] [Google Scholar]
- Yang N, Wang T, Li Q, Han F, Wang Z, Zhu R, et al. HBXIP drives metabolic reprogramming in hepatocellular carcinoma cells via METTL3-mediated m6A modification of HIF-1α. J Cell Physiol. 2021a;236(5):3863–80. [DOI] [PubMed] [Google Scholar]
- Yang Z, Quan Y, Chen Y, Huang Y, Huang R, Yu W, et al. Knockdown of RNA N6-methyladenosine methyltransferase METTL3 represses Warburg effect in colorectal cancer via regulating HIF-1α. Signal Transduct Target Ther. 2021b;6(1):89. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang K, Zhong Z, Zou J, Liao JY, Chen S, Zhou S, et al. Glycolysis and tumor progression promoted by the m(6)A writer VIRMA via m(6)A-dependent upregulation of STRA6 in pancreatic ductal adenocarcinoma. Cancer Lett. 2024;590:216840. [DOI] [PubMed] [Google Scholar]
- Ye K, Li L, Wu B, Wang D. METTL3 m6A-dependently promotes miR-21-5p maturation to accelerate choriocarcinoma progression via the HIF1AN-induced inactivation of the HIF1A/VEGF pathway. Genes Genomics. 2022;44(11):1311–22. [DOI] [PubMed] [Google Scholar]
- Yue Y, Liu J, Cui X, Cao J, Luo G, Zhang Z, et al. VIRMA mediates preferential m(6)A mRNA methylation in 3’UTR and near stop codon and associates with alternative polyadenylation. Cell Discov. 2018;4:10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zaccara S, Jaffrey SR. A unified model for the function of YTHDF proteins in regulating m(6)A-modified mRNA. Cell. 2020;181(7):1582-95.e18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zagórska A, Dulak J. HIF-1: the knowns and unknowns of hypoxia sensing. Acta Biochim Pol. 2004;51(3):563–85. [PubMed] [Google Scholar]
- Zhang C, Samanta D, Lu H, Bullen JW, Zhang H, Chen I, et al. Hypoxia induces the breast cancer stem cell phenotype by HIF-dependent and ALKBH5-mediated m⁶A-demethylation of NANOG mRNA. Proc Natl Acad Sci USA. 2016a;113(14):E2047–56. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang C, Zhi WI, Lu H, Samanta D, Chen I, Gabrielson E, et al. Hypoxia-inducible factors regulate pluripotency factor expression by ZNF217- and ALKBH5-mediated modulation of RNA methylation in breast cancer cells. Oncotarget. 2016b;7(40):64527–42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang B, Jiang H, Dong Z, Sun A, Ge J. The critical roles of m6A modification in metabolic abnormality and cardiovascular diseases. Genes Dis. 2021;8(6):746–58. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang X, Li S, He J, Jin Y, Zhang R, Dong W, et al. TET2 suppresses VHL deficiency-driven clear cell renal cell carcinoma by inhibiting HIF signaling. Cancer Res. 2022;82(11):2097–109. [DOI] [PubMed] [Google Scholar]
- Zhang H, Han B, Tian S, Gong Y, Liu L. ZNF740 facilitates the malignant progression of hepatocellular carcinoma via the METTL3/HIF-1A signaling axis. Int J Oncol. 2024;65(5):105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zheng G, Dahl JA, Niu Y, Fedorcsak P, Huang CM, Li CJ, et al. ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol Cell. 2013;49(1):18–29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhong J, Xu Z, Ding N, Wang Y, Chen W. The biological function of demethylase ALKBH1 and its role in human diseases. Heliyon. 2024;10(13): e33489. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
No datasets were generated or analysed during the current study.