Abstract
Human epidermal growth factor receptor 2 (HER2, also known as ERBB2) signaling promotes cell growth and differentiation, and is overexpressed in several tumor types, including breast, gastric and colorectal cancer. HER2-targeted therapies have shown clinical activity against these tumor types, resulting in regulatory approvals. However, the efficacy of HER2 therapies in tumors with HER2 mutations has not been widely investigated. SGNTUC-019 is an open-label, phase 2 basket study evaluating tucatinib, a HER2-targeted tyrosine kinase inhibitor, in combination with trastuzumab in patients with HER2-altered solid tumors. The study included a cohort of 31 heavily pretreated female patients with HER2-mutated metastatic breast cancer who were also HER2 negative per local testing. Hormone receptor (HR)-positive patients also received fulvestrant. The overall response rate (primary endpoint) was 41.9% (90% confidence interval (CI): 26.9–58.2). Secondary endpoints of duration of response and progression-free survival were 12.6 months (90% CI: 4.7 to not estimable) and 9.5 months (90% CI: 5.4–13.8), respectively. No new safety signals were detected. Responses were observed across various HER2 mutations, including mutations in the tyrosine kinase and extracellular domains. The chemotherapy-free regimen of tucatinib and trastuzumab showed clinically meaningful antitumor activity with durable responses and favorable tolerability in heavily pretreated patients with HER2 mutations. These data support further investigation of HER2-targeted therapies in this patient population. ClinicalTrials.gov registration: NCT04579380.
Subject terms: Breast cancer, Breast cancer
In this phase 2 basket trial, patients with heavily pretreated HER2-mutated metastatic breast cancer received the small-molecule HER2 inhibitor tucatinib and anti-HER2 trastuzumab, leading to encouraging clinical response rates.
Main
HER2 is a member of the epidermal growth factor receptor family of four related tyrosine kinases. When activated, HER2 signals through the phosphoinositide 3-kinase (PI3K) and mitogen-activated protein kinase (MAPK) pathways to promote cell growth and differentiation. In tumors in which HER2 is overexpressed or amplified (HER2-positive, HER2+), activation results in uncontrolled cell proliferation, inhibition of apoptosis and metastasis1–6. HER2 is a validated, actionable target, with HER2-targeted therapies showing clinical activity that resulted in regulatory approvals for the treatment of patients with HER2+ breast, gastric and colorectal cancers7–15.
Recently, with the increased utilization of genomic profiling through next-generation sequencing (NGS), there has been a growing interest in HER2 (also known as ERBB2) somatic mutations as oncogenic drivers, in the absence of HER2+ disease, in multiple solid tumors. In breast cancer, approximately 2–5% of tumors harbor HER2 mutations16–21. HER2-mutated (HER2-mut) breast cancer is more frequently associated with lobular histology and heavily pretreated patients with metastatic disease22–24. HER2-targeted agents have shown clinical activity in HER2-mut metastatic breast cancer (MBC) in basket trials and case reports25–27. However, there are currently no treatments that have been approved by the US Food and Drug Administration (FDA) or the European Medicines Agency specifically for patients with HER2-mut MBC.
Tucatinib is an oral tyrosine kinase inhibitor that is highly selective for HER2 (ref. 28) and approved in multiple geographic regions for treatment of previously treated HER2+ MBC (combined with trastuzumab and capecitabine)11,29–31 and previously treated HER2+ metastatic colorectal cancer (combined with trastuzumab)12. In addition, preclinical data have shown that tucatinib in combination with trastuzumab has antitumor activity in HER2-mut patient-derived xenograft tumor models of multiple solid tumors, suggesting that patients with activating HER2 mutations may derive clinical benefit from the combination32. SGNTUC-019 (NCT04579380) is an open-label phase 2 basket study evaluating tucatinib in combination with trastuzumab in patients with HER2-altered solid tumors. Here we report results of the efficacy, safety and exploratory biomarker analyses from the HER2-mut MBC cohort of tucatinib in combination with trastuzumab (and fulvestrant in patients who have hormone-receptor-positive (HR+) disease).
Results
Patient characteristics
The first patient was enrolled in the basket study on 18 January 2021, and the last patient enrolled on 14 April 2023. Thirty-one patients were enrolled in the HER2-mut MBC cohort from September 2021 to December 2022. Patients with HER2-mut MBC were enrolled if they had measurable disease, had progressed during or after at least one previous line of systemic therapy in the locally advanced or metastatic setting, and had progressed during or were intolerant of the most recent line of systemic therapy; patients with HER2+ MBC were excluded. Demographics and baseline characteristics of patients with HER2-mut MBC are shown in Table 1. The median age was 64.0 years (range 43–76 years). Tumor histology was 58% lobular and 42% ductal. Most patients were HR+ (87%). Seven patients (23%) had stage IV disease at initial diagnosis. The median number of previous lines of therapy in any setting was 4 (range 1–10) and 3 (range 1–7) in the locally advanced or metastatic setting. Seventeen patients (55%) had previously received fulvestrant before enrollment.
Table 1.
Patient demographics and baseline characteristics of the HER2-mut MBC cohort
| Characteristics | Total (N = 31) |
|---|---|
| Median age, years (range) | 64.0 (43, 76) |
| Age category, n (%) | |
| <65 years | 16 (52) |
| ≥65 years | 15 (48) |
| Sex, n (%) | |
| Male | 0 |
| Female | 31 (100) |
| Ethnicity, n (%) | |
| Hispanic, Latino or Latina, or of Spanish origin | 2 (6) |
| Not Hispanic, Latino or Latina, or of Spanish origin | 24 (77) |
| Unknown | 1 (3) |
| Not reportable | 4 (13) |
| Race, n (%) | |
| Asian | 11 (35) |
| Black or African American | 1 (3) |
| White | 15 (48) |
| Not reportable | 4 (13) |
| ECOG performance status score, n (%) | |
| 0 | 19 (61) |
| 1 | 12 (39) |
| Hormone receptor status | |
| Negative | 4 (13) |
| Positive | 27 (87) |
| Tumor type histology | |
| Ductal | 13 (42) |
| Lobular | 18 (58) |
| TNM stage at time of diagnosis | |
| I | 5 (16) |
| II | 6 (19) |
| III | 13 (42) |
| IV | 7 (23) |
| History of metastatic disease | |
| Yes | 31 (100) |
| No | 0 |
| Number of previous systemic therapy regimens, median (range) | 4 (1, 10) |
| Number of previous systemic therapy regimens in the locally advanced or metastatic setting, median (range) | 3 (1, 7) |
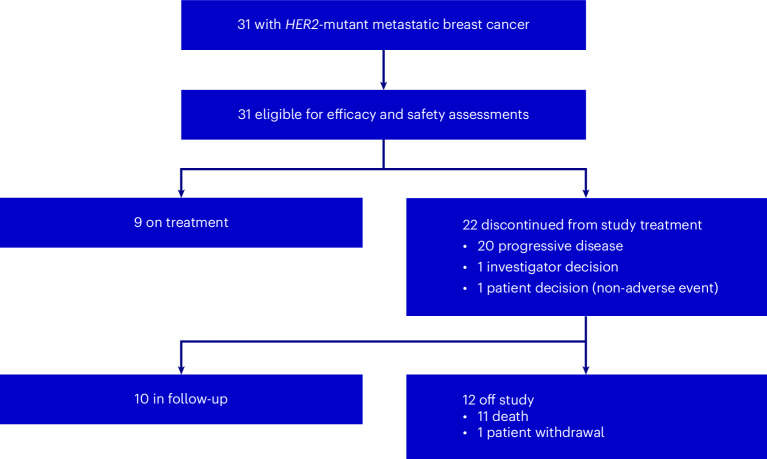
Patient disposition is summarized in Fig. 1. As of the data cutoff (1 November 2023), the median duration of follow-up was 15.0 months (90% CI: 13.1–17.5). Nine patients remain on study treatment, 10 are in long-term follow-up and 12 are off study. Of the 22 patients who discontinued study treatment, 20 discontinued because of progressive disease and one patient each stopped treatment because of investigator decision or patient decision. No patient discontinued all study treatments because of a treatment-emergent adverse event (TEAE). Of the 12 patients that were off study, 11 were off study owing to death.
Fig. 1. CONSORT diagram of the SGNTUC-019 study.
Flow diagram detailing patients enrolled in the SGNTUC-019 study (NCT04579380) HER2-mutated MBC cohort.
Efficacy
The confirmed objective response rate (cORR) per investigator assessment was 41.9% (90% CI: 26.9–58.2; Table 2), including 2 confirmed complete responses. Patients with HR+ or HR− MBC had a cORR of 44.4% (12 out of 27; 90% CI: 28.0–61.8) and 25.0% (1 out of 4; 90% CI: 1.3–75.1), respectively. Of the 31 patients, 13 were classified with ductal histology and 18 were classified with lobular histology. cORR was similar in both groups: 5 out of 13 for ductal histology (38.5% (90% CI: 16.6–64.5)) and 8 out of 18 for lobular histology (44.4% (90% CI: 24.4–65.9)). The median duration of response (DOR) was 12.6 months (90% CI: 4.7 to not estimable), and the median time to first response was 1.4 months (range 1.2–6.2). The disease control rate was 80.6% (90% CI: 65.3–91.2). Tumor size decreased in 22 patients (73%) of the 30 with a baseline and at least one post-baseline assessment (Fig. 2).
Table 2.
Summary of responses in the HER2-mut MBC cohort
| Total (N = 31) | |
|---|---|
| Best overall responsea, n (%) | |
| CR | 2 (6.5) |
| PR | 11 (35.5) |
| SD | 12 (38.7) |
| PD | 4 (12.9) |
| Not availableb | 2 (6.5) |
| cORR, n (%) | 13 (41.9) |
| 90% CIc for cORR | (26.9, 58.2) |
| Median duration of objective responsed (months) (90% CI)c | 12.6 (4.7, −) |
| Disease control ratee, n (%) | 25 (80.6) |
| 90% CIc for disease control rate | (65.3, 91.2) |
| Median PFS (months) (90% CI)f | 9.5 (5.4, 13.8) |
| Median OS (months) (90% CI)f | 20.1 (15.9, −) |
CI, confidence interval; cORR, confirmed objective response rate; CR, complete response; OS, overall survival; PFS, progression-free survival; PR, partial response; SD, stable disease; PD, progressive disease.
In the case of SD, follow-up measurements must have met the SD criteria at least once after study entry at a minimum interval of 5 weeks.
aConfirmed best overall response assessed per Response Evaluation Criteria in Solid Tumors (RECIST) v1.1.
bIncludes patients whose disease assessments were not evaluable and patients with no post-baseline response assessment.
cTwo-sided 90% exact CI, computed using the Clopper–Pearson method40.
dAs estimated using Kaplan–Meier methods.
eDisease control defined as confirmed CR or confirmed PR or SD.
fCalculated using the complementary log–log transformation method41.
Fig. 2. Response to study treatment and HER2 biomarker assessments.
a, Maximum percentage reduction in the sum of the diameters of target lesions per investigator and HER2 biomarker assessments. Only patients who had baseline and post-baseline target lesion measurements are included (n = 30). The asterisks indicate patients that were excluded from any analysis of local results because their eligibility (HER2-negative status and HER2 mutations) was determined by central blood-based NGS (B-NGS). T-NGS denotes tissue-based NGS testing by central lab assay. In all assays, ‘ND’ (not determined) indicates that there was no assay result, which may have been due to failed assay, failed quality control or no assay run due to sample issues, and ‘NE’ denotes non-evaluable. In the ‘Amp’ (amplification) panel, ‘central’ indicates overall HER2-amplification status as determined by IHC with FISH reflex, both by central lab testing; whereas ‘local’ denotes local testing results. In the ‘Domain’ panel, patients had mutations exclusively in the domains indicated (that is, the tyrosine kinase (TK) or extracellular domains of HER2). The dashed line at 20% indicates progressive disease (PD). The dashed line at −30% indicates the threshold for a partial response (PR). D, ductal; L, lobular. b, Duration of treatment of patients in the HER2-mut MBC cohort.
The median progression-free survival (PFS) was 9.5 months (90% CI: 5.4–13.8), with an estimated 6 month PFS of 65.0% (90% CI: 48.3–77.5) and an estimated 12 month PFS of 45.0% (90% CI: 28.8–60.0; Fig. 3a). The median overall survival (OS) was 20.1 months (90% CI: 15.9 to not estimable; Fig. 3b), with an estimated 12 month OS of 74.2% (90% CI: 58.6–84.7).
Fig. 3. Kaplan–Meier curves for PFS and OS per investigator in the HER2-mut MBC cohort.
The vertical lines indicate censoring.
Safety
The median treatment duration with tucatinib was 9.0 months (range 0.7–24.4); with trastuzumab, 9.0 months (range 0.7–24.4); and with fulvestrant, 7.4 months (range 0.9–24.4). TEAEs were reported in all patients (Table 3 and Extended Data Table 1). The most common TEAEs were diarrhea (20 patients, 65%), nausea (11, 35%), vomiting and pruritus (9, 29%), and infusion-related reaction (8, 26%; Table 3). Among the 20 patients reported to have diarrhea, 4 patients (13%) had maximum severity of grade 3, while 12 patients (39%) and 4 patients (13%) had grade 1 and grade 2 diarrhea, respectively. The median time to resolution was 7.0 days (range 1–530) for any-grade diarrhea and 2.0 days (range 1–4) for grade ≥3 diarrhea. The other most common grade ≥3 events were increased levels of alanine aminotransferase and hypertension, with 3 patients (10%) reported for each. A total of 8 patients (26%) had serious TEAEs, of which 3 (10%) were related to tucatinib and none were related to either trastuzumab or fulvestrant.
Table 3.
The most common TEAEs in the HER2-mut MBC cohort
| Preferred term | Total (N = 31) | |
|---|---|---|
| Any grade, n (%) | Grade ≥3a, n (%) | |
| Any TEAE | 31 (100) | 15 (48) |
| Diarrhea | 20 (65) | 4 (13) |
| Nausea | 11 (35) | 0 |
| Vomiting | 9 (29) | 0 |
| Pruritus | 9 (29) | 0 |
| Infusion-related reaction | 8 (26) | 0 |
| Back pain | 7 (23) | 2 (6) |
| Cough | 7 (23) | 0 |
| Edema peripheral | 7 (23) | 0 |
| Chills | 6 (19) | 0 |
| Pyrexia | 6 (19) | 0 |
| Appetite decreased | 6 (19) | 2 (6) |
| Fatigue | 6 (19) | 1 (3) |
| Arthralgia | 6 (19) | 1 (3) |
| Dry skin | 6 (19) | 0 |
| Alanine aminotransferase increased | 5 (16) | 3 (10) |
| Aspartate aminotransferase increased | 5 (16) | 2 (6) |
| Hypertension | 5 (16) | 3 (10) |
| Constipation | 4 (13) | 0 |
| Insomnia | 4 (13) | 0 |
| Blood creatinine increased | 3 (10) | 0 |
| Anemia | 3 (10) | 0 |
| Abdominal pain | 3 (10) | 0 |
| Dyspnea | 3 (10) | 0 |
| Headache | 3 (10) | 0 |
| COVID-19 | 3 (10) | 0 |
| Stomatitis | 3 (10) | 0 |
| Dizziness | 3 (10) | 0 |
| Gastroesophageal reflux disease | 3 (10) | 0 |
| Blood bilirubin increased | 3 (10) | 0 |
| Myalgia | 3 (10) | 0 |
| Onychoclasis | 3 (10) | 0 |
TEAEs reported in ≥10.0% of the patients who received at least a single dose of the study drug are listed here. The events are reported as per the preferred terms in the Medical Dictionary for Regulatory Activities.
aGrade ≥3 TEAEs not listed here included acute kidney injury, cellulitis, chronic kidney disease, hepatic function abnormal, hydronephrosis, hypokalemia, influenza, sepsis and systemic inflammatory response syndrome; each was reported in one patient (3%).
Extended Data Table 1.
Summary of safety events in the HER2-mut MBC cohort
No TEAEs resulted in the discontinuation of all study treatments. Tucatinib was discontinued because of TEAEs in 2 patients (6%), including an event of increased aspartate aminotransferase (grade 4) and an event of pseudocirrhosis (grade 2). There were no discontinuations of trastuzumab or fulvestrant due to TEAEs. TEAEs resulted in dose reduction of tucatinib in 7 patients (23%) and fulvestrant in 1 patient (3%). No TEAEs resulted in death. All 11 deaths (35%) were related to disease progression.
Exploratory biomarker analyses
Among the 31 patients enrolled in the HER2-mut MBC cohort, 28 had evaluable central immunohistochemistry (IHC) results, 27 had evaluable central fluorescence in situ hybridization (FISH) results, 24 had evaluable central tissue-based NGS results and 29 had evaluable blood-based NGS results. Responses were observed in patients with a range of HER2 mutations, including mutations in both the tyrosine kinase and the extracellular domains (Fig. 2a). A listing of individual responses and HER2 mutations as determined by local and central testing is found in Extended Data Table 2.
Extended Data Table 2.
Listing of responses, histology, and HER2 mutations by local and central testing
HER2 alterations and response
Among patients with a mutation identified by local testing (n = 28), central blood-based NGS (n = 23) or central tissue-based NGS (n = 20), HER2 mutations (both single nucleotide variants and insertions and deletions (indels)) were most often located exclusively in the tyrosine kinase domain or exclusively in the extracellular domain. Depending on the type of assay used, 69.6–85.0% of patients had mutations exclusively in the tyrosine kinase domain and 15.0–21.4% of patients had mutations exclusively in the extracellular domain (Fig. 2a and Extended Data Table 3). The range of response rates across assays was 36.4–41.2% in patients with mutations exclusively in the tyrosine kinase domain and 50.0–66.7% in patients with mutations exclusively in the extracellular domain (Fig. 2a and Extended Data Table 3).
Extended Data Table 3.
HER2 altered domains: prevalence and response rates
The most commonly mutated amino acid residues in the HER2-mut MBC cohort were at amino acid residues Leu755 (a tyrosine kinase domain residue) and Ser310 (an extracellular domain residue). Depending on the type of assay used, 40.0–47.8% of patients had a mutation at Leu755, and 15.0–26.1% of patients had a mutation at Ser310 (Extended Data Fig. 1 and Extended Data Table 4). As this analysis was based on the presence or absence of an individual mutation, some patients had co-occurring HER2 mutations. The range of response rates across assays was 33.3–37.5% in patients with a mutation in Leu755 and 50.0–66.7% in patients with a mutation in Ser310 (Extended Data Fig. 1 and Extended Data Table 4).
Extended Data Fig. 1. Response versus presence of L755 and/or S310F mutations.
B-NGS, blood-based NGS testing by central lab assay; CR, complete response; D, ductal; L, lobular; Local, local testing results; Mut, Mutation; ND, not determined; NE, non-evaluable; NGS, next-generation sequencing; PD, progressive disease; PR, partial response; SD, stable disease; T-NGS, tissue-based NGS testing by central lab assay. Only subjects who had baseline and post-baseline target lesion measurements are included (n = 30). * indicates patients that were excluded from any analysis of local results because their eligibility (HER2-negative status and HER2 mutations) was determined by central blood-based NGS.
Extended Data Table 4.
HER2 L755 and S310 mutations: prevalence and response rates
Results showed a roughly equal distribution of patients who were HER2-low (IHC 1+ or IHC 2+/FISH-negative, n = 12) and those without HER2 expression (IHC 0, n = 14). Based on central testing, the response rate was 50.0% (6 out of 12; 90% CI: 24.5–75.5) in HER2-low patients and 42.9% (6 out of 14; 90% CI: 20.6–67.5) in patients without HER2 expression.
HER2 assay agreement
In current practice, multiple types of assay are used to identify HER2 alterations. To determine whether patient outcomes were impacted by this variability, results were analyzed among patients with sufficient samples available for standardized central testing. The percentage agreement for detection of HER2 mutations (detected versus not detected) was 76.9% (90% CI: 59.5–89.4, n = 26) between local and central blood-based NGS, and 85.7% (90% CI: 67.1–96.0, n = 21) between local and central tissue-based NGS (Fig. 2a, Extended Data Fig. 1 and Extended Data Table 2). There was 100% agreement between any local test for HER2 overexpression or amplification status versus central blood-based NGS (90% CI: 89.1–100, n = 26) and central tissue-based NGS (90% CI: 86.7–100, n = 21) (Fig. 2a).
Concomitant non-HER2 alterations
Additional concomitant non-HER2 alterations (that is, amplified and/or with mutations or indels) identified using central tissue-based and blood-based NGS are shown in Extended Data Figs. 2 and 3. Owing to the small dataset (24 and 29 evaluable results in the tissue-based and blood-based NGS data, respectively), we examined genes that were altered in at least 25% of patients in a given assay. In the tissue-based NGS dataset, CDH1, characteristic of invasive lobular carcinoma, and PIK3CA were the only genes altered in at least 25% of patients. Responses were observed in patients with and without alterations in CDH1 and PIK3CA. Response rates among patients with and without CDH1 alterations were 62.5% (9 out of 15; 90% CI: 36.0–80.9) and 22.2% (2 out of 9; 90% CI: 4.1–55.0), respectively. Response rates among patients with and without PIK3CA alterations were 25.0% (2 out of 8; 90% CI: 4.6–60.0) and 56.3% (9 out of 16; 90% CI: 33.3–77.3), respectively. In the blood-based NGS dataset, only PIK3CA was altered in at least 25% of patients. Patients with and without alterations in PIK3CA showed responses of 36.4% (4 out of 11; 90% CI: 13.5–65.0) and 44.4% (8 out of 18; 90% CI: 24.4–65.9), respectively.
Extended Data Fig. 2. Concomitant genomic alterations from central tissue-based NGS.
CNV, copy number variant; D, ductal; INDEL, insertion and deletion; L, lobular; ND, not determined; NGS, next-generation sequencing; SNV, single nucleotide variant.
Extended Data Fig. 3. Waterfall plot vs concomitant gene alterations from central blood-based NGS.
CNV, copy number variant; CR, complete response; D, ductal; INDEL, insertion and deletion; L, lobular; ND, not determined; NE, non-evaluable; NGS, next-generation sequencing; PD, progressive disease; PR, partial response; SD, stable disease; SNV, single nucleotide variant. Only subjects who had baseline and post-baseline target lesion measurements are included (n = 30).
Discussion
Over the past 25 years, monoclonal antibodies, antibody–drug conjugates and tyrosine kinase inhibitors targeting HER2 have been developed that have resulted in notable improvements in PFS and OS for patients with HER2+ MBC7,8,11,13,33,34. In the past decade, the use of NGS assays to molecularly profile and identify other potential targetable alterations in patients with MBC has increased substantially35. HER2 somatic mutations occur in about 2–5% of all patients with breast cancer, with a higher prevalence in the advanced disease setting16–21,35. However, the benefit of using HER2-targeted therapies in patients with HER2-mut MBC, in the absence of HER2+ disease, is not well established; early data using HER2-targeted agents in this patient population have shown evidence of clinical activity25.
In the SGNTUC-019 study, the combination of tucatinib and trastuzumab (with fulvestrant in patients with HR+ disease) is active in patients with heavily pretreated HER2-mut MBC, with a cORR of 41.9%. Moreover, the responses were durable, with a median DOR of 12.6 months. The response to study treatment occurred early, with the median time to first response of 1.4 months corresponding with the first tumor assessment on treatment. In addition, the median PFS and OS were 9.5 months (90% CI: 5.4–13.8) and 20.1 months (90% CI: 15.9 to not estimable), respectively. This clinically meaningful activity was seen despite patients having been heavily pretreated with a median of four previous lines of systemic therapy in any setting and three previous lines in the locally advanced or metastatic setting. This further adds to the data from the SUMMIT trial, which showed that HER2 mutations are potentially actionable targets in MBC; treatment with neratinib in combination with fulvestrant and trastuzumab for patients with HR+, HER2-mut MBC (n = 57) showed similar clinical activity (cORR, 39%, and median PFS, 8.3 months)27.
In patients with tumor tissue available, central testing showed that almost all patients either were HER2-low (IHC 2+/FISH-negative or IHC 1+) or lacked expression of HER2 (IHC 0). Responses to study treatment were similar in both patient populations, suggesting that the HER2 mutation is an oncogenic driver in those patients.
The combination of tucatinib and trastuzumab, with fulvestrant for HR+ patients, was well tolerated and consistent with the established safety profile of the combination36,37. Diarrhea was the most commonly reported adverse event; however, most events were grade 1 in severity and manageable with standard clinical care, with a grade 3 occurrence of 13%. Furthermore, antidiarrheal prophylaxis was not required on the study and no discontinuations of tucatinib occurred because of diarrhea. By contrast, grade 3 diarrhea occurred in 53% (30 out of 57) of patients receiving the combination of neratinib, fulvestrant and trastuzumab in the SUMMIT trial, despite the patients receiving loperamide prophylaxis27. Only two patients (6%) discontinued tucatinib because of TEAEs, but remained on trastuzumab. No patients discontinued all study treatments because of an adverse event, highlighting the tolerability of this chemotherapy-free approach in patients with previous exposure to multiple lines of therapy and associated toxicities.
Exploratory biomarker analyses showed responses in patients harboring a variety of HER2 mutations. There was evidence of clinical activity in patients with mutations in the tyrosine kinase domain and extracellular domain of HER2, specifically with mutations at Leu755 and Ser310, two of the most frequently mutated amino acids in HER2 in multiple cancers, including breast cancers38,39. Moderate-to-strong agreement between local and central test methods was observed, suggesting that both tissue-based and blood-based NGS may be used to identify patients harboring HER2 mutations who may respond to treatment with tucatinib and trastuzumab. Finally, a variety of concomitant non-HER2 genomic alterations were observed, including alterations in CDH1 and PIK3CA. Clinical activity was observed across patients with a wide variety of alterations; however, larger datasets are required to draw any conclusions about the impact of concomitant non-HER2 mutations on response.
One limitation of this analysis is the small size of the cohort (31 patients in total) and lack of a comparator arm. In addition, the duration of follow-up was short; therefore, the OS data are immature with relatively few events and should be interpreted with caution. Conclusions based on the exploratory biomarker data are also limited by the relatively small size of the cohort and the availability of data across multiple central testing methods (24 patients with evaluable tissue-based NGS and 29 with blood-based NGS). Conclusions may also be affected by assay differences between tissue-based and blood-based NGS, notably by differences in the set of genes included in each assay panel, as well as differential timing of tissue and blood collection in many cases.
In summary, the combination of tucatinib and trastuzumab (with fulvestrant in HR+ disease) was well tolerated and clinically active in patients with previously treated HER2-mut MBC. These results support further investigation of HER2-directed therapies in patients with HER2 mutations.
Methods
Inclusion and ethics
The study was conducted in accordance with regulatory requirements, the International Council for Harmonisation Good Clinical Practice guidelines and the Declaration of Helsinki. All patients provided written informed consent. The protocol was approved by institutional review boards and ethics committees according to the practice at each participating study site (Supplementary Information).
Patients and study design
SGNTUC-019 (NCT04579380) is an open-label, phase 2 basket study of patients with previously treated, locally advanced, unresectable or metastatic HER2-altered solid tumors. The breast cancer cohort comprised patients with HER2-mut locally advanced or metastatic disease, without evidence of overexpression or amplification of HER2, and enrolled patients at academic hospitals or cancer centers in North America, Europe and Asia. Patients must have progressed during or after at least one previous line of systemic therapy in the locally advanced or metastatic disease setting. In addition, patients must have progressed or be intolerant of the most recent line of systemic therapy. Patients with HR+ disease must have received a cyclin-dependent kinase (CDK4/6) inhibitor in the metastatic setting. Patients with previous exposure to fulvestrant were eligible. All patients were required to have evidence of measurable disease as per Response Evaluation Criteria in Solid Tumors (RECIST) v1.1 on baseline imaging. Patients with an Eastern Cooperative Oncology Group (ECOG) performance status of 0 or 1 and adequate baseline cardiac, hepatic, renal and hematologic function were eligible. Patients previously treated with any systemic anticancer therapy, radiation therapy, major surgery or experimental agent within 3 weeks of the first dose of study treatment were excluded. In addition, patients must not have received previous HER2-directed therapy. Study enrollment criteria did not include any requirements regarding sex and/or gender. Patient sex was based on self-identification at baseline. Full inclusion and exclusion criteria are available in the protocol, which has been previously published37.
Eligible patients had HER2 mutations identified by local or central NGS. Local testing for identification of HER2 mutations could be via archival or fresh tumor tissue or blood. Central testing was through blood-based NGS (Guardant360 CDx Test, Guardant). HER2 mutations eligible for inclusion in the cohort were prespecified. Patients with HER2 mutations that were not prespecified were also potentially eligible, if supported by scientific literature.
Procedures
Patients in the HER2-mut MBC cohort were treated with tucatinib 300 mg orally twice daily and trastuzumab 8 mg kg−1 intravenously then 6 mg kg−1 every 3 weeks in a 21 day cycle. Patients with HR+ disease also received fulvestrant 500 mg intramuscularly once every 4 weeks starting from cycle 1 day 1, and once on cycle 1 day 15. Disease response to the study treatment and the occurrence of disease progression were determined according to RECIST v1.1, as assessed by the investigator. Disease assessments were performed at baseline, every 6 weeks for the first 24 weeks and then every 12 weeks until the occurrence of documented disease progression per RECIST v1.1, death, withdrawal of consent, loss to follow-up or study closure.
Safety was assessed by the incidence of TEAEs, graded according to the National Cancer Institute Common Terminology Criteria for Adverse Events v5.0, recording of concomitant medications, physical examination findings, vital signs, laboratory tests, pregnancy testing and cardiac function. Cardiac ejection fraction was assessed via echocardiogram or a multigated acquisition scan at screening and every 12 weeks thereafter.
For the exploratory biomarker assessments, central HER2 testing was performed in a laboratory accredited by Clinical Laboratory Improvement Amendments. Patients’ blood samples were collected during prescreening or on day 1 of cycle 1 for central testing using a 74-gene blood-based NGS assay (Guardant360 CDx Test, Guardant). Tumor tissue biopsies, archival or fresh (if available), were collected during screening. Central HER2 testing of tumor tissue was performed using IHC (PATHWAY HER-2, Roche), FISH (HER2 IQFISH pharmDx Assay, Agilent) and 505-gene tissue-based NGS assay (Elio Tissue Complete, PGDx). IHC and FISH results were evaluated using the American Society of Clinical Oncology and College of American Pathologists Breast Scoring criteria42.
Assessments
The primary endpoint was cORR, defined as the proportion of patients with confirmed complete response or partial response according to RECIST v1.1, per investigator assessment. Secondary endpoints included disease control rate, DOR, PFS by investigator assessment, OS and safety. The exploratory endpoints included biomarker assessments as described in the section ‘Exploratory biomarker data assessment and analyses’.
Statistical analysis
The HER2-mut MBC cohort aimed to enroll 30 response-evaluable patients, a number calculated per the 90% exact CI given a range of expected cORR of 10–30%. One more patient was accrued than planned because one patient was initially deemed not response evaluable but was later found to be response evaluable. An interim analysis was not planned for the HER2-mut MBC cohort. All enrolled patients received at least one dose of tucatinib and trastuzumab and were included in the evaluation for efficacy and safety. Two-sided 90% exact CIs for response rates were calculated using the Clopper–Pearson method. Median PFS and OS were estimated using the Kaplan–Meier method; the associated 90% CI was calculated on the basis of the complementary log–log transformation. Safety and concordance of local versus central HER2 testing results were assessed by descriptive statistics. All analyses except the biomarker analyses were performed with SAS, version 9.4 (SAS Institute). Biomarker analyses were performed with R, version 4.0.2 (R Core Team and the R Foundation for Statistical Computing). Because all patients in the study cohort were female, no sex- or gender-based analyses have been performed.
Exploratory biomarker data assessment and analyses
Central testing methods
Blood-based NGS analyses of amplifications and sequence alterations (mutations and indels) were from the standard analysis pipelines used by Guardant (Guardant360 CDx Test). Similarly, tissue-based NGS determination of amplifications and sequence alterations were from internal pipelines used by PGDx (Elio Tissue Complete). A further analysis step was the removal of synonymous mutations, except those indicated as potentially affecting a splice region. For blood-based NGS testing, we removed TP53 from the analyses as this gene’s mutations may be from clonal hematopoiesis of indeterminate potential as opposed to reflecting tumor state.
Assignment of domains for HER2 mutation
Consistent with previous approaches, the following definitions were used to assign mutations by amino acid residue number to protein in HER2: 52–643 for the extracellular domain and 694–883 for the tyrosine kinase domain39.
HER2 assay agreement
In the analyses of local versus central testing, local refers to any test to determine HER2 mutations, or overexpression or amplification status, performed at the local site. Three patients used results from central blood-based NGS rather than local testing to establish HER2 mutational and amplification status for eligibility. They were excluded from these analyses, and their local results are labeled as not determined when data are shown.
HER2 status, defined as the final HER2 overexpression or amplification status, could be either positive, negative or not determined. For central IHC and FISH testing, HER2 status was determined as follows: HER2-negative status (IHC 0, IHC 1 or IHC 2+ with FISH negative), HER2-positive status (IHC 2+ with FISH-positive or IHC 3+ with any FISH result) and HER2 status ‘not determined’ (IHC 2+ without evaluable FISH results).
To determine the HER2 overexpression or amplification status across assays, pairwise comparisons were performed using data from patients who had results (that is, positive or negative) from both assays. The results are reported as percentage agreement, with two-sided 90% exact CIs calculated using the Clopper–Pearson method40.
HER2 mutation detection agreement was calculated using the same methods described above for the HER2 overexpression or amplification status. These comparisons were based on simple HER2 ‘mutation detected’ versus HER2 ‘mutation not detected’.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Online content
Any methods, additional references, Nature Portfolio reporting summaries, source data, extended data, supplementary information, acknowledgements, peer review information; details of author contributions and competing interests; and statements of data and code availability are available at 10.1038/s41591-024-03462-0.
Supplementary information
List of institutional review boards and independent ethics committees.
Acknowledgements
We thank the patients who participated in this study, their families, and the investigators and staff at the SGNTUC-019 clinical sites. We also thank A. Drees, K. Earley and S. Lonning of Pfizer for critical review of the data and the paper and K. Garton of Pfizer for critical review of the paper. Finally, we thank the entire SGNTUC-019 study team for support. H. Brignull of Populus Group, supported by Pfizer, provided medical writing and editorial support in accordance with Good Publication Practice guidelines. This study was sponsored by Seagen, in collaboration with Merck Sharp & Dohme LLC, a subsidiary of Merck & Co. Seagen was acquired by Pfizer in December 2023. F. Hoffmann-La Roche provided trastuzumab for the study. The sponsor had final responsibility for the design of the trial, protocol, database maintenance, trial conduct, data analyses and confirmation of the accuracy of the data. We prepared and approved the paper and made the decision to submit it for publication.
Extended data
Author contributions
A.F.C.O., T.B.-S., B.J.M., Y.N., D.M.O., M.R., E.Y.Y., J.R., S. Tan, M.B., T.E.S. and P.R.P. contributed to the study conception or design. All authors contributed to the acquisition, analysis or interpretation of the data and the drafting or substantial revision of the paper.
Peer review
Peer review information
Nature Medicine thanks the anonymous reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Ulrike Harjes, in collaboration with the Nature Medicine team.
Data availability
Following requests and subject to review, Pfizer will provide the data that support the findings of this study. Subject to certain criteria, conditions and exceptions, Pfizer may also provide access to the related individual de-identified participant data. See https://www.pfizer.com/science/clinical-trials/trial-data-and-results for more information.
Competing interests
A.F.C.O. reports research funding from Pfizer and Roche; honoraria for presentations from AstraZeneca, Esai, Gilead, Lilly, Pfizer, Roche and Seagen; advisory board fees from AstraZeneca, Pfizer, Roche and Seagen; and conference support from AstraZeneca, Lilly, Novartis and Roche. G.C. reports consulting or advisory roles with AstraZeneca, Celucity, Daiichi Sankyo, Ellipsis, Exact Sciences, Gilead, Lilly, MBS, Menarini, Merck, Pfizer, Roche and Veracyte; institutional research funding from Astellas, AstraZeneca, Blueprint Medicine, BMS, Daiichi Sankyo, Kymab, Merck, Novartis, Philogen, Relay Therapeutics, Roche and Sanofi; speaker fees from AstraZeneca, Daiichi Sankyo, Novartis, Pfizer and Roche; and writing engagement with Pfizer. N.M. reports grants or contracts to the institution from AstraZeneca, Boehringer Ingelheim, Incyte, MSD, Novartis, Ono Pharmaceutical, Pfizer and Seagen; payment or honoraria for lectures, presentations, speakers’ bureaus, paper writing or educational events from AstraZeneca, FujiFilm Toyama Chemical, Miyarisan Pharmaceutical, Novartis, Taiho Pharmaceutical and Yakult Honsha; and data safety monitoring or advisory board roles with AstraZeneca and Boehringer Ingelheim. D.-Y.O. reports research funding from Array BioPharma, AstraZeneca, BeiGene, Eli Lilly, Handok, MSD, Novartis and Servier, and consulting or advisory roles with Arcus Biosciences, ASLAN Pharmaceuticals, AstraZeneca, Basilea, Bayer, BeiGene, Celgene, Genentech/Roche, Halozyme, IQVIA, Merck Serono, MSD Oncology, Novartis, Taiho Pharmaceutical, Turning Point Therapeutics, Yuhan and Zymeworks. H.S. reports institutional research funding from Amgen; consulting fees from AstraZeneca, Celgene, Eisai, Novartis, PUMA, Seattle Genetics, Sanofi and Sermonix; advisory board roles with AstraZeneca, Eisai, Eli Lilly, Novartis and PUMA; speaker fees from Merck; and licensing fees for intellectual property from Celyad Oncology. S. Takahashi reports grants and/or personal fees from AstraZeneca, Bayer, Chugai, Daiichi Sankyo, Eisai, MSD, Novartis and Taiho. T.B.-S. reports institutional research funding from Abgenomics, Agios, Arcus, Arys, Atreca, Bayer, BMS, Boston Biomedical, Celgene, Clovis, Eisai, Genentech, Incyte, Ipsen, Lilly, Merus, Mirati, Novartis, Pfizer and Seagen; institutional consulting fees from Arcus, Bayer, Eisai, Incyte, Ipsen, Genentech, Merck KGaA, Merck, Merus, Pfizer, Seagen and Servier; personal consulting fees from AbbVie, Aptitude Health, AstraZeneca, Beigene, Blueprint Medicines, Boehringer Ingelheim, Caladrius Biosciences, Celularity, Daiichi Sankyo, Deciphera, Exact Science, Exelixis, Foundation Medicine, GlaxoSmithKline, Illumina, Janssen, Kanaph, MJH Life Sciences, Natera, Sanofi, Sobi, Stemline, TreosBio, Xilio and Zai Labs; IDMC/DSMB roles with 1Globe, AstraZeneca, Exelixis, Fibrogen, Merck/Eisai, PanCan, Suzhou Kintor and The Valley Hospital; consulting or advisory board roles with Artiva, Bard, Immuneering, Imugene, Replimune, Sun Biopharma and Xilis; and patents WO/2018/183488: Human PD1 peptide vaccines and uses thereof, licensed to Imugene, and WO/2019/055687: Methods and compositions for the treatment of cancer cachexia, licensed to Recursion. M.E.B. reports consulting or advisory roles with LifeOmic, Novartis and Strata Oncology; institutional research funding from AbbVie, Apollomics, Arcus Ventures, Elevation Oncology, Endeavor BioMedicines, Genentech, Loxo, Merck, Puma Biotechnology, Seagen and Strata Oncology; travel, accommodations and expenses from LifeOmic; and patents, royalties and other intellectual property: patents for (1) an implantable and localized drug delivery device that can sample the tumor microenvironment and deliver the drug, (2) a method to detect recombination events with CRISPR-mediated editing and (3) expansion microscopy without specialized equipment. K.Y.C. reports institutional research funding from Epizyme+L20 and MedImmune. P.R.D. reports institutional grants from Pfizer; advisory board roles with Astellas Pharma, BMS, Ipsen, Merck and Pfizer; honoraria for lectures from Bayer; travel support from Janssen; and a role as a substitute board member for the Clinical Trials College, Federal Public Service, Kingdom of Belgium; and holds stocks in Alkermes, Mural Oncology PLC and Biocartis Group NV. V.G. reports research funding from Bayer, Boehringer and Roche and institutional funding from Amcure, Astelas, AstraZeneca, Bayer, BeiGene, BMS, FibroGen, Genentech, Lilly, Medimmune, Merck Serono, MSD, Natera, Novartis, Roche, Servier, Sierra Oncology and Takeda. M.G.-M. reports a consulting or advisory role with AstraZeneca and travel, accommodation and expenses from GlaxoSmithKline and MSD Oncology. E.P.H. reports advisory and consulting roles (all payments to the institution) with Accutar Biotechnology, AstraZeneca, Daiichi Sankyo, Eli Lilly, Ellipses Pharma, Entos, Fosun Pharma, Genentech/Roche, Gilead Sciences, Jazz Pharmaceuticals, Jefferies LLC, Medical Pharma Services, Mersana, Novartis, Olema Pharmaceuticals, Pfizer, Stemline Therapeutics, Tempus Labs, Theratechnologies, Tubulis, Verascity Science and Zentalis Pharmaceuticals and institutional research funding from Abbvie, Accutar Biotechnology, Acerta Pharma, ADC Therapeutics, AKESOBIO Australia, Amgen, Aravive, ArQule, Artios, Arvinas, AstraZeneca, AtlasMedx, BeiGene, Black Diamond, Bliss BioPharmaceuticals, Boehringer Ingelheim, Bristol Myers Squibb, Cascadian Therapeutics, Clovis, Compugen, Context Therapeutics, Cullinan, Curis, CytomX, Daiichi Sankyo, Dana-Farber Cancer Institute, Dantari, Deciphera, Duality Biologics, eFFECTOR Therapeutics, Eisai, Eli Lilly, Ellipses Pharma, Elucida Oncology, EMD Serono, Fochon Pharmaceuticals, FujiFilm, G1 Therapeutics, Genentech/Roche, Gilead Sciences, H3 Biomedicine, Harpoon, Hutchinson MediPharma, Immunogen, Immunomedics, Incyte, Infinity Pharmaceuticals, Inspirna, InventisBio, Jacobio, Karyopharm, K-Group Beta, Kind Pharmaceuticals, Leap Therapeutics, Loxo Oncology, Lycera, Mabspace Biosciences, Macrogenics, MedImmune, Mersana, Merus, Millennium, Molecular Templates, Myriad Genetic Laboratories, Novartis, Nucana, Olema, OncoMed, Oncothyreon, ORIC Pharmaceuticals, Orinove, Orum Therapeutics, Pfizer, PharmaMar, Pieris Pharmaceuticals, Pionyr Immunotherapeutics, Plexxikon, Prelude Therapeutics, ProfoundBio, Radius Health, Regeneron, Relay Therapeutics, Repertoire Immune Medicine, Rgenix, SeaGen, Sermonix Pharmaceuticals, Shattuck Labs, Silverback Therapeutics, StemCentRx, Stemline Therapeutics, Sutro, Syndax, Syros, Taiho, TapImmune, Tesaro, Tolmar, Torque Therapeutics, Treadwell Therapeutics, Verastem, Zenith Epigenetics and Zymeworks. B.J.M. reports honorarium for consulting or speaking from Acrivon, Adaptimmune, Agenus, Akeso Bio, Amgen, AstraZeneca, Biohaven, BMS, Corcept, Easai, Eli Lilly, Genalux, Genmab/Seagen/Pfizer, GOG Foundation, Gradalis, GSK, Heng Rui, Immunogen/Abbvie, Iovance, Karyopharm, Merck, Mersana, Mural/Alkermes, Myriad, Novartis, Novocure, OncoC4, Panavance, ProfoundBio, Regeneron, Roche/Genentech, Sutro, Verastem, Zentalis and Zymeworks. Y.N. reports consulting or advisory roles with Daiichi Sankyo, Exact Sciences, Gilead Sciences, Guardant Health, Natera, Premo Partners, Roche, Seagen and Takeda; participation in speakers’ bureau for Becton Dickinson, CareNet, Chugai Pharma, Daiichi Sankyo, Eisai, Guardant Health, Guardant Health Japan, Hisamitsu Pharmaceutical, Merck, Miyarisan Pharmaceutical, MSD KK, Taiho Pharmaceutical and Zeria Pharmaceutical; and institutional research funding from Chugai Pharma, Daiichi Sankyo, Genomedia, Guardant Health, Guardant Health AMEA, Roche Diagnostics KK, Seagen and Tempus. D.N. reports a consulting or advisory role with Janssen Oncology; stock and other ownership interests in Intuitive Surgical and Teladoc; and financial interests with Novartis and Takeda. D.M.O. reports consulting or advisory roles with Adaptimmune, Agenus, AstraZeneca, Clovis Oncology, Corcept Therapeutics, DualityBio, Eisai, Elevar Therapeutics, GlaxoSmithKline, GOG Foundation, Immunogen, Imvax, Laekna Therapeutics, Merck, Mersana, Novartis, Novocure, OncoC4, Onconova Therapeutics, Regeneron, Roche, Seagen, Sutro Biopharma, Umoja Biopharma and Verastem and institutional research funding from AbbVie, AbbVie/Stemcentrx, Acerta Pharma, Advaxis, Ajinomoto, Amgen, AstraZeneca, Arcus Biosciences, Array BioPharma, BBI Healthcare, BeiGene, Bristol Myers Squibb, Cerulean Pharma, Clovis Oncology, Deciphera, Eisai, EMD Serono, Ergomed, Exelixis, Genentech/Roche, Genmab, GlaxoSmithKline, OncoQuest, Pfizer, Precision Therapeutics, Immunogen, Incyte, Iovance Biotherapeutics, Janssen Research & Development, Karyopharm Therapeutics, Leap Therapeutics, Ludwig Institute for Cancer Research, Merck, Mersana, Novartis, NovoCure, PharmaMar, Regeneron, Roche, Sanofi, Seagen, Sumitomo Dainippon Pharma Oncology, Sutro Biopharma, Tesaro, TRACON Pharma and Verastem. A.B.O. reports consulting or advisory roles with AstraZeneca, Clovis Oncology, Genentech, GlaxoSmithKline, Novocure and Tesaro. M.R. reports honoraria for lectures and consultancy from Amgen, AstraZeneca, BMS, BeiGene, Boehringer Ingelheim, Daiichi Sankyo, GSK, Lilly, Merck, Mirati, MSD, Novartis, Pfizer, Regeneron and Sanofi; and compensation for membership in DMSB by Daiichi Sankyo and Sanofi. K.S. reports research funds from AstraZeneca, Amgen, Daiichi Sankyo, Gilead, Merck, NanoCarrier, PRA Health Sciences and Takeda; and personal fees from AstraZeneca, Bayer Yakuhin, Eisai, MSD, Nihon Medi-Physics and Pfizer. Y.S. reports institutional grants or contracts from Chugai and Taiho; payment or honoraria for lectures, presentations, speaker’s bureau, paper writing or educational events from Astellas, Bayer Yakuhin, Bristol Myers Squibb KK, Chugai, Daiichi Sankyo, Eli Lilly Japan KK, Guardant Health, Merck, MSD KK, Novartis, Ono, Sysmex, Taiho and Takeda; and roles on data safety monitoring or advisory boards with Guardant Health, Merck and Ono. C.V.M. reports institutional research funding from DigiCore and Gilead; consulting or advisory roles (to the institution) from AstraZeneca, Daiichi Sankyo, Lilly, MSD and Novartis; travel, accommodations and expenses from Amgen, Astellas, Bristol Myers Squibb, DigiCore, Gilead, Merck, MSD Oncology, Pfizer and Roche; and institutional research funding from all companies above. E.Y.Y. reports consulting or advisory roles with AADi, Advanced Accelerator Applications, Bayer, Janssen, Merck and Oncternal Therapeutics; institutional research funding from Agensys, Bayer, Blue Earth Diagnostics, Daiichi Sankyo, Dendreon, Lantheus Medical Imaging, Merck, Seagen, Surface Oncology, Taiho Pharmaceutical and Tyra Biosciences; personal fees from Aadi Bioscience, Advanced Accelerator Applications, Janssen and Oncterna; and grants and personal fees from Bayer and Merck. J.R. reports employment at Pfizer; stock and other ownership interests in Pfizer; and travel and accommodation expenses from Pfizer. S. Tan reports employment at Pfizer; stock and other ownership interests in Pfizer; and travel and accommodation expenses from Pfizer. M.B. reports employment at Pfizer, and stock and other ownership interests in Pfizer. T.E.S. reports consulting or advisory roles with Abbvie, AstraZeneca, Blueprint Medicines, Boehringer Ingelheim, Coherus Biosciences, G1 Therapeutics, Gilead Sciences, Pfizer, Spectrum Pharmaceuticals and Takeda and institutional research funding from AstraZeneca, Genentech/Roche, Mirati Therapeutics, Nuvalent, and Seagen. P.R.P. reports honoraria from Dava Oncology, OncLive and Frontiers; consulting or Advisory Role with Personalized Cancer Therapy, OncoPlex Diagnostics, Immunonet BioSciences, Pfizer, Heron, Puma Biotechnology, Sirtex Medical, Caris Life Sciences, Juniper Pharmaceuticals, Bolt Biotherapeutics and AbbVie; speakers’ bureau with Genentech/Roche; institutional research funding from Genentech/Roche, Fabre-Kramer, Advanced Cancer Therapeutics, Caris Centers of Excellence, Pfizer, Pieris Pharmaceuticals, Cascadian Therapeutics, Bolt Biotherapeutics, Byondis, Seagen, Orum and Carisma; and intellectual property US patent numbers 8,486,413, 8,501,417, 9,023,362 and 9,745,377. Non-financial interests: G.C. reports uncompensated relationships with Consiglio Superiore di Sanità (officer), Italian National Health Council (advisor for the Ministry of Health), ESMO (officer), ESMO Open (editor in chief), Europa Donna (advisory role), member of the Scientific Council Patient Advocacy Association, EUSOMA (leadership role) and Fondazione Beretta (advisory role) Cancer Research Foundation. A.R. reports research funding from Bristol Myers Squibb; payment or honoraria from Bristol Myers Squibb and Novartis; and support for attending meetings and/or travel from Bristol Myers Squibb, Merck Sharp & Dohme and Novartis. D.N. reports uncompensated relationships with Takeda and Novartis. C.V.M. reports a non-financial competing interest as a board member of BSMO. P.R.P. reports uncompensated relationships with Pfizer, Seagen and Jazz.
Footnotes
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
is available for this paper at 10.1038/s41591-024-03462-0.
Supplementary information
The online version contains supplementary material available at 10.1038/s41591-024-03462-0.
References
- 1.Riese, D. J. 2nd & Stern, D. F. Specificity within the EGF family/ErbB receptor family signaling network. Bioessays20, 41–48 (1998). [DOI] [PubMed] [Google Scholar]
- 2.Olayioye, M. A., Neve, R. M., Lane, H. A. & Hynes, N. E. The ErbB signaling network: receptor heterodimerization in development and cancer. EMBO J.19, 3159–3167 (2000). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Yarden, Y. & Sliwkowski, M. X. Untangling the ErbB signalling network. Nat. Rev. Mol. Cell Biol.2, 127–137 (2001). [DOI] [PubMed] [Google Scholar]
- 4.Schlessinger, J. Ligand-induced, receptor-mediated dimerization and activation of EGF receptor. Cell110, 669–672 (2002). [DOI] [PubMed] [Google Scholar]
- 5.Holbro, T. & Hynes, N. E. ErbB receptors: directing key signaling networks throughout life. Annu. Rev. Pharmacol. Toxicol.44, 195–217 (2004). [DOI] [PubMed] [Google Scholar]
- 6.Hynes, N. E. & Lane, H. A. ERBB receptors and cancer: the complexity of targeted inhibitors. Nat. Rev. Cancer5, 341–354 (2005). [DOI] [PubMed] [Google Scholar]
- 7.Slamon, D. J. et al. Use of chemotherapy plus a monoclonal antibody against HER2 for metastatic breast cancer that overexpresses HER2. N. Engl. J. Med.344, 783–792 (2001). [DOI] [PubMed] [Google Scholar]
- 8.Baselga, J. et al. Pertuzumab plus trastuzumab plus docetaxel for metastatic breast cancer. N. Engl. J. Med.366, 109–119 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Bang, Y. J. et al. Trastuzumab in combination with chemotherapy versus chemotherapy alone for treatment of HER2-positive advanced gastric or gastro-oesophageal junction cancer (ToGA): a phase 3, open-label, randomised controlled trial. Lancet376, 687–697 (2010). [DOI] [PubMed] [Google Scholar]
- 10.Nakamura, Y. et al. Circulating tumor DNA-guided treatment with pertuzumab plus trastuzumab for HER2-amplified metastatic colorectal cancer: a phase 2 trial. Nat. Med.27, 1899–1903 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Murthy, R. K. et al. Tucatinib, trastuzumab, and capecitabine for HER2-positive metastatic breast cancer. N. Engl. J. Med.382, 597–609 (2020). [DOI] [PubMed] [Google Scholar]
- 12.Strickler, J. H. et al. Tucatinib plus trastuzumab for chemotherapy-refractory, HER2-positive, RAS wild-type unresectable or metastatic colorectal cancer (MOUNTAINEER): a multicentre, open-label, phase 2 study. Lancet Oncol.24, 496–508 (2023). [DOI] [PubMed] [Google Scholar]
- 13.Cortés, J. et al. Trastuzumab deruxtecan versus trastuzumab emtansine for breast cancer. N. Engl. J. Med.386, 1143–1154 (2022). [DOI] [PubMed] [Google Scholar]
- 14.Shitara, K. et al. Trastuzumab deruxtecan in previously treated HER2-positive gastric cancer. N. Engl. J. Med.382, 2419–2430 (2020). [DOI] [PubMed] [Google Scholar]
- 15.Krop, I. E. et al. Trastuzumab emtansine versus treatment of physician’s choice in patients with previously treated HER2-positive metastatic breast cancer (TH3RESA): final overall survival results from a randomised open-label phase 3 trial. Lancet Oncol.18, 743–754 (2017). [DOI] [PubMed] [Google Scholar]
- 16.Endo, Y. et al. HER2 mutation status in Japanese HER2-positive breast cancer patients. Breast Cancer23, 902–907 (2016). [DOI] [PubMed] [Google Scholar]
- 17.Hyman, D. M. et al. HER kinase inhibition in patients with HER2- and HER3-mutant cancers. Nature554, 189–194 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Khoury, T. et al. ERBB2 juxtamembrane domain (trastuzumab binding site) gene mutation is a rare event in invasive breast cancers overexpressing the ERBB2 gene. Mod. Pathol.24, 1055–1059 (2011). [DOI] [PubMed] [Google Scholar]
- 19.Boulbes, D. R. et al. HER family kinase domain mutations promote tumor progression and can predict response to treatment in human breast cancer. Mol. Oncol.9, 586–600 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Zuo, W. J. et al. Dual characteristics of novel HER2 kinase domain mutations in response to HER2-targeted therapies in human breast cancer. Clin. Cancer Res.22, 4859–4869 (2016). [DOI] [PubMed] [Google Scholar]
- 21.Zhang, X. et al. Parallel analyses of somatic mutations in plasma circulating tumor DNA (ctDNA) and matched tumor tissues in early-stage breast cancer. Clin. Cancer Res.25, 6546–6553 (2019). [DOI] [PubMed] [Google Scholar]
- 22.Connell, C. M. & Doherty, G. J. Activating HER2 mutations as emerging targets in multiple solid cancers. ESMO Open2, e000279 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Deniziaut, G. et al. ERBB2 mutations associated with solid variant of high-grade invasive lobular breast carcinomas. Oncotarget7, 73337–73346 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Wynn, C. S. & Tang, S. C. Anti-HER2 therapy in metastatic breast cancer: many choices and future directions. Cancer Metastasis Rev.41, 193–209 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Li, B. T. et al. 654O efficacy and safety of trastuzumab deruxtecan (T-DXd) in patients (pts) with solid tumors harboring specific HER2-activating mutations (HER2m): primary results from the international phase II DESTINY-PanTumor01 (DPT-01) study. Ann. Oncol.34, S459–S460 (2023). [Google Scholar]
- 26.Tian, H. et al. Dramatic response to pyrotinib and T-DM1 in HER2-negative metastatic breast cancer with 2 activating HER2 mutations. Oncologist28, e534–e541 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Jhaveri, K. et al. Neratinib + fulvestrant + trastuzumab for HR-positive, HER2-negative, HER2-mutant metastatic breast cancer: outcomes and biomarker analysis from the SUMMIT trial. Ann. Oncol.34, 885–898 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Kulukian, A. et al. Preclinical activity of HER2-selective tyrosine kinase inhibitor tucatinib as a single agent or in combination with trastuzumab or docetaxel in solid tumor models. Mol. Cancer Ther.19, 976–987 (2020). [DOI] [PubMed] [Google Scholar]
- 29.Curigliano, G. et al. Tucatinib versus placebo added to trastuzumab and capecitabine for patients with pretreated HER2+ metastatic breast cancer with and without brain metastases (HER2CLIMB): final overall survival analysis. Ann. Oncol.33, 321–329 (2022). [DOI] [PubMed] [Google Scholar]
- 30.Lin, N. U. et al. Intracranial efficacy and survival with tucatinib plus trastuzumab and capecitabine for previously treated HER2-positive breast cancer with brain metastases in the HER2CLIMB trial. J. Clin. Oncol.38, 2610–2619 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Lin, N. U. et al. Tucatinib vs placebo, both in combination with trastuzumab and capecitabine, for previously treated ERBB2 (HER2)-positive metastatic breast cancer in patients with brain metastases: updated exploratory analysis of the HER2CLIMB randomized clinical trial. JAMA Oncol.9, 197–205 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Peterson, S., Rosler, R. & Klucher, K. Abstract 4222: Tucatinib, a selective small molecule HER2 inibitor, is active in HER2 mutant driven tumors. Cancer Res.80, 4222 (2020). [Google Scholar]
- 33.Verma, S. et al. Trastuzumab emtansine for HER2-positive advanced breast cancer. N. Engl. J. Med.367, 1783–1791 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Geyer, C. E. et al. Lapatinib plus capecitabine for HER2-positive advanced breast cancer. N. Engl. J. Med.355, 2733–2743 (2006). [DOI] [PubMed] [Google Scholar]
- 35.Sturgill, E. G. et al. Next-generation sequencing of patients with breast cancer in community oncology clinics. JCO Precis. Oncol.5, 1297–1311 (2021). [DOI] [PubMed] [Google Scholar]
- 36.Strickler, J. H. et al. HER2 testing in the MOUNTAINEER trial: analysis of treatment response based on central HER2 assessment using IHC/ISH and NGS. J. Clin. Oncol.41, abstr. 3528 (2023). [Google Scholar]
- 37.Nakamura, Y. et al. Tucatinib and trastuzumab for previously treated human epidermal growth factor receptor 2-positive metastatic biliary tract cancer (SGNTUC-019): a phase II basket study. J. Clin. Oncol.41, 5569–5578 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Pahuja, K. B. et al. Actionable activating oncogenic ERBB2/HER2 transmembrane and juxtamembrane domain mutations. Cancer Cell34, 792–806.e5 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Robichaux, J. P. et al. Pan-cancer landscape and analysis of ERBB2 mutations identifies poziotinib as a clinically active inhibitor and enhancer of T-DM1 activity. Cancer Cell36, 444–457.e447 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Clopper, C. J. & Pearson, E. S. The use of confidence or fiducial limits illustrated in the case of the binomial. Biometrika26, 404–413 (1934). [Google Scholar]
- 41.Collett, D. in Modelling Survival Data in Medical Research (ed. Collett, D.) 2nd edn, 237–251 (Chapman & Hall, 1994).
- 42.Wolff, A. C. et al. Human epidermal growth factor receptor 2 testing in breast cancer: American Society of Clinical Oncology/College of American Pathologists clinical practice guideline focused update. J. Clin. Oncol.36, 2105–2122 (2018). [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
List of institutional review boards and independent ethics committees.
Data Availability Statement
Following requests and subject to review, Pfizer will provide the data that support the findings of this study. Subject to certain criteria, conditions and exceptions, Pfizer may also provide access to the related individual de-identified participant data. See https://www.pfizer.com/science/clinical-trials/trial-data-and-results for more information.