Abstract
Exosomes are nanoscale extracellular vesicles that are shed from different cells in the body. Exosomes encapsulate several biomolecules including lipids, proteins, and nucleic acids, and can therefore play a key role in cellular communication. These vesicles can be isolated from different body fluids and their small sizes make them attractive in various biomedical applications. Here, we review state‐of‐the art approaches in exosome isolation and purification, and describe their potential use in cancer vaccines, drug delivery, and diagnostics.
Keywords: drug delivery, immunotherapies, nanobiology, patient‐targeted therapies
1. Introduction
Communication between cells is a critical process that occurs in all organisms. This sharing of information is facilitated either horizontal gene transfer by viruses1 and bacteriophages2 or by the secretion of soluble factors.3 Cells secrete extracellular vesicles (EVs), namely, apoptotic bodies, microvesicles, and exosomes,4 which can have significant impact on the local microenvironment as well as on distant tissues in the body.5 Apoptotic bodies are the largest among EVs, with sizes ranging from 50 to 5,000 nm, and contain DNA, RNA, and histone proteins. Apoptotic bodies are eventually removed by macrophages.6, 7 Microvesicles (50–1,000 nm) are generally smaller than apoptotic bodies and are also known as shedding vesicles or exovesicles. Exosomes (30–120 nm; Figure 1),8 are the smallest among secreted vesicles, and consist of a lipid bilayer membrane that surrounds cytosol and other contents. Exosomes derived from human embryonic kidney cells have been observed to shrink in size when stored under 4°C temperatures.9 Exosomes typically do not contain cellular organelles but can carry different molecular constituents, including proteins and nucleic acids, from the cell of origin.10 Proteins including those from the tetraspanin family (CD9, CD63, CD81, and CD82), ESCRT complex (TSG101, Alix)11 and heat shock proteins (Hsp 60, Hsp70, Hsp90) are known to be found in exosomes; the composition of these proteins differs based on the cell or tissue of origin.12, 13
Figure 1.

Schematic showing the release of exosomes and other vesicles from cells. Adapted from Ref. 8
Vesicles secreted by cells can meet one of the following fates: (a) internalization by other cells in the immediate proximity, (b) internalization by cells at a significant distance away from the cell of origin, and (c) removal by distant tissues following entry into systemic circulation.14 Exosomes carry proteins and nucleic acids from their cell of origin,15 and these contents can be delivered to a different recipient cell leading to intercellular communication that can impact different physiological process.5, 16, 17 Western blotting18, 19, 20, 21, 22, 23, 24 and fluorescence‐activated cell sorting analysis22, 25, 26, 27 of beads coated with exosomes have helped determine the presence of known cellular proteins in exosome preparations from different cells. In addition, mass spectrometry can be employed to identify unknown cellular proteins present in exosomes.27, 28, 29, 30 Exosome‐facilitated messages can regulate cellular growth, division, and apoptosis;31, 32 changes in gene expression in recipient cells can be induced following delivery of multiple miRNAs by exosomes.33 For example, exosomes derived from three different melanoma cell lines, while being morphologically very similar, had different effects on T‐cell proliferation/growth suppression.34 In this review, we first discuss cell sources that have been explored for obtaining exosomes and isolation methods that have been employed for obtaining enriched populations of these vesicles. We also discuss the use of exosomes as therapeutic carriers, disease biomarkers, and vaccines.
2. Cell sources for exosomes
Cell lines including bEND3 and EL‐4 are predominantly used for isolation of exosomes used in drug delivery due to the ease of their availability, the ability to scale up and the ability to generate exosomes of similar quality.35, 36 Mesenchymal stem cells or hESC‐MSCs are also excellent sources for isolation of exosomes,37 and it has been observed that exosomes isolated from these are tolerated well by the immune system thereby making them an excellent choice for drug delivery.15, 38, 39 These cell lines can also be used to isolate exosomes for the purpose of developing anti‐cancer vaccines. It is possible that primary cells isolated from patients/mice for developing these exosome‐based anti‐cancer vaccines might demonstrate a batch to batch variation in strength of the immune response.20, 40 The use of exosomes for identifying biomarker levels mandates that patient‐derived cells/body fluids are used. The presence/absence of disease can be determined by comparing exosomal biomarker levels of the patient with that of healthy controls.41, 42, 43, 44, 45
3. Isolation and purification of exosomes
Several methods have been investigated for the isolation and purification of exosomes from biological fluids.46, 47 Centrifugation, filtration, immunological separation, microfluidic isolation,48 and size‐exclusion chromatography can be effectively applied in both laboratory research and clinical medicine. Differential ultracentrifugation remains one of the most common techniques for exosome isolation49, 50, 51 and consists of several centrifugation steps that remove cells, large vesicles, and debris at lower centrifugation speeds. The pellets are discarded while the supernatant is subjected to higher centrifugation speeds in order to obtain exosomes as a pellet. Differential ultracentrifugation is commonly employed for the isolation of exosomes but the efficiency of the method is lower when plasma and serum are used due to higher viscosities of these fluids.24, 47, 52 Density gradient centrifugation combines differential ultracentrifugation with a sucrose density gradient. This method is mainly used for separating exosomes from nonvesicular particles including proteins and protein/RNA aggregates. The method allows separation of low‐density exosomes from other contaminants but is highly sensitive to the centrifugation time.24, 52, 53
Specific binding of antibodies to receptors present on the surface of exosome have been explored for isolating these vesicles from mixtures54, 55; in many cases isolated exosomes are subsequently analyzed for DNA or total RNA.56, 57, 58, 59, 60 As an application of this approach, antibodies are displayed onto magnetic beads to facilitate the specific binding and isolation of exosomes.61, 62 The advantage of this immunoaffinity technique is the presence of various tetraspanins on exosomes isolated from different cell types which can be targeted using antibodies. The immunoaffinity technique employed was reported to be more effective at isolating exosomes compated to ultracentrifugation and density gradient methods.53, 63 This method can be extremely effective for small‐scale applications with low volume samples but the costs associated with scaling up this approach may be prohibitive for large scale isolation procedures.
Ultrafiltration has been investigated for the separation of exosomes from proteins,39, 64, 65 although the efficacy of this method has not been fully established for clinical samples. A porous membrane can be used for trapping exosomes resulting in their isolation from cell culture media. The filtration membrane helps concentrate the exosomal population. Recovery of exosomes from the membranes is facilitated by using ethanol or NaOH, which is typically followed by rinsing with phosphate‐buffered saline.24 However, application of large external force in this approach can result in deformation or damage of exosomes.
Polymer‐facilitated precipitation, most commonly using polyethylene glycol, is also employed for recovering exosomes from mixtures (Figure 2).66, 67 The main advantage of this method is the use of neutral pH. Although, there are no adverse effects on the isolated exosomes, contamination of exosomes with other nonexosomal materials is a significant drawback of this method.68 In addition, the presence of the polymer may interfere with downstream analyses and/or usage.69 Polymer‐facilitated precipitation can be significantly improved by using methods for removing the polymer used in the operation. Size‐exclusion chromatography is commonly used to separate macromolecules based on size,70 and has also been investigated for exosome isolation (Figure 3).71, 72 Exosomes isolated using this method are subjected to minimal shear force, resulting in relatively low damage to the structure of these vesicles; however, deformation of larger vesicles has been reported.73
Figure 2.
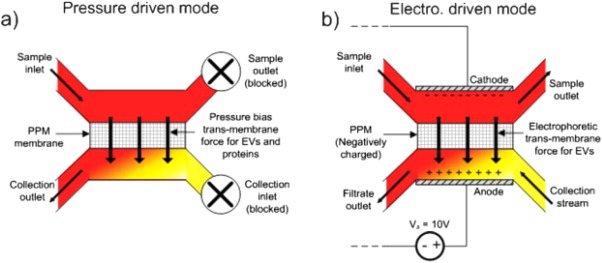
Schematic of the polymer‐based precipitation method used for the isolation of exosomes. Adapted from Ref. 54
Figure 3.

Schematic of the size‐exclusion chromatography approach employed for the isolation and purification of exosomes. Adapted from Ref. 58
Exosomes isolated using ultracentrifugation—for example, using the ExoQuick approach—can result in increased yields compared to other methods, while also maintaining the quality of the exosomes isolated.24 The swift isolation and higher exosome recovery enables analysis of protein expression in the recovered exosomes. Ultracentrifugation with density gradient centrifugation can improve the efficacy of the exosome purification without damaging the morphology. Variations in sizes of isolated exosomes have also been observed between fresh samples and those stored in DMSO and freezing also resulted in degradation of exosomal RNA over time.74
4. Applications of exosomes
4.1. Drug and nucleic acid delivery
The small sizes of exosomes make them attractive as vehicles in drug and nucleic acid delivery, although detailed molecular characterization of exosomes is necessary before adopting them in widespread applications. Yang et al. hypothesized that exosomes derived from brain cells displayed brain‐specific surface proteins which allowed them to pass through the blood‐brain barrier (BBB) and deliver drugs across this barrier.35 Exosomes were isolated from four different cell lines including brain endothelial cell line bEND.3, human glioblastoma cell line U‐87 MG, human brain neuroectodermal cell line PFSK‐1, and human brain glioblastoma A‐172 cells. Rhodamine‐123 (2 mg/ml), doxorubicin, or paclitaxel were independently loaded into different exosomes by mixing followed by incubation for 2 hr. Doxorubicin‐loaded exosomes demonstrated higher efficacies for inducing death in U87‐MG glioblastoma cells compared to paclitaxel‐loaded exosomes. Exosomes isolated from bEND.3 cells were able to deliver the rhodamine‐123 dye across the BBB following delivery via the cardinal vein of zebrafish embryos. Exosomes from bEND.3 cells were able to deliver drugs to a U‐87 MG tumor grown in the brain of zebrafish and were also observed to inhibit VEGF (vascular endothelial growth factor) levels in vivo (Figure 4).35 This, in turn, was able to induce a significant decrease in the tumor size, compared to treatments with the free drug. Intranasal delivery of curcumin or JSI‐124 (cucurbitacin I) inhibitor‐loaded exosomes was investigated as a potential therapeutic approach for brain inflammatory diseases.75 Three different mouse models exhibiting lipopolysaccharide (LPS)‐induced brain inflammation, autoimmune encephalatis, or the GL26 brain tumor model which exhibits inflammation due to infiltration of immune cells including macrophages and T‐cells, were used in the study.76 Exosomes were loaded by mixing curcumin with EL‐4 (mouse lymphoma cell)‐derived exosomes at a temperature of 22°C after which the loaded exosomes were separated using sucrose gradient centrifugation. Intranasally delivered exosomes were taken up by microglial cells, which are key mediators in neuro‐inflammatory diseases; delivery of curcumin‐loaded exosomes resulted in a reduction in activated microglial cells in both encephelatis and LPS‐induced brain inflammation models. Intranasal delivery of exosomes loaded with the STAT3 inhibitor JSI‐124 resulted in increased survival of mice with GL26 brain tumors. This study suggests that intranasal delivery of exosome‐encapsulated drugs could lead to a noninvasive approach for direct drug delivery to the CNS.75
Figure 4.
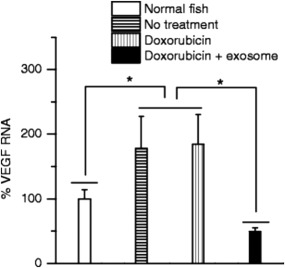
In vivo delivery of doxorubicin (Dox)‐loaded exosomes across the blood brain barrier in zebrafish model. Significant inhibition of VEGF was observed in Dox‐loaded exosomes compared to untreated controls and those treated with unencapsulated doxorubicin. Adapted from Ref. 35
Exosomes isolated from RAW264.7 macrophages were loaded with catalase as a potential therapeutic approach for Parkinson's disease; catalase was loaded in order to potentially degrade reactive oxygen species in order to protect from inflammation.77 Exosomes were loaded using four different methods—incubation with saponin, sonication, freeze‐thaw, and extrusion. Sonication‐loaded exosomes resulted in the highest uptake of catalase in PC12 neuronal cells in vitro as observed using fluorescence spectroscopy and confocal microscopy. Sonication was carried out using 20% power at 500V, 2kHz for 6 cycles pulsed for 4 s on and paused for 2 s. Extrusion was also employed for loading, and was performed by mixing catalase with exosome solution followed by 10 rounds of extrusion through a Avanti Lipids extruder.78 Catalase‐loaded exosomes, named exoCAT, protected neuronal cells from oxidative stress in vivo following intracranial injection into C57BL/6 mice. Biodistribution of exoCAT following intracranial injections indicated localization primarily in neuronal and microglial cells but also in astrocytes and endothelial cells.
Exosomes extracted from cow milk were employed for the delivery of therapeutic molecules against lung and breast cancer.79 Exosomes loaded with withaferin‐A were injected intraperitoneally into female athymic nude mice subcutaneously injected with A549 cells. A tumor inhibitory effect was observed with withaferin‐A loaded exosomes at doses lower than those observed with the unencapsulated drug.
Exosomes isolated from dendritic cells from the bone marrow of C57BL/6 mice were employed for delivering small interfering RNA (siRNA) to the brain. A Rabies viral glycoprotein (RVG) peptide (single letter amino acid sequence: YTIWMPENPRPGTPCDIFTNSRGKRASNG) was displayed onto the exosomal surface for targeting the acetylcholine receptor in the brain. Electroporation at 400 V and 125 μF was used to load these exosomes with siRNA against GAPDH, and the loaded vesicles were employed to investigate the knock down of GAPDH gene in C2C12 (murine muscle) and Neuro2A (neuronal cells). Gene knockdown efficacy using exosomes loaded with GAPDH siRNA was similar to that observed using lipofectamine. Exosomes were also well tolerated and did not induce strong immune responses in C57BL/6 and BALB/C mice. RVG exosomes were also able to deliver BACE1 siRNA across the BBB.80
Exosomes derived from human bone marrow mesenchymal stem cells were investigated for delivering functional anti‐miR‐9 to glioblastoma multiforme cells81; communication between MSCs and brain glioblastoma cells can be mediated by exosomes.82 Flow cytometry and quantitative PCR (qPCR) indicated that MSC‐derived exosomes, loaded with anti‐mir9, led to the reduction of MDR1 expression in T98G and U87 glioblastoma cells. Delivery of anti‐mir9 increased the sensitivity of these cells towards temozolomide, resulting in increased cancer cell death.81
Strategies for loading molecular cargo in exosomes and related efficacies differ based on the chemistry of the loaded molecule. The most common method for loading small‐molecule drugs involves mixing the drug solution with a suspension of exosomes and incubating them at 25–37°C. The drug loading efficiency is typically determined using liquid chromatrography.83, 84 Loading of DNA onto exosomes using electroporation methods is likely restricted by the size of the exosomes with only large exosomes being capable of carrying large linear/plasmid DNA85 The loading efficiencies achieved using electroporation varied between 15 and 30% of those achieved with small molecule drugs and siRNA,86, 87 Momen‐Heravi et al. were able to achieve up to 55% loading efficiency using electroporation for miRNA molecules.88 Smyth et al. achieved doxorubicin loading efficiencies of 5% by weight of exosomes using the mixing technique.89 Similarly, Sun et al. observed a binding capacity of 2.9 g of curcumin for every gram of exosome using the mixing technique followed by sucrose density gradient centrifugation.36 Kim et al. compared different techniques that is, mixing incubation, electroporation, and sonication for loading the drug paclitaxel into exosomes and observed 1.5, 5, and 29% loading efficiencies, respectively.90 In comparison, Yang et al. were able achieve 5% loading efficiency of siRNA in cationic lipopolymers with an encapsulation efficiency of 70%.91 and Cao et al. obtained 33% loading efficiency in calcium phosphate nanoparticles.92
Taken together, these studies indicate the potential utility of exosomes in drug and nucleic acid delivery. The choice of cells from which exosomes are isolated, yield of exosome vesicles, cargo loading procedures, selection of targeting biomolecules (e.g., peptides) on the surface, biodistribution, and immune response are key factors for consideration for exosome‐mediated delivery in future translational applications. Loading of small‐molecule drugs can be efficient, although there is room for improvement in case of loading DNA/siRNA.
5. Biomarkers
Exosomes play a vital role in cell‐cell communication by directly engaging with surface ligands and/or by transferring their contents between cells.93, 94, 95, 96 Presence of exosomal RNA was implicated as evidence for horizontal transfer of genetic information between various cell types.97, 98, 99 Exosomes are thought to also transfer cellular mRNA as well as microRNA which indicates that tumor exosomes are functional and could suppress mRNA that codes for signal transduction components within T‐cells.100, 101 The RNA population in tumor‐secreted exosomes includes microRNA and it is possible that this exosomal microRNA reflects the parental tumor signature. As a result, microRNA expression profiling can be useful as a diagnostic tool in diseases, including some cancers which lack definitive molecular biomarkers.
Figure 5.

miRNA biomarker expression levels in tumor and exosome‐derived samples showing comparable levels of miRNA between the two sample types. Adapted from Ref. 104
Exosome levels can be slightly elevated in benign tumors and highly elevated in cancerous patients as compared to normal controls; exosomes were isolated from the sera of patients/control subjects using magnetic activated cell sorting. Approximately, 175 different miRNA were found to be similar between tumor cells and exosomes. The up‐regulated miRNA profile from exosomes also matched up‐regulated miRNA profiles in ovarian cancer patients at different stages of the disease. However, this approach was not able to distinguish between different stages of cancer.102, 103, 104, 105 Expression of miRNA in circulating tumor‐derived exosomes derived from lung adenocarcinoma patients were similar to that seen in primary tumors, indicating the potential use of these tumor‐derived exosomes as biomarkers. In the future, it may be possible to analyze miRNA from circulating exosomes obviating the need for, or at least complementing, tumor biopsy samples10 in applications related to detecting disease, monitoring response to therapy, and investigating cancer recurrence.
The mRNA expression of two distinct biomarkers, PCA‐3 and TMPRSS2, is found in exosomes and can be used to provide a direct link to the incidence of prostate cancer.106 Proteins extracted from urinary exosomes can be of potential use in diagnostics of urinary tract diseases,45 and prostate and bladder cancers. Eight urinary exosomal proteins were identified as biomarkers when patients with prostate, bladder cancer cells, and healthy cells were compared.107 It has been reported in many cases that exosomal miRNA are prospective biomarkers for renal fibrosis and cardiovascular diseases.108, 109, 110
Potential advantages of using exosomes as biomarkers includes the ability to reduce the use of invasive surgery for monitoring disease. Exosomes can be isolated from serum samples and have been successfully used for biomarker detection in ovarian, lung and pancreatic cancers.111, 112, 113 Exosomal miRNA has also been found to be useful in distinguishing between pancreatic carcinomas from benign pancreatic tumors and chronic pancreatitis113 and in detection of miRNA (miR‐34A) responsible for conferring drug resistance to prostate cancer.41 Exosomal miRNA appear to be stable and can undergo multiple freeze‐thaw cycles, variations in pH and heating without undergoing degradation or loss of expression levels; the miRNA profile is also highly specific to cancer/tumor tissue.113 However, a key drawback of using exosomes as biomarkers markers is that they cannot determine the severity of the disease because the miRNA levels can be identical during different stages of the disease as shown by Taylor et al. for ovarian cancer and Rabinowitz et al. in the case of lung cancer.104, 112 The potential of using exosomal markers for clinical diagnostics needs to be further investigated in depth because various exosomal components including lipids, proteins and miRNAs can be promising in disease diagnostics.
Figure 6.

miRNA profile comparison between tumor and circulating exosomes. Adapted from Ref. 104
6. Anticancer vaccines
Exosomes derived from specific sites in the body can be promising candidates for anti‐cancer vaccines since they can present antigens against that specific type of cancer. Exosomes can be isolated from three sites of origin ascites (ascite‐derived exosomes or AEX), dendritic cells (dendritic cell‐derived exosomes or DEX), and tumors (tumor‐derived exosomes or TEX).114, 115, 116, 117 The antigens in these exosomes can elicit an immune response via MHC class I or class II molecules on CD8+ or CD4+ T cells. These exosomal MHC class I/II molecules on exosomes are likely used for communication with specific cell types.118, 119 Melanoma antigens in DEXs were used to prime cytotoxic T lymphocytes in order to elicit an immune response through the Mart1 specific pathway in patients with high‐grade melanoma.21, 120 In a different study, exosomes were isolated from four different pancreatic cancer cell lines, SOJ‐6, BxPC‐3, MiaPaCa‐2, and Panc‐1. Exosomes derived from SOJ‐6 and BxPC‐3 cells were more effective at reducing growth human pancreatic adenocarcinoma cells compared to those derived from MiaPaCa‐2 and Panc‐1 cells; regulation of Hes‐1 protein via downregulation of the Notch‐1 signaling pathway and activation of mitochondria‐dependent apoptosis pathway played a key role in the cell ablation efficacy.121
Dendritic cell‐derived exosomes were shown to elicit Natural Killer (NK) cell responses following intradermal injection in C57BL/6 mice. Exosomes isolated from normal volunteers were primed with melanoma associated antigens MAGE3.A1 and MAGE3.DP04, and were used as vaccines in stage IIIb and stage IV melanoma patients. The treatment resulted in a regression of the cancer; while no changes in cytotoxic T‐lymphocyte levels were observed, the level of NK cell increased with treatment. However, the NK cells obtained from patients that responded to the DEX treatment were more effective at killing K562 cells in culture compared to those obtained from nonresponders.115, 122
These studies indicate potential applications of exosomes in cancer immunotherapy. However, exosomes generally need mature DCs to elicit a T‐cell response, since antigens present on exosomes need to be taken up by the dendritic cells before they can activate T‐lymphocytes. This can be avoided by the use of adjuvants in some cases, which can allow exosomes to directly prime T‐lymphocytes. In addition, the reliance on an intact immune system can restrict their use in immunocompromised or immunosuppressed patients. However, this approach can elicit an immune response in already developed cancer which can induce regression of the tumor.122 This approach may also find less resistance from the suppressive nature of the tumor microenvironment,116, 117, 122, 123, 124, 125, 126 which can be a significant advantage.
Figure 7.

Interactions of dendritic cell‐derived exosomes and tumor‐derived exosomes with inflammatory cells. Reproduced with permission from Creative Commons. Adapted from Ref. 117
7. Future directions
The potential for exosomes in the field of drug delivery is significant owing to their ability to selectively express proteins like tetraspanins which may allow for cell targeting. It will be necessary to obtain highly pure formulations of exosomes with low amounts of protein aggregates and other microvascular particles. Optimization of ultracentrifugation coupled with density gradient centrifugation may offer one route toward obtaining enriched populations of exosomes. Further advancements that enhance the loading of drug/nucleic acid are necessary for effective delivery of these therapeutic molecules. Improvements in analytical methods and advances in biomarker discovery can facilitate the use of exosomes in disease detection.
8. Conclusions
Exosomes are starting to gather attention in cancer therapeutics and diagnostics, with several applications in drug delivery, tumor immunotherapy, and diagnostic biomarkers. Their unique strengths include enhanced passive targeting due to small size, indigenous nature, and the ability to cross biological barriers. However, the cumbersome nature of the methods required for isolation/purification, inability to distinguish between different cancer stages, and incomplete understanding of their impact on the immune system are some of the current limitations with this technology. It is anticipated that sophisticated engineering and detailed clinical studies that address these limitations will lead to the translation of exosome‐based technologies in the future.
Funding information National Science Foundation, Grant/Award Numbers: 1404084 and 1403860; Arizona Biomedical Research Commission, Grant/Award Number: ADHS14‐0829981; National Institutes of Health, Grant/Award Number: 1R01GM093229‐01A1
Literature cited
- 1. Walther W, Stein U. Viral vectors for gene transfer. Drugs. 2000;60 (2):249–271. [DOI] [PubMed] [Google Scholar]
- 2. Ochman H, Lawrence JG, Groisman EA. Lateral gene transfer and the nature of bacterial innovation. Nature. 2000;405(6784):299–304. [DOI] [PubMed] [Google Scholar]
- 3. El Andaloussi S, Mager I, Breakefield XO, Wood MJA. Extracellular vesicles: biology and emerging therapeutic opportunities. Nat Rev Drug Discov. 2013;12(5):347–357. [DOI] [PubMed] [Google Scholar]
- 4. Kooijmans SAA, Vader P, van Dommelen SM, van Solinge WW, Schiffelers RM. Exosome mimetics: a novel class of drug delivery systems. Int J Nanomedicine. 2012;7:1525–1541. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Lee Y, EL Andaloussi S, Wood MJA. Exosomes and microvesicles: extracellular vesicles for genetic information transfer and gene therapy. Hum Mol Genet. 2012;21(R1):R125–R134. [DOI] [PubMed] [Google Scholar]
- 6. Schiller M, Bekeredjian‐Ding I, Heyder P, Blank N, Ho AD, Lorenz HM. Autoantigens are translocated into small apoptotic bodies during early stages of apoptosis. Cell Death Differ. 2008;15(1):183–191. [DOI] [PubMed] [Google Scholar]
- 7. Simpson RJ, Jensen SS, Lim JWE. Proteomic profiling of exosomes: current perspectives. Proteomics. 2008;8(19):4083–4099. [DOI] [PubMed] [Google Scholar]
- 8. Lin J, Li J, Huang B, et al. Exosomes: novel biomarkers for clinical diagnosis. Sci World J. 2015;2015:8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Sokolova V, Ludwig A‐K, Hornung S, et al. Characterisation of exosomes derived from human cells by nanoparticle tracking analysis and scanning electron microscopy. Colloids Surf B: Biointerfaces. 2011;87(1):146–150. [DOI] [PubMed] [Google Scholar]
- 10. van den Boorn JG, Daßler J, Coch C, Schlee M, Hartmann G. Exosomes as nucleic acid nanocarriers. Adv Drug Deliv Rev. 2013;65(3):331–335. [DOI] [PubMed] [Google Scholar]
- 11. Simons M, Raposo G. Exosomes – vesicular carriers for intercellular communication. Curr Opin Cell Biol. 2009;21(4):575–581. [DOI] [PubMed] [Google Scholar]
- 12. Batista BS, Eng WS, Pilobello KT, Hendricks‐Muñoz KD, Mahal LK. Identification of a conserved glycan signature for microvesicles. J Proteome Res. 2011;10(10):4624–4633. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Kalani A, Tyagi A, Tyagi N. Exosomes: mediators of neurodegeneration, neuroprotection and therapeutics. Mol Neurobiol. 2014;49(1):590–600. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Pant S, Hilton H, Burczynski ME. The multifaceted exosome: biogenesis, role in normal and aberrant cellular function, and frontiers for pharmacological and biomarker opportunities. Biochem Pharmacol. 2012;83(11):1484–1494. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Lai RC, Yeo RWY, Tan KH, Lim SK. Exosomes for drug delivery—a novel application for the mesenchymal stem cell. Biotechnol Adv. 2013;31(5):543–551. [DOI] [PubMed] [Google Scholar]
- 16. van Niel G, Porto‐Carreiro I, Simoes S, Raposo G. Exosomes: a common pathway for a specialized function. J Biochem. 2006;140(1):13–21. [DOI] [PubMed] [Google Scholar]
- 17. Schorey JS, Bhatnagar S. Exosome function: from tumor immunology to pathogen biology. Traffic. 2008;9(6):871–881. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Raposo G, Nijman HW, Stoorvogel W, et al. B lymphocytes secrete antigen‐presenting vesicles. J Exp Med. 1996;183(3):1161–1172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Raposo G, Tenza D, Mecheri S, Peronet R, Bonnerot C, Desaymard C. Accumulation of major histocompatibility complex class II molecules in mast cell secretory granules and their release upon degranulation. Mol Biol Cell. 1997;8(12):2631–2645. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Zitvogel L, Regnault A, Lozier A, et al. Eradication of established murine tumors using a novel cell‐free vaccine: dendritic cell derived exosomes. Nat Med. 1998;4(5):594–600. [DOI] [PubMed] [Google Scholar]
- 21. Wolfers J, Lozier A, Raposo G, et al. Nat Med. 2001;7(3):297–303. [DOI] [PubMed] [Google Scholar]
- 22. Blanchard N, Lankar D, Faure F, et al. TCR activation of human T cells induces the production of exosomes bearing the TCR/CD3/ζ complex. J Immunol. 2002;168(7):3235–3241. [DOI] [PubMed] [Google Scholar]
- 23. Escola J‐M, Kleijmeer MJ, Stoorvogel W, Griffith JM, Yoshie O, Geuze HJ. Selective enrichment of tetraspan proteins on the internal vesicles of multivesicular endosomes and on exosomes secreted by human B‐lymphocytes. J Biol Chem. 1998;273(32):20121–20127. [DOI] [PubMed] [Google Scholar]
- 24. Lobb RJ, Becker M, Wen Wen S, Wong CSF, Wiegmans AP, Leimgruber A, Möller A. Optimized exosome isolation protocol for cell culture supernatant and human plasma. J Extracell Vesicles. 2015;4:27031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Rabesandratana H, Toutant J‐P, Reggio H, Vidal M. Decay‐accelerating factor (CD55) and membrane inhibitor of reactive lysis (CD59) are released within exosomes during in vitro maturation of reticulocytes. Blood. 1998;91(7):2573–2580. [PubMed] [Google Scholar]
- 26. Clayton A, Court J, Navabi H, et al. Analysis of antigen presenting cell derived exosomes, based on immuno‐magnetic isolation and flow cytometry. J Immunol Methods. 2001;247(1‐2):163–174. [DOI] [PubMed] [Google Scholar]
- 27. Théry C, Boussac M, Véron P, Ricciardi‐Castagnoli P, Raposo G, Garin J, Amigorena S. Proteomic analysis of dendritic cell‐derived exosomes: a secreted subcellular compartment distinct from apoptotic vesicles. J Immunol. 2001;166(12):7309–7318. [DOI] [PubMed] [Google Scholar]
- 28. Skokos D, Le Panse S, Villa I, et al. Mast cell‐dependent B and T lymphocyte activation is mediated by the secretion of immunologically active exosomes. J Immunol. 2001;166(2):868–876. [DOI] [PubMed] [Google Scholar]
- 29. Van Niel G, Raposo G, Candalh C, et al. Intestinal epithelial cells secrete exosome‐like vesicles. Gastroenterology. 2001;121(2):337–349. [DOI] [PubMed] [Google Scholar]
- 30. Théry C, Regnault A, Garin J, et al. Molecular characterization of dendritic cell‐derived exosomes. J Cell Biol. 1999;147(3):599–610. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Azmi AS, Bao B, Sarkar FH. Exosomes in cancer development, metastasis and drug resistance: a comprehensive review. Cancer Metastasis Rev. 2013;32:0–10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Braicu C, Tomuleasa C, Monroig P, Cucuianu A, Berindan‐Neagoe I, Calin GA. Exosomes as divine messengers: are they the Hermes of modern molecular oncology[quest]. Cell Death Differ. 2015;22(1):34–45. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Vlassov AV, Magdaleno S, Setterquist R, Conrad R. Exosomes: current knowledge of their composition, biological functions, and diagnostic and therapeutic potentials. Biochim Biophys Acta. 2012;1820(7):940–948. [DOI] [PubMed] [Google Scholar]
- 34. Wu Y. Melanoma exosomes deliver a complex biological payload that upregulates PTPN11 to suppress T lymphocyte function. Pigment Cell Melanoma Res. 2017;30:203–218. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Yang T, Martin P, Fogarty B, et al. Exosome delivered anticancer drugs across the blood‐brain barrier for brain cancer therapy in Danio Rerio. Pharm Res. 2015;32(6):2003–2014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Sun D, Zhuang X, Xiang X, et al. A novel nanoparticle drug delivery system: the anti‐inflammatory activity of curcumin is enhanced when encapsulated in exosomes. Mol Ther. 2010;18(9):1606–1614. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Chen TS, Arslan F, Yin Y, et al. Enabling a robust scalable manufacturing process for therapeutic exosomes through oncogenic immortalization of human ESC‐derived MSCs. J Transl Med. 2011;9:47‐47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Timmers L, Lim SK, Hoefer IE, et al. Human mesenchymal stem cell‐conditioned medium improves cardiac function following myocardial infarction. Stem Cell Res. 2011;6(3):206–214. [DOI] [PubMed] [Google Scholar]
- 39. Arslan F, Lai RC, Smeets MB, et al. Mesenchymal stem cell‐derived exosomes increase ATP levels, decrease oxidative stress and activate PI3K/Akt pathway to enhance myocardial viability and prevent adverse remodeling after myocardial ischemia/reperfusion injury. Stem Cell Res. 2013;10(3):301–312. [DOI] [PubMed] [Google Scholar]
- 40. André F, Chaput N, Schartz NEC, et al. Exosomes as potent cell‐free peptide‐based vaccine. I. Dendritic cell‐derived exosomes transfer functional MHC class I/peptide complexes to dendritic cells. J Immunol. 2004;172(4):2126–2136. [DOI] [PubMed] [Google Scholar]
- 41. Corcoran C, Rani S, O'Driscoll L. miR‐34a is an intracellular and exosomal predictive biomarker for response to docetaxel with clinical relevance to prostate cancer progression. Prostate. 2014;74(13):1320–1334. [DOI] [PubMed] [Google Scholar]
- 42. Chen X, Ba Y, Ma L, et al. Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res. 2008;18:997–1006. [DOI] [PubMed] [Google Scholar]
- 43. Kosaka N, Iguchi H, Ochiya T. Circulating microRNA in body fluid: a new potential biomarker for cancer diagnosis and prognosis. Cancer Sci. 2010;101:2087–2092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Brase JC, Wuttig D, Kuner R, Sültmann H. Serum microRNAs as non‐invasive biomarkers for cancer. Mol Cancer. 2010;9:1–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Zhou H, Cheruvanky A, Hu X, et al. Urinary exosomal transcription factors, a new class of biomarkers for renal disease. Kidney Int. 2008;74(5):613–621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Van Deun J, Mestdagh P, Sormunen R, et al. The impact of disparate isolation methods for extracellular vesicles on downstream RNA profiling. J Extracellular Vesicles. 2014;3:24858. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Momen‐Heravi F, Balaj L, Alian S, et al. Current methods for the isolation of extracellular vesicles. Biol Chem. 2013;394:1253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Chen C, Skog J, Hsu C‐H, et al. Microfluidic isolation and transcriptome analysis of serum microvesicles. Lab Chip. 2010;10(4):505–511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Escrevente C, Keller S, Altevogt P, Costa J. Interaction and uptake of exosomes by ovarian cancer cells. BMC Cancer. 2011;11:108‐108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Jeppesen DK, Nawrocki A, Jensen SG, et al. Quantitative proteomics of fractionated membrane and lumen exosome proteins from isogenic metastatic and nonmetastatic bladder cancer cells reveal differential expression of EMT factors. Proteomics. 2014;14(6):699–712. [DOI] [PubMed] [Google Scholar]
- 51. Østergaard O, Nielsen CT, Iversen LV, Jacobsen S, Tanassi JT, Heegaard NHH. Quantitative proteome profiling of normal human circulating microparticles. J Proteome Res. 2012;11(4):2154–2163. [DOI] [PubMed] [Google Scholar]
- 52. Greening DW, Xu R, Ji H, Tauro BJ, Simpson RJ. A protocol for exosome isolation and characterization: evaluation of ultracentrifugation, density‐gradient separation, and immunoaffinity capture methods In: Posch A, ed. Proteomic Profiling: Methods and Protocols. New York, NY: Springer; 2015:179–209. [DOI] [PubMed] [Google Scholar]
- 53. Théry C, Amigorena S, Raposo G, Clayton A. Isolation and characterization of exosomes from cell culture supernatants and biological fluids In: Bonifacino JS, Harford JB, Lippincott‐Schwartz J, Yamada KM, eds. Current Protocols in Cell Biology. Hoboken, NJ: John Wiley & Sons, Inc; 2001. [DOI] [PubMed] [Google Scholar]
- 54. Purushothaman A, Bandari SK, Liu J, Mobley JA, Brown EE, Sanderson RD. Fibronectin on the surface of myeloma cell‐derived exosomes mediates exosome‐cell interactions. J Biol Chem. 2016;291(4):1652–1663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55. Estelles A, Sperinde J, Roulon T, et al. Exosome nanovesicles displaying G protein‐coupled receptors for drug discovery. Int J Nanomed. 2007;2(4):751–760. [PMC free article] [PubMed] [Google Scholar]
- 56. van Balkom BWM, Eisele AS, Pegtel DM, Bervoets S, Verhaar MC. Quantitative and qualitative analysis of small RNAs in human endothelial cells and exosomes provides insights into localized RNA processing, degradation and sorting. J Extracell Vesicles. 2015;4:26760 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Li M, Zeringer E, Barta T, Schageman J, Cheng A, Vlassov AV. Analysis of the RNA content of the exosomes derived from blood serum and urine and its potential as biomarkers. Philos Trans R Soc B Biol Sci. 2014;369(1652):20130502. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. Guescini M, Genedani S, Stocchi V, Agnati LF. Astrocytes and glioblastoma cells release exosomes carrying mtDNA. J Neural Transm (Vienna). 2009;117(1):1–4. [DOI] [PubMed] [Google Scholar]
- 59. Balaj L, Lessard R, Dai L, et al. Tumour microvesicles contain retrotransposon elements and amplified oncogene sequences. Nat Comms. 2011;2:180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60. Thakur BK, Zhang H, Becker A, et al. Double‐stranded DNA in exosomes: a novel biomarker in cancer detection. Cell Res. 2014;24(6):766–769. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61. Pugholm LH, Revenfeld ALS, Søndergaard EKL, Jørgensen MM. Antibody‐based assays for phenotyping of extracellular vesicles. BioMed Res Int. 2015;2015:524817. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62. Zarovni N, Corrado A, Guazzi P, et al. Integrated isolation and quantitative analysis of exosome shuttled proteins and nucleic acids using immunocapture approaches. Methods. 2015;87:46–58. [DOI] [PubMed] [Google Scholar]
- 63. Mincheva‐Nilsson L, Baranov V, Nagaeva O, Dehlin E. Isolation and characterization of exosomes from cultures of tissue explants and cell lines In: Colgan JE, Bierer BE, Margulies DH, Shevach EM, Strober W, eds. Current Protocols in Immunology. Hoboken, NJ: John Wiley & Sons, Inc; 2001. [DOI] [PubMed] [Google Scholar]
- 64. Nordin JZ, Lee Y, Vader P, et al. Ultrafiltration with size‐exclusion liquid chromatography for high yield isolation of extracellular vesicles preserving intact biophysical and functional properties. Nanomedicine. 2015;11(4):879–883. [DOI] [PubMed] [Google Scholar]
- 65. Gercel‐Taylor C, Atay S, Tullis RH, Kesimer M, Taylor DD. Nanoparticle analysis of circulating cell‐derived vesicles in ovarian cancer patients. Anal Biochem. 2012;428(1):44–53. [DOI] [PubMed] [Google Scholar]
- 66. Rider MA, Hurwitz SN, Meckes DG. ExtraPEG: a polyethylene glycol‐based method for enrichment of extracellular vesicles. Sci Rep. 2016;6:23978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67. Davies RT, Kim J, Jang SC, Choi E‐J, Gho YS, Park J. Microfluidic filtration system to isolate extracellular vesicles from blood. Lab Chip. 2012;12(24):5202–5210. [DOI] [PubMed] [Google Scholar]
- 68. Taylor DD, Zacharias W, Gercel‐Taylor C. Exosome isolation for proteomic analyses and RNA profiling In Simpson JR, Greening WD, eds. Serum/Plasma Proteomics: Methods and Protocols. Totowa, NJ: Humana Press; 2011:235‐246. [DOI] [PubMed] [Google Scholar]
- 69. Taylor DD, Shah S. Methods of isolating extracellular vesicles impact down‐stream analyses of their cargoes. Methods. 2015;87:3–10. [DOI] [PubMed] [Google Scholar]
- 70. Hong P, Koza S, Bouvier ESP. Size‐exclusion chromatography for the analysis of protein biotherapeutics and their aggregates. J Liquid Chromatogr Relat Technol. 2012;35(20):2923–2950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71. Wang Z, Wu H‐j, Fine D, et al. Ciliated micropillars for the microfluidic‐based isolation of nanoscale lipid vesicles. Lab Chip. 2013;13(15):2879–2882. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72. Baranyai T, Herczeg K, Onódi Z, et al. Isolation of exosomes from blood plasma: qualitative and quantitative comparison of ultracentrifugation and size exclusion chromatography methods. PloS One. 2015;10(12):e0145686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73. Witwer KW, Buzás EI, Bemis LT, et al., Standardization of sample collection, isolation and analysis methods in extracellular vesicle research. J Extracell Vesicles. 2013;2:20360. doi:10.3402/jev.v2i0.20360 [DOI] [PMC free article] [PubMed]
- 74. Wu Y, Deng W, Klinke DJ. Exosomes: improved methods to characterize their morphology, RNA content, and surface protein biomarkers. Analyst. 2015;140(19):6631–6642. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75. Zhuang X, Xiang X, Grizzle W, et al. Treatment of brain inflammatory diseases by delivering exosome encapsulated anti‐inflammatory drugs from the nasal region to the brain. Mol Ther. 2011;19(10):1769–1779. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76. Candolfi M, Curtin JF, Stephen Nichols W, et al. Intracranial glioblastoma models in preclinical neuro‐oncology: neuropathological characterization and tumor progression. J Neurooncol. 2007;85(2):133–148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77. Haney MJ, Klyachko NL, Zhao Y, et al. Exosomes as drug delivery vehicles for Parkinson's disease therapy. J Control Release. 2015;207:18–30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78. Haney MJ, Klyachko NL, Zhao Y, et al. Exosomes as drug delivery vehicles for Parkinson's disease therapy. J Controlled Release. 2015;207:18–30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79. Munagala R, Aqil F, Jeyabalan J, Gupta RC. Bovine milk‐derived exosomes for drug delivery. Cancer Lett. 2016;371(1):48–61. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80. Alvarez‐Erviti L, Seow Y, Yin H, Betts C, Lakhal S, Wood MJA. Delivery of siRNA to the mouse brain by systemic injection of targeted exosomes. Nat Biotechnol. 2011;29(4):341–345. [DOI] [PubMed] [Google Scholar]
- 81. Munoz JL, Bliss SA, Greco SJ, Ramkissoon SH, Ligon KL, Rameshwar P. Delivery of functional anti‐miR‐9 by mesenchymal stem cell‐derived exosomes to glioblastoma multiforme cells conferred chemosensitivity. Mol Ther Nucleic Acids. 2013;2:e126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82. Biancone L, Bruno S, Deregibus MC, Tetta C, Camussi G. Therapeutic potential of mesenchymal stem cell‐derived microvesicles. Nephrol Dial Transplant. 2012;27(8):3037–3042. [DOI] [PubMed] [Google Scholar]
- 83. Zhuang X, Xiang X, Grizzle W, et al. Treatment of brain inflammatory diseases by delivering exosome encapsulated anti‐inflammatory drugs from the nasal region to the brain. Mol Ther. 2011;19(10):1769–1779. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84. Jang SC, Kim OY, Yoon CM, et al. Bioinspired exosome‐mimetic nanovesicles for targeted delivery of chemotherapeutics to malignant tumors. ACS Nano. 2013;7(9):7698–7710. [DOI] [PubMed] [Google Scholar]
- 85. Lamichhane TN, Raiker RS, Jay SM. Exogenous DNA loading into extracellular vesicles via electroporation is size‐dependent and enables limited gene delivery. Mol Pharmaceutics. 2015;12(10):3650–3657. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86. Banizs AB, Huang T, Dryden K, Berr SS, Stone JR, Nakamoto RK, Shi W, He J. In vitro evaluation of endothelial exosomes as carriers for small interfering ribonucleic acid delivery. Int J Nanomed. 2014;9:4223–4230. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87. Tian Y, Li S, Song J, et al. A doxorubicin delivery platform using engineered natural membrane vesicle exosomes for targeted tumor therapy. Biomaterials. 2014;35(7):2383–2390. [DOI] [PubMed] [Google Scholar]
- 88. Momen‐Heravi F, Bala S, Bukong T, Szabo G. Exosome‐mediated delivery of functionally active miRNA‐155 inhibitor to macrophages. Nanomedicine. 2014;10(7):1517–1527. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89. Smyth T, Kullberg M, Malik N, Smith‐Jones P, Graner MW, Anchordoquy TJ. Biodistribution and delivery efficiency of unmodified tumor‐derived exosomes. J Controlled Release. 2015;199:145–155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90. Kim MS, Haney MJ, Zhao Y, et al. Development of exosome‐encapsulated paclitaxel to overcome MDR in cancer cells. Nanomedicine. 2016;12(3):655–664. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91. Yang X‐Z, Dou S, Sun T‐M, Mao C‐Q, Wang H‐X, Wang J. Systemic delivery of siRNA with cationic lipid assisted PEG‐PLA nanoparticles for cancer therapy. J Control Release. 2011;156(2):203–211. [DOI] [PubMed] [Google Scholar]
- 92. Cao X, Deng W, Wei Y, et al. Encapsulation of plasmid DNA in calcium phosphate nanoparticles: stem cell uptake and gene transfer efficiency. Int J Nanomed. 2011;6:3335–3349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93. Polgar J, Matuskova J, Wagner DD. The P‐selectin, tissue factor, coagulation triad. J Thromb Haemost. 2005;3(8):1590–1596. [DOI] [PubMed] [Google Scholar]
- 94. Barry OP, Praticò D, Savani RC, FitzGerald GA. Modulation of monocyte‐endothelial cell interactions by platelet microparticles. J Clin Invest. 1998;102(1):136–144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95. Rozmyslowicz T, Majka M, Kijowski J, et al. Platelet‐and megakaryocyte‐derived microparticles transfer CXCR4 receptor to CXCR4‐null cells and make them susceptible to infection by X4‐HIV. Aids. 2003;17(1):33–42. [DOI] [PubMed] [Google Scholar]
- 96. Camussi G, Deregibus MC, Bruno S, Cantaluppi V, Biancone L. Exosomes/microvesicles as a mechanism of cell‐to‐cell communication. Kidney Int. 2010;78(9):838–848. [DOI] [PubMed] [Google Scholar]
- 97. Baj‐Krzyworzeka M, Szatanek R, Węglarczyk K, et al. Tumour‐derived microvesicles carry several surface determinants and mRNA of tumour cells and transfer some of these determinants to monocytes. Cancer Immunol Immunother. 2006;55(7):808–818. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98. Deregibus MC, Cantaluppi V, Calogero R, et al. Endothelial progenitor cell–derived microvesicles activate an angiogenic program in endothelial cells by a horizontal transfer of mRNA. Blood. 2007;110(7):2440–2448. [DOI] [PubMed] [Google Scholar]
- 99. Yuan A, Farber EL, Rapoport AL, et al. Transfer of microRNAs by embryonic stem cell microvesicles. PloS One. 2009;4(3):e4722. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100. Valenti R, Huber V, Filipazzi P, et al. Human tumor‐released microvesicles promote the differentiation of myeloid cells with transforming growth factor‐β–mediated suppressive activity on T lymphocytes. Cancer Res. 2006;66(18):9290–9298. [DOI] [PubMed] [Google Scholar]
- 101. Whiteside TL. Immune modulation of T‐cell and NK (natural killer) cell activities by TEXs (tumour‐derived exosomes). Biochm Soc Trans. 2013;41(1):245–251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102. Henderson MC, Azorsa DO. The genomic and proteomic content of cancer cell‐derived exosomes. Front Oncol. 2012;2:38. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103. Melo SA, Sugimoto H, O'Connell Joyce T, et al. Cancer exosomes perform cell‐independent MicroRNA biogenesis and promote tumorigenesis. Cancer Cell. 26(5):707–721. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104. Taylor DD, Gercel‐Taylor C. MicroRNA signatures of tumor‐derived exosomes as diagnostic biomarkers of ovarian cancer. Gynecol Oncol. 2008;110(1):13–21. [DOI] [PubMed] [Google Scholar]
- 105. Rosell R, Wei J, Taron M. Circulating MicroRNA signatures of tumor‐derived exosomes for early diagnosis of non–small‐cell lung cancer. Clin Lung Cancer. 2009;10(1):8–9. [DOI] [PubMed] [Google Scholar]
- 106. Mincheva‐Nilsson L, Baranov V. Cancer exosomes and NKG2D receptor–ligand interactions: impairing NKG2D‐mediated cytotoxicity and anti‐tumour immune surveillance. Semin Cancer Biol. 2014;28:24–30. [DOI] [PubMed] [Google Scholar]
- 107. Smalley DM, Sheman NE, Nelson K, Theodorescu D. Isolation and identification of potential urinary microparticle biomarkers of bladder cancer. J Proteome Res. 2008;7(5):2088–2096. [DOI] [PubMed] [Google Scholar]
- 108. Kuwabara Y, Ono K, Horie T, et al. Increased MicroRNA‐1 and MicroRNA‐133a levels in serum of patients with cardiovascular disease indicate myocardial damage. Circulation. 2011;4(4):446–454. [DOI] [PubMed] [Google Scholar]
- 109. Lv L‐L, Cao Y‐H, Pan M‐M, et al. CD2AP mRNA in urinary exosome as biomarker of kidney disease. Clin Chim Acta. 2014;428:26–31. [DOI] [PubMed] [Google Scholar]
- 110. Chen J‐F, Mandel EM, Thomson JM, et al. The role of microRNA‐1 and microRNA‐133 in skeletal muscle proliferation and differentiation. Nat Genet. 2006;38(2):228–233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111. Resnick KE, Alder H, Hagan JP, Richardson DL, Croce CM, Cohn DE. The detection of differentially expressed microRNAs from the serum of ovarian cancer patients using a novel real‐time PCR platform. Gynecol Oncol. 2009;112:55–59. [DOI] [PubMed] [Google Scholar]
- 112. Rabinowits G, Gerçel‐Taylor C, Day JM, Taylor DD, Kloecker GH. Exosomal MicroRNA: a diagnostic marker for lung cancer. Clin Lung Cancer. 2009;10(1):42–46. [DOI] [PubMed] [Google Scholar]
- 113. Que R, Ding G, Chen J, Cao L. Analysis of serum exosomal microRNAs and clinicopathologic features of patients with pancreatic adenocarcinoma. World J Surg Onc. 2013;11(1):219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114. Whiteside TL. Tumor‐derived exosomes and their role in tumor‐induced immune suppression. Vaccines. 2016;4(4):35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115. Viaud S, Terme M, Flament C, et al. Dendritic cell‐derived exosomes promote natural killer cell activation and proliferation: a role for NKG2D ligands and IL‐15Rα. PLoS One. 2009;4(3):e4942. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116. Runz S, Keller S, Rupp C, et al. Malignant ascites‐derived exosomes of ovarian carcinoma patients contain CD24 and EpCAM. Gynecol Oncol. 2007;107(3):563–571. [DOI] [PubMed] [Google Scholar]
- 117. Kunigelis KE, Graner MW. The dichotomy of tumor exosomes (TEX) in cancer immunity: is it all in the ConTEXt? Vaccines (Basel). 2015;3(4):1019–1051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118. Lynch S, Santos SG, Campbell EC, et al. Novel MHC class I structures on exosomes. J Immunol. 2009;183(3):1884–1891. [DOI] [PubMed] [Google Scholar]
- 119. LeBrasseur N. MHC and antigen in exosomes. J Cell Biol. 2007;179(1):4–4. [Google Scholar]
- 120. Escudier B, Dorval T, Chaput N, et al. Vaccination of metastatic melanoma patients with autologous dendritic cell (DC) derived‐exosomes: results of thefirst phase I clinical trial. J Transl Med. 2005;3(1):1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121. Ristorcelli E, Beraud E, Mathieu S, Lombardo D, Verine A. Essential role of Notch signaling in apoptosis of human pancreatic tumoral cells mediated by exosomal nanoparticles. Int J Cancer. 2009;125(5):1016–1026. [DOI] [PubMed] [Google Scholar]
- 122. Tan A, De La Peña H, Seifalian AM. The application of exosomes as a nanoscale cancer vaccine. Int J Nanomedicine. 2010;5:889–900. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123. Simona F, Laura S, Simona T, Riccardo A. Contribution of proteomics to understanding the role of tumor‐derived exosomes in cancer progression: state of the art and new perspectives. Proteomics. 2013;13(10‐11):1581–1594. [DOI] [PubMed] [Google Scholar]
- 124. Keller S, König A‐K, Marmé F, et al. Systemic presence and tumor‐growth promoting effect of ovarian carcinoma released exosomes. Cancer Lett. 2009;278(1):73–81. [DOI] [PubMed] [Google Scholar]
- 125. André F, Schartz NEC, Chaput N, et al. Tumor‐derived exosomes: a new source of tumor rejection antigens. Vaccine. 2002;20:A28–A31., Supplement 4, [DOI] [PubMed] [Google Scholar]
- 126. Kharaziha P, Ceder S, Li Q, Panaretakis T. Tumor cell‐derived exosomes: a message in a bottle. Biochim Biophys Acta. 2012;1826(1):103–111. [DOI] [PubMed] [Google Scholar]


