Abstract
Objectives
Periodontitis is a serious periodontal infection that damages the soft tissues and bone around teeth and is linked to systemic conditions. Accurate diagnosis and staging, complemented by radiographic evaluation, are vital. This systematic review (PROSPERO ID: CRD42023480552) explores artificial intelligence (AI) applications in assessing alveolar bone loss and periodontitis on dental panoramic and periapical radiographs
Methods
Five databases (Medline, Embase, Scopus, Web of Science, and Cochrane’s Library) were searched from January 1990 to January 2024. Keywords related to “artificial intelligence”, “Periodontal bone loss/Periodontitis”, and “Dental radiographs” were used. Risk of bias and quality assessment of included papers were performed according to the APPRAISE-AI Tool for Quantitative Evaluation of AI Studies for Clinical Decision Support. Meta analysis was carried out via the “metaprop” command in R V3.6.1.
Results
Thirty articles were included in the review, where 10 papers were eligible for meta-analysis. Based on quality scores from the APPRAISE-AI critical appraisal tool of the 30 papers, 1 (3.3%) were of very low quality (score < 40), 3 (10.0%) were of low quality (40 ≤ score < 50), 19 (63.3%) were of intermediate quality (50 ≤ score < 60), and 7 (23.3%) were of high quality (60 ≤ score < 80). No papers were of very high quality (score ≥ 80). Meta-analysis indicated that model performance was generally good, eg, sensitivity 87% (95% CI, 80%-93%), specificity 76% (95% CI, 69%-81%), and accuracy 84% (95% CI, 75%-91%).
Conclusion
Deep learning shows much promise in evaluating periodontal bone levels, although there was some variation in performance. AI studies can lack transparency and reporting standards could be improved. Our systematic review critically assesses the application of deep learning models in detecting alveolar bone loss on dental radiographs using the APPRAISE-AI tool, highlighting their efficacy and identifying areas for improvement, thus advancing the practice of clinical radiology.
Keywords: artificial intelligence, deep learning, panoramic radiographs, periapical radiographs, periodontitis
Introduction
Rationale
Periodontitis is a serious multifactorial periodontal infection that damages the soft tissues and bone around teeth and is linked to systemic conditions.1 The prevalence of periodontitis is estimated to be about 50% in the United States based on the American Academy of Periodontology,2 and around 10%-15% globally suffer from severe cases that cause loss of teeth.2 Periodontitis is a complex multifactorial process that is initiated with bacterial plaque accumulation, and biofilms, followed by a host immune reaction or inflammatory response.1,3 If not treated, periodontitis progression will eventually lead to teeth loss and impaired oral function.4 Although no conclusive cause-and-effect has been established, studies have correlated periodontitis as a possible predisposition for systematic conditions, such as cardiovascular diseases, respiratory tract infections, adverse pregnancy outcomes, Alzheimer’s disease, and oral and colorectal cancer.4,5
In addition to clinical findings of periodontal disease, dental radiography is an integral part of diagnosis and treatment planning as it provides a comprehensive evaluation of hard dento-alveolar structures, as well as calculus depositions, shape of roots, and alveolar bone level.6–8 Several radiographic techniques are used for periodontal examination. Bitewing provides limited details on the maxillary and mandibular teeth crowns and the alveolar crest level. Full mouth series of parallel periapical radiographs have been considered “the gold standard” for periodontal evaluation. This is because periapical radiographs provide information on teeth and supporting structures with relatively low-dose radiation, while still providing images of high resolution and that are of good quality.
Panoramic radiographs have become the most commonly used modality in dental examination and periodontal evaluation. Such radiographs provide a comprehensive view of the maxillofacial structures. They also capture maxillary and mandibular teeth and the alveolar bone in one image and in a few seconds. Furthermore, they involve a relatively low radiation dose and yet still give acceptable image quality.6,9 The introduction of cone beam computed tomography (CBCT) allowed the 3D evaluation of periodontal structures and a comprehensive evaluation of periodontal defects such as furcation defects, fenestration, and dehiscence and postsurgical evaluation of regenerative periodontal procedures.10,11 However, CBCT can lead to a high dose of radiation, as well as inherent artefacts, and so should not be used in routine examination procedures.12,13
Currently there is a surge in artificial intelligence (AI) applied to all aspects of dentistry, with a wide range of applications ranging from simple task management to complicated diagnostic evaluation and tools in decision making.14 A recent innovation has been the introduction of deep learning (DL), which is a form of AI that often involves the use of neural networks. Some dental/oral health examples in the literature are cancer cell detection and healing evaluation, enhanced restoration margins adaptation, caries detection and shade selection, pre-and post-orthognathic surgical evaluation, cephalometric analysis, periodontal evaluation, and bone loss detection, CAD-CAM and 3D printing for implant treatment, root canal morphology, canal length, and vertical root fracture.14,15 Additionally, Hunge et al16 highlighted AI applications in 3D diagnostic imaging in their narrative review, especially relating to multidetector CT and CBCT to discover and delineate jaw cysts and tumors, lymph node metastases, salivary glands diseases, temporo-mandibular joints (TMJs), maxillary sinuses, mandibular fracture, and dento-maxillofacial deformities. DL is extensively utilized for segmentation tasks,17,18 accurately delineating structures in dental radiographs.19 Additionally, DL models are applied for both segmentation and classification, enabling the precise identification and categorization of dental conditions.20–22
Objectives
Two systematic reviews23,24 have explored the application of DL in evaluating periodontal bone loss assessed via dental radiographs, excluding CBCT. There has been a noticeable increase in the number of publications since these systematic reviews were published. These papers addressed the different DL models and may provide additional information important in improving the diagnostic accuracy of these models and help in clinical implementation and support in the decision-making process. Therefore, this systematic review aims to explore, analyze, and summarize the application of DL models in evaluating periodontal bone loss using the newly developed APPRAISE-AI Tool for Quantitative Evaluation of AI Studies for Clinical Decision Support.25
Methods
Eligibility criteria
The articles were collected in January 2024 following the PICO/PIRO (Population, Intervention/Index Test [AI-Model], Comparison/Reference Standard, and Outcome) question format was followed during the search, P: Patient with periodontal bone loss/periodontitis; I: Radiographic image evaluation with AI; C: Radiographic evaluation of clinician or relative to an established ground truth and/or gold standard, which according to authors of the papers was established either from patients records or experts who pre-labelled the images; O: periodontal bone loss detection and classification accuracy. We used PICO/PIRO instead of PICOS as almost all studies included did not discuss the study design and all seemed cross-sectional. This highlights common problems related to transparency and reporting that are later discussed in the results and discussion section. This study included all articles published from January 1990 to January 2024; papers that were written in English; articles that applied any type of AI models, such as RCNN or SVM, to evaluate the periodontal bone level, periodontitis, and/or periodontal diseases on intraoral or extraoral radiographs, such as Periapical, Bitewings, and Panoramic radiographs. Although SVM is a machine learning (ML) algorithm, it was retained due to relatively little evidence found in the literature. It fits both inclusion criteria and keywords used in the search. We also noted that it has been used as a comparison model in some of the DL studies we included.26,27 We excluded studies published before 1990, non-English articles, and conference abstracts without fully published documents.
Information sources
The systematic review has been registered with PROSPERO (ID: CRD42023480552). Search parameters and keywords were developed by the authors. Keywords were set initially by using the Population, Intervention/Index Test (AI-Model), Comparison/Reference Standard, and Outcome (PICO/PIRO) approach. These keywords were then iterated between all authors until a mutually agreed set of terms was found.
Search strategy
The databases searched in January 2024 were Medline via Ovid (13 papers), Embase via Ovid (41 papers), Scopus through Elsevier (83 papers), Web of Science (62 papers), and Cochrane Library (5 papers). The total initial search yielded 204 that were exported to the EndNote Library. Terms used: (“Artificial intelligence” OR “AI” OR “Machine learning” OR “Neural network” OR “Deep Learning” OR “Convolutional neural networks”) AND (“Alveolar bone loss” OR “Periodontal bone loss” OR “Periodontal disease” OR “Periodontitis” OR “Diagnosis of periodontal bone loss” OR “Detect alveolar bone loss”) AND (Radiograph OR “Dental radiograph” OR “Periapical radiograph” OR “Panoramic radiograph” OR “Radiographic imaging” OR “Cone beam computed tomography” OR “CBCT”).
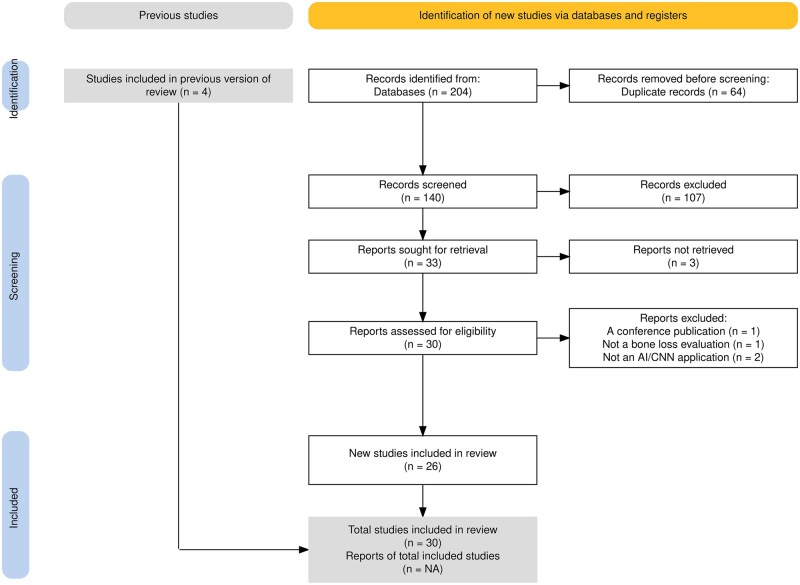
The selection process, data extraction, analysis, and reporting procedures in this review follow the PRISMA (Preferred Reporting Items for Systematic Reviews and Meta-Analysis) guidelines.28 A PRISMA chart29 shown in Figure 1 illustrates the stages of data extraction. The database search yielded 204 papers, 64 of which were removed during the removal of duplicate sources. An additional 107 articles were found to be irrelevant according to our exclusion criteria and were removed during initial screening. Thirty-three articles were included in the database search. Additionally, 4 papers not found in the database search were retrieved from the previous systematic reviews23,24 and included in the study.
Figure 1.
PRISMA flowchart for the study.
Data collection
Those 37 articles were uploaded to the Rayyan systematic review collaboration website to be evaluated by all other researchers. Two researchers critically appraised these articles. Three full article texts could not be retrieved and were removed from the study. Four papers were also excluded during the final evaluation as they did not meet the selection criteria as shown in the PRISMA flowchart. Finally, 30 eligible articles, on which all reviewers agreed, were included in this systematic review. Eleven of these 30 articles provided common outcomes that could be used in meta-analyses.
Risk of bias assessment
Critical appraisal of the final 30 papers was performed by 2 independent reviewers to reduce the risk of bias. The appraisal tool used in this study was published by Kwong et al25 (APPRAISE-AI Tool for Quantitative Evaluation of AI Studies for Clinical Decision Support). Each paper was scored independently using the APPRAISE-AI tool by 2 independent reviewers (2 of the authors) on the scale going from a minimum of 0 (extremely poor quality) to 100 (extremely good quality), according to APPRAISE-AI.25 The 2 raters agreed within 6 points for 25 out of 30 papers, which indicates good agreement between the 2 raters in absolute terms (with respect to overall scores in the range [0,100]), and a composite mark (ie, the mean of the 2 scores) was used as the final score for each paper in this case. For those cases where initial disagreement was greater than 6 marks, the 2 reviewers re-evaluated each paper and agreed on a final score mutually. Thus, a robust estimate of quality was determined via the APPRAISE-AI score.
Six domains were identified for the APPRAISE-AI critical appraisal tool [25]: clinical relevance (maximum domain score = 4), data quality (maximum domain score = 24), methodological conduct (maximum domain score = 20), robustness of results (maximum domain score = 20), reporting quality (maximum domain score = 12), and reproducibility (maximum domain score = 20). Scores for each of these domains could be determined readily also and they were found to be extremely useful in analyzing the strengths and weaknesses of the article used here. In order to facilitate comparison between these domain scores, they were also scaled linearly in the range minimum of 0 (extremely poor quality) to 100 (extremely good quality). The specific questions used for critical appraisal via the APPRAISE-AI tool are presented in the Supplementary Material to this article.
Effect measures
The performance of DL models in detecting periodontal bone loss/periodontitis was measured with different parameters across studies. The following performance measures were used: accuracy, sensitivity (recall), specificity, precision (positive predictive value, PPV), F1-scores, negative predictive value (NPV), and the area under the receiving characteristic curve (AUC/ROC). Segmentation of features and localization accuracy were evaluated with intercession over union (IoU), dice similarity coefficient (DSC), Jaccard Index (JI), pixel accuracy (PA), and mean average precision (mAP). Note however that there was not enough data for meta analysis to be carried out for segmentation tasks. Bone loss or periodontitis was generally measured on a binary scale (no bone loss or bone loss). However, some studies22,30 had an ordinal scale and this is dichotomized by us explicitly here to form a binary scale (eg, no bone loss or any bone loss).
Syntheses methods
Meta-analysis
Measures such as sensitivity, specificity, accuracy, PPV, NPV, and even F1-scores (etc.) are all ratios of 2 integers, where the numerator counts the number of “events” with respect to some (perhaps effective for F1-scores) sample size (ie, the denominator). (Note also that the units of sampling were images rather than subjects in all papers.) Each of these measures lies in the range [0,1] and (crucially) values for these measures have a common meaning across all articles, ie, values near to 0 indicate extremely poor performance and values near to 1 indicate extremely good performance. Thus, we can treat each measure as a simple proportion, and standard methods of meta-analysis for a proportion can be employed. (Note that there were no consistent control groups (or methods) and so meta-analyses via odds ratios or relative risks could not be carried out.) Here, pooled point estimates and 95% confidence intervals of a single proportion via meta-analysis were found using the “metaprop” command for the statistical software package “meta” in the statistical software environment R V3.6.1. The default “arcsine” transformation was used here to calculate an overall proportion, although other transformations (eg, logit, double arcsine, logarithm, etc) all gave similar results (this was tested explicitly in all cases), which is an excellent test of the method.
Note that some papers used multiple types of neural networks and results are quoted here for each type of network. Augmented data were used in some papers, which inflated the effective sample size by multiple orders of magnitude compared to the other studies and so strongly affected confidence intervals. Subgroup meta-analyses of augmented versus no-augmentation is carried out here in addition to overall meta-analyses including results from all papers. Funnel plots and statistical tests of bias did not indicate that bias was strong, where there were enough papers to allow this analysis to be carried out for all performance measures. Note finally that random-effects meta-analysis was carried out for those cases with larger amounts of heterogeneity (ie, I2 values greater than approximately 50% and P < .05 for tests of heterogeneity). Sensitivity analyses were carried out where obvious outliers were detected, ie, meta-analysis was repeated with any outliers removed and results were compared to the original analysis containing all studies.
Results
Study selection and study characteristics
Table 1 presents the evidence table for all 30 articles based on authors, year of publication, country, sample size and type of images, AI software, main findings, statistical analysis, and appraisal score. The results show an increase in the number of publications in the year 2023 compared to previous years, with 11 out of 30 papers published in 2023,3,4,9,26,31–37 8 papers in 2022,22,30,38–43 4 papers in 2021,20,44,49,50 4 papers in 2020,27,45,51,52 2 papers in 2019,46,47 and only one paper in 2018.48
Table 1.
Summary of all articles included in the systematic review.
| Authors year, country | Sample size and type | AI-software | Main study findings | Statistical analysis | Appraisal score of 100 |
|---|---|---|---|---|---|
| Vollmer et al 2023,3 Germany | 1755 panoramic radiographs | Key points RCNN (ResNet-50-FPN) |
|
|
53.5 |
| Saylan et al 2023,4 Turkey | 1543 panoramic radiographs | YOLO-v5x |
|
|
57 |
| Sameer et al 2023,31 India | 116 panoramic radiographs | Support vector machine (SVM) and random forest (RF) |
|
|
32.5 |
| Ryu et al 2023,32 Korea | 4083 panoramic radiographs | Faster R-CNN |
|
|
52.5 |
| Mao et al 2023,33 Taiwan | 300 periapical radiographs |
|
|
|
46.5 |
| Liu et al 2023,34 China | 2275 panoramic radiographs | Alexnet |
|
|
70.5 |
| Kong et al 2023,35 China | 1747 panoramic radiographs | 2-stage PDCNN |
|
|
59.5 |
| Karacaoglu et al 2023,26 Turkey | 87 periapical of dry mandible |
|
|
|
54 |
| Chen et al 2023,36 Taiwan | 336 periapical radiographs (PA) | U-Net and Mask-RCNN |
|
|
52 |
| Chen et al 2023,37 Taiwan | 8000 periapical radiographs | YOLOv5 |
|
|
51 |
| Amasya et al 2023,9 Turkey | 6000 panoramic and additional from archive 100 panoramic to compare 3 clinicians |
|
|
|
48.5 |
| Widyaningrum et al 2022,22 Indonesia | 100 panoramic radiographs | Multi-Label U-Net and Mask R-CNN |
|
|
54 |
| Tsoromokos et al 2022,38 The Netherlands | 446 periapical radiographs | CNN Model not specified. |
|
|
63 |
| Shon et al 2022,39 Korea |
|
U-Net and YOLOv5 |
|
|
52 |
| Lee et al 2022,40 USA | 1247 periapical radiographs | Variations of U-Net (U-Net with CNN, ResNet-34, and ResNet-50 encoder) |
|
|
62.5 |
| Kearney et al 2022,41 USA |
|
|
|
|
56.5 |
| Jiang et al 2022,42 China | 640 panoramic radiographs |
|
|
|
59 |
| Chang et al 2022,43 USA | 1836 periapical radiographs | Inception V3 |
|
|
63.5 |
| Alotaibi et al 2022,30 KSA | 1724 periapical radiographs, anterior maxillary and mandibular teeth. | VGG-16 |
|
|
55.5 |
| Li et al 2021,44 China | 506 panoramic radiographs; 2 hospitals, Suzhou and Zhongshan. | Deetal-Perio |
|
|
53.5 |
| Kabir et al 2021,20 USA |
|
|
|
|
54.5 |
| Danks et al 2021,49 UK | 340 periapical radiographs | Symmetric hourglass architecture with ISM model |
|
|
62 |
| Chen et al 2021,50 China | 2900 periapical radiographs | Faster R-CNN′ |
|
|
47.5 |
| Thanathornwong et al 2020,51 Thailand | 100 panoramic radiographs | Faster R-CNN (base CNN was ResNet-101). |
|
|
50 |
| Moran et al 2020,27 Brazil | 467 periapical radiographs |
|
|
|
54.5 |
| Kurt et al 2020,52 Turkey | 2276 panoramic radiographs. | Pretrained Google Net Inception v3 |
|
|
49.5 |
| Chang et al 2020,45 South Korea | 340 panoramic radiographs. |
|
|
|
54.5 |
| Krois et al 2019,47 Germany | 85 panoramic radiographs | Custom made CNN |
|
|
62 |
| Kim et al 2019,46 South Korea |
|
DCNN called DeNTNet |
|
|
66 |
| Lee et al 2018,48 Korea | 1740 periapical radiographs | Modified VGG-19 network architecture. |
|
|
61 |
Results of individual studies
The majority of the studies used panoramic radiographs to assess radiographic bone and periodontal disease through a DL approach.3,4,9,22,31,32,34,35,39,42,44–47,51,52 Periapical radiographs were used in 13 papers.20,26,27,30,33,36–38,40,41,43,48–50 Only one article used bitewing radiographs.41
Twenty-nine studies used AI models to assess radiographic bone loss on dental radiographs.3,4,9,20,22,26,27,30–40,42–52 Kearney et al41 used GANs to evaluate inpainted and non-inpainted methods, which were used to evaluate clinical attachment loss rather than radiographic bone loss. Although this study did not assess the bone level, it still aligns with the original inclusion criteria and keywords established at the start of the search, which were never modified after the search process. It evaluated periodontal disease/periodontitis on 2D dental radiographs using an AI model. Therefore, it was not excluded from the review.
As seen in Table 1, all studies included in this review have established ground truth through expert image annotations and labeling. However, only 14 studies compared model performance to clinical experts.9,20,26,32,34,36,38,40–43,46–48
Risk of bias assessment
Based on the quantitative analysis of the APPRAISE-AI tools,25 7 papers were considered high quality (scored 60-79): Liu et al 2023; Tsoromokos et al 2022; Chang et al 2022; Lee et al 2022; Danks et al 2021; Kim et al 2019; and Krois et al 2019.34,38,40,43,46,47,49 Nineteen papers were considered intermediate quality (score 50-59), and 4 papers scored below 50 and were considered low-quality papers. The lowest scoring paper (Sameer et al 202331) scored 35, which is considered very low quality. Table 2 summarizes the papers based on the AI-score rating. The mean score of all items was 55.3 (median = 54.5; SD = 7.2).
Table 2.
Number and percentage of articles in each quality category via overall scores from the APPRAISE-AI critical appraisal tool.25
| Quality category | Number (%) | Papers |
|---|---|---|
|
7 (23.3%) | Liu (2023), Tsoromokos (2022), Chang (2022), Lee (2022), Danks (2021), Kim (2019), Krois (2019). |
|
19 (63.3%) | Kong (2023), Karacaoglu (2023), Saylan (2023), Vollmer (2023), Ryu (2023), Chen (2023)—first, Chen (2023)—second, Alotaibi (2022), Li (2021), Kabir- (2021), Moran (2020), Chang (2020), Widyaningrum (2022), Shon (2022), Lee (2018), Kearney (2022), Jiang (2022), Kurt (2020), Thanathornwong (2020) |
|
3 (10.0%) | Mao (2023), Amaysa (2023), Chen (2021) |
|
1 (3.3%) | Sameer et al 2023 |
As noted above, the APPRAISE-AI tool splits the appraisal of each paper into distinct sections, namely, title/introduction, methods, results, conclusions, and others. Each APPRAISE-AI item was mapped to one of the following domains: clinical relevance, data quality, methodological conduct, robustness of results, reporting quality, and reproducibility. In order to compare results for domains against each other using the same scale, note again that we scale domain scores linearly to lie in the range [0,100], where: clinical relevance, mean = 97.1 and SD = 6.3; data quality, mean = 58.9 and SD = 9.4; methodological conduct, mean = 54.4 and SD = 11.6: robustness of results, mean = 42.7 and SD = 10.1; reporting quality, mean = 72.9 and SD = 16.1: and reproducibility, mean = 45.5 and SD = 11.4. Table 3 also summarizes the results of each domain. As seen, most papers are either intermediate or low-quality (ie, domain score < 60), and only 7 papers produced high-quality results (ie, 60 ≤ domain score < 80).
Table 3.
Domain scores scaled in the range 0 (extremely poor) to 100 (extremely good) using the APPRAISE-AI critical appraisal tool.25
| Scaled | All items | Clinical relevance | Data quality | Methodological conduct | Robustness of results | Reporting quality | Reproducibility |
|---|---|---|---|---|---|---|---|
| Mean | 55.3 | 97.1 | 58.9 | 54.4 | 42.7 | 72.9 | 45.5 |
| Median | 54.5 | 100.0 | 58.3 | 56.3 | 45.0 | 75.0 | 47.5 |
| SD | 7.2 | 6.3 | 9.4 | 11.6 | 10.1 | 16.1 | 11.4 |
Results of syntheses
Eleven papers4,9,22,27,30–34,40,52 were found to be eligible for the meta-analysis. However, Mao’s et al 202333 study was removed from the meta-analysis, because it used a training sample for testing model performance instead of a validation sample, indicating a high risk of bias. Therefore, 10 papers4,9,22,27,30–32,34,40,52 have been used in the analysis. There was enough data for 6 measures to be assessed through meta-analysis. Table 3 shows results for the (point) estimates (and 95% CI) for sensitivity (recall), specificity, accuracy, precision (PPV), NPV, and F1-score. Figures 2 and 3 show forest plots for sensitivity and specificity, respectively. Additional figures of forest plots are attached as Supplementary Material. All results show overall high values for all parameters (ie, sensitivity (recall), specificity, accuracy, PPV, NPV) and F1-score.
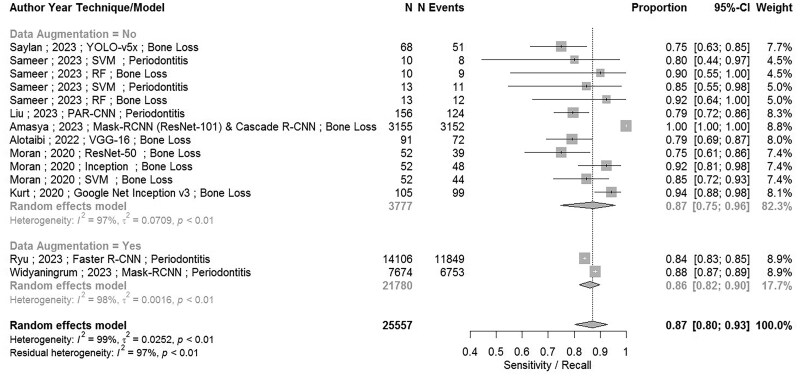
Figure 2.
Forest plot including meta-analysis for the model performance measure: sensitivity.
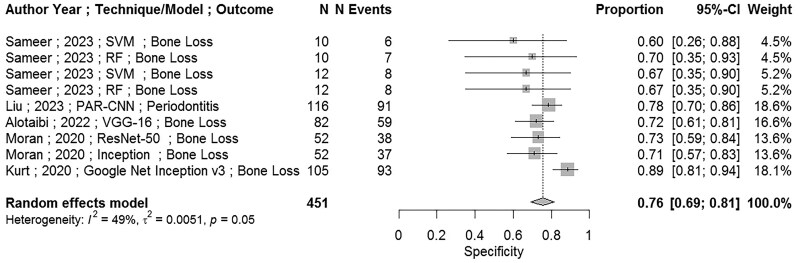
Figure 3.
Forest plot (excluding Moran [2020]) with meta-analysis for the model performance measure: specificity.
Results of meta-analysis for the sensitivity are shown in Figure 2 and Table 4, where sensitivity is given by 0.87 (95% CI, 0.75-0.96) for non-augmentation cases, 0.86 (95% CI, 0.82-0.90) for augmentation cases, and 0.87 (95% CI, 0.80-0.93) for both augmented and non-augmented cases combined. Note that we consider the data to be augmented when both training and testing data used in the model are augmented. Although confidence intervals are much narrower for the augmented data (as expected given that sample sizes are much larger), no compelling differences were identified in the point estimates between augmented and non-augmented data. There were inconclusive differences by outcome type and no difference by type of data measured for testing and training. As demonstrated, our approach acknowledges that finite test sample size itself impacts the confidence intervals in the meta-analysis. We demonstrated how augmentation reduced the confidence intervals by conducting a subgroup analysis on augmented versus non-augmented data.
Table 4.
Overview of the meta-analysis results.
| Measure | Description of measure | Point estimate and 95% CIs from meta-analysis (expressed as percentages here) |
|---|---|---|
| Sensitivity (also known as Recall) | Percentage of cases with periodontitis that were classified correctly as positive | 87% (95% CI, 80%-93%) |
| Specificity | Percentage of cases without periodontitis that were classified correctly as negative | 76% (95% CI, 69%-81%) |
| Accuracy | Percentage of cases both with and without periodontitis that were classified correctly | 84% (95% CI, 75%-91%) |
| PPV (also known as Precision) | Percentage of positive classifications (periodontitis etc.) that were correct | 81% (95% CI, 77%-84%) |
| NPV | Percentage of negative classifications (no periodontitis etc.) that were correct | 81% (95% CI, 73%-88%) |
| F1-score | Harmonic mean of the precision and sensitivity | 80% (95% CI, 74%-85%) |
Results of meta-analysis for the specificity are shown in Figure 3 and Table 4. None of the studies used in meta-analysis for the specificity employed data augmentation. Results of meta-analysis for the specificity were 0.69 (95% CI, 0.56-0.80). Moran et al’s (SVM)27 study was identified as an outlier and a sensitivity analysis was carried out. Removing this study reduced heterogeneity and changed the results of meta-analysis for the specificity slightly to 0.76 (95% CI, 0.69-0.81) and results are shown in Figure 3.
Meta-analysis was carried out for accuracy, PPV, NPV, and F1 score without data augmentation. Results for the accuracy were 0.82 (95% CI, 0.72-0.90) without augmentation across all studies. Removing the potential outlier of Moran et al’s (SVM; which did not use data augmentation),27 the accuracy changed slightly to 0.84 (95% CI, 0.75-0.91) across all studies, thus indicating the minimal impact of this potential outlier. Results for the precision (PPV) were 0.75 (95% CI, 0.67-0.83) without augmentation across all studies. Removing the potential outlier of Moran et al’s 2020 (SVM; which did not use data augmentation)27 reduced heterogeneity between studies and enabled the use of fixed-effects meta-analysis, PPV was adjusted to 0.81 (95% CI, 0.77-0.84) across all studies. Results for the NPV were 0.81 (95% CI, 0.73-0.88) across all studies. Results for the F1-score were 0.80 (95% CI, 0.74-0.85) across all studies. Again, note that results for all of these performance measures and across all studies are shown in Table 4 and Supplementary Materials.
Discussion
A systematic review was carried out here of the application of DL to detect periodontitis and periodontal bone loss from radiographic dental images. The review adhered to PRISMA standards, where 30 papers were used in this review. All articles were critically appraised using the APPRAISE-AI by 2 independent reviewers (ie, 2 authors of this article). Measures of model performance are often ratios of 2 integers, where this ratio lies in the range 0 to 1 and had a meaningful interpretation, namely: a value near to zero indicating extremely poor performance and near to 1 indicating extremely good performance. Standard methods of meta-analysis for a proportion using the “metaprop” command in R V3.6.1 could therefore be employed. Eleven papers provided quantitative evidence amenable to meta-analyses, and results were presented for the sensitivity (aka recall), specificity, accuracy, positive predictive value (aka precision), NPV, and F1 scores.
Using boundaries set by the APPRAISE-AI tool,25 critical appraisals indicated that 1 out of 30 papers (3.3%) were of very low quality (score < 40), 3 (10.0%) were of low quality (40 ≤ score < 50), 19 (63.3%) were of intermediate quality (50 ≤ score < 60), and 7 (23.3%) were of high quality (60 ≤ score < 80). No papers were of very high quality (score ≥ 80). This shows broadly that the quality of papers was adequate on the whole, although there was some variation in quality. The APPRAISE-AI tool subdivided the papers into 5 key areas/domains, namely, clinical relevance, data quality, methodological conduct, robustness of results, reporting, and reproducibility.
Not surprisingly, virtually all papers scored well on clinical relevance. This is probably because the maximum domain score was only 4 and so any attempt at the title, background, objective and problem, and clinical implementation was likely to receive a mark each. Previous systematic reviews show similar results as authors tend to have good reporting of clinical relevance and implementation and provide clear background and objectives.53,54 Similarly, reporting quality had a fairly high score compared to the other domains, and again its maximum score was only 12. Also, reporting cohort characteristics, limitations, and disclosures ought to be fairly straightforward tasks, and some form of “critical analysis” is a very common task when writing a paper, as noted in other reviews.24,54,55
Methodological conduct and data quality ought also to be straightforward tasks, but scoring for these domains was slightly lower. It is noticeable that many papers scored poorly on stating the sources of their data and also slightly less well on eligibility criteria for the data quality domain. Not surprisingly, authors tended to explain what the ground truths were and how data were abstracted and prepared, which are both “bread and butter” tasks in image analysis using DL. Similarly, data splitting and sample size calculations were explained adequately for the methodological conduct domain, although baseline models were explained less well. It seems that other reviews identified similar issues in explaining the methodological conduct with increased risk of bias despite using different critical appraisal tools.23,24,55
In addition, more than 50% of the included papers, in which experts annotated the images, did not include direct and blinded clinicians’ comparison.3,4,22,27,30,31,33,35,37,39,44,45,49–52 This limits the ability to validate the models’ performance in real-world experience. It might be argued that image annotation by experts can serve as a baseline to which an AI model is compared and validated. However, it is crucial to compare AI models’ performance directly with that of blinded experts, especially trained oral and maxillofacial radiologists, to ensure that they can complement and enhance clinicians’ diagnostic accuracy and improve trust and reliability.
Finally, robustness and reproducibility domains scored badly. For robustness, all items in this domain scored somewhat poorly, although error analysis was particularly poor where only a few papers even considered this. This reflects previous findings that highlight a lack of transparency and thoroughness in these areas.23,24,33,54,55 The poorer score for the reproducibility domain was driven by a lack of transparency (eg, authors not providing links to code or data) because model description and model specification tended to be reported extremely well. Overall, our analysis produced similar results to the APPRAISE-AI tool25 that showed the lowest domain scores were robustness of results, reproducibility, and methodological conduct.
Results of measures of model performance (sensitivity, specificity, F1 etc.) showed that overall performance was quite good, although there is quite a lot of variation between studies and so there is some “room for improvement,” which is consistent with previous studies.23,24,55 There was some evidence of a difference between sensitivity and specificity. Notably, results for the specificity of 76% (95% CI, 69%-81%) were somewhat lower than results for the sensitivity of 87% (95% CI, 80%-93%), indicating broadly that classifying negative cases correctly (ie, without disease) was a harder task than classifying positive cases correctly (ie, with disease). 95% confidence intervals were much smaller for augmented data compared to non-augmented data, which is exactly what one would expect as sample sizes have been increased synthetically compared to non-augmented data. Point estimates for these measures were broadly about the same (or perhaps slightly higher in some cases) for augmented versus non-augmented data, although this was inconclusive here. There was no evidence from this analysis that a particular type of neural network/DL model performs better than the others, although this might emerge in the future. Indeed, there appeared to be no other strong factor affecting results for measures of model performance, as far as we could tell.
One strength of the analyses carried out here is that we carry out meta-analysis for measures of model performance for AI applied to dental images. Furthermore, we have used an explicit critical appraisal tool to analyse our sources, which is another advantage. Weaknesses of our analyses are that there were relatively few studies for meta-analyses, although this is fast-moving field. Finally, we found high heterogeneity in our data, which makes the results of meta-analysis less reliable, even despite using random effects meta-analyses and sensitivity analyses. A common criticism of meta-analysis for experimental or lab-based studies is that the diverse setups (methods, populations, outcomes, etc.) render any average or composite value meaningless. However, our perspective is that meta-analysis remains valuable for gaining an overall understanding of results, as the data patterns for these measures generally show consistency across different studies.
In relation to clinical practice, rapid progress is clearly being made in this field. The results of meta-analyses for all of the measures of model performance indicate that (on average) models are not good enough as an automated screening tool as yet. Common acceptable performance cut-off values for screening tests are often cited as sensitivity and specificity roughly greater than or equal to 80%-90%,56 although one should note that the precise levels for these cut-offs are also strongly case-dependent and/or disease-dependent.56–58 However, we remark that some of the models in the papers used in this study might indeed perform well according to these criteria, but probably also require more (external) validation and testing. Critical appraisal carried out here indicates that a lack of transparency, absence of analysis of outliers and errors, and opacity regarding data sources in the articles considered here are potentially significant barriers to the subsequent adoption and translation by the dental community.
Future research should focus on transparency and rigorous explanation of study design and methods used in performing AI studies. We believe that the newly developed APPRSISE-AI tool by Kwong et al25 provides a useful tool to analyse the quality of future AI studies, including potential risks of bias. It can be used as a guideline in future research to create coherent, valid, and reproducible papers.
Conclusion
Studies showed various DL models can be developed and applied in dento-alveolar detection and segmentation and subsequent periodontal bone level evaluation with high accuracy. We applied the new APPRAISE-AI Tool in our study as it takes into consideration all necessary information that has to be reported in AI studies. Meta-analysis results indicate that model efficacy, averaged across included studies, is generally good to very good. Data augmentation appeared to enhance model performance, but this was not statistically significant. Despite literature heterogeneity and various performance parameters, AI models evaluated alveolar bone loss on 2D dental radiographs with high efficacy. However, it may not be good enough as an automated screening tool as yet; due to the lack of transparency, absence of analysis of outliers and errors, opacity, and discrepancy within studies. Finally, this systematic review highlights the need for more rigorous standards and clear guidelines in conducting, documenting, and reporting AI research in dentistry.
Supplementary Material
Contributor Information
Yahia H Khubrani, School of Dentistry, Cardiff University, The Annexe, University Dental Hospital, Heath Park, Cardiff CF14 4XY, United Kingdom; School of Dentistry, Jazan University, Jazan 82817, Saudi Arabia.
David Thomas, School of Dentistry, Cardiff University, The Annexe, University Dental Hospital, Heath Park, Cardiff CF14 4XY, United Kingdom.
Paddy J Slator, School of Computer Science and Informatics, Cardiff University, Cardiff CF24 4AG, United Kingdom; Cardiff University Brain Research Imaging Centre, School of Psychology, Cardiff University, Cardiff CF24 4HQ, United Kingdom.
Richard D White, Department of Clinical Radiology, University Hospital of Wales, Cardiff CF14 4XW, United Kingdom.
Damian J J Farnell, School of Dentistry, Cardiff University, The Annexe, University Dental Hospital, Heath Park, Cardiff CF14 4XY, United Kingdom.
Author contributions
Yahia H. Khubrani (Conceptualization, Methodology, Formal analysis, Data Curation, Writing original draft, Project administration), David Thomas (Conceptualization, Supervision, Writing—Review & Editing), Paddy Slator (Conceptualization, Supervision, Writing—Review & Editing), Richard D. White (Conceptualization, Writing—Review & Editing), and Damian J.J. Farnell (Conceptualization, Methodology, Supervision, Formal analysis, Data Curation)
Supplementary material
Supplementary material is available at Dentomaxillofacial Radiology online.
Funding
The first author was sponsored by Jazan University through the Saudi Arabia Cultural Bureau in the United Kingdom. However, “This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors”.
Conflicts of interest
The authors declare that they have no competing interests.
References
- 1. Nazir M, Al-Ansari A, Al-Khalifa K, Alhareky M, Gaffar B, Almas K. Global prevalence of periodontal disease and lack of its surveillance. Sci World J. 2020;2020:2146160-2146168. 10.1155/2020/2146160 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Kinane DF, Stathopoulou PG, Papapanou PN. Periodontal diseases. Nat Rev Dis Primers. 2017;3:17038. 10.1038/nrdp.2017.38 [DOI] [PubMed] [Google Scholar]
- 3. Vollmer A, Vollmer M, Lang G, et al. Automated assessment of radiographic bone loss in the posterior maxilla utilizing a multi-object detection artificial intelligence algorithm. Appl Sci. 2023;13(3):1858. 10.3390/app13031858 [DOI] [Google Scholar]
- 4. Uzun Saylan BC, Baydar O, Yeşilova E, et al. Assessing the effectiveness of artificial intelligence models for detecting alveolar bone loss in periodontal disease: a panoramic radiograph study. Diagnostics (Basel). 2023;13(10):1800. 10.3390/diagnostics13101800 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Bui FQ, Almeida-Da-Silva CLC, Huynh B, et al. Association between periodontal pathogens and systemic disease. Biomed J. 2019;42(1):27-35. 10.1016/j.bj.2018.12.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Corbet E, Ho D, Lai S. Radiographs in periodontal disease diagnosis and management. Aust Dent J. 2009;54(s1):S27-S43. 10.1111/j.1834-7819.2009.01141.x [DOI] [PubMed] [Google Scholar]
- 7. Tugnait A, Hirschmann PN, Clerehugh V. Validation of a model to evaluate the role of radiographs in the diagnosis and treatment planning of periodontal diseases. J Dent. 2006;34(7):509-515. 10.1016/j.jdent.2005.12.002 [DOI] [PubMed] [Google Scholar]
- 8. Weems RA, Manson-Hing LR, Jamison HC, Greer DF. Diagnostic yield and selection criteria in complete intraoral radiography. J Am Dent Assoc. 1985;110(3):333-338. 10.14219/jada.archive.1985.0320 [DOI] [PubMed] [Google Scholar]
- 9. Amasya H, Jaju PP, Ezhov M, et al. Development and validation of an artificial intelligence software for periodontal bone loss in panoramic imaging. Int J Imaging Syst Tech. 2024;34(1). 10.1002/ima.22973 [DOI] [Google Scholar]
- 10. Woelber J, Fleiner J, Rau J, Ratka-KrüGer P, Hannig C. Accuracy and usefulness of CBCT in periodontology: a systematic review of the literature. Int J Periodontics Restorative Dent. 2018;38(2):289-297. 10.11607/prd.2751 [DOI] [PubMed] [Google Scholar]
- 11. Revilla-León M, Gómez-Polo M, Barmak AB, et al. Artificial intelligence models for diagnosing gingivitis and periodontal disease: a systematic review. Rev J Prosth Dent. 2023;130(6):816-824. 10.1016/j.prosdent.2022.01.026 [DOI] [PubMed] [Google Scholar]
- 12. Misch KA, Yi ES, Sarment DP. Accuracy of cone beam computed tomography for periodontal defect measurements. J Periodontol. 2006;77(7):1261-1266. 10.1902/jop.2006.050367 [DOI] [PubMed] [Google Scholar]
- 13. Venkatesh E, Venkatesh Elluru S. Cone beam computed tomography: basics and applications in dentistry. J Istanb Univ Fac Dent. 2017;51(3 Suppl 1):S102-S121. 10.17096/jiufd.00289 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Thurzo A, Urbanová W, Novák B, et al. Where is the artificial intelligence applied in dentistry? Systematic review and literature analysis. Healthcare (Basel). 2022;10(7). 10.3390/healthcare10071269 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Ghods K, Azizi A, Jafari A, Ghods K. Application of artificial intelligence in clinical dentistry, a comprehensive review of literature. J Dent (Shiraz). 2023;24(4):356-371. 10.30476/dentjods.2023.96835.1969 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Hung KF, Ai QYH, Wong LM, Yeung AWK, Li DTS, Leung YY. Current applications of deep learning and radiomics on CT and CBCT for maxillofacial diseases. Diagnostics (Basel). 2022;13(1). 10.3390/diagnostics13010110 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Bayrakdar IS, Orhan K, Celik O, et al. A U-net approach to apical lesion segmentation on panoramic radiographs. Biomed Res Int. 2022;2022:7035367. 10.1155/2022/7035367 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Duong DQ, Nguyen KCT, Kaipatur NR, et al. Fully automated segmentation of alveolar bone using deep convolutional neural networks from intraoral ultrasound images. Annu Int Conf IEEE Eng Med Biol Soc. 2019;6632-6635. [DOI] [PubMed] [Google Scholar]
- 19. Cui Z, Fang Y, Mei L, et al. A fully automatic AI system for tooth and alveolar bone segmentation from cone-beam CT images. Nat Commun. 2022;13(1):2096. 10.1038/s41467-022-29637-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Kabir T, Lee CT, Nelson J, et al. An end-to-end entangled segmentation and classification convolutional neural network for periodontitis stage grading from periapical radiographic images. BBIM. 2021;1370-1375. [Google Scholar]
- 21. Lakshmi TK, Dheeba J. Classification and segmentation of periodontal cyst for digital dental diagnosis using deep learning. Comput Assist Methods Eng Sci. 2023;30(2):131-149. 10.24423/cames.505 [DOI] [Google Scholar]
- 22. Widyaningrum R, Candradewi I, Aji N, Aulianisa R. Comparison of multi-label U-net and mask R-CNN for panoramic radiograph segmentation to detect periodontitis. Imaging Sci Dent. 2022;52(4):383-391. 10.5624/isd.20220105 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Patil S, Joda T, Soffe B, et al. Efficacy of artificial intelligence in the detection of periodontal bone loss and classification of periodontal diseases: a systematic review. Rev J Am Dent Assoc (1939). 2023;154(9):795-804.e1. 10.1016/j.adaj.2023.05.010 [DOI] [PubMed] [Google Scholar]
- 24. Tariq A, Bin Nakhi F, Salah F, et al. Efficiency and accuracy of artificial intelligence in the radiographic detection of periodontal bone loss: a systematic review. Rev Imag Sci Dent. 2023;53(3):193-198. 10.5624/isd.20230092 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Kwong JCC, Khondker A, Lajkosz K, et al. APPRAISE-AI tool for quantitative evaluation of AI studies for clinical decision support. JAMA Netw Open. 2023;6(9):e2335377. 10.1001/jamanetworkopen.2023.35377 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Karacaoglu F, Kolsuz ME, Bagis N, Evli C, Orhan K. Development and validation of intraoral periapical radiography-based machine learning model for periodontal defect diagnosis. Proc Inst Mech Eng H J Eng Med. 2023;237(5):607-618. 10.1177/09544119231162682 [DOI] [PubMed] [Google Scholar]
- 27. Moran MBH, Faria M, Giraldi G, Bastos L, Da Silva Inacio B, Conci A. On using convolutional neural networks to classify periodontal bone destruction in periapical radiographs. Instit Electr Electron Eng Inc. 2020;2036-2039. [Google Scholar]
- 28. Page MJ, McKenzie JE, Bossuyt PM, et al. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ. 2021;372:n71. 10.1136/bmj.n71 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Haddaway NR, Page MJ, Pritchard CC, McGuinness LA. PRISMA2020: an R package and Shiny app for producing PRISMA 2020-compliant flow diagrams, with interactivity for optimised digital transparency and Open Synthesis. Campbell Syst Rev. 2022;18(2):e1230. 10.1002/cl2.1230 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Alotaibi G, Awawdeh M, Farook FF, Aljohani M, Aldhafiri RM, Aldhoayan M. Artificial intelligence (AI) diagnostic tools: utilizing a convolutional neural network (CNN) to assess periodontal bone level radiographically—a retrospective study. BMC Oral Health. 2022;22(1):399. 10.1186/s12903-022-02436-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Sameer S, Karthik J, Teja MS, Vardhan PA. Estimation of periodontal bone loss using SVM and random forest. Institute of Electrical and Electronics Engineers Inc., 2023. 2nd International Conference for Innovation in Technology, INOCON 2023, Bangalore, India. 10.1109/INOCON57975.2023.10101155 [DOI]
- 32. Ryu J, Lee DM, Jung YH, et al. Automated detection of periodontal bone loss using deep learning and panoramic radiographs: a convolutional neural network approach. Appl Sci (Switzerland). 2023;13(9):5261. 10.3390/app13095261 [DOI] [Google Scholar]
- 33. Mao YC, Huang YC, Chen TY, et al. Deep learning for dental diagnosis: a novel approach to furcation involvement detection on periapical radiographs. Bioengineering. 2023;10(7):802. 10.3390/bioengineering10070802 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Liu Q, Dai F, Zhu H, et al. Deep learning for the early identification of periodontitis: a retrospective, multicentre study. Clin Radiol. 2023;78(12):e985-e992. 10.1016/j.crad.2023.08.017 [DOI] [PubMed] [Google Scholar]
- 35. Kong Z, Ouyang H, Cao Y, et al. Automated periodontitis bone loss diagnosis in panoramic radiographs using a bespoke two-stage detector. Comput Biol Med. 2023;152:106374. 10.1016/j.compbiomed.2022.106374 [DOI] [PubMed] [Google Scholar]
- 36. Chen IH, Lin CH, Lee MK, et al. Convolutional-neural-network-based radiographs evaluation assisting in early diagnosis of the periodontal bone loss via periapical radiograph. J Dent Sci. 2024;19(1):550-559. 10.1016/j.jds.2023.09.032 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Chen CC, Wu YF, Aung LM, et al. Automatic recognition of teeth and periodontal bone loss measurement in digital radiographs using deep-learning artificial intelligence. J Dent Sci. 2023;18(3):1301-1309. 10.1016/j.jds.2023.03.020 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Tsoromokos N, Parinussa S, Claessen F, Moin DA, Loos BG. Estimation of alveolar bone loss in periodontitis using machine learning. Int Dent J. 2022;72(5):621-627. 10.1016/j.identj.2022.02.009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Shon HS, Kong V, Park JS, et al. Deep learning model for classifying periodontitis stages on dental panoramic radiography. Appl Sci (Switzerland). 2022;12(17):8500. 10.3390/app12178500 [DOI] [Google Scholar]
- 40. Lee CT, Kabir T, Nelson J, et al. Use of the deep learning approach to measure alveolar bone level. J Clin Periodontol. 2022;49(3):260-269. 10.1111/jcpe.13574 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Kearney VP, Yansane AIM, Brandon RG, et al. A generative adversarial inpainting network to enhance prediction of periodontal clinical attachment level. J Dent. 2022;123:104211. 10.1016/j.jdent.2022.104211 [DOI] [PubMed] [Google Scholar]
- 42. Jiang L, Chen D, Cao Z, Wu F, Zhu H, Zhu F. A two-stage deep learning architecture for radiographic staging of periodontal bone loss. BMC Oral Health. 2022;22(1):106. 10.1186/s12903-022-02119-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Chang J, Chang MF, Angelov N, et al. Application of deep machine learning for the radiographic diagnosis of periodontitis. Clin Oral Investig. 2022;26(11):6629-6637. 10.1007/s00784-022-04617-4 [DOI] [PubMed] [Google Scholar]
- 44. Li H, Zhou J, Zhou Y, et al. An interpretable computer-aided diagnosis method for periodontitis from panoramic radiographs. Front Physiol. 2021;12:655556. 10.3389/fphys.2021.655556 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Chang HJ, Lee SJ, Yong TH, et al. Deep learning hybrid method to automatically diagnose periodontal bone loss and stage periodontitis. Sci Rep. 2020;10(1):7531. 10.1038/s41598-020-64509-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Kim J, Lee HS, Song IS, Jung KH. DeNTNet: deep neural transfer network for the detection of periodontal bone loss using panoramic dental radiographs. Research support, non-U.S. Gov’t validation study. Science. 2019;9(1):17615. 10.1038/s41598-019-53758-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Krois J, Ekert T, Meinhold L, et al. Deep learning for the radiographic detection of periodontal bone loss. Research support, non-U.S. Gov’t. Sci. 2019;9(1):8495. 10.1038/s41598-019-44839-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Lee J-H, Kim D-H, Jeong S-N, Choi S-H. Diagnosis and prediction of periodontally compromised teeth using a deep learning-based convolutional neural network algorithm. J Periodontal Implant Sci. 2018;48(2):114-123. 10.5051/jpis.2018.48.2.114 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Danks RP, Bano S, Orishko A, et al. Automating periodontal bone loss measurement via dental landmark localisation. Int J Comput Assist Radiol Surg. 2021;16(7):1189-1199. 10.1007/s11548-021-02431-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Chen H, Li H, Zhao Y, Zhao J, Wang Y. Dental disease detection on periapical radiographs based on deep convolutional neural networks. Int J Comput Assist Radiol Surg. 2021;16(4):649-661. 10.1007/s11548-021-02319-y [DOI] [PubMed] [Google Scholar]
- 51. Thanathornwong B, Suebnukarn S. Automatic detection of periodontal compromised teeth in digital panoramic radiographs using faster regional convolutional neural networks. Imaging Sci Dent. 2020;50(2):169-174. 10.5624/isd.2020.50.2.169 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52. Kurt S, Çelik Ö, Bayrakdar İŞ, et al. Success of artificial intelligence system in determining alveolar bone loss from dental panoramic radiography images. Cumhuriyet Dent J. 2020;23(4):318-324. 10.7126/cumudj.777057 [DOI] [Google Scholar]
- 53. Patil S, Albogami S, Hosmani J, et al. Artificial intelligence in the diagnosis of oral diseases: applications and pitfalls. Diagnostics (Basel). 2022;12(5):1029. 10.3390/diagnostics12051029 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54. Sivari E, Senirkentli GB, Bostanci E, Guzel MS, Acici K, Asuroglu T. Deep learning in diagnosis of dental anomalies and diseases: a systematic review. Diagnostics (Basel). 2023;13(15):2512. 10.3390/diagnostics13152512 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55. Turosz N, Chęcińska K, Chęciński M, Brzozowska A, Nowak Z, Sikora M. Applications of artificial intelligence in the analysis of dental panoramic radiographs: an overview of systematic reviews. Dentomaxillofac Radiol. 2023;52(7):20230284. 10.1259/dmfr.20230284 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56. Maxim LD, Niebo R, Utell MJ. Screening tests: a review with examples. Inhal Toxicol. 2014;26(13):811-828. 10.3109/08958378.2014.955932 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Habibzadeh F, Habibzadeh P, Yadollahie M. On determining the most appropriate test cut-off value: the case of tests with continuous results. Biochem Med (Zagreb). 2016;26(3):297-307. 10.11613/bm.2016.034 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. Lalkhen AG, McCluskey A. Clinical tests: sensitivity and specificity. Contin Edu Anaesthesia Crit Care Pain. 2008;8(6):221-223. 10.1093/bjaceaccp/mkn041 [DOI] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.