SUMMARY
The B cell response to Salmonella typhimurium (STm) occurs massively at extrafollicular sites, without notable germinal centers (GCs). Little is known in terms of its specificity. To expand the knowledge of antigen targets, we screened plasmablast (PB)-derived monoclonal antibodies (mAbs) for Salmonella specificity, using ELISA, flow cytometry, and antigen microarray. Only a small fraction (0.5%–2%) of the response appeared to be Salmonella-specific. Yet, infection of mice with limited B cell receptor (BCR) repertoires impaired the response, suggesting that BCR specificity was important. We showed, using laser microdissection, that somatic hypermutation (SHM) occurred efficiently at extrafollicular sites leading to affinity maturation that in turn led to detectable STm Ag-binding. These results suggest a revised vision of how clonal selection and affinity maturation operate in response to Salmonella. Clonal selection initially is promiscuous, activating cells with virtually undetectable affinity, yet SHM and selection occur during the extrafollicular response yielding higher affinity, detectable antibodies.
INTRODUCTION
The immune response to microorganisms is an interplay between aspects of innate and adaptive immunity. Successful pathogens often have multiple mechanisms to evade or subvert the immune response. Furthermore, pathogens contain molecular patterns that stimulate a wide variety of innate immune pattern-recognition receptors, whether expressed on innate cells such as macrophages, dendritic cells (DCs), or on adaptive cells such as B lymphocytes. These pathways and innate receptor ligands in turn shape adaptive immunity.
Currently, our knowledge of B cell responses is mostly based on artificial systems that lack these natural innate immune cues. Instead, they use non-replicating antigens given in adjuvant. The “canonical” response described in these models includes a rapid transient extrafollicular (EF) plasmablast (PB) response followed by germinal center (GC) appearance (Shlomchik and Weisel, 2012). While some pathogen responses follow this progression (e.g., the response to influenza [Coro et al., 2006; Moyron-Quiroz et al., 2004; Onodera et al., 2012; Rothaeusler and Baumgarth, 2010]), there is increasing evidence that in many other infections this is not the case. During Ehrlichia muris infection splenic GC formation is suppressed (Racine et al., 2010). Similarly, Borrelia burgdorferi interferes with the B cell response by affecting its quality and kinetics, delaying GC appearance and instead stimulating immunoglobulin M (IgM) antibody-forming cells (AFCs) (Hastey et al., 2012).
Though the immune response to Salmonella enterica serovar Typhimurium (STm), a facultative intracellular gram negative bacterium, is relatively well studied (Dougan et al., 2011), information on the B cell response is limited. This is a major omission, considering that STm is a clinically relevant microorganism and that live attenuated strains have been proposed and are in phase I clinical trials as vectors for vaccines (Kong et al., 2012). Furthermore, STm and related serovars are a major cause of infectious diarrhea in the developed world and they are also responsible for serious disseminated infections in Africa and Asia. It is highly homologous to Salmonella Typhi, and considered a murine model for the study of this pervasive human pathogen. The B cell response to STm can be protective in both mice and humans, via antibodies or other mechanisms (Nanton et al., 2012).
STm induces a massive extrafollicular AFC response in the spleen, while GC formation is greatly delayed (Cunningham et al., 2007). Both T-dependent (TD) and T-independent (TI) components contribute to the response (Gil-Cruz et al., 2009). The mechanisms that shape this type of B cell response remain to be elucidated, whereas parameters of virulence and protection have received greater attention. Deletion of the signaling adaptor MyD88 appeared to favor, rather than inhibit, STm virulence (Arpaia et al., 2011; Barr et al., 2010; Neves et al., 2010).
A number of studies have addressed the targets of the B cell response, yet overall these remain poorly defined. LPS, outer membrane proteins (OMPs) and possibly flagellin are identified as primary Ags of the switched Ab response (Bobat et al., 2011; Calderón et al., 1986; Cunningham et al., 2007; Ortiz et al., 1989; Singh et al., 1992). Recently, some of the authors of the present work have screened immune sera on antigen (Ag) microarrays, thus identifying antibody (Ab) signatures of human and murine Salmonellosis (Lee et al., 2012). Serum signatures can partly describe the status of the Ab response, but they do not reveal its ontogeny; moreover, serum Ab profiles might be discordant with memory or effector cell specificities (Guan et al., 2009). Knowing antigenic targets is certainly important for vaccine design, yet further research is necessary to understand the underlying mechanisms of response and protection; for instance, to explain why vaccines to Salmonella have only moderate, transient efficacy (McGregor et al., 2013).
Here we focused both on defining the specificities of the B cell response and addressing why it follows an extrafollicular pathway rather than a GC one. Our initial hypothesis was that the massive plasmablast response was polyclonal and non-specific, owing to innate immune receptor stimulation of B cells. Initial evidence indicated that the response was apparently non-specific. However, a series of experiments using a variety of approaches ultimately revealed a process in which very low affinity, yet specific, B cells—found at unexpectedly high precursor frequency—join the initial proliferative plasmablast response, and in the absence of developed GCs eventually acquired somatic mutations which in turn led to sufficient affinity maturation for the ultimate detection of conventional “specificity” for the immunizing bacteria. These results reveal an unappreciated pathway of response to a gram-negative bacterial pathogen and in addition lead to a revised view of the nature of clonal selection, specificity, affinity, and humoral immune response evolution.
RESULTS
STm Infection Induces Rapid AFC Accumulation, but Not GC Formation, in the Spleen
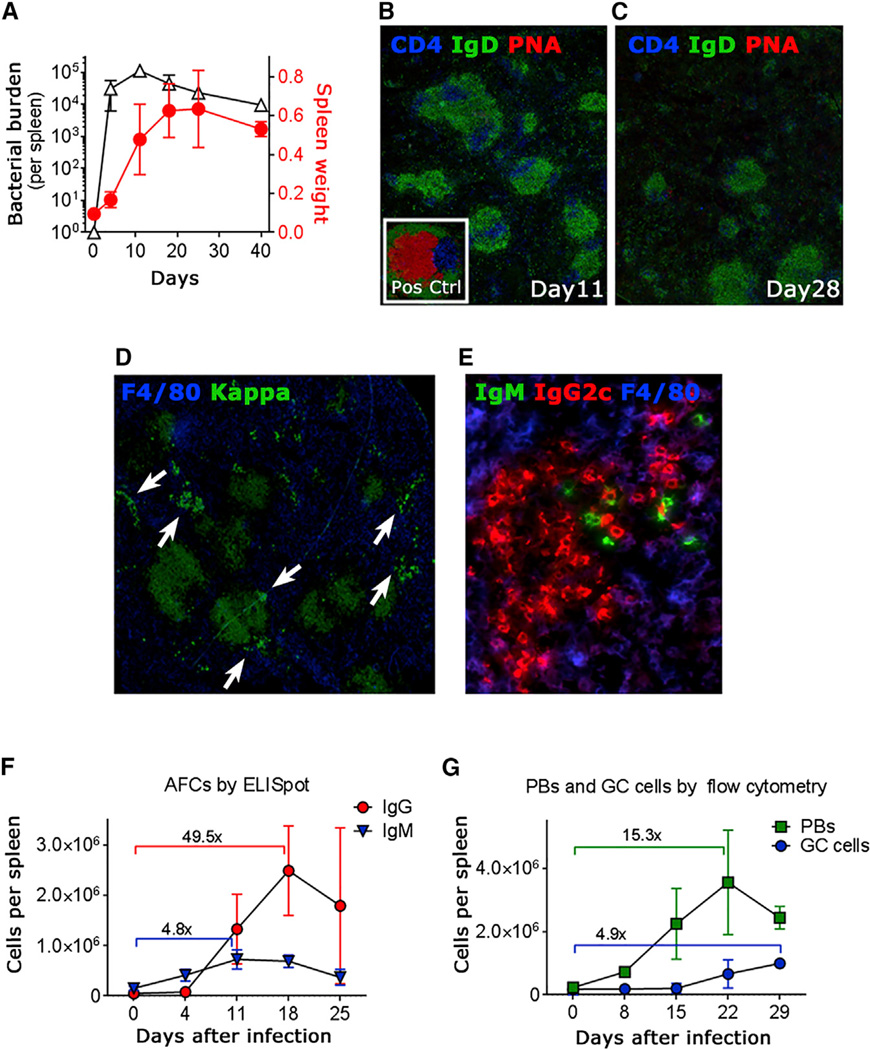
Following intraperitoneal (i.p.) administration of an aroA attenuated STm strain (Hoiseth and Stocker, 1981), rapid spreading of bacteria to several organs was observed, including to the liver, the gut, and especially the spleen, which rapidly increased in size (Figure 1A). Consistently with previous reports (Cunningham et al., 2007), there were few if any detectable GCs (Figure 1B and 1C) but there was massive accumulation of AFCs at EF sites (bright cell patches positive for intracellular Ig, Figure 1D). Of note, extensive class switching to IgG was detected at early stages of the response, with IgG2c as the dominant isotype (Figure 1E and [Cunningham et al., 2007]). We determined the kinetics of AFC accumulation and GC formation (Figure 1F and 1G) during the early to mid-phase (up to 30 days) of the immune response. The IgM AFC response was present at 4 days post-infection and peaked before 2 weeks, while the IgG response appeared and peaked slightly later but was remarkably larger in size, showing about a 50-fold expansion from baseline. The flow cytometry analysis showed a slight increase in the frequency of cells with a GC phenotype starting 3 weeks after infection, in contrast to immunofluorescence analysis, which we further addressed below.
Figure 1. STm Infection Induces a Rapid, Massive AFC Response without GCs.
(A) Spleen weight (red line) and splenic bacterial burden (black line) in infected mice over time.
(B–E) Immunofluorescence staining of cry-osections from spleens of infected mice for GCs (B, day 11; C, day 28), AFC patches (D, day 4, arrows) and isotype-switched IgG2c AFCs (E, day 11).
(F and G) Analysis of the abundance of AFCs by ELISpot (F) or of plasmablasts and GC cells by flow cytometry (G, gating as reported in Figure S1).
Data shown here are representative of at least six independent experiments. In (A), (F), and (G), data are represented as mean ± SD of groups of at least five mice. See also Figure S1.
Only a Minority of the AFCs Can Be Assigned to Salmonella Specificity
The massive AFC response led us to investigate the specificity of it. Given the limited knowledge and availability of STm Ags, we used a multi-pronged approach. First, we performed ELISpot assays using some of the few known Ags which are thought to be major targets, such as LPS and the outer membrane protein OmpA (Lee et al., 2012). In order to enumerate Ag-specific cells more broadly, we also used a lysate of STm as target. The percentage of AFCs specific for all of these Ags together was less than 0.5% of IgG and 2% of IgM (Figure 2A and 2B); this low frequency suggests that the vast majority of the STm AFC response is non-specific. However, ELISpot assays could lack sensitivity if the Ag concentration or the Ab affinities are low, although it must be said that even very low affinity hapten Abs can be detected by ELISpot, provided that the Ab is IgM (Dal Porto et al., 2002).
Figure 2. Screening of the Specificity of the AFC Response.
(A and B) ELISpot assays for Ag-specific IgG AFCs (Ags: LPS, OmpA, STm lysate) shown as total number of cells per spleen (A) or as percentage of the IgG and IgM AFC compartment (B). (C) Schematic panel illustrating the hybridoma production and screening strategy and methodologies. In the example showed here, an IgG mAb which was selected in the first screening is then probed on the Ag microarray, where LppA and LppB (proteins with high homology) were identified as the antigenic targets. (D and E)Summary of the frequency of Ag-specific IgG (D) or IgM (E) mAbs from hybridomas identified after the screening. Data shown in (A) and (B) are representative of at least six independent experiments and are represented as mean ± SD of groups of at least five mice. See also Figure S2.
Some of these issues can be overcome by producing mAbs and investigating the specificity at the single cell level. Using the traditional hybridoma technique (Figure 2C) we produced mAbs at an early (day 7) and late (day 21) time point, obtaining an extensive panel of both IgM (1,110) and IgG (650) lines. We initially screened all lines by ELISA (on the same Ags described above) and 480 of them by both ELISA and a flow cytometry assay that uses whole STm cells as a source of native Ag (Figure 2C). This process identified several lines that were either LPS-specific, STm-specific (mAbs reactive either to the lysate in ELISA or in the flow cytometry assay), or polyreactive (mAbs binding to more than one Ag, including control proteins such as GST and BSA in ELISA). A Salmonella Ag microarray has recently become available (Lee et al., 2012) and represents a powerful tool to dissect the specificity of the response. Because the multiplexing capability of the Ag microarray screening is currently relatively low, we tested all STm-specific clones (n = 20, 8 IgG and 12 IgM), and representative randomly selected “negative” lines (n = 55, 31 IgG and 24 IgM). For IgG mAbs, of the 8 assayed that were positive in the first screening 2 were further assigned to an Ag specificity. Of the 31 additional IgG mAbs, only one identified a target on the microarray. For the IgM, of the 12 positive mAbs, 3 identified a target on the array whereas 7 were shown to be polyreactive. Of the 24 randomly selected IgM mAbs, 6 bound an Ag on the microarray and 5 more were polyreactive. Representative profiles of polyreactivity, as well as of identified targets, that included NhaA, LppA/B, YbhC for IgG, and YjcS, HybG, FadD, DnaJ, OmpA, HemB, AphA for IgM, are shown in Figure S2. In summary (Figure 2D and 2E), by combining ELISA, flow cytometry, and microarray screening, we identified 21 STm-specific and 10 LPS-specific along with 31 polyreactive (30 IgM and 1 IgG) mAbs. It is striking that only 1 in 31 randomly selected IgG mAbs found a microarray hit, and that in general our extensive screening of 650 IgG mAbs led to the identification of only 11 STm-specific (1.7%) and 8 LPS-specific (1.2%) mAbs. This is consistent with the ELISpot, ELISA, and flow cytometry data, and strongly suggest that the great majority of IgG mAbs lack either specificity or sufficient affinity to be assigned to STm specificity.
TLR and IL-1R Signaling, and to Some Degree T Cells, Are Dispensable for AFC Responses
The specificity screening raised the question of whether STm induces a non-specific polyclonal plasmablast response; for instance, signaling via Toll-like receptors (TLRs) could be responsible for non-specific expansion of AFCs, such as in the case of the TI Ag LPS known in some cases to generate large amounts of isotype switched Abs (Koga et al., 1985). To test this possibility, we infected mice deficient for TLR2 and TLR4, as well as mice deficient for the adaptor protein MyD88 that signals downstream of all but one TLR (Figure 3). In both cases, a large switched and unswitched plasmablast response was still generated (Figure 3A), to an amount comparable to or higher than wild-type mice. Consistently, LPS-, lysate-, and OmpA-specific AFCs were detected in all three strains (Figure 3B–3D). Because interleukin-1 receptor (IL-1R)-family members also signal via MyD88, these data also exclude this axis as a likely source of non-specific B cell activation. This was confirmed in mice deficient for IL-1R as well as ASC (data not shown), excluding these canonical inflammasome-mediated mechanisms. It has been suggested that T cell signals contribute to selection; however, in the STm infection model, at least part of the EF response is independent of T cells (Figure S3 and [Gil-Cruz et al., 2009]). Together these data exclude key innate immune pathways, while suggesting that a substantial though not complete response can occur in the absence even of T cell signals.
Figure 3. Infection of Genetically Modified Mouse Strains.
(A–D) Assessment of the TLR-dependent component of the response. Wild-type B6, Myd88−/− B6, and TLR2-TLR4−/− B6 mice were infected and the AFC response was evaluated by Elispot for total IgG (solid lines in A) and IgM (dotted lines in A), LPS (B), STm lysate (C), and OmpA (D). Because infection with the standard dose used throughout our study resulted in high mortality rate in Myd88-deficient mice (not shown), these were infected with a lower amount of STm (2 × 104), which permitted 100% survival after 30 days. (E–G) Mice with engineered Ab genes/BCR were infected and the splenic AFC response was measured for: total IgG in wild-type C57BL/6, B1.8+/+(C57BL/6), or B1.8+/+ Jk−/− mice (C57BL/6) (E); total IgG (F) or OmpA-specific IgG (G) in wild-type Balb/c, Vk8R+/+ (BALB/c), or Jk−/− (Balb/c) mice. In all panels data are representative of at least two independent experiments and are represented as mean ± SD of groups of at least four mice.
A Diverse BCR Repertoire Is Required for Mounting the AFC Response
We then investigated the B cell receptor (BCR) specificity requirement for the response to take place. We reasoned that, if specific sensing of Ags via the BCR were required for B cell activation, then limiting the BCR repertoire should affect the response; whereas, if the response were truly non-specific (i.e., independent of BCR specificity), then it should be unaffected by restricting the BCR repertoire. This can be accomplished in various mice with engineered Ab genes (Figure 3E–3G). First, we infected mice homozygous for the B1–8 IgH site-directed transgene, which have a single VH and variable VL genes. These mice showed a dramatic reduction in the IgG AFC response (Figure 3E). The response could be further curtailed to background amounts in B1–8+/+ Jk−/− mice, where the VL repertoire was restricted to the λ locus. Similarly, infection of Vk8R+/+ mice and Jk−/− mice, both of which have variable VH but limited VL repertoires, showed a marked reduction in the quantity (Figure 3F) and quality (Figure 3G) of the IgG AFC response. Thus, a limited BCR repertoire resulted in a markedly diminished total and Ag-specific IgG response. These results indicate that indeed certain BCRs are selected by STm Ag, consistent with standard clonal selection theory.
The B Cell Response to STm Comprises a Subset of the Repertoire Undergoing Vigorous Somatic Hypermutation
It was conceivable that the inability to detect Ag specificity in the early response was due to low affinity of the great majority of responding B cells. If this were true, specificity would mainly be detected as a result of affinity maturation driven by somatic hypermutation (SHM). This scenario, which requires time, was suggested by the fact that LPS- or lysate-specific AFCs appeared later in the response. Furthermore, if the response were indeed specific at the outset, then the responding repertoire of BCRs should be narrower than the overall repertoire.
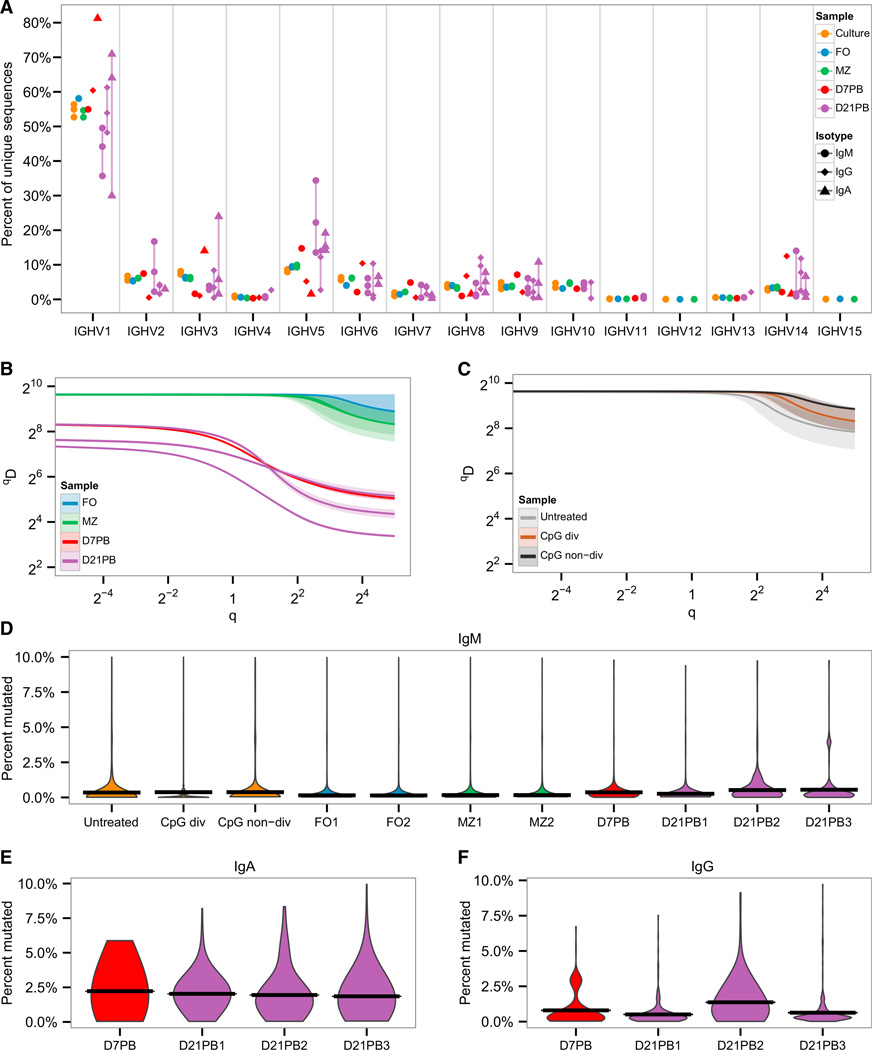
To assess to what extent SHM takes place, as well as the breadth and clonality of the responding repertoire of B cells, we performed high throughput sequencing of heavy chain VDJ mRNA. We employed a single-molecule barcoding strategy to tag each starting mRNA with a unique molecular identifier (UID), enabling computational assembly of unique mRNA amplification products into low-error consensus sequences (Shugay et al., 2014; Vander Heiden et al., 2014; Vollmers et al., 2013). This mitigates the impact of sequencing error, and improves the accuracy of mutation identification (Figure S4). The repertoires of plasmablast populations, sorted at either an early (day 7) or a late (day 21) time point, were compared to naive B cells, further divided in follicular (FO) and marginal zone (MZ) B cells. We analyzed approximately 14 million raw reads, yielding 346,349 and 3,885 unique, high-confidence VDJ arrangements from all naive and PB samples, respectively (Figure S4 and Table S1). The response was diverse (Figure 4A), with a representation of all VH families and very limited evidence for preferential usage of VH or JH genes by plasmablasts (Figure S4). Nonetheless, when examined at the level of individual sequence diversity, the plasmablast response showed clear signs of expansion and selection (Figure 4B and Figure S4). Clonal diversity was characterized using the general diversity index (qD) proposed by Hill (Hill, 1973), which encompasses a range of commonly used diversity measures as a smooth curve over a single varying parameter q (see Supplemental Information). According to this approach, the overall diversity of the naive population was similar between marginal zone and follicular B cell samples (Figure 4B). In contrast, the plasmablast populations at both day 7 and day 21 post infection had a lower qD index for all q indicating that they were significantly more focused (i.e., exhibited reduced diversity) compared to the naive repertoire. This reduction in diversity was apparent in terms of both the total number of clones (q → 0) and clonal dominance (q → ∞). This pattern was unchanged when considering diversity for each isotype separately (data not shown).
Figure 4. Repertoire Characteristics of the Plasmablast Response to STm Infection.
IgM sequences of naive follicular (FO) and marginal zone (MZ) B cells from uninfected mice were compared to IgM, IgG, and IgA sequences from plasmablasts 7 days (D7PB) and 21 days (D21PB) post-infection.
(A) V family usage (x axis) shown for different isotypes as the percentage (y axis) of processed sequences within each sample.
(B) Comparison of the Hill diversity index (qD, y axis) over varying diversity orders q (see Supplemental Information), between IgM sequences from naive samples and IgM, IgG, and IgA sequences from samples of infected mice. The median diversity score over all resampling realizations is plotted as a line with the middle 95% percentile of the resampled distribution indicated by a shaded background. All samples were randomly downsampled to 800 sequences for each resampling realization to correct for variations in sequencing depth.
(C) Comparison of Hill diversity indices between IgM sequences grown in cell culture under three conditions: untreated, cells that proliferated after CpG treatement (CpG div), and cells that did not proliferate after CpG treatment (CpG non-div). All samples were uniformly downsampled to 800 sequences for comparison to (C).
(D–F) Violin plots showing the distribution of mutations from the germline (y axis) for each sample (x axis) and isotype. Only sequences with ≤ 10% mutation frequencies are shown. See also Figure S4 and Table S1.
In order to assess whether the reduced diversity of the plasmablast response was evidence for antigen specificity, rather than a result of stochastic expansion of non-specific clones, we compared the repertoire diversity shift induced in vitro by a non-antigen-specific stimulus, such as CpG, which elicits proliferation of both follicular and MZ B cells. Figure 4C shows that stimulated and divided B cells had very similar repertoire diversity as untreated or non-divided cells, which was different from what was observed for the STm plasmablast repertoire (Figure 4B).
Finally, we analyzed the mutational content of the sorted populations (Figure 4D–4F and Table 1). Three weeks after infection, the average number of mutations per sequence was 5.6 (3.4, 6.7, and 8.6 for IgM, IgG, and IgA, respectively), rising from 3.7 (3.0, 3.8 and 8.8) at day 7 and a background amount in naive cells of 0.9. Mutations in the naive sequences are a consequence, at least in part, of sequencing and germline database errors, as well as possibly mutations in the preimmune repertoire and the inclusion of natural IgM memory B cells that share surface markers with naive B cells (Tomayko et al., 2010). Taking this into account, even when only sequences with more than one mutation were included in the analysis, the average frequency of mutated Ab genes at 21 days post-infection was 50% (24%, 47%, and 83% for IgM, IgG, and IgA, respectively), compared to 31% (23%, 40%, and 73% for IgM, IgG, and IgA, respectively) at day 7 and 7% in naive samples. Hence, STm-elicited plasmablasts contain numerous mutations and the mutational content increases over time, while the diversity of the responding plasmablasts is significantly narrower than either the FO or MZ preimmune repertoire. Both of these are consistent with selection despite the apparent lack of specificity.
Table 1.
SHM in the Ab Repertoire as Determined by High-Throughput Sequencing
| Sample | Isotype | Mean Mutation Count |
Mean Mutation Frequency |
Sequences with >0 Mutations |
Sequences with >1 Mutation |
Sequences with Mutation Frequency ≥0.01 |
|---|---|---|---|---|---|---|
| Untreated | IgM | 2.900 | 0.008 | 26.3% | 15.7% | 10.3% |
| CpG div | IgM | 3.250 | 0.009 | 29.4% | 14.2% | 9.0% |
| CpG non-div | IgM | 3.230 | 0.009 | 25.4% | 13.1% | 9.3% |
| FO1 | IgM | 0.831 | 0.002 | 18.0% | 6.4% | 2.8% |
| FO2 | IgM | 0.807 | 0.002 | 17.2% | 6.0% | 2.6% |
| MZ1 | IgM | 1.021 | 0.003 | 20.8% | 8.1% | 3.4% |
| MZ2 | IgM | 0.911 | 0.003 | 21.4% | 8.4% | 3.3% |
| D7PB | IgM | 3.088 | 0.009 | 66.5% | 23.3% | 6.0% |
| D7PB | IgG | 3.854 | 0.011 | 77.6% | 39.6% | 21.4% |
| D7PB | IgA | 8.859 | 0.025 | 93.8% | 73.4% | 71.9% |
| D21PB1 | IgM | 1.914 | 0.005 | 49.2% | 14.4% | 4.4% |
| D21PB1 | IgG | 2.683 | 0.007 | 75.7% | 35.6% | 12.0% |
| D21PB1 | IgA | 8.913 | 0.025 | 97.5% | 90.7% | 76.6% |
| D21PB2 | IgM | 3.700 | 0.010 | 67.0% | 31.7% | 18.5% |
| D21PB2 | IgG | 8.043 | 0.022 | 93.5% | 67.3% | 51.8% |
| D21PB2 | IgA | 7.575 | 0.021 | 98.2% | 80.2% | 71.3% |
| D21PB3 | IgM | 4.645 | 0.013 | 65.4% | 26.6% | 13.1% |
| D21PB3 | IgG | 4.538 | 0.013 | 87.2% | 36.4% | 17.8% |
| D21PB3 | IgA | 9.281 | 0.026 | 94.9% | 78.1% | 65.3% |
For naïve B cells, follicular and marginal zone B cells from two mice were analyzed separately. No significant differences were seen, and data are pooled here.
SHM Takes Place in Follicles and at Extrafollicular Sites
Given evidence for extensive SHM, it was compelling to investigate where it takes place. The appearance of specific AFC occurring prior to GC formation would be consistent with extra-GC SHM and Ag-driven affinity maturation. Although SHM does not canonically happen at EF sites, it does occur in murine spleen in the autoimmune setting (William et al., 2002), showing that GCs are not strictly necessary for this process; however, a physiological counterpart to extra-GC mutation has to date not been revealed in a murine pathogen-specific response. The rapid appearance of isotype-switched plasmablasts post-infection shows that activation-induced cytidine deaminase (AID) is activated very early in the response among extrafollicular B cells, making it plausible to think that SHM could also be occurring.
To resolve this, we first searched for GC-like structures that might have been an alternative site for SHM. The apparent contrast between the lack of GCs as assessed by immunofluorescence and the flow cytometric detection of cells with a GC phenotype (Figure 1), the frequencies of which admittedly did not rise above baseline until day 22, could be reconciled by the finding, starting 3 weeks after infection, of scattered small PNA-positive cell aggregates in some sections. The cells in these small clusters expressed the PNA target weakly, as they were only found by overexposing PNA; they also expressed low, but detectable, amounts of the GC marker Bcl6 (Figure S5). These clusters, which we refer to as “GC-like,” were found at an atypical site—the interface between the B and T cell zones—and did not appear to develop past this stage into proper GCs for the duration of our studies, up to 6 weeks after infection. A similar response was observed in BALB/c mice, with lack of proper GCs and a remarkable EF plasmablast response (Figure 3; data not shown). Together these data suggest that GCs could potentially form at late time points, but do not mature. We considered whether innate immune signals in this context could in fact suppress GC formation. Indeed, when BALB/c mice deficient for MyD88 were infected, fully developed GCs were observed as early as 10 days after infection, accompanied by EF plasmablasts, and were more numerous by day 15 (Figure S5). However, this was not observed in C57BL/6 MyD88-deficient animals, indicating that additional factors may suppress GC formation in a strain-specific manner.
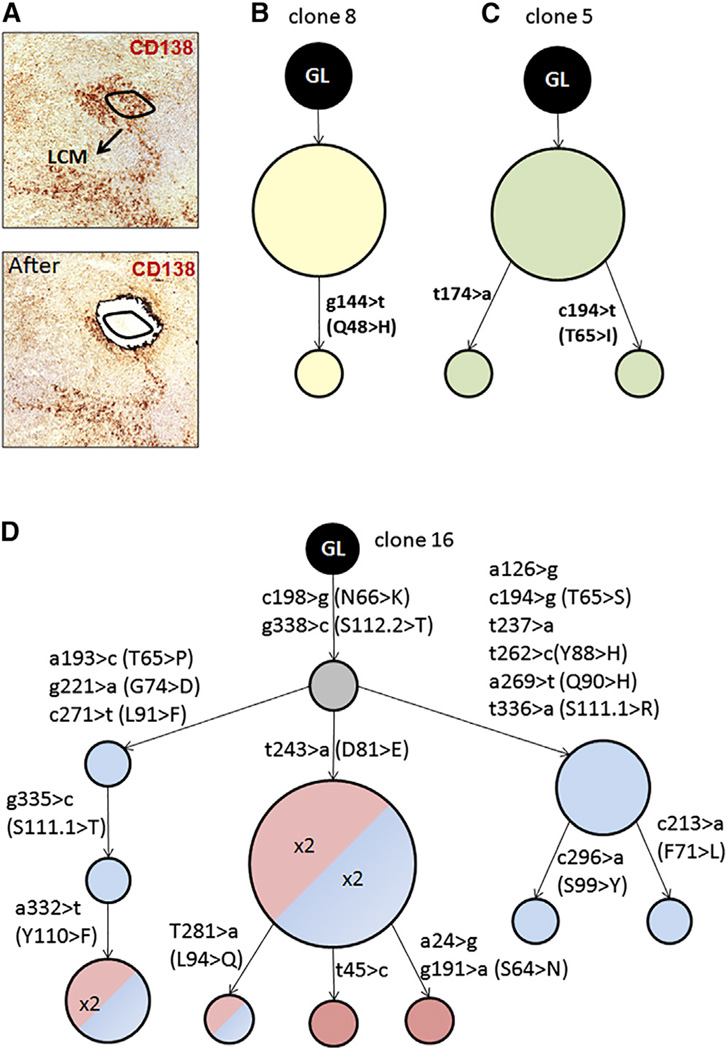
To address whether SHM takes place in “GC-like” structures and/or at EF sites, we performed laser capture microdissection (LCM) and V region sequencing of both structures (example, Figure 5A) at 3 weeks post-infection. Picks comprised 20 to 30 cells and in the case of plasmablasts, whenever possible, the same patches were taken from two or three consecutive slides. We dissected both adjacent and distant patches (Table S2). 14 plasmablast picks yielded a PCR product that was then cloned, followed by sequencing of V gene inserts in multiple colonies derived from each product. 79% of the unique sequences obtained showed some somatic mutations, averaging 3.2 mutations per mutated sequence (Tables S3 and S4). When clonally related sequences (with the same VDJ rearrangement) were found, then a lineage tree was built that describes the evolution of the clone. 11 of 14 picks gave clones to generate lineage trees. Such trees demonstrate that ongoing V region diversification was taking place among the few cells captured in each such microdissected patch. Figure 5B–5D shows representative trees (see also Figure S6); in some cases (Figure 5B and 5C) the trees were fairly simple, with two or three clonally related sequences that were exclusively derived from the same pick. In other cases, there was higher complexity, with sequences from different, though adjacent, picks (Figure 5D). The mutations observed in the lineage trees were authentic and not a result of PCR error: the rates are far higher than that expected from PCR error using high-fidelity polymerases (William et al., 2002). Moreover, the presence of shared mutations is not expected from PCR error, but is expected from clonal expansion and selection of authentic SHM. The isolation of the same mutation from different picks of the same geographic patch (i.e., taken from different serial sections), and hence different PCR amplifications fully excludes PCR error as the explanation. Rather, when one considers that it takes only 4 divisions to create a cluster of 16 cells, and that SHM in GCs introduces about 0.25 mutations/V region/division, then the extent of mutation is consistent with this high rate, as there should be roughly 1 difference between each sequence (Kleinstein et al., 2003). In parallel, we dissected and sequenced some of the GC-like structures. A similar degree of SHM was observed in these structures, with 70% of the sequences being mutated with an average of 2.3 mutations per mutated sequence (Table S3). Overall, these data show that there is robust diversification through SHM, and as extensively shown and discussed below, that this is likely to occur locally and determine affinity maturation.
Figure 5. Ongoing SHM Takes Place in Plasmablast Patches.
(A) Example of a plasmablast patch pick by laser capture and microdissection (LCM); the same patch is shown before and after the pick. The black line indicates the area of ~20 cells that was dissected. The plasmablast patches are identified with anti-CD138 staining by immunohistochemistry. (B-D) Examples of three clonal trees of different complexity derived from the analysis of the Ab gene sequences obtained by LCM. The size of each node indicates the number of identical sequences found. In (D), a more complex tree is shown that was composed of sequences that derived from several nearby picks; different colors of the nodes denote different, but adjacent, picks from which the sequence was derived, while the number within the node indicates in how many serial slides (always from the same patch) the same sequence was found. The gray circle indicates an inferred intermediate. The position of the mutated nucleotides and aminoacids (in the case of replacement mutations) are shown along the branches. See also Figure S6 and Tables S2–S4.
SHM Results in Efficient Affinity Maturation
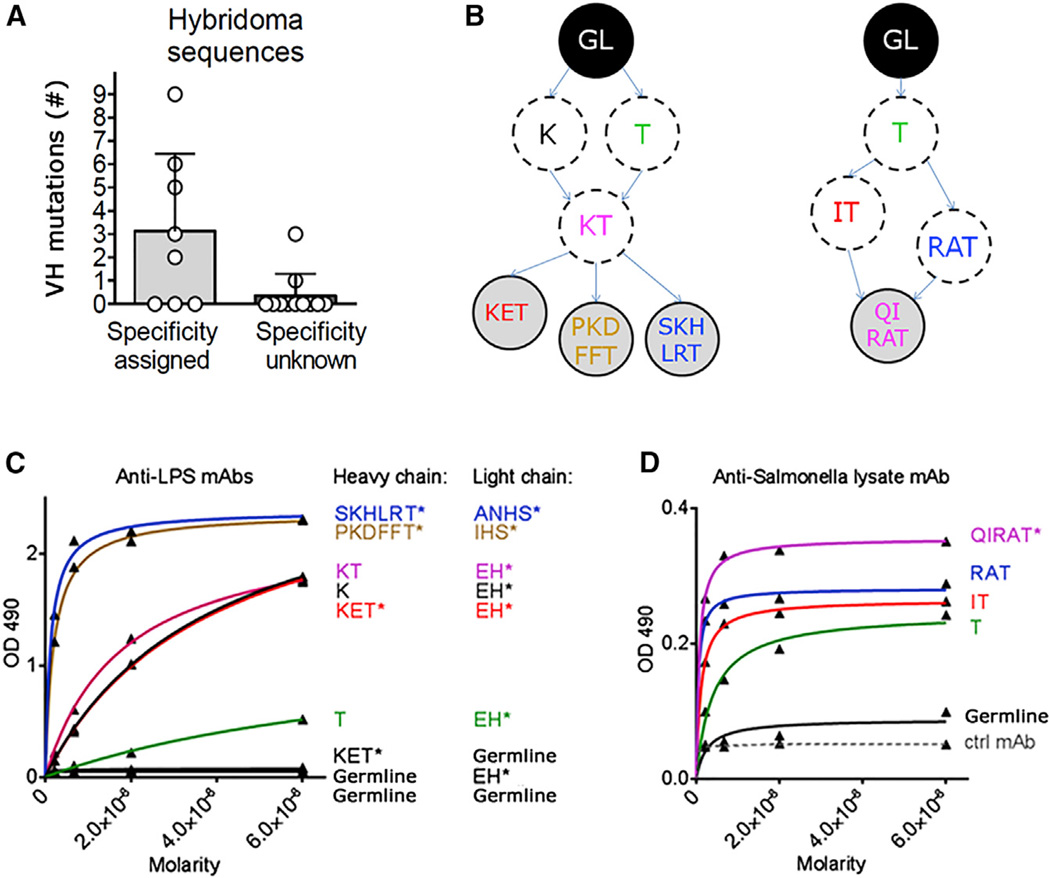
If SHM-driven affinity maturation promotes development of apparent Ag-specificity, then Ag-specific hybridomas should have more mutations and these mutations should directly promote Ag-binding. Indeed, hybridomas that were assigned to STm specificity had on average more than three mutations/ sequence, more than those with unknown specificity (Figure 6A). To address the function of these mutations, we selected four hybridomas with known specificity and that carried several mutations. We then cloned and expressed their V regions in recombinant form (Wardemann et al., 2003) and generated complete germline mutational revertants, as well as some partially mutated intermediates (Figure 6B). First, we investigated LPS-specific mAbs; three different but clonally related hybridomas were isolated that carried both shared and unique mutations. Affinity decreased markedly as mutations were removed, both for the heavy and the light chain (Figure 6C). In particular, a mutation leading to lysine (K) in position 66, which was shared by all isolated hybridomas and hence arose early in the lineage, appeared to have a large effect, as an artificial intermediate version without it (green line) showed a dramatic reduction in affinity. Importantly, when either the heavy or the light chains were reverted to germline, these mAbs showed no detectable binding to LPS. The remaining portion of the spleen that was not used to generate these hybridomas was used for LCM experiments; remarkably, this same LPS-specific clone was also identified in one of the largest dissected patches (Table S3). Hence, in this case we could link affinity maturation to the EF site. The correlation between SHM and affinity was confirmed for another mAb, specific for the STm lysate (Figure 6D), that showed stepwise loss of affinity as mutations were reverted, with binding of the germline revertant only slightly above background. This mAb carried several mutations in the heavy chain only, but had a germline light chain. Thus, we found that SHM via a series of steps had increased the affinity for STm antigens from barely or not detectable to a high degree of binding.
Figure 6. Effect of SHM on Affinity in Cloned mAbs.
(A) Number of mutations in Ab sequences from hybridoma clones.
(B) Diagram indicating the relationships between the mAbs we isolated and the mutants we made, with respect to the heavy chain. The letters inside the circles represent amino acid substitutions as compared to the germline (GL, black circle on top). Circles on a gray background represent Abs that were found in the hybridoma screening; dotted circles on white background represent artificial intermediate versions.
(C and D) Binding curves of recombinant mAbs expressed in their original or germline form, as well as in artificially mutated intermediate versions. The asterisks indicate the original sequences isolated from the hybridoma clones. An IgG sandwich ELISA was performed in parallel to confirm that all Abs were used in equal amounts in the assay (data not shown).
DISCUSSION
Given that STm was reported to induce only delayed GC responses, we sought to determine the nature of the alternative B cell response and whether this could create affinity matured Ab and memory. Our initial observation was that the massive AFC response to STm appeared to be largely non-specific. The lack of apparent specificity was determined by multiple methods, perhaps most convincingly by screening large panels of mAbs on a Salmonella Ag microarray, as well as against STm lysate and intact bacteria. We sought possible explanations for the broad B cell response seen in the STm infection model. Despite having tested key elements of TLR, IL-1R, and inflammasome pathways, we have not yet uncovered a receptor that could explain non-specific activation of B cells, though we cannot exclude that one exists.
Rather, several lines of evidence suggested that the response is in fact specific, but of very low affinity. A critical test of this idea was the assessment of the response in a variety of different mice with restricted BCR repertoires. Restriction of the BCR repertoire greatly reduced the response, which diminished according to the degree of repertoire restriction, and almost disappeared when both VH and VL were limited. In contrast, a truly non-specific response would have been unaffected by B cell repertoire restriction. We propose a model in which promiscuously selected, yet specifically stimulated, B cells then undergo mutation and selection at EF sites to eventually generate what appears to be Ag-specific cells at late time points.
Consistent with the model Ag-specific cells indeed emerged for the most part only late during the response. This would be expected if the response were of such low affinity that standard immunoassays could not usually detect the specificity until and unless somatic mutation and selection improved the affinities. In line with this, most of the mAbs with demonstrable STm specificity were encoded by mutated genes, while most “non-specific” mAbs were unmutated. This supports either of two scenarios: that most Abs are in fact specific for STm, but it is necessary that they mutate and mature in order to gain sufficient affinity to be detected; or alternatively, that most Abs are not specific and that only those few with some low degree of affinity in their germline configuration can enter a pathway that allow them to mature. To help distinguish these scenarios, we reverted mutations observed in specific Abs. In the two cases tested, the binding of Abs to their target was strongly dependent on affinity gains determined by mutated amino acids, as shown by the stepwise loss of reactivity as such mutated genes were progressively reverted to their germline. This result strongly argues that a broad repertoire of B cells that have a very low initial affinity for the antigen get activated and only later in the response become detectable as Ag-specific as a consequence of affinity maturation.
A key finding of our work is that the process that generates higher affinity Abs from these low-affinity precursors does not appear to depend on GCs, which have been canonically thought to be the exclusive site for SHM. Two pathways might contribute to this. The first involves small “GC-like” structures that occasionally appeared about 3 weeks after infection; these consisted of a small number of weakly PNA+ and Bcl6+ cells at the B-T cell border rather than the follicular center site of classical GCs. These cells might represent partially differentiated or immature GC B cells. These structures remained at a primitive stage without fully developing to GCs, even at later time points. The second pathway involves the EF site, where indeed V region mutations do occur during the expansion of the response. Notably, a similar phenomenon was described before in autoimmune lupus-prone mice (William et al., 2002) and to some extent in humans in non-lymphoid sites such as synovia or rheumatoid arthritis patients (Kim et al., 1999; Scheel et al., 2011; Schröder et al., 1996; Stott et al., 1998).
In fact, SHM took place vigorously within both pathways, and especially at EF sites. The mutational process itself could occur in plasmablasts, as well as in B blasts, both of which were shown in other contexts to express AID (Marshall et al., 2011; Odegard et al., 2008; Toellner et al., 1996). Clearly, isotype switching, mainly to IgG2c, another AID-dependent process, was also occurring at these sites. The extent of SHM in GC-like areas was not higher than in EF patches, and 25% to 40% of sequences within the GC-like structures (but not in EF patches) harbored stop codons in the complementarity determining region-3 (CDR3) region. Importantly, GC-like structures and adjacent EF locations did not share B cell clones, thus further pointing to a different origin of these two structures. As was shown originally in classic work proving that GCs were sites of SHM (Jacob et al., 1991), identical or clonal sequences sharing common mutations and VDJ junctions were found within small clusters of 10–20 cells. This could only arise if mutations were actively being incorporated into V regions of dividing cells at that site. In our dataset, clonally related sequences were found in the same patch dissected from consecutive serial sections or, in some cases, from adjacent patches, but never among sequences derived from more distant sites. This further indicates that a substantial proportion of the mutations we observed occurred locally, because clonal sequences would not be confined to local picks had mutations happened elsewhere. Though SHM actively takes place at both locations, based on the earlier appearance and much greater overall size of the EF patches, we suggest that these are likely the source of the majority of the affinity-matured “specific” AFCs.
This conclusion is directly supported by one case in point, in which the spleen from an infected mouse was used to perform both LCM and hybridoma production. Sequences were found among the plasmablast LCM picks that were clonally related to LPS-specific mAbs identified in the hybridoma screening. Importantly, this clone was shown in our mAb revertant experiments to have strong binding to LPS that was very much dependent on somatic mutations. Hence, this allowed us to directly show affinity maturation within an EF plasmablast-derived clone that was actively undergoing SHM.
At the broadest level, our results suggest a revised view of how clonal selection and affinity maturation operate in the response to a pathogen. The response is dominated by plasmablast formation at EF sites for at least several weeks. Plasmablasts uniquely can both expand and generate effector function, secreting Ig. We propose that such a response evolved to ensure protective function to rapidly replicating pathogens and needs to be generated as quickly as possible; failure to do so might result in death.
A major insight from this report is that these responses achieve great magnitude by engaging extremely low affinity B cells; yet the response is far from “non-specific.” Moreover, to a substantial degree this early response is independent of T cells and hence needn’t wait for effective T cell priming. This strategy would not only provide a measure of immediately effective pathogen protection, it also would generate a large number of clones that can subsequently be optimized. As we and others (Marshall et al., 2011; Toellner et al., 1996) show, plasmablasts and B blasts can quickly undergo isotype switch, a process independent of GCs and even to some degree of T cells; thus, a variety of effector functions aside from IgM are rapidly engaged. Isotype switching is a process that requires AID activity, which is also necessary and sufficient for SHM (Muramatsu et al., 2000). The fact that extensive isotype switching takes place in the EF response thus unequivocally indicates that functional AID is being expressed. It has further been established that AID is expressed in EF responses, at least in some models (Marshall et al., 2011; Odegard et al., 2008). Indeed, a second important finding is the unexpected discovery that the EF plasmablast response is permissive for SHM and in fact affinity-based selection. The evidence from the microdissection data, the high-throughput sequencing data, and the hybridoma sequencing data, together demonstrate that at least some and probably most of the mutations do occur at EF sites. In conclusion, SHM and affinity selection serve to improve the initial, GC-independent, low affinity response in a progressive way, as our mAb mutation and re-expression analysis demonstrated.
While at present this pathway has been demonstrated for Salmonella, as we have noted, a large, potentially “non-specific” early plasmablast response is a feature of immune responses to many pathogens; hence, it is likely that other aspects such as low starting affinity, early expression of AID, and GC-independent onset of SHM and affinity maturation, will be featured in responses to other pathogens. If so, then the textbook depiction of the humoral immune response might need to be modified to include this physiologically important alternative to the GC-dependent affinity maturation pathway.
EXPERIMENTAL PROCEDURES
Mice, Bacteria, and Infection Procedures
The mice strains used in this study were obtained as described in the Supplemental Information and bred under specific pathogen free conditions in the animal facility at Yale University. The aroA attenuated Salmonella Typhimurium strain SL3261 (Hoiseth and Stocker, 1981) was kindly provided by Roy Curtiss III, Arizona State University. For infection, 105 bacteria (unless otherwise specified) in PBS were injected i.p. The bacterial burden was assessed by plating serial dilutions of tissue homogenates, prepared by 1 mm-beads disruption in a FastPrep-24 instrument (MP Biomedicals). All mouse work was according to protocols approved by the Yale Institutional Animal Care and Use Committee.
Immunofluorescence Assays
7 µm spleen sections were prepared from OCT-frozen tissues, fixed in acetone for 10 min, and stored at −80°C. Antibodies and procedures used are described in Supplemental Information.
Antigens
Salmonella Typhimurium LPS was purchased from Sigma. A STm lysate was obtained by cell disruption using 0.1 mm beads in a FastPrep-24 instrument (MP Biomedicals). Recombinant antigens were generated by cloning STm genes in the pGEX-6P-1 vector (GE Life Sciences) vector. The following genes of interest were amplified directly from the same SL3261 STm strain used for the following infections: ompA, ompD, ybhC, FliC, and FljB. The recombinant plasmids were transformed in E. coli and GST-tagged recombinant proteins were produced and purified with Glutathione Sepharose 4B (GE Life Sciences) according to standard protocols.
ELISpot and Flow Cytometry Analysis of Splenocytes
Single cell suspensions from spleen were obtained by mechanical disruption of the tissue, followed by treatment with ACK buffer for lysis of red blood cells. For ELISpot assay, Immulon 4-HBX plates were coated with the following antigens: anti-kappa at 5 µg/ml, LPS at 5 µg/ml, STm lysate at 50 µg/ml, recombinant antigens at 10 µg/ml. Non-specific binding was blocked with 1% bovine serum albumin in PBS. Splenocytes were incubated for 6 hr at 37°C. AFCs were detected by using alkaline phosphatase-conjugated secondary antibodies (to IgG or IgM, Southern Biotech) and 5-bromo-4-chloro-3-indolyl-phosphate in agarose. For flow cytometry experiments, antibodies were purified in the laboratory or purchased as indicated in the Supplemental Information. Nonspecific binding to Fcg receptors was blocked with the anti-FcR clone 2.4G2; dead cells were excluded by ethidium monoazide staining, and doublets by scatter analysis. Cells were analyzed on a LSRII instrument (BD) and data analyzed by FlowJo software (TreeStar).
mAb Production and Screening
Hybridoma lines and clones were generated as reported in Supplemental Information. The original Ab genes were amplified (Tiller et al., 2009) from cDNA generated from hybridoma clones. Briefly, RNA was isolated using the RNeasy Mini kit (QIAGEN) per the manufacturer protocol, and cDNA prepared with the iScript cDNA synthesis kit (Bio-Rad) using random hexamers. Germline revertants or intermediate versions were generated by oligonucleotide-directed mutagenesis and then assembled by PCR. Recombinant mAbs were produced as described elsewhere (Wardemann et al., 2003). The amount of each recombinant mAb was estimated by an IgG sandwich ELISA and compared to a standard. For ELISA, antigens were coated O/N on Immulon 2-HBX plates at the same concentrations reported above and as described in Supplemental Information. Live bacterial flow cytometry was performed as follows: STm was grown O/N in LB media, aliquoted in a 96-well plate at approximately 106 bacteria per well and spun down. Bacteria were resuspended in undiluted hybridoma supernatant and incubated for 1 hr. After washing with PBS, an Alexa-488-labeled anti-kappa secondary Ab was added. After 30 min, bacteria were washed, fixed in 1.5% PFA, and analyzed on a FACSCalibur (BD). Microarrays were fabricated and probed with undiluted hybridoma supernatants as described previously (Lee et al., 2012).
Cell Sorting and High-Throughput Repertoire Sequence Analysis
Plasmablasts, naive follicular B cells, marginal zone B cells, and the cells for CpG DNA stimulation were obtained and treated as described in detail in Supplemental Information. For all cell types, after sorting cells were spun down and RNA isolated as described above. 250 ng of RNA was reverse-transcribed into cDNA using a biotinylated oligo dT primer. Libraries were created and sequenced as described in detail in the Supplemental Information. Raw high-throughput sequencing reads were quality controlled, assembled and filtered using pRESTO (Vander Heiden et al., 2014). V(D)J germline segments were determined using IMGT/HighV-QUEST (Alamyar et al., 2012). Clonal clustering was carried out and lineage inference was performed on each cluster using the software package PHYLIP 3.6 (N.T.G., unpublished data).
Laser Capture Microdissection
7 µm spleen sections were prepared from OCT-frozen tissues on the membrane-coated PEN slides (Leica). Plasmablast patches were detected using anti-CD138 and GC-like structures using PNA and anti-IgD in immunohistochemistry as reported in Supplemental Information. Microdissections were performed using a Leica LMD6500 instrument equipped with an optical microscope. Dissected patches were collected in the cap of PCR microtubes in 10 ml of digestion buffer (50 mM Tris-HCl, 50 mM KCl, 0.63 mM EDTA, 0.22% Igepal, 0.22% Tween20, 0.8 mg/ml proteinase K). Patches were digested at 55°C for 2 hr, then at 90°C for 5 min, and used for PCR amplification of Ab genes. Primers, PCR procedures, and data analysis are described in Supplemental Information.
Supplementary Material
Highlights.
Salmonella (STm) induces a large plasmablast response that is seemingly non-specific
The response is actually specific with affinities that are too low to detect
Extrafollicular SHM leads to affinity maturation and detectable affinities
The STm response differs markedly from the classical GC response to model antigens
ACKNOWLEDGMENTS
S.H.K. was supported by NIH Grant R01-AI104739. J.A.V.H. and N.T.G. were supported by NIH Grant T15-LM07056 from the National Library of Medicine. This work was funded by NIH grants R01-AI043603, R01-AR044077, and R01-AI 073722.
Footnotes
SUPPLEMENTAL INFORMATION
Supplemental Information includes six figures, four tables, and Supplemental Experimental Procedures and can be found with this article online at http://dx.doi.org/10.1016/j.immuni.2015.06.013.
AUTHOR CONTRIBUTIONS
R.D.N. designed and performed experiments, analyzed data, and wrote the manuscript. R.A.E., NT., S.-J.L., J.M.B., and T.J.G. performed experiments and analyzed data. J.A.V.H. and N.T.G. analyzed the sequencing data. S.H.K., F.V., E.M., and S.J.M. supervised experiments. M.J.S. designed experiments, supervised the study, and wrote the manuscript.
REFERENCES
- Alamyar E, Duroux P, Lefranc MP, Giudicelli V. IMGT(®) tools for the nucleotide analysis of immunoglobulin (IG) and T cell receptor (TR) V-(D)-J repertoires, polymorphisms, and IG mutations: IMGT/V-QUEST and IMGT/HighV-QUEST for NGS. Methods Mol. Biol. 2012;882:569–604. doi: 10.1007/978-1-61779-842-9_32. [DOI] [PubMed] [Google Scholar]
- Arpaia N, Godec J, Lau L, Sivick KE, McLaughlin LM, Jones MB, Dracheva T, Peterson SN, Monack DM, Barton GM. TLR signaling is required for Salmonella typhimurium virulence. Cell. 2011;144:675–688. doi: 10.1016/j.cell.2011.01.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barr TA, Brown S, Mastroeni P, Gray D. TLR and B cell receptor signals to B cells differentially program primary and memory Th1 responses to Salmonella enterica. J. Immunol. 2010;185:2783–2789. doi: 10.4049/jimmunol.1001431. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bobat S, Flores-Langarica A, Hitchcock J, Marshall JL, Kingsley RA, Goodall M, Gil-Cruz C, Serre K, Leyton DL, Letran SE, et al. Soluble flagellin, FliC, induces an Ag-specific Th2 response, yet promotes T-bet-regulated Th1 clearance of Salmonella typhimurium infection. Eur. J. Immunol. 2011;41:1606–1618. doi: 10.1002/eji.201041089. [DOI] [PubMed] [Google Scholar]
- Calderón I, Lobos SR, Rojas HA, Palomino C, Rodríguez LH, Mora GC. Antibodies to porin antigens of Salmonella typhi induced during typhoid infection in humans. Infect. Immun. 1986;52:209–212. doi: 10.1128/iai.52.1.209-212.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coro ES, Chang WL, Baumgarth N. Type IIFN receptor signals directly stimulate local B cells early following influenza virus infection. J. Immunol. 2006;176:4343–4351. doi: 10.4049/jimmunol.176.7.4343. [DOI] [PubMed] [Google Scholar]
- Cunningham AF, Gaspal F, Serre K, Mohr E, Henderson IR, Scott-Tucker A, Kenny SM, Khan M, Toellner KM, Lane PJ, MacLennan IC. Salmonella induces a switched antibody response without germinal centers that impedes the extracellular spread of infection. J. Immunol. 2007;178:6200–6207. doi: 10.4049/jimmunol.178.10.6200. [DOI] [PubMed] [Google Scholar]
- Dal Porto JM, Haberman AM, Kelsoe G, Shlomchik MJ. Very low affinity B cells form germinal centers, become memory B cells, and participate in secondary immune responses when higher affinity competition is reduced. J. Exp. Med. 2002;195:1215–1221. doi: 10.1084/jem.20011550. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dougan G, John V, Palmer S, Mastroeni P. Immunity to salmonellosis. Immunol. Rev. 2011;240:196–210. doi: 10.1111/j.1600-065X.2010.00999.x. [DOI] [PubMed] [Google Scholar]
- Gil-Cruz C, Bobat S, Marshall JL, Kingsley RA, Ross EA, Henderson IR, Leyton DL, Coughlan RE, Khan M, Jensen KT, et al. The porin OmpD from nontyphoidal Salmonella is a key target for a protective B1b cell antibody response. Proc. Natl. Acad. Sci. USA. 2009;106:9803–9808. doi: 10.1073/pnas.0812431106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guan Y, Sajadi MM, Kamin-Lewis R, Fouts TR, Dimitrov A, Zhang Z, Redfield RR, DeVico AL, Gallo RC, Lewis GK. Discordant memory B cell and circulating anti-Env antibody responses in HIV-1 infection. Proc. Natl. Acad. Sci. USA. 2009;106:3952–3957. doi: 10.1073/pnas.0813392106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hastey CJ, Elsner RA, Barthold SW, Baumgarth N. Delays and diversions mark the development of B cell responses to Borrelia burgdorferi infection. J. Immunol. 2012;188:5612–5622. doi: 10.4049/jimmunol.1103735. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hill MO. Diversity and evenness: a unifying notation and its consequences. Ecology. 1973;54:427–432. [Google Scholar]
- Hoiseth SK, Stocker BA. Aromatic-dependent Salmonella typhimurium are non-virulent and effective as live vaccines. Nature. 1981;291:238–239. doi: 10.1038/291238a0. [DOI] [PubMed] [Google Scholar]
- Jacob J, Kelsoe G, Rajewsky K, Weiss U. Intraclonal generation of antibody mutants in germinal centres. Nature. 1991;354:389–392. doi: 10.1038/354389a0. [DOI] [PubMed] [Google Scholar]
- Kim HJ, Krenn V, Steinhauser G, Berek C. Plasma cell development in synovial germinal centers in patients with rheumatoid and reactive arthritis. J. Immunol. 1999;162:3053–3062. [PubMed] [Google Scholar]
- Kleinstein SH, Louzoun Y, Shlomchik MJ. Estimating hyper-mutation rates from clonal tree data. J. Immunol. 2003;171:4639–4649. doi: 10.4049/jimmunol.171.9.4639. [DOI] [PubMed] [Google Scholar]
- Koga T, Nishihara T, Fujiwara T, Nisizawa T, Okahashi N, Noguchi T, Hamada S. Biochemical and immunobiological properties of lipopolysaccharide (LPS) from Bacteroides gingivalis and comparison with LPS from Escherichia coli. Infect. Immun. 1985;47:638–647. doi: 10.1128/iai.47.3.638-647.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kong W, Brovold M, Koeneman BA, Clark-Curtiss J, Curtiss R., 3rd Turning self-destructing Salmonella into a universal DNA vaccine delivery platform. Proc. Natl. Acad. Sci. USA. 2012;109:19414–19419. doi: 10.1073/pnas.1217554109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee SJ, Liang L, Juarez S, Nanton MR, Gondwe EN, Msefula CL, Kayala MA, Necchi F, Heath JN, Hart P, et al. Identification of a common immune signature in murine and human systemic Salmonellosis. Proc. Natl. Acad. Sci. USA. 2012;109:4998–5003. doi: 10.1073/pnas.1111413109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marshall JL, Zhang Y, Pallan L, Hsu MC, Khan M, Cunningham AF, MacLennan IC, Toellner KM. Early B blasts acquire a capacity for Ig class switch recombination that is lost as they become plasmablasts. Eur. J. Immunol. 2011;41:3506–3512. doi: 10.1002/eji.201141762. [DOI] [PubMed] [Google Scholar]
- McGregor AC, Waddington CS, Pollard AJ. Prospects for prevention of Salmonella infection in children through vaccination. Curr. Opin. Infect. Dis. 2013;26:254–262. doi: 10.1097/QCO.0b013e32835fb829. [DOI] [PubMed] [Google Scholar]
- Moyron-Quiroz JE, Rangel-Moreno J, Kusser K, Hartson L, Sprague F, Goodrich S, Woodland DL, Lund FE, Randall TD. Role of inducible bronchus associated lymphoid tissue (iBALT) in respiratory immunity. Nat. Med. 2004;10:927–934. doi: 10.1038/nm1091. [DOI] [PubMed] [Google Scholar]
- Muramatsu M, Kinoshita K, Fagarasan S, Yamada S, Shinkai Y, Honjo T. Class switch recombination and hypermutation require activation-induced cytidine deaminase (AID), a potential RNA editing enzyme. Cell. 2000;102:553–563. doi: 10.1016/s0092-8674(00)00078-7. [DOI] [PubMed] [Google Scholar]
- Nanton MR, Way SS, Shlomchik MJ, McSorley SJ. Cutting edge: B cells are essential for protective immunity against Salmonella independent of antibody secretion. J. Immunol. 2012;189:5503–5507. doi: 10.4049/jimmunol.1201413. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Neves P, Lampropoulou V, Calderon-Gomez E, Roch T, Stervbo U, Shen P, Kühl AA, Loddenkemper C, Haury M, Nedospasov SA, et al. Signaling via the MyD88 adaptor protein in B cells suppresses protective immunity during Salmonella typhimurium infection. Immunity. 2010;33:777–790. doi: 10.1016/j.immuni.2010.10.016. [DOI] [PubMed] [Google Scholar]
- Odegard JM, Marks BR, DiPlacido LD, Poholek AC, Kono DH, Dong C, Flavell RA, Craft J. ICOS-dependent extrafollicular helper T cells elicit IgG production via IL-21 in systemic autoimmunity. J. Exp. Med. 2008;205:2873–2886. doi: 10.1084/jem.20080840. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Onodera T, Takahashi Y, Yokoi Y, Ato M, Kodama Y, Hachimura S, Kurosaki T, Kobayashi K. Memory B cells in the lung participate in protective humoral immune responses to pulmonary influenza virus reinfection. Proc. Natl. Acad. Sci. USA. 2012;109:2485–2490. doi: 10.1073/pnas.1115369109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ortiz V, Isibasi A, García-Ortigoza E, Kumate J. Immunoblot detection of class-specific humoral immune response to outer membrane proteins isolated from Salmonella typhi in humans with typhoid fever. J. Clin. Microbiol. 1989;27:1640–1645. doi: 10.1128/jcm.27.7.1640-1645.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Racine R, Jones DD, Chatterjee M, McLaughlin M, Macnamara KC, Winslow GM. Impaired germinal center responses and suppression of local IgG production during intracellular bacterial infection. J. Immunol. 2010;184:5085–5093. doi: 10.4049/jimmunol.0902710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rothaeusler K, Baumgarth N. B-cell fate decisions following influenza virus infection. Eur. J. Immunol. 2010;40:366–377. doi: 10.1002/eji.200939798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scheel T, Gursche A, Zacher J, Häupl T, Berek C. V-region gene analysis of locally defined synovial B and plasma cells reveals selected B cell expansion and accumulation of plasma cell clones in rheumatoid arthritis. Arthritis Rheum. 2011;63:63–72. doi: 10.1002/art.27767. [DOI] [PubMed] [Google Scholar]
- Schröder AE, Greiner A, Seyfert C, Berek C. Differentiation of B cells in the nonlymphoid tissue of the synovial membrane of patients with rheumatoid arthritis. Proc. Natl. Acad. Sci. USA. 1996;93:221–225. doi: 10.1073/pnas.93.1.221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shlomchik MJ, Weisel F. Germinal center selection and the development of memory B and plasma cells. Immunol. Rev. 2012;247:52–63. doi: 10.1111/j.1600-065X.2012.01124.x. [DOI] [PubMed] [Google Scholar]
- Shugay M, Britanova OV, Merzlyak EM, Turchaninova MA, Mamedov IZ, Tuganbaev TR, Bolotin DA, Staroverov DB, Putintseva EV, Plevova K, et al. Towards error-free profiling of immune repertoires. Nat. Methods. 2014;11:653–655. doi: 10.1038/nmeth.2960. [DOI] [PubMed] [Google Scholar]
- Singh SP, Upshaw Y, Abdullah T, Singh SR, Klebba PE. Structural relatedness of enteric bacterial porins assessed with monoclonal antibodies to Salmonella typhimurium OmpD and OmpC. J. Bacteriol. 1992;174:1965–1973. doi: 10.1128/jb.174.6.1965-1973.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stott DI, Hiepe F, Hummel M, Steinhauser G, Berek C. Antigen-driven clonal proliferation of B cells within the target tissue of an autoimmune disease. The salivary glands of patients with Sjögren’s syndrome. J. Clin. Invest. 1998;102:938–946. doi: 10.1172/JCI3234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tiller T, Busse CE, Wardemann H. Cloning and expression of murine Ig genes from single B cells. J. Immunol. Methods. 2009;350:183–193. doi: 10.1016/j.jim.2009.08.009. [DOI] [PubMed] [Google Scholar]
- Toellner KM, Gulbranson-Judge A, Taylor DR, Sze DM, MacLennan IC. Immunoglobulin switch transcript production in vivo related to the site and time of antigen-specific B cell activation. J. Exp. Med. 1996;183:2303–2312. doi: 10.1084/jem.183.5.2303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tomayko MM, Steinel NC, Anderson SM, Shlomchik MJ. Cutting edge: Hierarchy of maturity of murine memory B cell subsets. J. Immunol. 2010;185:7146–7150. doi: 10.4049/jimmunol.1002163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vander Heiden JA, Yaari G, Uduman M, Stern JN, O’Connor KC, Hafler DA, Vigneault F, Kleinstein SH. pRESTO: a toolkit for processing high-throughput sequencing raw reads of lymphocyte receptor repertoires. Bioinformatics. 2014;30:1930–1932. doi: 10.1093/bioinformatics/btu138. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vollmers C, Sit RV, Weinstein JA, Dekker CL, Quake SR. Genetic measurement of memory B-cell recall using antibody repertoire sequencing. Proc. Natl. Acad. Sci. USA. 2013;110:13463–13468. doi: 10.1073/pnas.1312146110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wardemann H, Yurasov S, Schaefer A, Young JW, Meffre E, Nussenzweig MC. Predominant autoantibody production by early human B cell precursors. Science. 2003;301:1374–1377. doi: 10.1126/science.1086907. [DOI] [PubMed] [Google Scholar]
- William J, Euler C, Christensen S, Shlomchik MJ. Evolution of autoantibody responses via somatic hypermutation outside of germinal centers. Science. 2002;297:2066–2070. doi: 10.1126/science.1073924. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.