Through genetic analysis, this work identifies a growth function for a subset of ligands (CHALf) of the EPFL family. The results indicate that different EPFL members modulate discrete functions of the ER receptor family and that EPFLf signal specificity is achieved through spatial restriction of the TMM receptor, which potentiates stomatal EPFL signals and dampens CHALf growth signals.
Abstract
Core signaling pathways function in multiple programs during multicellular development. The mechanisms that compartmentalize pathway function or confer process specificity, however, remain largely unknown. In Arabidopsis thaliana, ERECTA (ER) family receptors have major roles in many growth and cell fate decisions. The ER family acts with receptor TOO MANY MOUTHS (TMM) and several ligands of the EPIDERMAL PATTERNING FACTOR LIKE (EPFL) family, which play distinct yet overlapping roles in patterning of epidermal stomata. Here, our examination of EPFL genes EPFL6/CHALLAH (CHAL), EPFL5/CHALLAH-LIKE1, and EPFL4/CHALLAH-LIKE2 (CLL2) reveals that this family may mediate additional ER-dependent processes. chal cll2 mutants display growth phenotypes characteristic of er mutants, and genetic interactions are consistent with CHAL family molecules acting as ER family ligands. We propose that different classes of EPFL genes regulate different aspects of ER family function and introduce a TMM-based discriminatory mechanism that permits simultaneous, yet compartmentalized and distinct, function of the ER family receptors in growth and epidermal patterning.
INTRODUCTION
The orchestration of programs that produce, pattern, and protect developing tissues is a complex regulatory problem. Not only must individual cells execute complex genetic programs to obtain distinctive identities and morphologies, but they must do so in concert with their neighbors. Both plants and animals use peptide ligands and cell surface receptor-based signaling to establish cell fate and pattern and to regulate self-renewal. In plants, this signaling is often mediated by leucine-rich repeat, single transmembrane pass receptor-like kinases (LRR-RLKs; Shiu and Bleecker, 2001). Putative peptide ligands for these LRR-RLKs are structurally diverse, ranging in size from modified pentapeptides to small (50 to 100 amino acids long) proteins and are often part of large gene families (reviewed in Butenko et al., 2009; Jun et al., 2010; Rowe and Bergmann, 2010; Rychel et al., 2010). The roles and receptor interactions of these putative ligands remain largely unknown, as do the mechanisms that enable related ligands to function in distinct processes and interact with different receptors.
One of the most extensively studied LRR-RLKs is that encoded by the Arabidopsis thaliana ERECTA (ER) gene, mutations in which result in altered plant stature and morphology (Torii et al., 1996). Demonstration of ER functions in development (cell proliferation, cell fate specification, and tissue patterning in most aerial organs), in homeostasis, and in immunity indicate that ER is a central player in multiple signaling contexts (Godiard et al., 2003; Xu et al., 2003; Shpak et al., 2004, 2005; Masle et al., 2005; Pillitteri et al., 2007). ER is partially redundant with two close paralogs, ERECTA-LIKE1 (ERL1) and ERL2, and these three genes are known as the ER family (ERf). Although ERL1 and ERL2 are capable of substituting for ER when expressed in its place, their endogenous expression patterns are distinct from that of ER, and they confer clear loss-of-function stomatal and growth phenotypes only in an er background (Shpak et al., 2004, 2005).
The patterning of stomata (epidermal pores that facilitate plant/atmosphere gas exchange) provides a simple, sensitive, and quantifiable system for the study of basic developmental phenomena. In Arabidopsis, asymmetric, stem cell–like divisions and cell–cell signaling are required for the arrangement of stomata in stereotyped patterns, and stomata and precursors can be analyzed to provide qualitative and quantitative measures of signaling outputs (Bergmann and Sack, 2007). For example, elimination of the entire ER family (er erl1 erl2) results in dramatic overproduction of mispatterned stomata (Shpak et al., 2005). Examination of lineage progression defects in single and double mutants suggested that ER is primarily responsible for restricting entry divisions, whereas ERL1 and ERL2 maintain the proliferative capacity of intermediate precursor stages (Shpak et al., 2005). Stomatal production and pattern are also regulated by a LRR receptor-like protein (LRR-RLP), TOO MANY MOUTHS (TMM) (Nadeau and Sack, 2002). Elimination of TMM leads to stomatal overproliferation and mispatterning phenotypes in leaves similar to, but less severe than, those of er erl1 erl2 (Geisler et al., 2000; Nadeau and Sack, 2002). Curiously, loss of TMM has an opposite phenotype in stems; here, it results in a lack of stomata (Geisler et al., 1998; Bhave et al., 2009). TMM is expressed in the epidermis of both leaves and stems and in both organs is restricted to stomatal precursor cells and their sisters (Nadeau and Sack, 2002).
Whereas TMM and the ERf are involved in overlapping developmental processes, the biochemical bases of their activities are likely to be different. All four proteins possess extracellular LRR repeats, but TMM, unlike the ERf receptors, lacks an intracellular kinase domain. By analogy with signaling pathways in the shoot meristem, it was hypothesized that TMM could participate in active signal transduction only though physical interactions with proteins that possess such domains (Nadeau and Sack, 2002; Shpak et al., 2005). Alternatively, TMM might modulate receptor kinase signaling by forming inactive associations with RLKs or altering availability of an RLK’s ligand (Shpak et al., 2005; Abrash and Bergmann, 2010).
Strong candidates to be ligands of the ERf and TMM receptors are members of the EPIDERMAL PATTERNING FACTOR LIKE (EPFL) family. The EPFLs are small proteins predicted (and in one case, demonstrated) to be processed into ~45– to 75–amino acid long secreted peptides with compact Cys knot structures (Kondo et al., 2009; Sugano et al., 2009). Three members of the gene family, EPF1, EPF2, and EPFL9/STOMAGEN, have clear roles in stomatal development. EPF1 and EPF2 are expressed specifically within subsets of stomatal lineage cells, and STOMAGEN is expressed in leaf mesophyll; all three depend largely upon TMM for their ability to affect stomatal development (Hara et al., 2007, 2009; Hunt and Gray, 2009; Kondo et al., 2009; Sugano et al., 2009). A fourth member of the family, EPFL6/CHALLAH (CHAL), however, exhibits different behaviors. CHAL was initially identified in a screen for suppressors of tmm-1 tissue-specific phenotypes, and consistent with this genetic relationship (and in contrast with other EPFLs), phenotypic effects of CHAL loss or overexpression are revealed or enhanced in a tmm mutant background. CHAL does not need TMM to signal and, in fact, appears to be inhibited by it (Abrash and Bergmann, 2010).
The diverse behaviors of EPFL family members, when considered in combination with TMM and the ER family and quantified using the discrete and sensitive output of stomatal development, provide a system in which to design and test models of signaling specificity during plant development. Here, we identify phenotypes for two additional EPFL family members that align them with ER functions and that uncover redundancy among related ligands. Our data suggest that different EPFL family members mediate different ERf roles, with CHAL-related family members dedicated for growth regulatory processes and EPF1/2-related members specialized for stomatal regulation. Together, our results point to a buffering mechanism that permits simultaneous, yet compartmentalized and distinct, function of a common receptor set during tissue patterning and organ growth.
RESULTS
CHAL Family Genes Are Involved in Developmental Processes outside of the Stomatal Lineage
To understand the mechanisms underlying CHAL’s unique and tissue-specific relationship to the ERf and TMM, we asked whether other EPFL family genes might be acting redundantly with CHAL, thereby obscuring CHAL’s true functions. Among the 11 members of the EPFL family, EPFL4 and EPFL5 share with CHAL a highly conserved sequence of amino acids (17/18 identical) located at the C terminus of the protein (Figure 1A). Because of this sequence similarity and functional similarities later revealed by our studies, it became useful to distinguish this subfamily of EPFLs as CHAL family (CHALf) ligands; we will refer to the CHAL paralogs as CHALLAH-LIKE1 (CLL1; EPFL5/At3g22820) and CLL2 (EPFL4/At4g14723). We obtained and characterized T-DNA mutant lines for each gene; cll1-1 (SALK_005080) bears a T-DNA insertion in the 5′ untranslated region and produces reduced transcript, whereas cll2-1 (SALK_071065) bears a T-DNA insertion in the second exon and appears to be a null allele (Figures 1B to 1E). Neither single mutant displayed obvious phenotypes related to stomatal development (data not shown; Hara et al., 2009). The structural similarities among the three CHALf members might lead to redundancy, such that a phenotype would be revealed only in multiple mutants; however, even in chal cll1 cll2 triple homozygotes, stomatal pattern appeared normal, and we observed no change in stomatal density (Figure 1F).
Figure 1.
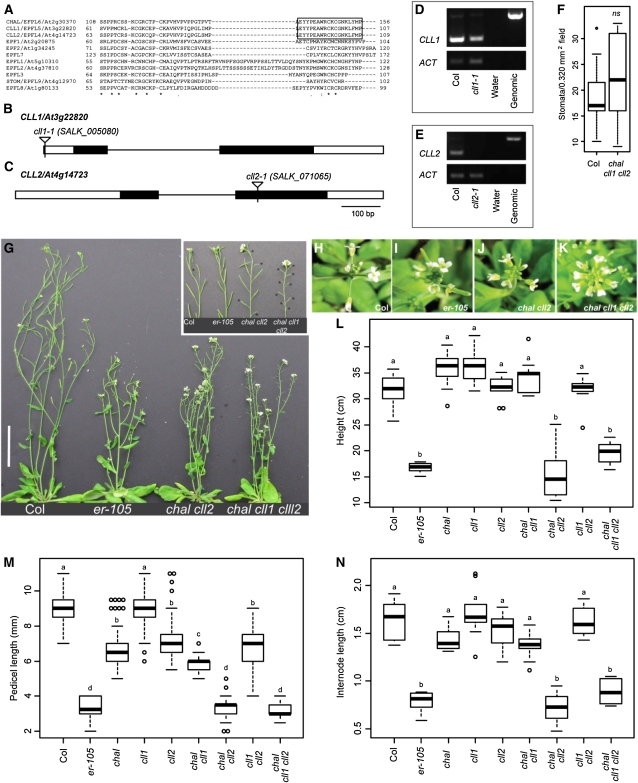
Mutations in CHAL and Paralogs CLL1 and CLL2 Generate Nonstomatal Growth Phenotypes Resembling er Mutants.
(A) ClustalW2 (Chenna et al., 2003) alignment of C-terminal region of EPF-like proteins. The box indicates the region highly conserved between CHAL, CLL1, and CLL2; asterisk, identical residue; colon, conserved substitution; dot, semiconserved substitution. Numbers immediately preceding and following the sequence indicate the amino acid position within each respective protein.
(B) and (C) Schematics of CLL1 and CLL2 loci, respectively. Empty boxes, annotated untranslated regions; filled boxes, exons; lines, introns; triangles, positions of T-DNA insertions.
(D) and (E) RT-PCR analysis of cll1-1 and cll2-1 alleles, respectively (ACT is ACTIN control; “Genomic” indicates genomic DNA template).
(F) Quantification of stomatal density in 8 DAG adaxial cotyledons. Data are displayed as boxplot graphs following standard conventions. Bold lines indicate medians, boxes indicate quartiles above and below median, whiskers extend to most extreme value no more than 1.5 interquartile ranges from box, and circles indicate values beyond whiskers. n ≥ 14 plants per genotype. ns, not significantly different from the wild type by Wilcoxon two-sample test (P > 0.2).
(G) Aerial portions of 48 DAG Col, er-105, chal cll2, and chal cll1 cll2 plants. Inset of primary inflorescences shows sterility of chal cll1 cll2, and asterisks mark sterile siliques. Bar = 5 cm.
(H) to (K) Inflorescence stem apices ~1 week after bolting illustrating reduction in pedicel elongation in er-105 (I), chal cll2 (J), and chal cll1 cll2 (K) relative to Col (H).
(L) Quantification of plant height at 48 DAG; n = 10 plants per genotype.
(M) Quantification of pedicel lengths at 6.5 weeks of age; n = 50 pedicels per genotype, as five pedicels sampled from each of 10 individuals.
(N) Quantification of internode lengths at 6.5 weeks of age; n = 10 individuals per genotype. Values represent average internode length calculated over four adjacent internodes.
In (L) to (N), letters a to d indicate maximal nonsignificant sets (MNS) to which the genotype belongs. Genotypes found in the same MNS are not significantly different from one another, whereas genotypes not found in a common MNS are significantly different from one another, at the specified significance level (α = 0.01 by nonparametric multiple comparisons by STP).
[See online article for color version of this figure.]
These results do not support a role for CHAL or its related genes as stomatal development regulators in a wild-type background. While constructing multiple CHALf mutants, however, we observed that a fraction of plants in segregating populations displayed distinctive alterations in growth. Given that ER functions in growth as well as stomatal patterning (Shpak et al., 2004, 2005) and that EPFLs were previously shown to act through the ER family (Hara et al., 2007, 2009; Hunt and Gray, 2009), we conducted a more comprehensive analysis of growth and development in single, double, and triple mutant CHALf lines. We found that plants bearing mutations in both CHAL and CLL2 (chal cll2 and chal cll1 cll2) are defective in growth, as evident from their short stature, shortened pedicels, and compact inflorescences (Figures 1G to 1K). Several CHALf mutant lines also display varying degrees of sterility, with chal cll1 cll2 producing mostly sterile siliques (Figure 1G, inset). We quantified plant height, pedicel length, and internode length in 7-week-old plants, comparing CHALf mutants to both wild-type and er-105 (null) mutants. In each of these characters, chal cll2 and chal cll1 cll2 mutants aligned with er, and both ligand and receptor mutants were statistically different from the wild type (Figures 1L to 1N). Having revealed new phenotypes for EPFL family members outside of the stomatal lineage, we measured height and pedicel and internode length in plants deficient for EPF1, EPF2, and STOMAGEN. For none of these other EPFL genes, however, did mutation or RNA interference (RNAi) knockdown produce consistent growth differences from wild-type controls (see Supplemental Figure 1 online).
The shortened pedicels and stems of chal cll2 and chal cll1 cll2 could be due to a defect in cell expansion or, like er, in cell proliferation (Shpak et al., 2004). To distinguish between these possibilities, we examined the cortex cells of pedicels using differential interference contrast (DIC) microscopy. We quantified the longitudinal length of adjacent cells in at least eight pedicels per genotype. The average longitudinal length of 10 cells from the wild type was 226.63 ± 33.5 μm; for er, 228.09 ± 41.78 μm; and for chal cll1 cll2, 198.43 ± 34.07 μm (mean ± sd). Although chal cll1 cll2 cortex cells were slightly shorter in length than the wild type (nonparametric multiple comparisons by simultaneous test procedure (STP), significant at α = 0.05 but not at α = 0.025), this difference is not sufficient to account for a reduction of pedicel length from 8.9 ± 0.88 mm (wild type) to 3.13 ± 0.39 mm (chal cll1 cll2, mean ± sd). We therefore conclude that, similar to er, the shortened pedicels of chal cll2 arise primarily through a defect in cell proliferation.
CHALf Genes Have Overlapping yet Distinct Expression Patterns
Receptor–ligand interactions require common temporal expression and overlapping (or adjacent) spatial expression. We created transcriptional reporters for CLL1 and CLL2 using 2.6 to 2.7 kb of 5′ sequence driving β-glucuronidase (GUS) and assayed GUS activity among progeny of at least four independent transformed lines alongside previously published CHAL (Abrash and Bergmann, 2010) and ERf transcriptional reporters (Shpak et al., 2004) at various developmental stages to determine the extent of their spatiotemporal overlap (Figure 2).
Figure 2.
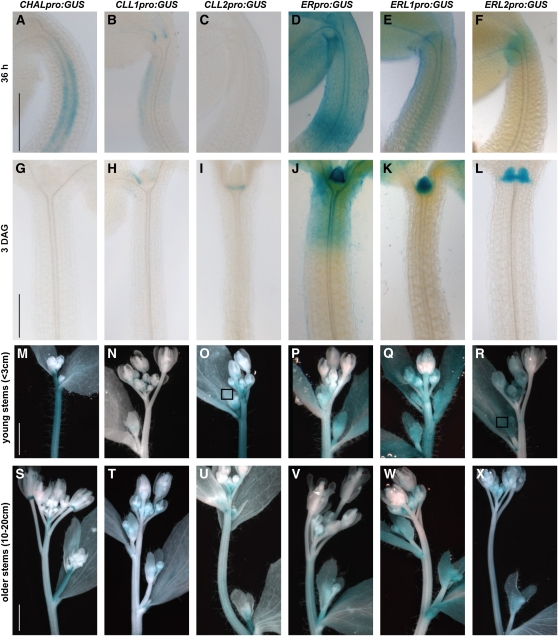
Transcriptional Reporters for CHALf and ERf Are Coexpressed in Tissues Consistent with Their Loss-of-Function Phenotypes.
All images are of reporters based on promoters of indicated genes fused to GUS (Pro:GUS) and are incubated for times described in Methods.
(A) to (F) Pro:GUS expression at 36 h after germination; seedlings are oriented to provide view of hypocotyl, cotyledon bases, and tips of first leaves and to capture the strongest expression levels. Specific reporters are indicated above each image, and each column represents the same reporter construct.
(G) to (L) Pro:GUS expression at 3 DAG.
(M) to (R) Pro:GUS expression of primary inflorescence soon after bolting (plant height < 3 cm). Boxes mark nonstaining leaf in CLL2 (O) and staining leaf in ERL2 (R).
(S) to (X) Pro:GUS expression in older primary inflorescence (plant height 10 to 20 cm).
Bars = 200 μm in (A) and (G) and 2 mm in (M) and (S). All images in a row are at the same magnification.
CHAL was initially characterized for its effects on hypocotyls, and, consistent with previous reports (Abrash and Bergmann, 2010), we found CHAL was expressed primarily in interior tissues of this organ. In this study, we observed high expression at 36 h after germination (Figure 2A) that ceased by 3 d after germination (DAG) (Figure 2G). In three independent lines, CLL1 was also observed in the hypocotyl, but its pattern was distinct from CHAL; its expression was asymmetric and observed on the side proximal to the folded cotyledons at germination (Figure 2B). CLL2 expression was not observed in the hypocotyl (Figure 2C) but was first seen at the base of the apical meristem at 3 DAG (Figure 2I). When compared with ERf expression patterns, CLL2 is coincident with ERL2 expression (Figure 2L), and CHAL and CLL1 expression in the young hypocotyl coincides with ER and ERL1 expression (Figures 2D and 2E).
Given their mutant phenotypes in plant growth, we anticipated that CHAL and CLL2 would be expressed in developing stems. We observed strong expression of both CHAL and CLL2 in developing stems soon after bolting (in plants <3 cm in height; Figures 2M and 2O). In older plants, 10 to 20 cm in height, the strongest expression was observed near the apex and in nascent branches (Figures 2S and 2U), suggesting that expression within an organ decreases as the organ ages and cells exit proliferative stages. CLL1, by contrast, was generally not expressed at high levels in stems (Figures 2N and 2T). These expression patterns are consistent with publicly available tiling array data on the CHALf genes (see Supplemental Figure 2J’ online). ERf genes (Figures 2P to 2R and 2V to 2X) were also expressed in stems and pedicels to varying degrees, with ERL2 showing the weakest expression. ERf expression noticeably overlapped with CHALf expression (Figures 2O, 2Q, 2S, and 2V).
CHALf genes are also expressed in the embryonic hypocotyl (CHAL and CLL1; see Supplemental Figures 2D and 2E online), the base of flowers (CLL1; see Supplemental Figure 2Q online), young siliques (CLL2, two lines; see Supplemental Figure 2R online), the chalazal region of ovules (CHAL and CLL1; see Supplemental Figures 2J and 2K online), and near the root apex (CHAL and CLL1; see Supplemental Figures 2V and 2W online). We confirmed earlier reports of ERf expression in embryos and ovule chalazal regions (Pillitteri et al., 2007) (see Supplemental Figures 2G and 2H online) and also observed ERf expression in the root (see Supplemental Figures 2S and 2U online). Previous studies implicated ERf in integument proliferation, defects in which lead to gametophyte abortion in erf mutants (Pillitteri et al., 2007). The strong ovule expression of CLL1 (and to a lesser extent, of CHAL) may explain the sterility observed when CLL1 is removed in combination with other CHALf genes (Figure 1G, inset). The GUS expression patterns are consistent with the CHALf genes mediating a subset of the ERf’s diverse growth functions, though notably, unlike the ERf (and other EPFLs), CHALf genes do not appear to be expressed in cotyledons or cauline leaves (Figures 2M to 2O and 2S to 2U; see Supplemental Figures 2Y to 2B’ online). Examination of stomatal precursors and guard cells indicates that, unlike ERfs, CHALf genes do not exhibit elevated stomatal lineage expression (see Supplemental Figures 2C’ to 2I’ online).
CLL1 Can Compensate for CLL2 Growth Activity
CLL1 and CLL2 are very similar in sequence, but loss-of-function and GUS expression studies suggest that these two proteins have different endogenous roles. To determine whether expression pattern or biochemical activity is the primary difference between CLL1 and CLL2, we tested whether CLL1 promotes growth when expressed under control of the CLL2 promoter (CLL2pro:CLL1). CLL2pro:CLL1 in chal cll2 qualitatively resembled the wild type (19/21 T1s), whereas a negative control (CLL2pro:GFP [for green fluorescent protein]) did not. We confirmed rescue through quantification of plant height, pedicel length, and internode length in 12 CLL2pro:CLL1 T1s (see Supplemental Figures 1A to 1C online) and three CLL2pro:CLL2 T1s (positive control; see Supplemental Table 1 online). These results indicate that both CLL1 and CLL2 have growth-promoting activity and that expressional differences likely account for the behavior of the paralogs. They further demonstrate that the CLL2 promoter fragment used in the GUS reporter constructs is sufficient for rescue of the chal cll2 growth phenotype, confirming the validity of this reporter as a proxy for CLL2 expression.
CHAL Family Overexpression Phenotypes Are ERf Dependent
The above analyses indicate that CHALf genes can function as growth regulators, likely acting through the ERf. However, the CHALf proteins are similar in sequence to stomatal regulators EPF1, EPF2, and STOMAGEN (Hara et al., 2009; Kondo et al., 2009), and CHAL itself was originally identified in a stomatal screen (Abrash and Bergmann, 2010). In overexpression assays, CHAL confers distinct, quantifiable stomatal phenotypes (Abrash and Bergmann, 2010), and Hara et al. (2009) report in an EFPL family survey that CLL1 and CLL2 can similarly repress stomatal production. We therefore used the stomatal phenotypes of overexpressors to measure CHALf activity and its mediation by the ERf genes (and TMM).
First, we confirmed that CLL1 and CLL2 can inhibit stomatal production in multiple independent transformants. We found that 35Spro:CLL1 and 35Spro:CLL2 markedly inhibit stomatal production when overexpressed in a wild-type background (Figure 3A), whereas 35Spro:CHAL T1 seedlings do not consistently differ from the wild type in their stomatal production (Figure 3A). In this regard, CLL1 and CLL2 bear some resemblance to the stomatal EPFL family members EPF1 and EPF2, which are also thought to act through the ERf.
Figure 3.
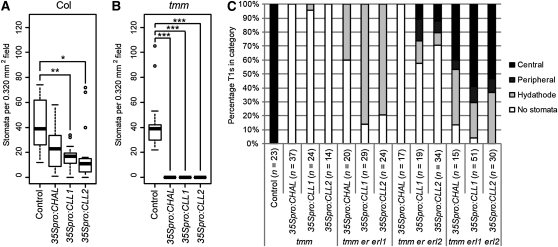
The Dependence of CHALf Overexpression Phenotypes on Receptor Genotypes Is Revealed through Stomatal Phenotype Assays.
(A) to (B) Quantification of effects of CHALf overexpression on stomatal density in wild-type and tmm adaxial cotyledons, respectively (10 DAG T1s). Control = transformation with empty vector. Bold lines indicate medians, boxes indicate quartiles above and below median, and whiskers extend to most extreme value no more than 1.5 interquartile ranges from box. n = 15 T1s per genotype in (A); n = 14 T1s per genotype in (B). *, significant at α = 0.05; **, significant at α = 0.01; ***, significant at α = 0.001 by nonparametric multiple comparisons by STP.
(C) Phenotypes of CHALf-overexpressing T1s in tmm and ERf mutant backgrounds (10 DAG). Each T1 was scored for rescue of stomatal differentiation (for details, see Methods). From least to most rescued, categories were defined as the following: no stomata = no stomata in cotyledons; hydathode = one or more stomata in hydathode region of cotyledon; peripheral = one or more stomata in cotyledon periphery; central = one or more stomata in central cotyledon. The plants scored quantitatively in (B) are a subset of those scored qualitatively in (C).
When overexpressed in a tmm background, however, CLL1 and CLL2 behave similarly to CHAL (and unlike EPF1 and EPF2). Rather than losing their activity in tmm, all three CHALf genes confer a striking reduction in stomatal density, such that cotyledons of T1 plants are typically devoid of stomata and display few or no asymmetric cell divisions (Figure 3B; see Supplemental Figures 3B, 3F, and 3J online). Thus, rather than mediating CHALf signaling, TMM actually functions to reduce or dampen this signaling.
Using the sensitized tmm background, we then conducted a series of experiments to assess whether one or more ERf members mediate CHALf activity and whether receptor specificity varies among family members. To do so, we overexpressed CLL1 and CLL2 in tmm backgrounds with reduced ERf activity (specifically, backgrounds in which two of the three ERf genes were mutated). Loss of any pair of ERf members was sufficient to noticeably mitigate the overexpression phenotypes of CLL1 and CLL2 in tmm, but the type and degree of phenotypic mitigation was dependent upon the specific receptor remaining (Figure 3C; see Supplemental Figure 3 online). In some backgrounds, only asymmetric division was extensively rescued, whereas in other backgrounds, differentiation was also considerably rescued. To represent asymmetric division phenotypes, we show images of typical cell division patterns for each genotype and present a table of observed phenotypes (see Supplemental Figure 3 online). To capture the range of differentiation phenotypes observed, we scored T1s by placing them into categories corresponding to the cotyledon regions in which they produced stomata (most to least rescued: central, peripheral, hydathode, no stomata; Figure 3C). We observed the greatest rescue of stomatal differentiation in tmm erl1 erl2 mutants (active receptor: ER), with a substantial fraction of CLL1 or CLL2 overexpressors producing stomata in the periphery or center of the cotyledon (Figure 3C; see Supplemental Figures 3E, 3I, and 3M online).
The phenotypes observed for CLL1 and CLL2 overexpression in tmm backgrounds with ERf mutations are similar to those observed for CHAL (Figure 3C; see Supplemental Figures 3B to 3E online; Abrash and Bergmann, 2010) and are consistent with all three genes exerting their effects through the ERf. Although CHAL, CLL1, and CLL2 overexpression phenotypes were not identical, there were no clear-cut differences in receptor specificity among the three ligands, and, given the variability we have observed in this assay, we do not think it can reveal subtle differences in specificity. Similarly, though our findings show that different ERf mutations modify CHALf overexpression phenotypes in distinct ways, these differences may reflect the nonidentical endogenous activities of the receptor genes (e.g., ER inhibits entry divisions, and ERL1 and ERL2 inhibit differentiation; Shpak et al., 2005) rather than differences in their capacity to be activated by the CHALf. Thus, our data suggest that CHALf members can likely act through all three members of the ERf but do not allow us to assign specific receptor-ligand pairings.
CHAL Family Mutations Affect Stomatal Production in tmm Hypocotyls
Based on overexpression assays, CHALf members appear able to activate signaling through the ERf receptors involved in stomatal production. So why doesn’t endogenous CHALf activity seem to influence stomatal development? Based on the expression patterns of CHALf reporters, a simple lack of expression can account for the absence of phenotypes in many tissues, including cotyledons. Then why aren’t clear stomatal phenotypes observed in organs with CHALf expression, such as the hypocotyl? Two basic possibilities come to mind: (1) CHALf members never reach the stomatal lineage, due to mistimed and/or subepidermal expression or (2) stomatal lineage-specific factors actively block CHALf effects. The latter possibility was of particular interest given that TMM, encoding a stomatal lineage-specific coreceptor, seemed to strongly dampen CHALf effects in overexpression assays (Figure 3).
If part of TMM’s role in the endogenous system is, in fact, to exclude CHALf signaling from the stomatal lineage, then two predictions should be fulfilled. First, reduction of CHALf activity should have little impact on stomatal patterning in a wild-type (TMM+) background because, if TMM prevents CHALf molecules from affecting the epidermis, it would simply have nothing to block when they were absent. Second, loss of TMM should enable the CHALf genes to influence stomatal production, resulting in fewer stomata where these genes are highly expressed.
To test these predictions, and to assess whether endogenous CHALf members can reach and affect the stomatal lineage, we examined CHALf mutant stomatal phenotypes in the presence and absence of TMM. We scored phenotypes in hypocotyls because the original chal phenotype was revealed in this organ (Abrash and Bergmann, 2010), CHAL and CLL1 are expressed here, and the number of stomata produced in the entire organ can be evaluated. In a wild-type (TMM+) background, chal cll1 cll2 hypocotyls resembled the wild type in stomatal number and, if anything, produced fewer stomata (perhaps due to growth alterations; Figure 4A). In a tmm background, by contrast, reduction of CHALf activity conferred a strong increase in hypocotyl stomatal production. tmm single mutants produce virtually no hypocotyl stomata, but tmm chal cll1 cll2 mutants produced routinely over 100 per hypocotyl, often arranged in large clusters (Figures 4B, 4C, and 4J). This result indicates that CHALf members can indeed reach and affect the stomatal lineage when present at their endogenous levels, in their endogenous spatiotemporal patterns.
Figure 4.
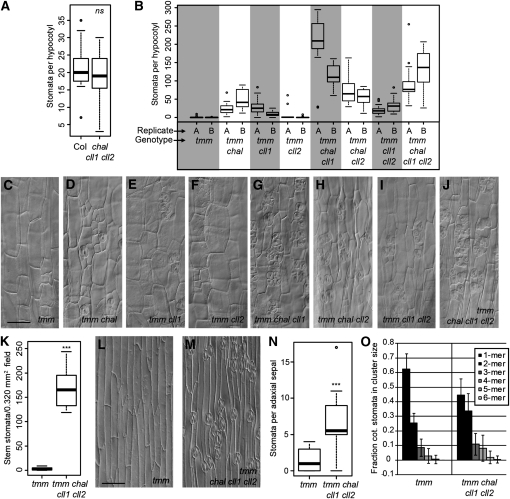
The Dependence of CHALf Loss-of-Function Phenotypes on TMM Is Revealed through Stomatal Phenotype Assays.
(A) Quantification of hypocotyl stomata in Col and chal cll1 cll2 (12 DAG, n = 20 plants per genotype). Bold lines indicate medians, boxes indicate quartiles above and below median, and whiskers extend to most extreme value no more than 1.5 interquartile ranges from box. ns, not significantly different from the wild type by Welch two-sample t test (P > 0.2).
(B) Quantification of hypocotyl stomata in CHALf mutants combined with tmm (12 DAG, n ≥ 10 plants per genotype). Each shaded or unshaded block denotes a genotype, while letters denote replicates. Plants were grown in two replicate batches A and B, each of which included all eight genotypes.
(C) to (J) DIC images of 12 DAG hypocotyls with genotypes indicated in each panel.
(K) Quantification of inflorescence stem stomatal density (n ≥ 9 plants per genotype). ***, P < 0.001 by Welch two-sample t test.
(L) and (M) DIC images of inflorescence stems with genotypes indicated in each panel.
(N) Quantification of stomata in adaxial sepals (n ≥ 10 sepals per genotype). ***P < 0.001 by Wilcoxon two-sample test.
(O) Extent of stomatal clustering in adaxial cotyledons (8 DAG, n ≥ 20 plants per genotype). Clusters are categorized by the number of stomata (“mers”) they contain; unclustered stomata are 1-mers. Shaded bars indicate mean; error bars are ± sd.
Bars = 50 μm in (C) and (L); (C) to (J) are at the same scale, and (L) and (M) are at the same scale.
Based on analysis of single and multiple CHALf mutants in tmm, CHAL and CLL1 appear primarily responsible for repression of stomatal production in tmm hypocotyls, consistent with their expression in this organ (Figures 2A, 2B, and 4B to 4J). CLL2 may have some effect, but a consistent pattern was difficult to discern due to high overall variability in phenotypic expressivity and penetrance. To capture this variability as faithfully as possible, and because no single replicate was adequately representative of our observations, we chose to show two replicates for each genotype (Figure 4B).
CHAL Family Activity Explains a Subset of tmm Phenotypes
The suite of distinct and sometimes opposing tissue-specific tmm phenotypes was recognized many years ago, but its underlying cause has been elusive (Geisler et al., 1998). Our finding that loss of TMM allows the CHALf genes to inappropriately repress stomatal production in hypocotyls provides a plausible explanation for phenotypes in this organ. Because CHALf genes are also expressed outside of hypocotyls, we asked whether CHALf activity contributes to tmm stomatal underproduction phenotypes in other organs. In stems, much as in hypocotyls, tmm chal cll1 cll2 mutants produced large numbers of stomata, dramatically more than did tmm (Figure 4K). The number of stomata in tmm chal cll1 cll2 stems was considerably greater than in the wild type (see Supplemental Figure 4 online), suggesting that loss of CHALf function does not restore wild-type stem stomatal patterning but rather causes stems to resemble other tmm organs that overproduce stomata (e.g., rosette leaves). In adaxial sepals, however, only a weak increase in stomatal density was observed upon loss of CHALf activity, such that tmm chal cll1 cll2 had increased stomatal number relative to tmm but still bore far fewer stomata than the wild type (Figure 4N; data not shown). tmm chal cll1 cll2 sepals also appeared larger than tmm sepals, potentially contributing to the observed difference.
We also examined the effects of reduced CHALf activity on an organ in which tmm plants show increased stomatal density, the adaxial cotyledon. We have not detected measureable CHALf expression in this organ (Figures 2A to 2C; see Supplemental Figures 2Z to 2B’ online), and in accord with this, did not detect a consistent difference in density between tmm and tmm chal cll1 cll2 (see Supplemental Table 2 online). We did consistently observe a shift in the size distribution of stomatal clusters in tmm chal cll1 cll2, such that the fraction of stomata contacting other stomata was higher than in tmm alone (0.377 ± 0.106 in tmm versus 0.554 ± 0.112 in tmm chal cll1 cll2, mean ± sd; P < 0.001 by Wilcoxon two-sample test; Figure 4O). As was noticed in the sepals, in general, the tmm lines appeared less healthy than the tmm chal cll1 cll2 lines, and this could indirectly affect some measurements. Ultimately, it appears that CHALf activity can largely explain loss of stomata from tmm stems and may contribute to stomatal reduction phenotypes in sepals but, as expected from expression patterns, does not contribute substantially to phenotypes in cotyledons.
Testing a Model for CHAL Family and EPF1/2 Interactions with TMM
Our loss-of-function analyses suggest a model in which TMM expression prevents stomatal lineage cells from perceiving growth-regulatory CHALf signals. At the same time, TMM also permits or enhances perception of EPF1 and EPF2 signals, which are produced in stomatal precursors and enforce stomatal pattern (Hara et al., 2007, 2009; Hunt and Gray, 2009). The relationship of TMM to these two classes of genes, together with the expression of TMM exclusively in the stomatal lineage, might permit efficient segregation of growth- and stomatal-regulatory signaling despite the spatial overlap of the respective ligands. In this model, two key factors (interactions with TMM and expression patterns) distinguish growth- and stomatal-regulatory ligand classes. If these factors were reversed, could these ligands substitute for one another? We addressed this question by rewiring the stomatal system to use growth-regulatory ligands. To do so, we removed TMM from the epidermis, expressed CHAL under an EPF1 promoter (EPF1pro:CHAL), and assayed CHAL’s ability to perform EPF1’s endogenous role of enforcing stomatal spacing.
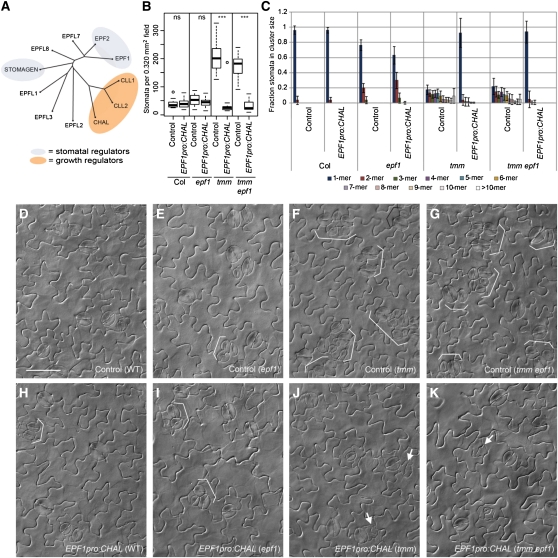
We transformed EPF1pro:CHAL into wild-type, epf1, tmm, and tmm epf1 backgrounds. As previously reported, the EPF1 promoter drives strong yellow fluorescent protein expression in stomatal precursors and young stomata (Hara et al., 2007; see Supplemental Figure 5 online) accompanied by weak, ubiquitous fluorescence (see Supplemental Figure 5 online). As predicted by our model, EPF1pro:CHAL did not decrease stomatal density or reduce stomatal clustering in backgrounds with functional TMM (wild type Columbia [Col] and epf1; Figures 5B and 5C, and 5H and 5I versus 5D and 5E). In backgrounds lacking TMM activity (tmm and tmm epf1), however, EPF1pro:CHAL conferred striking alterations in stomatal density and spacing (Figures 5B and 5C, and 5J and 5K versus 5F and 5G). Median stomatal density was reduced to roughly wild-type levels, and the clustering profile was strongly shifted toward single stomata and small clusters (Figures 5B and 5C). Plants displaying a high degree of rescue looked similar to the wild type in their overall pattern of differentiated stomata, although small cells likely representing arrested, transdifferentiated stomatal precursors were sometimes interspersed with the stomata (Figures 5J and 5K, white arrows). These results suggest that CHAL, when expressed in stomatal precursors and in the absence of a TMM buffer, can indeed function to enforce stomatal one-cell spacing.
Figure 5.
CHAL Can Enforce One-Cell Spacing When Expressed under the EPF1 Promoter in a tmm Background.
(A) Neighbor-joining tree depicting relationships between EPFL family proteins (modified from Rowe and Bergmann, 2010). Shading indicates proteins of known function.
(B) Quantification of the effects of EPF1pro:CHAL on stomatal density (versus empty vector) in 10 DAG abaxial cotyledons. Bold lines indicate medians, boxes indicate quartiles above and below median, and whiskers extend to most extreme value no more than 1.5 interquartile ranges from box. n = 15 T1s for each line. ns, not significant (P > 0.30 by Wilcoxon two-sample test); ***, P < 0.001 by Wilcoxon two-sample test.
(C) Quantification of the effects of EPF1pro:CHAL on stomatal clustering (versus empty vector) in 10 DAG abaxial cotyledons. Shaded bars indicate mean; error bars are ± sd.
(D) to (K) Representative DIC images of EPF1pro:CHAL and empty vector T1s. Genotypes and transgenes are indicated on each panel. White brackets indicate clustered stomata; white arrows indicate apparent transdifferentiated cells observed in strongly rescued lines.
Bar = 50 μm in (D); (D) to (K) are at the same scale.
[See online article for color version of this figure.]
DISCUSSION
Reuse of key signaling cascades in diverse processes and contexts is a common theme in the development of multicellular organisms. When multiple processes regulated by a single set of receptors occur in close spatiotemporal proximity, signaling specificity requires mechanisms that prevent spillover of activation from one process into another. The ERf receptors function in a diverse range of developmental and physiological processes in Arabidopsis, including growth regulation and patterning of stomata. Here, we identified three EPFL family members, CHAL, CLL1, and CLL2, as probable ERf agonists in regulation of growth. Furthermore, we presented evidence that TMM, which encodes a coreceptor expressed specifically in the stomatal lineage, acts to block CHALf effects on stomatal development, enabling segregation of the spatiotemporally overlapping, ERf-regulated processes of growth and stomatal patterning. While we cannot completely exclude the possibility that CHALf members act in parallel to the ERf pathway, the hypothesis that CHALf members act as growth-regulatory ligands for ERf receptors provides a simple, unifying explanation for the mutant phenotypes and genetic and physical interactions we observed.
Having established that CHALf members can function as growth regulators in pedicels and stems, we must reevaluate the relationship of these genes to stomatal development networks. Single or higher order mutations in CHALf members can generate strong stomatal phenotypes in tmm hypocotyls but fail to confer such phenotypes in wild-type plants, indicating that TMM functions endogenously to dampen or block CHALf signaling (in addition to transducing appropriate stomatal-regulatory signals; see below). As TMM expression is specific to, but widespread within, the stomatal lineage, such a mechanism would effectively prevent CHALf signals from affecting stomatal development. This in itself is an elegant solution to the problem of specificity. By producing a regulatory cofactor (TMM) in a spatially restricted domain (the stomatal lineage), the plant prevents ligands appropriate to one developmental process (growth) from interfering with a spatiotemporally overlapping process (stomatal development) that shares the same receptors.
When previous work on stomatal-regulatory EPFL family members is considered, TMM emerges as an even more ingenious and economical innovation. Not only does this coreceptor dampen growth-regulatory CHALf signals, but it also potentiates (and, indeed, is largely necessary for) transduction of stomatal-regulatory EPF1, EPF2, and STOMAGEN signals (Hara et al., 2007, 2009; Hunt and Gray, 2009; Kondo et al., 2009; Sugano et al., 2009). Thus, just as growth-regulatory signals will not aberrantly affect epidermal development because TMM is in the stomatal lineage, so stomatal-regulatory signals will not aberrantly affect growth because TMM is absent from the internal tissues. The relationship of TMM to these two classes of ligands, together with the expression of TMM exclusively in the stomatal lineage, could thus permit the efficient segregation of growth- and stomatal-regulatory signaling despite the spatial overlap of the respective ligands (Figure 6).
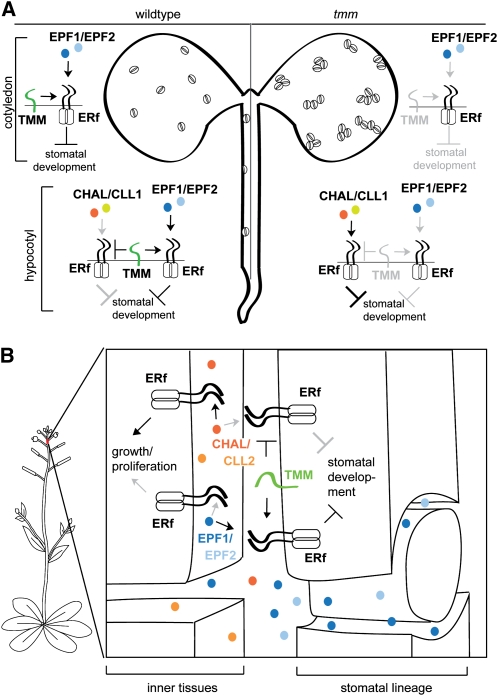
Figure 6.
Models in Which CHALf Acting through ERf Receptors in a Pathway Inhibited by TMM Can Explain Observed Phenotypes.
(A) Model for CHALf and EPF1/2 interactions with TMM and ERf receptors in different regions of wild-type and tmm seedlings. Arrows represent positive influences on signaling or developmental processes; T-bars represent negative influences. Grayed-out elements are inactive or not present in a particular organ or genotype. The positive regulator of stomatal development EPFL9/STOMAGEN also requires TMM for activity, but for simplicity, is not shown here. The left half of the seedling depicts the wild type (TMM+). In both the cotyledon and hypocotyl, stomatal lineage ligands EPF1/2 signal through ERf and TMM receptors to inhibit stomatal development for proper patterning. CHALf ligands in the hypocotyl do not influence stomatal patterning due to the presence of TMM, which dampens CHALf signaling. The right half of the seedling depicts the tmm mutant: Without TMM, EPF1/2 do not inhibit stomatal development, leading to the formation of stomatal clusters in cotyledons. The opposite effect is observed in the tmm hypocotyl, however, due to the presence of CHALf ligands in this organ. Without TMM’s dampening effect, CHALf ligands overactivate ERf receptors, resulting in a strong inhibition of stomatal development.
(B) Model for TMM compartmentalization of ERf receptor functions in growth and stomatal development in stems. Apoplast-mobile CHALf and EPF1/2 ligands can encounter ERf receptors both in the stomatal lineage and in nonstomatal cells, such as the inner tissues. TMM, which is present only in the stomatal lineage, differentially regulates these two ligand classes. Thus, CHALf signaling does not affect stomatal development because it is dampened by TMM, while EPF1/2 signaling does not affect growth because it is potentiated by TMM. Arrows represent positive influences on signaling pathways or developmental processes; T-bars represent negative influences. Grayed-out elements represent inactive pathways.
Among animal signaling pathways, it is possible to find examples in which cofactors modulate ligand activity in a manner similar to TMM. For instance, the glycosylphosphatidylinositol-anchored protein Cripto plays a seemingly TMM-like role in the interaction between the activin type II receptor and two classes of ligands in the TGF-β superfamily: it is required for Nodal-class ligands to signal through this receptor but impedes the ability of activin-class ligands to do the same (Kirkbride et al., 2005; Kelber et al., 2008). As these relationships have been worked out largely in tissue culture, it would be interesting to examine whether Cripto has an in vivo role similar to that of TMM (i.e., whether it functions to segregate activin II–dependent signaling pathways that operate in close proximity).
Our findings that characterized EPFL family members cluster into process-related subgroups raises the question of what roles the remaining EPFL genes (Figure 5A) might play. The five uncharacterized EPFL family members could encode stomatal regulators, growth regulators, or ligands active in some other ERf-dependent process, such as bacterial or fungal pathogen resistance (Godiard et al., 2003; Llorente et al., 2005). In neither growth nor stomatal development does loss of the currently characterized EPFLs fully recapitulate the suite of defects generated by loss of the ERf. chal cll1 cll2 triple mutants do not have the blunted siliques and altered leaf morphology characteristic of er mutants and are clearly not as dwarfed as er erl1 erl2. This suggests that additional EPFL genes may regulate growth and may do so redundantly with the CHALf. In support of this possibility, we found that two more EPFL family members, EPFL1 (At5g10310) and EPFL2 (At4g37810), display expression patterns roughly similar to the CHALf in publicly available tiling array data (see Supplemental Figure 2K' online; Laubinger et al., 2008) and display CHAL-like behaviors in overexpression assays: No obvious stomatal phenotypes were reported for these genes in a wild-type background (Hara et al., 2009), but we find that they noticeably reduce stomatal density in tmm (see Supplemental Figure 6 online). Activity of these genes might also account for tmm phenotypes not explained by CHALf activity, such as loss of stomata from the adaxial sepal.
Activity of additional EFPLs could also contribute to the substantial phenotypic variability seen for CHALf mutants in a tmm background. Despite care in standardizing growth conditions and seedling selection, we observed considerable variability in stomatal phenotypes among seedlings of the same genotype (Figure 4B). Such variability could be explained if expression of CHALf members (and of spatially overlapping, TMM-inhibited EPFLs) were constrained within a range appropriate to growth regulation but not conducive to consistent levels of stomatal production in the absence of TMM. This hypothesis might also account for the basal phenotypic variability observed in tmm single mutants. Alternatively, this variability might reflect the capacity of ERf receptors and/or their downstream mitogen-activated protein kinase cascade to transduce environmental cues and stresses (Godiard et al., 2003; Llorente et al., 2005; Nakagami et al., 2005), perhaps coupled with enhanced sensitivity to such cues and stresses in the absence of TMM. Independent of its cause, this variability must be considered when evaluating published experiments and designing new ones with EPFL genes in a tmm background.
Characterization of the diverse functions of EPFL family members represents one challenge for the future. Another such challenge will be to clarify the biochemical interactions among EPFL family ligands, ERf receptors, and TMM and to identify downstream outputs to measure the signaling capacities of specific ligand and receptor complexes. Genetic and phenotypic evidence suggests that EPFLs act through the ERf; we would therefore expect the ligands to physically interact with the extracellular domains of ERf receptors. TMM’s dampening effect on CHALf signaling, however, could arise via several different mechanisms. In one scenario, TMM might bind CHALf members directly, either titrating them away from the ERf or engaging them in unproductive ERf/TMM/CHALf complexes. Alternatively, TMM might not bind CHALf ligands directly, but instead tie up ERf receptors in TMM/ERf complexes incapable of CHALf binding. Rigorous demonstration of the mode of TMM activity will require purification of endogenous complexes from relevant tissues and awaits future studies. Binding of CHALf members to specific ER and TMM extracellular fragments can be observed in yeast two-hybrid assays (see Supplemental Figure 7 online), consistent with models based on ligand titration or formation of unproductive ERf/TMM/CHALf complexes. In stomatal lineage cells, the abundance of receptors and their differential affinities for ligands may further constrain system behavior. Modeling approaches such as those recently used to evaluate alternative models for shoot meristem LRR-RLK–mediated signaling (Sahlin et al., 2011) will also be important hypothesis testing tools, especially as the number of potential ligand and receptor players increases beyond what can be easily tested empirically.
The function of TMM as a pathway buffering/signaling specificity element represents, thus far, a unique role for an RLP in plant LRR-RLK signaling. What is the potential for this to be a widespread phenomenon? Among the ~220 Arabidopsis LRR-RLKs (Shiu and Bleecker, 2001) and 57 LRR-RLPs (Wang et al., 2008), only two developmental partnerships have been described: that of the ERf and TMM in growth and stomatal development and that of RLK CLAVATA1 (CLV1) and RLP CLV2 in shoot meristem maintenance. The latter appears to follow fundamentally different rules than the former: CLV2 likely forms an active signaling complex with a unique intracellular kinase (CORYNE; Muller et al., 2008), may signal largely in parallel to CLV1 (Nimchuk et al., 2011), and is expressed in a broader domain than is CLV1. Thus, it is unlikely that RLPs as a category are dominated by TMM-like behaviors. Rather, such coreceptor behaviors might be expected in cases where: (1) a core set of receptors is used in multiple contexts during the life of the organism, (2) the processes regulated by the receptors occur in close spatiotemporal proximity, and (3) ligands are mobile, such that those intended to act in one process could bleed over into another. It will be intriguing to see if other systems with these characteristics indeed have TMM-like mechanisms for ligand discrimination and, if not, even more intriguing to see what other paths evolution has found to achieve compartmentalized signaling in close quarters.
METHODS
Plant Materials and Growth Conditions
Col-0 or Col-0 containing the E1728 guard cell GFP marker (Gardner et al., 2009) was used as the wild type. Alleles used were cll1-1 (SALK_005080), cll2-1 (SALK_071065), chal-2 (Abrash and Bergmann, 2010), er-105, erl1-2, and erl2-1 (Shpak et al., 2004), tmm-1 (Nadeau and Sack, 2002), epf1-1 (Hara et al., 2007), and epf2-1 (Hunt and Gray, 2009). The left border junction of the cll1 T-DNA has sequence 5′-AATTTGATTACTCTTTGACTTTGGACCACT-3′, while the left border junction of the cll2 T-DNA has sequence 5′-TCAATTTGTTAGTAATAAGCGGTTTGGTGG-3′ (underline indicates T-DNA; plain text indicates genomic DNA). With the exception of er-105, the allele number is dropped when these mutations are referred to in the text. tmm er erl1, tmm er erl2, and tmm erl1 erl2 mutants (Shpak et al., 2005) were gifts of K.U. Torii. tmm er erl2 is also glabrous. epf1 epf2 was a gift of J.E. Gray. STOMAGEN RNAi lines (Sugano et al., 2009) were a gift of I. Hara-Nishimura. tmm epf1, multiple CHALf mutants, and CHALf mutants in tmm backgrounds were generated by crossing and isolated based on phenotypes and/or PCR genotyping (see below). tmm, tmm epf1, and some CHALf mutant lines in tmm backgrounds contain the E1728 guard cell GFP marker. ERpro:GUS, ERL1pro:GUS, and ERL2pro:GUS lines (Shpak et al., 2004) were gifts of K.U. Torii. CHALpro:GUS lines have been previously described (Abrash and Bergmann, 2010) and also contain the E1728 guard cell GFP marker. Seeds were stratified at 4°C for ≥2 d. Seedlings were grown on 0.5 Murashige and Skoog (MS) medium (Murashige and Skoog, 1962), 1% agar plates at 22°C under constant light in an incubator (I36LL; Percival). For later analyses, seedlings were transferred to soil and maintained in a growth chamber (AR66; Percival) at 22 ± 1°C with a 16-h-light/8-h-dark cycle.
Genotyping
Representative, functional primer pairs for genotyping (a subset of those used over the course of the experiment) and sequences of gene-specific primers are provided in Supplemental Table 3 online. T-DNA internal primers LBa1 and LBb1.3 (Alonso et al., 2003; http://signal.salk.edu) and JL-202 (Young et al., 2001) have been previously described.
RT-PCR
For analysis of cll1 and cll2, RNA was extracted from ~13 DAG whole seedlings using the RNeasy plant mini kit (Qiagen) with on-column DNase digestion. cDNA was synthesized with oligo(dT) primers using the Superscript III first-strand cDNA synthesis kit (Invitrogen), followed by amplification using Accuprime Pfx DNA polymerase (Invitrogen) and gene-specific primers (see Supplemental Table 3 online) with program 95°C for 2:00, 35 cycles of 95°C for 0:15, 54°C (for CLL2 and ACT; 58°C for CLL1) for 0:30, 68°C for 1:00, and final extension of 68°C for 5:00. Bands were visualized via ethidium bromide staining.
Plant Vector Construction and Transformation
Most manipulations involved Gateway-mediated cloning following the manufacturer’s instructions. Primers used for each step are given in Supplemental Table 3 online. For 35Spro:CLL1, the CLL1 coding sequence was amplified from clone U63004 (ABRC), cloned into pENTR/dTOPO (Invitrogen), and recombined into pH35GS (Kubo et al., 2005) using LR Clonase II (Invitrogen). For 35Spro:CLL2, cDNA clone DQ487600 (ABRC) was recombined into pH35GS. For 35Spro:EPFL1, clone GC63002 (ABRC) was recombined into pH35GS. For 35Spro:EPFL2, the EPFL2 coding sequence was cloned into pENTR and recombined into pH35GS. 35Spro:CHAL was described (Abrash and Bergmann, 2010). The empty vector control for overexpression assays, pH35S, was generated from pH35GS by XbaI/XhoI excision of the Gateway followed by blunting and self-ligation. For CLL1pro:GUS and CLL2pro:GUS, 2.6- to 2.7-kb 5′ regions were amplified from BAC MIWI23 (ABRC) for CLL1 and BAC T2D4 (ABRC) for CLL2. Fragments were cloned into pENTR/dTOPO and recombined into pBGGUS (Kubo et al., 2005). For EPF1pro:YFP, the promoter region defined by Hara et al. (2007) was cloned into pENTR/dTOPO and recombined into pHGY (Kubo et al., 2005). For EPF1pro:CHAL, the same EPF1 promoter was inserted in the NotI site of pENTR-CHAL (generated by amplifying the CHAL cDNA out of clone DQ446581 [ABRC] and cloning into pENTR/dTOPO) and the EPF1pro:CHAL fusion recombined into pHGS (a pH35GS derivative generated by HindIII/XbaI excision of 35Spro followed by blunting and self-ligation). The empty vector control for the EPF1pro assay, pHS, was made from pH35GS by HindIII/XhoI excision of 35Spro and the Gateway followed by blunting and self-ligation. For CLL2pro:CLL2 and CLL2pro:CLL1, CLL2pro was excised from pENTR/dTOPO using flanking NotI sites. CLL2 was amplified and cloned into pENTR/dTOPO. CLL1 and CLL2 in pENTR/dTOPO were NotI digested, and CLL2pro was inserted into the 5′ NotI site. Constructs were then recombined into pGWB1 (Nakagawa et al., 2007). CLL2pro:GFP was generated by recombining CLL2pro pENTR/dTOPO into pGWB4 (Nakagawa et al., 2007).
Stably transgenic plants were generated by Agrobacterium tumefaciens–mediated transformation using the floral dip protocol (Clough and Bent, 1998). Transgenic seedlings were selected either by growth on 1% agar + 0.5 MS plates supplemented with 50 μg/mL hygromycin (pH35GS, pHGY, pGWB1, and pGWB4 based constructs) or in soil treated with Basta (Finale 1:2000 dilution; pBGGUS-based constructs).
Analysis of Stomatal Phenotypes
Preparation of Tissue
Multiple plant tissues were assayed for stomatal production. For hypocotyls and cotyledons, plants were grown on 0.5× MS, 1% agar plates until time of collection (noted for each experiment in the main text). For analysis of inflorescence stem stomata, seedlings were moved from plates to soil at 10 DAG (Figure 4K) or 11 DAG (see Supplemental Figure 4 online) and the third most basal internode of the main stem was scored at 37 DAG (Figure 4K) or 46 DAG (see Supplemental Figure 4 online). For sepals, seedlings were moved from plates to soil at 11 DAG and scored at 46 DAG. Sepals were collected from the oldest pedicel on the main stem that still bore nonsenescent sepals.
Fixation and Mounting
Tissue samples were processed by one of three methods, depending on tissue type and age: (1) Samples were collected into 70% ethanol, cleared overnight at room temperature, and stored at 4°C as needed and then rinsed in water and mounted in Hoyer’s medium (Liu and Meinke, 1998). For stem samples, an epidermal peel was taken from the cleared sample following the water rinse. (2) Samples were cleared and stored in 7:1 ethanol:acetic acid at room temperature, treated with 1 n potassium hydroxide (30 min to overnight), rinsed in water, and mounted in Hoyer’s medium. (3) Fresh tissue was mounted directly in Hoyer’s medium and placed in a vacuum chamber for ~1 week or imaged directly if epidermal peel. The second method was typically most effective in preserving cell morphology.
Analysis and Scoring
Most images of stomatal phenotypes were obtained from samples mounted in Hoyer’s medium and visualized using DIC microscopy on a Leica DM2500 microscope. For quantitative analysis of hypocotyl and adaxial sepal stomatal number, scoring was performed on the microscope, and results reflect analysis of the entire organ or tissue. For analysis of cotyledon stomatal density and clustering, and of inflorescence stem stomatal density, images were taken at ×20 magnification (0.320-mm2 field), and stomata and/or clusters were counted manually from images using the Cell Counter in ImageJ (NIH). Only those stomata/clusters contained fully within the frame of the image were counted. For cotyledons, images were taken in the central region overlying the distal vascular loop. If a cotyledon was too small to fill the ×20 field, it was positioned so as to best fill the field and the stomata within the frame were counted. For inflorescence stems, images were randomly positioned within the regions of sample that were wide enough to fill the camera field. Except in cases where differences between genotypes were very obvious, images were randomized before scoring and scored blind.
For qualitative (category) scoring of CHALf overexpressing T1s in tmm and ERf mutant backgrounds, the adaxial and abaxial sides of all cotyledons present on the plant were examined for stomata, and each plant was placed into the “most rescued” category to which it belonged (most to least rescued: central stomata, peripheral stomata, hydathode stomata, no stomata). A single stoma in a given category was sufficient to place a plant into that category. Stomata were scored as “central” if contained within the vascular loops, “peripheral” if outside the vascular loops but not in the hydathode, and “hydathode” if present at the distal tip of the leaf, in or associated with the hydathode. Plants with stomata placing them in a higher category (e.g., central) often, but not always, also possessed stomata corresponding to lower categories (e.g., hydathode). One to three batches of a given genotype + construct combination were scored and pooled to give the final data presented.
Except in analyses of CHALf overexpressing T1s, which were often unhealthy due to lack of stomata, only seedlings that met a minimum health cutoff (based primarily on the presence and size of true leaves or on the size of cotyledons) were included in assays.
Analysis of Growth Phenotypes
Plant growth was quantified at ~7 weeks after germination. Plants were grown on 0.5× MS, 1% agar plates for 2 weeks and then transplanted to soil. Height measurement was recorded as the distance from rosette to the flower tips of the longest inflorescence. For chalf lines, the length of five pedicels per plant (pedicels 3 to 7 from the base of the primary inflorescence) was measured. For STOMAGEN RNAi lines, epf1 epf2, and rescue/complementation experiments (CLL2pro:CLL1, CLL2pro:GFP, and CLL2pro:CLL2 in chal cll2), the length of two pedicels per plant (pedicels 6 and 7 from the base of the primary inflorescence) was measured. For internodes, the distance between pedicels 3 and 7 for each plant was recorded and then divided by 4 to determine a single average internode length per plant. For chalf analysis, 10 plants were analyzed per genotype for all measurements. For epf1 epf2 and STOMAGEN RNAi analysis, nine plants were analyzed per genotype/independent line for all measurements (except STOMAGEN RNAi pedicels; eight plants per line were examined). MUTEpro:nucGFP (MacAlister et al., 2007) in Col was used as a BASTA-resistant control to compare with STOMAGEN RNAi lines. For rescue of chal cll2 by CLL2pro:CLL1 (and the CLL2pro:GFP control), 12 independent T1s were analyzed for height, and 11 independent T1s were analyzed for pedicel and internode length. For complementation of chal cll2 by CLL2pro:CLL2, three independent T1s were analyzed for each measurement.
To analyze cortex cell length, pedicels were analyzed at 6 to 7 weeks using DIC at ×40 on a Leica DM2500 microscope by focusing on the subepidermal cortex plane. The length of a longitudinal cell file (10 cells per cell file for data shown) was measured using ImageJ software (NIH). Two measurements were taken per pedicel, and two pedicels of four plants per genotype were analyzed, for a total of 16 measurements per genotype. Two replicates were examined.
Statistical Analysis
Most statistical analyses were performed in R (http://www.r-project.org/). For statistical analyses, each data point was treated as a separate observation, even when multiple data points were obtained from a single plant (e.g., in measurement of pedicel lengths). Data were checked for normality using the Shapiro-Wilk test [shapiro.test() function]. If all samples in a given data set were normally distributed, parametric analyses based on the normal distribution were used. For comparison of two samples, the Welch two-sample t test [t.test() function, unpaired, two-tailed] was used. If not all samples in a given set were normally distributed [or, in the case of multiple comparisons, if samples were normally distributed but differed significantly in variance by Bartlett’s test using the bartlett.test() function], nonparametric analyses were used instead. For comparison of two samples, the Wilcoxon two-sample test [wilcox.test() function, unpaired, two-tailed] was used. The reported P value is that returned by the function, but it should be noted that this is not an exact P value when ties are present (as they typically were). For multiple comparisons, nonparametric multiple comparisons by STP were performed, as described by Sokal and Rohlf (1995). Additional tables of critical values of the studentized range were obtained from Harter (1960). Because nonparametric multiple comparisons by STP require equal sample sizes, samples were randomized and then truncated to the size of the smallest sample included in the analysis. As no R function is available for nonparametric multiple comparisons by STP, a simple Java program was written to calculate the test statistics, while cutoff values were calculated by hand. Test statistics were not corrected for between-sample ties as the correction is typically small and uncorrected values are conservative. α Values indicate the experiment-wise error rate at which the comparison is significant. When maximal nonsignificant sets were reported, they were determined according to the definition of Rafter et al. (2003). Throughout, the statistical method used for each comparison is noted in the caption or text accompanying the data.
Boxplots were generated in R using the boxplot() function. The boxplot is a standard graphical display format (Sokal and Rohlf, 1995) in which a central horizontal line indicates the median, boxes above and below the line indicate quartiles above and below the median, and whiskers extend to the most extreme value no more than 1.5 interquartile ranges from the box. Observations outside the whiskers (sometimes called outliers) are depicted individually as circles. We treated these points as outliers for purposes of graphical display only and did not exclude them in statistical tests of significance.
Statistical comparisons between different CHALf mutant genotypes in a tmm background were not performed for hypocotyl data due to the high degree of within-genotype variation (as illustrated by the presentation of data in Figure 4B).
Yeast Two-Hybrid Assays
Yeast two-hybrid analysis was performed with the matchmaker Two-Hybrid System 3 (Clontech) using plasmids and conditions specified by the manufacturer. cDNAs corresponding to ligands were PCR amplified and cloned into pGBKT7 and receptor fragments into pGADT7 (primers in Supplemental Table 3 online). TPD1 cDNA was a gift of Dazhong Zhao (Jia et al., 2008). Pairwise interactions were conducted in triplicate at 1:100 and 1:1000 dilutions after incubation for 2, 3, and 4 d. Interactions were evaluated as positive if significant growth (~50% saturation) was observed by day 3.
Accession Numbers
The Arabidopsis Genome Initiative identifiers for the genes mentioned in this article are as follows: CHAL/EPFL6 (At2g30370), CLL1/EPFL5 (At3g22820), CLL2/EPFL4 (At4g14723), EPF1 (At2g20875), EPF2 (At1g34245), EPFL1 (At5g10310), EPFL2 (At4g37810), STOMAGEN (At4g12970), ER (At2g26330), ERL1 (At5g62230), ERL2 (At5g07180), TMM (At1g80080), TPD1 (At4g24972), and MUTE (At3g06120).
Supplemental Data
The following materials are available in the online version of this article.
Supplemental Figure 1. Redundancy and Range of Nonstomatal Phenotypes within the EPFL Family.
Supplemental Figure 2. Pro:GUS and Tiling Array Expression of the CHALf and ERf in Additional Tissues.
Supplemental Figure 3. Qualitative Phenotypes of CHALf Overexpression in Receptor Mutant Backgrounds.
Supplemental Figure 4. Stem Stomatal Density in tmm, tmm chal cll1 cll2, and Wild-Type Plants.
Supplemental Figure 5. Expression Pattern of EPF1pro:YFP.
Supplemental Figure 6. Overexpression Phenotypes of Additional EPFL Family Members in tmm.
Supplemental Figure 7. Yeast Two-Hybrid Analysis of Interactions between EPFL Ligands and Receptor Fragments.
Supplemental Table 1. CLL2pro:CLL2 Complementation of chal cll2.
Supplemental Table 2. Stomatal Density in tmm and tmm chal cll1 cll2 Adaxial Cotyledons.
Supplemental Table 3. Primers Used in This Work.
Supplementary Material
Acknowledgments
We thank D. Zhao, I. Hara-Nishimura, K. Torii, and J. Gray for reagents, Akhila Bettadapur for lab assistance, E. Shpak for insights, and members of our lab for discussions and critical reading of the manuscript. This work was supported by a Stanford Terman fellowship, by National Science Foundation Grant IOS-0844521, and by a Stanford Vice Provost for Undergraduate Education Major Grant to E.B.A. K.A.D. and E.B.A. are National Science Foundation Graduate Research Fellows.
AUTHOR CONTRIBUTIONS
E.B.A., K.A.D., and D.C.B. designed research and wrote the article. E.B.A. and K.A.D. performed research and analyzed data.
References
- Abrash E.B., Bergmann D.C. (2010). Regional specification of stomatal production by the putative ligand CHALLAH. Development 137: 447–455 [DOI] [PubMed] [Google Scholar]
- Alonso J.M., et al. (2003). Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301: 653–657 [DOI] [PubMed] [Google Scholar]
- Bergmann D.C., Sack F.D. (2007). Stomatal development. Annu. Rev. Plant Biol. 58: 163–181 [DOI] [PubMed] [Google Scholar]
- Bhave N.S., Veley K.M., Nadeau J.A., Lucas J.R., Bhave S.L., Sack F.D. (2009). TOO MANY MOUTHS promotes cell fate progression in stomatal development of Arabidopsis stems. Planta 229: 357–367 [DOI] [PubMed] [Google Scholar]
- Butenko M.A., Vie A.K., Brembu T., Aalen R.B., Bones A.M. (2009). Plant peptides in signalling: Looking for new partners. Trends Plant Sci. 14: 255–263 [DOI] [PubMed] [Google Scholar]
- Chenna R., Sugawara H., Koike T., Lopez R., Gibson T.J., Higgins D.G., Thompson J.D. (2003). Multiple sequence alignment with the Clustal series of programs. Nucleic Acids Res. 31: 3497–3500 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clough S.J., Bent A.F. (1998). Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16: 735–743 [DOI] [PubMed] [Google Scholar]
- Gardner M.J., Baker A.J., Assie J.M., Poethig R.S., Haseloff J.P., Webb A.A. (2009). GAL4 GFP enhancer trap lines for analysis of stomatal guard cell development and gene expression. J. Exp. Bot. 60: 213–226 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Geisler M., Nadeau J., Sack F.D. (2000). Oriented asymmetric divisions that generate the stomatal spacing pattern in Arabidopsis are disrupted by the too many mouths mutation. Plant Cell 12: 2075–2086 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Geisler M., Yang M., Sack F.D. (1998). Divergent regulation of stomatal initiation and patterning in organ and suborgan regions of the Arabidopsis mutants too many mouths and four lips. Planta 205: 522–530 [DOI] [PubMed] [Google Scholar]
- Godiard L., Sauviac L., Torii K.U., Grenon O., Mangin B., Grimsley N.H., Marco Y. (2003). ERECTA, an LRR receptor-like kinase protein controlling development pleiotropically affects resistance to bacterial wilt. Plant J. 36: 353–365 [DOI] [PubMed] [Google Scholar]
- Hara K., Kajita R., Torii K.U., Bergmann D.C., Kakimoto T. (2007). The secretory peptide gene EPF1 enforces the stomatal one-cell-spacing rule. Genes Dev. 21: 1720–1725 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hara K., Yokoo T., Kajita R., Onishi T., Yahata S., Peterson K.M., Torii K.U., Kakimoto T. (2009). Epidermal cell density is autoregulated via a secretory peptide, EPIDERMAL PATTERNING FACTOR 2 in Arabidopsis leaves. Plant Cell Physiol. 50: 1019–1031 [DOI] [PubMed] [Google Scholar]
- Harter H.L. (1960). Tables of range and studentized range. Ann. Math. Statist. 4: 1123–1147 [Google Scholar]
- Hunt L., Gray J.E. (2009). The signaling peptide EPF2 controls asymmetric cell divisions during stomatal development. Curr. Biol. 19: 864–869 [DOI] [PubMed] [Google Scholar]
- Jia G., Liu X., Owen H.A., Zhao D. (2008). Signaling of cell fate determination by the TPD1 small protein and EMS1 receptor kinase. Proc. Natl. Acad. Sci. USA 105: 2220–2225 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jun J., Fiume E., Roeder A.H., Meng L., Sharma V.K., Osmont K.S., Baker C., Ha C.M., Meyerowitz E.M., Feldman L.J., Fletcher J.C. (2010). Comprehensive analysis of CLE polypeptide signaling gene expression and overexpression activity in Arabidopsis. Plant Physiol. 154: 1721–1736 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kelber J.A., Shani G., Booker E.C., Vale W.W., Gray P.C. (2008). Cripto is a noncompetitive activin antagonist that forms analogous signaling complexes with activin and nodal. J. Biol. Chem. 283: 4490–4500 [DOI] [PubMed] [Google Scholar]
- Kirkbride K.C., Ray B.N., Blobe G.C. (2005). Cell-surface co-receptors: Emerging roles in signaling and human disease. Trends Biochem. Sci. 30: 611–621 [DOI] [PubMed] [Google Scholar]
- Kondo T., Kajita R., Miyazaki A., Hokoyama M., Nakamura-Miura T., Mizuno S., Masuda Y., Irie K., Tanaka Y., Takada S., Kakimoto T., Sakagami Y. (2009). Stomatal density is controlled by a mesophyll-derived signaling molecule. Plant Cell Physiol. 51: 1–8 [DOI] [PubMed] [Google Scholar]
- Kubo M., Udagawa M., Nishikubo N., Horiguchi G., Yamaguchi M., Ito J., Mimura T., Fukuda H., Demura T. (2005). Transcription switches for protoxylem and metaxylem vessel formation. Genes Dev. 19: 1855–1860 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laubinger S., Zeller G., Henz S.R., Sachsenberg T., Widmer C.K., Naouar N., Vuylsteke M., Scholkopf B., Ratsch G., Weigel D. (2008). At-TAX: A whole genome tiling array resource for developmental expression analysis and transcript identification in Arabidopsis thaliana. Genome Biol. 9: R112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu C.M., Meinke D.W. (1998). The titan mutants of Arabidopsis are disrupted in mitosis and cell cycle control during seed development. Plant J. 16: 21–31 [DOI] [PubMed] [Google Scholar]
- Llorente F., Alonso-Blanco C., Sanchez-Rodriguez C., Jorda L., Molina A. (2005). ERECTA receptor-like kinase and heterotrimeric G protein from Arabidopsis are required for resistance to the necrotrophic fungus Plectosphaerella cucumerina. Plant J. 43: 165–180 [DOI] [PubMed] [Google Scholar]
- MacAlister C.A., Ohashi-Ito K., Bergmann D.C. (2007). Transcription factor control of asymmetric cell divisions that establish the stomatal lineage. Nature 445: 537–540 [DOI] [PubMed] [Google Scholar]
- Masle J., Gilmore S.R., Farquhar G.D. (2005). The ERECTA gene regulates plant transpiration efficiency in Arabidopsis. Nature 436: 866–870 [DOI] [PubMed] [Google Scholar]
- Murashige T., Skoog F. (1962). A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol. Plant. 15: 473–497 [Google Scholar]
- Muller R., Bleckmann A., Simon R. (2008). The receptor kinase CORYNE of Arabidopsis transmits the stem cell-limiting signal CLAVATA3 independently of CLAVATA1. Plant Cell 20: 934–946 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nadeau J.A., Sack F.D. (2002). Control of stomatal distribution on the Arabidopsis leaf surface. Science 296: 1697–1700 [DOI] [PubMed] [Google Scholar]
- Nakagami H., Pitzschke A., Hirt H. (2005). Emerging MAP kinase pathways in plant stress signalling. Trends Plant Sci. 10: 339–346 [DOI] [PubMed] [Google Scholar]
- Nakagawa T., Kurose T., Hino T., Tanaka K., Kawamukai M., Niwa Y., Toyooka K., Matsuoka K., Jinbo T., Kimura T. (2007). Development of series of gateway binary vectors, pGWBs, for realizing efficient construction of fusion genes for plant transformation. J. Biosci. Bioeng. 104: 34–41 [DOI] [PubMed] [Google Scholar]
- Nimchuk Z.L., Tarr P.T., Ohno C., Qu X., Meyerowitz E.M. (2011). Plant stem cell signaling involves ligand-dependent trafficking of the CLAVATA1 receptor kinase. Curr. Biol. 21: 345–352 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pillitteri L.J., Bemis S.M., Shpak E.D., Torii K.U. (2007). Haploinsufficiency after successive loss of signaling reveals a role for ERECTA-family genes in Arabidopsis ovule development. Development 134: 3099–3109 [DOI] [PubMed] [Google Scholar]
- Rafter J.A., Abell M.L., Braselton J.P. (2003). Statistics with Maple. (San Diego, CA: Academic Press; ). [Google Scholar]
- Rowe M.H., Bergmann D.C. (2010). Complex signals for simple cells: The expanding ranks of signals and receptors guiding stomatal development. Curr. Opin. Plant Biol. 13: 548–555 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rychel A.L., Peterson K.M., Torii K.U. (2010). Plant twitter: Ligands under 140 amino acids enforcing stomatal patterning. J. Plant Res. 123: 275–280 [DOI] [PubMed] [Google Scholar]
- Sahlin P., Melke P., Jönsson H. (2011). Models of sequestration and receptor cross-talk for explaining multiple mutants in plant stem cell regulation. BMC Syst. Biol. 5: 2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sessions A., Weigel D., Yanofsky M.F. (1999). The Arabidopsis thaliana MERISTEM LAYER 1 promoter specifies epidermal expression in meristems and young primordia. Plant J. 20: 259–263 [DOI] [PubMed] [Google Scholar]
- Shiu S.H., Bleecker A.B. (2001). Receptor-like kinases from Arabidopsis form a monophyletic gene family related to animal receptor kinases. Proc. Natl. Acad. Sci. USA 98: 10763–10768 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shpak E.D., Berthiaume C.T., Hill E.J., Torii K.U. (2004). Synergistic interaction of three ERECTA-family receptor-like kinases controls Arabidopsis organ growth and flower development by promoting cell proliferation. Development 131: 1491–1501 [DOI] [PubMed] [Google Scholar]
- Shpak E.D., McAbee J.M., Pillitteri L.J., Torii K.U. (2005). Stomatal patterning and differentiation by synergistic interactions of receptor kinases. Science 309: 290–293 [DOI] [PubMed] [Google Scholar]
- Sokal R.R., Rohlf F.J. (1995). Biometry: The Principles and Practice of Statistics in Biological Research, 3rd ed (New York: W.H. Freeman and Company; ). [Google Scholar]
- Sugano S.S., Shimada T., Imai Y., Okawa K., Tamai A., Mori M., Hara-Nishimura I. (2009). Stomagen positively regulates stomatal density in Arabidopsis. Nature 463: 241–244 [DOI] [PubMed] [Google Scholar]
- Torii K.U., Mitsukawa N., Oosumi T., Matsuura Y., Yokoyama R., Whittier R.F., Komeda Y. (1996). The Arabidopsis ERECTA gene encodes a putative receptor protein kinase with extracellular leucine-rich repeats. Plant Cell 8: 735–746 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang G., et al. (2008). A genome-wide functional investigation into the roles of receptor-like proteins in Arabidopsis. Plant Physiol. 147: 503–517 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu L., Xu Y., Dong A., Sun Y., Pi L., Xu Y., Huang H. (2003). Novel as1 and as2 defects in leaf adaxial-abaxial polarity reveal the requirement for ASYMMETRIC LEAVES1 and 2 and ERECTA functions in specifying leaf adaxial identity. Development 130: 4097–4107 [DOI] [PubMed] [Google Scholar]
- Young J.C., Krysan P.J., Sussman M.R. (2001). Efficient screening of Arabidopsis T-DNA insertion lines using degenerate primers. Plant Physiol. 125: 513–518 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.