Abstract
Introduction
Anifrolumab is a type I interferon (IFN) receptor 1 (IFNAR1) blocking antibody approved for treating patients with systemic lupus erythematosus (SLE). Here, we investigated the immunomodulatory mechanisms of anifrolumab using longitudinal transcriptomic and proteomic analyses of the 52-week, randomised, phase 3 TULIP-1 and TULIP-2 trials.
Methods
Patients with moderate to severe SLE were enrolled in TULIP-1 and TULIP-2 and received intravenous anifrolumab or placebo alongside standard therapy. Whole-blood expression of 18 017 genes using genome-wide RNA sequencing (RNA-seq) (pooled TULIP; anifrolumab, n=244; placebo, n=258) and 184 plasma proteins using Olink and Simoa panels (TULIP-1; anifrolumab, n=124; placebo, n=132) were analysed. We compared treatment groups via gene set enrichment analysis using MetaBase pathway analysis, blood transcriptome modules, in silico deconvolution of RNA-seq and longitudinal linear mixed effect models for gene counts and protein levels.
Results
Compared with placebo, anifrolumab modulated >2000 genes by week 24, with overlapping results at week 52, and 41 proteins by week 52. IFNAR1 blockade with anifrolumab downregulated multiple type I and II IFN-induced gene modules/pathways and type III IFN-λ protein levels, and impacted apoptosis-associated and neutrophil extracellular traps-(NET)osis-associated transcriptional pathways, innate cell activating chemokines and receptors, proinflammatory cytokines and B-cell activating cytokines. In silico deconvolution of RNA-seq data indicated an increase from baseline of mucosal-associated invariant and γδT cells and a decrease of monocytes following anifrolumab treatment.
Discussion
Type I IFN blockade with anifrolumab modulated multiple inflammatory pathways downstream of type I IFN signalling, including apoptotic, innate and adaptive mechanisms that play key roles in SLE immunopathogenesis.
Keywords: Biological Therapy; Lupus Erythematosus, Systemic; Autoimmune Diseases
Video Abstract.
Disclaimer: this video summarises a scientific article published by BMJ Publishing Group Limited (BMJ). The content of this video has not been peer-reviewed and does not constitute medical advice. Any opinions expressed are solely those of the contributors. Viewers should be aware that professionals in the field may have different opinions. BMJ does not endorse any opinions expressed or recommendations discussed. Viewers should not use the content of the video as the basis for any medical treatment. BMJ disclaims all liability and responsibility arising from any reliance placed on the content.
WHAT IS ALREADY KNOWN ON THIS TOPIC
Impaired cell death processes and increased type I interferon (IFN) signalling play central roles in the loss of immune tolerance in patients with systemic lupus erythematosus (SLE).
Analyses of data from a phase 2 trial revealed that type I IFN signalling blockade with anifrolumab suppressed inflammatory proteins associated with disease activity, improved markers of cardiometabolic disease and reversed SLE-related lymphopenia, neutropenia and thrombocytopaenia.
WHAT THIS STUDY ADDS
We provide key insights into the immunomodulatory mechanisms underpinning type I IFN signalling blockade with anifrolumab treatment, in the first transcriptomic and proteomic analysis performed on data from two phase 3 trials of patients with moderate to severe SLE despite standard therapy.
Type I IFN signalling blockade with anifrolumab suppressed apoptosis and neutrophil extracellular traps- (NET)osis cell death pathways, which contribute to the complex immunopathogenesis of SLE.
Anifrolumab treatment impacted both innate and adaptive immunity; it downmodulated multiple plasma cell and monocyte gene modules, upregulated T-cell and lymphocyte modules, and was associated with increased lymphocyte counts, suggesting that IFNAR1 inhibition with anifrolumab may help reverse the imbalances in immune cell composition observed in patients with SLE.
HOW THIS STUDY MIGHT AFFECT RESEARCH, PRACTICE OR POLICY
Our findings provide valuable insights into the mechanisms by which inhibition of type I IFN signalling leads to favourable clinical outcomes through broad anti-inflammatory effect on multiple pathways downstream of IFNAR1, which are key drivers of SLE pathogenesis
Introduction
Systemic lupus erythematosus (SLE) is a complex autoimmune disease in which patients experience diverse organ involvement and recurrent flares, making diagnosis and treatment challenging.1 2 SLE immunopathogenesis is characterised by increased circulating levels of cell death debris, caused by impaired clearance of apoptotic cells and neutrophil extracellular traps (NETs).1 2 The subsequent build-up of and increased sensitivity to self-antigens leads to an increase in type I interferon (IFN-I) pathway activation and promotes the formation of immune complexes, which together cause an inflammatory response that contributes to tissue damage.2 These impaired innate and adaptive immunological processes are reflected in dysregulation of the phenotype and function of a wide range of cell types, including B cells, T cells, monocytes, neutrophils and plasmacytoid dendritic cells (pDCs).2
Genomic, transcriptomic and proteomic analyses have provided evidence for the key role of type I IFN pathway activation in SLE pathogenesis. High serum IFN-α is thought to be an hereditable risk factor for SLE, and there are multiple genetic variants in type I IFN pathway genes that are associated with SLE risk.3 Elevated type I IFN signalling, measured using IFN gene signatures (IFNGS), is associated with disease activity, organ involvement and the activity of other immune pathways in patients with SLE.4–7 Transcriptomic analyses have identified additional dysregulated biological pathways in SLE, including B-cell activating factor (BAFF) signalling and modules of genes associated with inflammation and lymphoid signalling.6 Proteomic analyses have shown that elevated type I IFN proteins are associated with SLE inflammatory markers.5 8
The key role of type I IFN in SLE pathogenesis led to development of anifrolumab, a human Ig G1κ antibody that binds to the type I IFN receptor subunit 1 (IFNAR1) to inhibit type I IFN signalling.9 Anifrolumab has been approved as a treatment for adult patients with moderate to severe SLE9 based on results from the phase 2 MUSE trial and two phase 3 trials, TULIP-1 and TULIP-2.10–12 In these studies, anifrolumab treatment reduced overall disease activity compared with placebo.10–12 Anifrolumab also had an acceptable safety and tolerability profile.10–12
Analyses of data from the MUSE trial revealed that anifrolumab suppressed inflammatory proteins associated with disease activity, improved markers of cardiometabolic disease and reversed SLE-related lymphopenia, neutropenia and thrombocytopaenia, supporting a wide-reaching effect of IFNAR1 blockade.8 13 In this study, we used transcriptomic and proteomic analyses of the TULIP-1 and TULIP-2 patient population to further explore the immunomodulatory mechanisms underlying the effects of IFNAR1 blockade with anifrolumab in SLE.
Methods
Study design and patients
Details of the TULIP-1 (NCT02446912) and TULIP-2 (NCT02446899) trials have been previously published.11 12 Briefly, both studies were multicentre, double-blind, randomised, placebo-controlled, 52-week phase 3 trials that enrolled adult patients with moderate to severe SLE despite standard therapy. Patients were randomised to receive intravenous anifrolumab (300 mg in TULIP-1 and TULIP-2; 150 mg in TULIP-1 only) or placebo every 4 weeks for 48 weeks alongside standard therapy. The trial design included mandatory attempts to taper glucocorticoids. In both studies, randomisation was stratified by SLE Disease Activity Index 2000 (SLEDAI-2K) score at screening (<10 vs ≥10 points14), glucocorticoid dosage at baseline (<10 vs ≥10 mg/day prednisone/equivalent) and type I 4-gene (IFN-α-inducible protein 27 (IFI27), IFN-induced protein 44 (IFI44), IFN-induced protein 44 like (IFI44L) and radical s-adenosyl methionine domain containing 2 (RSAD2)) IFNGS test at screening (high vs low status).11 12 Analyses described here used data only from the anifrolumab 300 mg (the approved dose) and placebo groups.
Whole blood transcriptomics
Whole blood samples (n=1506 samples; n=502 patients) were collected in PAXgene tubes at baseline, week 24, and week 52, and were stored at −80°C at a central laboratory. All samples were shipped together and whole blood genome-wide RNA sequencing (RNA-seq) was performed by Q2 Solutions Expression Analysis for each sample using Illumina’s TruSeq Stranded Total RNA protocol (Illumina, San Diego, California, USA) as per manufacturer’s instructions. Paired-end sequencing with 100 bp reads was performed, with 40M reads per sample generated.
Raw sequencing data in FASTQ format were processed using an in-house RNA-seq pipeline. Briefly, read quality for all samples was assessed using FastQC (V.0.11.7),15 QualiMap (V.2.2.2c)16 and SAMtools-stats (V.1.9).17 Quality control (QC) metrics for QualiMap were based on a STAR (V.2.7.2b)18 alignment against the human genome (GRCh38.p13, Ensembl V.100). QC metrics were summarised using MultiQC (V.1.9).19 Sequencing adapters were then trimmed from the remaining libraries using NGmerge (V.0.3).20 A human transcriptome index consisting of cDNA and ncRNA entries from Ensembl (V.100) was generated and reads were mapped to the index using Salmon (V.1.1.0).21 The bioinformatics workflow was organised using Nextflow workflow management system (V.20.01)22 and Bioconda software management tools.23
RNA-seq samples were assessed for outliers using the R package ClassDiscovery (V.3.4.0).24 Outlier samples (n=13) were iteratively removed based on their Mahalanobis distance to centroid by principal component analysis, using the top 5000 variable genes. Downstream analysis was conducted using only patients with baseline, week 24 and week 52 samples that passed QC measures (n=1473 samples; n=491 patients; anifrolumab 300 mg, n=241; placebo, n=250).
Low count genes (<5 counts in >50% of the samples) were excluded from differential expression analysis, which was performed on 18 017 genes and conducted with a longitudinal linear mixed effect model adjusted for randomisation stratification factors (glucocorticoids dosage </≥10 mg/day, IFNGS high/low status at screening and SLEDAI-2K score 0–10/≥10) using R package variancePartition (V.1.20.0, dream function) with repeated measures designs.25 No significant batch effects were observed. The estimated coefficients and p values for the treatment-time interaction term were displayed in volcano plots, using the R package EnhancedVolcano (V.1.13.2).26 Gene functions were confirmed using the National Center for Biotechnology Information (NCBI) library.27
Gene set enrichment analysis (GSEA) was performed using the R package clusterProfiler (V.4.3.3),28 MetaBase pathway database (V.22.2.7),29 and a fixed repertoire of blood transcriptome modules previously developed to support analysis and interpretation of blood transcriptome data.30 GSEA was conducted on all ranked genes using the z-standard scores from differential expression analysis. This method returns a Normalised Enrichment Score (NES), to allow for comparisons across gene sets of different sizes, and a false discovery rate (FDR)-adjusted p value (PFDR) for multiple testing.
In silico deconvolution of RNA-seq data was used for estimating cell type abundances. This was performed by applying support vector regression (SVR) using a reference signature matrix from 13 immune cell types isolated from healthy donor peripheral blood mononuclear cells (PBMC).31 SVR was implemented using the R package granulator (V.1.1.0).32
Plasma proteomics
Whole blood for plasma samples (n=768 samples; n=317 patients) were collected using EDTA-treated tubes during the TULIP-1 trial at baseline, week 12 and week 52. Plasma samples were stored at −80°C until analysis and were assessed for 184 protein analytes using two targeted Olink platform panels (Olink Target 96 Cardiovascular III, V.6113 and Olink Target 96 Immuno-Oncology, V.3112; Olink, Upsala, Sweden). Five protein analytes with low serum concentrations (IFN lambda 1 (IFNL1), interleukin-1 beta (IL1B), IFN alpha (IFNA1/IFNA13), interleukin-17A (IL17A) and interleukin-21 (IL21)) were also assessed in the same samples using ultra-sensitive Simoa technology (RBM, Austin, Texas, USA).
Downstream proteomics analysis was conducted using only patients with baseline, week 12 and week 52 samples that passed QC assessments, and for proteins above the limit of detection (LOD) threshold in ≥50% of samples (n=169 proteins; n=256 patients; anifrolumab 300 mg, n=124; placebo, n=132). Data from Olink-assessed proteins (n=164) were presented in relative Normalised Protein eXpression (NPX), and data from Simoa (n=5) were presented in pg/mL. A longitudinal linear mixed effect model adjusted for randomisation stratification factors was used to compare the 1-year trajectories of protein levels between patients receiving anifrolumab 300 mg versus placebo.
Lymphocyte cell count
Blood samples from patients were collected at baseline and every 4 weeks. Lymphocyte numbers were derived from the complete blood count with the differential cell count.
Statistics analysis
Detailed statistical analyses are described alongside the transcriptomics-related and proteomics-related methodologies above. When applicable, p values were adjusted for multiple testing using the Benjamini-Hochberg procedure for calculating FDR.
Patient and public involvement
Patients and/or the public were not involved in the design, conduction, reporting or dissemination of this research.
Results
Patient demographics and baseline characteristics
Of 726 patients who received anifrolumab 300 mg or placebo in the TULIP-1 or TULIP-2 trials, 502 patients comprised the transcriptomic dataset (anifrolumab 300 mg, n=244; placebo, n=258) and 256 patients comprised the proteomics dataset (from the TULIP-1 trial only, anifrolumab 300 mg, n=124; placebo, n=132; online supplemental figure S1). Generally, demographics and baseline characteristics for patients in the transcriptomics and proteomics datasets were consistent with the overall study populations of the TULIP trials.11 12 Over 90% of patients were female, and the majority of patients were White (table 1). Approximately 50% of patients in the transcriptomics dataset and 71% of patients in the proteomics dataset had SLEDAI-2K≥10 at baseline. In the transcriptomics and proteomic datasets, 82% and 78% of patients were IFNGS-high at screening, respectively. At baseline, ~50% of patients in the transcriptomics set and 56% of patients in the proteomics set were receiving ≥10 mg/day of glucocorticoids.
Table 1.
Demographics and baseline characteristics of patients with moderate to severe SLE who participated in the TULIP-1 or TULIP-2 trials and had available transcriptomics or proteomics data
| Transcriptomic dataset | Proteomic dataset* | ||||
| Anifrolumab 300 mg | Placebo | Anifrolumab 300 mg | Placebo | ||
| Patients | n | 244 | 258 | 124 | 132 |
| Age, years | Median (range) | 44.0 (18–69) | 42.5 (18–68) | 40.0 (18–68) | 42.0 (18–68) |
| Sex, n (%) | Female | 224 (91.8) | 239 (92.6) | 113 (91.1) | 124 (93.9) |
| Male | 20 (8.2) | 19 (7.4) | 11 (8.9) | 8 (6.1) | |
| Race, n (%)† | American Indian/Alaska Native | 4 (1.7) | 1 (0.4) | 0 | 0 |
| Asian | 12 (5.0) | 16 (6.3) | 8 (6.5) | 4 (3.0) | |
| Black/African American | 29 (12.0) | 35 (13.8) | 16 (12.9) | 17 (12.9) | |
| Other | 20 (8.3) | 22 (8.7) | 1 (0.8) | 2 (1.5) | |
| White | 176 (73.0) | 179 (70.8) | 99 (79.8) | 109 (82.6) | |
| GCs dosage, n (%)ठ| <10 mg/day | 119 (48.8) | 132 (51.2) | 54 (44.3) | 58 (43.9) |
| ≥10 mg/day | 125 (51.2) | 126 (48.8) | 68 (55.7) | 74 (56.1) | |
| SLEDAI-2K score, categorical, n (%)‡ | 0–10 | 119 (48.8) | 132 (51.2) | 36 (29.0) | 38 (28.8) |
| ≥10 | 125 (51.2) | 126 (48.8) | 88 (71.0) | 94 (71.2) | |
| IFN 4-gene test status‡¶, n (%) | High | 199 (81.6) | 211 (81.8) | 95 (77.9) | 103 (78.0) |
| Low | 45 (18.4) | 47 (18.2) | 27 (22.1) | 29 (22.0) | |
*The proteomics dataset comprised patients from TULIP-1 only.
†Race group was reported by the patients.
‡The baseline characteristics presented here were stratification factors used for randomisation of patients at the start of the TULIP-1 and TULIP-2 trials.
§Oral prednisone or equivalent.
¶IFNGS test status at screening.
GCs, glucocorticoids; IFN, interferon; n, number of patients; SLE, systemic lupus erythematosus; SLEDAI-2K, SLE Disease Activity Index 2000.
ard-2023-225445supp001.pdf (5.2MB, pdf)
Differentially expressed genes in patients receiving anifrolumab 300 mg versus placebo
Differential expression analysis was performed on 18 017 genes in samples from 491 patients. The effect of anifrolumab versus placebo on gene expression was in agreement between patients in the TULIP-1 and TULIP-2 data sets at week 24 (figure 1A) and at week 52 (figure 1B). The impact of anifrolumab versus placebo was also consistent between weeks 24 and 52 for pooled TULIP data (figure 1C).
Figure 1.
Effects of anifrolumab treatment on gene regulation in TULIP-1 and TULIP-2. Linear mixed effect model (z-score) for the effect of anifrolumab versus placebo in TULIP-1 plotted against TULIP-2 at (A) week 24 and (B) week 52 and (C) in week 24 vs week 52 in the pooled TULIP-1 and TULIP-2.a Each gene (individual point) could be downregulated in both trials/time points with FDR-adjusted p≤0.05 (blue), upregulated in both trials/visits with FDR-adjusted p≤0.05 (red) or not significantly different between treatment arms (grey). (D) Volcano plot for the effect of anifrolumab versus placebo at weeks 24 (left) and 52 (right) in pooled TULIP-1 and TULIP-2 data. Negative log10 of nominal p value versus log2 fold change (FC) were calculated for each gene using a linear mixed effect model. Genes could be significantly upregulated (blue points right of vertical dash line, FDR-adjusted p<0.05 and log2 FC>0.5) or significantly downregulated (blue points left of vertical dash line, FDR-adjusted p<0.05 and log2 FC <−0.5) by anifrolumab 300 mg versus placebo, or not significantly different between anifrolumab 300 mg versus placebo groups (purple and grey points; FDR-adjusted p>0.05). For all panels, anifrolumab n=241 and placebo n=250. aDifferentially modulated genes in patients receiving anifrolumab versus placebo at week 24 or 52 can be found in online supplemental table S1. FC, fold change; FDR, false discovery rate; NS, not significant; SLE, systemic lupus erythematosus.
At week 24, 1681 genes were downregulated (PFDR<0.05) and 1170 were upregulated (PFDR<0.05) by anifrolumab versus placebo (figure 1D and online supplemental table S1). At week 52, 2092 genes were downregulated and 1645 were upregulated by anifrolumab versus placebo (figure 1D and online supplemental table S1). Of the genes that were upregulated by anifrolumab compared with placebo, 789 were common between weeks 24 and 52; 1420 genes were downregulated at both weeks 24 and 52. As expected, type I IFN inducible genes11 12 27 33 were among those most significantly downregulated by anifrolumab versus placebo (eg, IFI27, week 24: log2FC=−4.5, week 52: log2FC=−4.18; IFI44, week 24: log2FC=−2.59, week 52: log2FC=−2.43; IFI44L, week 24: log2FC=−3.28, week 52: log2FC=−3.13; RSAD2, week 24: log2FC=−3.27, week 52: log2FC=−3.06; ISG15, week 24: log2FC=−2.51; week 52: log2FC=−2.30). Type II and combined type I/II IFN inducible genes27 were also significantly downregulated by anifrolumab (eg, IFI16, week 24: log2FC=−0.62, week 52: log2FC=−0.59; MX dynamin like GTPase 1 (MX1), week 24: log2FC=−2.38, week 52: log2FC=−2.27; signal transducer and activator of transcription 1 (STAT1), week 24: log2FC=−0.54, week 52: log2FC=−0.47; STAT2, week 24: log2FC=−0.75, week 52: log2FC=−0.71). Several other genes involved in innate and adaptive immune response27 34 35 were also downregulated at weeks 24 and 52 (eg, sialic acid binding Ig like lectin 1 (SIGLEC1), week 24: log2FC=−3.69, week 52: log2FC=−3.56; tumour necrosis factor (TNF) ligand superfamily 13B (TNFSF13B), also known as B-cell activating factor (BAFF), week 24: log2FC=−0.58, week 52: log2FC=−0.52; cluster of differentiation 38 (CD38), week 24: log2FC=−0.52, week 52: log2FC=−0.47). Overall, similarities observed between follow-up time points and the consistency observed between trials supported pooling the TULIP data for further transcriptomic analysis.
Differentially regulated pathways in patients receiving anifrolumab 300 mg versus placebo
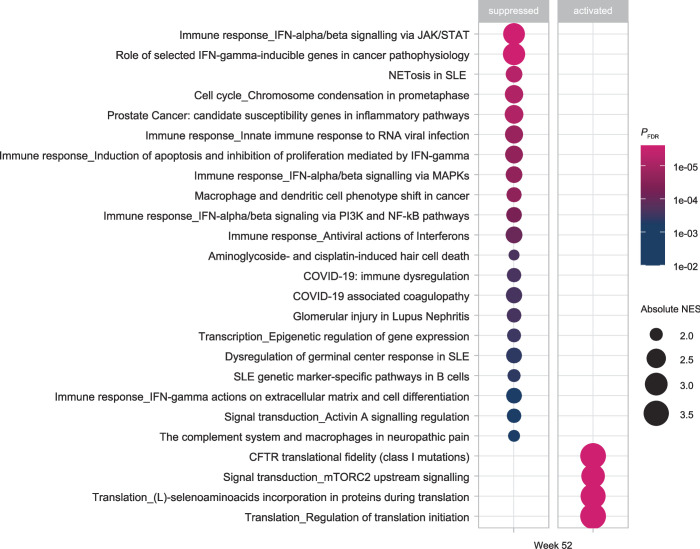
GSEA using the MetaBase database on pooled TULIP data revealed the top 25 pathways that were significantly downregulated or upregulated by anifrolumab compared with placebo at week 52 (upregulated: NES>0; downregulated: NES<0; PFDR≤0.001; figure 2 and online supplemental table S2). Several MetaBase intracellular signalling pathways downstream of IFN-α/-β29 were suppressed, including via Janus kinases/signal transducer and activator of transcription (JAK/STAT; NES=−2.84), via phosphoinositide 3-kinases and nuclear factor kappa light chain enhancer of activated B cells (NF-kB; NES=−2.13), and via mitogen-activated protein kinase (MAPK; NES=−2.21) pathways. Multiple MetaBase pathways associated with IFN-γ29 were downregulated, including IFN-γ mediated apoptosis induction and inhibition of proliferation (NES=−2.32) and IFN-γ actions on extracellular matrix and cell differentiation (NES=−2.13). A NETosis-associated MetaBase pathway29 was also downregulated (NES=−2.21). Several MetaBase pathways associated with protein synthesis29 were upregulated (eg, regulation of translation initiation, NES=3.59).
Figure 2.
Pathway analysis for genes that were downregulated or upregulated by anifrolumab. The top 25 most significantly dysregulated pathways in patients receiving anifrolumab (n=241) vs placebo (n=250) at week 52 in pooled TULIP-1 and TULIP-2 data (PFDR≤0.001).a GSEA using the MetaBase pathway database was conducted on a list of genes ranked by the z-standard score calculated using a longitudinal linear mixed effect model. Each suppressed or activated pathway is represented by an individual point (larger size for greater absolute NES; lighter shade for greater significance of FDR-adjusted p value). aDifferentially regulated pathways in patients receiving anifrolumab versus placebo can be found in online supplemental table S2. FDR, false discovery rate; GSEA, gene set enrichment analysis; NES, normalised enrichment score; PFDR, FDR-adjusted p value; SLE, systemic lupus erythematosus.
Impact of anifrolumab 300 mg versus placebo on blood transcriptome modules and cell counts
Additional GSEA using fixed repertoires of transcriptional modules30 revealed several modules that were significantly downregulated or upregulated by anifrolumab at week 52 (figure 3A; online supplemental figure S2 and online supplemental table S3). Modules (M) that were significantly (PFDR≤0.01) downregulated by anifrolumab versus placebo at week 52 included those associated with IFN activity (eg, M10.1, NES=−2.68; M8.3, NES=−2.66; M15.127, NES=−2.38; M13.17, NES=−2.84; M15.64, NES=−2.62). Downregulated non-IFN pathways included modules for antigen presentation (M15.47, NES=−2.03), neutrophils (M15.26, NES=−1.88) and monocytes (M15.7, NES=−1.86). Several modules associated with lymphocytes (M13.27, NES=2.10) and T cells (M12.6, NES=2.14) were significantly upregulated, as were modules associated with protein modification and synthesis (M11.1, NES=2.84; M13.28, NES=2.73) (PFDR≤0.01).
Figure 3.
Effect of anifrolumab on lymphocyte modules and peripheral blood lymphocyte count. (A) Gene set enrichment analysis of blood transcriptome modules at week 52 that were upregulated or downregulated by anifrolumab versus placebo, ordered by NES and categorised according to immune pathway (PFDR≤0.01).a (B) In-silico RNA-seq deconvolution for predicting cell type abundance performed by applying support vector regression; plotted for baseline, week 24 and week 52 for anifrolumab or placebo as mean estimated proportion ±95% CI, in reference to a signature matrix of 13 cell types from healthy individuals.b (C) Lymphocyte counts from a complete blood count assay, plotted as mean GI/L ±95% CI from baseline to week 52 for anifrolumab and placebo. For (A, B), anifrolumab n=241 and placebo n=250 .c aAnifrolumab 300 mg versus placebo on each blood transcriptome module can be found in online supplemental table S3. bIn-silico RNA-seq deconvolution data for all cell type predictions can be found in online supplemental table S4 and online supplemental figure S3. cLymphocyte counts for all time points can be found in online supplemental table S5. FDR, false discovery rate; MAIT, mucosal-associated invariant T; NES, normalised enrichment score; PFDR, FDR-adjusted p value.
These results were supported by in silico deconvolution RNA-seq analysis to estimate cell type abundance of 13 cell types in bulk RNA-seq data (figure 3B; online supplemental figure S3 and online supplemental table S4). In patients receiving anifrolumab, the estimated proportion of mucosal-associated invariant T (MAIT) cells and γδT cells increased, and the estimated proportion of monocytes decreased from baseline to week 52, while they remained stable in patients receiving placebo. Deconvolution data did not show clear differences between treatment groups for the estimated abundance of B cells, basophils, pDCs, neutrophils, memory CD4 or CD8 T cells, or naïve T cells (online supplemental figure S3 and online supplemental table S4).
As anifrolumab treatment was associated with upregulation of lymphocyte-associated gene modules, next we compared lymphocyte trajectories over time in the anifrolumab and placebo groups as part of a complete blood count (figure 3C and online supplemental table S5). Lymphocyte counts rapidly and consistently increased from baseline to week 8 and remained stable until week 52 on treatment with anifrolumab, but not with placebo.
Proteins longitudinally modulated by anifrolumab 300 mg versus placebo
Of the 169 proteins from 256 patients that measured above the LOD threshold (Olink: 164; Simoa: 5), 1-year trajectories of 41 were modulated by anifrolumab versus placebo (longitudinal mixed effect model, figure 4; online supplemental table S6 and online supplemental table S7). Anifrolumab downmodulated 34 proteins (PFDR≤0.05), including: IFN-λ (log2FC=−0.75), IFN-γ-induced chemokines35 36 (eg, C-X-C motif chemokine ligand (CXCL) 10, log2FC=−1.05; CXCL16, log2FC=−0.34), chemokines known to regulate and recruit monocytes35 (monocyte chemotactic protein (MCP)-2, log2FC=−0.62; MCP-1 or chemokine (CC-motif) ligand 2 (CCL2), log2FC=−0.39), cytokines and chemokines with B-cell and T-cell regulatory functions35 37 (BAFF, log2FC=−0.40; TNF, log2FC=−0.41) and T-cell regulatory molecules35 38 (Galectin-9, log2FC=−0.61; lymphocyte activation gene three protein (LAG3), log2FC=−0.77). Seven proteins were significantly (PFDR≤0.05) upmodulated, including TNF-related apoptosis-inducing ligand receptor 3 (TRAIL-R3, log2FC=0.37). IFN-α protein (IFNA1/IFNA13) was not significantly impacted by anifrolumab (log2FC=0.40; PFDR=0.51).
Figure 4.
Effect of anifrolumab treatment versus placebo on protein expression in the TULIP-1 trial. (A) Longitudinal linear mixed effect models comparing the 1-year trajectories of protein levels between patients receiving anifrolumab 300 mg versus placebo.a,b Proteins are ordered on the x-axis and coloured according to their FDR-adjusted p value. Mean log2 fold changes between the anifrolumab and placebo groups at week 52 were adjusted for baseline stratification factors. 41 of the 169 proteins tested had significantly different 1-year trajectories between anifrolumab and placebo (PFDR≤0.05). (B) Relative Normalised Protein eXpression (NPX) log2 scale (or pg/mL for IFN-λ) of the top 10 longitudinally impacted proteins on treatment with anifrolumab from baseline to week 52 (PFDR<0.00001).b For all panels, anifrolumab n=124 and placebo n=132. aProteins longitudinally modulated by anifrolumab 300 mg versus placebo can be found in online supplemental table S6. bRelative NPX of the top 10 longitudinally impacted proteins on treatment with anifrolumab from baseline to week 52 can be found in online supplemental table S7. CCL, chemokine (C-C motif) ligand; CXCL, chemokine (C-X-C motif) ligand; FC, fold change; FDR, false discovery rate; GNR, granulin precursor; IFNL, interferon-λ; IP, IFN-γ inducible protein; LAG, lymphocyte-activation gene; LGALS, galectin; LAMP, lysosomal associated membrane protein; MCP, Monocyte chemotactic protein; PFDR, FDR-adjusted p value; TRAIL-R, tumour necrosis factor-related apoptosis-inducing ligand-receptor; TNFRSF, TNF receptor superfamily.
Discussion
Despite its complex immunopathology, SLE has consistent immune features, including the chronic activation of type I IFNs in patients with moderate to severe disease.1 Anifrolumab binds to IFNAR1 to inhibit type I IFN signalling, leading to strong suppression of IFNGS which is not observed in patients receiving placebo11 12 39; however, it was not previously known how anifrolumab impacts the diverse immune mechanisms downstream of the type I IFN pathway. This extensive transcriptomic and proteomic analysis of data from two phase 3 trials reveals important new insights into the immunomodulatory mechanisms underlying type I IFN blockade with anifrolumab treatment.
First, in addition to the expected modulation of type I IFN-induced genes, anifrolumab suppressed multiple genes and pathways annotated by the NCBI and Metabase databases as regulated by type II IFN (IFN-γ),27 29 and decreased levels of type III IFN (IFN-λ) protein. Anifrolumab may have downstream inhibitory effects on the targets of type II and III IFN signalling due to the considerable overlap of IFN-dependent transcriptional elements, including activation of a variety of STAT and interferon-regulatory factors by both IFNAR and IFN-γ receptors, as well as the presence of interferon-sensitive response elements and gamma-activated site elements in many interferon-stimulated genes.40 Although the role of type II and III IFN pathways remains unresolved in SLE, they have been implicated in disease pathogenesis owing to the association between IFN-γ and clinical disease onset and flares,1 and the role of type III IFN in tissue-specific inflammation,41 suggesting that their inhibition may benefit patients with SLE. Anifrolumab did not impact IFN-α protein levels in this study, contrary to what might be predicted from models of SLE that include indirect feed-forward effects of IFN-α on pDCs.1 Of note, substantial IFN production may occur from non-circulating sources,42 therefore, we cannot rule out tissue-specific changes in IFN-α protein levels.
Elevated apoptosis and NETosis are key drivers of SLE pathogenesis, with levels of apoptotic cells and NETosis correlating with disease activity in patients with SLE.43 44 Here, we show that anifrolumab modulated apoptosis and NETosis-related pathways, which have been hypothesised to lead to IFN-α production.2 GSEA revealed downregulation of Metabase gene pathways for NETosis in SLE and IFN-γ-induced apoptosis (eg, FAS, CASP1, CASP3 and CASP7 genes) with anifrolumab. Additionally, genes and proteins associated with inhibition of apoptosis were upregulated, including TRAIL-R3, a decoy receptor that opposes TRAIL-mediated death45; normalisation of TRAIL-R3 concentrations with anifrolumab were also observed in a previous study of patients from the phase 2 MUSE trial.13 The impact of anifrolumab on NETosis gene pathways shown here is also in agreement with another study with patients from the MUSE trial; markers of NETosis known as NET complexes (CitH3-DNA, MPO-DNA, NE-DNA) positively correlated with a 21-gene IFNGS, and anifrolumab treatment significantly reduced the levels of all three NET complexes after 1 year of treatment, whereas no reductions were observed in the placebo group.8 These changes in apoptosis and NETosis pathways suggest that anifrolumab treatment may improve dysregulated cell death processes that play a role in lupus pathology.
Our results suggest that IFNAR1 inhibition with anifrolumab also helps to reverse the changes in immune cell composition commonly observed in patients with SLE, particularly in the lymphoid compartment.2 Anifrolumab downmodulated multiple plasma cell and monocyte gene modules and upregulated lymphocyte and T-cell modules (analysed through transcriptome module enrichment analysis), reduced estimated levels of monocytes (analysed through RNA-seq deconvolution), and levels of proteins involved in the recruitment of monocytes. Specifically, anifrolumab downmodulated several molecules that contribute to monocyte and macrophage dysregulation in autoimmunity, including the SIGLEC1 gene and the CD163 and MCP-1 (CCL2) proteins. The expression of SIGLEC1 gene, an IFN-regulated cell-adhesion molecule exclusively expressed in blood monocytes, is increased in several autoimmune diseases.34 Findings from a single-centre flow cytometry study indicated that anifrolumab treatment suppresses intermediate monocyte numbers and SIGLEC1 gene expression, further supporting our findings.46 CD163 protein is a marker for alternatively activated M2 macrophages, and is overexpressed in the blood and skin of patients with SLE.47 48 MCP-1 is involved in the regulation and recruitment of monocytes and other immune cells to tissue; levels of MCP-1 protein were shown to correlate with renal involvement and disease activity in patients with SLE and,49 consistent with our results, can be reduced with anifrolumab treatment.13 Overall, the observed downregulation of these molecules supports the hypothesis that anifrolumab may restore monocyte function and immune cell homoeostasis in the blood of patients with SLE.
Other tissue-recruiting chemokines were also downmodulated by anifrolumab treatment, including the IFN-γ responsive chemokine CXCL10. CXCL10 is secreted by monocytes, fibroblasts and endothelial cells and is involved in inflammatory cell recruitment, particularly to the kidney.35 Serum CXCL10 protein is elevated in patients with SLE and has been shown to correlate with both disease activity and type I IFN pathway activity, further implicating a role in lupus pathogenesis.35 Reduction of CXCL10 and other chemokines, like the aforementioned MCP-1, following IFNAR1 blockade may contribute to the observed improvement in disease activity.
In addition to effects on innate immune cells and chemokines, we found that anifrolumab treatment increased the estimated blood numbers of MAIT cells and γδT cells compared with placebo. Blood MAIT cell counts are significantly lower in patients with SLE compared with healthy individuals.50 MAIT cells can be activated by IFN-α, and increased MAIT cell activation correlates with SLE disease activity.50 γδT cells share many transcriptional characteristics with MAIT cells,51 and evidence suggests that blood γδ T cells are decreased in patients with active SLE,52 which could potentially be restored to normal levels with anifrolumab treatment.
Anifrolumab treatment also downmodulated several molecules with key roles in B-cell and T-cell activation or recruitment, including gene and protein levels of BAFF, protein levels of CXCL13, LAG3 and galactin-9, and CD38 gene expression. BAFF is a B-cell activating factor typically elevated in patients with SLE.53 CXCL13, a B-cell chemoattractant, has been implicated in the development of lupus nephritis owing to its role in B cell kidney infiltration, and its levels correlate with disease activity in patients with SLE.54 Expression of LAG3, a CD4-related activation-induced molecule that negatively regulates the expansion of activated T cells and pDCs, increases on T-cell and pDC activation and is elevated in patients with SLE.38 55 Pathogenic LAG3+T cell counts positively associate with SLE disease activity.56 Galectin-9 is a ligand for CD40 on type 1 T helper cells that is typically overexpressed in patients with SLE.35 Serum galactin-9 was shown to correlate with IFNGS and measures of SLE disease activity.35 Many of the immune cells that contribute to SLE pathogenesis, including CD4+ and CD8+ memory T cells, may express the multifunctional cell-surface protein CD38, which is considered a potential therapeutic target in patients with SLE.57 In support of the potential for blocking CD38, treatment with anti-CD38 antibody, daratumumab, in two patients with refractory SLE, led to improved lupus serological biomarkers, reduced type I IFN signalling, and improved clinical outcomes assessed across multiple lupus manifestations.58 These findings, alongside the sustained increase in lymphocyte counts seen with anifrolumab treatment, support the downstream effects of IFNAR1 blockade on adaptive immune pathways and the normalisation of peripheral blood leucocyte subset composition with anifrolumab treatment.
Limitations of this study include those inherent to the deconvolution analysis, including the lack of convergence for some cell populations. No statistical tests were performed on values from deconvolution; these results should be further validated using flow cytometry. The correlation between RNA sequencing-based gene expression and plasma proteomics was not assessed to confirm biological validity, because plasma proteomics samples were available only from the TULIP-1 trial, and sample collection timings differed for the proteomics and transcriptomics analyses. Additionally, this study did not include healthy controls, and therefore, comparisons were limited to the available data. This study also did not describe results according to IFNGS status, nor according to race/ethnicity, given the small size of some subgroups; patients participating in this study were predominantly White, which limits the applicability of these findings to other races/ethnicities who may have a higher prevalence of high IFNGS scores. However, within the limitations of the population studied, the association between INFGS status and race with clinical response has been published previously.59 Furthermore, approximately half of patients were receiving glucocorticoids at baseline and mandatory taper attempts led to reduced glucocorticoid dosages throughout the trial in the anifrolumab versus placebo groups, which may have impacted gene expression.60 Finally, additional granularity for understanding the changes of the immune cell subpopulations in response to IFNAR1 blockade would have been achieved through the use of a single-cell RNA preparation, rather than with bulk RNA extracted from whole blood.
Taken together, our findings provide key insights into the immunomodulatory mechanisms underpinning IFNAR1 blockade with anifrolumab treatment, in the first transcriptomics and proteomics analyses performed on data from phase 3 trials in patients with SLE. Our findings support the broad anti-inflammatory effects downstream of IFNAR1 blockade with anifrolumab, including on pathways associated with apoptosis and NETosis, as well as multiple pathways affecting both innate and adaptive immunity, suggesting that IFNAR1 inhibition with anifrolumab may help reverse the imbalances in immune cell composition observed in patients with SLE.
Acknowledgments
The authors would like to thank the patients and healthcare providers who participated in this study. The authors would also like to thank the investigators and research staff who contributed to the analysis and interpretation of the data, in particular Karl Nordström, Manasa Surakala, Abirami Pandian, Priyabrata Panigrahi, Mridul Chaudhary for their contributions to the processing and transferring RNA-seq data, and Dr Deepak Rao for his contribution to the manuscript development. The views expressed in this publication are those of the author(s). Medical writing support was provided by Sofia Fernandes, PhD, of JK Associates, part of Avalere Health. This support was funded by AstraZeneca.
Footnotes
Handling editor: Josef S Smolen
@edvital
Correction notice: This article has been corrected since it published Online First. The first author statement has been added as well as amendments to the abstract and table 1.
Contributors: All authors were involved in drafting the article or revising it critically for important intellectual content, and all authors approved the final version to be published. Substantial contributions to the conception or design of study: MR, WIW, RT, EFM, RF, EMV, AP, HA-M and PZB; substantial contributions to data acquisition: TB, MR, WIW, RT, RAF and PZB; substantial contributions to data analysis and/or interpretation: TB, HS, PJN, PGG, MNL, WIW, RT, NF, DM, EFM, EMV, CC, AP, HA-M, PZB and EC; guarantor: EC.
Funding: This study was funded by AstraZeneca.
Competing interests: TB is an employee and stock holder of AstraZeneca; HS is an employee and stock holder of AstraZeneca; PJN was an employee of AstraZeneca at the time the study was being conducted and is an employee and stock holder of GSK; PGG is an employee and stock holder of AstraZeneca; MNL was an employee of AstraZeneca at the time the study was being conducted; MR was an employee and stock holder of AstraZeneca at the time the study was being conducted and is an employee and stock holder of GSK; WIW is an employee and stock holder of AstraZeneca; is applying for two patents (No. 17/999257; No. 18/556733); NF is an employee and stock holder of AstraZeneca; DM is an employee and stock holder of AstraZeneca; RT is an employee and stock holder of AstraZeneca; EFM received grants/contracts from AbbVie, Amgen, AstraZeneca, Biogen, BMS, Eli Lilly, EMD Serono, Genentech, GSK, Janssen, Novartis, Takeda and UCB; received consulting feed from AbbVie, AstraZeneca, Capella, Eli Lilly, EMD Serono, Galapagos, IGM, Novartis, Servier, Wolf, and Zenas; received payment/honoraria from AstraZeneca, BMS, GSK, and Roche; received support for meetings/travel from AstraZeneca and Roche; participated on Data Safety Monitoring/Advisory Boards of AstraZeneca, EMD Serono, Galapagos, Janssen, Novartis, and Takeda; is the Board Director of Rare Voices Australia and Exosome Biosciences Pty; RAF received grants/contracts, consulting fees, payment/honoraria, and support for attending meetings/travel from AstraZeneca and participated on a Data Safety Monitoring/Advisory Board of AstraZeneca; EMV received grants/contracts from AstraZeneca and Sandoz; consulting fees from AbbVie, AstraZeneca, CESAS, Elli Lilly, Novartis, Otsuka, Pfizer, Roche, UCB; payment/honoraria from AstraZeneca, Novastis and Otsuka; support for attending meetings/travel from Otsuka; participated on Data Safety Monitoring/Advisory Boards of Aurinia; is the General Secretary in SLEuro; CC is an employee and stock holder of AstraZeneca; AP is an employee and stockholder of AstraZeneca; HA-M was an employee and stock holder of AstraZeneca at the time the study was being conducted; PZB is an employee and stock holder of AstraZeneca; EC is an employee and stock holder of AstraZeneca.
Patient and public involvement: Patients and/or the public were not involved in the design, or conduct, or reporting, or dissemination of this research.
Provenance and peer review: Not commissioned; externally peer reviewed.
Supplemental material: This content has been supplied by the author(s). It has not been vetted by BMJ Publishing Group Limited (BMJ) and may not have been peer-reviewed. Any opinions or recommendations discussed are solely those of the author(s) and are not endorsed by BMJ. BMJ disclaims all liability and responsibility arising from any reliance placed on the content. Where the content includes any translated material, BMJ does not warrant the accuracy and reliability of the translations (including but not limited to local regulations, clinical guidelines, terminology, drug names and drug dosages), and is not responsible for any error and/or omissions arising from translation and adaptation or otherwise.
Data availability statement
Data underlying the findings described here may be obtained in accordance with AstraZeneca’s data sharing policy described at https://www.astrazenecaclinicaltrials.com/our-transparency-commitments/. Reuse is permitted only with permission from AstraZeneca.
Ethics statements
Patient consent for publication
All patients provided informed consent for the publication of results from TULIP-1 and TULIP-2 trials. Not applicable for this study.
Ethics approval
This study involves human participants and as this was a post hoc analysis of anonymised data, no ethics committee or institutional review board approvals were required; all such approvals were obtained in the original trials (Furie RA, 2019; Morand EF, 2020). This study was conducted in accordance with principles of the Declaration of Helsinki and the International Conference on Harmonisation Guidance for Good Clinical Practice. Participants gave informed consent to participate in the study before taking part.
References
- 1. Caielli S, Wan Z, Pascual V. Systemic lupus erythematosus pathogenesis: interferon and beyond. Annu Rev Immunol 2023;41:533–60. 10.1146/annurev-immunol-101921-042422 [DOI] [PubMed] [Google Scholar]
- 2. Gupta S, Kaplan MJ. Bite of the wolf: innate immune responses propagate autoimmunity in lupus. J Clin Invest 2021;131:e144918. 10.1172/JCI144918 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Ghodke-Puranik Y, Niewold TB. Genetics of the type I interferon pathway in systemic lupus erythematosus. Int J Clin Rheumatol 2013;8. 10.2217/ijr.13.58 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Brohawn PZ, Streicher K, Higgs BW, et al. Type I interferon gene signature test-low and -high patients with systemic lupus erythematosus have distinct gene expression signatures. Lupus 2019;28:1524–33. 10.1177/0961203319885447 [DOI] [PubMed] [Google Scholar]
- 5. Smith MA, Chiang C-C, Zerrouki K, et al. Using the circulating proteome to assess type I interferon activity in systemic lupus erythematosus. Sci Rep 2020;10:4462. 10.1038/s41598-020-60563-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Lindblom J, Toro-Domínguez D, Carnero-Montoro E, et al. Distinct gene dysregulation patterns herald precision medicine potentiality in systemic lupus erythematosus. J Autoimmun 2023;136:103025. 10.1016/j.jaut.2023.103025 [DOI] [PubMed] [Google Scholar]
- 7. Rodríguez-Carrio J, Burska A, Conaghan PG, et al. Association between type I interferon pathway activation and clinical outcomes in rheumatic and musculoskeletal diseases: a systematic literature review informing EULAR points to consider. RMD Open 2023;9:e002864. 10.1136/rmdopen-2022-002864 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Casey KA, Smith MA, Sinibaldi D, et al. Modulation of cardiometabolic disease markers by type I interferon inhibition in systemic lupus erythematosus. Arthritis Rheumatol 2021;73:459–71. 10.1002/art.41518 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. AstraZeneca . SAPHNELO. Prescribing Information FDA, 2021. [Google Scholar]
- 10. Furie R, Khamashta M, Merrill JT, et al. Anifrolumab, an anti–Interferon‐α receptor monoclonal antibody, in moderate‐to‐severe systemic lupus erythematosus. Arthritis & Rheumatology 2017;69:376–86. 10.1002/art.39962 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Furie RA, Morand EF, Bruce IN, et al. Type I interferon inhibitor anifrolumab in active systemic lupus erythematosus (TULIP-1): a randomised, controlled, phase 3 trial. Lancet Rheumatol 2019;1:e208–19. 10.1016/S2665-9913(19)30076-1 [DOI] [PubMed] [Google Scholar]
- 12. Morand EF, Furie R, Tanaka Y, et al. Trial of anifrolumab in active systemic lupus erythematosus. N Engl J Med 2020;382:211–21. 10.1056/NEJMoa1912196 [DOI] [PubMed] [Google Scholar]
- 13. Casey KA, Guo X, Smith MA, et al. Type I interferon receptor blockade with anifrolumab corrects innate and adaptive immune perturbations of SLE. Lupus Sci Med 2018;5:e000286. 10.1136/lupus-2018-000286 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Fanouriakis A, Tziolos N, Bertsias G, et al. Update on the diagnosis and management of systemic lupus erythematosus. Ann Rheum Dis 2021;80:14–25. 10.1136/annrheumdis-2020-218272 [DOI] [PubMed] [Google Scholar]
- 15. FastQC: A Quality Control Tool for High Throughput Sequence Data, Available: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ [Accessed 1 Sep 2023].
- 16. Okonechnikov K, Conesa A, García-Alcalde F. Qualimap 2: advanced multi-sample quality control for high-throughput sequencing data. Bioinformatics 2016;32:292–4. 10.1093/bioinformatics/btv566 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Li H, Handsaker B, Wysoker A, et al. The sequence alignment/map format and SAMtools. Bioinformatics 2009;25:2078–9. 10.1093/bioinformatics/btp352 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Dobin A, Davis CA, Schlesinger F, et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 2013;29:15–21. 10.1093/bioinformatics/bts635 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Ewels P, Magnusson M, Lundin S, et al. MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics 2016;32:3047–8. 10.1093/bioinformatics/btw354 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Gaspar JM. NGmerge: merging paired-end reads via novel empirically-derived models of sequencing errors. BMC Bioinformatics 2018;19:536. 10.1186/s12859-018-2579-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Patro R, Duggal G, Love MI, et al. Salmon provides fast and bias-aware Quantification of transcript expression. Nat Methods 2017;14:417–9. 10.1038/nmeth.4197 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Nextflow. Available: https://www.nextflow.io/ [Accessed 1 Sep 2023].
- 23. Grüning B, Dale R, Sjödin A, et al. Bioconda: sustainable and comprehensive software distribution for the life sciences. Nat Methods 2018;15:475–6. 10.1038/s41592-018-0046-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Coombes KR. ClassDiscovery: Classes and Methods for “Class Discovery” with Microarrays or Proteomics, Available: https://cran.r-project.org/web/packages/ClassDiscovery/index.html [Accessed 11 Jul 2023].
- 25. Hoffman GE, Roussos P. Dream: powerful differential expression analysis for repeated measures designs. Bioinformatics 2021;37:192–201. 10.1093/bioinformatics/btaa687 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Blighe K RS, Lewis M. EnhancedVolcano: Publication-ready volcano plots with enhanced colouring and labeling, Available: https://github.com/kevinblighe/EnhancedVolcano [Accessed 11 Jul 2023].
- 27. National Library of Medicine (US), National Center for Biotechnology Information. Available: https://www.ncbi.nlm.nih.gov/gene/ [Accessed 4 Dec 2023].
- 28. Wu T, Hu E, Xu S, et al. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation (Camb) 2021;2:100141. 10.1016/j.xinn.2021.100141 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Metacore - integrated pathway analysis for multi-Omics data. Available: https://portal.genego.com/ [Accessed 6 Dec 2023].
- 30. Altman MC, Rinchai D, Baldwin N, et al. Development of a fixed module repertoire for the analysis and interpretation of blood transcriptome data. Nat Commun 2021;12:4385. 10.1038/s41467-021-24584-w [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Monaco G, Lee B, Xu W, et al. RNA-Seq signatures normalized by mRNA abundance allow absolute deconvolution of human immune cell types. Cell Rep 2019;26:1627–40. 10.1016/j.celrep.2019.01.041 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Pfister S KV, Ferrero E. granulator: Rapid benchmarking of methods for *in silico* deconvolution of bulk RNA-seq data, Available: https://github.com/xanibas/granulator [Accessed 11 Jul 2023].
- 33. Perng Y-C, Lenschow DJ. ISG15 in antiviral immunity and beyond. Nat Rev Microbiol 2018;16:423–39. 10.1038/s41579-018-0020-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Oliveira JJ, Karrar S, Rainbow DB, et al. The plasma biomarker soluble SIGLEC-1 is associated with the type I interferon transcriptional signature, ethnic background and renal disease in systemic lupus erythematosus. Arthritis Res Ther 2018;20:152. 10.1186/s13075-018-1649-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Fasano S, Milone A, Nicoletti GF, et al. Precision medicine in systemic lupus erythematosus. Nat Rev Rheumatol 2023;19:331–42. 10.1038/s41584-023-00948-y [DOI] [PubMed] [Google Scholar]
- 36. Abel S, Hundhausen C, Mentlein R, et al. The transmembrane CXC-chemokine ligand 16 is induced by IFN-γ and TNF-α and shed by the activity of the disintegrin-like metalloproteinase ADAM10. J Immunol 2004;172:6362–72. 10.4049/jimmunol.172.10.6362 [DOI] [PubMed] [Google Scholar]
- 37. Mehta AK, Gracias DT, Croft M. TNF activity and T cells. Cytokine 2018;101:14–8. 10.1016/j.cyto.2016.08.003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Workman CJ, Wang Y, El Kasmi KC, et al. LAG-3 regulates plasmacytoid dendritic cell homeostasis. J Immunol 2009;182:1885–91. 10.4049/jimmunol.0800185 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Riggs JM, Hanna RN, Rajan B, et al. Characterisation of anifrolumab, a fully human anti-interferon receptor antagonist antibody for the treatment of systemic lupus erythematosus. Lupus Sci Med 2018;5:e000261. 10.1136/lupus-2018-000261 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Barrat FJ, Crow MK, Ivashkiv LB. Interferon target-gene expression and epigenomic signatures in health and disease. Nat Immunol 2019;20:1574–83. 10.1038/s41590-019-0466-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Goel RR, Wang X, O’Neil LJ, et al. Interferon lambda promotes immune dysregulation and tissue inflammation in TLR7-induced lupus. Proc Natl Acad Sci U S A 2020;117:5409–19. 10.1073/pnas.1916897117 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Psarras A, Alase A, Antanaviciute A, et al. Functionally impaired plasmacytoid dendritic cells and non-haematopoietic sources of type I interferon characterize human autoimmunity. Nat Commun 2020;11:6149. 10.1038/s41467-020-19918-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Moore S, Juo HH, Nielsen CT, et al. Role of neutrophil extracellular traps regarding patients at risk of increased disease activity and cardiovascular comorbidity in systemic lupus erythematosus. J Rheumatol 2020;47:1652–60. 10.3899/jrheum.190875 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Courtney PA, Crockard AD, Williamson K, et al. Increased apoptotic peripheral blood neutrophils in systemic lupus erythematosus: relations with disease activity, antibodies to double stranded DNA, and neutropenia. Ann Rheum Dis 1999;58:309–14. 10.1136/ard.58.5.309 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Baetu TM, Hiscott J. On the TRAIL to apoptosis. Cytokine Growth Factor Rev 2002;13:199–207. 10.1016/s1359-6101(02)00006-0 [DOI] [PubMed] [Google Scholar]
- 46. Carter LM, Wigston Z, Laws P, et al. Rapid efficacy of anifrolumab across multiple subtypes of recalcitrant cutaneous lupus erythematosus parallels changes in discrete subsets of blood transcriptomic and cellular biomarkers. Br J Dermatol 2023;189:210–8. 10.1093/bjd/ljad089 [DOI] [PubMed] [Google Scholar]
- 47. Nakayama W, Jinnin M, Makino K, et al. CD163 expression is increased in the involved skin and sera of patients with systemic lupus erythematosus. Eur J Dermatol 2012;22:512–7. 10.1684/ejd.2012.1756 [DOI] [PubMed] [Google Scholar]
- 48. Kwiecień I, Polubiec-Kownacka M, Dziedzic D, et al. CD163 and CCR7 as markers for macrophage polarization in lung cancer microenvironment. Cent Eur J Immunol 2019;44:395–402. 10.5114/ceji.2019.92795 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Živković V, Cvetković T, Mitić B, et al. Monocyte chemoattractant protein-1 as a marker of systemic lupus erythematosus: an observational study. Rheumatol Int 2018;38:1003–8. 10.1007/s00296-017-3888-x [DOI] [PubMed] [Google Scholar]
- 50. Chiba A, Tamura N, Yoshikiyo K, et al. Activation status of mucosal-associated invariant T cells reflects disease activity and pathology of systemic lupus erythematosus. Arthritis Res Ther 2017;19:58. 10.1186/s13075-017-1257-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Provine NM, Binder B, FitzPatrick MEB, et al. Unique and common features of innate-like human Vδ2(+) ΓδT cells and mucosal-associated invariant T cells. Front Immunol 2018;9:756. 10.3389/fimmu.2018.00756 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52. Paredes JL, Fernandez-Ruiz R, Niewold TB. T cells in systemic lupus erythematosus. Rheum Dis Clin North Am 2021;47:379–93. 10.1016/j.rdc.2021.04.005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Stohl W, Metyas S, Tan S-M, et al. B lymphocyte stimulator overexpression in patients with systemic lupus erythematosus: longitudinal observations. Arthritis Rheum 2003;48:3475–86. 10.1002/art.11354 [DOI] [PubMed] [Google Scholar]
- 54. Schiffer L, Worthmann K, Haller H, et al. CXCL13 as a new biomarker of systemic lupus erythematosus and lupus nephritis - from bench to bedside? Clin Exp Immunol 2015;179:85–9. 10.1111/cei.12439 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55. Yu S, Fujio K, Ishigaki K, et al. Increased concentration of serum soluble LAG3 in systemic lupus erythematosus. Arthritis Res Ther 2012;14:1–54. 10.1186/ar3617 22393579 [DOI] [Google Scholar]
- 56. Kato R, Sumitomo S, Tsuchida Y, et al. CD4+ CD25+ LAG3+ T cells with a feature of Th17 cells associated with systemic lupus erythematosus disease activity. Front Immunol 2019;10:1619. 10.3389/fimmu.2019.01619 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Burns M, Ostendorf L, Biesen R, et al. Dysregulated CD38 expression on peripheral blood immune cell subsets in SLE. Int J Mol Sci 2021;22:2424. 10.3390/ijms22052424 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. Ostendorf L, Burns M, Durek P, et al. Targeting CD38 with daratumumab in refractory systemic lupus erythematosus. N Engl J Med 2020;383:1149–55. 10.1056/NEJMoa2023325 [DOI] [PubMed] [Google Scholar]
- 59. Vital EM, Merrill JT, Morand EF, et al. Anifrolumab efficacy and safety by type I interferon gene signature and clinical subgroups in patients with SLE: post hoc analysis of pooled data from two phase III trials. Ann Rheum Dis 2022;81:951–61. 10.1136/annrheumdis-2021-221425 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60. Kino T, Chrousos GP. Chapter 3.2 - Glucocorticoid effects on gene expression. In: Steckler T, Kalin NH, Reul JMHM, eds. Techniques in the Behavioral and Neural Sciences. Elsevier, 2005: 295–311. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
ard-2023-225445supp001.pdf (5.2MB, pdf)
Data Availability Statement
Data underlying the findings described here may be obtained in accordance with AstraZeneca’s data sharing policy described at https://www.astrazenecaclinicaltrials.com/our-transparency-commitments/. Reuse is permitted only with permission from AstraZeneca.