Abstract
Transposons are found in virtually all organisms and play fundamental roles in genome evolution. They can also acquire new functions in the host organism and some have been developed as incisive genetic tools for transformation and mutagenesis. The hAT transposon superfamily contains members from the plant and animal kingdoms, some of which are active when introduced into new host organisms. We have identified two new active hAT transposons, AeBuster1, from the mosquito Aedes aegypti and TcBuster from the red flour beetle Tribolium castaneum. Activity of both transposons is illustrated by excision and transposition assays performed in Drosophila melanogaster and Ae. aegypti and by in vitro strand transfer assays. These two active insect transposons are more closely related to the Buster sequences identified in humans than they are to the previously identified active hAT transposons, Ac, Tam3, Tol2, hobo, and Hermes. We therefore reexamined the structural and functional relationships of hAT and hAT-like transposase sequences extracted from genome databases and found that the hAT superfamily is divided into at least two families. This division is supported by a difference in target-site selections generated by active transposons of each family. We name these families the Ac and Buster families after the first identified transposon or transposon-like sequence in each. We find that the recently discovered SPIN transposons of mammals are located within the family of Buster elements.
TRANSPOSONS are fundamental agents of genome evolution and can acquire functions independent of transposition in genomes (Britten 2005; Britten 2006; Volff 2006). The abundance of whole-genome sequence data has greatly increased our ability to identify and characterize the complete complement of transposons within genomes, leading to a deeper understanding of the origins of transposons and their dynamic relationships within the genomes they inhabit. Transposons are classified as either RNA transposons that transpose through an RNA intermediate or as DNA transposons that transpose via a DNA intermediate (Finnegan 1990).
DNA transposons in the hAT superfamily are widespread in plants and animals and include a number of active, well-studied elements such as the Ac transposon of Zea mays, the hobo transposon of Drosophila melanogaster, the Hermes transposon of the housefly Musca domestica, and the Tol2 transposon of the Japanese Medaka fish, Oryzias latipes (McClintock 1950; Blackman et al. 1989; O'Brochta et al. 1996; Kawakami and Shima 1999). hAT transposons are also found in mammalian genomes including humans where they are the most abundant DNA transposons and comprise 1.55% (195 Mb) of the total genome (Lander et al. 2001). None of the hAT elements in the human genome, however, are known to have been active during the past 50 million years (Lander et al. 2001). In contrast, in the genome of the little brown bat, Myotis lucifugus, there has been a marked expansion of DNA transposons with hAT transposons being the most abundant and recent component of this expansion (Ray et al. 2008). No active hAT transposons have been recovered from the genome of M. lucifugus, but the abundance of these and other DNA transposons indicates that some mammals have retained the ability to support high DNA transposon activity (Ray et al. 2008).
SPIN (space invaders) transposons are hAT transposons that have been found in opossum, bushbaby, tenrec, and little brown bat as well as in the African clawed frog, the green anole lizard, murine rodents, and a triatomid bug (Pace et al. 2008; Gilbert et al. 2010). In contrast SPIN transposons are absent from cats, dogs, cattle, armadillo, treeshrew, and humans (Pace et al. 2008). Their discontinuous distribution combined with their sequence conservation and abundance in some species has led to the hypothesis that SPIN transposons have been repeatedly horizontally transferred within deuterostomes; however, as yet, no active native SPIN transposon has been found, nor has the likely vector of these horizontal transfers been identified, although host–parasite relationships have been invoked as possible platforms for enabling this to occur (Pace et al. 2008; Gilbert et al. 2010et al.). We show here that SPINs are closely related to likely exapted and highly conserved proteins present in some mammalian species.
DATA ACCESSION NUMBERS
Protein sequences used in the article were collected from several databases and one manuscript. These are listed below as well as sequence names as they appear in Figure 1. Accession numbers for the appropriate database are show in parentheses if they differ from the sequence name. The sequence SPIN_Ml_a is a variant of the SPIN_Ml element, and the sequence of SPIN_Ml_a is available upon request.
GENBANK (http://www.ncbi.nlm.nih.gov)
Ac-like (AAC46515), Ac (CAA29005), AeBuster1 (ABF20543), AeBuster2 (ABF20544), AmBuster1 (EFB22616), AmBuster2(EFB25016), AmBuster3 (EFB20710), AmBuster4 (EFB22020), BtBuster1 (ABF22695), BtBuster2 (ABF22700), BtBuster3 (ABF22697), CfBuster1 (ABF22696), CfBuster2 (ABF22701), CfBuster3 (XP_854762), CfBuster4 (XP_545451), CsBuster (ABF20548), Daysleeper (CAB68118), DrBuster1 (ABF20549), DrBuster2 (ABF20550), EcBuster1 (XP_001504971), EcBuster3(XP_001503499), EcBuster4 (XP_001504928), Hermes (AAC37217>), hermit (LCU22467), Herves (AAS21248), hobo (A39652), Homer (AAD03082), hopper-we (AAL93203), HsBuster1 (AAF18454), HsBuster2 (ABF22698), HsBuster3 (NP_071373), HsBuster4 (AAS01734), IpTip100 (BAA36225), MamBuster2 (XP_001108973), MamBuster3 (XP_001084430), MamBuster3 (XP_001084430), MamBuster4 (XP_001101327), MmBuster2 (AAF18453), PtBuster2 (ABF22699), PtBuster3 (XP_001142453), PtBuster4 (XP_527300), Restless (CAA93759), RnBuster2 (NP_001102151), SpBuster1 (ABF20546), SpBuster2 (ABF20547), SsBuster4 (XP_001929194), Tam3 (CAA38906), TcBuster (ABF20545), Tol2 (BAA87039), tramp (CAA76545), XtBuster (ABF20551)
ENSEMBL (http://www.ensembl.org)
PtBuster1 (ENSPTRG00000003364)
REPBASE (http://www.girinst.org)
Ac-like2 (hAT-7_DR), Ac-like1 (hAT-6_DR), hAT-5_DR, MlBuster1 (hAT-4_ML), Myotis-hAT1, SPIN_Et, SPIN_Ml, SPIN-Og
TEFam (tefam.biochem.vt.edu)
AeHermes1 (TF0013337), AeBuster3 (TF001186), AeBuster4 (TF001187), AeBuster5 (TF001188), AeBuster7 (TF001336), AeHermes2 (TF0013338), AeTip100-2 (TF000910), Cx-Kink2 (TF001637), Cx-Kink3 (TF001638), Cx-Kink4 (TF001639), Cx-Kink5 (TF001640), Cx-Kink7 (TF001636), Cx-Kink8 (TF001635)
SPIN_Md, SPIN_Xt
The hAT superfamily of DNA transposons also provides a number of examples of elements being exapted to essential functions within the host genome, a process sometimes referred to as domestication. Sinzelle et al. (2009) list 10 examples of domesticated hAT transposons with 8 of these retaining both their DNA binding and catalytic domains. These include the DAYSLEEPER gene from Arabidopsis thaliana, the GARY genes from cereal grasses, the DREF gene from D. melanogaster, the GON-14 and LIN-15B genes of Caenorhabditis elegans, and the TRAMP and Buster genes from Homo sapiens (Sinzelle et al. 2009). Feschotte and Pritham (2007) list 11 examples of hAT transposon domestication with 3 of these possibly also involving the codomestication of P-element-like sequences. hAT transposon sequences are present in the human genome with the Charlie/MER1 hAT elements accounting for approximately 90% of hAT sequences; however, an active hAT transposase consensus sequence for these has not been clearly identified, suggesting that these may be inactive, “dead” transposons (Smit and Riggs 1996; Lander et al. 2001). Similarities in the mechanism of excision and the generation of circular DNA products, or episomes, between the Hermes hAT transposase and the human RAG1 recombinase indicate that both share a common mechanism of transposition (Zhou et al. 2004; O'brochta et al. 2009). However, comparisons of the primary structures of the RAG1 transposase with all eukaryotic transposases suggests that these are more closely related to the Transib transposases (Zhou et al. 2004; Kapitonov and Jurka 2005; O'Brochta et al. 2009).
hAT transposons and related domesticated sequences are thus a very large superfamily both in their diversity and in their absolute numbers and there is an increasing interest in the role that this superfamily has played in the evolution of the plant and animal kingdoms. A handful of different active members of this superfamily have been discovered; indeed Ac, the first transposon to be identified, is a hAT transposon discovered by McClintock on the basis of its mutagenic ability in maize (McClintock 1950). Much remains to be learned about the activity and regulation of hAT elements, particularly when active forms are introduced into new hosts.
Employing a bioinformatic methodology that was previously successful in identifying the active Herves hAT transposon from the mosquito Anopheles gambiae (Arensburger et al. 2005), we sought to determine whether new active hAT transposons were present in two recently sequenced insect genomes—those of the mosquito Aedes aegypti and the red flour beetle, Tribolium castaneum (Nene et al. 2007; Richards et al. 2008). We identified two new active hAT transposons, AeBuster1 from Ae. aegypti and TcBuster from T. castaneum. Sequence comparison with existing hAT transposons and hAT-like sequences showed that both were surprisingly closely related to the Buster genes of deuterostomes yet were clearly active transposons. We demonstrate that both AeBuster1 and TcBuster are active in interplasmid transposition assays performed in D. melangaster and Ae. aegypti embryos. Transposition of these transposons into target DNA generates 8-bp target-site duplications as is true of other hAT elements but with a consensus sequence consistently different to those generated by the Hermes and hobo transposons. These are the first Buster transposons shown to be active.
The activity of both AeBuster1 and TcBuster led us to reexamine the hAT transposon superfamily phylogeny. Previous studies have focused only on those transposons and sequences closely related to the Ac, hobo, Tol2, and Hermes elements and concluded that, while there was evidence that this superfamily is very ancient, the grouping of these elements into clusters consistent with the evolution of their hosts suggested that horizontal transfer had not played a role in their evolution (Simmons 1992; Kempken and Windhofer 2001; Rubin et al. 2001). However, a more recent analysis of hAT elements detected in 12 Drosophila species concluded that four clades (or families) of hAT elements could be identified (Ortiz and Loreto 2009). Furthermore, this phylogeny was not completely congruent with the phylogeny of the host species, indicating a possible role for horizontal transfer in the distribution of some of these hAT transposons, a possibility that was originally conceived when the distribution of hobo elements was first analyzed by DNA sequencing (Ortiz and Loreto 2009). We find that, on the basis of the primary sequence of their transposases, the hAT superfamily is composed of at least two families that we classify as the Ac family and the Buster family, named after the first elements identified in each. The structural distinction between the Ac and Buster families is further supported by the functional difference in target-site selection of members of each upon insertion into target DNA. We find that the recently discovered SPIN elements are members of the Buster family.
MATERIALS AND METHODS
Discovery of novel transposable elements and mammalian exapted genes:
We refined the bioinformatic methods previously used to identify the active Herves hAT element from the mosquito A. gambiae (Arensburger et al. 2005). In brief, whole-genome sequences from Ae. aegypti (Liverpool) and T. castaneum (Georgia 2) were searched for regions showing similarity to known hAT transposase sequences (transposase sequences collected from the following databases: Repbase http://www.girinst.org/, Tefam http://tefam.biochem.vt.edu/tefam/, and NCBI http://www.ncbi.nih.gov), using the TBLASTN program (cutoff value e < 10−4) (Nene et al. 2007; Richards et al. 2008). Coordinates of all identified regions were used as input for a set of custom written PERL scripts for further analysis. These scripts searched each coordinate region and surrounding sequences for nucleotide regions with the following characteristics: (1) presence of an intact ORF, (2) 8-bp target-site duplications (TSDs), (3) terminal inverted repeats (TIRs) ≥10 bp in size immediately adjacent and internal to the TSDs, and (4) the same transposon flanked by different TSDs in the genome and so most likely present at multiple loci. The output of these scripts was manually reviewed. Scripts are available upon request.
In the course of this search a surprising similarity was observed between the AeBuster1 and TcBuster transposase sequences and translated regions of the human genome. These similarities were further investigated using various BLAST programs, leading to the discovery of four putative exapted mammalian genes (described below).
Consensus TSD sequences:
Target-site sequence preferences of 299 putatively full-length hAT transposons were examined, using the following criteria for inclusion into the analysis. The Repbase database was queried for all hAT repeat class entries that had a predicted transposase sequence and belonged to a species with an available whole-genome sequence; this resulted in the inclusion of 273 elements (supporting information, Table S1). Two additional SPIN elements not included in Repbase but previously described were also added (Pace et al. 2008). A total of 24 additional transposons discovered during the course of this analysis were also included.
All potential 8-bp TSD sequences were identified by repeating the following steps for each transposon. First, the genome of origin was searched using the BLAST algorithm (cutoff value e < 10−4) for any region that matched the first 50 bp of the transposon. The genome was then searched again for matches to the last 50 bp of the transposon using the same method. Second, using these matching regions, the 8 bp immediately adjacent and outside of the transposon sequence was identified and stored. Third, to prevent overrepresentation of elements that were likely duplicated through a nontransposase-mediated mechanism, any duplicate 8-bp sequence was removed, leaving only a single representative sequence for each potential TSD. Finally, a 50% consensus sequence was generated.
Sequence selection, alignment, and phylogenetic analysis:
Three amino acid transposase sequence data sets were created and used for phylogenetic analysis. The first database consisted of 50 transposons selected (on the basis of a survey of prior publications) to represent the phylogenetic diversity of the hAT superfamilly. Emphasis for selection was placed on full-length and potentially active transposons rather than on consensus sequences. The number of taxa included in this data set was limited to allow more in-depth phylogenetic analysis (see below). A second database of 78 sequences included the 50 previously identified sequences as well as the amino acid sequences of 28 exapted genes with known sequence similarity to hAT transposons (Table S2). These exapted gene sequences were identified from the literature and as part of this study (see below). The third database of 192 transposase sequences was created by using the transposons used for the consensus TSD analysis (see above), selecting those sequences that had at least 10 identified, unique TSD sequences.
The 50-sequence data set was aligned using the program M-Coffee, which computes a consensus alignment from several multiple sequence alignment programs (PCMA, mafft, clustalw, dialign, poa, muscle, probsons, and T-Coffee) (Moretti et al. 2007). The remaining two data sets were aligned using the multiple sequence alignment program T-Coffee (Notredame 2010). Furthermore, this 50-sequence data set was used as input for the program ProtTest (Abascal et al. 2005) to identify the best-suited amino acid substitution matrix for these data, which was WAG (Whelan and Goldman 2001) with among-site rate variation accommodated by a gamma shape parameter and invariant sites. On the basis of these results, two phylogenetic trees were created using this data set and amino acid substitution matrix: (1) a tree based on the maximum-likelihood optimality criterion using the program TREE-PUZZLE v. 5.2 (Schmidt et al. 2002), and (2) a tree based on a Bayesian estimation of phylogeny using the program MRBAYES (Ronquist and Huelsenbeck 2003). Phylogenetic trees of the other two data sets were constructed using the program FastTree2 using default parameters (Price et al. 2010).
Clones and plasmids:
pBSAeBusterLR:
AeBuster1 was amplified in sections from Ae. aegypti (Orlando). Genomic DNA was purified using a Wizard Genomic DNA Kit (Promega), and 360 ng of DNA was used as template in 50-μl PCR reactions using a TripleMaster PCR System (Eppendorf). The pGT-AeBusterL2 clone was made by PCR amplification using primers that encompassed the region from the TSD of one copy of AeBuster1 to the sequence immediately upstream of the ATG of the AeBuster ORF. The primers used were AeBuster L2 For: 5′-AATGGTACCGCTTATGGCATAGATTCCCAAACTGTG-3′ (TIR in boldface type), and AeBuster L Rev: 5′-GATCTCGAGATCTGAAATTATCAAATAATGAATCGCATATTCTG-3′, with the following PCR program: 94° 2′, 4 × (94° 20′′, 60° 20′′, 72° 30′′), 25 × (94° 20′′, 69° 20′′, 72° 30′′), 72° 5′, 4°. Following amplification, the DNA was purified using a Qiaquick PCR purification kit (Qiagen), quantified on an agarose gel, and ligated into the pGEM-T Easy vector (Promega). Inserts were sequenced, and then clones were digested with KpnI and XhoI (New England Biolabs). Gel-purified fragments (Zymoclean Gel DNA Recovery Kit, Zymo Research) were cloned into pBluescript SK+ digested with the same enzymes to make the clone pBSAeBL2. The right end of AeBuster1 was amplified in a similar manner using the primers AeBuster R For: 5′-GATTCTAGATGCGCATCGAACAACATTTTTAGTGAG-3′ and AeBuster R2 Rev: 5′-AATGAGCTCCCATAAGCCATAGGTTCCCAAACTTTTC-3′ (TIR in boldface type), which encompassed the region immediately 3′ of the stop codon through the target-site duplication. The PCR program was: 94° 2′, 4 × (94° 20′′, 60° 20′′, 72° 30′′), 25 × (94° 20′′, 72° 30′′), 72° 5′, 4°. The right end PCR product was cloned as above, first into pGEM-T Easy and then into the left end clone following digestion with SacI and XbaI (New England Biolabs, NEB) to yield the clone pBSAeBusterLR.
pBSAeBuster donor 1:
SmaI-digested pBSAeBLR was ligated with XmnI-digested pGENToriAlpha, which was derived from pK19 and in which the kanamycinR gene was replaced with the gentamycinR gene from pFastBac HTb (Invitrogen) to generate a donor element, which has a replication origin, a gentamycinR gene, and a lacZ-alpha gene.
pKhsp70AeBuster1:
For cloning the AeBuster1 ORF, PCR was carried out as above, but with the primers AeBuster ORF For, 5′-GATGATATCAGATATGGATAAATGGTTGTTGAAGAAGC-3′, and Ae Buster ORF Rev, 5′-GATGATATCTTAGTGTGATGGATGGCTTGCTTG-3′. The PCR program was: 94° 2′, 3 × (94° 20′′, 63° 20′′, 72° 2′), 25 × (94° 20′′, 68° 20′′, 72° 2′), 72° 5′, 4°. DNA was purified as above, ligated into pGEM-T Easy, and sequenced. One clone, pGEMT-AeBuster clone 9 ORF, contained a sequence with two silent mutations, but no differences in protein sequence from AeBuster1 transposase copies 1 and 2, and was used for the helper plasmid containing the AeBuster1 transposase. pGEMT-AeBuster clone 9 ORF was digested with EcoRV (NEB) and cloned into the SmaI site of pKhsp70 to make the helper pKhsp70AeBuster1.
pBADAeBuster1:
pBAD/Myc-His A (Invitrogen) digested with NcoI and ApaI (NEB) was ligated to a PCR product (digested with NcoI and ApaI) amplified as above using pGEMT-AeBuster clone 9 ORF as the template. The primers used were AeBuster coli F, 5′-GATCCATGGATAAATGGTTGTTGAAGAAGCCC-3′, and AeBuster coli R (AATGGGCCCGTGTGATGGATGGCTTTGCTTG). The same PCR program was used as for the ORF amplification described above. Purification of the His-tagged AeBuster1 transposase was performed using the same protocol as described for the His-tagged Hermes transposase (Zhou et al. 2004).
pBSTcBuster LR1 and pBSTcBusterLR2: TcBuster was amplified from T. castanaeum Ga-2 DNA (obtained from Dr. Susan Brown, Kansas State University, Manhattan, KS). Approximately 30 ng of DNA was used as template in 50 μl PCR reactions using the TripleMaster PCR System (Eppendorf). The clone pBSTcBusterL was made by PCR amplification using primers that encompassed the region from the TSD of TcBuster1 to the sequence immediately upstream of the ATG of the TcBuster1 ORF. The primers used were TcBuster1 L For (AATGGTACCCTTTAGGCCAGTGTTCTTCAACCTG (TIR in boldface type) and TcBuster1 L Rev (CATCTCGAGATTTCTGAACGATTCTAGGTTAGGATCAAAC) with the following PCR program: 94° 2 min, 3 × (94° for 20 sec, 62° for 20 sec, 72° for 30 sec), 26 × (94° for 20 sec, 70° for 20 sec, 72° for 20 sec), 72° for 5 min, 4°. Following amplification the DNA was purified using the QIAquick PCR Purification Kit (Qiagen), quantified on an agarose gel, digested with KpnI and XhoI (NEB), repurified, and ligated into pBluescript SK+ (Stratagene), which had been digested with KpnI and XhoI and gel purified (Zymoclean Gel DNA Recovery Kit, Zymo Research). Inserts were sequenced, and then clones were digested with SacI and XbaI (NEB). Gel-purified vector was ligated to TcBuster1 right end PCR product and digested with SacI and XbaI.
The right end of TcBuster1 was amplified in a similar manner using the primers TcBuster1 R1 For (GATTCTAGACCAAAGCACGGGCTCACCTTGTTC) or TcBuster1 R2 for (GATTCTAGACAACTGATCCATCCCGATATTGATAATTTGTGC) and TcBuster1 R Rev (AATGAGCTCGTATAAGCAGTGTTTTCAACCTTTGCCATCC), using the PCR program described above. The R1 end contained only 145 bp of the transposon, while the R2 end contained 313 bp of transposon, and included some ORF sequence. Following amplification, the PCR products were purified, digested with SacI and XbaI, quantified, and ligated to the left end clone to make clones pBSTcBusterLR1 and pBSTcBusterLR2.
pTcBDG2 and pTcBDG3:
Donor plasmids pTcBDG2 and pTcBDG3 were made by ligation of XbaI-digested pBSTcBusterLR1 or pBSTcBusterLR2 with NheI-digested pGENToriAlpha.
pKhsp70TcBuster1:
The TcBuster helper plasmid was made as follows: PCR was performed with the primers TcBuster1 ORF For
(AATGATATCAGAAATATGATGCTGAATTGGCTCAAAAGTGG) and TcBuster1 ORF Rev (GATGATATCTTAATGACTTTTTTGCGCTTGCTTATTATTGCAC). The PCR program was: 94° for 2 min, 3 × (94° for 20 sec, 67.5° for 20 sec, 72° for 2 min), 26 × (94° for 20 sec, 70° for 20 sec, 72° for 2 min), 72° for 5 min, 4°.
PCR product was purified as above, digested with EcoRV, cloned into the plasmid pKhsp70 (Arensburger et al. 2005), and digested with SmaI to generate pKHsp70TcBuster1. The sequence of the ORF was identical to the sequence from the bioinformatic analysis described above except for a single base change that resulted in a Ser–Thr substitution at aa 551. Thr at this position was found in all of the clones sequenced for the TcBuster1 ORF.
pGDV1:
Is a Bacillus subtilus plasmid incapable of replicating in Escherichia coli and used as a target plasmid in plasmid-based transposition assays (Sarkar et al. 1997a).
Plasmid-based transposition and excision assays:
A mixture of three plasmids, “donor” (pBSAeBuster-GenOriLacZ or pBSTcBuster-GenOriLacZ), “target” (pGDV1), and “helper” (pKhsp70AeBuster1 or pKhsp70TcBuster1) were introduced into insect cells either by transfection of Schneider 2 cells or by microinjection into D. melanogaster and Ae. aegypti preblastoderm embryos as previously described (Arensburger et al. 2005; Sarkar et al. 1997b). Transfection of donor, target, and helper (or negative control plasmids pK19 or Ac5cEGFP) into Schneider 2 cells was performed in six-well plates as previously described (Arensburger et al. 2005). For some experiments the quantity of donor was decreased and the helper increased (1.25 and 3.75 μg). Plasmids were recovered with the Wizard Genomic DNA kit and used to transform DH10 cells by electroporation. Transposition of donor elements into the target plasmids resulted in recombinant plasmids expressing resistance genes for gentamycin and chloramphenicol. To select for transformed DH10 cells with recombinant plasmids resulting from transposition-electroporated cells were plated on LB plates containing gentamycin (7 μg/ml) and chloramphenicol (10 μg/ml). After 3 days incubation at 37°, resistant colonies were picked and grown in LB media containing only gentamycin. Plasmid DNA was purified from these cells using the Wizard Plus miniprep kit (Promega). The presence of a recombinant plasmid arising from transposition was verified by digesting the plasmid DNA with EcoRV to check for a diagnostic pattern of bands (1.1, 1.5, and 3.2 kb). Plasmids passing this initial test were confirmed as transposition events by DNA sequencing.
Alternatively, AeBuster1 excision assays were performed on plasmids recovered from transposition assays performed in Ae. aegypti embryos. Recovered plasmid DNA was digested with EcoRV and the resulting DNA was used to transform DH10 cells by electroporation followed by selection on LB plates containing ampicillin and X-gal (20 mg/liter). Because EcoRV cuts only within the AeBuster1 donor element, plasmids arising as a result of excision are resistant to EcoRV linearization. Uncut excision products efficiently transform E. coli while linearized donor plasmids do not. Putative excision events were confirmed by restriction digestion and DNA sequencing. AmpicillinR, LacZ− colonies were selected, mapped and then sequenced across the empty excision site using the primers 5′-CGTCCCATTCGCCATTCAGG-3′.
Cell-free transposition reactions:
The methods used here are essentially identical to those described for a number of other elements (Zhou et al. 2004). Short (40 bp) DNA oligonucleotides containing the terminal sequences of the left and right ends of AeBuster1 were radiolabeled using T4 kinase and [γ-P32]ATP. Labeled oligonucleotides were annealed to their unlabeled complementary strands in 10 mM Tris-HCl, pH 7.5, by heating at 90° for 5 min and then cooling the reaction at room temperature. The annealed left and right ends were used directly in strand-transfer reactions. The AeBuster1 L-40mers were 5′-CATAGATTCCCAAACTGTGGGTCGCGACCCCCTGGGGGGT and 3′-GTATCTAAGGGTTTGACACCCAGCGCTGGGGGACCCCCCA. The AeBuster1 R-40mers were 5′-CATAGGTTCCCAAACTTTTCGGATGCGCGACCCCCCTAGC and 3′-GTATCCAAGGGTTTGAAAAGCCTACGCGCTGGGGGGATCG. TIRs are in boldface type. Strand-transfer reactions contained 400 nM AeBuster1 transposase and 10 nM AeBuster1 ends in 25 mM MOPS pH 7.6, 5% (v/v) glycerol, 2 mM DTT, 10 mM MgCl2, 100 μg/ml BSA, and 10 nM pUC19 plasmid as target DNA in a final volume of 20 μl. The reaction mixtures were incubated 30° for 2 hr, stopped by adding SDS and EDTA to 1% SDS and 20 mM EDTA and incubated for 1 hr at 37°. The products were displayed on 1% agarose/TAE gels.
RESULTS
The hAT superfamily consists of at least two families of transposons:
We used an algorithm that searched the Ae. aegypti and T. castaneum genomes for hAT transposons on the basis of the presence of TIRs ≥10 bp in length and 8 bp TSDs flanking a sequence with at least one ORF that has similarity to hAT transposases (Arensburger et al. 2005). The conceptual ORF translation of these new elements, as well as selected previously described hAT transposase sequences, was used to construct an unrooted maximum-likelihood transposase tree that revealed two major clusters of related sequences shown in the gray regions of Figure 1 that we have designated the Ac family and the Buster family. The hAT transposase sequences used in this analysis were selected on the basis of the phylogenetic diversity of the superfamily and on their annotation as full-length sequences, thus increasing the likelihood that they are, or recently were, mobile in their host organism. A second tree using the same transposase sequences but using an optimality criterion based on Bayesian inference rather than maximum likelihood was consistent with the tree shown in Figure 1 (the Bayesian tree shown in Figure S1). The relationship between these hAT transposases and known related exapted genes is also shown (Feschotte and Pritham 2007; Sinzelle et al. 2009) (Figure S2).
Figure 1.—
Consensus phylogenetic tree showing the relationship of amino acid transposase sequences, between 50 selected full-length hAT elements based on a maximum-likelihood optimality criterion (50% majority rule consensus). Numbers next to most nodes show quartet puzzling reliability based on 10,000 puzzling steps, a measure of nodal support similar to bootstrapping, produced by the program TREE-PUZZLE. Nodal support inside the SPIN, AeBuster3, SPIN_Ml_a clade is not shown for purposes of clarity. The shaded areas indicate the proposed division of the hAT superfamily into the Buster and Ac families. The scale bar represents a phylogenetic distance of 1 amino acid substitution per site.
The majority of the amino acid sequence variation contributing to this distribution was found to reside in two domains of these transposases. We used the crystal structure of the Hermes transposase and T-coffee alignments of hAT and Buster transposases to reexamine the domain structure of these transposases (Figure S3). We found that both families of hAT transposases have four domains, an N-terminal domain containing a BED zinc finger, a second DNA-binding domain that, at least in the Hermes transposase, also plays a role in oligomerization, a catalytic domain containing the first two carboxylates of the catalytic triad, and then a long insertion domain that contains numerous α-helices (Zhou et al. 2004; Hickman et al. 2005) (Figure S3 and Figure S4). Then follows an α-helix containing a glutamate residue that is in close proximity to the catalytic domain and so completes the catalytic DDE triad (Hickman et al. 2005). This final region has been termed the hAT domain on the basis of its conservation between hAT transposases and its role in oligomerization; however, the crystal structure of the Hermes transposase indicates that multiple regions of this protein contribute to oligomerization (Hickman et al. 2005). The most significant variations among members of the Ac and Buster families lie in the second DNA-binding domain and in the insertion domain.
The Buster transposon family contains a large number of new transposon and transposon-like sequences, including the transposons AeBuster1 from the mosquito Ae. aegypti and TcBuster from T. castaneum whose activity we describe below. The Buster family also includes its namesake, the Buster/Charlie nonfunctional transposon-like sequences originally described in humans (Smit 1999) that we have found to be extremely highly conserved and consider more in detail below.
Buster transposons are widely distributed and were found in aquatic and terrestrial protostomes and deuterostomes. The Buster family contains the recently described transposons from the bat, M. lucifgus (Ray et al. 2006). Myotis hAT forms a clade with Buster transposase sequences from another bat transposase (MlBuster1), a mosquito (Ae. aegypti), a sea urchin Strongylocentrotus purpuratus, and zebrafish (Danio rerio). The newly discovered bat transposase Myotis hAT SPIN_Ml_a forms a clade with AeBuster3 from Ae. aegypti and the recently discovered SPIN transposons in some deuterostomes, some of which have been proposed to have been horizontally transferred between species to best explain their distribution (Pace et al. 2008). The placement of the SPIN transposons within the Buster family reveals their relationship to the Buster transposase sequences found in deuterostomes such as the sea squirt, Ciona savignyi, as well as to those found in the mosquitoes, Ae. aegypti and Culex quinquefasciatus.
Family-specific target-site duplications:
A further sequence difference separates transposon members of the Ac and Buster families although this is based on the mobility properties of the transposase rather than its structure. For the majority of members of the Ac family, the consensus TSD generated upon transposon integration is 5′-nTnnnnAn-3′. The Tol2 transposon, which we classify as being a member of the Ac family, has a slightly altered TSD of 5′-STTATAAS-3′ (where S stands for C or G), which still conforms to the preference of members of this family to insert at sites containing a T and A at the second and seventh positions (Kondrychyn et al. 2009). The Buster transposons in contrast have the TSD consensus of 5′-nnnTAnnn-3′, which we derived both from sequences flanking Buster transposons in their native genomes and from the transposition assays described below. This TSD was also reported flanking hAT sequences in M. lucifugus (Ray et al. 2006), which we now classify as being members of the Buster family. It remains to be determined which part of the hAT transposase mediates target-site recognition.
To determine if this difference in target-site selection was conserved between other members of the Ac and Buster families, we extracted 273 hAT sequences from RepBase using the criteria described in materials and methods, combined them with two recently described SPINs (these two additional SPINs were not deposited into RepBase) and 24 hAT sequences from the current study, and examined their flanking TSDs (Table S1). Those hAT sequences that were annotated with a transposase sequence in RepBase or for which the transposase sequence was known (192 hAT sequences) were used to construct a more extensive tree and the distribution of the two types of TSDs assigned to each sequence (Figure S5). In this tree, members of the Buster family were found within a single clade as was expected from our previous results. Furthermore, 38/45 sequences assigned to this Buster clade were flanked by TSDs that contained the dinucleotide TA in the fourth and fifth position. Of the remaining seven, two (hAT-2 ET, hAT-41 SM) had the dinucleotide NA at the fourth and fifth positions (N indicating that a 50% consensus nucleotide sequence could not be identified for that position), two (SPIN Md, hAT1 MD) contained a nucleotide other than TA at the fourth and fifth positions so did not conform to the Buster target-site consensus, and the remaining three contained NN at these two positions. In contrast only 13/147 hAT sequences in the remaining clades had TSDs with TA at the fourth and fifth positions. Five of these (homo4, hAT-52 HM, hATm-3 HM, hATm-4 HM, hATm-56 HM) also contained T and A at the second and seventh positions, respectively, which, taken alone, are diagnostic of the Ac family TSD consensus sequence; three (hAT-53 HM, hATm34-HM, hATm26-HM) contained either T at the second position or A at the seventh position, while the remaining five did not show any similarities to the Ac family TSD consensus.
Further support for SPINs being members of the Buster family and generating TSDs containing TA at the fourth and fifth positions was obtained through the construction of a functional mammalian SPIN transposon on the basis of consensus sequence of inactive SPINs from mammals and their comparison with the active Buster transposons described here (X. Li, H. Ewis, R. H. Hice, N. Parker, L. Zhou, N. Malani, F. Bushman, C. Feschotte, P. W. Atkinson, N. L. Craig, unpublished results). Analysis of the TSDs generated by transposition of this resurrected SPIN in human cell revealed a consensus sequence of 5′-NNNTANNN-3′ (X. Li, H. Ewis, R. H. Hice, N. Parker, L. Zhou, N. Malani, F. Bushman, C. Feschotte, P. W. Atkinson, N. L. Craig, unpublished results).
The three Tip transposons do not provide sufficient data to enable the placement of Tip transposons into either the Ac or Buster family or into a separate, third family of the hAT superfamily. Previously, sequence similarities between the Tip100 ORFs and Charlie sequences were identified (Robertson 2002). However, at the present time our small sample size does not allow further clarification of their position relative to other hAT transposons.
TcBuster and AeBuster1 are active Buster transposons:
The distribution of Buster transposons across invertebrates and vertebrates led us to ask whether any were indeed active transposons. The Buster transposons from two insects, Ae. aegypti and T. castaneum, contain intact ORFs flanked by perfect TIRs and TSDs. The TcBuster transposon is 2489 bp in length, contains 18-bp TIRs, and has a 2110-bp ORF encoding a transposase 636 amino acids long. The AeBuster1 transposon is 2459 bp in length, contains 15-bp TIRs, and has a 1919-bp ORF encoding a transposase 639 amino acids long. Both transposons thus possessed the structural attributes of active hAT transposases.
Both transposons were cloned from mosquito and beetle genomic DNA and were initially tested for activity in excision assays in Ae. aegypti embryos using established techniques (Atkinson et al. 1993). Upon excision from their sites in the donor plasmid both Buster transposons left footprint sequences consistent with what has been observed for other active hAT transposons such as hobo, Ac, and Tam3 (Coen et al. 1989; Federoff 1989; Atkinson et al. 1993) (Figure 2). Typically, additional nucleotides are found at the excision site and arise following the resolution of hairpin structures formed during excision followed by nonhomologous end joining (Zhou et al. 2004). In some cases the addition of DNA is associated with small deletions of the flanking DNA; in other cases no additional DNA remains and deletions into flanking DNA occur. These types of excision footprints were observed following excision of both AeBuster1 and TcBuster in Ae. aegypti embryos; however, from the small sample size examined, it appeared as if more complete templated addition of extra DNA occurred following TcBuster excision.
Figure 2.—
Footprint sequences remaining at empty sites following excision of either AeBuster1 (A) or TcBuster (B) from donor plasmids in Ae. aegypti embryos. TSDs are underlined and the transposon is shown within the block arrows. The sequences of five empty sites arising from the excision of AeBuster1 and six empty sites arising from the excision of TcBuster are shown with additional DNA that is inserted into the empty sites shown between the TSDs. Smaller arrows show how this is related to sequences within the TSDs.
We next examined the ability of both Buster transposons to transpose in Ae. aegypti and D. melanogaster embryos and in D. melanogaster S2 cells. Interplasmid transposition assays were performed according to previous protocols (Sarkar et al. 1997a). Both Buster transposons were capable of accurate cut-and-paste transposition in D. melanogaster and Ae. aegypti (Tables 1 and 2). As expected for hAT transposons, an 8-bp TSD was generated upon integration (Tables 3, 4, and 5) and, as seen when the endogenous TSDs of these transposons were sequenced, these Busters generated a TSD consensus sequence of 5′-nnnTAnnn-3′, which differs from TSDs generated by the Ac family members (Figure 3). The frequencies of transposition of both transposons in these two insects was within the same order of magnitude recorded for Hermes in both species (Sarkar et al. 1997a,b).
TABLE 1.
In vivo transposition frequencies of TcBuster and AeBuster1 in D. melanogaster and Ae. aegypti embryos
| Transposon/transposase | Species | No. of experiments | No. of donor plasmids | No. of independent transpositions | Transposition frequency (×10−4) |
| TcBuster | D. melanogaster | 1 | 134,600 | 35 | 2.6 |
| 0 | D. melanogaster | 2 | 145,000 | 0 | 0 |
| TcBuster | Ae. aegypti | 3 | 514,000 | 40 | 0.78 |
| 0 | Ae. aegypti | 3 | 382,200 | 0 | 0 |
| AeBuster1 | D. melanogaster | 2 | 171,000 | 26 | 1.52 |
| 0 | D. melanogaster | 3 | 98,400 | 0 | 0 |
| AeBuster1 | Ae. aegypti | 4 | 596,400 | 10 | 0.17 |
| 0 | Ae. aegypti | 3 | 268,000 | 0 | 0 |
TABLE 2.
In vivo transposition frequencies of TcBuster and AeBuster1 in D. melanogaster S2 cells in the presence and absence of transposase
| Transposon/transposase | Species | No. of donor plasmids | No. of independent transpositions | Transposition frequency (×10−4) |
| TcBuster | D. melanogaster/S2 cells | 633,000 | 9 | 0.14 |
| 0 | D. melanogaster/S2 cells | 65,000 | 0 | 0 |
| AeBuster1 | D. melanogaster/S2 cells | 2,413,000 | 7 | 0.03 |
TABLE 3.
TSDs generated by the transposition of TcBuster into developing embryos of D. melanogaster and Ae. aegypti
| Species | TSD | No. of Independent Events | Insertion position |
| D. melanogaster | CCTTAAAC | 3 | 83(+) |
| GTTTAAGG | 3 | 83(−) | |
| TCATATTC | 2 | 398(−) | |
| GACTACAT | 1 | 509(+) | |
| CAGTAATC | 2 | 785(−) | |
| ATCAAAGC | 1 | 801(−) | |
| GTGTAAAT | 2 | 810(−) | |
| AAATAAAC | 1 | 869(−) | |
| TGTTACTG | 1 | 876(+) | |
| GATTAAAG | 1 | 943(+) | |
| CTTTAATC | 2 | 943(−) | |
| CGTTAAAT | 2 | 966(−) | |
| ATCTAAAT | 1 | 1045(−) | |
| CTCTAGCC | 1 | 1965(−) | |
| CTCTAGAG | 2 | 1993(+) | |
| ACATAGCC | 2 | 2207(−) | |
| GAATAAAG | 1 | 2325(+) | |
| CTTTATTC | 1 | 2325(−) | |
| GCTTAAAT | 1 | 2445(+) | |
| GATTAGAC | 1 | 2470(+) | |
| GTCTAATC | 2 | 2470(−) | |
| TGTTATAT | 1 | 2493(+) | |
| ATATAACA | 1 | 2493(−) | |
| Ae. aegypti | ATTTAGAT | 1 | 25(+) |
| CCTTAAAC | 1 | 83(+) | |
| GAATATGA | 1 | 398(+) | |
| GGATAGAC | 2 | 427(+) | |
| GGTTAAAA | 1 | 489(+) | |
| GACTACAT | 2 | 509(+) | |
| GATTACTG | 1 | 785(+) | |
| ATTGCTTT | 1 | 798(+) | |
| GCTTTGAT | 1 | 801(+) | |
| ATTTACAC | 2 | 810(+) | |
| GATTAAAG | 3 | 943(+) | |
| ATTTAACG | 1 | 966(+) | |
| CTATACGT | 1 | 1268(−) | |
| GCATAGGT | 1 | 1925(+) | |
| GGCTAGAG | 4 | 1965(+) | |
| CTCTAGAG | 7 | 1993(+) | |
| CGTTAGCA | 1 | 2050(+) | |
| AGATAGAG | 3 | 2146(+) | |
| GGCTATGT | 1 | 2207(+) | |
| ACATAGCC | 1 | 2207(−) | |
| GAATAAAG | 2 | 2325(+) | |
| CTTTATTC | 1 | 2325(−) | |
| TGTTATAT | 1 | 2493(+) |
The 8-bp TSDs are shown with the conserved TA at positions 4 and 5 in boldface type. Insertion site into the pGDV1 target plasmid (2575 bp in length) and the orientation of insertion are shown.
TABLE 4.
TSDs generated by the transposition of AeBuster1 into developing embryos of D. melanogaster and Ae. aegypti
| Species | TSD | No. of independent events | Insertion position |
| D. melanogaster | CATCAAGA | 1 | 54(+) |
| GTTTAAGG | 1 | 83(−) | |
| ATTCAAAT | 1 | 144(+) | |
| ATTTAGAC | 1 | 168(+) | |
| ACTTAACC | 1 | 207(−) | |
| ATTTACAC | 1 | 810(+) | |
| GATTAAAG | 3 | 943(+) | |
| CTTTAATC | 1 | 943(−) | |
| CTCTAGAG | 2 | 1993(+) | |
| ACTTATAG | 1 | 2160(+) | |
| CTATAAGT | 1 | 2160(−) | |
| ACATAGCC | 1 | 2207(−) | |
| AGCTAACA | 2 | 2214(−) | |
| TTATAACA | 2 | 2381(+) | |
| GTTTACCA | 1 | 2425(+) | |
| TACCAGAT | 1 | 2428(+) | |
| GCTTAAAT | 2 | 2445(+) | |
| GATTAGAC | 1 | 2470(+) | |
| GTCTAATC | 1 | 2470(−) | |
| Ae. aegypti | AAATAAAC | 1 | 460(+) |
| ATTTACAC | 1 | 810(+) | |
| ATATAAAT | 1 | 925(+) | |
| CTTTAATC | 1 | 943(−) | |
| CTCTAGAG | 3 | 1993(+) | |
| GGCTATGT | 1 | 2207(+) | |
| ACATAGCC | 1 | 2207(−) | |
| TGTTAGCG | 1 | 2226(+) |
The 8-bp TSDs are shown with the conserved TA at positions 4 and 5 in boldface type. Insertion site into the pGDV1 target plasmid (2575 bp in length) and the orientation of insertion are shown.
TABLE 5.
TSDs generated by the transposition of AeBuster1and TcBuster into D. melanogaster S2 cells
| Transposon | TSD | No. of independent events | Insertion position |
| AeBuster1 | GTTTAATA | 1 | 32(−) |
| TATTAGAA | 1 | 679(+) | |
| GATTACGT | 1 | 785(+) | |
| ATTTACAC | 1 | 810(+) | |
| GATTAAAG | 1 | 943(+) | |
| CTCTAGAG | 1 | 1993(+) | |
| GATTAGAC | 1 | 2470(−) | |
| TcBuster | ATTTAGAT | 1 | 25(+) |
| CCTTAAAC | 1 | 83(+) | |
| TTTTAACC | 1 | 489(−) | |
| GACTACAT | 1 | 509(+) | |
| GCTTTGAT | 1 | 801(+) | |
| AAATAAAC | 1 | 869(−) | |
| GATTAAAG | 1 | 943(+) | |
| GTTTACCA | 1 | 2425(−) | |
| GCTTAAAT | 1 | 2445(+) |
The 8-bp TSDs are shown with the conserved TA at positions 4 and 5 in boldface type. Insertion site into the pGDV1 target plasmid (2575 bp in length) and the orientation of insertion are shown.
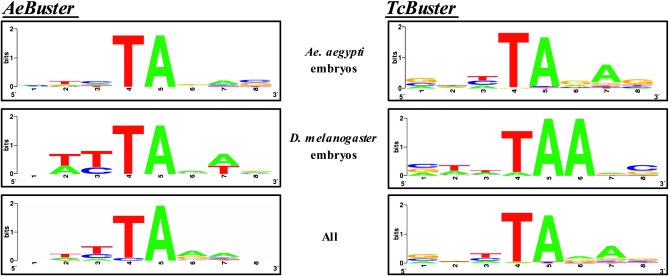
Figure 3.—
WebLogo (http://weblogo.berkeley.edu/logo.cgi) TSDs generated by AeBuster 1 and TcBuster in developing embryos of Ae. aegypti and D. melanogaster.
The AeBuster1 transposase was His-tagged and purified from E. coli using protocols developed for the related Hermes transposase (Zhou et al. 2004). This catalyzed in vitro the strand transfer of a 40 mer containing either end of the AeBuster1 element to a target molecule as was seen in similar experiments with the Hermes transposase (Zhou et al. 2004) (Figure 4). Strand transfer occurred with equal efficiency in the presence of Mg2+ or Mn2+. These data show that at least one type of Buster transposase is amenable to purification from an E. coli expression system and that, in vitro, only 40 bp of the AeBuster1 ends are necessary for recognition and binding of the transposase leading to strand transfer.
Figure 4.—
Strand transfer reactions using precleaved left and right AeBuster1 ends.5′-end-labeled 40mer oligonucleotides containing the L and R ends with their 16-bp TIRs were incubated with AeBuster1 transposase and a pUC19 target DNA and then displayed on an agarose gel. In a single end join (SEJ), a single transposon end oligonucleotide is joined to the target plasmid; in a double end join (DEJ), to transposon ends oligonucleides join to the target DNA at the same position, linearizing the plasmid. Lane 1, left end–no transposase; lane 2, left end–plus transposase; lane 3, right end–no transposase; lane 4, right end–plus transposase. All the reactions shown were run on the same gel and are from the same gel image but have been cropped and arranged for easier viewing.
Conservation of Buster genes in mammals:
The Buster family consists of both active transposons and very similar transposase-like sequences that have lost their TIRs, are highly conserved across species both in sequence and genomic location, and so appear to be important domesticated, or exapted, genes. As shown in Table S3A and Table S3B there is a high degree of conservation of Buster gene ORFs between mammalian species but clearly differences between the Buster genes themselves.
Buster1 genes retain a number of features important to active transposases such as the carboxylate DDE triad that forms the catalytic domain of active hAT transposases and the RW motif located downstream from the second aspartate of this triad. On the basis of the analysis of Hermes transposase this motif is believed to stabilize the penultimate, displaced nucleotide of flanking DNA upon hairpin formation during transposon excision (Zhou et al. 2004). The TcBuster and AeBuster1 transposases contain the CxxH motif also found in the Hermes transposase and many other hAT transposases, and substitution of this histidine with alanine in Hermes severely limits in vitro transposition, indicating its requirement for function (Zhou et al. 2004).
The mammalian Buster1 genes have the CxxH motif expanded to CLLYRH with histidine being replaced by the aromatic amino acid tyrosine but in all other Busters the CxxH motif is conserved. In TcBuster, AeBuster1 transposase, and the mammalian Buster1 proteins, this CxxH or CLLYRH motif is part of a larger motif with the sequence CxxxC(28–29x)CxxH/CLLYRH, which, while similar to the THAP-type Zn and MYM-type Zn finger domains (Bessiere et al. 2008; Gocke and Yu 2008), appears to be a novel type of zinc finger that could, in principle, coordinate a divalent cation. The second C of this expanded motif is two residues upstream from the central D of the catalytic triad of these transposases, raising the question as to whether this region of the molecule can actually participate in two different chelations of a divalent cation. The conservation of these motifs in all these mammalian Buster1 proteins suggests that Buster1 remains functional in all these species and is supported by the low (dN/dS) values of Buster1 across these species (Tables S3A and Tables S3B).
The Buster2, 3, and 4 sequences are present in many eutherian species and are also highly conserved (Tables S3A, Table S3B, and Table S4). They are all found in humans, chimps, rhesus monkeys, dogs, and cattle with Busters2 and 3 previously being identified in humans (Smit 1999). Buster3, also found in horses, is as conserved among these species as is Buster1; however, its DDE motif has been replaced by ADE that most likely renders it incapable of transposase-like catalytic activity. Buster2 is also present in these four species and, in rats and mice, it is part of a fusion with a long N-terminal region with its own additional zinc finger motif. Rodent Buster2 is highly conserved and is found in Rattus and Mus EST libraries. In humans, chimps, dogs, and cattle this additional N-terminal domain is present at the Buster2 locus and conceptual translations indicate that the same protein could be synthesized in these species. The annotation of Buster 4 in the human genome shows that it also contains an additional long N-terminal domain that contains a leucine-rich domain and an integrase domain. This has previously been deposited into GenBank as a SCAN domain containing protein 3 (NP_443155.1|), the carboxy end of which is identical to the Charlie 10 hAT element.
DISCUSSION
We have presented phylogenetic and integration site analyses that support the hypothesis that the hAT superfamily is composed of two distinct families of transposons. Previous analyses of hAT transposon phylogeny focused only on transposons within what we now define as the Ac branch of the hAT element superfamily (Kempken and Windhofer 2001; Rubin et al. 2001). The members of this branch known to be active are Ac, Tam3, hobo, Hermes, Herves, and Tol2 with all also being active in new host species (Martin et al. 1989; Haring et al. 1991; O'brochta et al. 1994; Sarkar et al. 1997a; Kawakamai and Noda 2004; Arensburger et al. 2005; Evertts et al. 2007)
Our data show that members of the Buster family, which include the recently discovered SPIN transposons, are in fact capable of transposition upon introduction to new host species. Our data also illustrate the strong conservation of several domesticated Buster genes within several mammalian species including primates, dogs, horses, and cattle, strongly suggesting an essential, but as yet, unknown function for these genes. Most probably, especially for Buster1, this involves recognizing and binding to DNA and then possibly modifying it similar in principle to the example of how the RAG1 protein, which is related to the Transib transposase, rearranges specific chromosomal DNA as part of the process of V(D)J recombination (Kapitonov and Jurka 2005). We speculate the true roles of these Buster proteins may therefore be closely related to their original roles as recombinases.
The organization of the Buster family is complex with close relationships between Buster sequences from mammals and Buster transposons from invertebrates. The M. lucifugus Ml Buster1 and Myotis-hAT transposons are within a clade shared with Buster transposon sequences from sea urchin, zebrafish, and Ae. aegypti, while the Myotis SPIN_MI_a transposon is closely related to the AeBuster3 transposon of Ae. aegypti and the newly discovered SPIN transposons from mammals and the pipid frog, Xenopus tropicalis. On separate branches are Buster transposons from two mosquitoes (Ae. aegypti, Cx. quinquefasciatus), a beetle (T. castaneum), a tunicate (Ciona savigny), and the highly conserved domesticated Buster1, 2, 3, and 4 sequences from mammals.
The breadth, yet discontinuity, in the distribution of Buster family sequences across vertebrates and invertebrates is intriguing (Figure 1). In cases where domestication has occurred in the mammals, the Buster sequences can be highly conserved (Table S3A, Table S3B, and Table S4). As noted by Pace et al. (2008), the SPIN transposons are also highly conserved among vertebrates as well as with an insect species, leading these authors to propose that they have been horizontally transferred between species (Gilbert et al. 2010). Given this hypothesis of horizonatal transfer of SPIN transposons between mammals, it is possible that the related Buster transposons may also be capable of transposition in vertebrates, including humans.
AeBuster1 is the first active DNA transposon to be isolated from the human disease vector mosquito Ae. aegypti. TcBuster is also an active transposon yet appears to be present in the sequenced T. castaneum genome only once. Given its activity in D. melanogaster and Ae. aegypti, this suggests that TcBuster may be a new introduction into T. castaneum or, alternatively, it has been subjected to fairly efficient negative regulation in this beetle.
Comparison of the TSDs generated by AeBuster1 and TcBuster with those generated by hAT transposons of the Ac family show a clear difference in target-site specificity. In all cases 8-bp TSDs are generated but the consensus sequence between TcBuster, AeBuster1, and the other hAT transposons differs in that there is a strong preference for TA in the central positions of the 8-bp TSD. A more direct comparison between AeBuster1, TcBuster, and Hermes transposition into the pGDV1 target plasmid in transposition assays performed in D. melanogaster clearly illustrates this difference (Sarkar et al. 1997a). The hot spots of Hermes transposition into pGDV1 at sites 318 bp, 736 bp, 1254 bp, and 2303 bp are never targeted by AeBuster1 or TcBuster, while the hot spots of both AeBuster1 and TcBuster transposition into pGDV1 at sites 943 bp, 1993 bp, and 2470 bp are, with the exception of a single Hermes transposition into 1993 bp in Ae. aegypti, not targeted by Hermes (Sarkar et al. 1997a,b). AeBuster1 and TcBuster therefore display a markedly different target-site preference from the Hermes transposon and other members of the Ac family.
The structural basis for this difference in TSD preference is not known but it is reasonable to speculate that this consistent difference between members of the Ac and Buster families resides in an as-yet unidentified region of the transposase that recognizes target DNA. The structure of the Hermes transposase reveals two DNA binding domains that, based on secondary structure alignments, are also present in the two active Buster transposases although it is not known if either plays a role in target-site recognition. Numerous amino acid differences exist between these regions. The difference in TSDs between the two active Buster transposases and the active members of the Ac family provides a means by which the target-site recognition sequence could be clearly identified through mutagenesis followed by assays that would reveal a change in TSD consensus from the nTnnnnAn form to the nnnTAnnn form or vice versa.
There are clear similarities between the TSDs generated by AeBuster1 and TcBuster and those surrounding a subset of hAT-like MER1 elements previously identified in the human genome (Smit and Riggs 1996). More recently, the insertion-site preferences of MER1A, MER1B, MER20, MER30, and MER58A sequences were determined through the bioinformatic analysis of their positions within other transposons in the human genome and found to have the same preferences for T and A at the fourth and fifth positions of the 8-bp TSD (Levy et al. 2009). We propose that these MER1 elements are actually remnants of once active Buster transposons, the only extant members of which are now the four domesticated Buster genes, which, on the basis of their conservation between species, clearly have important, but as-yet unknown, functions.
Our assignment of the hAT superfamily into at least two families on the basis of the sequence of their transposases and target-site selections is necessarily constrained by the number of completely sequenced genomes and our ability to locate full-length hAT transposases that are most likely functional. Similarly, the TIRs of hAT transposons can exhibit much variation in their sequence, making the identification of internally deleted hAT transposons and hAT MITEs problematic. Flanking 8-bp TSDs conforming to one of two consensus sequences described here are informative but not necessarily definitive since other transposons, for example the P element of D. melanogaster, also generate 8-bp TSDs upon insertion into DNA. Our data do not allow the assignment of Tip100 transposons to one of the two families. We are also unable to assign nDart hAT elements from rice, which have been shown to generate GC-rich TSDs when they are mobilized through crossing with strains containing active Dart transposons (Takagi et al. 2010). The absence of an identified functional Dart transposase also currently prevents the true assignment of these transposons within the hAT superfamily.
The discovery of active AeBuster1 and TcBuster transposons and their ability to transpose in new hosts offers support for the proposition that members of this family of transposons have been, at some stage, transferred between deuterostomes and protostomes. Indeed the presence of Buster sequences in a single clade composed of such diverse host species as mosquito, sea urchin, zebrafish, and bat illustrate the lack of concordance between the phylogenies of these sequences and their hosts.
The size and diversity of the hAT superfamily is somewhat unique among eukaryote transposons. On the basis of the structure and target-site recognition, it contains clearly distinguishable families with multiple active members from each available for modification and study both in vitro and in vivo. Active members from the Ac and Buster families are capable of high levels of transpositional activity in new hosts in different phyla and, within each family, clear examples of transpoase domestication are becoming more apparent as more genomes continue to be annotated. The presence and distribution of Buster transposons in mosquitoes, known vectors of pathogens between vertebrates and invertebrates, offers one conceivable route of horizontal transfer of these transposons and may, in part, explain the distribution of these transposons and domesticated sequences that is emerging. As such the ability to modify and exquisitely control their activity and target-site specificity should enable them to be become efficient genetic tools in medical and agricultural applications.
Acknowledgments
This research was supported by National Institutes of Health grants AI45741 to PWA and NLC and GM48102 to D.A.O. and P.W.A. N.L.C. is an Investigator at the Howard Hughes Medical Institute.
LITERATURE CITED
- Abascal F., Zardoya R., Posada D., 2005. ProtTest: selection of best-fit models of protein evolution. Bioinformatics 21: 2104–2105 [DOI] [PubMed] [Google Scholar]
- Arensburger P., O'Brochta D. A., Orsetti J., Atkinson P. W., 2005. Herves, a new active hAT transposable element from the African malaria mosquito Anopheles gambiae. Genetics 169: 697–708 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Atkinson P. W., Warren W. D., O'Brochta D. A., 1993. The hobo transposable element of Drosophila can be cross-mobilized in houseflies and excises like the Ac element of maize. Proc. Natl. Acad. Sci. USA 83: 9693–9697 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bessiere D., Lacroix C., Campagne S., Ecochard V., Guillet V., et al. , 2008. Structure-function analysis of the THAP zinc finger of THAP1, a large C2H2 DNA binding module linked to Rb/E2F pathways. J. Biol. Chem. 283: 4352–4363 [DOI] [PubMed] [Google Scholar]
- Blackman R. K., Macy M., Koehler D., Grimaila R., Gelbart W. M., 1989. Identification of a fully functional hobo transposable element and its use for germ line transformation of Drosophila. EMBO J. 8: 211–217 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Britten R., 2005. Transposable elements have contributed to thousands of human proteins. Proc. Natl. Acad. Sci. USA 103: 1798–1803 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Britten R., 2006. Almost all human genes resulted from ancient duplication. Proc. Natl. Acad. Sci. USA 193: 19027–19032 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coen E. S., Robbins T. P., Almeida J., Hudson A., Carpenter R., 1989. Consequences and mechanisms of transposition in Antirrhinum majus., pp. 413–436 in Mobile DNA, edited by Berg D. E., Howe M. M. American Society for Microbiology, Washington, DC [Google Scholar]
- Evertts A. G., Plymire C., Craig N. L., Levin H. L., 2007. The Hermes transposon of Musca domestica is an efficient tool for the mutagenesis of Schizosaccharomyces pombe. Genetics 177: 2519–2523 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Federoff N. V., 1989. Maize transposable elements, pp. 375–411 in Mobile DNA, edited by Berg D. E., Howe M. M. ASM Press, Washington, DC [Google Scholar]
- Feschotte C., Pritham E. J., 2007. DNA transposons and the evolution of eukaryotic genomes. Annu. Rev. Genet. 41: 331–368 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Finnegan D. J., 1990. Transposable elements and DNA transposition in eukaryotes. Curr. Op. Cell. Biol. 2: 471–477 [DOI] [PubMed] [Google Scholar]
- Gilbert C., Schaack S., Pace J. K. I., Brindley P. J., Feshotte C., 2010. A role for host-parasite interactions in the horizontal transfer of transposons across phyla. Nature 464: 1347–1350 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gocke C. B., Yu H., 2008. ZNF198 stabilizes the LSD1-coREST-HDCAC1 complex on chromatin through its MYM-type zinc fingers. PLoS One 3: e3255. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haring M. A., Teeuwen-de Vroomen M. J., Nijkamp H. J., Hille J., 1991. Trans-activation of an artificial dTam3 transposable element in transgenic tobacco plants. Plant Mol. Biol. 16: 39–47 [DOI] [PubMed] [Google Scholar]
- Hickman A. B., Perez Z. N., Zhou L., Musingarimi P., Ghirlando R., et al. , 2005. Molecular architecture of a eukaryotic transposase. Nat. Struct. Mol. Biol. 12: 715–721 [DOI] [PubMed] [Google Scholar]
- Kapitonov V. V., Jurka J., 2005. RAG1 core and V(D)J recombination signal sequnces were derived from Transib transposons. PLoS Biol. 3: e181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kawakamai K., Noda T., 2004. Transposition of the Tol2 element, an Ac-like element from the Japanese medaka fish, Oryzias latipes, in mouse embryonic stem cells. Genetics 166: 895–899 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kawakami K., Shima A., 1999. Identification of the Tol2 transposase of the medaka fish Oryzias latipes that catalyzes excision of a nonautonomous Tol2 element in zebrafish Danio rerio. Gene 240: 239–244 [DOI] [PubMed] [Google Scholar]
- Kempken F., Windhofer F., 2001. The hAT family: a versatile transposon group common to plants, fungi, animals, and man. Chromosoma 110: 1–9 [DOI] [PubMed] [Google Scholar]
- Kondrychyn I., Garcia-Lecea M., Emelyanov A., Parinov S., Korzh V., 2009. Genome-wide analysis of Tol2 transposon reintegration in zebrafish. BMC Genomics 10: 418. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lander E. S., Linton L. M., Birren B., Nusbaum C., Zody M. C., et al. , 2001. Initial sequencing and analysis of the human genome. Nature 409: 860–921 [DOI] [PubMed] [Google Scholar]
- Levy A., Schwartz S., Ast G., 2009. Large-scale discovery of insertion hotspots and preferential integration sites of human transposed elements. Nucleic Acids Res. 38: 1515–1530 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martin C., Prescott A., Lister C., MacKay S., 1989. Activity of the transposon Tam3 in Antirrhinum and tobacco: possible role of DNA methylation. EMBO J. 8: 997–1004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- McClintock B., 1950. The origin and behavior of mutable loci in maize. Proc. Natl. Acad. Sci. USA 36: 344–355 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moretti S., Armougom F., Wallace I. M., Higgins D. G., Jongeneel C. V., et al. , 2007. The M-Coffee web server: a meta-method for computing multiple sequence alignments by combining alternative alignment methods. Nucleic Acids Res. 35: W645–W648 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nene V., Wortman J. R., Lawson D., Haas B., et al. , 2007. Genome sequence of Aedes aegypti, a major arbovirus vector. Science 316: 1718–1723 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Notredame C., 2010. Computing mutiple sequence/structure alignments with the T-coffee package. Curr. Protoc. Bioinformatics 3.8: 1–25 [DOI] [PubMed] [Google Scholar]
- O'Brochta D. A., Warren W. D., Saville K. J., Atkinson P. W., 1994. Interplasmid transposition of Drosophila hobo elements in non-drosophilid insects. Mol. Gen. Genet. 244: 9–14 [DOI] [PubMed] [Google Scholar]
- O'Brochta D. A., Warren W. D., Saville K. J., Atkinson P. W., 1996. Hermes, a functional non-drosophilid gene vector from Musca domestica. Genetics 142: 907–914 [DOI] [PMC free article] [PubMed] [Google Scholar]
- O'Brochta D. A., Stosic C. D., Pilitt K., Subramanian R. A., Hice R., et al. , 2009. Transpositionally active episomal hAT elements. BMC Mol. Biol. 14: 108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ortiz M., Loreto E. L. S., 2009. Characterization of new hAT transposable elements in 12 Drosophila genomes. Genetica 135: 67–75 [DOI] [PubMed] [Google Scholar]
- Pace J. K. I., Glibert C., Clark M. S., Feschotte C., 2008. Repeated horizontal transfer of a DNA transposon in mammals and other tetrapods. Proc. Natl. Acad. Sci. USA 105: 17023–17028 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Price M. N., Dehal P. S., Arkin A. P., 2010. FastTree 2: approximately maximum-likelihood trees for large alignments. PLoS One 5: e9490. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ray D. A., Pagan H. J. T., Thompson M. L., Stevens R. D., 2006. Bats with hATs: Evidence for recent DNA transposon activity in genus Myotis. Mol. Biol. Evol. 24: 632–639 [DOI] [PubMed] [Google Scholar]
- Ray D. A., Feschotte C., Pagan H. J. T., Smith J. D., Pritham E. J., et al. , 2008. Multiple waves of recent DNA transposon activity in the bat, Myotis lucifugus. Genome Research 18: 717–728 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Richards S., Gibbs R. A., Weinstock G. M., Brown S. J., Dennell R., et al. , 2008. The genome of the modern beetle and pest Tribolium castaneum. Nature 452: 949–955 [DOI] [PubMed] [Google Scholar]
- Robertson H. M., 2002. Evolution of DNA transposons in eukaryotes, pp. 1093–1110 in Mobile DNA II, edited by Craig N. L., Craigie R., Gellert M., Lambowitz A. M. ASM Press, Washington, DC [Google Scholar]
- Ronquist F., Huelsenbeck J. P., 2003. MRBAYES 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19: 1572–1574 [DOI] [PubMed] [Google Scholar]
- Rubin E., Lithwick G., Levy A. A., 2001. Structure and evolution of the hAT transposon superfamily. Genetics 158: 949–957 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sarkar A., Coates C. J., Whyard S., Willhoeft U., Atkinson P. W., et al. , 1997a. The Hermes element from Musca domestica can transpose in four families of cylorrhaphan flies. Genetica 99: 15–29 [DOI] [PubMed] [Google Scholar]
- Sarkar A., Yardley K., Atkinson P. W., James A. A., O' Brochta D. A., 1997b. Transposition of the Hermes element in embryos of the vector mosquito, Aedes aegypti. Insect Biochem. Mol. Biol. 27: 359–363 [DOI] [PubMed] [Google Scholar]
- Schmidt H. A., Strimmer K., Vingron M., von Haeseler A., 2002. TREE-PUZZLE: maximum likelihood phylogenetic analysis using quartets and parallel computing. Bioinformatics 18: 502–504 [DOI] [PubMed] [Google Scholar]
- Simmons G. M., 1992. Horizontal transfer of hobo transposable elements within the Drosophila melanogaster species complex: evidence from DNA sequencing. Mol. Biol. Evol. 9: 1050–1060 [DOI] [PubMed] [Google Scholar]
- Sinzelle L., Izsvak Z., Ivics Z., 2009. Molecular domestication of transposable elements: from detrimental parasites to useful host genes. Cell. Mol. Life Sci. 66: 1073–1093 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smit A. F. A., 1999. Interspersed repeats and other mementos of transposable elements in Mamm. Genomes. Curr. Opinion Genet. Dev. 9: 657–663 [DOI] [PubMed] [Google Scholar]
- Smit A. F. A., Riggs A. D., 1996. Tiggers and other DNA transposon fossils in the human genome. Proc. Natl. Acad. Sci. USA 93: 1443–1448 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Takagi K., Maekawa M., Tsugane K., Iida S., 2010. Transposition and traget preferences of an active nonautonmous DNA transposon nDart1 and its relatives belonging to the hAT superfamily in rice. Mol. Genet. Genomics 284: 343–355 [DOI] [PubMed] [Google Scholar]
- Volff J.-N., 2006. Turning junk into gold: domestication of transposable elements and the creation of new genes in eukaryotes. BioEssays 28: 913–922 [DOI] [PubMed] [Google Scholar]
- Whelan S., Goldman N., 2001. A general empirical model of protein evolution derived from multiple protein families using a maximum likelihood approach. Mol. Biol. Evol. 18: 691–699 [DOI] [PubMed] [Google Scholar]
- Zhou L., Mitra R., Atkinson P. W., Hickman A. B., Dyda F., et al. , 2004. Transposition of hAT elements links transposable elements and V(D)J recombination. Nature 432: 995–1001 [DOI] [PubMed] [Google Scholar]