Abstract
Triterpenoid saponins are bioactive metabolites that have evolved recurrently in plants, presumably for defense. Their biosynthesis is poorly understood, as is the relationship between bioactivity and structure. Barbarea vulgaris is the only crucifer known to produce saponins. Hederagenin and oleanolic acid cellobioside make some B. vulgaris plants resistant to important insect pests, while other, susceptible plants produce different saponins. Resistance could be caused by glucosylation of the sapogenins. We identified four family 1 glycosyltransferases (UGTs) that catalyze 3-O-glucosylation of the sapogenins oleanolic acid and hederagenin. Among these, UGT73C10 and UGT73C11 show highest activity, substrate specificity and regiospecificity, and are under positive selection, while UGT73C12 and UGT73C13 show lower substrate specificity and regiospecificity and are under purifying selection. The expression of UGT73C10 and UGT73C11 in different B. vulgaris organs correlates with saponin abundance. Monoglucosylated hederagenin and oleanolic acid were produced in vitro and tested for effects on P. nemorum. 3-O-β-d-Glc hederagenin strongly deterred feeding, while 3-O-β-d-Glc oleanolic acid only had a minor effect, showing that hydroxylation of C23 is important for resistance to this herbivore. The closest homolog in Arabidopsis thaliana, UGT73C5, only showed weak activity toward sapogenins. This indicates that UGT73C10 and UGT73C11 have neofunctionalized to specifically glucosylate sapogenins at the C3 position and demonstrates that C3 monoglucosylation activates resistance. As the UGTs from both the resistant and susceptible types of B. vulgaris glucosylate sapogenins and are not located in the known quantitative trait loci for resistance, the difference between the susceptible and resistant plant types is determined at an earlier stage in saponin biosynthesis.
Triterpenoid saponins are a heterogeneous group of bioactive metabolites found in many species of the plant kingdom. The general conception is that saponins are involved in plant defense against antagonists such as fungi (Papadopoulou et al., 1999), mollusks (Nihei et al., 2005), and insects (Dowd et al., 2011). Saponins consist of a triterpenoid aglycone (sapogenin) linked to usually one or more sugar moieties. This combination of a hydrophobic sapogenin and hydrophilic sugars makes saponins amphiphilic and enables them to integrate into biological membrane systems. There, they form complexes with membrane sterols and reorganize the lipid bilayer, which may result in membrane damage (Augustin et al., 2011).
However, our knowledge of the biosynthesis of saponins, and the genes and enzymes involved, is limited. The current conception is that the precursor 2,3-oxidosqualene is cyclized to a limited number of core structures, which are subsequently decorated with functional groups, and finally activated by adding glycosyl groups (Augustin et al., 2011). These key steps are considered to be catalyzed by three multigene families: (1) oxidosqualene cyclases (OSCs) forming the core structures, (2) cytochromes P450 adding the majority of functional groups, and (3) family 1 glycosyltransferases (UGTs) adding sugars. This allows for a vast structural complexity, some of which probably evolved by sequential gene duplication followed by functional diversification (Osbourn, 2010). A major challenge is thus to understand the processes of saponin biosynthesis, which structural variants of saponins play a role in defense against biotic antagonists, and how saponin biosynthesis evolved in different plant taxa. This knowledge is also of interest for biotechnological production and the use of saponins as protection agents against agricultural pests as well as for pharmacological and industrial uses as bactericides (De Leo et al., 2006), anticancerogens (Musende et al., 2009), and adjuvants (Sun et al., 2009).
Barbarea vulgaris (winter cress) is a wild crucifer from the Cardamineae tribe of the Brassicaceae family. It is the only species in this economically important family known to produce saponins. B. vulgaris has further diverged into two separate evolutionary lineages (types; Hauser et al., 2012; Toneatto et al., 2012) that produce different saponins, glucosinolates, and flavonoids (Agerbirk et al., 2003b; Dalby-Brown et al., 2011; Kuzina et al., 2011). Saponins of the one plant type make plants resistant to the yellow-striped flea beetle (Phyllotreta nemorum), diamondback moth (Plutella xylostella), and other important crucifer specialist herbivores (Renwick, 2002); therefore, it has been suggested to utilize such plants as a trap crop to diminish insect damage (Badenes-Perez et al., 2005). The other plant type is not resistant to these herbivores. B. vulgaris, therefore, is ideal as a model species to study saponin biosynthesis, insect resistance, and its evolution, as we can contrast genes, enzymes, and their products between closely related but divergent plant types.
Insect resistance of the one plant type, called G because it has glabrous leaves, correlates with the content of especially hederagenin cellobioside, oleanolic acid cellobioside, 4-epi-hederagenin cellobioside, and gypsogenin cellobioside (Shinoda et al., 2002; Agerbirk et al., 2003a; Kuzina et al., 2009; Fig. 1). These saponins are absent in the susceptible plant type, called P because it has pubescent leaves, which contains saponins of unknown structures and function (Kuzina et al., 2011). The sapogenins (aglycones) of the resistance-causing saponins hederagenin and oleanolic acid cellobioside do not deter feeding by P. nemorum, which highlights the importance of glycosylation of saponins for resistance (Nielsen et al., 2010). Therefore, the presence or absence of sapogenin glycosyltransferases could be a determining factor for the difference in resistance between the insect resistant G-type and the susceptible P-type of B. vulgaris.
Figure 1.
Chemical structures of the four known G-type B. vulgaris saponins that correlate with resistance to P. nemorum and other herbivores. The cellobioside and sapogenin parts of the saponin are underlined, and relevant carbon positions are numbered.
Some P. nemorum genotypes are resistant to the saponin defense of B. vulgaris (Nielsen, 1997b, 1999). Resistance is coded by dominant R genes (Nielsen et al., 2010; Nielsen 2012): larvae and adults of resistant genotypes (RR or Rr) are able to feed on G-type foliage and utilize B. vulgaris as host plant (de Jong et al., 2009), whereas larvae of the susceptible genotype (rr) die and adult beetles stop feeding on G-type foliage. Larvae and adults of all known P. nemorum genotypes can feed on P-type B. vulgaris (Fig. 2).
Figure 2.
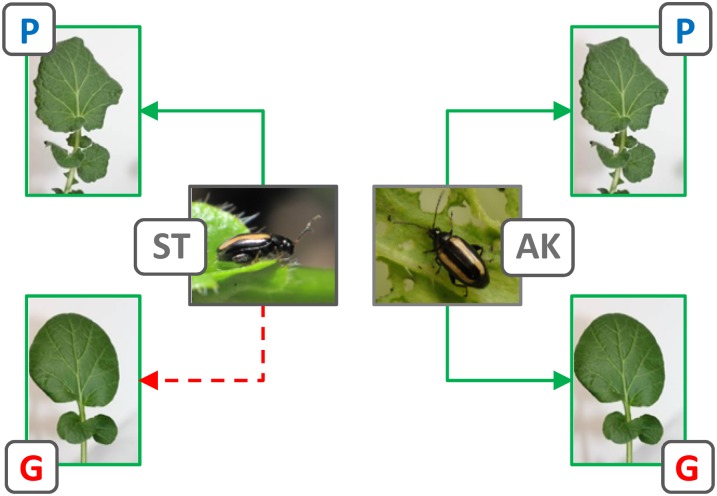
Feeding behavior of adult P. nemorum that are either susceptible (ST) or resistant (AK) toward the saponin-based defense of G-type B. vulgaris; the P-type produces different saponins and is not resistant against P. nemorum. Potential feeding is shown by green arrows, and termination of feeding briefly after initiation is indicated by a red dashed arrow. Larvae of the ST line die if fed on G-type plants.
In this study, we asked which enzymes are involved in glucosylation of sapogenins in B. vulgaris, whether saponins with a single C3 glucosyl group are biologically active, and whether the difference between the insect resistant and susceptible types of B. vulgaris is caused by different glucosyltransferases.
We report the identification of two UDP-glycosyltransferases, UGT73C10 and UGT73C11, which have high catalytic activity and substrate specificity and regiospecificity for catalyzing 3-O-glucosylation of the sapogenins oleanolic acid and hederagenin. The products, 3-O-β-d-glucopyranosyl hederagenin and 3-O-β-d-glucopyranosyl oleanolic acid, are predicted precursors of hederagenin and oleanolic acid cellobioside, respectively. The expression patterns of UGT73C10 and UGT73C11 in different organs of B. vulgaris correlate with saponin abundance, and monoglucosylated sapogenins, especially 3-O-β-d-glucopyranosyl hederagenin, deter feeding by P. nemorum. Our results thus show that glucosylation with even a single glucosyl group activates the resistance function of these sapogenins. However, since the UGTs are present and active in both the insect-resistant and -susceptible types of B. vulgaris, we cannot explain the difference in resistance by different glucosylation abilities. Instead, the difference between the susceptible and resistant types must be determined at an earlier stage in saponin biosynthesis.
RESULTS
Identification of a Sapogenin UDP-Glycosyltransferase by Activity-Based Screening of a cDNA Expression Library
To identify enzymes that glycosylate sapogenins (aglycones of saponins) from B. vulgaris, a complementary DNA (cDNA) expression library was generated from B. vulgaris var variegata, a commercial B. vulgaris variety with a saponin profile similar to the insect-resistant G-type. The library was screened by activity assays using UDP-Glc and oleanolic acid as donor and acceptor substrate, respectively. A single cDNA clone was identified, of which the encoded enzyme glucosylated oleanolic acid, as evidenced by comigration with authentic 3-O-Glc oleanolic acid on thin-layer chromatography (TLC) analysis. The clone was designated BvUGT1 and found to contain a 1,566-bp cDNA with an open reading frame (ORF) of 495 amino acids. BLAST analyses identified Arabidopsis thaliana UGT73C5 as its closest homolog. BvUGT1 has 88% nucleotide identity to UGT73C5, and the encoded amino acid sequence, BvUGT1, is 83% identical to UGT73C5. In addition to oleanolic acid, BvUGT1 also glucosylated hederagenin and echinocystic acid.
Identification of BvUGT1 Homologs in G- and P-Type B. vulgaris
Putative BvUGT1 homologs in the resistant G-type and susceptible P-type were searched by mining a 454 transcriptome data set from the G-type (Kuzina et al., 2011) and the P-type. Based on the identified singlets and contigs, two different full-length ORFs from G-type plants and three from P-type plants were isolated by PCR. The genomic sequences were identified by PCR and shown to be intronless, which is also the case for the seven UGT73Cs in the A. thaliana genome (Paquette et al., 2003). Thus, putative BvUGT1 homologs are not only present in both the G- and P-type B. vulgaris genomes, but they are also expressed. The three P-type UGTs were named UGT73C9, UGT73C10, and UGT73C12, and the two G-type sequences were named UGT73C11 and UGT73C13 (Fig. 3), by the UGT nomenclature committee (Mackenzie et al., 1997). The ORFs of the five UGTs each span 1,488 bp and encode proteins consisting of 495 amino acids.
Figure 3.
Maximum likelihood phylogeny of UGT73Cs described in this study and from online databases. Species are indicated as prefixes to the UGT name: Bv, B. vulgaris; At, A. thaliana; Al, A. lyrata; Br, B. rapa. UGT73C9, UGT73C10, and UGT73C12, shown in blue, are from P-type B. vulgaris, while UGT73C11 and UGT73C13, shown in red, are from the G-type. AtUGT73B5 is included as an outgroup. Bootstrap values (100 iterations) are shown next to the corresponding nodes.
Of the five sequences, UGT73C11 is most identical to BvUGT1 from B. vulgaris var variegata, differing in only three nucleotides, which causes a conservative amino acid substitution of Asp-338 to Glu in UGT73C11. Based on a reconstruction of the phylogeny of the UGTs (Fig. 3), UGT73C9 and UGT73C10 from the P-type and UGT73C11 from the G-type form a discrete cluster, as does UGT73C12 from the P-type and UGT73C13 from the G-type. UGTs in the first cluster are more than 95% identical to each other, and those in the second cluster are more than 97% identical (Supplemental Table S1). Accordingly, UGT73C9/UGT73C10 from the P-type correspond to UGT73C11 from the G-type and UGT73C12 from the P-type corresponds to UGT73C13 from the G-type. In comparison with UGT73C homologs from A. thaliana, Arabidopsis lyrata, and Brassica rapa, the five B. vulgaris sequences are most closely related to A. thaliana UGT73C5 and UGT73C6 and a UGT73C5 homolog in A. lyrata.
The UGTs described in the phylogeny have been exposed to different levels of selection since they diverged, as indicated by the significantly better fit of a model with independent ω (ratio of the number of nonsynonymous substitutions per nonsynonymous site to the number of synonymous substitutions per synonymous site [dN/dS ratios]) for each branch compared with a single common ω ratio for all branches (2ΔlnL = 13.9; P < 0.001). Positive selection among branches was further indicated by the better fit of a model including positive selection (model M3) than a model without M0 (2ΔlnL = 304.7; P < 0.001); 4.3% of the codons were estimated to have been under positive selection. Only the branches leading to UGT73C9, UGT73C10, and UGT7311 showed signs of positive selection; branches leading to UGT73C12 and UGT73C13 as well as A. thaliana, A. lyrata, and B. rapa have ω < 1, showing that these branches are under purifying selection.
All five UGT sequences were mapped to an existing linkage map of B. vulgaris (Kuzina et al., 2011) and found to be located in a region that corresponds to A. thaliana chromosome 2 between 13.5 and 19.6 Mb. None of the UGTs lie within previously reported regions containing quantitative trait loci (QTL) for resistance toward P. nemorum larvae feeding (Kuzina et al., 2011). In A. thaliana, six out of the seven UGT73C genes are positioned in a tandem repeat cluster at 15.4 Mb on chromosome 2. Therefore, it is likely that the identified B. vulgaris UGT73C genes are located in a similar UGT73C cluster in the B. vulgaris genome.
Heterologous Expression and in Vitro Activities of the UGT73Cs
To determine if the five UGTs isolated from G- and P-type B. vulgaris have similar catalytic activities as BvUGT1 from B. vulgaris var variegata, they were heterologously expressed in Escherichia coli. The corresponding crude protein extracts were assayed with different sapogenins as putative sugar acceptors and UDP-Glc as sugar donor. UGT73C10, UGT73C11, UGT73C12, and UGT73C13 catalyzed transfer of a Glc moiety from UDP-Glc to the oleanane sapogenins oleanolic acid and hederagenin and to the lupane sapogenin betulinic acid (Fig. 4). In addition, their precursors β-amyrin and lupeol were glucosylated, but with lower efficiency (Fig. 5). In contrast, UGT73C9 from the P-type appeared inactive toward the compounds tested.
Figure 4.
Activity of the heterologously expressed B. vulgaris UGT73Cs toward sapogenins. Enzyme assays contained 750 ng of recombinant UGT in 50 µL and 50 µm oleanolic acid (oa), hederagenin (he), or betulinic acid (be) as acceptor substrates and 1 mm UDP-Glc as donor substrate. The assays were incubated for 60 min at 30°C and analyzed by TLC. Compounds were visualized by spraying with 10% sulfuric acid in methanol and subsequent heating. The (inverted) image was taken at long-wave UV (366 nm) excitation. Migration of authentic oleanolic acid, hederagenin, 3-O-β-Glc oleanolic acid (oa-Glc), and 3-O-β-Glc hederagenin (he-Glc) is shown in the reference lane (Ref.). Positions of aglycones (aglc), monoglucosides (m-Glc), and diglucosides (di-Glc) are indicated on the left side.
Figure 5.
Substrate specificity of UGT73C10 and UGT73C12. TLC analyses of activity assays with recombinant UGT73C10 or UGT73C12 using 14C-labeled UDP-Glc as donor substrate are shown. Substrates tested were oleanolic acid (oa), hederagenin* (he), β-amyrin (βa), betulinic acid (be), kaempferol (ka), quercetin (qu), and TCP, applied at either 100 or 10 µm concentration. *The hederagenin batch contained a low amount of oleanolic acid.
The glucosylation positions of the two oleanane sapogenins produced by the UGTs were determined by NMR spectroscopy. Based on one-dimensional (1-D) 1H- and 13C- as well as two-dimensional (2-D) Correlation Spectroscopy (COSY)-, Total Correlation Spectroscopy (TOCSY)-, and Heteronuclear Single Quantum Coherence (HSQC)-NMR analyses (Supplemental Data Set S1), the glucosides were concluded to be 3-O-β-d-glucopyranosyl oleanolic acid and 3-O-β-d-glucopyranosyl hederagenin. This is in agreement with these monoglucosides as predicted precursors of oleanolic acid cellobioside and hederagenin cellobioside, respectively.
In addition to the 3-O-monoglucosides, UGT73C12 and UGT73C13 also formed low amounts of diglucosides, while this activity was barely detectable for UGT73C10 and UGT73C11. Based on retention times and fragmentation patterns in liquid chromatography -mass spectrometry analyses, these diglucosides could not be oleanolic acid and hederagenin cellobioside, respectively, but represent bidesmosidic glucosylation (i.e. glycosylation at two different positions; Supplemental Fig. S1). A diglucosylated betulinic acid was, in addition to two different betulinic acid monoglucosides, produced in detectable amounts after 30 min of incubation when using betulinic acid concentrations as low as 10 µm (Fig. 5). After alkaline hydrolysis (saponification), which cleaves the ester but not the ether bonds in glucosylated products, the betulinic acid diglucoside and one of the two betulinic acid monoglucosides were no longer detectable (Supplemental Fig. S2). Therefore, the degraded monoglucoside must be 28-O-glucosylated betulinic acid and the diglucoside must be 3,28-O-diglucosylated betulinic acid. Similarly, the diglucosidic forms of oleanolic acid and hederagenin would represent 3,28-O-diglucosides. Under assay conditions with high amounts of enzyme, increased incubation time, and elevated incubation temperature, UGT73C13 also produced an oleanolic acid triglucoside (Supplemental Fig. S1), which further demonstrates the lower substrate specificity and regiospecificity of UGT73C13. However, the low in vitro production of these glucosides suggests that these additional activities only play a minor role, if any, in planta.
Other members of the UGT73C subfamily have been assigned to be involved in flavonoid and brassinosteroid metabolism (Jones et al., 2003; Poppenberger et al., 2005; Modolo et al., 2007). Glycosylated flavonols derived from quercetin and kaempferol are present in B. vulgaris (Senatore et al., 2000; Dalby-Brown et al., 2011). Consequently, the flavonols quercetin and kaempferol, the phytosterols obtusifoliol, campesterol, sitosterol, and stigmasterol, and the brassinosteroid 24-epi-brassinolide were tested as substrates. 2,4,5-Trichlorophenol (TCP) was included as a positive control, as it can be glycosylated by several different plant UGTs (Messner et al., 2003; Brazier-Hicks and Edwards, 2005). Of the compounds tested, UGT73C9 only showed weak activity toward TCP when applied in 1 mm concentration. In contrast, UGT73C10, UGT73C11, UGT73C12, and UGT73C13 glucosylated TCP at 10 µm concentration (Fig. 5). The levels of oleanolic acid, hederagenin, and betulinic acid glucosides produced by these four UGTs were constantly higher than the levels of TCP glucosides, showing that sapogenins are better substrates. UGT73C10 and UGT73C11 showed weak activity toward quercetin and kaempferol at 100 µm concentration, while at 10 µm, glucosides could not be detected. In contrast, UGT73C12 and UGT73C13 clearly produced flavonol glucosides in assays with 100 µm quercetin or kaempferol, while at 10 µm, the glucosides were hardly detectable (Fig. 5). 24-Epi-brassinolide glucoside(s) were not observed with UGT73C11, whereas UGT73C13 catalyzed glucosylation of 24-epi-brassinolide to a product that comigrated with 24-epi-brassinolide glucoside, produced by A. thaliana UGT73C5 (Supplemental Fig. S3). None of the B. vulgaris UGTs glucosylated the phytosterols. A. thaliana UGT73B5 was included to represent a UGT73 from a different subfamily than UGT73C. UGT73B5 glucosylated TCP but neither of the sapogenins or other compounds tested (Supplemental Figs. S3 and S13).
UDP-Gal and UDP-GlcA were tested as alternative sugar donors. No glucuronides could be detected with any of the B. vulgaris UGTs when UDP-GlcA was used as sugar donor, but low activity was observed for UDP-Gal (Supplemental Fig. S4). 1H-NMR analysis revealed that the UDP-Gal stock contained traces of UDP-Glc, suggesting that the activity observed most likely originates from the UDP-Glc contamination (Thorsøe et al., 2005).
In summary, UGT73C10, UGT73C11, UGT73C12, and UGT73C13 preferentially glucosylate different oleanane and lupane sapogenins. Both UGT73C10 and UGT73C11 show high regiospecificity and substrate specificity by predominantly glucosylating the C3-hydroxyl group of sapogenins via an ether linkage. In comparison, UGT73C12 and UGT73C13 show lower substrate specificity and also glucosylate the sapogenin C28-carboxyl group via an ester bond. However, the ability to glucosylate at the C28-carboxyl group varied strongly: C28 glucosylation was abundant for betulinic acid and to a lesser extent for oleanolic acid and weakly for hederagenin. The similar enzymatic characteristics of UGT73C10 from the P-type and UGT73C11 from the G-type corroborate the phylogenetic reconstruction (Fig. 3), as do the characteristics of UGT73C12 from the P-type and UGT73C13 from the G-type. UGT73C9 apparently does not glucosylate any of the tested compounds besides the positive control substrate TCP, despite clustering with UGT73C10 and UGT73C11.
Kinetic Parameters of UGT73C11 and UGT73C13
Enzymes in the biosynthesis of plant specialized metabolism are generally characterized by low Km and high turnover rates. To evaluate the affinity and catalytic efficiencies of the two UGT clusters (Fig. 3), the kinetic parameters of UGT73C11 and UGT73C13 (both from the G-type) were determined toward hederagenin and oleanolic acid (Table I). Optimal assay conditions were at pH 8.6 for UGT73C11 and pH 7.9 for UGT73C13, with 1 mm dithiothreitol (DTT) as reductant. Purification of the recombinant UGTs was omitted due to decreasing specific activity upon metal chelate affinity-based purification. Instead, recombinant UGT amounts were quantified directly in crude E. coli protein extracts by taking advantage of an introduced N-terminal fused S-tag.
Table I. Kinetic parameters of UGT73C11 and UGT73C13 toward oleanolic acid and hederagenin.
| UGT | Sapogenin | Km | kcat | kcat/Km | Ki | Vmax |
|---|---|---|---|---|---|---|
| µm | s−1 | s−1 µm−1 | µm | nmol min−1 mg−1 | ||
| UGT73C11 | Oleanolic acid | 9.7 ± 2.2 | 0.816 | 0.084 | 817 ± 118 | |
| Hederagenin | 3.3 ± 0.8 | 0.389 | 0.118 | 390 ± 38 | ||
| UGT73C13 | Oleanolic acida | 12.5 ± 2.1 | 0.231 | 0.019 | 262 | 231 ± 21 |
| Oleanolic acidb | 7.6 ± 1.2 | 0.176 | 0.023 | 176 ± 7 | ||
| Hederagenin | 22.9 ± 4.8 | 0.131 | 0.006 | 131 ± 10 |
Kinetic parameters based on fit to the substrate inhibition equation.
Kinetic parameters based on fit to the Michaelis-Menten equation.
Most of the saturation curves (Supplemental Fig. S6) were hyperbolic and could be described by the Michaelis-Menten equations (for estimates, see Table I). However, for UGT73C13, the reaction velocities decreased when oleanolic acid concentrations exceeded 50 µm, indicating that it inhibits enzyme activity beyond this concentration. Similar substrate inhibition has previously been reported for other family 1 UDP-glycosyltransferases (Luukkanen et al., 2005; Ono et al., 2010). UGT73C11 has a 7-fold lower Km value and a 3-fold higher turnover rate (kcat value) with hederagenin than UGT73C13. The two UGTs have comparable Km values with oleanolic acid, but UGT73C11 has a 3.5-fold higher kcat value. The kinetic parameters, therefore, corroborate that UGT73C10 and UGT73C11 have higher affinity for sapogenins and more efficiently catalyze 3-O-glucosylation of oleanolic acid and hederagenin than UGT73C12 and UGT73C13. The low Km (less than 10 μm) and high kcat values of UGT73C11 are in comparable ranges to flavonol UGTs with their in planta acceptor substrates (Noguchi et al., 2007; Ono et al., 2010). The 1.4-fold higher catalytic efficiency (kcat/Km) for hederagenin than for oleanolic acid indicates that hederagenin is the preferred substrate for UGT73C11. Interestingly, UGT73C13 shows opposite substrate preference, as it has a 3-fold higher kcat/Km value for oleanolic acid than for hederagenin. The Km for UDP-Glc was estimated to be around 95 μm for UGT73C11 and 25 μm for UGT73C12 (Supplemental Fig. S5).
In Vitro Activities of the UGT73Cs toward B. vulgaris Sapogenin Mixtures
The saponin composition of B. vulgaris is complex, with more than 40 putative saponins detected in liquid chromatography-mass spectrometry analyses (Supplemental Figs. S7 and S8). The majority of these appear specific for either one of the two plant types, while others are present in variable amounts in both types. To evaluate if the UGTs can glucosylate other B. vulgaris sapogenins than oleanolic acid and hederagenin, crude saponin-containing extracts of both plant types were subjected to acidic hydrolysis to O-deglycosylate the saponins. Tandem mass spectrometry to n-fold (MSn) fragmentation analyses showed that the saccharide side chains of saponins in both B. vulgaris types consist of one to four hexosyl moieties, as concluded from the sequential loss of fragments with a mass of 162 D. The MSn fragmentation patterns of the most intense putative saponins in the G-type extract further indicate that they are derived from sapogenins with masses of 456 and 472 D, corresponding to oleanolic acid and hederagenin, as well as 458 and 488 D. In addition, a few less intense putative saponins appear to be derived from sapogenins with masses of 470, 474, and 476 D. In metabolite extracts of the P-type, the most abundant putative saponins originate from sapogenins with a mass of 474 D, followed by saponins derived from 458- and 488-D sapogenins. Only a few putative saponins based on sapogenins with masses of 456 and 472 D occur in this plant type.
After acid hydrolyzation, the putative saponins could not be detected, which confirms complete deglycosylation (Supplemental Figs. S9 and S10). The hydrolyzed G-type extract contained at least 40 structurally distinct compounds that are likely to be sapogenins, while in the P-type extract, 13 putative sapogenins were detected. Incubation of these extracts with UGT73C10, UGT73C11, UGT73C12, and UGT73C13 and UDP-Glc as sugar donor yielded numerous compounds that, based on MSn fragmentation patterns, were putative sapogenin monoglucosides (Supplemental Fig. S11). For both the G- and P-type sapogenin extracts, incubation with UGT73C10 and UGT73C11 reduced peak intensities of all putative sapogenins and resulted in the formation of the corresponding monoglucosides. In contrast, UGT73C12 and UGT73C13 appeared restricted to glucosylate only a subset of the putative sapogenins. Moreover, monoglucosides were produced at lower rates by UGT73C12 and UGT73C13 compared with UGT73C10 and UGT73C11. As expected, 3-O-β-d-Glc hederagenin (compound G27 in Supplemental Fig. S11) and 3-O-β-d-Glc oleanolic acid (compound G35 in Supplemental Fig. S11) were among the products formed from the G-type extract by UGT73C10 and UGT73C11. Surprisingly, only trace amounts of these two sapogenin monoglucosides were observed upon incubation of the G-type extract with UGT73C12 and UGT73C13. These UGTs additionally produced low amounts of diglucosides and compounds that may be kaempferol glucosides (according to their MSn fragmentation patterns). These findings corroborate that UGT73C12 and UGT73C13 have lower substrate specificity toward sapogenins than UGT73C10 and UGT73C11, which was also concluded from the in vitro enzyme assays (Fig. 5).
In Planta Saponin Accumulation Correlates with Organ-Specific Expression of the UGT73Cs
Steady-state transcript levels of the UGT73Cs were determined in leaves, petioles, and roots of 2-month-old G- and P-type B. vulgaris plants and compared with saponin accumulation in these organs. Metabolite extracts were evaluated by liquid chromatography-mass spectrometry and revealed a characteristic organ-specific saponin relative abundance in both plant types. Relative accumulation was highest in leaves, intermediate in petioles, and widely absent in roots (Fig. 6A; Supplemental Fig. S12). This pattern was consistent across the different plants tested.
Figure 6.
Comparison of relative saponin abundance and expression of the UGTs in different B. vulgaris organs. A, Relative saponin abundance in leaf, petiole, and root extracts of three G-type plants (G1–G3), based on the mean peak areas ± sd of the extracted ion chromatograms from liquid chromatography-mass spectrometry of the four insect resistance-correlated G-type saponins: hederagenin cellobioside (he-cell), oleanolic acid cellobioside (oa-cell), gypsogenin cellobioside (gy-cell), and 4-epi-hederagenin cellobioside (4e-cell). Overlaid base peak chromatograms of all liquid chromatography-mass spectrometry runs are provided in Supplemental Figure S12. B, Expression of UGT73C11 in the three G-type plants (G1–G3) and combined expression of UGT73C9 and UGT73C10 in three P-type plants (P1–P3), determined with primer set 1 relative to actin (ACT2). Values are means ± sd of technical duplicates. C, Corresponding expression analysis of UGT73C13 in G1 to G3 and UGT73C12 in P1 to P3, determined with primer set 2.
Two primer sets were used to quantify steady-state transcription levels of the UGTs by quantitative real-time PCR (Fig. 6, B and C). Due to the high sequence identities between UGT73C11 in the G-type and UGT73C10 and UGT73C9 in the P-type, it was not possible to design a primer that could differentiate between these three genes. Accordingly, primer set 1 amplifies UGT73C11 in the G-type, while in the P-type it amplifies simultaneously UGT73C9 and UGT73C10. Similarly, primer set 2 amplifies UGT73C13 from the G-type and UGT73C12 from the P-type. All plants showed the highest expression of UGT73C11 and UGT73C9/C10 in leaves, an up to 10-fold lower expression in petioles, and up to 200-fold lower expression in roots, despite some variation among individual plants tested. A similar expression pattern was observed for UGT73C13 and UGT73C12. In general, UGT73C11 and UGT73C9/C10 were expressed at a higher level than UGT73C13 and UGT73C12. The highest expression level of UGT73C13 was observed in plants with the lowest UGT73C11 expression. Since those plants were in a more progressed developmental stage (Supplemental Fig. S12), this suggests alternating expression regulation of the two genes during plant ontogenesis.
3-O-β-d-Glc Hederagenin Is a Feeding Deterrent against P. nemorum
The two diglucosides hederagenin and oleanolic acid cellobioside have previously been shown to deter feeding by P. nemorum (Nielsen et al., 2010). To determine if the corresponding monoglucosides have a similar effect, approximately 12.5 mg of 3-O-β-d-Glc hederagenin and 8.5 mg of 3-O-β-d-Glc oleanolic acid were produced in vitro with UGT73C10 (see above). Both compounds were painted on 92-mm2 radish (Raphanus sativus) leaf discs in doses of 3.75, 15, and 60 nmol and presented to P. nemorum adults of either the susceptible (ST; rr genotype) or resistant (AK; Rr genotype) line, and the area consumed was evaluated after 24 h (Fig. 1).
3-O-β-d-Glc hederagenin significantly reduced the leaf consumption by susceptible ST beetles, with dose-dependent reductions of 26%, 55%, and 92% in response to 3.75, 15, and 60 nmol per leaf disc, respectively (Fig. 7A; the reduction by 15 and 60 nmol was statistically significant [P < 0.005] when tested separately). A dose-dependent reduction of leaf consumption was also observed for the resistant AK line, with 16% and 67% reduction in response to 15 and 60 nmol, respectively (only the reduction by 60 nmol was significant when tested separately).
Figure 7.
Consumption of radish leaf discs painted with different amounts of 3-O-β-Glc hederagenin (A) and 3-O-β-Glc oleanolic acid (B) by susceptible ST and resistant AK lines of P. nemorum. Consumption is shown as mean total area consumed from two leaf discs (total area, 92 mm2) that were presented to one beetle (±1.96 se corresponding to a confidence interval of 95%). Assays with 3.75 nmol of 3-O-β-Glc oleanolic acid were omitted due to the low efficacy at higher doses.
3-O-β-d-Glc oleanolic acid had a significantly weaker effect on leaf consumption for both P. nemorum lines (Fig. 7B). Only the high dose of 60 nmol reduced consumption by the ST line (45% reduction), whereas there was no effect on the AK line at any dose. Feeding assays with 3.75 nmol were not conducted, as there was no significant effect with 15 nmol.
When tested in a joint linear mixed-effect model, there was a significant three-way interaction between sapogenin monoglucosides, their doses, and the P. nemorum lines, with a significance level of P < 0.0001. Thus, (1) 3-O-β-d-Glc hederagenin is more effective than 3-O-β-d-Glc oleanolic acid, (2) the feeding deterrence of the sapogenin monoglucosides is dose dependent, and (3) the efficacy toward the susceptible P. nemorum line is higher than toward the resistant line.
DISCUSSION
Saponin biosynthesis is not fully understood, nor is the relationship between the different chemical structures and their roles in plant defense. Here, we have identified two UGTs that specifically glucosylate sapogenins in the wild crucifer B. vulgaris. These UGTs have evolved to be specific for 3-O-glucosylation of sapogenins. Previously, UGTs that glucosylate sapogenins at the C28 carboxylic groups have been identified in Medicago truncatula (UGT73F3; Naoumkina et al., 2010) and in Saponaria vaccaria (UGT74M1; Meesapyodsuk et al., 2007). Monoglucosylated 3-O-β-d-Glc hederagenin, produced in vitro by one of the UGTs identified here, UGT73C10, is a strong feeding deterrent against P. nemorum, demonstrating that 3-O-glucosylation of saponins is essential for bioactivity. The UGTs are expressed in both a P. nemorum resistant and a susceptible type of B. vulgaris, which fits our observation that most, if not all, saponins in the P and G-types are 3-O-glucosylated. The presence of UGTs in both the plant types catalyzing 3-O-glucosylation sapogenins, and the genomic locations of genes coding for these UGTs outside QTL associated with resistance to P. nemorum, suggest that the difference in resistance between the two B. vulgaris types is determined by an earlier enzymatic step in saponin biosynthesis.
UGT73C10/C11: Two Neofunctionalized UDP-Glc:Sapogenin 3-O-Glucosyltransferases
Of the five UGTs we identified in B. vulgaris ssp. arcuata, UGT73C10 from the insect-susceptible P-type and UGT73C11 from the resistant G-type showed highest activity and specificity toward a wide range of sapogenins. Both enzymes exhibit high regiospecificity by preferably glucosylating the C3 hydroxyl group, which is in agreement with structures of saponins in both B. vulgaris types. Both enzymes, in contrast, were essentially inactive toward the flavonols and phytosterols tested. Their acceptor substrate specificity thus differs substantially from other characterized members of the UGT73C subfamily. UGT73C8 from M. truncatula glucosylates several (iso)flavonoids in vitro (Modolo et al., 2007). A. thaliana UGT73C6 was suggested to be a UDP-Glc:flavonol-3-O-glycoside-7-O-glucosyltransferase by Jones et al. (2003), based on in vitro activities and T-DNA knockout lines. Recent studies show that UGT73C6 is functionally similar to the well-studied UGT73C5, also from A. thaliana, in its ability to glucosylate brassinosteroids in overexpression lines (Husar et al., 2011). UGT73C5 in addition glucosylates numerous structurally diverse acceptor substrates (Lim et al., 2003, 2004; Poppenberger et al., 2003, 2005, 2006; Hou et al., 2004; Weis et al., 2006; Caputi et al., 2008). It was originally identified as a mycotoxin-detoxifying enzyme (Poppenberger et al., 2003), but recently, it was suggested to be involved in brassinosteroid homeostasis (Poppenberger et al., 2005). In our study, A. thaliana UGT73C5 also glucosylated oleanolic acid, hederagenin, and betulinic acid in vitro, providing further evidence for the promiscuity of this enzyme (Supplemental Fig. S13). However, it had substantially lower catalytic efficiency and regiospecificity toward oleanolic acid and hederagenin than UGT73C11 and UGT73C13 from B. vulgaris (Supplemental Fig. S13). A. thaliana is not known to produce triterpenoid saponins or sapogenins, although triterpenoids such as β-amyrin and lupeol accumulate in cuticular waxes of stems, siliques, and buds (Shan et al., 2008). Therefore, it is unlikely that the in vitro activities of UGT73C5 with sapogenins reflect an in planta function.
The broad substrate affinity commonly found for some UGTs has been proposed to enable flexibility in response to changes in metabolite profiles (Vogt and Jones, 2000). Specialized enzymes for new biosynthetic pathways may originate from broad progenitor enzymes and are generally characterized by having a lower Km (and thus higher substrate specificity) and higher catalytic efficiency (kcat/Km) than their more promiscuous progenitors (Jensen, 1976; Aharoni et al., 2005; Khersonsky and Tawfik, 2010). Ancestors of UGT73C10/C11 from B. vulgaris could thus have been promiscuous UGT73C5-like enzymes that evolved a more narrow specificity and higher efficiency for catalyzing sapogenin 3-O-glucosylation. Based on our analyses, UGT73C12/C13 have broader substrate and product specificities and could represent evolutionary intermediates to UGT73C10/C11 or UGTs specialized in glucosylation of yet unknown sapogenins in B. vulgaris.
Our phylogenetic reconstruction shows that the five B. vulgaris UGT73Cs indeed cluster separately from the UGT73Cs in A. thaliana, A. lyrata, and B. rapa (Fig. 3). It further suggests that UGT73C10, UGT73C11, and UGT73C9 originate from a gene duplication event after the split from A. thaliana and B. rapa and before the P and G-types separated. Another gene duplication separated UGT73C9 from UGT73C10, probably in the P-type after the P- and G-types split. Alternatively, this duplication occurred before the P-G bifurcation and the UGT73C9 copy was lost subsequently in the G-type.
Of the UGTs in our phylogenetic analysis, UGT73C9, UGT73C10, and UGT7311 showed clear signs of positive selection during their differentiation. This corroborates our biochemical data, which show that UGT73C10 and UGT73C11 have evolved to a new specialized function. In contrast, UGT73C12 and UGT73C13 showed no signs of selection, corroborating that they have not evolved new biochemical functions; this further suggests that they may be orthologs of A. thaliana UGT73C5 or UGT73C6. The observation that UGT73C9 is under positive selection questions the function of this UGT in saponin biosynthesis. Based on our biochemical data, UGT73C9 appears as an expressed pseudogene; however, the phylogenetic analysis indicates that the gene has been under positive selection. An alternative hypothesis is that the substrate for UGT73C9 was not included in our analysis. As the saponin profiles of P- and G-type B. vulgaris differ, UGT73C9 could possibly be involved in the differentiation of these.
Genes for the B. vulgaris UGTs were located in a genomic region syntenic to a part of A. thaliana chromosome 2, which contains a tandem repeat cluster of UGT73Cs. Our recent genome sequencing indicates that the B. vulgaris UGT73Cs identified here are also part of a repetitive cluster containing several UGT-like repeats and in higher number than the corresponding UGT73C cluster in A. thaliana. This supports that UGT73C10/C11 evolved via gene duplications from a broad-spectrum UGT73C in a common ancestor shared with A. thaliana, as discussed above. It further supports the idea that the evolution of novel bioactive metabolites often occurs via gene duplication and neofunctionalization (Osbourn, 2010; Weng et al., 2012) followed by increased specialization (Jensen, 1976; Aharoni et al., 2005; Khersonsky and Tawfik, 2010).
3-O-Glucosylation of Hederagenin Deters Feeding by P. nemorum
Monoglucosylation of hederagenin into 3-O-β-d-Glc hederagenin clearly suppressed feeding by P. nemorum. A similar but lower suppression was found for 3-O-β-d-Glc oleanolic acid. The diglucosylated forms of hederagenin and oleanolic acid (hederagenin cellobioside and oleanolic acid cellobioside) have previously been found to suppress feeding (Nielsen et al., 2010), in contrast to the aglycones (hederagenin and oleanolic acid). Our results now show that glucosylation with only a single glucosyl group is enough to affect herbivores. The amount of monoglucosides used in our feeding assays was comparable to natural levels of hederagenin cellobioside in B. vulgaris leaves (Shinoda et al., 2002), and our results thus demonstrate that 3-O-β-d-Glc hederagenin and 3-O-β-d-Glc oleanolic acid are biologically relevant feeding deterrents. Furthermore, the higher efficiency of hederagenin than oleanolic acid, in both their monoglycosylated and diglycosylated forms, shows that C23 hydroxylation in the hederagenin backbone increases this antifeedant effect.
The precise mechanism that enables glucosylated sapogenins to deter insects is not known. The dependency on glycosylation indicates that membrane perturbation plays a role, at least for P. nemorum. In agreement with this, saponins have been shown to damage the midgut epithelium of pea aphids (Acyrthosiphon pisum; De Geyter et al., 2012). Alternatively, glucosylated saponins may have a more adverse taste for insects than the corresponding sapogenins (Glendinning, 2002); however, P. nemorum larvae die from exposure to G-type leaves (Nielsen, 1997a).
Nielsen et al. (2010) suggested that cleavage of theβ-1,4-glycosidic bond in the cellobiosides by β-glucosidases allows resistant P. nemorum lines to feed on G-type B. vulgaris. This mechanism would be similar to what has been found for fungal adaptation to saponins (Osbourn et al., 1991; Wubben et al., 1996; Pareja-Jaime et al., 2008). Our findings, however, show that the monoglucosides of the saponins are also active and that resistance must be based on the ability to hydrolyze the glycosidic bond between the aglycone and the first linked sugar at the C3 position.
The resistance of G-type B. vulgaris against herbivorous insects, such as P. xylostella and susceptible P. nemorum, has previously been shown to depend on the presence of saponins, and especially hederagenin and oleanolic acid cellobioside, which are absent in the susceptible P-type (Shinoda et al., 2002; Agerbirk et al., 2003a; Kuzina et al., 2009; Nielsen et al., 2010). Therefore, the synthesis of saponins was initially thought to be unique to the G-type. However, saponins were recently also discovered in the susceptible P-type (Kuzina et al., 2011), and we are now pursuing their structure and identity. The presence of closely related UGTs in the G- and P-types of B. vulgaris, which have the same substrate specificity and regiospecificity, strongly indicates that the difference between resistance and susceptibility of the two B. vulgaris types is not caused by different UGTs, despite their obvious role in activating sapogenins by glucosylation. This is further substantiated by results from our QTL analysis, where the UGTs described here do not colocalize with resistance to P. nemorum or saponin identity (Kuzina et al., 2011). Instead, the difference in resistance between the G- and P-types must be determined at an earlier step in saponin biosynthesis, presumably during cyclation by OSCs or backbone decoration by cytochromes P450.
Evolution of Saponin Biosynthesis in Barbarea Species
The multitude of different putative sapogenins in the G- and P-types indicates that OSCs and P450s are responsible for much of the saponin diversity in this species and probably for the differences between the two plant types. The phylogeny of OSCs (Phillips et al., 2006; Augustin et al., 2011) suggests frequent changes in product spectra during evolution, which is supported by the drastic spectrum changes that may arise from only a few amino acid substitutions (Lodeiro et al., 2005). Changes in cytochrome P450 activity are also known to affect saponin profiles and activity. Carelli et al. (2011) showed that lack of a functional CYP716A12, which catalyzes C28 carboxylation of triterpenoid sapogenins, results in a complete loss of hemolytic saponins in M. truncatula. In contrast, nonhemolytic saponins were unaffected. The nonhemolytic saponins are derived from sapogenins that are not carboxylated at the C28 position, and MSn fragmentation of these revealed an aglycone fragment ion with a deduced mass of 474 D (Pollier et al., 2011). A similar fragmentation product was observed for P-type saponins and suggests that structurally similar sapogenins, with four hydroxyl groups but no C28 carboxylation, are present in this plant type. Different abilities to catalyze C28 oxygenation by cytochromes P450 could thus be involved in determining the different structures of G- and P-type saponins and thus their effect on insect herbivores.
The current hypothesis for the evolution of insect resistance in B. vulgaris suggests that it took place after the first species of the Barbarea genus had emerged (Agerbirk et al., 2003b; the age of this split is unknown at present). An OSC probably mutated to be able to catalyze the conversion of oxidosqualene into saponin precursors, which is in agreement with the presence of triterpenoids in A. thaliana. Later, UGTs must have evolved to become specific to the novel sapogenins produced by the resistant Barbarea species, as we have shown here. Whether the cytochromes P450 involved in saponin biosynthesis of Barbarea species have also specialized is not known. Much later, B. vulgaris differentiated into the G- and P-types, possibly during one of the last ice ages (Hauser et al., 2012; Toneatto et al., 2012). Thus, the two plant types are genetically and geographically differentiated, reproductively somewhat incompatible, and differ for several traits apart from insect resistance and saponin structure (Toneatto et al., 2010; Dalby-Brown et al., 2011). Thus, the most likely scenario suggests that the P-type lost resistance to P. nemorum during this allopatric separation. Our results here clearly show that this loss of insect resistance was not caused by a loss of UGT function. Instead, we have shown that UGTs of B. vulgaris have adapted to the earlier evolutionary gain of saponins in this species.
MATERIALS AND METHODS
Activity-Based cDNA Library Screening
Barbarea vulgaris var variegata (Chiltern Seeds) leaf RNA was used for first-strand synthesis with the ZAP-cDNA Synthesis Kit (Stratagene). The resulting cDNA was digested with XhoI, ligated into the predigested Uni-ZAP XR vector (Stratagene), and transformed into the Escherichia coli strain XL1-Blue MRF′ (Stratagene). After in vivo excision of pBluescript SK− phagemids from the Uni-ZAP XR vectors, the obtained E. coli colonies were combined in terrific broth (TB) medium and transferred to 96-well plates (approximately 100 colonies per well). The E. coli suspensions were incubated with shaking at 37°C for 3 h and then for 3 h with 0.1 mm isopropylthio-β-galactoside (IPTG). Cultures of individual wells were combined into batches (four wells per batch), and the bacterial cells were harvested by centrifugation. The bacterial cells were resuspended in 20 mm Tris-HCl, pH 7.5, and 2 mm DTT and lysed by sonication. Enzymatic activity was tested by incubating the lysates overnight at 30°C with 200 µm UDP-Glc and 175 µm oleanolic acid. Ethyl acetate extracts of the activity assays were analyzed by TLC on Silica Gel 60 F254 plates (5554; Merck), using chloroform:methanol:water (32:9:1) as mobile phase, and stained by spraying with 10% sulfuric acid in methanol followed by heating. Batches that showed oleanolic acid glucosylation activity were in additional screening rounds stepwise further diluted until a single active clone designated BvUGT1 was identified.
Cloning of BvUGT1 Homologs from B. vulgaris ssp. arcuata
Contigs representing fragments of BvUGT1 homologs were identified in a 454 pyrosequencing-generated transcriptomic G-type data set (Kuzina et al., 2011) using local BLASTX. Total RNA was extracted from leaves of G- and P-type B. vulgaris using the NucleoSpin RNA Plant kit (Macherey-Nagel) and 3′ RACE performed with the FirstChoice RLM-RACE kit (Ambion) according to the manufacturer’s protocol. The applied primers are listed in Supplemental Table S2.
The nucleotide sequences of UGT73C9, UGTC10, UGT73C11, UGT73C12, and UGT73C13 were cloned from genomic DNA of an F1 hybrid plant, which originated from crossings between G- and P-type plants (Kuzina et al., 2009), and ligated into pGEM-T Easy for sequencing.
PCRs for cloning were performed with Phusion High-Fidelity DNA Polymerase (Finnzymes), and PCRs for screening and A-tailing reactions were performed with Hotmaster Taq DNA Polymerase (5prime). A-tailing reactions were set up according to the pGEM-T Easy manual (Promega). Sequencing was performed by Eurofins MWG Operon.
Phylogenetic Analysis
UGT73 amino acid sequences were aligned (Supplemental Data Set S2) using MUSCLE and used to construct a maximum likelihood bootstrapped phylogenetic tree using MEGA (version 5.05; Jones, Taylor, and Thornton substitution model, uniform rates among sites, 100 bootstrap replications; Tamura et al., 2011). The A. thaliana lyrata and Brassica rapa UGTs, identified by BLAST searches at www.phytozome.net and www.brassica-rapa.org, have not been officially named and therefore are named here according to their grouping with Arabidopsis thaliana.
To test for signs of past selection on the UGTs, branch and site models were estimated using codeml in the PAML package (http://abacus.gene.ucl.ac.uk/software/paml.html). For positive selection between branches, the free-ratio model was compared with the one-ratio model and tested by comparing the twice log-likelihood difference between models to an χ2 distribution with 18 degrees of freedom. Seven site models were estimated: M0 (one ratio); M1 (nearly neutral; two categories); M2 (positive selection; three categories); M3 (discrete; three categories); M5 (γ; 10 categories); M7 (β; 10 categories); and M8 (β&ω > 1; 11 categories); these were tested as above with degrees of freedom corresponding to the differences in the number of parameters for the models tested.
Locating UGT73C9, UGT73C10, UGT73C11, UGT73C12, and UGT73C13 on the B. vulgaris Linkage Map
The five UGTs were mapped using the derived cleaved amplified polymorphic sequences or cleaved amplified polymorphic sequences technique. PCR was performed using genomic DNA of an F2 segregating population generated from a cross between P- and G-type B. vulgaris (Kuzina et al., 2009). PCR products obtained using primers mapPSfor and sepSrev (UGT73C9 to -C11), mapPSfor and sepIIrev (UGT73C12/C13), or Inf and dCapsAvaII (UGT73C11) were digested with EcoRV, BsaJI, AvaII, or PciI to discriminate between UGT73C9, UGT73C10, UGT73C11, and UGT73C13, respectively. Data were scored and analyzed as described by Kuzina et al. (2011).
Heterologous Expression of B. vulgaris UGT73Cs
N-terminally His-tagged expression constructs of UGT73C9, UGT73C10, UGT73C11, UGT73C12, and UGT73C13 were obtained by subcloning into the NheI and BamHI restriction sites of the pET28c vector (Novagen). N-terminally S-tag expression constructs of the five UGT73C ORFs were achieved by Gateway cloning into pJAM1786 (Luo et al., 2007).
For heterologous expression of the His-tag and S-tag constructs, expression vectors were transformed into the E. coli strain XJb(DE3) (Zymo Research). Expression was carried out in 25-mL Erlenmeyer flasks and started by inoculating 2 mL of Luria-Bertani medium, containing either 50 µg mL−1 kanamycin (His-tag constructs) or 100 µg mL−1 carbenicillin (S-tag constructs), with a single colony. A 12-h incubation phase at 30°C and 220 rpm was followed by the addition of 4 mL of TB medium containing appropriate selection antibiotics. Ara and IPTG were added to final concentrations of 3 and 0.1 mm, respectively, and the cultures were incubated for 24 h at 15°C and 220 rpm. For expression of the S-tag constructs, 1 µL of 50 mg mL−1 carbenicillin mL−1 culture was added approximately 12 h after the addition of TB medium.
Bacteria were harvested in aliquots corresponding to 2 mL of culture with an optical density of 8.0, resuspended in 750 µL aliquot−1 10 mm HEPES, pH 7.8, and stored at −80°C. Bacteria were lysed by thawing aliquots at room temperature. The viscosity of lysates was lowered by incubation with DNaseI (AppliChem) treatment (1 µg mL−1). Cell debris were removed by centrifugation, and supernatants were used as crude protein extracts for enzyme assays. Quantification of heterologously expressed enzymes, fused to an S-tag within E. coli crude protein extracts, was carried out using the FRETWorks S-tag assay kit (Novagen) according to the manufacturer’s protocol.
Substrate Specificity Assays
Enzyme assays to determine substrate specificity were performed in a final volume of 20 µL, containing 2 µL of E. coli crude protein extract with recombinant UGT73C9, UGT73C10, UGT73C11, UGT73C12, or UGT73C13 coupled to an S-tag. Reaction conditions were 25 mm TAPS-HCl, pH 8.6, 1 mm DTT, 7 µm UDP-Glc (Sigma-Aldrich), and 3.31 µm (0.74 kBq) UDP-[14C]Glc (Perkin-Elmer). Ethanol was removed from the UDP-[14C]Glc stock by evaporation prior to setting up the assays. Enzyme assays were started by addition of the acceptor substrates solubilized in dimethyl sulfoxide (DMSO) to final concentrations of 1 mm (only TCP), 100 µm, or 10 µm of the acceptor substrate and 6.25% to 10% (v/v) DMSO, respectively. Reactions were incubated for 30 min at 30°C and stopped by the addition of 130 µL of methanol. Precipitated proteins were removed by centrifugation. Solvent from the supernatant was removed with a vacuum concentrator, and metabolites were dissolved in 20 µL of 50% ethanol and analyzed by TLC. TLC plates were developed in ethyl acetate:methanol:formic acid:water (7.5:0.5:1:1), and radioactive bands were visualized using a STORM 840 PhosphorImager (Molecular Dynamics).
Acceptor substrates in this study were as follows: oleanolic acid (ICN Biomedical), hederagenin (Carl Roth), betulinic acid (Carl Roth), β-amyrin (Sigma-Aldrich), lupeol (Sigma-Aldrich), quercetin (Sigma-Aldrich), kaempferol (Fluka), and obtusifoliol, campesterol, sitosterol, stigmasterol, and 2,4,5-trichlorophenol (Sigma-Aldrich).
Determination of Enzyme Kinetic Parameters
Freshly lysed E. coli crude protein extracts were diluted in 10 mm TAPS-HCl, pH 8.0, and 10 mg mL−1 bovine serum albumin (BSA) to final concentrations of 5 ng µL−1 S-tag UGT73C11 and 45 ng µL−1 S-tag UGT73C13. The diluted crude protein extracts were applied in master mixtures with final reaction conditions as follows: 25 mm TAPS-HCl, pH 8.6 (UGT73C11) or pH 7.9 (UGT73C13), 1 mm DTT, 500 µm UDP-Glc, 2 mg mL−1 BSA, and 0.5 ng µL−1 UGT73C11 or 4.5 ng µL−1 UGT73C13. Enzyme assays were performed in a volume of 20 µL. Concentrations of UDP-[14C]Glc (Perkin-Elmer) in the total amount of UDP-Glc ranged from 3.31 µm (0.04 kBq µL−1) to 33.12 µm (0.37 kBq µL−1) to ensure sufficient signal intensity. Oleanolic acid and hederagenin were dissolved in 100% DMSO and assayed in duplicate in final concentrations ranging from 0.125 to 8 µm for UGT73C11 and 1.56 to 100 µm for UGT73C13, but with a constant final DMSO concentration of 6.25%. Reactions were preincubated for 3 min at 30°C prior to addition of the acceptor substrate. After incubation for 3 min at 30°C, enzymatic activities were stopped by the addition of 50 µL of ethyl acetate. Assays were extracted four times with 50 µL of ethyl acetate, and the solvent from the combined extractions was removed by evaporation in a vacuum concentrator. Metabolites were dissolved in 96% ethanol and analyzed by TLC. TLC plates were developed using dichloromethane:methanol:water (80:19:1) as mobile phase and visualized as described above. Products were quantified by codeveloping TLC plates with a defined oleanolic acid or hederagenin [14C]monoglucoside dilution series. Signal intensities were quantified using ImageQuant 5.0 (Molecular Dynamics). Km and Vmax values were calculated using SigmaPlot 11.0 (Systat Software) for nonlinear regression according to the Michaelis-Menten equation or the velocity equation for substrate inhibition.
14C-labeled monoglucosides were obtained by overnight incubation of 20 nmol of oleanolic acid and hederagenin with UGT73C11 at reaction conditions similar to those applied for the actual enzyme assays (500 µm UDP-Glc including 33.12 µm UDP-[14C]Glc [0.37 kBq µL−1]). Complete conversion of the aglycones was confirmed by TLC analysis of aliquots of these reactions.
Plant Material
B. vulgaris ssp. arcuata seeds were collected in natural populations in Denmark: G-type (Amager; 55°38′N, 12°34′E) and P-type (Tissø; 55°36′N, 11°18′E). Plants were grown at 20°C, 16 h of light/8 h of darkness, and 70% to 75% air humidity, fertilized once a week, and the soil was treated with Bactimos L (Abbott Laboratories) whenever necessary.
Comparison of Saponin Levels and in Planta Expression of UGT73Cs
To determine saponin levels, metabolites were extracted from 20 to 30 mg of ground, lyophilized leaf, petiole, and root material by boiling for 10 min with 37.5 µL of 55% ethanol per mg of tissue powder. Samples were cooled on ice and centrifuged to remove insoluble particles. Supernatants were kept for more than 2 h at −20°C and centrifuged to remove precipitates. Extracts were filtered (polyvinylidene difluoride; 0.45 µm) and transferred to glass sample vials for liquid chromatography-mass spectrometry analysis. An Agilent 1100 Series LC device (Agilent Technologies), equipped with a Gemini NX column (35°C; 2.0 × 150 mm, 3.5 µm; Phenomenex) and coupled to a Bruker HCT-Ultra ion-trap mass spectrometer (Bruker Daltonics), was used for spectrometric analysis. Mobile phases were eluent A, water with 0.1% (v/v) formic acid, and eluent B, acetonitrile with 0.1% (v/v) formic acid. The gradient program was as follows: 0 to 1 min, isocratic 12% B; 1 to 33 min, linear gradient 12% to 80% B; 33 to 35 min, linear gradient 80% to 99% B; 35 to 38 min, isocratic 99% B; 38 to 45 min, isocratic 12% B at a constant flow rate of 0.2 mL min−1. The detector was operated in negative electrospray mode and included tandem mass spectrometry to two stages (MS2) and three stages (MS3). Chromatograms were analyzed with DataAnalysis 4.0 (Bruker Daltonics), and saponin abundance was calculated based on summed extracted ion chromatograms of all adduct ions.
RNAwas extracted from 100 to 150 mg of ground leaf, petiole, and root material by incubation for 10 min with 900 µL of prewarmed hexadecyltrimethylammonium bromide extraction buffer (Chang et al., 1993) at 65°C and 660 rpm. After 2-fold extraction with 900 µL of chloroform-isoamyl alcohol, RNA was precipitated overnight (4°C) from the supernatant by the addition of LiCl to a final concentration of 2 m. Pellets were dissolved in 500 µL of sodium chloride-Tris-EDTA buffer (le Provost et al., 2007; prewarmed to 65°C) containing 0.1% SDS. RNA was extracted with chloroform-isoamyl alcohol and precipitated from the aqueous phase by adjusting the NaCl concentration to 0.67 m, adding 1 volume of isopropanol, and subsequent incubation for 5 h at −20°C. RNA pellets were washed with 70% ethanol (−20°C), dried, and redissolved in 30 µL of diethyl pyrocarbonate-treated water. The remaining genomic DNA was removed by on-column DNase treatment using the RNeasy Mini Kit (Qiagen). RNA extracts were assessed for purity and quantified with a NanoDrop ND-1000 (NanoDrop Technologies) and a 2100 Bioanalyzer (Agilent Technologies).
Reference gene sequences were obtained by mapping the 454 pyrosequencing-derived reads of G- and P-type leaf RNA preparations (V. Kuzina and S. Bak, unpublished data) to a data set consisting of all A. thaliana cDNA sequences (TAIR9_cdna_20090619) using the CLC Genomics Workbench (CLC bio). Two primer pairs, ACT2_for1/ACT2_rev1 and ACT2_for2/ACT2_rev2, were designed from reads mapped to A. thaliana ACT2 (AT3G18780). With the exception of four single-nucleotide polymorphisms in an intron region of the ACT2_for1/ACT2_rev1 product from the P-type, sequences derived for each primer set from the two plant types were 100% identical. The sequence identity of the two PCR products to the A. thaliana ACT2 ORF were 91% and 96%, respectively, while the encoded protein sequences were 100% identical to A. thaliana ACT2. Threshold cycle values of the two primer sets were almost identical in quantitative real-time PCR tests on leaf, petiole, and root tissues from a single G-type plant (±0.08–0.26). In addition, threshold cycle values across the three investigated tissues were found widely constant, with a range of ±0.31.
Five micrograms of RNA from each leaf, petiole, and root extract was applied in 100-µL reactions for cDNA synthesis using the iScript cDNA Synthesis Kit (Bio-Rad) according to the manufacturer’s instructions. quantitative real-time PCR experiments were performed with the DyNAmo Flash SYBR Green quantitative real-time PCR Kit (Finnzymes) in 20-µL reactions according to the manufacturer’s instructions by adding 1 µL of the cDNA preparations as template per reaction. Primer pairs were RTS_for and RTS_rev (UGT73C9 to -C11), RTII_for and RTII_rev (UGT73C12/C13), as well as ACT2_for1 and ACT2_rev1 (ACT2). Duplicates of each setup were run on a Qiagen Rotor-Gene Q Real-Time PCR cycler with settings for melting, annealing, extension, and acquiring of 10 s at 95°C, 10 s at 65°C, 20 s at 72°C, and 1 s at 76°C, respectively.
Quantitative real-time PCR experiments were analyzed using LinRegPCR (version 12.7; Ramakers et al., 2003; Ruijter et al., 2009). Relative expression values were calculated as the ratios of the starting concentrations (N0) given for the ACT2 reference and the corresponding UGT73C primer sets in the LinRegPCR output.
Extraction and Reglucosylation of B. vulgaris Sapogenins
Crude saponin extracts from the G- and P-type were obtained by boiling freshly harvested leaves for 10 min with 5 mL of 55% ethanol g−1 fresh leaf material. Extracts were cooled on ice, centrifuged to remove insoluble particles, and the cleared supernatant was stored at −20°C for more than 4 h. Precipitates were removed by centrifugation, and HCl was added to a final concentration of 1 m followed by incubation for 24 h at 99°C and 1,400 rpm. A 1.2-fold volume of 1 m Tris base was added to shift the pH to basic conditions, and ethanol concentrations were adjusted to 14%. Polyvinylpolypyrrolidone and BSA were added to final concentrations of 50 mg mL−1 and 10 mg mL−1, respectively, followed by six extractions each with one-tenth volume of ethyl acetate. The ethyl acetate fractions were combined, and solvent was removed in a vacuum concentrator. Metabolites were redissolved in 96% ethanol, and the polyvinylpolypyrrolidone/BSA-based purification step was repeated in one-tenth scale. Finally, the sapogenin-containing extracts were dissolved in 1 mL of 96% ethanol per initially applied 2.5 mL of hydrolyzed leaf extract.
Enzymatic activity assays were performed in a volume of 50 µL with reaction conditions of 25 mm TAPS, pH 8.6 (UGT73C9 to -C11), pH 7.9 (UGT73C12/C13), or pH 8.2 (combination of UGT73C9, UGT73C10, or UGT73C11 with UGT73C12 or UGT73C13), 1 mm DTT, 1 mm UDP-Glc, and with diluted E. coli crude protein extracts containing in total 750 ng of the recombinant UGT73C(s). Aliquots of the sapogenin-containing extracts were dried in a vacuum concentrator and redissolved in 1 µL of DMSO per 6.4 µL of the initial sapogenin-containing ethanol solution. Addition of 3.13 µL of the sapogenin-containing DMSO solutions was used to start reactions after 3 min of preincubation at 30°C. Reactions were incubated for 30 or 120 min at 30°C, and enzymatic activities were subsequently stopped by the addition of 325 µL of ice-cold methanol. Precipitated proteins were removed by centrifugation, and the supernatant was evaporated to dryness in a vacuum concentrator. The dried extracts were redissolved in 60 µL of 50% methanol, filtered (polyvinylidene difluoride; 0.45-µm pore diameter), and subjected to liquid chromatography-mass spectrometry analysis (see above).
Production of Hederagenin and Oleanolic Acid Monoglucosides for NMR and Bioassays
For large-scale production of hederagenin and oleanolic acid monoglucoside, four 2-L Erlenmeyer flasks, containing 250 mL of TB medium with 50 µg mL−1 kanamycin, were inoculated with fresh XJb(DE3) colonies harboring the pET28::UGT73C10 plasmid and incubated for 12 h at 30°C and 180 rpm. Addition of 500 mL of TB medium and adjustment of the final concentrations of kanamycin, Ara, and IPTG to 50 µg mL−1, 3 mm, and 0.1 mm, respectively, were followed by further incubation at 15°C and 140 rpm for 24 h. The bacteria were harvested by centrifugation, resuspended in 10 mm HEPES, pH 7.9, and frozen at −80°C. Lysis was achieved by thawing bacteria in a water bath at room temperature. DNA was degraded by treatment with DNase I (0.01 mg mL−1, 5 mm MgCl2, and 1 mm CaCl2). Cell debris were removed by centrifugation, and the supernatant was adjusted to 20 mm HEPES, pH 7.9, and 500 mm NaCl prior to the addition of 3 mL of equilibrated HIS-Select Nickel Affinity Gel (Sigma-Aldrich). One hour of incubation at 4°C was followed by removal of the supernatant and three times washing of the affinity gel with 20 mm HEPES, pH 7.9, and 500 mm NaCl and once with 25 mm TAPS, pH 8.6, and 1 mm DTT. Enzymatic reactions were set up in 100-mL glass flasks at a final volume of 50 mL. The reaction conditions were 25 mm TAPS, pH 8.6, 1 mm DTT, and 750 µm UDP-Glc. Approximately 1.5 mL of UGT73C10-loaded affinity gel was added to each reaction mixture, and enzymatic reactions were started by the addition of 10 mg of hederagenin (Extrasynthese) and oleanolic acid (Extrasynthese) dissolved in 3.125 mL of DMSO. Reaction mixtures were incubated at 37°C and 150 rpm, and progressing glucosylation of the two sapogenins was monitored by TLC analysis of 20-µL aliquots.
Hederagenin and oleanolic acid monoglucosides were extracted with ethyl acetate and, after evaporation of the solvent in a vacuum concentrator, dissolved in 60% to 70% DMSO prior to application to preparative HPLC for further purification. An Agilent 1200 series preparative HPLC system (Agilent Technologies), fitted with a Phenomenex Synergi 4µ Hydro-RP column (21.2 × 250 mm, 4 μm, 80 Å; Phenomenex), was used for this. Elution was carried out using a mobile phase containing acetonitrile and water with 0.01% trifluoroacetic acid. The gradient protocol was as follows: 5% acetonitrile for 5 min, linear gradient from 5% to 30% acetonitrile for 5 min, linear gradient from 30% to 100% acetonitrile for 50 min, and 100% acetonitrile for 5 min, at a constant flow rate of 15 mL min−1. A diode array detector was used to monitor the elution of compounds by their UV absorption at 200 nm. Fractions containing oleanolic acid and hederagenin glucosides were collected and evaporated to dryness using a vacuum concentrator.
The purified hederagenin and oleanolic acid monoglucosides were dissolved in NMR-suitable methanol-d4 (Sigma-Aldrich), and NMR spectra were recorded at room temperature on a Bruker Avance DSX 500-MHz NMR spectrometer (Bruker Daltonics) equipped with a broadband inverse probe. Acquired data were calibrated according to the residual solvent peaks at 3.31 ppm for 1H spectra and 49.01 ppm for 13C spectra. For structural elucidation of the two monoglucosides, 1-D 1H and 13C as well as 2-D COSY, TOCSY, and HSQC experiments were performed and compared with corresponding spectra of oleanolic acid and hederagenin and reported NMR data of structurally related compounds (Supplemental Data Set S1).
Phyllotreta nemorum Feeding Assays
Nonchoice feeding assays were performed as described previously by Nielsen et al. (2010). Briefly, purified 3-O-β-d-Glc hederagenin and 3-O-β-d-Glc oleanolic acid were in final concentrations of 2, 0.5, and 0.125 mm dissolved in 75% ethanol. Sapogenin monoglucoside solution (15 µL) was painted on both sides of 95-mm2 radish (Raphanus sativus) leaf discs, which resulted in doses of 60 nmol (632 pmol mm−2), 15 nmol (158 pmol mm−2), and 3.75 nmol (39 pmol mm−2) sapogenin monoglucoside per leaf disc. Control leaf discs were treated with solvent only. Two identically treated leaf discs were exposed to one beetle for 24 h. Consumed leaf area was measured with a stereomicroscope. For the origin and maintenance of the two flea beetle (P. nemorum) lines, see Nielsen et al. (2010).
Results were analyzed using the R software package (www.r-project.org). The linear effect model allowed for a possible correlation between measurements from the same beetle. The starting model included a three-way interaction between beetle line, compound type, and dose; a 5% significance level was used for model reduction tests.
Sequence data from this article can be found in the GenBank/EMBL data libraries under accession numbers JQ291611 (BvUGT1), JQ291612 (UGT73C9), JQ291613 (UGT73C10), JQ291614 (UGT73C11), JQ291615 (UGT73C12), and JQ291616 (UGT73C13).
Supplemental Data
The following materials are available in the online version of this article.
Supplemental Figure S1. Comparison of oleanolic acid glucosylation products after long-term incubation of oleanolic acid with UGT73C10, UGT73C11, UGT73C12, and UGT73C13 with oleanolic acid cellobioside.
Supplemental Figure S2. Alkaline hydrolysis (saponification) of betulinic acid glucosylation products derived from UGT73C13 activity.
Supplemental Figure S3. Comparison of UGTs from B. vulgaris (UGT73C11, UGT73C13, and UGT73C9) and A. thaliana (UGT73C5 and UGT73B5) in their activity toward hederagenin, 24-epi-brassinolide, and TCP.
Supplemental Figure S4. Comparison of UDP-Glc and UDP-Gal as sugar donor substrates of UGT73C10.
Supplemental Figure S5. Determination of Km values of UDP-Glc for UGT73C11 and UGT73C12.
Supplemental Figure S6. Kinetics of UGT73C11 and UGT73C13 with oleanolic acid and hederagenin as acceptor substrates.
Supplemental Figure S7. Liquid chromatography-mass spectrometry analysis of a G-type B. vulgaris metabolite extracted with 55% ethanol.
Supplemental Figure S8. Liquid chromatography-mass spectrometry analysis of a P-type B. vulgaris metabolite extracted with 55% ethanol.
Supplemental Figure S9. Liquid chromatography-mass spectrometry analysis of an acidic hydrolyzed G-type B. vulgaris metabolite extract.
Supplemental Figure S10. Liquid chromatography-mass spectrometry analysis of an acidic hydrolyzed P-type B. vulgaris metabolite extract.
Supplemental Figure S11. Glucosylation activity of UGT73C9 to UGT73C13 toward G-type and P-type B. vulgaris sapogenin extracts.
Supplemental Figure S12. Overlaid Liquid chromatography-mass spectrometry analyses of metabolite extracts from the B. vulgaris plants used for the saponin abundance and UGT73C9 to -C13 expression correlation analysis.
Supplemental Figure S13. Comparison of UGTs from B. vulgaris (UGT73C11, UGT73C13, and UGT73C9) and A. thaliana (UGT73C5 and UGT73B5) in their activity toward sapogenins.
Supplemental Table S1. Amino acid and nucleotide sequence identities of UGT73s used in the phylogenetic analysis.
Supplemental Table S2. Primers used in this study.
Supplemental Data Set S1. Structure elucidation of hederagenin and oleanolic acid monoglucosides based on 1-D 1H- and 13C- and 2-D TOCSY-, COSY-, and HSQC-NMR data.
Supplemental Data Set S2. Multiple sequence alignment, amino acid sequences, and nucleotide sequences used for the phylogenetic analysis.
Acknowledgments
We are grateful to Rubini Kannangara for helpful discussions and commenting on the manuscript. Mohammed Saddik Motawie and Henrik Toft Simonsen are thanked for consulting in chemical aspects, and Tamara van Mölken is thanked for P. nemorum images. Peter McKenzie is acknowledged for naming the UGTs according to the UGT nomenclature. Mika Zagrobelny is thanked for help and discussions on the use of codeml in the PAML package.
Glossary
- ORF
open reading frame
- TLC
thin-layer chromatography
- QTL
quantitative trait loci
- 1-D
one-dimensional
- 2-D
two-dimensional
- LC
liquid chromatography
- MS
mass spectrometry
- TCP
2,4,5-trichlorophenol
- DTT
dithiothreitol
- qPCR
quantitative PCR
- OSC
oxidosqualene cyclase
- IPTG
isopropylthio-β-galactoside
- DMSO
dimethyl sulfoxide
- BSA
bovine serum albumin
- cDNA
complementary DNA
References
- Agerbirk N, Olsen CE, Bibby BM, Frandsen HO, Brown LD, Nielsen JK, Renwick JAA. (2003a) A saponin correlated with variable resistance of Barbarea vulgaris to the diamondback moth Plutella xylostella. J Chem Ecol 29: 1417–1433 [DOI] [PubMed] [Google Scholar]
- Agerbirk N, Ørgaard M, Nielsen JK. (2003b) Glucosinolates, P. nemorum resistance, and leaf pubescence as taxonomic characters in the genus Barbarea (Brassicaceae). Phytochemistry 63: 69–80 [DOI] [PubMed] [Google Scholar]
- Aharoni A, Gaidukov L, Khersonsky O, McQ Gould S, Roodveldt C, Tawfik DS. (2005) The ‘evolvability’ of promiscuous protein functions. Nat Genet. 371: 73–76 [DOI] [PubMed] [Google Scholar]
- Augustin JM, Kuzina V, Andersen SB, Bak S. (2011) Molecular activities, biosynthesis and evolution of triterpenoid saponins. Phytochemistry 72: 435–457 [DOI] [PubMed] [Google Scholar]
- Badenes-Perez FR, Shelton AM, Nault BA. (2005) Using yellow rocket as a trap crop for diamondback moth (Lepidoptera: Plutellidae). J Econ Entomol 98: 884–890 [DOI] [PubMed] [Google Scholar]
- Brazier-Hicks M, Edwards R. (2005) Functional importance of the family 1 glucosyltransferase UGT72B1 in the metabolism of xenobiotics in A. thaliana thaliana. Plant J 42: 556–566 [DOI] [PubMed] [Google Scholar]
- Caputi L, Lim E-K, Bowles DJ. (2008) Discovery of new biocatalysts for the glycosylation of terpenoid scaffolds. Chemistry 14: 6656–6662 [DOI] [PubMed] [Google Scholar]
- Carelli M, Biazzi E, Panara F, Tava A, Scaramelli L, Porceddu A, Graham N, Odoardi M, Piano E, Arcioni S, et al. (2011) Medicago truncatula CYP716A12 is a multifunctional oxidase involved in the biosynthesis of hemolytic saponins. Plant Cell 23: 3070–3081 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chang S, Puryear J, Cairney J. (1993) A simple and efficient method for isolating RNA from pine trees. Plant Mol Biol Rep 11: 113–116 [Google Scholar]
- Dalby-Brown L, Olsen CE, Nielsen JK, Agerbirk N. (2011) Polymorphism for novel tetraglycosylated flavonols in an eco-model crucifer, Barbarea vulgaris. J Agric Food Chem 59: 6947–6956 [DOI] [PubMed] [Google Scholar]
- De Geyter E, Smagghe G, Rahbé Y, Geelen D. (2012) Triterpene saponins of Quillaja saponaria show strong aphicidal and deterrent activity against the pea aphid Acyrthosiphon pisum. Pest Manag Sci 68: 164–169 [DOI] [PubMed] [Google Scholar]
- de Jong PW, Breuker CJ, Vos H, Vermeer KMCA, Oku K, Verbaarschot P, Nielsen JK, Brakefield PM. (2009) Genetic differentiation between resistance phenotypes in the phytophagous P. nemorum, Phyllotreta nemorum. J Insect Sci 9: 1–8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- De Leo M, De Tommasi N, Sanogo R, D’Angelo V, Germanò MP, Bisignano G, Braca A. (2006) Triterpenoid saponins from Pteleopsis suberosa stem bark. Phytochemistry 67: 2623–2629 [DOI] [PubMed] [Google Scholar]
- Dowd PF, Berhow MA, Johnson ET. (2011) Differential activity of multiple saponins against omnivorous insects with varying feeding preferences. J Chem Ecol 37: 443–449 [DOI] [PubMed] [Google Scholar]
- Glendinning JI. (2002) How do herbivorous insects cope with noxious secondary plant compounds in their diet? Entomol Exp Appl 104: 15–25 [Google Scholar]
- Hauser TP, Toneatto F, Nielsen JK. (2012) Genetic and geographic structure of an insect resistant and a susceptible type of Barbarea vulgaris in western Europe. Evol Ecol 26: 611–624 [Google Scholar]
- Hou B, Lim E-K, Higgins GS, Bowles DJ. (2004) N-Glucosylation of cytokinins by glycosyltransferases of A. thaliana thaliana. J Biol Chem 279: 47822–47832 [DOI] [PubMed] [Google Scholar]
- Husar S, Berthiller F, Fujioka S, Rozhon W, Khan M, Kalaivanan F, Elias L, Higgins GS, Li Y, Schuhmacher R, et al. (2011) Overexpression of the UGT73C6 alters brassinosteroid glucoside formation in A. thaliana thaliana. BMC Plant Biol 11: 51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jensen RA. (1976) Enzyme recruitment in evolution of new function. Annu Rev Microbiol 30: 409–425 [DOI] [PubMed] [Google Scholar]
- Jones P, Messner B, Nakajima J, Schäffner AR, Saito K. (2003) UGT73C6 and UGT78D1, glycosyltransferases involved in flavonol glycoside biosynthesis in A. thaliana thaliana. J Biol Chem 278: 43910–43918 [DOI] [PubMed] [Google Scholar]
- Khersonsky O, Tawfik DS. (2010) Enzyme promiscuity: a mechanistic and evolutionary perspective. Annu Rev Biochem 79: 471–505 [DOI] [PubMed] [Google Scholar]
- Kuzina V, Ekstrøm CT, Andersen SB, Nielsen JK, Olsen CE, Bak S. (2009) Identification of defense compounds in Barbarea vulgaris against the herbivore Phyllotreta nemorum by an ecometabolomic approach. Plant Physiol 151: 1977–1990 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuzina V, Nielsen JK, Augustin JM, Torp AM, Bak S, Andersen SB. (2011) Barbarea vulgaris linkage map and quantitative trait loci for saponins, glucosinolates, hairiness and resistance to the herbivore Phyllotreta nemorum. Phytochemistry 72: 188–198 [DOI] [PubMed] [Google Scholar]
- le Provost G, Herrera R, Paiva JA, Chaumeil P, Salin F, Plomion C. (2007) A micromethod for high throughput RNA extraction in forest trees. Biol Res 40: 291–297 [PubMed] [Google Scholar]
- Lim E-K, Ashford DA, Hou B, Jackson RG, Bowles DJ. (2004) A. thaliana glycosyltransferases as biocatalysts in fermentation for regioselective synthesis of diverse quercetin glucosides. Biotechnol Bioeng 87: 623–631 [DOI] [PubMed] [Google Scholar]
- Lim E-K, Baldauf S, Li Y, Elias L, Worrall D, Spencer SP, Jackson RG, Taguchi G, Ross J, Bowles DJ. (2003) Evolution of substrate recognition across a multigene family of glycosyltransferases in A. thaliana. Glycobiology 13: 139–145 [DOI] [PubMed] [Google Scholar]
- Lodeiro S, Schulz-Gasch T, Matsuda SPT. (2005) Enzyme redesign: two mutations cooperate to convert cycloartenol synthase into an accurate lanosterol synthase. J Am Chem Soc 127: 14132–14133 [DOI] [PubMed] [Google Scholar]
- Luo J, Nishiyama Y, Fuell C, Taguchi G, Elliott K, Hill L, Tanaka Y, Kitayama M, Yamazaki M, Bailey P, et al. (2007) Convergent evolution in the BAHD family of acyl transferases: identification and characterization of anthocyanin acyl transferases from A. thaliana thaliana. Plant J 50: 678–695 [DOI] [PubMed] [Google Scholar]
- Luukkanen L, Taskinen J, Kurkela M, Kostiainen R, Hirvonen J, Finel M. (2005) Kinetic characterization of the 1A subfamily of recombinant human UDP-glucuronosyltransferases. Drug Metab Dispos 33: 1017–1026 [DOI] [PubMed] [Google Scholar]
- Mackenzie PI, Owens IS, Burchell B, Bock KW, Bairoch A, Bélanger A, Fournel-Gigleux S, Green M, Hum DW, Iyanagi T, et al. (1997) The UDP glycosyltransferase gene superfamily: recommended nomenclature update based on evolutionary divergence. Pharmacogenetics 7: 255–269 [DOI] [PubMed] [Google Scholar]
- Meesapyodsuk D, Balsevich J, Reed DW, Covello PS. (2007) Saponin biosynthesis in Saponaria vaccaria: cDNAs encoding β-amyrin synthase and a triterpene carboxylic acid glucosyltransferase. Plant Physiol 143: 959–969 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Messner B, Thulke O, Schäffner AR. (2003) A. thaliana glucosyltransferases with activities toward both endogenous and xenobiotic substrates. Planta 217: 138–146 [DOI] [PubMed] [Google Scholar]
- Modolo LV, Blount JW, Achnine L, Naoumkina MA, Wang X, Dixon RA. (2007) A functional genomics approach to (iso)flavonoid glycosylation in the model legume Medicago truncatula. Plant Mol Biol 64: 499–518 [DOI] [PubMed] [Google Scholar]
- Musende AG, Eberding A, Wood C, Adomat H, Fazli L, Hurtado-Coll A, Jia W, Bally MB, Guns ET. (2009) Pre-clinical evaluation of Rh2 in PC-3 human xenograft model for prostate cancer in vivo: formulation, pharmacokinetics, biodistribution and efficacy. Cancer Chemother Pharmacol 64: 1085–1095 [DOI] [PubMed] [Google Scholar]
- Naoumkina MA, Modolo LV, Huhman DV, Urbanczyk-Wochniak E, Tang Y, Sumner LW, Dixon RA. (2010) Genomic and coexpression analyses predict multiple genes involved in triterpene saponin biosynthesis in Medicago truncatula. Plant Cell 22: 850–866 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nielsen JK. (1997a) Variation in defences of the plant Barbarea vulgaris and in counteradaptations by the P. nemorum Phyllotreta nemorum. Entomol Exp Appl 82: 25–35 [Google Scholar]
- Nielsen JK. (1997b) Genetics of the ability of Phyllotreta nemorum larvae to survive in an atypical host plant, Barbarea vulgaris ssp. arcuata. Entomol Exp Appl 82: 37–44 [Google Scholar]
- Nielsen JK. (1999) Specificity of a Y-linked gene in the P. nemorum Phyllotreta nemorum for defences in Barbarea vulgaris. Entomol Exp Appl 91: 359–368 [Google Scholar]
- Nielsen JK. (2012) Non-random segregation of an autosomal gene in males of the P. nemorum, Phyllotreta nemorum: implications for colonization of a novel host plant. Entomol Exp Appl 143: 301–312 [Google Scholar]
- Nielsen JK, Nagao T, Okabe H, Shinoda T. (2010) Resistance in the plant, Barbarea vulgaris, and counter-adaptations in P. nemorum mediated by saponins. J Chem Ecol 36: 277–285 [DOI] [PubMed] [Google Scholar]
- Nihei K-I, Ying B-P, Murakami T, Matsuda H, Hashimoto M, Kubo I. (2005) Pachyelasides A-D, novel molluscicidal triterpene saponins from Pachyelasma tessmannii. J Agric Food Chem 53: 608–613 [DOI] [PubMed] [Google Scholar]
- Noguchi A, Saito A, Homma Y, Nakao M, Sasaki N, Nishino T, Takahashi S, Nakayama T. (2007) A UDP-glucose:isoflavone 7-O-glucosyltransferase from the roots of soybean (Glycine max) seedlings: purification, gene cloning, phylogenetics, and an implication for an alternative strategy of enzyme catalysis. J Biol Chem 282: 23581–23590 [DOI] [PubMed] [Google Scholar]
- Ono E, Homma Y, Horikawa M, Kunikane-Doi S, Imai H, Takahashi S, Kawai Y, Ishiguro M, Fukui Y, Nakayama T. (2010) Functional differentiation of the glycosyltransferases that contribute to the chemical diversity of bioactive flavonol glycosides in grapevines (Vitis vinifera). Plant Cell 22: 2856–2871 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Osbourn A. (2010) Gene clusters for secondary metabolic pathways: an emerging theme in plant biology. Plant Physiol 154: 531–535 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Osbourn AE, Clarke BR, Dow JM, Daniels MJ. (1991) Partial characterization of avenacinase from Gaeumannomyces graminis var. avenae. Physiol Mol Plant Pathol 38: 301–312 [Google Scholar]
- Papadopoulou K, Melton RE, Leggett M, Daniels MJ, Osbourn AE. (1999) Compromised disease resistance in saponin-deficient plants. Proc Natl Acad Sci USA 96: 12923–12928 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paquette S, Møller BL, Bak S. (2003) On the origin of family 1 plant glycosyltransferases. Phytochemistry 62: 399–413 [DOI] [PubMed] [Google Scholar]
- Pareja-Jaime Y, Roncero MIG, Ruiz-Roldán MC. (2008) Tomatinase from Fusarium oxysporum f. sp. lycopersici is required for full virulence on tomato plants. Mol Plant Microbe Interact 21: 728–736 [DOI] [PubMed] [Google Scholar]
- Phillips DR, Rasbery JM, Bartel B, Matsuda SPT. (2006) Biosynthetic diversity in plant triterpene cyclization. Curr Opin Plant Biol 9: 305–314 [DOI] [PubMed] [Google Scholar]
- Pollier J, Morreel K, Geelen D, Goossens A. (2011) Metabolite profiling of triterpene saponins in Medicago truncatula hairy roots by liquid chromatography Fourier transform ion cyclotron resonance mass spectrometry. J Nat Prod 74: 1462–1476 [DOI] [PubMed] [Google Scholar]
- Poppenberger B, Berthiller F, Bachmann H, Lucyshyn D, Peterbauer C, Mitterbauer R, Schuhmacher R, Krska R, Glössl J, Adam G. (2006) Heterologous expression of A. thaliana UDP-glucosyltransferases in Saccharomyces cerevisiae for production of zearalenone-4-O-glucoside. Appl Environ Microbiol 72: 4404–4410 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Poppenberger B, Berthiller F, Lucyshyn D, Sieberer T, Schuhmacher R, Krska R, Kuchler K, Glössl J, Luschnig C, Adam G. (2003) Detoxification of the Fusarium mycotoxin deoxynivalenol by a UDP-glucosyltransferase from A. thaliana thaliana. J Biol Chem 278: 47905–47914 [DOI] [PubMed] [Google Scholar]
- Poppenberger B, Fujioka S, Soeno K, George GL, Vaistij FE, Hiranuma S, Seto H, Takatsuto S, Adam G, Yoshida S, et al. (2005) The UGT73C5 of A. thaliana thaliana glucosylates brassinosteroids. Proc Natl Acad Sci USA 102: 15253–15258 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramakers C, Ruijter JM, Deprez RHL, Moorman AFM. (2003) Assumption-free analysis of quantitative real-time polymerase chain reaction (PCR) data. Neurosci Lett 339: 62–66 [DOI] [PubMed] [Google Scholar]
- Renwick JAA. (2002) The chemical world of crucivores: lures, treats and traps. Entomol Exp Appl 104: 35–42 [Google Scholar]
- Ruijter JM, Ramakers C, Hoogaars WMH, Karlen Y, Bakker O, van den Hoff MJB, Moorman AFM. (2009) Amplification efficiency: linking baseline and bias in the analysis of quantitative PCR data. Nucleic Acids Res 37: e45. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Senatore F, D’Agostino M, Dini I. (2000) Flavonoid glycosides of Barbarea vulgaris L. (Brassicaceae). J Agric Food Chem 48: 2659–2662 [DOI] [PubMed] [Google Scholar]
- Shan H, Wilson WK, Phillips DR, Bartel B, Matsuda SPT. (2008) Trinorlupeol: a major nonsterol triterpenoid in A. thaliana. Org Lett 10: 1897–1900 [DOI] [PubMed] [Google Scholar]
- Shinoda T, Nagao T, Nakayama M, Serizawa H, Koshioka M, Okabe H, Kawai A. (2002) Identification of a triterpenoid saponin from a crucifer, Barbarea vulgaris, as a feeding deterrent to the diamondback moth, Plutella xylostella. J Chem Ecol 28: 587–599 [DOI] [PubMed] [Google Scholar]
- Sun H-X, Xie Y, Ye Y-P. (2009) Advances in saponin-based adjuvants. Vaccine 27: 1787–1796 [DOI] [PubMed] [Google Scholar]
- Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S. (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28: 2731–2739 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thorsøe KS, Bak S, Olsen CE, Imberty A, Breton C, Lindberg Møller B. (2005) Determination of catalytic key amino acids and UDP sugar donor specificity of the cyanohydrin glycosyltransferase UGT85B1 from Sorghum bicolor: molecular modeling substantiated by site-specific mutagenesis and biochemical analyses. Plant Physiol 139: 664–673 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Toneatto F, Hauser TP, Nielsen JK, Ørgaard M. (2012) Genetic diversity and similarity in the Barbarea vulgaris complex (Brassicaceae). Nord J Bot 30: 506–512 [Google Scholar]
- Toneatto F, Nielsen JK, Ørgaard M, Hauser TP. (2010) Genetic and sexual separation between insect resistant and susceptible Barbarea vulgaris plants in Denmark. Mol Ecol 19: 3456–3465 [DOI] [PubMed] [Google Scholar]
- Vogt T, Jones P. (2000) Glycosyltransferases in plant natural product synthesis: characterization of a supergene family. Trends Plant Sci 5: 380–386 [DOI] [PubMed] [Google Scholar]
- Weis M, Lim E-K, Bruce N, Bowles D. (2006) Regioselective glucosylation of aromatic compounds: screening of a recombinant glycosyltransferase library to identify biocatalysts. Angew Chem Int Ed Engl 45: 3534–3538 [DOI] [PubMed] [Google Scholar]
- Weng JK, Philippe RN, Noel JP. (2012) The rise of chemodiversity in plants. Science 336: 1667–1670 [DOI] [PubMed] [Google Scholar]
- Wubben JP, Price KR, Daniels MJ, Osbourn AE. (1996) Detoxification of oat leaf saponins by Septoria avenae. Phytopathology 86: 986–992 [Google Scholar]