Abstract
Methylation of 3′-terminal nucleotides of miRNA/miRNA* is part of miRNAs biogenesis in plants but is not found in animals. In Arabidopsis thaliana this reaction is carried out by a multidomain AdoMet-dependent 2′-O-methyltransferase HEN1. Using deletion and structure-guided mutational analysis, we show that the double-stranded RNA-binding domains R1 and R2 of HEN1 make significant but uneven contributions to substrate RNA binding, and map residues in each domain responsible for this function. Using GST pull-down assays and yeast two-hybrid analysis we demonstrate direct HEN1 interactions, mediated by its FK506-binding protein-like domain and R2 domain, with the microRNA biogenesis protein HYL1. Furthermore, we find that HEN1 forms a complex with DICER-LIKE 1 (DCL1) ribonuclease, another key protein involved in miRNA biogenesis machinery. In contrast, no direct interaction is detectable between HEN1 and SERRATE. On the basis of these findings, we propose a mechanism of plant miRNA maturation which involves binding of the HEN1 methyltransferase to the DCL1•HYL1•miRNA complex excluding the SERRATE protein.
INTRODUCTION
MicroRNAs (miRNAs) are small 20–24 nucleotide RNAs involved in post-transcriptional regulation of gene expression in higher eukaryotes. The majority of proteins responsible for biogenesis of active microRNAs are strongly conserved between animals and plants; however, the maturation pathway has specific features in each kingdom (1). In animals, long primary miRNAs are successively cleaved by Drosha and Dicer ribonucleases to yield small ∼22 nt miRNA/miRNA* duplexes bearing 3′-dinucleotide overhangs. In contrast, a single DICER-LIKE 1 (DCL1) ribonuclease performs the cleavage of both arms of the predecessor stem-loop RNA in plant cells (2). The excision of proper miRNA duplexes involves physical interaction with additional RNA-binding partners, such as the zinc finger protein SERRATE (SE, a homolog of animal ARS2) and the double-stranded RNA-binding protein HYL1, also known as DRB1, a homolog of animal RDE-4 and R2D2 (3–6), in nuclear dicing or D-bodies (7–9) where they form a plant miRNA processing complex, called microprocessor (10). In contrast to animals, nearly all plant miRNA/miRNA* duplexes are methylated at their 3′-termini by the small RNA 2′-O-methyltransferase HEN1 (11–14), which protects them from HESO1-mediated 3′-end uridylation and subsequent degradation (15–17). Finally, the plant miRNA/miRNA*s are sorted and the majority of them loaded to effector protein ARGONAUTE 1 (AGO1), the catalytic component of a RNA-induced silencing complex (RISC), where a miRNA* passenger strand is selectively degraded (18,19). Since the asymmetric assembly of guide miRNA into RISC is abolished in a hyl1 mutant, the HYL1 protein presumably aids the correct strand selection from the duplex (10,20). However, it is not clear how HYL1 guides the miRNA/miRNA* duplex for the selective cleavage of its passenger strand by ARGONAUTE and what role, if any, in this process is played by the HEN1-mediated modification.
The HEN1 methyltransferase catalyzes the transfer of a methyl group from the cofactor S-adenosyl-L-methionine (AdoMet) onto the 2′-hydroxyl of the 3′-terminal nucleotide of small RNAs such as miRNA/miRNA* and siRNA/siRNA* (21,22). The X-ray crystallographic study of the ternary complex with double-stranded miRNA/miRNA* and cofactor product S-adenosyl-L-homocysteine (AdoHcy) revealed five structural domains of Arabidopsis HEN1 (23). The N-terminal part of 2′-O-methyltransferase harboring two double-stranded RNA-binding domains (dsRBDs) R1 and R2, as well as a La-motif-containing domain, L, is thought to be responsible for binding and sizing of the target RNA strand (Figure 1A). These domains cooperate to bind the central part and the non-target end of dsRNA and assist in proper positioning of the bound duplex in the C-terminal catalytic domain. However, the roles of the second RNA-binding motif R2 and the central region of HEN1, homologous to FK506-binding proteins, still remain obscure (23).
Figure 1.
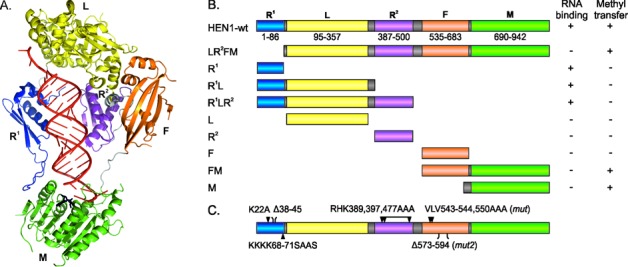
Functional analysis of HEN1 domains. (A) Representation of domains on the structure of HEN1 in complex with the miR173/miR173* duplex (red) and AdoHcy (black) (23). (B) Schematic view of the HEN1 deletion variants. RNA binding was assessed in vitro by EMSA using the miR173/miR173* substrate; methyl transfer activity was studied by monitoring incorporation of [3H]-methyl groups from labeled AdoMet into miR173/miR173*. R1 and R2 (blue and purple, correspondingly): double-stranded RNA-binding domains; L (yellow): La-motif-containing domain; F (orange): FK506-binding protein-like domain; M (green): methyltransferase domain. (C) Positions of HEN1 mutations.
Here, we performed a series of mapping experiments to further dissect the roles of the individual domains of the HEN1 methyltransferase. Functional characterization of the dsRBDs provided evidence for their significant, but unequal contribution to HEN1 interactions with the RNA substrate, and revealed a principal role of the FK506-binding protein-like domain of previously unknown function in binding the HYL1 protein. We also examined the interactions of HEN1 with three proteins that form a plant miRNA processing complex predicted by Manavella et al. (10). We found that HEN1 physically interacts with the double-stranded RNA-binding protein HYL1 (DRB1) and DCL1 protein, but shows no interaction with either SERRATE or SE-core protein. Based on these findings, we propose a new mechanism of plant miRNA maturation which envisions binding of the HEN1 methyltransferase to the DCL1•HYL1•miRNA complex excluding the SERRATE protein.
MATERIALS AND METHODS
Construction of plasmids and purification of recombinant proteins
Detailed procedures regarding construction of plasmids, expression and purification of recombinant proteins used in this study as well as HEN1 and its derivatives activity assays are described in Supplementary materials and methods.
Electrophoretic mobility shift assay
The equilibrium dissociation (Kd) and dissociation rate (koff) constants of the binary (protein•dsRNA) and ternary (protein•dsRNA•AdoHcy) complexes of HEN1 or its variants were determined as previously described (22). The electrophoretic mobility shift assay (EMSA) experiments with HYL1 and its mutants were performed with minor modifications. In 20 μl reaction volumes containing 10 mM Tris–HCl (pH 7.4), 50 mM NaCl, 0.1 mg/ml bovine serum albumin (BSA), 5% (v/v) glycerol and 0.05 u/μl RiboLock (Thermo Scientific), 45 nM 32P-labeled siR173/siR173* (siR173 5′-UUAACGCUUGCAGAGAGAAUCAC-3′, siR173* 5′-GAUUCUCUCUGCAAGCGUUAAAG-3′) or miR173/miR173* (miR173 5′-UUCGCUUGCAGAGAGAAAUCAC-3′, miR173* 5′-GAUUCUCUGUGUAAGCGAAAG-3′) were incubated with 0.01–1 μM purified recombinant proteins for 20 min at 25°C. If the dissociation rate of the complex was analyzed, 45 nM 32P-labeled siR173/siR173* was incubated with 250 nM HYL1 for 20 min at 25°C, then diluted with 2.5 μM of identical unlabeled RNA and left to equilibrate for 0–45 min. In EMSA experiments, where protein•RNA complexes were titrated with increasing concentration of the second protein, initial complexes were formed by mixing 45 nM 32P-labeled siR173/siR173* with 94 nM HYL1 or 1 μM HEN1 for 10 min at 25°C. After addition of 0.03–1 μM HEN1 or 0.05–0.9 μM HYL1 incubation continued for 20 min. The products of binding and titration reactions were electrophoresed on a 6% native polyacrylamide gel (19:1 acrylamide/bisacrylamide) in 0.5× Tris-borate buffer and autoradiographed. Fractions of bound and free RNA were evaluated using Multi Gauge v.3.0 software (Fujifilm) and the Kd values were obtained using the GraFit software version 5 (Erithacus Software) by fitting single site saturation curve to experimental data.
RNA methylation assay using the periodate oxidation method
Methylation activity of HEN1 and its mutants with unlabeled AdoMet were studied by the periodate oxidation method (24). Methyltransferases reactions were carried out with a 1 μM enzyme, 100 μM AdoMet and 0.1 μM miR173/miR173*, of which one strand was 5′-phosphorylated using [γ-33P]ATP (PerkinElmer). The reaction products were treated with sodium periodate, pre-heated with an equal volume of denaturing 2× RNA Loading Dye (Thermo Scientific), resolved on 15% polyacrylamide gel electrophoresis (PAGE) with 7 M urea and analyzed using FLA-5100 Image Readerand MultiGauge v.3.0 software package (Fujifilm).
Pre-steady-state kinetics analysis of HEN1 and its variants
The methyl group transfer under [E]>[S] conditions was studied using 33P-labeled miR173/miR173*. Reaction component (A), 0.2 μM RNA and 0.5 μM HEN1 or its mutant in Reaction buffer [10 mM Tris–HCl (pH 7.5), 50 mM NaCl, 10 mM KCl and 0.1 mg/ml BSA] was mixed by a Rapid Chemical Quench-Flow appliance RQF-3 (KinTek) with component (B), containing 200 μM AdoMet in the same buffer. The final concentrations of RNA, protein and AdoMet after combining equal volumes (15 μl of each) of components A and B were 0.1, 0.25 and 100 μM, respectively. Proteinase K in Stop buffer [7 mM Tris–HCl (pH 7.4), 0.17 mM ethylenediaminetetraacetic acid, 3 mM NaCl and 0.5% sodium dodecyl sulfate (SDS)] was added to a final concentration of 0.4 mg/ml to stop the reaction after certain period of incubation at 37°C. Sodium periodate treated samples were processed and analyzed as described above (RNA methylation assay using periodate oxidation method). Kinetic parameters of guide and passenger strands of RNA duplex were obtained by fitting experimental data to a single- or two-exponential equation using the GraFit5 software (Erithacus Software).
Detection of protein–protein interactions via the yeast two-hybrid system
Two-hybrid assays were carried out using the DupLEX-A system (OriGene Technologies, Rockville, MD, USA). A yeast strain EGY48 holding a LexA operon–LEU2 reporter was transformed with the reporter plasmid pSH18–34 containing the GAL1-lacZ gene under the control of eight LexA operators. EGY48 carrying pSH18–34 was co-transformed with the appropriate combination of ‘bait’ and ‘prey’ plasmids, respectively, pEG202-NLS (binding domain, BD) and pJG4–5 (activation domain, AD) fusion plasmids. Only bait plasmids, which passed recommended control tests (the auto-activation tests and the repression assay), were used. The transformation reactions were spread on YNB (glu)-his-ura-trp plates, which were incubated at 30°C 2–3 days until colonies appeared. From each interaction four independent colonies were streaked onto fresh YNB (glu)-his-ura-trp plates and incubated at 30°C for 2 days. Cells of each streak were suspended in 100 μl of sterile water and 2.5 μl of each cell suspension was spotted on 5-bromo-4-chloro-3-indolyl-β-d-galactopyranoside (X-Gal) containing YNB (glu)-his-ura-trp, YNB (glu)-his-ura-trp-leu, YNB (gal)-his-ura-trp and YNB (gal)-his-ura-trp-leu plates to screen for expression of LEU2 and lacZ reporters. The plates were then incubated at 30°C for 3–4 days.
GST pull-down experiments
One micromolar of glutathione S-transferase (GST) or GST-fused proteins were incubated with 2 μM of His-tagged proteins in the Binding buffer containing 10 mM Tris–HCl (pH 7.4), 50 mM NaCl, 5% (v/v) glycerol, 0.1% (v/v) Triton X-100 and 0.1 mM dithiothreitol (DTT) for 20 min at 22°C. The preformed protein complexes were added to 15 μl of Glutathione Sepharose 4B resin (GE Healthcare) prepared according to the manufacturer's recommendations and incubated for 2 h at room temperature with constant agitation. Reaction mixtures were washed three times with Binding buffer. Glutathione Sepharose 4B bound proteins were eluted by heating for 5 min at 95°C in reducing SDS-PAGE sample buffer, resolved on SDS-PAGE and transferred onto Hybond ECL nitrocellulose membrane (GE Healthcare) with Protein transfer buffer [5 mM Tris–HCl (pH 8.3), 192 mM glycine and 15% (v/v) methanol] at 100 V for 2 h in the cold room. Membranes were blocked with 3% BSA in PBS-T [80 mM Na2HPO4, 20 mM NaH2PO4 (pH 7.4), 0.1 M NaCl and 0.1% (v/v) Tween-20] for 1 h at room temperature and washed two times with PBS-T. Membranes were incubated with primary anti-GST (GE Healthcare) (1:4000) or tetra-His (Qiagen) (1:4000) antibodies overnight at 4°C, washed three times with PBS-T and incubated with horseradish peroxidase-conjugated secondary antibodies for 2 h at room temperature. After three washes with PBS-T and two with phosphate buffered saline (PBS), proteins signals were detected using 3,3′,5,5′-tetramethylbenzidine liquid substrate system (Sigma-Aldrich) or ECL western blotting detection reagents (GE Healthcare).
RESULTS
Functional differences of HEN1 double-stranded RNA-binding domains
Protein crystallography and bioinformatics studies identified five structural domains (R1, L, R2, F and M in Figure 1A) in the small RNA methyltransferase HEN1 (23,25). To determine the exact roles of the individual domains in catalysis and interactions with RNA substrates, we constructed a series of truncated HEN1 variants (Figure 1B). We first looked at their involvement in RNA binding and catalysis by using EMSAs and methylation activity/kinetic experiments. We found that HEN1 variants with N-terminal deletions showed no detectable RNA binding (Figure 2) whereas the wild-type protein assembled into a complex with miR173/miR173* RNA. Thus, similarly to the previously characterized catalytic domain HEN1-M (13,22), HEN1-LR2FM does not form a stable complex with the substrate (Figure 2) although is able to completely modify both RNA strands in vitro (Supplementary Figure S1A). On the other hand, the short N-terminal constructs encompassing the R1 domain (R1 or R1L) led to the formation of highly immobile protein•RNA complexes (retained in gel wells) (Figure 2); bound RNA was released after proteolysis with Proteinase K (data not shown) suggesting that the truncated proteins aggregated with the bound substrate during electrophoresis.
Figure 2.
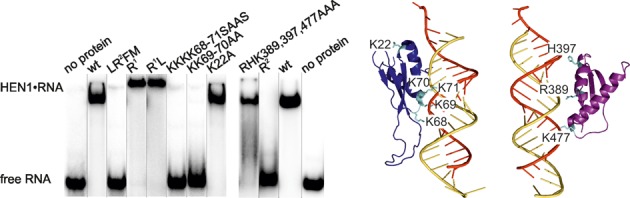
Distinct contributions of the double-stranded RNA-binding domains R1 and R2 to miRNA/miRNA* binding. Left, the RNA-binding capacity of the truncated and mutation variants of HEN1 was assessed by EMSA using 0.25 μM protein and 0.05 μM miR173/miR173* duplex. Right, amino acid residues selected for mutagenesis in the dsRNA-binding domains R1 (blue) and R2 (purple).
To further delineate the binding capacity of the R1 domain toward miRNA, we performed a structure-guided mutational analysis of the N-terminal domain. We selected lysine residues K68, K69, K70 and K71 which, according to the X-ray structure, are potentially involved in substrate interaction (23) and are conserved in plant HEN1 homologues. Two mutant variants, KK69–70AA and KKKK68–71SAAS, were produced and examined. Both mutants proved to be deficient in RNA binding (Figure 2), confirming that the key interactions with dsRNA occur at the N-terminal double-stranded RNA-binding domain. Yet, in the context of a large number of inferred contacts from different HEN1 domains to the RNA (23) such a dramatic impact of only two residues was quite unexpected. In contrast, a control mutant (K22A), in which lysine-22 was replaced with alanine, exhibited normal affinity toward miR173/miR173*.
Similarly to the wild-type enzyme, HEN1-LR2FM and the full-length proteins with mutations in the R1 domain, KK69–70AA and KKKK68–71SAAS, were able to methylate both strands of the miR173/miR173* duplex (Supplementary Figure S1A). Moreover, multiple-turnover rate kobs of the HEN1-LR2FM protein under steady-state conditions was commensurable and accounted for approximately half that of the WT enzyme, 1.1 versus 2.7 per min correspondingly. Quantitative comparison of the methyl transfer activities toward individual strands of miR173/miR173* duplex was done under single-turnover ([E]>[S]) conditions using 100 μM concentration of S-adenosyl-L-methionine. The HEN1-LR2FM-driven rates of methyl transfer to individual strands of miR173/miR173* decreased 2.5–35-fold, kchemmiR173 = 1.8 ± 0.1 versus 4.5 ± 0.2 and kchemmiR173* = 1.0 ± 0.1 versus 35 ± 3, compared with full-length protein under the same conditions (this paper and (22)). Our data show that the removal of the N-terminal dsRBD1 domain leads to a slight and asymmetric reduction in the methylation activity of HEN1 (Supplementary Figure S1C). Altogether our data imply that the N-terminal dsRBD R1 is essential for substrate binding but is not critical for catalysis.
A peculiarity of the Arabidopsis HEN1 R1 and R2 domains as compared to typical dsRBDs is a 15–40 residue insertion between β-strands β1 and β2 (Supplementary Figure S2A) (23). Similar insertions are found in other plant small RNA methyltransferases suggesting their significance for protein function. The insertion loop in the R2 domain is disordered in crystal structure (23) precluding a sensible prediction of its possible functional role. In contrast, the shorter loop in domain R1 appears to be responsible for making direct contacts to the C-terminal methyltransferase domain M and thus possibly participates in communication between the R1 and M subunits which have to act in concert to achieve RNA modification. To examine the importance of this insertion for HEN1 function, we generated a deletion mutant variant lacking residues 38–45 which are located at the tip of the loop (Supplementary Figure S2B). Detailed kinetic studies of the mutant protein HEN1Δ38–45 revealed that the deletion does not affect affinity toward RNA or methylation activity: Kd values of binary and ternary complexes or the methyl group transfer rate kchem are indistinguishable from the wild-type protein (Table 1 and Supplementary Table S1). However, unlike the wild-type methyltransferase, which formed well-defined complexes with RNA in gels, HEN1Δ38–45 formed smeared bands, especially with the fully methylated substrate, presumably due to partial release of the bound RNA during analysis (Supplementary Figure S3A). A detailed analysis of the dissociation kinetics of the binary HEN1•RNA complexes with unmethylated miR173/miR173* confirmed a destabilizing effect of the deletion (Supplementary Figure S3B, Table 2). A more substantial loss in the decay rate was seen using the fully methylated substrate, which represents a final reaction product (Supplementary Figure S3C). The binary complex completely disintegrated before the first experimental time point of 10 min, and even AdoHcy supplement barely increased the overall durability of HEN1Δ38–45•miR173CH3/miR173*CH3•AdoHcy complex (Table 2). Altogether, our experiments revealed that the insertion in the R1 domain plays no important role in RNA recognition or methyl group transfer, but it is essential for stabilization of the ternary complex after the methylation reaction.
Table 1. Interaction of HEN1 and its mutants with RNA duplexes.
| RNA duplex | Binary HEN1•RNA Kd value (M×10−10) | Ternary HEN1•RNA•AdoHcy Kd value (M×10−10) | ||||
|---|---|---|---|---|---|---|
| wt | Δ38–45 | RHK389,397,477AAA | wt | Δ38–45 | RHK389,397,477AAA | |
| Unmethylated miR173/miR173* | 4.3 ± 0.5 | 7.9 ± 0.5 | 3020 ± 290 | 3.2 ± 0.3 | 3.2 ± 0.4 | 650 ± 120 |
| Fully methylated miR173CH3/miR173*CH3 | 72 ± 1 | 21 ± 1 | Complex was not identified | 18 ± 2 | 10 ± 1 | Complex was not identified |
| Unmethylated siR173/siR173* | 5.0 ± 0.5 | 2930 ± 270 | 2.1 ± 0.4 | 680 ± 170 | ||
Reaction mixtures of ternary complexes contained 100 μM AdoHcy. wt denotes full-length HEN1 and Δ38–45, RHK389,397,477AAA denote mutant methyltransferases HEN1Δ38–45, HEN1-RHK389,397,477AAA, respectively. Binding constants were obtained by fitting the data of 3–9 experiments with a single-exponential equation.
Table 2. Dissociation kinetics of complexes of RNA duplexes with HEN1 variants.
| RNA duplex | Binary HEN1•RNA koff value (per min × 10−3 amplitude) | Ternary HEN1•RNA•AdoHcy koff value (per min × 10−3) | ||||
|---|---|---|---|---|---|---|
| wt | Δ38–45 | RHK389,397,477AAA | wt | Δ38–45 | RHK389,397,477AAA | |
| Unmethylated miR173/miR173* | 4.8 ± 0.2 (67%) | 5.2 ± 0.4 (44%) | 1.8 ± 0.2 (44%) | 2.0 ± 0.1 | 4.1 ± 0.1 (84%) | 2.1 ± 0.2 (76%) |
| 87 ± 14 (33%) | >> 70 (56%)a | >> 70 (56%)a | >> 70 (16%)a | >> 70 (24%)a | ||
| Fully methylated miR173CH3/miR173*CH3 | 11.6 ± 0.4 (26%) | >> 70a | 4.3 ± 0.2 | 14 ± 2 (11%) | ||
| >> 70 (74%)a | >> 70 (89%)a | |||||
| Unmethylated siR173/siR173* | 3.1 ± 0.2 (75%) | 7.1 ± 0.3 (53%) | 1.3 ± 0.1 | 2.1 ± 0.1 (83%) | ||
| 45 ± 15 (25%) | >> 70 (47%)a | >> 70 (17%)a | ||||
aEntire or major part of complex decays during 10 min.
Reaction mixtures of ternary complexes contained 100 μM AdoHcy. Abbreviations for proteins are given in Table 1. Binary and ternary complex decay data sets were fitted using a single- or two-exponential decay models. The amplitude of the dissociation rate is expressed as a percentage of the total amplitude. Results are means of 3–10 experiments ± S.D.
To understand the importance of the second dsRBD for substrate binding, we examined the ability of isolated R2 domain to associate with miR173/miR173* by EMSA. In contrast to domain R1, the sole R2 formed no stable complexes with RNA (Figure 2). To further seek possible roles for R2, three positively charged amino acids R389, H397 and K477, which could potentially form hydrogen bonds/salt bridges with the bound RNA, were changed to Ala. The triple alanine mutant was indistinguishable from the wild-type HEN1 in catalytic activity assays (Supplementary Figure S1B) as well as in kinetic parameters (Supplementary Table S1). Although HEN1-RHK389,397,477AAA was capable of RNA binding as assayed by EMSA (Figure 2), detailed analysis revealed that the mutations had a considerable effect on the stability of the binary HEN1•RNA and ternary HEN1•RNA•AdoHcy complexes (Figure 3D and E, Table 2). No significant difference in HEN1-RHK389,397,477AAA binding to completely complementary siR173/siR173* or partially unpaired miR173/miR173* was observed (Figure 3B and C, Table 1). In contrast, the triple mutations abolished the HEN1 ability to associate with the fully methylated substrate miR173CH3/miR173*CH3 (Figure 3A). Since only the M domain interacts directly with the target nucleotide, this result implicates the R2 domain in governing substrate association with the M domain.
Figure 3.
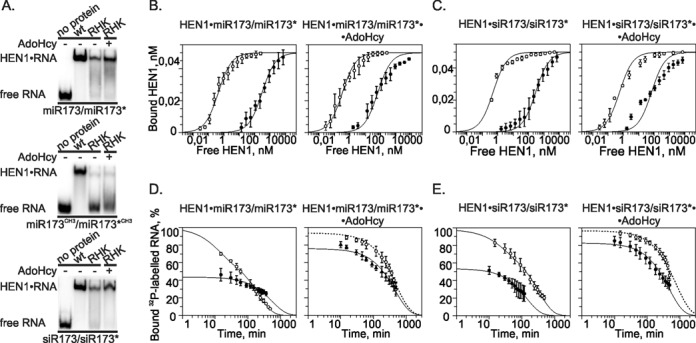
Interaction of the HEN1-RHK389,397,477AAA mutant with miR173/miR173* and siR173/siR173* substrates. (A) Native PAA gel EMSAs of 0.25 μM HEN1 or 7 μM HEN1-RHK389,397,477AAA binding to 0.05 μM unmethylated miR173/miR173*, fully methylated miR173CH3/miR173*CH3 and siR173/siR173* RNA duplexes in the absence (−) and presence (+) of 100 μM AdoHcy. wt: wild-type HEN1; RHK: mutant HEN1-RHK389,397,477AAA. (B), (C) Comparison of proteins’ affinity toward different RNA substrates. HEN1 (○) and mutant HEN1-RHK389,397,477AAA (•) mixed with 50 pM miR173/miR173* (B) or siR173/siR173* (C) RNA was incubated without (the left diagram) or with (the right diagram) 100 μM AdoHcy. Fits to a single-site saturation equation are shown as solid lines. (D), (E) Dissociation analysis of HEN1 (○) and its mutant HEN1-RHK389,397,477AAA (•) binary HEN1•RNA (the diagram at the left) and ternary HEN1•RNA•AdoHcy (the diagram at the right) complexes with miR173/miR173* (D) and siR173/siR173* (E) duplexes. Full-length HEN1 (0.25 μM) or mutant (7 μM) were incubated for 30 min with 0.15 nM 32P-labeled RNA duplexes in the absence of the cofactor or in the presence of 100 μM AdoHcy. A 13 000-fold excess of competitor unlabeled RNA substrate was added and aliquots were withdrawn at specified time points for immediate analysis by EMSA. Decay time courses of the binary and ternary complexes along with single-exponential fits (dotted lines) or two-exponential fits (solid lines) are shown.
Overall, the performed functional analysis revealed uneven roles of the R1 and R2 dsRBDs. R1 is a key factor for tight binding of small RNA duplexes, while R2 domain, in cooperation with M domain, further enhances the stability of the HEN1•miRNA/miRNA* complex.
HEN1 methyltransferase and double-stranded RNA-binding protein HYL1 interact physically
Available data suggest that domains R1, L, R2 and M collectively control key steps of the methyltransferase activity, namely, RNA binding, sizing of the target strand and methyl group transfer (13,23). However, the function of the F domain, the only HEN1 domain pointing away from the bound RNA, remained unknown. Therefore, the central part of HEN1 may be suspected to play a role in non-enzymatic processes such as interaction with other proteins involved in plant miRNA biogenesis. As the double-stranded RNA-binding protein HYL1 affects the strand selection in RISC, the process following miRNA/miRNA* methylation (10,20), we hypothesized that HYL1 might be a partner of HEN1. We first carried out EMSAs of HYL1 with miR173/miR173* and siR173/siR173* and found that HYL1 binds both types of double-stranded small RNAs with fairly similar affinity (Supplementary Figure S4). Consistent with previous findings by Yang et al. (26), initially HYL1 binds RNA as a monomer, but dimerizes as the protein concentration increases, resulting in a higher molecular mass complex. Upon titration of monomeric HYL1•siR173/siR173*complex (formed by using fixed 94 nM concentration of protein and 45 nM of RNA duplex) with increasing amounts of HEN1, 33P-labeled RNA bands were shifted up (Figure 4A). The band showed a lower mobility than the HYL1•RNA or HEN1•RNA structures, likely reflecting the formation of a ternary HEN1•RNA•HYL1 complex. As expected, reciprocal titration of HEN1•RNA with HYL1 also showed the appearance of a lower-mobility complex in a gel (Supplementary Figure S5).
Figure 4.
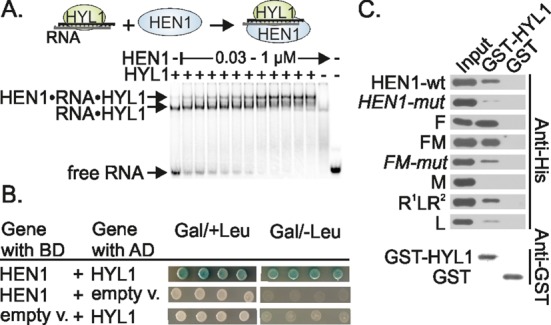
HEN1 interacts with HYL1 in vitro and in vivo. (A) Titration of RNA•HYL1 with HEN1 reveals a high molecular weight complex. Varying amount of HEN1 (from 0.03 to 1 μM) was added to the RNA•HYL1 complex, formed after preincubation of 94 nM of protein with 45 nM of siR173/siR173*. (+) and (−) indicates presence and absence of particular proteins. (B) Validation of the interaction between HEN1 and HYL1 using a two-hybrid assay. Full-length cDNA of HEN1 and HYL1 were fused to the DNA encoding the LexA DNA-binding domain (BD) and B42 transcriptional activation domain (AD), respectively. Four independent co-transformants in yeast strain EGY48 were screened for the lacZ and Leu reporter genes on plates containing selective X-Gal medium in the absence (Gal/–Leu) or with leucine (Gal/+Leu). (C) Analysis of protein–protein interaction among HYL1 and HEN1 domains by GST pull-down assay. The experiments were performed using 68 pmol of GST or GST-fused HYL1 (GST-HYL1) and 136 pmol of His-tagged HEN1 proteins. The pull-down fractions were analyzed by protein blotting with anti-His antibodies. Presence of GST-HYL1 and GST proteins in pull-down samples was confirmed by western blot with anti-GST antibodies. The input fractions represent 20% of the total amount of His-tagged proteins used in pull-down assays. Schemes of truncated HEN1 proteins are depicted in Figure 1. HEN1-mut and FM-mut: the full-length HEN1 and truncated protein composed of HEN1 F (FK506-binding protein-like) and M (methyltransferase) domains, respectively, which have three hydrophobic residues V543, L544 and V550 changed to alanine.
To reliably confirm the occurrence of specific interaction between the proteins in vivo, the yeast LexA two-hybrid system was exploited. The Arabidopsis HEN1 in-frame fused to the LexA DNA-binding domain (HEN1-BD) was used as prey to test for interaction with HYL1 fused with the LexA activation domain (HYL1-AD) in the strain EGY48[p8op-lacZ] containing LEU2 and lacZ reporters with upstream LexA operators. As shown in Figure 4B, the yeast cells co-expressing both chimeric genes were able to grow on medium without leucine supplement and developed a blue color in the presence of X-Gal, reflecting activation of the LEU2 and lacZ genes mediated by the bait–prey interactions. This in vivo result provides firm evidence that HEN1 is a binding partner of the HYL1. The specificity of the observed protein–protein interaction was controlled using self-activation tests with yeast cells carrying either HEN1-BD or HYL1-AD (Figure 4B).
Finally, to demonstrate a direct physical protein–protein interaction, a HYL1 fusion protein with GST was prepared and used as a bait for GST pull-down assays with immobilized reduced glutathione. The presence of captured His6-tagged HEN1 was observed using anti-His antibodies. Data presented in Figure 4C revealed efficient pull-down of the full-length HEN1 by the GST-HYL1 protein. At the same time the binding of HEN1 to GST alone was not observed excluding the possibility of non-specific HEN1–wt interactions with either GST or Glutathione Sepharose 4B. Thus the in vitro GST pull-down assays confirmed the in vivo results of the yeast two-hybrid analyses. Notably, addition of miR173/miR73* did not affect the strength of the HEN1–wt signal in a pull-down sample (data not shown) indicating that the presence of the small RNA duplex is not essential for the HEN1•HYL1 interaction.
To delineate individual domains involved in this interaction, the GST-HYL1 pull-down experiments were performed with the truncated variants of HEN1. The strongest histidine signal was observed with the deletion derivatives HEN1-F and HEN1-FM possessing the FK506-binding protein-like domain (Figure 4C). However, domain M alone did not co-purify with HYL1 disproving any role for the C-terminal part of HEN1 in HYL1 binding. On the other hand, the HEN1-R1LR2 derivative (residues 1–519) lacking the F and M domains showed a clear co-precipitation signal suggesting that the N-terminal part of HEN1 may also contribute to the binding. To further refine the N-terminal interacting regions of HEN1, we examined individual dsRBDs, R1 and R2, as well as the La-motif-containing domain, L. Since only minor amounts of HEN1-L were extracted by GST-HYL1, we conclude that the tight interaction is unlikely to result from the La-motif-containing domain. Unfortunately, the HEN1 variants representing sole R1 or R2 domains were highly prone to aggregation (were present in the glutathione sepharose pellet even in samples without GST, data not shown). To circumvent this limitation, the NusA protein, which is known to enhance solubility and stability of heterologous proteins in Escherichia coli, was fused to the N-terminus of the target domains. Although this approach was unsuccessful for R2 (data not shown), the non-specific signal in the controls was eliminated in the case of R1 (Supplementary Figure S6). However, the pull-down experiments showed no appearance of R1 in GST-HYL1 fractions. Taking into account that the contribution of R1 and L domains to the binding is negligible, it could be indirectly assumed that the major determinant of the observed HEN1-R1LR2•HYL1 interaction is R2. Altogether, our analysis suggests that HYL1 extensively recognizes the central part of HEN1, containing R2 and F domains.
To define regions of the F domain that mediate its interaction with HYL1, we performed a structure-guided mutational analysis. In general, sites of protein–protein interaction often correspond to disordered protein regions which become structured upon complex formation (27,28). Therefore, we considered two disordered loops of the F domain in the HEN1 crystal structure (Supplementary Figure S7) as potential candidates for the HYL1-binding role. The shorter loop (residues 540–553) links fairly distant structural regions and its removal or even shortening could thus substantially alter the overall structure of the F domain. In contrast, the longer loop (residues 570–601) connects adjacent β-strands and its shortening would be expected to be harmless for the structural integrity of the domain. Based on these considerations, we constructed mutants either by introducing residue substitutions within the shorter loop or by deleting the longer loop. The deletion of residues 573–594 of the long loop had no detectable effect on the protein–protein interaction (Supplementary Figure S7, lane mut2). In contrast, a triple substitution VLV543–544,550AAA in the short disordered region significantly reduced binding of HEN1-FM to HYL1 (Figure 4C, lane FM-mut). Moreover, introduction of the same mutations into the full-length protein significantly reduced the amount of the GST-HYL1-bound HEN1, supporting the principal role of the F domain in the HEN1•HYL1 complex formation (Figure 4C, lane HEN1-mut). Importantly, the methyl group transfer experiments with miR173/miR173* RNA confirmed that the loop mutants retained complete wild-type methylation capacity (data not shown) consistent with our observed dispensability of the F domain for catalytic activity (Figure 1B).
dsRNA-binding domain R2 of HYL1 directly binds F domain of HEN1
HYL1 consists of two dsRNA-binding domains, nuclear localization signal and a long C-terminal extension of unknown function (Figure 5A). To identify which part of the protein binds to HEN1, we produced three truncated HYL1 variants containing individual dsRBDs (R1 and R2) each in separate proteins or both combined in a single protein for reciprocal GST pull-down assays. As shown in Figure 5A, R1 did not interact with HEN1, although it was functionally active and capable of RNA binding in the EMSA analysis (Supplementary Figure S8). In contrast, R2 alone or the tandem R1R2 protein bound both the full-length HEN1 and HEN1-F proteins, indicating that HYL1 interacts with the FK506-binding protein-like domain of HEN1 via the R2 domain. It is known that this non-canonical dsRBD also participates in protein–protein interactions with other proteins involved in miRNA biogenesis (5,31). Interestingly, the triple alanine mutation VLV543–544,550AAA significantly reduced the affinity of HEN1-FM for GST-HYL1-R2 suggesting the importance of hydrophobic interactions mediated by these residues for R2 domain binding. HEN1-R1LR2 interacts with HYL1-R2 but not HYL1-R1 (data not shown), indicating that the HYL1•HEN1 interaction is largely mediated by R2 of HYL1. Direct contacts of the HEN1-F domain with the second double-stranded RNA domain of HYL1 was further confirmed by yeast two-hybrid analysis (Figure 5B). It should be noted that we reversed the components of two-hybrid systems as compared with the above described experiments involving full-length proteins: HYL1-R2 and HEN1-F were fused to LexA binding and activation domain, respectively. Neither HEN1-F nor HYL1-R2 alone did auto-activate the reporter genes. In conclusion, both GST pull-down and yeast two-hybrid assays proved that the F domain of HEN1 and the R2 domain of HYL1 are both necessary and sufficient to mediate interaction between the two proteins.
Figure 5.
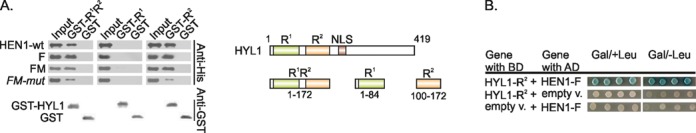
HYL1 interacts with HEN1 via its second dsRNA-binding domain R2. (A) GST pull-down assays indicating HEN1 binds to HYL1-R1R2 and R2 but not to HYL1-R1. On the right schematic representation of full-length HYL1 and shorter proteins of separate domains is given. R1 and R2: double-stranded RNA-binding domains, NLS: nuclear localization signal. (B) Two-hybrid interaction between FK506-binding protein-like domain HEN1-F and the second dsRNA-binding domain HYL1-R2. All experiments were performed as described in Figure 4.
HEN1 binds the microprocessor complex component DCL1 but not SERRATE
To gain further insights into possible in vivo roles of HEN1, we tested if HEN1 directly interacts with other plant microprocessor complex proteins, namely, SERRATE and DCL1. The full-length and truncated SE-core (amino acids 194–543) variants of SERRATE were fused to the LexA binding and activation domain, respectively, and examined using yeast two-hybrid system. We found no detectable HEN1-dependent reporter signal with any of the analyzed proteins (Figure 6A). Furthermore, no co-purification of the full-length HEN1 was detected in pull-down experiments containing GST-SE-core (Figure 6B). Previous in vitro pull-down studies have identified HYL1 as a binding partner of the SE-core consisting of the two N-terminal alpha-helices, the middle α-helix dominant and the C-terminal non-canonical C2H2 zinc-finger domains (6). As expected, this shorter SERRATE isoform also showed interaction with the full-length HYL1 in our two-hybrid assay thereby verifying its functionality in yeast cells (Figure 6A). Altogether these results convincingly demonstrate that SERRATE does not directly interact with the HEN1 methyltransferase and at the same time confirm the previously reported interaction between SE-core and HYL1.
Figure 6.
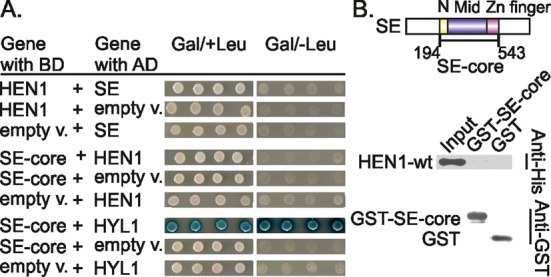
Lack of detectable interactions between SERRATE and HEN1 methyltransferase. (A) Yeast two-hybrid analysis revealed no in-cell interactions between HEN1 and SERRATE (SE). HYL1, a known SE-core-interacting partner (6), was used as a positive control for yeast assay. (B) GST pull-down experiment shows no interaction between central part of SERRATE (SE-core) and full-length HEN1 proteins in vitro. Experiments were carried out as depicted in Figure 4. Schematic representation of the domains in SE is made according to Machida et al. (6). SE-core construct is comprised of N-terminal (N), middle (Mid) and Zinc finger domains.
The ribonuclease III-type enzyme DCL1, the principal player of miRNA biogenesis in plants, promotes different steps of miRNA maturation. DCL1 Helicase and PAZ domains interact with other proteins involved in miRNA biogenesis—SERRATE (6), DDL (29) and NOT2 (30), while its DUF283 and R1R2 domains specifically bind HYL1 (31,32). We thus prepared the three domains, Helicase, DUF283 and R1R2, as separate truncation proteins and examined their interactions with HEN1 using the yeast two-hybrid system. Data presented in Figure 7A show that at least two of them, the Helicase and the tandem of dsRBDs, R1R2, contribute to methyltransferase binding. Remarkably, we detected no HEN1 binding in the case of the DCL1-DUF283 domain, which was shown to be essential for HYL1 binding (31). To further confirm the observed interaction pattern we performed similar pull-down assays as shown in Figure 7B. We found that HEN1 co-purified with GST-DCL1-Helicase, GST-DCL1-PAZ and GST-DCL1-R1R2, although the DCL1-R1R2 fragment pulled down more methyltransferase than either DCL1-Helicase or DCL1-PAZ domains. HEN1 was not detected upon incubation with GST-DCL1-DUF283 or GST proteins, although the GST signal was detected in samples. Due to previously reported poor solubility of protein fragment containing the DCL1 ribonuclease domain (6), this domain has not been selected for analysis. In contrast to HYL1, the HEN1 mutation VLV543–544,550AAA did not reduce binding of DCL1 to HEN1 suggesting that these amino acids are not essential for direct contacts between DCL1 domains and HEN1.
Figure 7.
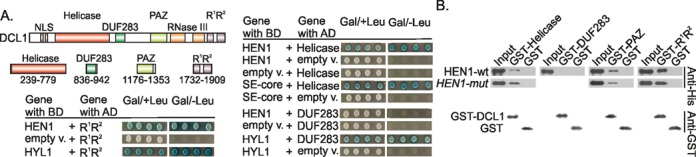
Interaction of HEN1 with individual domains of DICER-LIKE 1 (DCL1). (A) Yeast two-hybrid analysis of the interaction between HEN1 and DCL1 domains. Abbreviations of domains are as follows: NLS: nuclear localization signal, Helicase: DExD/H-box RNA helicase, DUF283: domain of unknown function 283, PAZ: Piwi/Argonaute/Zwille domain, RNase III: ribonuclease III, R1, R2: dsRNA-binding domains. Interactions between SE-core with Helicase and HYL1 with DUF283 or R1R2 domains of DCL1 served as positive controls (5,6,31). (B) Detection of the interaction between DCL1 domains and HEN1 using GST pull-down. GST-Helicase, GST-DUF283, GST-PAZ and GST-R1R2 denote GST-DCL1-Helicase, GST-DCL1-DUF283, GST-DCL1-PAZ and GST-DCL1-R1R2, respectively. Experiments were carried out as described in Figure 4.
DISCUSSION
The molecular basis of double-stranded microRNA recognition by HEN1
The elongated N-terminal part upstream of the catalytic domain, which folds into four domains, distinguishes the plant small RNA 2′-O-methyltransferase HEN1 from bacterial, protozoan or animal homologues (23,33). In this study, we found that the dsRBDs, R1 and R2 of HEN1, considerably but unequally contribute to the binding of mature miRNA/miRNA* duplexes to the protein. The site-specific mutagenesis of lysine residues K69 and K70 in the R1 domain completely abolished formation of detectable HEN1•RNA complex. By contrast, the R2 domain is not so strictly required for RNA recognition since changes of three amino acids (RHK389,397,477AAA) important for substrate recognition lead to 200–600-fold decreased (but detectable in PAA gels) association with the substrate (Table 1). It is noteworthy that the binding affinities of the RHK389,397,477AAA mutant were similarly reduced with respect to miRNA/miRNA* and siRNA/siRNA*, both in binary and in ternary complexes. This result is in conflict with the previous hypothesis that the R2 domain favors binding of miRNA/miRNA*-type small RNAs duplexes containing mispairs and bulges (23). Rather, the R2 domain is important for the stabilization of methyltransferase complexes with RNA since the mutant releases unmethylated substrate faster than the wild-type protein (Table 2). We presume that R2 together with the catalytic domain M stabilizes the HEN1•RNA complex and preserves its integrity after the methylation reaction, such that the methylated RNA remains in assembly with the other proteins, HYL1 and DCL1 (see below).
Interplay of HEN1 with other microRNAs biogenesis proteins
Plant endogenous small RNAs, such as miRNAs, ta-siRNAs, nat-siRNA, hc-siRNA, ra-siRNA, lsiRNA and exogenous small RNAs are 2′-O-methylated at their 3′-ends (34). Hence, HEN1 is an essential constituent in biogenesis of all the classes of small RNAs. However, the interactions of the methyltransferase with other proteins participating in small RNAs biogenesis remained obscure even for the best studied miRNA maturation pathway. The microprocessor complex comprising DCL1, HYL1 and SERRATE processes pri-miRNAs to mature miRNA/miRNA* duplexes in nuclear dicing bodies (D-bodies) (10,32). We found that HEN1, the protein modifying miRNA/miRNA*, interacts with the microprocessor constituents, DCL1 and HYL1, but does not bind the SERRATE protein. Remarkably, the functional evaluation of domain–domain interactions revealed that both HEN1 and SERRATE physically bind to identical regions of DCL1 (Helicase, PAZ and R1R2) and HYL1 (R2) (Figure 8A). This finding implies that HEN1 can assemble the particular DCL1•HYL1 complex only when SERRATE is released from the microprocessor complex and disengages its interaction sites on DCL1 and HYL1. Based on collective results, we propose a model in which, following excision of miRNA/miRNA* from primary RNA by the microprocessor, the HEN1 methyltransferase takes over the place of the SERRATE protein, which leaves the complex (Figure 8B). Then the RNA duplex is methylated and presumably the new complex selectively loads the guide strand of the mature miRNA/miRNA* into the RISC complex. This model is supported by the observations obtained by other groups: (i) there is no evidence of SE activity in processes following mature miRNA/miRNA* formation. In contrast, the precise selection of the guide strand in RISC, the process following miRNA/miRNA* methylation, is impaired in the absence of HYL1 (10,20); (ii) the studies in animal systems have shown that the association of the Dicer ribonuclease with the specific dsRNA-binding proteins, TRBP in human or R2D2 in flies, is required for the recruitment of Argonaute to the small RNAs; this suggests a role of Dicer•dsRNA-binding protein heterodimer not only in small RNA processing but also as a platform for RISC assembly (35–37). By analogy, the small RNA duplexes may be assisted by supporting proteins to be effectively recruited to Argonaute in plants. Direct interaction between cellular HYL1 and AGO1 displayed by bimolecular fluorescence complementation in living plants is consistent with this assumption (7). Whereas HYL1 forms short-lived complexes with RNA at least in vitro (Supplementary Figure S4C), it is very likely that DCL1 and HEN1 either jointly or individually aid the retention of the miRNA/miRNA* duplex in the complex.
Figure 8.
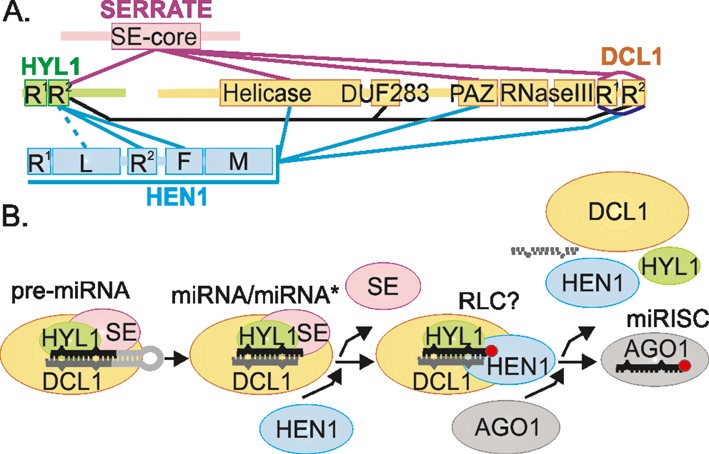
HEN1 interaction network and the proposed model of late stages of miRNA biogenesis. (A) Protein–protein interaction network involving SE, HYL1, DCL1 and HEN1. Cyan lines show interactions experimentally determined in this work, purple and black lines depict those reported previously (6,31,32). (B) Proposed model of miRNA biogenesis envisions that after the miRNA/miRNA* duplex is cut out of its precursor, SE is expelled and HEN1 methyltransferase is bound in the microprocessor complex to form a HYL1•HEN1•DCL1 complex, which might represent the still unidentified plant RISC-loading complex (RLC) anticipated by Eamens et al. (20). We hypothesize that this complex could direct HEN1 methylation (red circle) to the target miRNA strand (black) thus labeling it for incorporation into AGO1 complex.
To date it is not clear if the small RNA methyltransferase modifies both strands or only one particular strand of the mature miRNA/miRNA* duplex in a plant cell. Although HEN1 can methylate each strand in a non-processive manner in vitro (22), we suppose that in vivo the HYL1•DCL1 complex sterically orientates HEN1 and thereby predetermines which strand will be modified. The 3′-end of the passenger strand most probably is shielded from methylation and consequently remains intact. Normally only methylated microRNAs are involved in the specific Arabidopsis RISC complexes (11), whereas unmethylated strands undergo uridylation and degradation by miRNA nucleotidyl transferase HESO1 (16,38). We speculate that HEN1 marks the correct guide strand of miRNA/miRNA* duplex by methylation whereas HESO1 performs ‘quality control’ of the small RNA in RISC to ensure that it carries an attached methyl group. Alternatively, the methylation pattern could determine which strand will be removed during RISC maturation, however, further studies will have to address this intriguing possibility.
SUPPLEMENTARY DATA
Supplementary Data are available at NAR Online.
Acknowledgments
The authors would like to thank Satoru Machida and Adam Yuan for providing vectors carrying DCL1, SERRATE or HYL1 coding sequences and Alma Gedvilaitė, Rasa Rakauskaitė for help with two-hybrid analysis.
Footnotes
These authors contributed equally to the paper as first authors.
FUNDING
The Research Council of Lithuania [MIP-028/2012 to G.V.]. Funding for open access charge: The Research Council of Lithuania.
Conflict of interest statement. None declared.
REFERENCES
- 1.Bologna N.G., Schapire A.L., Palatnik J.F. Processing of plant microRNA precursors. Brief. Funct. Genomics. 2013;12:37–45. doi: 10.1093/bfgp/els050. [DOI] [PubMed] [Google Scholar]
- 2.Rogers K., Chen X. Biogenesis, turnover, and mode of action of plant microRNAs. Plant Cell. 2013;25:2383–2399. doi: 10.1105/tpc.113.113159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Kurihara Y., Takashi Y., Watanabe Y. The interaction between DCL1 and HYL1 is important for efficient and precise processing of pri-miRNA in plant microRNA biogenesis. RNA. 2006;12:206–212. doi: 10.1261/rna.2146906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Dong Z., Han M.-H., Fedoroff N. The RNA-binding proteins HYL1 and SE promote accurate in vitro processing of pri-miRNA by DCL1. Proc. Natl. Acad. Sci. U.S.A. 2008;105:9970–9975. doi: 10.1073/pnas.0803356105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Hiraguri A., Itoh R., Kondo N., Nomura Y., Aizawa D., Murai Y., Koiwa H., Seki M., Shinozaki K., Fukuhara T. Specific interactions between Dicer-like proteins and HYL1/DRB-family dsRNA-binding proteins in Arabidopsis thaliana. Plant Mol. Biol. 2005;57:173–188. doi: 10.1007/s11103-004-6853-5. [DOI] [PubMed] [Google Scholar]
- 6.Machida S., Chen H.-Y., Adam Yuan Y. Molecular insights into miRNA processing by Arabidopsis thaliana SERRATE. Nucleic Acids Res. 2011;39:7828–7836. doi: 10.1093/nar/gkr428. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Fang Y., Spector D.L. Identification of nuclear dicing bodies containing proteins for microRNA biogenesis in living Arabidopsis plants. Curr. Biol. 2007;17:818–823. doi: 10.1016/j.cub.2007.04.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Song L., Han M.-H., Lesicka J., Fedoroff N. Arabidopsis primary microRNA processing proteins HYL1 and DCL1 define a nuclear body distinct from the Cajal body. Proc. Natl. Acad. Sci. U.S.A. 2007;104:5437–5442. doi: 10.1073/pnas.0701061104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Fujioka Y., Utsumi M., Ohba Y., Watanabe Y. Location of a possible miRNA processing site in SmD3/SmB nuclear bodies in Arabidopsis. Plant Cell Physiol. 2007;48:1243–1253. doi: 10.1093/pcp/pcm099. [DOI] [PubMed] [Google Scholar]
- 10.Manavella P.A., Hagmann J., Ott F., Laubinger S., Franz M., Macek B., Weigel D. Fast-forward genetics identifies plant CPL phosphatases as regulators of miRNA processing factor HYL1. Cell. 2012;151:859–870. doi: 10.1016/j.cell.2012.09.039. [DOI] [PubMed] [Google Scholar]
- 11.Yu B., Yang Z., Li J., Minakhina S., Yang M., Padgett R.W., Steward R., Chen X. Methylation as a crucial step in plant microRNA biogenesis. Science. 2005;307:932–935. doi: 10.1126/science.1107130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Abe M., Yoshikawa T., Nosaka M., Sakakibara H., Sato Y., Nagato Y., Itoh J. WAVY LEAF1, an ortholog of Arabidopsis HEN1, regulates shoot development by maintaining MicroRNA and trans-acting small interfering RNA accumulation in rice. Plant Physiol. 2010;154:1335–1346. doi: 10.1104/pp.110.160234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Vilkaitis G., Plotnikova A., Klimasauskas S. Kinetic and functional analysis of the small RNA methyltransferase HEN1: the catalytic domain is essential for preferential modification of duplex RNA. RNA. 2010;16:1935–1942. doi: 10.1261/rna.2281410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Ji L., Chen X. Regulation of small RNA stability: methylation and beyond. Cell Res. 2012;22:624–636. doi: 10.1038/cr.2012.36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Li J., Yang Z., Yu B., Liu J., Chen X. Methylation protects miRNAs and siRNAs from a 3′-end uridylation activity in Arabidopsis. Curr. Biol. 2005;15:1501–1507. doi: 10.1016/j.cub.2005.07.029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Ren G., Chen X., Yu B. Uridylation of miRNAs by hen1 suppressor1 in Arabidopsis. Curr. Biol. 2012;22:695–700. doi: 10.1016/j.cub.2012.02.052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Zhao Y., Yu Y., Zhai J., Ramachandran V., Dinh T.T., Meyers B.C., Mo B., Chen X. The Arabidopsis nucleotidyl transferase HESO1 uridylates unmethylated small RNAs to trigger their degradation. Curr. Biol. 2012;22:689–694. doi: 10.1016/j.cub.2012.02.051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Baumberger N., Baulcombe D.C. Arabidopsis ARGONAUTE1 is an RNA Slicer that selectively recruits microRNAs and short interfering RNAs. Proc. Natl. Acad. Sci. U.S.A. 2005;102:11928–11933. doi: 10.1073/pnas.0505461102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Mi S., Cai T., Hu Y., Chen Y., Hodges E., Ni F., Wu L., Li S., Zhou H., Long C., et al. Sorting of small RNAs into Arabidopsis argonaute complexes is directed by the 5′ terminal nucleotide. Cell. 2008;133:116–127. doi: 10.1016/j.cell.2008.02.034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Eamens A.L., Smith N.A., Curtin S.J., Wang M.-B., Waterhouse P.M. The Arabidopsis thaliana double-stranded RNA binding protein DRB1 directs guide strand selection from microRNA duplexes. RNA. 2009;15:2219–2235. doi: 10.1261/rna.1646909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Yang Z., Ebright Y.W., Yu B., Chen X. HEN1 recognizes 21–24 nt small RNA duplexes and deposits a methyl group onto the 2′ OH of the 3′ terminal nucleotide. Nucleic Acids Res. 2006;34:667–675. doi: 10.1093/nar/gkj474. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Plotnikova A., Baranauskė S., Osipenko A., Klimašauskas S., Vilkaitis G. Mechanistic insights into small RNA recognition and modification by the HEN1 methyltransferase. Biochem. J. 2013;453:281–290. doi: 10.1042/BJ20121699. [DOI] [PubMed] [Google Scholar]
- 23.Huang Y., Ji L., Huang Q., Vassylyev D.G., Chen X., Ma J.-B. Structural insights into mechanisms of the small RNA methyltransferase HEN1. Nature. 2009;461:823–827. doi: 10.1038/nature08433. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Yang Z., Vilkaitis G., Yu B., Klimasauskas S., Chen X. Approaches for studying microRNA and small interfering RNA methylation in vitro and in vivo. Methods Enzymol. 2007;427:139–154. doi: 10.1016/S0076-6879(07)27008-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Tkaczuk K.L., Obarska A., Bujnicki J.M. Molecular phylogenetics and comparative modeling of HEN1, a methyltransferase involved in plant microRNA biogenesis. BMC Evol. Biol. 2006;6:6. doi: 10.1186/1471-2148-6-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Yang S.W., Chen H.-Y., Yang J., Machida S., Chua N.-H., Yuan Y.A. Structure of Arabidopsis HYPONASTIC LEAVES1 and its molecular implications for miRNA processing. Structure. 2010;18:594–605. doi: 10.1016/j.str.2010.02.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Fong J.H., Shoemaker B.A., Garbuzynskiy S.O., Lobanov M.Y., Galzitskaya O.V., Panchenko A.R. Intrinsic disorder in protein interactions: insights from a comprehensive structural analysis. PLoS Comput. Biol. 2009;5:e1000316. doi: 10.1371/journal.pcbi.1000316. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Wright P.E., Dyson H.J. Linking folding and binding. Curr. Opin. Struct. Biol. 2009;19:31–38. doi: 10.1016/j.sbi.2008.12.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Yu B., Bi L., Zheng B., Ji L., Chevalier D., Agarwal M., Ramachandran V., Li W., Lagrange T., Walker J.C., et al. The FHA domain proteins DAWDLE in Arabidopsis and SNIP1 in humans act in small RNA biogenesis. Proc. Natl. Acad. Sci. U.S.A. 2008;105:10073–10078. doi: 10.1073/pnas.0804218105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Wang L., Song X., Gu L., Li X., Cao S., Chu C., Cui X., Chen X., Cao X. NOT2 proteins promote polymerase II-dependent transcription and interact with multiple MicroRNA biogenesis factors in Arabidopsis. Plant Cell. 2013;25:715–727. doi: 10.1105/tpc.112.105882. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Qin H., Chen F., Huan X., Machida S., Song J., Yuan Y.A. Structure of the Arabidopsis thaliana DCL4 DUF283 domain reveals a noncanonical double-stranded RNA-binding fold for protein–protein interaction. RNA. 2010;16:474–481. doi: 10.1261/rna.1965310. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Liu Q., Yan Q., Liu Y., Hong F., Sun Z., Shi L., Huang Y., Fang Y. Complementation of HYPONASTIC LEAVES1 by double-strand RNA-binding domains of DICER-LIKE1 in nuclear dicing bodies. Plant Physiol. 2013;163:108–117. doi: 10.1104/pp.113.219071. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Huang R.H. Unique 2′-O-methylation by Hen1 in eukaryotic RNA interference and bacterial RNA repair. Biochemistry. 2012;51:4087–4095. doi: 10.1021/bi300497x. [DOI] [PubMed] [Google Scholar]
- 34.Katiyar-Agarwal S., Jin H. Role of small RNAs in host-microbe interactions. Annu. Rev. Phytopathol. 2010;48:225–246. doi: 10.1146/annurev-phyto-073009-114457. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Chendrimada T.P., Gregory R.I., Kumaraswamy E., Norman J., Cooch N., Nishikura K., Shiekhattar R. TRBP recruits the Dicer complex to Ago2 for microRNA processing and gene silencing. Nature. 2005;436:740–744. doi: 10.1038/nature03868. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Liu X., Jiang F., Kalidas S., Smith D., Liu Q. Dicer-2 and R2D2 coordinately bind siRNA to promote assembly of the siRISC complexes. RNA. 2006;12:1514–1520. doi: 10.1261/rna.101606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Okamura K., Robine N., Liu Y., Liu Q., Lai E.C. R2D2 organizes small regulatory RNA pathways in Drosophila. Mol. Cell. Biol. 2011;31:884–896. doi: 10.1128/MCB.01141-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Ren G., Xie M., Zhang S., Vinovskis C., Chen X., Yu B. Methylation protects microRNAs from an AGO1-associated activity that uridylates 5′ RNA fragments generated by AGO1 cleavage. Proc. Natl. Acad. Sci. U.S.A. 2014;111:6365–6370. doi: 10.1073/pnas.1405083111. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


