Abstract
Background and Aims
Asexual reproduction is a prominent evolutionary process within land plant lineages and especially in ferns. Up to 10 % of the approx. 10 000 fern species are assumed to be obligate asexuals. In the Asplenium monanthes species complex, previous studies identified two triploid, apomictic species. The purpose of this study was to elucidate the phylogenetic relationships in the A. monanthes complex and to investigate the occurrence and evolution of apomixis within this group.
Methods
DNA sequences of three plastid markers and one nuclear single copy gene were used for phylogenetic analyses. Reproductive modes were assessed by examining gametophytic and sporophyte development, while polyploidy was inferred from spore measurements.
Key Results
Asplenium monanthes and A. resiliens are confirmed to be apomictic. Asplenium palmeri, A. hallbergii and specimens that are morphologically similar to A. heterochroum are also found to be apomictic. Apomixis is confined to two main clades of taxa related to A. monanthes and A. resiliens, respectively, and is associated with reticulate evolution. Two apomictic A. monanthes lineages, and two putative diploid sexual progenitor species are identified in the A. monanthes clade.
Conclusions
Multiple origins of apomixis are inferred, in both alloploid and autoploid forms, within the A. resiliens and A. monanthes clades.
Keywords: Apomixis, Asplenium monanthes, asexual reproduction, apogamy, diplospory, reticulate evolution, hybridization, polyploidy
INTRODUCTION
Asexually reproducing organisms are at the forefront of evolutionary theory and continue to be the subject of new inquiries (Judson and Normark, 1996; Fontaneto et al., 2007; Schwander et al., 2011). Although the evolutionary transition to obligate asexual reproduction is relatively rare across eukaryotes, it appears to be an important evolutionary process in plants (Mogie, 1990, 1992; Asker and Jerling, 1992). However, the taxonomic distribution of apomictic (asexual) taxa across plant lineages is uneven, with estimations of 0·1 % in angiosperms, up to 10 % in ferns, and with little or no evidence in gymnosperms, mosses, liverworts or hornworts (Walker, 1966a; Lovis, 1977; Asker and Jerling, 1992; Mogie, 1992; Pichot et al., 2001; Park and Kato, 2003).
Sexual reproduction is ancestral in homosporous ferns, and the transition to apomixis requires the evolution and alternation of two distinct phenomena: diplospory and apogamy. They represent the avoidance of meiotic reduction during sporogenesis (diplospory), and the spontaneous development of a sporophyte without fertilization (apogamy) (Lovis, 1977). All except one of the documented apomict fern taxa are obligate (Lloyd, 1973; Lovis, 1977), and the reverse transition from apomixis to sexual reproduction has not been observed.
The transition to apomixis has long been associated with hybridization, polyploidy and female sterility (Mogie, 1992). This is due to the meiotic chromosome behaviour observed in apomictic taxa, the high proportion of triploid apomictic taxa and because apomictic taxa are usually found in complex reticulation networks with other apomictic and sexually reproducing lineages (Manton, 1950; Lovis, 1977; Walker, 1979; Grusz et al., 2009). A recent study by Beck et al. (2011a) supports the role of hybridization in apomictic origins, showing that it drives the transition to apomixis in diploid Boechera species. The interpretation of genetic patterns in other apomictic lineages often reveals a cryptic range of genotypes and morphotypes (Grusz et al., 2009; Schneller and Krattinger, 2010). These patterns can be caused by a number of factors, including: ongoing apomict formation caused by coexisting sexual progenitor species (clonal turnover); extinction or rarity of sexual progenitors; hybridization between apomicts and closely related sexual taxa through functional antheridia; ongoing somatic mutation; and homoeologous chromosome pairing (Klekowski, 1970; Klekowski and Hickok, 1974; Ishikawa et al., 2003). Crucial to the understanding of these patterns is the level of gene flow to and from apomictic lineages. In elucidating origins of apomixis, it is important therefore to consider two hypotheses: a single origin and the subsequent diversification of apomictic lineages, or multiple origins by hybridization between different sexual relatives.
In this study, we aim to investigate the occurrence and origins of apomixis in the Asplenium monanthes complex, for which a modern taxonomic treatment is overdue. The complex consists of up to 22 terrestrial and saxicolous species that occur mainly in Mesoamerica, with its centre of diversity in southern Mexico (based on Mickel and Smith, 2004). The complex is part of the ‘black-stemmed’ spleenwort group that also includes the A. normale complex, A. trichomanes complex, A. viride and the Hawaiian Diellia complex (Schneider et al., 2004, 2005). The A. monanthes complex roughly corresponds to ‘Grupo 5’ of Adams (1995), and contains two documented triploid apomictic species, A. monanthes (n = 2n = 108 and 2n = 144) (Manton, 1950; Manton and Vida, 1968; Manton et al., 1986; Wagner et al., 1970; Tryon et al., 1973; Lovis et al., 1977; Smith and Mickel, 1977) and A. resiliens (n = 2n = 108). It also contains a rare pentaploid apomict species, A. heteroresiliens. The latter is thought to be a hybrid between the apomictic triploid A. resiliens and the sexual tetraploid/hexaploid A. heterochroum (2n = 144 or 216) (Morzenti and Wagner, 1962; Wagner, 1963, 1966; Morzenti, 1966; Wagner and Wagner, 1966; Walker, 1966b; Wagner et al., 1970; Windham, 1983; Haufler and Soltis, 1986). Besides these species, the complex includes the diploid sexual A. formosum (2n = 72) (Manton, 1959; Walker, 1966b; Ghatak, 1977; Ammal and Bahavanandan, 1991; Guillén and Daviña, 2005), and several less well-studied taxa such as A. castaneum, A. blepharodes, A. fibrillosum, A. hallbergii, A. palmeri, A. polyphyllum and A. soleirolioides (Mickel and Smith, 2004). Little or no evidence exists concerning the reproductive biology and ploidy level of these taxa.
Asplenium monanthes is a widespread and presumed long-lived apomictic species that forms 32 unreduced spores via the Döpp–Manton scheme of sporogenesis (Manton, 1950). Currently the complex lacks comprehensive revision, and so here we follow the treatment of the fern flora of Mexico by Mickel and Smith (2004). Although Mexico is the centre of species diversity for this complex, species are distributed from southern North America, thorough Central and South America (Stolze, 1981; Tryon et al., 1993; Wagner et al., 1993; Stolze et al., 1994; Adams, 1995; Smith et al., 1999; Mickel and Smith, 2004; Zuloaga et al., 2008; Jørgensen et al., 2010; Kessler and Smith, 2011). Apomictic triploid A. monanthes and the sexual diploid A. formosum show a wider distribution than the other species within this range, and are distinct as they also occur in other localities throughout the sub-tropics, including Africa and various islands in the Atlantic and Indian oceans. Current knowledge makes this group a prime candidate to explore the origins and long-term fate of apomictic ferns, along with aspects of their biogeography and ecology.
Here, we use plastid and nuclear sequence data to reconstruct species relationships within the A. monanthes complex. This phylogenetic framework is used in combination with some experiments to determine the reproductive modes and ploidy levels in the taxa. Then, we evaluate the following hypotheses: (1) apomixis has multiple origins within this complex; (2) apomixis is associated with reticulate evolution; and (3) apomictic lineages have sexual progenitors present in the complex.
MATERIALS AND METHODS
Taxonomic sampling
In total, 140 specimens were studied, representing approx. 50 % of the species in the Asplenium monanthes complex (Mickel and Smith, 2004; Supplementary Data, Table S1). The approximated taxonomic coverage may be inflated as the result of the limited knowledge on the taxonomy of this group. First, some unsampled taxa were erroneously considered as members of the lineages based on morphology alone. Secondly, several of the unsampled taxa were known from a single or a few samples in herbaria. Studies on the putative sister lineage, the A. trichomanes complex, showed the limitations of morphological-based studies in assessing species diversity of Asplenium species complexes (Lovis, 1977; H. Schneider, unpublished). Most specimens were collected during fieldwork in Mexico, El Salvador and Costa Rica. DNA samples were preserved in silica gel, and voucher specimens were deposited at the herbarium of the Natural History Museum in London (BM), as well as at MEXU, LAGU and INBIO. Additional silica-dried material was provided by colleagues (collectors, localities and additional voucher information are given in Supplementary Data, Table S1). Some samples were obtained from herbarium specimens stored at BM. Unfortunately we were unable to obtain material for A. gentryi, A. heteroresiliens, A. nesioticum, A. oligosorum, A. olivaceum, A. pringlei, A. sanchezii, A. stolonipes, A. tryonii, A. underwoodii and A. vespertinum. These species are rare and/or have restricted geographic ranges that were not visited (Mickel and Smith, 2004). All included samples were carefully identified using the keys provided in Mickel and Smith (2004), and by critical comparison with herbarium collections held at the BM. If needed, specimen identifications were restudied in the context of the obtained DNA sequences, to address inconsistencies in results and identifications. Outgroup taxa were chosen based on Schneider et al. (2004, 2005).
DNA extraction, PCR amplification and sequencing
Total genomic DNA was extracted from leaf tissue using cetyltrimethylammonium bromide (CTAB) (Doyle, 1987). Three plastid markers commonly used in ferns were successfully amplified using polymerase chain reactions (PCRs): (1) the psbA–trnH intergenic spacer (IGS) region (approx. 600 nucleotides) (Aldrich et al., 1988); (2) the rps4 plus rps4–trnS IGS region (approx. 1000 nucleotides) (Smith and Cranfill, 2002); and (3) the trnL–trnF region including the trnL intron and the trnL–trnF IGS region (approx. 900 nucleotides) (Taberlet et al., 1991; Trewick et al., 2002). Samples from A. formosum proved difficult to amplify for trnL–trnF, so only the trnL IGS was amplified using E and F primers (Taberlet et al., 1991; Trewick et al., 2002).
PCR was carried out using Promega Gotaq® Green Master Mix, and amplification was performed using an ABI Veriti Thermo Cycler (Applied Biosystems, Warrington, Cheshire, UK) using protocols from Shaw et al. (2007). We also sequenced a fragment of the low copy nuclear gene pgiC (approx. 600 nucleotides) for a sub-set of specimens. Specimens were chosen in order to attain sufficient phylogenetic coverage, based on the relationships observed in the plastid data set. The generated fragment included the introns spanning exons 14 and 16. The region was amplified using primers pgiC14FN–16RN (Ishikawa et al., 2002), and PCR conditions as above were used. Successfully amplified products were isolated by gel extraction using a QIAquick gel extraction kit (www.qiagen.com). Isolated bands were then cloned using the Promega pGEM®-T Vector system to isolate multiple copies present within the nuclear genome. Only one PCR amplicon per individual was cloned, which unfortunately increases the potential effect of PCR copy preference (Brysting et al., 2011). Where possible, eight colonies were randomly chosen from the plated transformants and amplified by diluting colonies in 30 µL of sterile water and using 1·5 µL of this colony suspension as DNA template. The plasmid-specific primers, puc/M13, were used to amplify the inserted fragments using the thermocycling conditions: 3 min initial denaturation at 94 °C, followed by 35 cycles with 15 s denaturation at 94 °C, 30 s annealing at 55 °C, 1 min elongation at 72 °C and no final extension. Products were eluted in 40 µL of deionized water and then purified using Montage PCR micro 96 Plates. Purified products were quantified using NanoDrop ND-8000 (Thermo Scientific). Sequencing reactions were set up using 2 ng of purified PCR product per 100 bp, 3·5 µl of buffer (2×), 1 µL of 1 µm stock solution of primer, 0·5 µL of Big Dye version 1·1, and the final volume made up to 10 µL with deionized water. Cycle sequencing reactions were run on an Applied Biosystems® 9700 96-well Dual Block Thermal Cycler (Life Technologies Ltd, Paisley, UK) as follows: 28 cycles of 10 s denaturation at 96 °C, 5 s annealing at 50 °C, and 4 min elongation at 60 °C. Both forward and reverse strands were sequenced. The fragments were assembled and edited using Sequencher v4·8 (Gene Codes Corporation) and aligned manually using MacClade 4·08 (Maddison and Maddison, 1989, 2005). All newly generated sequences were confirmed using BLAST searches in GenBank (Benson et al., 2011). The three plastid regions were analysed both separately and in combination. All alignments were checked visually for ambiguous regions, which were excluded from all analyses. Gaps in the alignment were treated as missing data. All sequences are available from GenBank (see Supplementary Data, Tables S1 and S2 for accession numbers).
Phylogenetic analyses
Topological congruence among the three plastid regions was evaluated using bootstrap resampling with maximum parsimony (MP) (Felsenstein, 1985) using PAUP* 4·0b8 (Swofford, 1993, 2002). The bootstrap heuristic searches were as follows: 1000 replicates, ten random sequence additions per replicate, TBR branch swapping and MULTrees option on, collapse zero-length branches off, and saving all trees. The strict bootstrap consensus trees of the individual regions were compared by eye to detect topological conflicts among the three regions using the criteria outlined by Mason-Gamer and Kellogg (1996). We did not find any major topological incongruence between regions (Supplementary Data, Figs S1–S3); therefore, the three plastid regions were compiled into a single matrix for analysis. Specimens that did not have sequence data for all three regions were not included in the combined analysis. This data set was then reduced to represent unique haplotypes only (Supplementary Data, Table S3); this resulted in a data set of 54 specimens. The pgiC data set was reduced to represent unique clones per specimen. To designate unique clones we had to consider the possibility of polymerase error, which can result in two clones of the same allele being amplified with some base pair differences. In order not to confuse these for different alleles, clones that had very similar sequences were only treated as unique if they were separated by significant branch support (Bayesian posterior probability ≥0·6). If sequence differences between these clones had no significant branch support then they were treated as being the same allele, and any observed base differences were inferred to be due to polymerase error. The disadvantage of this method is that real alleles that are recovered only once (due to PCR bias) may be disregarded if supported within a clade of other alleles. No chimeric sequences were observed. According to these criteria, the full data set was reduced from 385 clones to 141, representing only one sequence per unique clone per specimen (Supplementary Data, Table S2).
Maximum likelihood (ML) analyses on the combined plastid and nuclear data sets were carried out in PhyML 3·0 with standard search options and parameters estimated simultaneously from the data (Guindon et al., 2005, 2010). The nucleotide substitution model with the least number of parameters that best fit the data was determined using a likelihood ratio test and the Akaike information criterion (AIC) as implemented in jModeltest version 3·7 (Posada and Crandall, 1998; Posada, 2008), and branch support was estimated using 300 bootstrap replicates (Supplementary Data, Table S4).
Bayesian analysis (BY) was performed on the same two data sets using substitution models according to the Bayesian information criterion (BIC) determined in jModeltest (Supplementary Data, Table S4). Analyses were run in MrBayes 3·1 (Huelsenbeck and Ronquist, 2001; Ronquist and Huelsenbeck, 2003), with Markov Chain Monte Carlo (MCMC) run for 5 million generations and sampled every 500 generations to approximate the posterior probabilities of trees. The two analyses were run simultaneously and a conservative burn-in phase of 25 % was implemented in both data sets to disregard trees prior to convergence on the ML. The remaining trees were then compiled to give 7500 trees for each run, from which a 50 % majority rule consensus was calculated.
Reproductive modes
Diplospory involves a departure from the normal pathway of sporogenesis in ferns, resulting in the production of unreduced (2n) spores (Döpp, 1932; Manton, 1950; Braithwaite, 1964). In derived ferns (see Pryer et al., 2004), the number of spores per sporangium often can be used as a reliable indicator for the occurrence of diplospory because the number of spore mother cells and the number of cell divisions are highly conserved (Schneider et al., 2009). Thus, the vast majority of sexual ferns produce 64 haploid spores per sporangium. In contrast, apomicts exhibiting diplospory produce 32 unreduced spores per sporangium (Döpp, 1932; Manton, 1950; Tryon, 1956; Knobloch, 1966; Vida, 1970).
We determined the number of spores per sporangia for most of the collected specimens. Where possible, whole sporangia were isolated, but in some instances no unopened sporangia were found due to the age of some of the specimens. The sporangia/spore samples were mounted using glycerine jelly, and spore number per sporangia was counted for up to ten sporangia per specimen using a translucent light microscope.
Apogamy involves the initiation of sporophytic growth from a gametophyte without fertilization and indeed without change in chromosome numbers (Manton, 1950; Regalado Gabancho et al., 2010). To confirm the presence/absence of apogamous growth, we carried out experiments in which prothalli were grown from the spore, allowing direct observations on the gametophyte stage of the fern life cycle. Spores were filtered and sterilized using diluted HCl and then applied to the surface of sterilized potted compost and sealed in a re-sealable plastic bag. The pots were kept in dappled light to shaded conditions at 10–20 °C. The pots were monitored regularly and growth typically began after 4 weeks and took between 8 and 24 weeks until prothalli were mature. For determination of reproductive mode, select mature prothalli were mounted on slides and observed in water. Euparal mounting medium (ANSCO Laboratories, Manchester, UK) was used to obtain images. The prothalli were then examined for the presence of antheridia and archegonia and evidence of sporophyte emergence (see examples in Fig. 1). Where sporophytes occurred, their emergent position, character and shape were recorded. Observation of the outgrowth of the sporophyte from vegetative cells of the gametophyte is considered as the best approach to identify apogamous reproduction in ferns (Huang et al., 2011), although apogamy can be induced in sexual fern gametophytes under some conditions (Somer et al., 2009; Kawakami et al., 2011).
Fig. 1.
Sexual vs. apogamous prothalli. Pictures of cultivated prothalli from different specimens (Supplementary Data, Table S7): (A) sexual A. formosum, RD28 – the dashed arrow show the archegonia; (B) apogamous A. monanthes, RD110; (C) apogamous A. monanthes, RD101b; (D) apogamous A. resiliens, RD127; (E) apogamous A. resiliens, RD128; (F) apogamous A. monanthes, RD132; (G) apogamous A. resiliens RD107; (H) apogamous A. monanthes, RD17. Filled circles indicate apical meristems, and sporophytes are indicated by filled squares. Solid arrows indicate apogamous growth.
Ploidy level
In ferns, the size of spores and stomatal guard cells can often be employed to infer differences in ploidy levels among closely related species (Barrington et al., 1986; Huang et al., 2006). Spore measurements were successfully employed in several recent studies on the evolution of apomixis in ferns (Beck et al., 2010; Sigel et al., 2011) despite the occurrence of aborted spores, which may add some error to the estimate range (Huang et al., 2011). In the A. trichomanes complex, the sister group to the A. monanthes complex, it was possible to distinguish ploidy level according to spore size classes, with a mean spore length of 29·4 µm in diploids and 41·6 µm in tetraploids (Moran, 1982; Tutin et al., 1993). Here, spores were measured for a representative sampling of 49 specimens (i.e. maximizing phylogenetic coverage). Spores from individual specimens were mounted using glycerine jelly and measured for length and width using AxioVision on a calibrated light microscope (v4·8.2, www.zeiss.com). An average of 25 spores per specimen were measured, although with some specimens very few spores were recovered. Box plots of width and length were plotted separately to determine the variance within each. Ploidy level was inferred based on statistical analysis of spore length variation. Analyses included a one-way analysis of variance (ANOVA) and Tukey–Kramer analyses at 95 % confidence levels using R (R Development Core Team, 2011). Ploidy level was also estimated based upon calibrations of the mean values and variation in spore measurements to known spore measurements and ploidy levels in the A. trichomanes complex (Moran, 1982; Tutin et al., 1993).
Reticulate evolution
In order to illustrate reticulate evolution (see Hörandl et al., 2005; Pirie et al., 2009; Huson and Scornavacca, 2011), we reconstructed a phylogenetic network using SplitsTree version 4·12·6 (Huson and Bryant, 2006). Plastid and nuclear data sets were reduced to a sub-set of compatible samples (Supplementary Data, Tables S5 and S6). One sample for each unique combination of plastid vs. nuclear loci was included. However, unique combinations did not include those that varied in nuclear copy number, only those that varied in nuclear copy distribution, i.e. nuclear copies that occurred in different clades. The nuclear data set was further divided into four sub-sets, so that each specimen was represented only once in a data set, i.e. multiple copies were present in separate data sets. The optimal trees resulting from ML bootstrap analyses of the plastid and nuclear data sets were used as input files. A reticulation network was computed using the ‘RECOMB2007’ method as implemented in SplitsTree (Klöpper and Huson, 2008). The resulting network was then used to inform the manual construction of a reticulation cladogram, which summarizes the reticulate relationships between and within species of this complex.
RESULTS
Plastid phylogenetic analyses
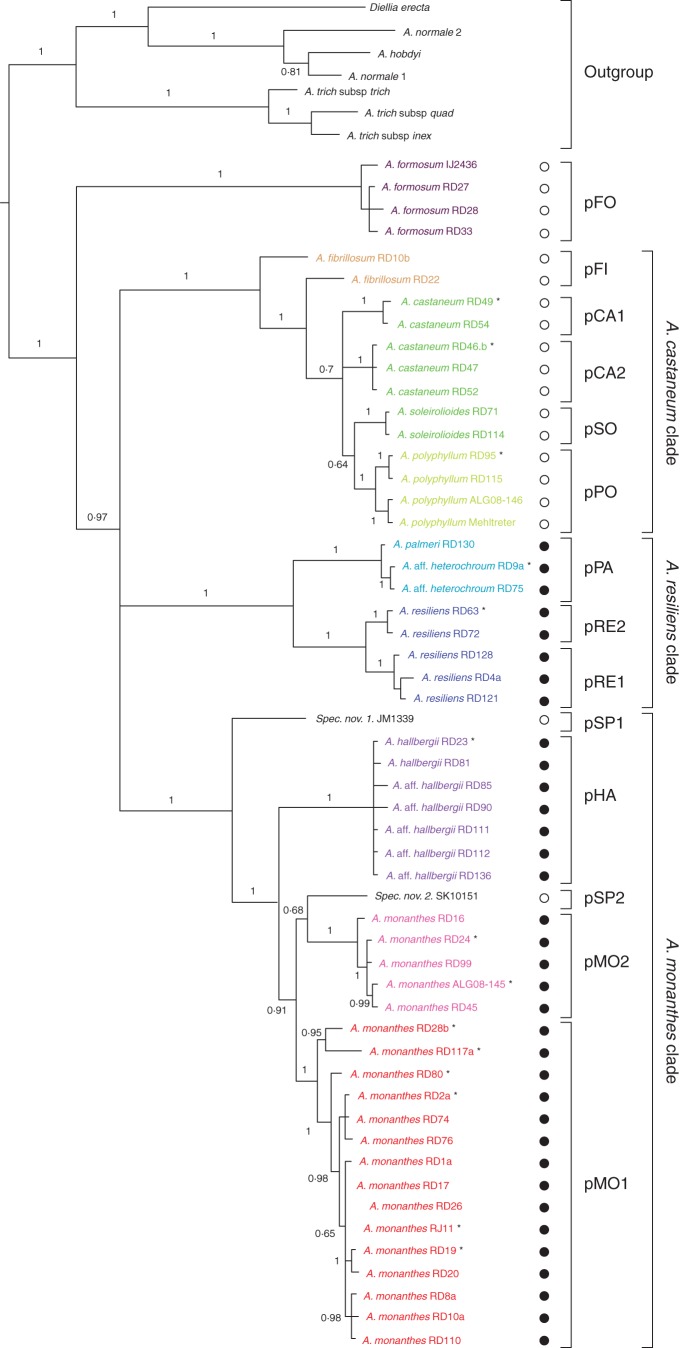
Analysis of the combined plastid data set using BY (Fig. 2) and ML (Supplementary Data, Fig. S4) resulted in trees with similar topology: see Fig. 2 for posterior support values, and Supplementary Data, Fig. S4 for bootstrap support values. Asplenium formosum (pFO) was found to be the sister to the remainder of the A. monanthes complex, which formed a polytomy consisting of three well-supported clades: the A. castaneum clade, the A. resiliens clade and the A. monanthes clade (Fig. 2). The A. castaneum clade consisted of distinct lineages of A. polyphyllum (pPO) and A. soleirolioides (pSO), two A. castaneum sub-clades (pCA1 and pCA2), and two lineages containing a paraphyletic A. fibrillosum (pFI) (Fig. 2). In analysis of the psbA–trnH data set, A. blepharodes is recovered within the A. castaneum clade (Supplementary Data, Fig. S1). The A. resiliens clade consisted of three lineages: two distinct A. resiliens sub-clades (pRE1 and pRE2), and one mixed sub-clade consisting of A. palmeri and A. aff. heterochroum specimens (pPA) (Fig. 2). Three exceptions were observed, but not present in the combined analysis: two A. palmeri specimens (CJR2494 and ES486) were recovered in the pRE2 sub-clade (Supplementary Data, Figs S1–S3), and one A. resiliens specimen (CJR2504) was recovered in sub-clade pPA (Supplementary Data, Fig. S3). The A. monanthes clade consisted of two isolated accessions designated spec.nov.1 (pSP1) and spec.nov.2 (pSP2), and three well-supported monophyletic groups. These included two sub-clades of A. monanthes (pMO1 and pMO2), and one sub-clade of A. hallbergii (pHA) (Fig. 2).
Fig. 2.
Bayesian phylogenetic tree based on the combined plastid data. Posterior support values of P > 0·6 are shown. Tips are labelled with species names followed by a voucher accession, and colour-coded according to species. Sub-clades are abbreviated and summarized to the right of the tree. Abbreviations for sub-clades correspond to those used in the text. Asterisks indicate tips representing multiple haplotypes (Supplementary Data, Table S3). Filled black circles indicate apomictic specimens, and open circles indicate specimens inferred to have a sexual mode of reproduction.
Nuclear phylogenetic analyses
The BY (Fig. 3) and ML (Supplementary Data, Fig. S5) analysis of the pgiC 14FN–16RN data set (approx. 600 bp) resulted in trees of similar topology. However, the sub-clades recovered were not analogous to the sub-clades present in the plastid-based phylogenetic trees. There was evidence of reticulate evolution as individual specimens often had multiple copies of nuclear sequences that were distributed in different sub-clades. Reticulate evolution generally was restricted to species within the three major clades (A. castaneum, A. resiliens and A. monanthes) identified in the plastid analysis, although one A. fibrillosum accession (RD10b) shared one nuclear copy with the A. palmeri/A. aff. heterochroum sub-clade (nPA).
Fig. 3.
Bayesian phylogenetic tree based on the nuclear data. Posterior support values of P > 0·6 are shown. Tips are labelled with species names, followed by a voucher accession (e.g. RD112), and indication of the clone identifier out of the total number of clones (e.g. C2/3 = clone no. 2 out of three unique clones; see text for details). Tips are colour-coded according to species. Sub-clades are abbreviated and summarized to the right of the tree.
Sequences obtained from A. castaneum specimens were recovered in three different sub-clades, two of which (nPO and nSO) were sister to the remainder of the complex. Asplenium polyphyllum sequences were also found in these two sub-clades, and A. soleirolioides sequences were present in sub-clade nSO. The remainder of the complex formed an unresolved polytomy. The third sub-clade of A. castaneum (nCA) was recovered within this polytomy as the weakly supported (posterior probability = 0·75) putative sister to a clade comprised of A. hallbergii clone sequences, and one A. monanthes sequence (nHA). The polytomy also included: one clade of A. fibrillosum clone sequences (nFI); one clade comprising sequences of A. monanthes and spec.nov.1 (nMO1); one clade comprising sequences of A. monanthes, A. hallbergii and one sequence of spec.nov.2 (nMO2); and a clade comprising sequences of A. formosum, A. resiliens, A. palmeri, A. aff. heterochroum and one sequence of A. fibrillosum. The latter clade was a polytomy consisting of four sub-clades: nFO, formed of A. formosum sequences only; nRE1, formed of A. resiliens sequences and one A. palmeri sequence; nRE2, formed of A. resiliens sequences only; and nPA, formed of A. palmeri and A. aff. heterochroum sequences, one sequence of A. resiliens and one sequence of A. fibrillosum (see Fig. 3 for details).
Evidence for reticulate evolution
The unique clones (i.e. nuclear copies) within individual specimens were interpreted as different alleles. The distribution of these unique clones showed evidence of reticulation between clades (Fig. 3; Supplementary Data, Table S5). These reticulate relationships are summarized in a hybridization network (Fig. 4; Supplementary Data, Fig. S6 and Table S6).
Fig. 4.
A reticulation network illustrating the evolutionary history of the A. monanthes complex. This summarizes a hybridization network (Supplementary Data, Fig. S6) performed in SplitsTree (Huson and Bryant, 2006) and illustrates hypothetical hybrid relationships observable by comparison between nuclear and plastid trees (Figs 1 and 2). The terminals illustrated are inferred as different species forms. Where copy numbers for one-species forms are variable, we have included the range of values in parentheses after the species label. Branches are colour-coded according to species. Filled black circles indicate apomictic species, and open circles indicate species inferred to have a sexual mode of reproduction. The plastid group and nuclear copy distribution are summarized adjacent to terminal labels. The dashed red line indicates inferred association of this spec.nov.1 with the MO1 A. monanthes lineage.
The nuclear copy distribution of A. formosum specimens and A. fibrillosum specimen RD22 showed no evidence of reticulation, and these specimens only had one or two copies. The A. fibrillosum accession designated RD10b had four copies, one of which was recovered in the nPA clade. Asplenium castaneum specimens appeared in three forms according to copy number and distribution. One showed evidence of reticulation, sharing one copy in sub-clade nCA and the other in sub-clade nPO. Of the remaining inferred A. castaneum forms, one had only a single clone sequenced (present in sub-clade nCA), and the other showed no evidence of reticulation, having a single copy in sub-clade nSO. Asplenium polyphyllum specimens were present in two forms: one with a single copy in sub-clade nPO, whereas the other showed reticulation with one copy in sub-clade nPO and a second copy in sub-clade nSO. Asplenium soleirolioides showed no reticulation. Asplenium resiliens specimens showed clear evidence of reticulation, with all except one specimen having one, two or three copies distributed between sub-clades nRE1 and nRE2. The exception was specimen CJR2504, which had one sequence in sub-clade nPA, and two in sub-clade nRE1. Most specimens of A. palmeri and A. aff. heterochroum showed no evidence of reticulation. There was one exception, whereby one A. palmeri specimen (CJR2494) had three copies, two distributed in sub-clade nPA and one in sub-clade nRE1. Details are shown in Fig. 3 and a summary is presented in Fig. 4.
Extensive reticulation was observed between the A. monanthes and A. hallbergii clades designated nMO1, nMO2 and nHA. Asplenium monanthes specimens with the plastid pMO1 haplotype had up to two nuclear copies in clade nMO1 and a third in nMO2. Asplenium hallbergii specimens showed reticulation, having up to two nuclear copies in the nHA clade, and a single copy in clade nMO2. Specimen RD23 was the exception, having three copies only in the nHA clade. One of the A. aff. hallbergii specimens (RD85) had four nuclear copies, two of which were derived from the nHA clade and two from the nMO2 clade. In addition to these clear cases of reticulate evolution, there were a few specimens that showed no evidence of reticulation. Asplenium monanthes specimens with the plastid haplotype pMO2 were monophyletic (2–3 copies) in clade nMO2, and spec.nov.2 (of sub-clade nMO2) and spec.nov.1 (of sub-clade nMO1) had one and two copies of pgiC, respectively. The exception in pMO2 was specimen RD99, which had a nuclear copy in each of the clades nMO1, nMO2 and nHA.
Evidence for diplospory and apogamy
Diplospory was inferred for taxa producing 32 spores per sporangium; these included A. monanthes, A. hallbergii, A. aff. hallbergii, A. resiliens, A. palmeri and A. aff. heterochroum (Table 1; Supplementary Data, Table S5). No diplospory was evident in A. formosum, A. fibrillosum, A. castaneum, A. polyphyllum, A. soleirolioides, spec.nov.1 or spec.nov.2, all of which had counts of 64 spores per sporangium. One specimen of A. aff. hallbergii (RD85) yielded counts of both 32 and 64 spores per sporangium from the same individual; all of these spores appeared viable.
Table 1.
Spore characteristics and apogamy as recovered in this study by investigating spore characters, and cultivation and observations of gametophytes
| Species | Spore number | Apogamy | Mean spore length range (μm) | Mean spore width range (μm) |
|---|---|---|---|---|
| A. castaneum | 64 | – | 36·54–46·53 | 28·39–37·69 |
| A. fibrillosum | 64 | – | 44·34–47·90 | 34·85–37·97 |
| A. formosum* | 64 | No | 27·85–32·20 | 20·64–25·02 |
| A. hallbergii | 32 | Yes | 39·15–46·83 | 27·07–32·84 |
| A. aff. hallbergii | 32/64 | Yes | 43·32–44·97 | 30·45–31·86 |
| A. aff. heterochroum | 32 | Yes | 37·43–38·62 | 25·89–27·83 |
| A. monanthes (MO1)* | 32 | Yes | 38·02–47·38 | 26·68–32·08 |
| A. monanthes (MO2)* | 32 | Yes | 38·45–48·54 | 26·63–34·72 |
| A. palmeri† | 32 | – | – | – |
| A. polyphyllum | 64 | – | 35·87–39·87 | 25·06–30·06 |
| A. resiliens† | 32 | Yes | 39·78–44·70 | 28·17–31·10 |
| A. soleirolioides | 64 | – | 36·88 | 27·55 |
| Spec.nov.1 | 64 | – | 31·26 | 21·42 |
| Spec.nov.2 | 64 | – | 29·22 | 20·77 |
Mean spore length and width are reported as ranges where multiple specimens were measured per species, and as single values where only one specimen was measured per species.
* Taxa for which chromosome counts are available: A. formosum 2n = 72 (Manton, 1959; Walker, 1966b; Ghatak, 1977; Ammal and Bahavanandan, 1991; Guillén and Daviña, 2005); A. monanthes n = 2n = 108 and 2n = 144 (Manton, 1950; Manton and Vida, 1968; Wagner et al., 1970; Tryon et al., 1973; Smith and Mickel, 1977; Manton et al., 1986) and A. resiliens n = 2n = 108 (Morzenti and Wagner, 1962; Wagner, 1963, 1966; Morzenti, 1966; Wagner and Wagner, 1966; Walker, 1966b; Wagner et al., 1970; Windham, 1983).
† Wagner et al. (1993) reported 64 spores for A. palmeri whereas Mickel and Smith (2004) show no spore count.
Our spore germination experiments showed that A. formosum produced normal sexual organs, and showed no evidence of apogamous growth (Fig. 1A). All studied specimens of A. hallbergii, A. aff. heterochroum, A. monanthes and A. resiliens produced gametophytes without sexual organs (see Fig. 1B–H). Sporophytes in these taxa emerged either through a callus-like sporophytic growth from the central or lower region of the gametophyte (Fig. 1B–E), or by extension of the apical meristem (Fig. 1E–G). In some cases, prothalli were deformed where the apical cell appears to have become detached (Fig. 1H). Apomictic reproduction appeared to be obligate in all cases, with apogamous prothalli exclusive to specimens exhibiting evidence of diplospory (Table 1; Supplementary Data, Table S7).
Inferring ploidy level
Analysis of spore length variation between the different species showed that the spores of A. formosum are significantly smaller than those of all other species (P < 0·001), with the exception of spec.nov.1 and spec.nov.2 (Table 1, Fig. 5; Supplementary Data, Table S8 and Fig. S7). Spec.nov.1 and spec.nov.2 had significantly smaller spore sizes (P < 0·001) than all other specimens within the A. monanthes MO1 clade and the A. monanthes MO2 clade, respectively. Mean spore lengths in A. formosum, spec.nov.1 and spec.nov.2 were <32 µm, comparable with those of sexual diploids in the related A. trichomanes complex (see Fig. 5). We infer from these results that these three taxa represent sexually reproducing diploids.
Fig. 5.
Boxplots illustrating variation in spore length of species/specimens of the A. monanthes complex. Each boxplot is labelled below with its corresponding species name and specimen voucher, and coloured according to species as in Fig. 2. Boxplots are grouped together according to clades and ordered from lowest to highest mean lengths. Each boxplot represents the variance of measurements of spores within each specimen, the thick horizontal line is the median, the box indicates the variation observed between the 25th and 75th percentiles, the whiskers show the variance range, and small circles identify extreme outliers. Dashed lines running across the graph illustrate average spore measurements in comparison with the ploidy levels of the spores, based on average counts of triploid A. resiliens (spore ploidy = 3x) specimens, hexaploid A. heterochroum (spore ploidy = 3x) specimens (Morzenti, 1966) and diploids and tetraploids of the A. trichomanes complex (spore ploidy = x and 2x, respectively) (Tutin et al., 1993). Filled black circles indicate apomictic specimens that produce 32 spores per sporangium. Open circles indicate specimens that produce 64 spores per sporangium and are inferred to have a sexual mode of reproduction. The asterisk indicates a specimen that produces both 32 and 64 spores per sporangium.
The remainder of the A. monanthes complex showed significant spore size variation between (<0·001) and within species (<0·05, with the exception of A. aff. heterochroum, A. soleirolioides and A. fibrillosum). However, this variation was difficult to classify into distinguishable groups and so inference of ploidy level was not possible. All remaining species did have large mean spore sizes (35–52 µm), and were significantly larger (P < 0·001) than inferred diploid species (A. formosum, spec.nov.1 and spec.nov.2) (Table 1, Fig. 5; Supplementary Data, Table S8 and Fig. S7). We therefore infer that the remaining species were all polyploids.
DISCUSSION
Evolutionary relationships and reticulate evolution
Plastid sequence data identified four main lineages within this complex, and some of these lineages have also been recovered in the pgiC data set. The backbone of the pgiC-based tree is a polytomy, and this nuclear marker only provides limited evidence to reconstruct relationships among these species. This region has not commonly been used in ferns, and other studies have encountered similar problems resolving species evolutionary relationships (Juslén et al., 2011). However, pgiC is better suited to trace reticulate evolution when compared with the plastid marker data (see also Juslén et al., 2011).
In the plastid phylogenetic tree, A. formosum was recovered as the putative sister to three clades that broadly correspond to taxonomic species assemblages of A. castaneum, A. resiliens and A. monanthes. Relationships among these three clades are unresolved, though the lineages themselves are strongly supported. The nuclear phylogenetic tree shows evidence of extensive reticulate evolution within these clades but not between them, except for one hybrid specimen originally identified as A. fibrillosum (Figs 3 and 4). We are able to infer parentage of putative hybrids based on the phylogenetic distribution of nuclear clones. However, it is important to consider that the power of these inferences is potentially limited by PCR copy preference (Brysting et al., 2011), or processes such as diploidization, which could produce misleading results.
According to spore number and cultivation experiments, apomixis has evolved in the A. monanthes and A. resiliens clades but appears to be absent in the A. castaneum clade. Several previously published chromosome counts indicate that A. formosum is a sexual diploid (2n = 72) (Manton, 1959; Walker, 1966b; Ghatak, 1977; Ammal and Bahavanandan, 1991; Guillén and Daviña, 2005), which is consistent with the spore measurements and gametophyte experiments reported here. We did not find any evidence to support suggestions of A. formosum as one parent of A. monanthes.
Fern spore size has been successfully used to determine ploidy level in several recent studies on apomixis in xeric ferns (Beck et al., 2010; Sigel et al., 2011). Also, studies correlating cytofluorometry and spore measurements in the A. trichomanes complex have indicated that spore size provides a useful estimator of ploidy level among the black-stemmed rock spleenworts (Ekrt and Stech, 2008). However, it is not always clear if spore size is reliable for inferring ploidy level (Barrington et al., 1986), and in this study we were only able to use spore size measurements to distinguish between diploid and polyploid specimens, and were not able to infer polyploid levels.
The A. resiliens clade
We report that A. palmeri and A. heterochroum-like morphotypes produce 32 spores per sporangium and show an apomictic mode of reproduction. An unpublished chromosome count of n = 2n = 108 for A. palmeri (M. D. Windham, pers. comm.) confirms that at least some specimens assigned to this taxon are apomictic triploids. Wagner et al. (1993) reported 64 spore counts per sporangium in the Flora of North America treatment of this species, whereas Mickel and Smith (2004) did not record the spore number for this species. Asplenium heterochroum was previously known as a sexual polyploid (4x and 6x) with 64 spores (Morzenti and Wagner, 1962; Wagner, 1963, 1966; Morzenti, 1966; Wagner and Wagner, 1966; Walker, 1966b; Wagner et al., 1970). Our conflicting results may reflect reported taxonomic difficulties (Stolze, 1981; Tryon et al., 1993; Stolze et al., 1994) and suggest that these species require further cytological study. This complex was not exhaustively sampled, and inference of reticulate relationships and apomictic origins calls for caution.
The A. castaneum clade
The distinction between A. polyphyllum and A. castaneum has previously posed taxonomic difficulties (Mickel and Smith, 2004), and our reconstruction of phylogenetic relationships was unable to resolve these. Our spore size data provide the first evidence for polyploidy in this species complex. Spore data indicate that A. fibrillosum, A. polyphyllum, A. soleirolioides and A. castaneum are polyploids.
The plastid phylogenetic tree identifies species lineages broadly corresponding to proposed morphological groups. Asplenium castaneum comprises two sub-clades, with members of the pCA1 sub-clade more similar to A. fibrillosum in appearance, and representatives of the pCA2 sub-clade more likely to be misidentified as A. polyphyllum. However, the nuclear phylogenetic tree indicates extensive reticulation and multiple hybrid formation (Fig. 4). The observed pattern is similar in complexity to the pervasive reticulation observed in the A. trichomanes polyploid complex (Lovis, 1977). Denser sampling of specimens and additional cytological information (e.g. chromosome counts) are required to reconstruct the relationships within this widespread complex.
The A. monanthes clade
This clade comprises two A. monanthes lineages (MO1 and MO2) and one A. hallbergii (HA) lineage, which are supported in both plastid and nuclear genomes. It also contains two accessions (spec.nov.1 and spec.nov.2), which are isolated in the plastid genome but associated with MO1 and MO2 in the nuclear genome. Specimens of A. monanthes and A. hallbergii are apomictic and appear to be triploid and possibly tetraploid, as indicated by previous chromosome counts for A. monanthes (Manton, 1950; Smith and Mickel, 1977). Both the MO1 and A. hallbergii lineages consist of allopolyploid hybrids formed with the MO2 lineage, which itself appears to be autoploid (Figs 4 and 6). Specimen RD99 is an exception to the inference of autoploidy in lineage MO2, as it contains clones from three lineages. This could indicate PCR bias in the cloning method, but we would expect that the high sample number within this lineage would overcome this bias. RD99 could therefore represent a case of gene flow from an apomictic lineage, through hybridization with a sexual relative, via functional antheridia.
Fig. 6.

A reticulogram illustrating the apomictic polyploid species relationships and hypothetical origins of apomixis within the A. monanthes clade. Extant cytotypes are used to illustrate hypothesized lineage origins and parental relationships. The inferred ploidy levels within the three lineages are represented using circles for diploids, triangles for triploids and squares for tetraploids. The diploids are sexually reproducing species, whereas the triploids and tetraploid forms are apomictic. The dashed circle represents a hypothesized (either unsampled or extinct) parent cytotype. Dashed arrows indicate paternal parent relationships. Solid arrows indicate maternal parents (i.e. unreduced gametophytes).
The A. aff. hallbergii specimens (RD85 and RD90) are nested within the A. hallbergii lineage, but appear physically larger in size and show up to four nuclear copies, implying that they could be tetraploid variants (Figs 4 and 5). One of these specimens (RD90) produced both 32- and 64-spored sporangia. Both types of spores appeared normal and viable. This would suggest that RD90 is producing both reduced and unreduced spores, and thus might be a facultative apomict, though separate cultivation experiments are necessary to test this hypothesis.
Spec.nov.1 (JM1339) and spec.nov.2 (SK10151) are inferred to be sexual diploids, and have affinities for the nuclear A. monanthes lineages nMO1 and nMO2, respectively (Figs 4 and 6). Although no morphological analysis was performed, some patterns were observed during visual identification of these specimens. Spec.nov.2 is morphologically similar to some A. monanthes specimens, but spec.nov.1 appears distinct, having previously been identified as A. polyphyllum (Monterrosa et al., 2009). Further study confirmed the morphological differences of the later specimen from A. monanthes and A. polyphyllum (as defined in Mickel and Smith, 2004). The A. hallbergii group is morphologically distinct from the two A. monanthes groups (Mickel and Smith, 2004). Morphological distinctions between the MO1 and MO2 lineages are unclear, but further investigation may uncover correlations between morphological and genetically identified taxa.
In general, our approach using chloroplast DNA (cpDNA) and nuclear ribosomal DNA (nrDNA) markers improved our understanding of the relationships within the group, especially when polyploidy is considered. Further improvements will probably be achieved by expanding the taxonomic sampling, using a more sensitive method to determine polyploidy (e.g. flow cytometry) and by using several nuclear markers. Single nuclear markers can result in misleading interpretations as a result of factors such as incomplete lineage sorting which resemble patterns of ongoing hybridization (Nitta et al., 2011).
Origins of apomixis
The occurrence of apomixis in this complex can be explained in two ways: (1) a single origin with a complex history of reversals and post-origin differentiation/divergence of apomictic lineages; or (2) multiple independent origins from more than one sexual progenitor species (e.g. Beck et al., 2011a, b; Sigel et al., 2011).
The weak resolution among clades and putative sister relationships of the A. monanthes clade and A. resiliens clade may be seen as arguments for a single origin of apomixis in these ferns. This would then imply that the various apomictic lineages were the result of diversification within an asexual lineage (Fontaneto et al., 2007). However, this would require the sexual taxa (spec.nov.1 and spec.nov.2) within the A. monanthes clades to have undergone evolutionary reversals from asexual to sexual reproduction (Figs 1 and 4). To our knowledge, this process has not been documented for apomictic ferns yet.
The alternative hypothesis of multiple origins is supported by evidence for the association of apomixis with reticulate evolution in both the A. resiliens and A. monanthes clades. Here, apomictic taxa are present in multiple polyploid and hybrid forms, including putative autoploid and alloploid lineages. Although reticulation is common throughout the complex (including sexual taxa), and there are some non-hybrid apomictic lineages present, the pattern of hybridization observed does not correspond to the single origin hypothesis.
We therefore favour this second scenario of multiple origins within the whole complex. Within the A. monanthes clade in particular, there is evidence for three lineages. The pattern of nuclear copy distribution between these lineages indicates three origins of apomixis from up to three different sexual progenitor species (Fig. 6; Supplementary Data, Tables S5 and S6). Under the assumption of rarity of regaining sexual reproduction in obligate apomictic lineages, the observed pattern does not support a single origin, and gene flow between asexual and sexual lineages by hybridization (via functional antheridia) with sexual relatives. This inference is strengthened by the lack of antheridia observations in gametophyte studies. However, one specimen (RD99) has nuclear copies from all three apomictic lineages. This indicates that apomixis may have spread via functional hybridization in some instances.
Nuclear copies from the MO2 lineage are present in both the A. monanthes MO1 and the A. hallbergii apomictic lineages (Figs 4 and 6). This indicates that the A. monanthes MO2 lineage is crucial in alloploid apomictic origin. The A. monanthes MO2 lineage itself is a putative autoploid lineage. Although the precise pathways to autoploid apomict origin are difficult to infer, this process almost certainly involved the cross-fertilization of normal haploid gametophytes and unreduced gametes, the latter produced by either sexual diploids or autotetraploids (Fig. 6).
The affinity of the sexual diploids, spec.nov.1 and spec.nov.2, for the A. monanthes MO1 and MO2 lineages, respectively, indicates that these accessions may represent sexual progenitor species for each group. No putative diploid sexual progenitor specimen was observed for the A. hallbergii lineage but, due to the clear difference in morphology and distinction in both the plastid and nuclear genomes, we suggest the participation of at least one additional undetected sexual progenitor. The A. castaneum sub-clade nCA (weakly supported as sister to A. hallbergii by the nuclear gene analysis) and the putative facultative tetraploid A. aff. hallbergii are potential progenitors (Fig. 4).
Similar patterns are observed within the A. resiliens clade, although we make no inferences here, as more exhaustive sampling of species within this clade is necessary.
These hypotheses of origin need to consider the limits of single gene markers; additional studies would benefit from the use of several nuclear genes or a better marker system (such as co-dominant microsatellites) that sample a larger portion of the genomes. We also need to be aware that the dynamics of genomic re-arrangement in apomictic polyploids may create patterns similar to those observed by entirely different processes such as homoeologous chromosome pairing (Ishikawa et al., 2003). The large ranges of spore size in some specimens such as RD24 and RD99 (Fig. 5) suggest that such processes may be present in this group and may also affect spore development.
These caveats aside, our results are congruent to those of several recent studies on cheilanthoid ferns (Grusz et al., 2009; Beck et al., 2011b; Sigel et al., 2011) and filmy ferns (Nitta et al., 2011) that reported multiple recent origins of apomixis (both alloploidy and autoploid) from a variety of ancestral diploid progenitor species.
Conclusions
Within this complex there appear to be a number of relatively well-established apomictic lineages of independent origins. However, we find no strong evidence for long-lived apomictic lineages that outlast or replace their diploid progenitors completely. Sexual progenitors are rare, and the lack of evidence for functional antheridia in apomictic prothalli provides little evidence for ongoing clonal turnover. Apomictic lineages are associated with reticulate evolution. However, we are as yet unable to determine the precise mechanisms of origin or, for that matter, whether this association of apomixis with reticulation is the cause or the effect of apomictic origins within this complex.
Our study is limited by the fact that nuclear copy distribution is not the best indicator of parentage/reticulation patterns, and our conclusions should be viewed primarily as a foundation for further investigation. Our findings also call for some taxonomic studies in the complex. For example, following the collection of two previously unknown sexual diploid taxa for the flora of El Salvador (Monterrosa et al., 2009), more extensive sampling is needed and may lead to the discovery of several new taxa that are either rare and/or have small distribution ranges. We believe that future work would benefit from using population genetic approaches and more in-depth cytological analysis to disentangle fully the mechanisms by which asexuality evolves and is maintained.
SUPPLEMENTARY DATA
ACKNOWLEDGEMENTS
We would like to thank A. Aitken, A. Hall and S. Russell for help in the laboratory; the DNA sequencing services at the Natural History Museum; P. Acock, M. Christenhusz, M. Dickinson, A. Grusz, S. Hennequin, I. Jimenez, R. Jonas, P. Kamau, M. Kessler, S. Knapp, T. Kroemer, A. Larsson, J. Launert, M. Lehnert, T. Lemieux, C. Marsh, K. Mehlttreter, A. Monro, J. Monterosa, H. Muth, E. Otto, C. Rothfels, P. Rothfels, E. Schuettpelz, D. Tejero-Diez, M. Windham and G. Yatskievych for providing samples and helping in the field; S. Ansell, T. Barraclough, M. Carine, A. Dyer, A. de Groot, C. Gueiden, S. Hennequin, H. Hipperson, M. Lewis, J. Marquardt, C. Múrria, A. Papadopulos, N. Robson, A. Smith, J. Vogel, L. Wang and Y. Yu for helpful discussions; and M. D. Windham, an anonymous referee and the editor for helpful comments. This work was supported by the Natural History Museum in London, the Chinese Academy of Science, the Royal Society and the European Research Council.
LITERATURE CITED
- Adams CD. In: Asplenium. In. Davidse G, Sousa M, Knapp S, editors. Mexico: Universidad Nacional Autónoma de México; 1995. pp. 290–324. Flora Mesoamericana. [Google Scholar]
- Aldrich J, Cherney BW, Merlin E, Christopherson L. The role of insertions/deletions in the evolution of the intergenic region between psbA and trnH in the chloroplast genome. Current Genetics. 1988;14:137–146. doi: 10.1007/BF00569337. [DOI] [PubMed] [Google Scholar]
- Ammal LS, Bahavanandan KV. Cytological studies on the genus Asplenium Linn. Indian Fern Journal. 1991;8:69–73. [Google Scholar]
- Asker SE, Jerling L. Apomixis in plants. London: CRC Press; 1992. [Google Scholar]
- Barrington DS, Paris CA, Ranker TA. Systematic inferences from spore and stomate size in the ferns. American Fern Journal. 1986;76:149–159. [Google Scholar]
- Beck JB, Windham MD, Yatskievych G, Pryer KM. A diploids-first approach to species delimitation and interpreting polyploid evolution in the fern genus Astrolepis (Pteridaceae) Systematic Botany. 2010;35:223–234. [Google Scholar]
- Beck JB, Alexander PJ, Allphin L, Al-Shehbaz IA, Rushworth C, Bailey CD, Windham MD. Does hybridization drive the transition to asexuality in diploid Boechera. Evolution. 2011a;66:985–995. doi: 10.1111/j.1558-5646.2011.01507.x. [DOI] [PubMed] [Google Scholar]
- Beck JB, Windham MD, Pryer KM. Do asexual polyploid lineages lead short evolutionary lives? A case study from the fern genus Astrolepis. Evolution. 2011b;65:3217–3229. doi: 10.1111/j.1558-5646.2011.01362.x. [DOI] [PubMed] [Google Scholar]
- Benson DA, Karsch-Mizrachi I, Lipman DJ, Ostell J, Sayers EW. GenBank. Nucleic Acids Research. 2011;39(suppl):D32–D37. doi: 10.1093/nar/gkq1079. http://dx.doi.org/10.1093/nar/gkq1079 . [DOI] [PMC free article] [PubMed] [Google Scholar]
- Braithwaite AF. A new type of apogamy in ferns. New Phytologist. 1964;63:293–305. [Google Scholar]
- Brysting AK, Mathiesen C, Marcussen T. Challenges in polyploid phylogenetic reconstruction: a case story from the arctic-alpine Cerastium alpinum complex. Taxon. 2011;60:333–347. [Google Scholar]
- Döpp W. Die Apogamie bei Aspidium remotum Al.Br. Planta. 1932;17:86–152. [Google Scholar]
- Doyle JJ. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochemical Bulletin. 1987;19:11–15. [Google Scholar]
- Ekrt E, Stech M. A morphometric study and revision of the Asplenium trichomanes group in the Czech Republic. Preslia. 2008;80:325–347. [Google Scholar]
- Felsenstein J. Confidence-limits on phylogenies: an approach using the bootstrap. Evolution. 1985;39:783–791. doi: 10.1111/j.1558-5646.1985.tb00420.x. [DOI] [PubMed] [Google Scholar]
- Fontaneto D, Herniou EA, Boschetti C, Caprioli M, Melone G, Ricci C, Barraclough TG. Independently evolving species in asexual bdelloid rotifers. PLoS Biology. 2007;5:e87. doi: 10.1371/journal.pbio.0050087. http://dx.doi.org/10.1371/journal.pbio.0050087 . [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ghatak J. Biosystematic survey of Pteridophytes from Shevaroy Hills, South India. The Nucleus. 1977;20:105–108. [Google Scholar]
- Grusz AL, Windham MD, Pryer KM. Deciphering the origins of apomictic polyploids in the Cheilanthes yavapensis complex (Pteridaceae) American Journal of Botany. 2009;96:1636–1645. doi: 10.3732/ajb.0900019. [DOI] [PubMed] [Google Scholar]
- Guillén RH, Daviña JR. Estudios cromosómicos en especies de Asplenium (Aspleniaceae) de la Argentina. Darwiniana. 2005;43:44–51. [Google Scholar]
- Guindon S, Lethiec F, Duroux P, Gascuel O. PHYML Online: a web server for fast maximum likelihood-based phylogenetic inference. Nucleic Acids Research. 2005;33:W557–W559. doi: 10.1093/nar/gki352. http://dx.doi.org/10.1093/nar/gki352 . [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guindon S, Dufayard JF, Lefort V, Anisimova M, Hordijk W, Gascuel O. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3·0. Systematic Biology. 2010;59:307–321. doi: 10.1093/sysbio/syq010. [DOI] [PubMed] [Google Scholar]
- Haufler CH, Soltis DE. Genetic evidence suggests that homosporous ferns with high chromosome numbers are diploid. Proceedings of the National Academy of Sciences, USA. 1986;88:4389–4393. doi: 10.1073/pnas.83.12.4389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hörandl E, Paun O, Johansson JT, Lehnebach C, Armstrong T, Chen L, Lockhart P. Phylogenetic relationships and evolutionary traits in Ranunculus s.l. (Ranunculaceae) inferred from ITS sequence analysis. Molecular Phylogenetics and Evolution. 2005;36:305–327. doi: 10.1016/j.ympev.2005.02.009. [DOI] [PubMed] [Google Scholar]
- Huang Y-M, Chou H-M, Hsieh T-H, Wang J-C, Chiou W-L. Cryptic characteristics distinguish diploid and triploid varieties of Pteris fauriei (Pteridaceae) Canadian Journal of Botany. 2006;84:261–268. [Google Scholar]
- Huang YM, Hsu SM, Hsieh TS, Chou HU, Chiou WL. Three Pteris species (Pteridacae, Pteridophyta) reprodcue by apogamy. Botanical Studies. 2011;52:79–87. [Google Scholar]
- Huelsenbeck JP, Ronquist F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics. 2001;17:754–755. doi: 10.1093/bioinformatics/17.8.754. [DOI] [PubMed] [Google Scholar]
- Huson DH, Bryant D. Application of phylogenetic networks in evolutionary studies. Molecular Biology and Evolution. 2006;23:254–267. doi: 10.1093/molbev/msj030. [DOI] [PubMed] [Google Scholar]
- Huson DH, Scornavacca C. A survey of combination methods for phylogenetic networks. Genome Biology Evolution. 2011;3:23–35. doi: 10.1093/gbe/evq077. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ishikawa H, Watano Y, Kano K, Ito M, Kurita S. Development of primer sets for PCR amplification of the pgiC gene in ferns. Journal of Plant Research. 2002;115:65–70. doi: 10.1007/s102650200010. [DOI] [PubMed] [Google Scholar]
- Ishikawa H, Ito M, Watano Y, Kurita S. Electrophoretic evidence for homoeologous chromosome pairing in the apogamous fern species Dryopteris nipponensis (Dryopteridaceae) Journal of Plant Research. 2003;116:165–167. doi: 10.1007/s10265-003-0082-x. [DOI] [PubMed] [Google Scholar]
- Jørgensen PM, Nee M, Beck SG. Catálogo de las plantas vasculares de Bolivia. 2010 http://www.tropicos.org/Project/BC . [Google Scholar]
- Judson OP, Normark BB. Ancient asexual scandals. Trends in Ecology and Evolution. 1996;11:41–46. doi: 10.1016/0169-5347(96)81040-8. [DOI] [PubMed] [Google Scholar]
- Juslén A, Väre H, Wikström N. Relationships and evolutionary origins of polyploid Dryopteris (Dryopteridaceae) from Europe inferred using nuclear pgiC and plastid trnL-F sequence data. Taxon. 2011;60:1284–1294. [Google Scholar]
- Kawakami SM, Kawakami S, Katsuhiko K, Shmakov A. Cytological studies of Russian Altai ferns and the haploid sporophyte formation. Chromosome Botany. 2011;6:21–23. [Google Scholar]
- Kessler M, Smith AR. Ferns of Bolivia. 2011 http://www.systbot.uzh.ch/static/fernsofbolivia/ [Google Scholar]
- Klekowski EJ., Jr. Populational and genetic studies of a homosporous fern – Osmunda regalis. American Journal of Botany. 1970;57:1122–1138. [Google Scholar]
- Klekowski EJ, Jr, Hickok LG. Nonhomologous chromosome pairing in the fern Ceratopteris. American Journal of Botany. 1974;61:422–432. [Google Scholar]
- Klöpper TH, Huson DH. Drawing explicit phylogenetic networks and their integration into SplitsTree. BMC Evolutionary Biology. 2008;8:22. doi: 10.1186/1471-2148-8-22. http://dx.doi.org/10.1186/1471-2148-8-22 . [DOI] [PMC free article] [PubMed] [Google Scholar]
- Knobloch IW. A preliminary review of spore number and apogamy within the genus Cheilanthes. American Fern Journal. 1966;56:163–167. [Google Scholar]
- Lloyd RM. Facultative apomixis and polyploidy in Matteuccia orientalis. American Fern Journal. 1973;63:43–48. [Google Scholar]
- Lovis JD. Evolutionary patterns and processes in ferns. Advances in Botanical Research. 1977;4:230–415. [Google Scholar]
- Lovis JD, Rasbach H, Rasbach K, Reichstein T. Asplenium azoricum and other ferns of the A. trichomanes group from the Azores. American Fern Journal. 1977;67:81–93. [Google Scholar]
- Maddison DR, Maddison WP. MacClade 4: analysis of phylogeny and character evolution. 2005 doi: 10.1159/000156416. Version 4·08a. http://macclade.org . [DOI] [PubMed] [Google Scholar]
- Maddison WP, Maddison DR. Interactive analysis of phylogeny and character evolution using the computer program MacClade. Folia Primatologica. 1989;53:190–202. doi: 10.1159/000156416. [DOI] [PubMed] [Google Scholar]
- Manton I. Problems of cytology and evolution in the Pteridophyta. Cambridge: Cambridge University Press; 1950. [Google Scholar]
- Manton I. Cytological information of the ferns of West Tropical Africa. In: Alson AHG, editor. The ferns and fern allies of West Tropical Africa. Cambridge: Cambridge University Press; 1959. pp. 75–81. [Google Scholar]
- Manton I, Vida G. Cytology of the ferns flora of Tristan da Cunha. Proceedings of the Royal Society of London B: Biological Sciences. 1968;170:361–371. [Google Scholar]
- Manton I, Vida G, Gibby M. Cytology of the fern flora of Madeira. Bulletin of the British Museum (Natural History). Botany. 1986;15:123–161. [Google Scholar]
- Mason-Gamer RJ, Kellogg EA. Testing for phylogenetic conflict among molecular data sets in the tribe Triticeae (Gramineae) Systematic Biology. 1996;45:524–545. [Google Scholar]
- Mickel JT, Smith AR. The Pteridophytes of Mexico. New York: New York Botanical Gardens Press; 2004. [Google Scholar]
- Mogie M. Homospory and the cost of asexual reproduction. Evolution. 1990;44:1707–1710. doi: 10.1111/j.1558-5646.1990.tb03858.x. [DOI] [PubMed] [Google Scholar]
- Mogie M. The evolution of asexual reproduction in plants. London: Chapman and Hall; 1992. [Google Scholar]
- Monterrosa JS, Pena-chocarro MC, Knapp S, Lechuga RE. Guia de indentificacion de helechos de El Salvador. Antiguo Cuscatlan. 2009 [Google Scholar]
- Moran RC. The Asplenium trichomanes complex in the United States and adjacent Canada. American Fern Journal. 1982;72:5–11. [Google Scholar]
- Morzenti VM. Morphological and cytological data on southeastern United States species of the Asplenium heterochroum-resiliens complex. American Fern Journal. 1966;56:167–177. [Google Scholar]
- Morzenti VM, Wagner WH. Southeastern American blackstem spleenworts of the Asplenium heteochroum-resilines complex. Association of Southeastern Biologists Bulletin. 1962;9:40–41. [Google Scholar]
- Nitta JH, Ebihara A, Ito M. Reticulate evolution in the Crepidomanes minutum species complex (Hymenophyllaceae) American Journal of Botany. 2011;98:1782–1800. doi: 10.3732/ajb.1000484. [DOI] [PubMed] [Google Scholar]
- Park CH, Kato M. Apomixis in the interspecific triploid hybrid fern Cornopteris christenseniana (Woodsiaceae) Journal of Plant Research. 2003;116:93–103. doi: 10.1007/s10265-003-0081-y. [DOI] [PubMed] [Google Scholar]
- Pichot C, Maataoui M, Raddi S, Raddi P. Surrogate mother for endangered Cupressus. Nature. 2001;412:39–39. doi: 10.1038/35083687. [DOI] [PubMed] [Google Scholar]
- Pirie MD, Humphreys AM, Barker NP, Linder HP. Reticulation, data combination, and inferring evolutionary history: an example from Danthonioideae (Poaceae) Systematic Biology. 2009;58:612–628. doi: 10.1093/sysbio/syp068. [DOI] [PubMed] [Google Scholar]
- Posada D. jModelTest: phylogenetic model averaging. Molecular Biology and Evolution. 2008;25:1253–1256. doi: 10.1093/molbev/msn083. [DOI] [PubMed] [Google Scholar]
- Posada D, Crandall KA. MODELTEST: testing the model of DNA substitution. Bioinformatics. 1998;14:817–818. doi: 10.1093/bioinformatics/14.9.817. [DOI] [PubMed] [Google Scholar]
- Pryer KM, Schuettpelz E, Wolf PG, Schneider H, Smith AR, Cranfill R. Phylogeny and evolution of ferns (monilophytes) with a focus on the early leptosporangiate divergences. American Journal of Botany. 2004;91:1582–1598. doi: 10.3732/ajb.91.10.1582. [DOI] [PubMed] [Google Scholar]
- R Development Core Team. R: a language and environment for statistical computing. 2011 http://www.R-project.org/ [Google Scholar]
- Regalado Gabancho L, Prada C, Gabriel y Galán JM. Sexuality and apogamy in the Cuban Asplenium auritum–monodon complex (Aspleniaceae) Plant Systematics and Evolution. 2010;289:137–146. [Google Scholar]
- Ronquist F, Huelsenbeck JP. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics. 2003;19:1572–1574. doi: 10.1093/bioinformatics/btg180. [DOI] [PubMed] [Google Scholar]
- Schneider H, Russell SJ, Cox CJ, et al. Chloroplast phylogeny of asplenioid ferns based on rbcL and trnL-F spacer sequences (Polypodiidae, Aspleniaceae) and its implications for biogeography. Systematic Botany. 2004;29:260–274. [Google Scholar]
- Schneider H, Ranker TA, Russell SJ, et al. Origin of the endemic fern genus Diellia coincides with the renewal of Hawaiian terrestrial life in the Miocene. Proceedings of the Royal Society B: Biological Sciences. 2005;272:455–460. doi: 10.1098/rspb.2004.2965. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schneider H, Smith AR, Pryer KM. Is morphology really at odds with molecules in estimating fern phylogeny. Systematic Botany. 2009;34:455–475. [Google Scholar]
- Schneller J, Krattinger K. Genetic composition of Swiss and Austrian members of the apogamous Dryopteris affinis complex (Dryopteridaceae, Polypodiopsida) based on ISSR markers. Plant Systematics and Evolution. 2010;286:1–6. [Google Scholar]
- Schwander T, Henry L, Crespi Bernard J. Molecular evidence for ancient asexuality in Timema stick insects. Current Biology. 2011;21:1129–1134. doi: 10.1016/j.cub.2011.05.026. [DOI] [PubMed] [Google Scholar]
- Shaw J, Lickey EB, Schilling EE, Small RL. Comparison of whole chloroplast genome sequences to choose noncoding regions for phylogenetic studies in angiosperms: the tortoise and the hare III. American Journal of Botany. 2007;94:275–288. doi: 10.3732/ajb.94.3.275. [DOI] [PubMed] [Google Scholar]
- Sigel EM, Windham MD, Huiet L, Yatskievych G, Pryer KM. Species relationships and farina evolution in the Cheilanthoid fern genus Argyrochosma (Pteridaceae) Systematic Botany. 2011;36:554–564. [Google Scholar]
- Smith AR, Cranfill RB. Intrafamilial relationships of the thelypteroid ferns (Thelypteridaceae) American Fern Journal. 2002;92:131–149. [Google Scholar]
- Smith AR, Mickel JT. Chromosome counts for Mexican ferns. Brittonia. 1977;29:391–398. [Google Scholar]
- Smith AR, Kessler M, Gonzales J. New records of pteridophytes from Bolivia. American Fern Journal. 1999;89:244–266. [Google Scholar]
- Somer M, Arbesú R, Menéndez V, Revilla MA, Fernández H. Sporophyte induction studies in ferns in vitro. Euphytica. 2009;171:203–210. [Google Scholar]
- Stolze RG. Ferns and fern allies of Guatemala. Part II. Polypodiaceae. Fieldiana, Botany. 1981;6:1–522. [Google Scholar]
- Stolze RG, Pacheco L, Øllgaard B. Polypodiaceae – Dryopteridoideae – Physematieae. In: Harling G, Andersson L, editors. Flora of Ecuador. Goteborg: University of Goteborg, 108; 1994. 14(5B) [Google Scholar]
- Swofford DL. PAUP: a computer-program for phylogenetic inference using maximum parsimony. Journal of General Physiology. 1993;102:A9–A9. [Google Scholar]
- Swofford DL. PAUP* phylogenetic analysis using parsimony (* and other methods). Version 4·0b10. Sunderland, MA: Sinauer Associates; 2002. [Google Scholar]
- Taberlet P, Gielly L, Pautou G, Bouvet J. Universal primers for amplification of 3 non-coding regions of chloroplast DNA. Plant Molecular Biology. 1991;17:1105–1109. doi: 10.1007/BF00037152. [DOI] [PubMed] [Google Scholar]
- Trewick SA, Morgan-Richards M, Russell SJ, et al. Polyploidy, phylogeography and Pleistocene refugia of the rockfern Asplenium ceterach: evidence from chloroplast DNA. Molecular Ecology. 2002;11:2003–2012. doi: 10.1046/j.1365-294x.2002.01583.x. [DOI] [PubMed] [Google Scholar]
- Tryon RM. A revision of the American species of Notholaena. Contributions from the Gray Herbarium. 1956;179:1–106. [Google Scholar]
- Tryon R, Voeller B, Tryon A, Riba R. Fern biology in Mexico (a class field program) BioScience. 1973;23:28–33. [Google Scholar]
- Tryon RM, Stolz RG, Leon B. Pteridophyta of Peru. Part V. 18: Aspleniaceae – 21. Polpodiaceae. Fieldiana, Botany. 1993;32:1–190. [Google Scholar]
- Tutin TG, Burges NA, Chater AO, et al. Flora Europea. Cambridge: Cambridge University Press; 1993. [Google Scholar]
- Vida G. The nature of polyploidy in Asplenium ruta-muraria L. and A. lepidium C. Presl. Caryologia. 1970;23:525–547. [Google Scholar]
- Wagner WH. A biosystematic study of United States ferns – preliminary abstract. American Fern Journal. 1963;53:1–16. [Google Scholar]
- Wagner WH. Two new species of ferns from the United States. American Fern Journal. 1966;56:3–17. [Google Scholar]
- Wagner WH, Wagner FS. Castanea. Vol. 31. Virginia: Biosystematic Studies 1964–65; 1966. Pteridophytes of the mountain lake aras, Gilles Co; pp. 121–140. [Google Scholar]
- Wagner WH, Farrar DR, McAlpin BW. Pteridology of the Highlands Biological Station Area, Southern Appalachians. Journal of the Elisha Mitchell Scientific Society. 1970;86:1–24. [Google Scholar]
- Wagner WH, Moran RC, Werth CR. Flora of North America. New York: Oxford University Press; 1993. Aspleniaceae; pp. 237–238. [Google Scholar]
- Walker TG. Apomixis and vegetative reproduction in ferns. In: Hawkes JG, editor. Reproductive biology and taxonomy of vascular plants. The report of the Conference held by the Botanical Society of the British Isles, at Birmingham University in 1965. Oxford: Pergamon Press; 1966a. pp. 152–161. [Google Scholar]
- Walker TG. A cytotaxonomic survey of the pteridophytes of Jamaica. Transactions of the Royal Society of Edinburgh. 1966b;66:169–237. [Google Scholar]
- Walker TG. The cytogenetics of ferns. In: Dyer AF, editor. The experimental biology of ferns. New York: Academic Press; 1979. pp. 87–132. [Google Scholar]
- Windham MD. The ferns of Elden Mountain, Arizona. American Fern Journal. 1983;73:85–93. [Google Scholar]
- Zuloaga FO, Morrone O, Belgrana MJ. Catalogo de las plantas vasculares del Cono Sur. St. Louis: MBG Press; 2008. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.