Abstract
The mechanism of colistin resistance (Colr) in Acinetobacter baumannii was studied by selecting in vitro Colr derivatives of the multidrug-resistant A. baumannii isolate AB0057 and the drug-susceptible strain ATCC 17978, using escalating concentrations of colistin in liquid culture. DNA sequencing identified mutations in genes encoding the two-component system proteins PmrA and/or PmrB in each strain and in a Colr clinical isolate. A colistin-susceptible revertant of one Colr mutant strain, obtained following serial passage in the absence of colistin selection, carried a partial deletion of pmrB. Growth of AB0057 and ATCC 17978 at pH 5.5 increased the colistin MIC and conferred protection from killing by colistin in a 1-hour survival assay. Growth in ferric chloride [Fe(III)] conferred a small protective effect. Expression of pmrA was increased in Colr mutants, but not at a low pH, suggesting that additional regulatory factors remain to be discovered.
Among gram-negative pathogens that are reported as “multidrug resistant” (MDR), Acinetobacter baumannii is rapidly becoming a focus of significant attention (1, 7, 25, 32, 38, 39, 46, 51). In intensive care units, up to 30% of A. baumannii clinical isolates are resistant to at least three classes of antibiotics, often including fluoroquinolones and carbapenems (25).
The emergence of MDR gram-negative pathogens, including A. baumannii, has prompted increased reliance on the cationic peptide antibiotic colistin (12). Regrettably, increasing colistin use has led to the discovery of resistant strains (10, 11, 22, 26). For example, in a recent study, 12% of carbapenemase-producing Enterobacteriaceae were found to be colistin resistant (Colr) (6). Although still uncommon, A. baumannii isolates resistant to all available antimicrobial agents have been reported (26, 45) and are of enormous concern, given their potential to spread in the critical care environment.
Colistin and other polymyxins are cyclic cationic peptides produced by the soil bacterium Bacillus polymyxa that act by disrupting the negatively charged outer membranes of gram-negative bacteria (37, 50). The following three distinct mechanisms that give rise to colistin resistance are known: (i) specific modification of the lipid A component of the outer membrane lipopolysaccharide, resulting in a reduction of the net negative charge of the outer membrane; (ii) proteolytic cleavage of the drug; and (iii) activation of a broad-spectrum efflux pump (13, 14, 49). The mechanism of colistin resistance in Acinetobacter spp. is not yet known. Heteroresistance to colistin in A. baumannii has been described (17, 24), but it is uncertain whether the basis for this resistance is the presence of a genetically distinct population of cells or whether variation in the regulatory program among genetically identical cells may be sufficient for the expression of resistance.
In Salmonella enterica, the two-component signaling systems PmrAB and PhoPQ are involved in sensing environmental pH, Fe3+, and Mg2+ levels, leading to altered expression of a set of genes involved in lipid A modification (14, 43, 53). A small adapter protein, PmrD, serves as an interface between the two-component systems by stabilizing the activated form of PmrA in S. enterica (19), but other mechanisms of coordinated regulation are described for other species (52). Mutations causing constitutive activation of PmrA and PmrB are associated with colistin resistance (31, 33). Interestingly, the phoPQ and pmrD genes do not appear to be present in Acinetobacter spp., based on computational analysis of the genome sequences (2).
PmrA-regulated resistance to colistin in S. enterica and P. aeruginosa results from modification of lipid A with 4-deoxy-aminoarabinose (Ara4N) or phosphoethanolamine via activation of ugd, the pmrF (or pbgP) operon, and pmrC, which encode UDP-glucose dehydrogenase (the first step in Ara4N biosynthesis), Ara4N biosynthetic enzymes, and lipid A phosphoethanolamine transferase, respectively (8, 15, 21, 41, 48). The Ara4N biosynthesis and attachment genes are not present in A. baumannii or Neisseria meningitidis (36, 47). N. meningitidis is intrinsically resistant to polymyxins, demonstrating that Ara4N modification of lipid A is not required for resistance. Mutations in the pmrC ortholog lptA, encoding the lipid A phosphoethanolamine transferase, reduce colistin resistance in N. meningitidis, suggesting that this modification alone may be sufficient for conferring colistin resistance (49). Here we show that the PmrAB system is involved in regulating colistin resistance in A. baumannii by identification of mutations in resistant isolates that exhibit constitutive expression of pmrA.
MATERIALS AND METHODS
Strains of A. baumannii.
AB060 and AB0057 are colistin-susceptible MDR strains isolated from patients at Walter Reed Army Medical Center in 2004, and ATCC 17978 is a largely drug-susceptible strain isolated from a patient in the late 1950s in New York City (2, 18, 44). The clinical isolate ACCA152 is a Colr strain isolated from the respiratory tract of a patient at Cedars-Sinai Medical Center. Identification as A. baumannii was confirmed with API 20NE kits (bioMérieux, Durham, NC). Relevant features of each strain are given in Table 1.
TABLE 1.
Strains used in this studya
| Strain | Parent | Colistin susceptibility | pmr genotype | Reference |
|---|---|---|---|---|
| ATCC 17978* | NA | S | Wild type | 44 |
| AB0057* | NA | S | Wild type | 2, 18 |
| MAC101 | AB0057 | R | PmrA P102H, PmrB A262P | This study |
| MAC102 | AB0057 | R | PmrB A227V | This study |
| MAC103 | MAC101 | S | Same as MAC101 | This study |
| MAC201 | ATCC 17978 | R | PmrB T13N, PmrB P233S | This study |
| MAC203 | MAC201 | S | PmrB T13N, PmrB P233S, PmrB Δ411-444 | This study |
| ACCA152* | NA | R | PmrB P233T | |
| AB060* | NA | S | PmrB FS209 (frameshift) | 18 |
| ATCC 27853 | NA | S | ND | 28 |
All strains are A. baumannii, except for P. aeruginosa ATCC 27853. *, clinical isolate. NA, not applicable; R, resistant; S, susceptible.
Susceptibility testing.
MICs were determined by broth microdilution in cation-adjusted Mueller-Hinton broth (MHB) according to Clinical and Laboratory Standards Institute methods (34), using in-house trays for colistin (Sigma-Aldrich, St. Louis, MO) and commercial trays (Sensititre [Trek Diagnostics, Cleveland, OH] and MicroScan [Siemens Healthcare Diagnostics, West Sacramento, CA]) for other agents. MICs were determined for ampicillin, piperacillin-tazobactam, cefuroxime, cefotaxime, ceftriaxone, ceftazidime, cefepime, cefoxitin, meropenem, imipenem, levofloxacin, ciprofloxacin, gentamicin, amikacin, and tetracycline. Colistin MICs were also determined by Etest (bioMérieux, Durham, NC) on Mueller-Hinton agar plates (BBL, Sparks, MD). Pseudomonas aeruginosa ATCC 27853 was used for quality control on each day of testing. All assays used colistin sulfate (referred to as “colistin” hereafter; Sigma-Aldrich, Inc.), and concentrations are reported as the concentration of colistin base, after adjusting for potency based on information from the manufacturer.
Selection of Colr mutants.
Colr derivative clones were obtained by growth in liquid culture (lysogeny broth [LB]) with colistin at 1.0 μg/ml, which is above the MIC. Successive passage in increasing colistin concentrations demonstrated a high level of resistance in each derivative. Mutation detection was performed by PCR amplification of the entire pmrCAB operon, followed by DNA sequencing using primers distributed approximately every 500 bases; both strands were completely sequenced. Mutations were confirmed to be specific to the Colr derivatives by PCR and sequencing of selected regions of the parental isolates. Primer sequences are reported in Table 2.
TABLE 2.
Primer sequences used for pmrCAB sequencing and quantitative PCR
| Primer | Sequence |
|---|---|
| pmrBAC_L | GCGAGGAGCACATTTCCTAA |
| pmrBAC_R | TGTAGTCACTCACGATGCTGAA |
| pmrBAC_L2 | TTAAAGTTACATCTTGCTTTGCC |
| pmrBAC_L2c | GGCAAAGCAAGATGTAACTTTAA |
| pmrBAC_R2 | TCGATGAAATTCTAGATACTCAAATG |
| pmrBAC_L3 | CCCAAATATCGATAAACAGATCTTC |
| pmrBAC_R3 | TTGAAGCAGATCCGTCAAAG |
| pmrBAC_L4 | TGCACCCAAATTTAAACCATC |
| pmrBAC_R4 | CCGACTTGTGATACGAATGC |
| pmrBAC_L5 | TCATTTGGCTTAATACATGGTCTG |
| pmrBAC_R5 | GGATGATTTCACCGCAAAATAG |
| pmrBAC_R5c | CTATTTTGCGGTGAAATCATCC |
| pmrA_1 | GGTGTTGCTGCTCTTTGACG |
| pmrA_2 | GGTGGAATGGGTCAATAACG |
| 16S_rRNA_1 | CAGCTCGTGTCGTGAGATGT |
| 16S_rRNA_2 | CGTAAGGGCCATGATGACTT |
Survival analysis assays were performed by growth in LB to an optical density of ∼0.6, followed by dilution into medium containing colistin. After 1 hour at 37°C, cultures were diluted and plated on LB plates without colistin. The percentage of surviving cells was determined. Population analysis profiles were determined by plating dilutions of a mid-log-phase culture (optical density, ∼0.6) onto LB plates, with and without the addition of 7 μg/ml colistin; the percentage of Colr cells in the original culture was determined based on the ratio of the number of colonies on colistin plates to that on LB plates without colistin.
Quantitative PCR.
Total RNA was isolated from bacterial cells grown in LB to mid-log phase (A600 = 0.5). Colistin (7 μg/ml) or ferric chloride (1 mM) was added as appropriate. Bacterial RNA was stabilized using RNAprotect bacterial reagent (Qiagen) and purified using an RNeasy kit (Qiagen) and nuclease-free water. The RNA concentration was determined using a NanoDrop 1000 spectrophotometer (Thermo Scientific).
Before reverse transcription (RT), RNA samples were treated with Turbo DNase (Ambion) to digest genomic DNA, followed by treatment with RNaseOUT recombinant RNase inhibitor (Invitrogen) to prevent RNA degradation. The RT reaction was performed using Moloney murine leukemia virus reverse transcriptase (Invitrogen). Negative control reactions were performed using equal concentrations of RNA without RT reagents.
Primers for RT-PCR were designed using Primer3 and are listed in Table 2. Specificity was evaluated by melting curve analysis, and only artifact-free primers were used for RT-PCR. 16S rRNA was used as a housekeeping gene for normalization.
Real-time PCR amplification was carried out on a Chromo4 continuous fluorescence detector (MJ Research), using Power SYBR green PCR master mix (Applied Biosystems) as directed by the manufacturer. In each run, a blank sample (distilled water) and a no-reverse-transcriptase control were run to evaluate DNA contamination. The critical cycle threshold was determined by Opticon Monitor software, version 2.03 (MJ Research). Relative gene expression differences were calculated using the standard curve method. The quantity of the target transcript, pmrA, was determined from the standard curve of the 16S rRNA housekeeping gene, and the appropriate wild-type strain (AB0057 or ATCC 17978) was used as the calibrator sample to which differences were compared. For each sample, at least three biological replicates (from separate initial cultures) were performed, and the expression level in each replicate was measured a minimum of six times.
RESULTS
Survival analysis of strains with reference genome sequences.
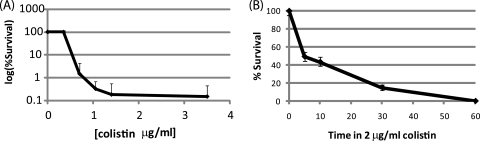
The origins and genotypes of the parent strains and derived mutants used in this study are shown in Table 1 and Fig. 1, and the colistin susceptibility of these isolates is shown in Table 3. We tested two clinical isolates of A. baumannii (AB0057 and ATCC 17978) for colistin susceptibility in rich media (LB and MHB). Using colistin in a survival analysis assay, the MDR isolate AB0057 was killed in a concentration- and time-dependent manner (Fig. 2). Fewer than 1% of cells survived a 1-hour incubation in 1.4 μg/ml colistin. Bactericidal activity was time dependent, with a half-life of approximately 5 minutes at 1.4 μg/ml colistin. A population analysis profile showed that within a log-phase culture, <0.00001% of AB0057 cells were able to grow on LB agar containing 1.4 μg/ml colistin. Similar results were obtained with A. baumannii ATCC 17978 (data not shown).
FIG. 1.
Selection of Colr and Cols mutants. The scheme for selection or isolation of Colr mutants and revertants of each mutant is shown. Colr strains are indicated by bold circles.
TABLE 3.
Antimicrobial susceptibilities of parent and mutant strains of A. baumannii
| Isolate | Colistin MIC (μg/ml)
|
|
|---|---|---|
| Etesta | Broth microdilution | |
| AB0057 | 1.5 | 1-2 |
| MAC101 | 32 | >128 |
| MAC102 | 4-8 | 64 |
| MAC103 | 0.5 | 1 |
| ATCC 17978 | 0.75 | 2 |
| MAC201 | 64 | 128 |
| MAC203 | 0.5 | 2 |
| ACCA152 | 24 | >128 |
| AB060 | 0.5 | 1 |
For Etest, susceptible MICs were ≤2 μg/ml and resistant MICs were ≥4 μg/ml.
FIG. 2.
Survival of strain AB0057 in colistin. (A) Mid-log-phase cultures of AB0057 in LB were treated with the indicated concentration of colistin for 1 h at 37°C, and then quantitative counts of live cells were determined by plating 0.1-ml samples of serial 10-fold dilutions on LB agar plates. (B) Cells were grown in LB for 4 hours, and then colistin was added at the indicated concentration. Aliquots were removed at the given intervals, and quantitative counts were performed as described above. “Percent survival” corresponds to 100 × the ratio of the number of CFU in the presence of colistin to the number of CFU in the absence of colistin at each time/concentration point.
Selection of Colr derivatives of isolates with completely sequenced genomes.
In order to define genetic loci responsible for colistin resistance, we selected Colr derivatives of A. baumannii isolates AB0057 and ATCC 17978, for which complete genome sequences are available (2, 44). Overnight cultures were diluted 1:100 in LB containing 0, 0.1, 0.5, 1.0, 2.0, and 4.0 μg/ml colistin. The cells in cultures containing 0, 0.1, and 0.5 μg/ml colistin (i.e., below the MIC) demonstrated robust growth, while the cells in cultures containing ≥1 μg/ml colistin did not grow. Cultures containing higher concentrations of colistin were left to grow for 18 to 24 h. At 24 h, the cultures with 1.0 μg/ml colistin demonstrated growth. These cultures were plated onto LB agar containing 7 μg/ml colistin, and individual colonies were propagated for further analysis. These cells were confirmed to be A. baumannii by PCR, using several primer pairs that had been used for gap closure during genome sequencing (2), and by biochemical identification on API 20NE panels. Two independent Colr derivatives of AB0057 (MAC101 and MAC102) and one of ATCC 17978 (MAC201) were obtained.
Colr derivative strains were tested for resistance to increasing concentrations of colistin and were able to grow in up to 64 μg/ml colistin in liquid culture. Colistin MICs determined by Etest for the Colr derivative strains ranged from 8 to 64 μg/ml (Table 3). Cells were routinely grown in and maintained on LB agar or agar containing 7 μg/ml colistin. The antibiotic susceptibility profiles of the parental and Colr derivative strains showed no changes for any of the agents tested other than colistin (data not shown). In contrast to the results of other studies (23), acquisition of colistin resistance did not increase susceptibility to other antibiotics.
The proportion of resistant cells declined when Colr strains were grown in the absence of colistin. Colr strains were passaged in LB every 12 hours. By the ninth passage, only about 1% of cells were able to grow on plates containing 7 μg/ml colistin (Fig. 3). Thus, Colr strains exhibit a modest growth disadvantage compared with their respective Cols parents. A single colony from an LB plate at the ninth passage was isolated and maintained as an example of a revertant for each parental strain (MAC103 and MAC203). These revertant clones were Cols (Table 3).
FIG. 3.
Reversion of colistin resistance. Each clone was passaged in successive overnight and daily cultures. Each daily culture was plated on LB agar with and without 10 μg/ml colistin. The ratio of CFU from the two plates is plotted.
Mutational analysis of Colr derivative strains and revertants.
The two-component system proteins PmrB (sensor kinase) and PmrA (response regulator) are implicated in polymyxin resistance in several genera (31, 42, 52). The A. baumannii orthologs of these genes are not obvious based solely on pairwise alignments with the P. aeruginosa PmrB and PmrA sequences. AB57_3174 is a clear ortholog of PmrC, however, strongly implicating the adjacent genes, encoding the two-component system proteins AB57_3173 and AB57_3172, as PmrA and PmrB, respectively. The ATCC 17978 proteins encoded by these genes are A1S_2752, A1S_2751, and A1S_2750, respectively.
In order to understand the genetic basis for this phenotype, the complete coding region and promoter of the pmrCAB operon were sequenced for MAC101, MAC102, and MAC201. Mutations in pmrB and pmrA that are not present in the parental isolates were found in the derivative strains (Fig. 4). MAC101 has mutations in both pmrA and pmrB, and MAC201 has two point mutations in pmrB. In each case, at least one of the mutations falls within a conserved functional domain. Supporting the importance of the pmrAB system in clinically relevant resistance to colistin, the Colr A. baumannii isolate ACCA152 carries the pmrB mutation P233T. A mutation at the same codon was found in MAC201.
FIG. 4.
Mutational analysis of the pmr operon. Mutations in bold affect conserved residues in functional domains. All strains are Colr, except for AB060 and MAC203, which are Cols.
Sequencing of the pmr operon in MAC203 revealed a deletion in pmrB that removes the C-terminal 32 amino acids, including a portion of the histidine kinase domain. The sequence of the pmrCAB region in the revertant MAC103 is 100% identical to the MAC101 sequence, so the suppressing mutation maps outside the pmrCAB locus.
Expression analysis of pmrA.
The identification of several independent mutations in pmrB and pmrA demonstrates the importance of these genes as regulators of colistin resistance in A. baumannii. Based on prior analysis of PmrA function, we hypothesized that these mutations result in activation of PmrA (33). To evaluate the effects of these mutations in the Colr strains, quantitative RT-PCR was performed using primers designed to target the pmrA transcript. The Colr strains (MAC101, MAC102, and MAC201) exhibited 5- to 40-fold increased expression of pmrA (Fig. 5A). This is consistent with activating mutations and PmrA auto-regulation (27). Growth in the presence of colistin was not necessary for high-level pmrA expression (data not shown). In contrast, expression levels in the ΔpmrB revertant strain MAC203 were similar to those in the wild-type parental strain. Interestingly, pmrA expression in the Cols revertant MAC103 was also at approximately baseline levels, despite the observation that the pmr locus is identical in sequence to that in MAC101. Further analysis will be necessary to identify the mutation in this strain and to explain the reduction in pmr expression.
FIG. 5.
Gene expression analysis of pmrA. The level of expression of pmrA was measured by real-time quantitative RT-PCR. Expression levels were normalized to 16S rRNA. (A) Expression analysis of wild-type and mutant strains. Changes with respect to wild-type expression levels are shown. (B) Expression analysis at pH 5.5 and 7.7. Changes are expressed with respect to expression levels measured at pH 7.7. (C) Expression analysis in the presence of 1 mM FeCl3. Changes are expressed with respect to expression levels of the wild-type strain grown without iron. Each value represents the mean ± standard deviation for at least three independent cultures. An asterisk indicates that the difference is significant (P < 0.01) by the t test.
Induction of colistin resistance by environmental factors.
Growth of the two wild-type A. baumannii strains at pH 5.5 resulted in a remarkable increase in the MIC for colistin, to ≥64 μg/ml (Table 4). The MIC was significantly attenuated for the pmrB deletion mutant MAC203, suggesting that PmrB is required for the induction of colistin resistance, as seen with S. enterica (40). A second clinical isolate (AB060), harboring a frameshift mutation at codon 209 of PmrB, was also unable to grow in colistin-containing medium at pH 5.5. Thus, PmrB seems to be required for acid pH-induced colistin resistance. Contrary to expectations, the level of expression of pmrA was unchanged at pH 5.5 compared with that at pH 7.7 (Fig. 5B). The broth microdilution assay measures the effect of chronic exposure of cells to colistin. We also performed a 1-hour survival analysis assay, as described above, to assess the short-term effect of colistin on cells grown at acid pH. In this assay, MAC203 exhibited a level of protection from the bactericidal activity of colistin that was equivalent to that of both wild-type strains (Table 4).
TABLE 4.
Effects of acid pH on colistin susceptibility
| Strain | Broth microdilution colistin MIC (μg/ml) (MHB)
|
% Survival following 1-h colistin challenge (LB)a
|
||
|---|---|---|---|---|
| pH 5.5 | pH 7.7 | pH 5.5 | pH 7.7 | |
| AB0057 | 64 | 1 | 70 ± 11 | <1 |
| ATCC 17978 | 64 | 1-2 | 75 ± 26 | <1 |
| MAC103 | 128 | 1 | ND | ND |
| MAC203 | 4 | 1 | 68 ± 52 | <1 |
| AB060 | 1 | 0.25 | ND | ND |
| P. aeruginosa ATCC 27853 | 1 | 2 | ND | ND |
Values are means for at least two independent cultures ± standard deviations. ND, not done.
The addition of 1 mM ferric chloride [iron(III)] increased the MIC for colistin ∼2- to 4-fold (Table 5). Ferric chloride precipitates in MHB at this concentration, so assays were performed in LB. AB0057 and ATCC 17978 cells grown in LB in the presence of 1 mM FeCl3 were able to resist lysis by colistin during a 1-hour survival assay better than cells grown without iron (Table 5), although the effect was variable. Fe3+-induced colistin resistance is rapid: pretreatment of cells for 1 hour resulted in essentially the same level of resistance as that observed in cells grown overnight in LB supplemented with 1 mM FeCl3 (data not shown). pmrA expression was unaffected by Fe3+ exposure (Fig. 5C). MAC203 cells, carrying a partial deletion of pmrB, exhibited protection from colistin in the presence of ferric chloride equivalent to that of the wild-type parent. In addition, the Colr strain MAC201 showed an increase in colistin resistance in the presence of iron. Taken together, these observations suggest that the Pmr system may not be the only contributor to regulation of colistin resistance.
TABLE 5.
Effects of Fe3+ on colistin susceptibility in LB
| Strain | Broth microdilution colistin MIC (μg/ml)
|
% Survival following 1-h colistin challengea
|
||
|---|---|---|---|---|
| 0 mM FeCl3 | 1 mM FeCl3 | 0 mM FeCl3 | 1 mM FeCl3 | |
| AB0057 | 0.25 | 0.5 | 1.6 ± 0.2 | 16 ± 8 |
| ATCC 17978 | 0.25 | 0.5 | 0.2 ± 0.4 | 56 ± 65 |
| MAC203 | 0.5 | 4 | 0.1 ± 0.05 | 15 ± 10 |
| MAC201 | 16 | 128 | ND | ND |
| P. aeruginosa ATCC 27853 | 0.25 | 0.25 | ND | ND |
Values are means for at least two independent cultures ± standard deviations. ND, not done.
DISCUSSION
We show that mutations in the genes encoding the two-component signaling proteins PmrB and PmrA are linked to colistin resistance in A. baumannii. In the Colr mutants, expression of pmrA is increased, suggesting that the mutations cause constitutive activation of PmrA, which in turn autoregulates the pmrCAB promoter. A partial deletion of pmrB in a Colr background results in reversion to a colistin-susceptible phenotype, reinforcing the importance of the PmrAB system in regulating colistin resistance in A. baumannii. In addition, colistin resistance can be induced by acid pH and, to a lesser extent, Fe3+ supplementation. This induction is rapid. Thus, it is likely to involve solely changes in the cellular regulatory program rather than selection of rare spontaneous mutations. Acid pH- and Fe3+-induced resistance appears to be at least partially independent of PmrA. Expression of pmrA is unchanged during growth at acid pH or in ferric chloride. The ΔpmrB strains MAC203 and AB060 exhibit markedly reduced pH-induced resistance but no difference in Fe3+-induced resistance. It is possible that PmrB interacts with other response regulator proteins (3). This would explain both the lack of pmrA upregulation and the ability of ΔpmrB cells to respond to Fe3+ treatment.
The downstream targets of PmrA that are responsible for colistin resistance still remain to be determined. Autoregulation of the pmrCAB operon is observed in S. enterica and P. aeruginosa, suggesting that increased PmrC-mediated transfer of phosphoethanolamine to lipid A may be involved. A recent study of proteins that are differentially expressed in a Colr derivative of A. baumannii ATCC 19606 found ∼35 protein expression changes but not PmrC (9). The pmr operon was not sequenced for this mutant, and it remains to be determined whether these proteins may be encoded by PmrA target genes. Knowledge of the mechanism(s) of resistance in A. baumannii will be informative for consideration of the development of strategies to prevent and combat it. For example, inhibition of Ara4N biosynthesis and/or attachment to lipid A has been proposed as a means of disabling polymyxin resistance (20), but this approach would likely be completely ineffective for Acinetobacter spp., which do not contain the target enzymes.
The regulation of colistin resistance observed in A. baumannii differs in substantive ways from that in P. aeruginosa, the closest relative that has been studied in detail. Unlike the case in A. baumannii, there is no evidence of acid pH- or Fe3+-induced colistin resistance in P. aeruginosa, and this was confirmed by the use of a P. aeruginosa control strain in the analyses presented here. The role of a limiting Mg2+ concentration in the regulation of colistin resistance in A. baumannii has not yet been evaluated. Regulation of colistin resistance has been studied most extensively in S. enterica, where low magnesium, high iron, and low pH can induce resistance, mediated by the PhoPQ (Mg2+) and PmrAB (Fe3+ and pH) two-component systems (16, 43). Fe3+-mediated changes in lipid A are PmrA dependent and are the same as those associated with colistin resistance (35). Thus, both S. enterica and P. aeruginosa exhibit regulation of colistin resistance in response to environmental factors, although in different ways (27).
In addition to regulating resistance to cationic antimicrobial peptides, the PmrAB two-component system is linked to the control of genes associated with virulence in S. enterica and Legionella pneumophila (3, 14). The role, if any, of the A. baumannii PmrAB system in other aspects of its pathology remains to be determined. The Colr isolates exhibited differences in protein expression compared to their Cols progenitors (data not shown), and identification of these differences may prove useful for dissecting the genetic and biochemical features of colistin resistance.
Heteroresistance to colistin was previously observed in A. baumannii, with the percentage of Colr cells ranging from 0.00001% to 0.0002% (17, 24). Heteroresistance has been defined as “the presence of one or several bacterial subpopulations at a frequency of 10−7 to 10−3, which can grow at higher antibiotic concentrations than predicted by the MIC for the majority of cells” (30). These subpopulations may reflect genetic heterogeneity in a mixed population or genetically identical cells that express different gene sets in response to divergent regulatory programs (4, 5, 29). The clinical significance of colistin heteroresistance in A. baumannii is not yet known, but it is quite worrisome because (i) it is possible that selection of resistant strains could lead to treatment failure and (ii) the inadvertent transmission of Colr strains of Acinetobacter spp. could have a significant impact by necessitating additional isolation and decontamination regimens. We have shown that growth conditions can modulate the MIC and affect the percentage of cells that are able to survive a 1-hour colistin challenge. Stable colistin resistance was correlated with the presence of genetic mutations, but high-level resistance could also be induced by growth at pH 5.5.
If colistin resistance can readily arise based on regulatory changes in response to environmental factors or selection of a rare mutation in the presence of colistin, the phenotype of colistin susceptibility will be subject to significant changes over the course of an infection. Rapid determination of the presence of a subpopulation of Colr cells within a sample from an infected site, for example, by enzyme-linked immunosorbent assay or genetic assay, could assist in treatment decisions and infection control.
Acknowledgments
We thank Joseph Schinaman for technical assistance with survival assays.
R.A.B. was supported by grants from the National Institutes of Health (RO1 AI072219), the Veterans Affairs Merit Review Program, and Geriatric Research, Education and Clinical Care VISN 10. M.D.A. was supported by startup funds from CWRU.
Footnotes
Published ahead of print on 15 June 2009.
REFERENCES
- 1.Abbott, A. 2005. Medics braced for fresh superbug. Nature 436:758. [DOI] [PubMed] [Google Scholar]
- 2.Adams, M. D., K. Goglin, N. Molyneaux, K. M. Hujer, H. Lavender, J. J. Jamison, I. J. MacDonald, K. M. Martin, T. Russo, A. A. Campagnari, A. M. Hujer, R. A. Bonomo, and S. R. Gill. 2008. Comparative genome sequence analysis of multidrug-resistant Acinetobacter baumannii. J. Bacteriol. 190:8053-8064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Al-Khodor, S., S. Kalachikov, I. Morozova, C. T. Price, and Y. Abu Kwaik. 2009. The PmrA/PmrB two-component system of Legionella pneumophila is a global regulator required for intracellular replication within macrophages and protozoa. Infect. Immun. 77:374-386. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Avery, S. V. 2006. Microbial cell individuality and the underlying sources of heterogeneity. Nat. Rev. Microbiol. 4:577-587. [DOI] [PubMed] [Google Scholar]
- 5.Booth, I. R. 2002. Stress and the single cell: intrapopulation diversity is a mechanism to ensure survival upon exposure to stress. Int. J. Food Microbiol. 78:19-30. [DOI] [PubMed] [Google Scholar]
- 6.Castanheira, M., H. S. Sader, L. M. Deshpande, T. R. Fritsche, and R. N. Jones. 2008. Antimicrobial activities of tigecycline and other broad-spectrum antimicrobials tested against serine carbapenemase- and metallo-beta-lactamase-producing Enterobacteriaceae: report from the SENTRY Antimicrobial Surveillance Program. Antimicrob. Agents Chemother. 52:570-573. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Dijkshoorn, L., A. Nemec, and H. Seifert. 2007. An increasing threat in hospitals: multidrug-resistant Acinetobacter baumannii. Nat. Rev. Microbiol. 5:939-951. [DOI] [PubMed] [Google Scholar]
- 8.Ernst, R. K., E. C. Yi, L. Guo, K. B. Lim, J. L. Burns, M. Hackett, and S. I. Miller. 1999. Specific lipopolysaccharide found in cystic fibrosis airway Pseudomonas aeruginosa. Science 286:1561-1565. [DOI] [PubMed] [Google Scholar]
- 9.Fernandez-Reyes, M., M. Rodriguez-Falcon, C. Chiva, J. Pachon, D. Andreu, and L. Rivas. 2009. The cost of resistance to colistin in Acinetobacter baumannii: a proteomic perspective. Proteomics 9:1632-1645. [DOI] [PubMed] [Google Scholar]
- 10.Gales, A. C., R. N. Jones, and H. S. Sader. 2006. Global assessment of the antimicrobial activity of polymyxin B against 54 731 clinical isolates of gram-negative bacilli: report from the SENTRY antimicrobial surveillance programme (2001-2004). Clin. Microbiol. Infect. 12:315-321. [DOI] [PubMed] [Google Scholar]
- 11.Gilad, J., S. Eskira, K. Riesenberg, F. Schlaeffer, E. Hyam, and A. Borer. 2005. Emergence of nosocomial colistin-resistant Acinetobacter baumannii, abstr. K-1292, p. 333. Abstr. 45th Annu. Intersci. Conf. Antimicrob. Agents Chemother. American Society for Microbiology, Washington, DC.
- 12.Go, E. S., C. Urban, J. Burns, B. Kreiswirth, W. Eisner, N. Mariano, K. Mosinka-Snipas, and J. J. Rahal. 1994. Clinical and molecular epidemiology of acinetobacter infections sensitive only to polymyxin B and sulbactam. Lancet 344:1329-1332. [DOI] [PubMed] [Google Scholar]
- 13.Guina, T., E. C. Yi, H. Wang, M. Hackett, and S. I. Miller. 2000. A PhoP-regulated outer membrane protease of Salmonella enterica serovar Typhimurium promotes resistance to alpha-helical antimicrobial peptides. J. Bacteriol. 182:4077-4086. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Gunn, J. S. 2008. The Salmonella PmrAB regulon: lipopolysaccharide modifications, antimicrobial peptide resistance and more. Trends Microbiol. 16:284-290. [DOI] [PubMed] [Google Scholar]
- 15.Gunn, J. S., K. B. Lim, J. Krueger, K. Kim, L. Guo, M. Hackett, and S. I. Miller. 1998. PmrA-PmrB-regulated genes necessary for 4-aminoarabinose lipid A modification and polymyxin resistance. Mol. Microbiol. 27:1171-1182. [DOI] [PubMed] [Google Scholar]
- 16.Gunn, J. S., S. S. Ryan, J. C. Van Velkinburgh, R. K. Ernst, and S. I. Miller. 2000. Genetic and functional analysis of a PmrA-PmrB-regulated locus necessary for lipopolysaccharide modification, antimicrobial peptide resistance, and oral virulence of Salmonella enterica serovar Typhimurium. Infect. Immun. 68:6139-6146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Hawley, J. S., C. K. Murray, and J. H. Jorgensen. 2008. Colistin heteroresistance in acinetobacter and its association with previous colistin therapy. Antimicrob. Agents Chemother. 52:351-352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Hujer, K. M., A. M. Hujer, E. A. Hulten, S. Bajaksouzian, J. M. Adams, C. J. Donskey, D. J. Ecker, C. Massire, M. W. Eshoo, R. Sampath, J. M. Thomson, P. N. Rather, D. W. Craft, J. T. Fishbain, A. J. Ewell, M. R. Jacobs, D. L. Paterson, and R. A. Bonomo. 2006. Analysis of antibiotic resistance genes in multidrug-resistant Acinetobacter sp. isolates from military and civilian patients treated at the Walter Reed Army Medical Center. Antimicrob. Agents Chemother. 50:4114-4123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Kato, A., A. Y. Mitrophanov, and E. A. Groisman. 2007. A connector of two-component regulatory systems promotes signal amplification and persistence of expression. Proc. Natl. Acad. Sci. USA 104:12063-12068. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Kline, T., M. S. Trent, C. M. Stead, M. S. Lee, M. C. Sousa, H. B. Felise, H. V. Nguyen, and S. I. Miller. 2008. Synthesis of and evaluation of lipid A modification by 4-substituted 4-deoxy arabinose analogs as potential inhibitors of bacterial polymyxin resistance. Bioorg. Med. Chem. Lett. 18:1507-1510. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Lee, H., F. F. Hsu, J. Turk, and E. A. Groisman. 2004. The PmrA-regulated pmrC gene mediates phosphoethanolamine modification of lipid A and polymyxin resistance in Salmonella enterica. J. Bacteriol. 186:4124-4133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Li, J., and R. L. Nation. 2006. Old polymyxins are back: is resistance close? Clin. Infect. Dis. 43:663-664. [DOI] [PubMed] [Google Scholar]
- 23.Li, J., R. L. Nation, R. J. Owen, S. Wong, D. Spelman, and C. Franklin. 2007. Antibiograms of multidrug-resistant clinical Acinetobacter baumannii: promising therapeutic options for treatment of infection with colistin-resistant strains. Clin. Infect. Dis. 45:594-598. [DOI] [PubMed] [Google Scholar]
- 24.Li, J., C. R. Rayner, R. L. Nation, R. J. Owen, D. Spelman, K. E. Tan, and L. Liolios. 2006. Heteroresistance to colistin in multidrug-resistant Acinetobacter baumannii. Antimicrob. Agents Chemother. 50:2946-2950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Lockhart, S. R., M. A. Abramson, S. E. Beekmann, G. Gallagher, S. Riedel, D. J. Diekema, J. P. Quinn, and G. V. Doern. 2007. Antimicrobial resistance among gram-negative bacilli causing infections in intensive care unit patients in the United States between 1993 and 2004. J. Clin. Microbiol. 45:3352-3359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Matthaiou, D. K., A. Michalopoulos, P. I. Rafailidis, D. E. Karageorgopoulos, V. Papaioannou, G. Ntani, G. Samonis, and M. E. Falagas. 2008. Risk factors associated with the isolation of colistin-resistant gram-negative bacteria: a matched case-control study. Crit. Care Med. 36:807-811. [DOI] [PubMed] [Google Scholar]
- 27.McPhee, J. B., S. Lewenza, and R. E. Hancock. 2003. Cationic antimicrobial peptides activate a two-component regulatory system, PmrA-PmrB, that regulates resistance to polymyxin B and cationic antimicrobial peptides in Pseudomonas aeruginosa. Mol. Microbiol. 50:205-217. [DOI] [PubMed] [Google Scholar]
- 28.Medeiros, A. A., T. F. O'Brien, W. E. Wacker, and N. F. Yulug. 1971. Effect of salt concentration on the apparent in-vitro susceptibility of Pseudomonas and other gram-negative bacilli to gentamicin. J. Infect. Dis. 124(Suppl.):S59-S64. [DOI] [PubMed] [Google Scholar]
- 29.Megerle, J. A., G. Fritz, U. Gerland, K. Jung, and J. O. Radler. 2008. Timing and dynamics of single cell gene expression in the arabinose utilization system. Biophys. J. 95:2103-2115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Morand, B., and K. Muhlemann. 2007. Heteroresistance to penicillin in Streptococcus pneumoniae. Proc. Natl. Acad. Sci. USA 104:14098-14103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Moskowitz, S. M., R. K. Ernst, and S. I. Miller. 2004. PmrAB, a two-component regulatory system of Pseudomonas aeruginosa that modulates resistance to cationic antimicrobial peptides and addition of aminoarabinose to lipid A. J. Bacteriol. 186:575-579. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Munoz-Price, L. S., and R. A. Weinstein. 2008. Acinetobacter infection. N. Engl. J. Med. 358:1271-1281. [DOI] [PubMed] [Google Scholar]
- 33.Murray, S. R., R. K. Ernst, D. Bermudes, S. I. Miller, and K. B. Low. 2007. pmrA(Con) confers pmrHFIJKL-dependent EGTA and polymyxin resistance on msbB Salmonella by decorating lipid A with phosphoethanolamine. J. Bacteriol. 189:5161-5169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.National Committee for Clinical Laboratory Standards. 2004. Performance standards for antimicrobial susceptibility testing. M100-S14. Fourteenth informational supplement. NCCLS, Wayne, PA.
- 35.Nishino, K., F. F. Hsu, J. Turk, M. J. Cromie, M. M. Wosten, and E. A. Groisman. 2006. Identification of the lipopolysaccharide modifications controlled by the Salmonella PmrA/PmrB system mediating resistance to Fe(III) and Al(III). Mol. Microbiol. 61:645-654. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Parkhill, J., M. Achtman, K. D. James, S. D. Bentley, C. Churcher, S. R. Klee, G. Morelli, D. Basham, D. Brown, T. Chillingworth, R. M. Davies, P. Davis, K. Devlin, T. Feltwell, N. Hamlin, S. Holroyd, K. Jagels, S. Leather, S. Moule, K. Mungall, M. A. Quail, M. A. Rajandream, K. M. Rutherford, M. Simmonds, J. Skelton, S. Whitehead, B. G. Spratt, and B. G. Barrell. 2000. Complete DNA sequence of a serogroup A strain of Neisseria meningitidis Z2491. Nature 404:502-506. [DOI] [PubMed] [Google Scholar]
- 37.Paulus, H., and E. Gray. 1964. The biosynthesis of polymyxin B by growing cultures of bacillus polymyxa. J. Biol. Chem. 239:865-871. [PubMed] [Google Scholar]
- 38.Peleg, A. Y., H. Seifert, and D. L. Paterson. 2008. Acinetobacter baumannii: emergence of a successful pathogen. Clin. Microbiol. Rev. 21:538-582. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Perez, F., A. M. Hujer, K. M. Hujer, B. K. Decker, P. N. Rather, and R. A. Bonomo. 2007. Global challenge of multidrug-resistant Acinetobacter baumannii. Antimicrob. Agents Chemother. 51:3471-3484. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Perez, J. C., and E. A. Groisman. 2007. Acid pH activation of the PmrA/PmrB two-component regulatory system of Salmonella enterica. Mol. Microbiol. 63:283-293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Raetz, C. R., and C. Whitfield. 2002. Lipopolysaccharide endotoxins. Annu. Rev. Biochem. 71:635-700. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Roland, K. L., L. E. Martin, C. R. Esther, and J. K. Spitznagel. 1993. Spontaneous pmrA mutants of Salmonella typhimurium LT2 define a new two-component regulatory system with a possible role in virulence. J. Bacteriol. 175:4154-4164. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Shi, Y., M. J. Cromie, F. F. Hsu, J. Turk, and E. A. Groisman. 2004. PhoP-regulated Salmonella resistance to the antimicrobial peptides magainin 2 and polymyxin B. Mol. Microbiol. 53:229-241. [DOI] [PubMed] [Google Scholar]
- 44.Smith, M. G., T. A. Gianoulis, S. Pukatzki, J. J. Mekalanos, L. N. Ornston, M. Gerstein, and M. Snyder. 2007. New insights into Acinetobacter baumannii pathogenesis revealed by high-density pyrosequencing and transposon mutagenesis. Genes Dev. 21:601-614. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Taccone, F. S., H. Rodriguez-Villalobos, D. De Backer, V. De Moor, J. Deviere, J. L. Vincent, and F. Jacobs. 2006. Successful treatment of septic shock due to pan-resistant Acinetobacter baumannii using combined antimicrobial therapy including tigecycline. Eur. J. Clin. Microbiol. Infect. Dis. 25:257-260. [DOI] [PubMed] [Google Scholar]
- 46.Talbot, G. H., J. Bradley, J. E. Edwards, Jr., D. Gilbert, M. Scheld, and J. G. Bartlett. 2006. Bad bugs need drugs: an update on the development pipeline from the Antimicrobial Availability Task Force of the Infectious Diseases Society of America. Clin. Infect. Dis. 42:657-668. [DOI] [PubMed] [Google Scholar]
- 47.Tettelin, H., N. J. Saunders, J. Heidelberg, A. C. Jeffries, K. E. Nelson, J. A. Eisen, K. A. Ketchum, D. W. Hood, J. F. Peden, R. J. Dodson, W. C. Nelson, M. L. Gwinn, R. DeBoy, J. D. Peterson, E. K. Hickey, D. H. Haft, S. L. Salzberg, O. White, R. D. Fleischmann, B. A. Dougherty, T. Mason, A. Ciecko, D. S. Parksey, E. Blair, H. Cittone, E. B. Clark, M. D. Cotton, T. R. Utterback, H. Khouri, H. Qin, J. Vamathevan, J. Gill, V. Scarlato, V. Masignani, M. Pizza, G. Grandi, L. Sun, H. O. Smith, C. M. Fraser, E. R. Moxon, R. Rappuoli, and J. C. Venter. 2000. Complete genome sequence of Neisseria meningitidis serogroup B strain MC58. Science 287:1809-1815. [DOI] [PubMed] [Google Scholar]
- 48.Trent, M. S., A. A. Ribeiro, S. Lin, R. J. Cotter, and C. R. Raetz. 2001. An inner membrane enzyme in Salmonella and Escherichia coli that transfers 4-amino-4-deoxy-l-arabinose to lipid A: induction on polymyxin-resistant mutants and role of a novel lipid-linked donor. J. Biol. Chem. 276:43122-43131. [DOI] [PubMed] [Google Scholar]
- 49.Tzeng, Y. L., K. D. Ambrose, S. Zughaier, X. Zhou, Y. K. Miller, W. M. Shafer, and D. S. Stephens. 2005. Cationic antimicrobial peptide resistance in Neisseria meningitidis. J. Bacteriol. 187:5387-5396. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Vaara, M., and P. Viljanen. 1985. Binding of polymyxin B nonapeptide to gram-negative bacteria. Antimicrob. Agents Chemother. 27:548-554. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Villegas, M. V., and A. I. Hartstein. 2003. Acinetobacter outbreaks, 1977-2000. Infect. Control Hosp. Epidemiol. 24:284-295. [DOI] [PubMed] [Google Scholar]
- 52.Winfield, M. D., T. Latifi, and E. A. Groisman. 2005. Transcriptional regulation of the 4-amino-4-deoxy-l-arabinose biosynthetic genes in Yersinia pestis. J. Biol. Chem. 280:14765-14772. [DOI] [PubMed] [Google Scholar]
- 53.Wosten, M. M., L. F. Kox, S. Chamnongpol, F. C. Soncini, and E. A. Groisman. 2000. A signal transduction system that responds to extracellular iron. Cell 103:113-125. [DOI] [PubMed] [Google Scholar]