Summary
We examined the evolutionary history of leading multidrug resistant hospital pathogens, the enterococci, to their origin hundreds of millions of years ago. Our goal was to understand why, among the vast diversity of gut flora, enterococci are so well adapted to the modern hospital environment. Molecular clock estimation, together with analysis of their environmental distribution, phenotypic diversity and concordance with host fossil records, place the origins of the enterococci around the time of animal terrestrialization – 425 – 500 MYA. Speciation appears to parallel the diversification of hosts, including the rapid emergence of new enterococcal species following the End Permian Extinction. Major drivers of speciation include changing carbohydrate availability in the host gut. Life on land would have selected for the precise traits that now allow pathogenic enterococci to survive desiccation, starvation and disinfection in the modern hospital; foreordaining their emergence as leading hospital pathogens.
eTOC
Why, among the vast diversity of gut microbiota, has enterococci become so well adapted to the modern hospital environment?

Introduction
Enterococci, typically less than 0.1% of the core gut microbiota of humans (Schloissnig et al., 2013), emerged from that vast diversity to become leading multidrug resistant, hospital-adapted pathogens (Van Tyne and Gilmore, 2014). This happened twice – multidrug resistant (MDR), pathogenic lineages of both Enterococcus faecalis and distantly related E. faecium have emerged. Enterococci serve as collection and distribution points for mobile elements, transmitting a variety of antibiotic resistances to both gram-positive and gram-negative species (Courvalin, 1994) including the transmission of vancomycin resistance to methicillin-resistant strains of Staphylococcus aureus (Weigel et al., 2003).
Most MDR enterococci belong to highly hospital-adapted lineages (Lebreton et al., 2013a; Raven et al., 2016). MDR enterococci differ from commensals in lacking CRISPR defenses (Palmer et al., 2010), and possess genomes that are often >25% larger than commensals through accretion of mobile elements –phages, pathogenicity islands and resistance genes (Shankar et al., 2002; Paulsen et al., 2003). Resistant strains are transmitted through cycles of: 1) low level oral acquisition from the contaminated hospital environment, 2) survival of transit through the digestive tract, 3) establishment and expansion in the antibiotic perturbed colon (leading to recontamination of the hospital environment), and 4) translocation into the bloodstream (or less direct infection of the urinary tract or surgical sites) (Gilmore et al. 2013). Properties that underlie the emergence of enterococci as hospital pathogens include environmental persistence and resistance to disinfection protocols, efficient colonization of a new host, and the intrinsic ability to broadly withstand antimicrobials that eliminate gut competitors. In the antibiotic treated colon, intrinsic resistance concentrates enterococci with overtly resistant microbes harboring mobile elements, which they readily acquire and further transmit. Infection depends on the strain’s ability to invade and evade host defenses (Van Tyne and Gilmore, 2014; Arias and Murray, 2012). The challenge in therapeutically resolving infection derives from intrinsic and acquired antibiotic resistance. This raises fundamental questions: 1) Do all enterococcal species possess such attributes?; 2) What genes were gained since divergence from ancestral microbes that endow them with environmental persistence and tolerance to disinfection?; and 3) Since enterococci are widespread in nature and colonize the gastrointestinal tracts of most land animals, including insects and invertebrates (Mundt, 1963; Martin and Mundt, 1972; Van Tyne and Gilmore, 2014), what events selected for the emergence of this genus, and diversification within it?
To answer these questions, we selected 24 enterococcal species representing all major phylogenetic branches (Van Tyne and Gilmore, 2014) (Fig. S1). We determined and compared genotype to phenotype, and, where possible, host association. We further compared traits found in all enterococci to those in commensal and multidrug resistant enterococci most commonly associated with humans. The data collectively indicate that traits that now facilitate enterococcal survival and transmission in hospitals were of selective advantage early in the Paleozoic Era, and support a model for the origin of the enterococci around the time of terrestrialization of their animal hosts.
Results
Phylogeny of enterococci, and host association
An existing Enterococcus 16S rDNA-based phylogeny was used to select representative species for comparative genome analysis. The genomes of 24 species spanning the diversity of the genus (Fig. S1) were sequenced and compared to those of 5 closely related species: Melissococcus plutonius (Cai and Collins, 1994), Tetragenococcus halophilus (Collins et al., 1990), Vagococcus lutrae (Collins et al., 1989), Carnobacterium maltaromaticum (Collins et al., 1987) and Lactococcus garviae (Wallbanks et al., 1990). Very high quality draft genome sequences (2 to 13 contigs) were generated from hybrid assemblies of paired end reads of fragment (180 bp) and mate-pair (mean 2.8 kb) libraries (Illumina), which were then connected by long sequence reads (mean 1.1 kb, PacBio) (Table S1). Previously sequenced genomes of E. faecalis and E. faecium associated with multidrug resistant hospital infection were included in the comparison (Bourgogne et al., 2008; Lam et al., 2012; Lebreton et al., 2013a; Palmer et al., 2012; Paulsen et al., 2003; Sahm et al., 1989).
Enterococcal genome sizes span a surprisingly wide range – from 2.3 Mb (E. sulfureus) to 5.4 Mb (E. pallens) (Fig. 1A, Table S1). Enterococcal and outgroup genomes share 526 single copy orthologous genes in common (Fig. S2A), from which a SNP-based phylogeny (Stamatakis, 2006) was constructed (Fig. 1B). Four main lineages are evident and supported by BAPS (Cheng et al., 2013) clustering (Table S1). Compared to the mean (3.2 Mb), E. pallens and E. columbae groups possess significantly larger (mean 4.6 Mb, p<0.01) or smaller (mean 2.7 Mb, p<0.01) genomes, respectively. As expected, L. garviae, C. maltaromaticum and Vagococcus lutrae connect outside of the Enterococcus genus; however, M. plutonius and T. halophilus are solidly embedded within it (Fig. 1B). Therefore, for subsequent analyses, the latter were included as enterococci.
Figure 1. Genome-based phylogeny of the Enterococcus genus.
A: Distribution of Enterococcus species genome sizes (color coded by phylogenetic group). B: Phylogenomic tree of Enterococcus and outgroup species (grey) based on alignment of 526 single copy core genes. Species groups: E. faecalis group (blue), E. columbae group (green), E. pallens group (red) and E. faecium group (yellow). Each node is assigned a unique numeric identifier. Icons represent host associations inferred from both qualitative and quantitative (Table S2) analyses of the literature. C: Prevalence of Enterococcus and Vagococcus in the GI tract of land (n= 492) or aquatic hosts (n= 1300) isolated from the wild. Presence or absence was determined by 16S rDNA analysis of microbiome samples available in public databases (Table S3). Mean and SEM are indicated. Statistical significance was evaluated by Fisher’s Exact Test.
Pioneering work in the 60’s and 70’s by Mundt and colleagues (Martin and Mundt, 1972; Mundt, 1963), found a wide distribution of enterococci in land animals – birds, insects, mammals and reptiles. To gain additional insight into host association, we examined the literature for other individual reports of enterococci within a host, and performed a quantitative term analysis of PubMed and Web of Science (Table S2). Host tropisms were operationally defined when two or more publications reported a given host association. Assuming similar reporting biases for all microbes queried, enterococci were reported mainly in association with land animals (97% of reports), whereas outgroup species of Vagococcus and Carnobacterium were mainly reported in association with aquatic animals (87% of reports) (Table S2). Others previously noted the association of ancestral microbes, vagococci and carnobacteria, with marine life (compiled by Buller, 2014): Vagococcal species occur in European seabream and river water, otter, shrimp, trout and Atlantic salmon (and animals that consume them including seals and porpoises) (compiled in Buller, 2014); and Carnobacterium species are found in various fish, including trout, salmon, wild bass and catfish (compiled in Buller, 2014). E. faecalis is most ubiquitously reported. Enterococcal species reported in insects are often found in the gastrointestinal (GI) tracts of other animals up the food chain (Fig. 1B). M. plutonius is reported mainly in bees, where it causes European foulbrood (Bailey, 1983). Two closely related species, E. cecorum and E. columbae appear to have a strong tropism for birds; with the latter occurring only in birds of the Columbidae family (Baele et al., 2002).
To substantiate the apparent occurrence of enterococci mainly in land animals, and ancestral vagococci in aquatic life, we quantified enterococcal or vagococcal 16S rDNA within reported gut metagenomes of land and marine animals. Enterococcal 16S DNA is highly enriched in metagenomes from land animals (Fisher’s exact test, p < 10−51), and vagococcal sequences are highly enriched in marine life (p < 10−21), independently supporting inferences from the literature (Table S3, Fig. 1C).
Genes and traits that distinguish enterococci from other microbes
Although the Enterococcus pan-genome is large (Fig. S2AB), a core of 1037 genes occurs in >90% (allowing for genes missed by sequence gaps) of enterococcal species (including M. plutonius and T. halophilus) (Table S4, tab 1). Of this 1037 core gene set, it was of interest to determine whether any were unique among eubacteria. To identify possibly unique genes, we compared the core set of 1037 enterococcal genes to 100 genomes from representative branches of the eubacterial domain (Table S4, tab 2). Candidate genes absent from those genomes, and hence potentially unique to enterococci, were further interrogated for novelty by BLASTn and BLASTx (Altschul et al., 1990) comparison to the NCBI non-redundant database. Genes lacking identifiable homologs elsewhere (>60% amino acid identity over >40% of gene product length), were defined as Enterococcus-unique. This identified a set of 10 unique genes (Fig. 2A, Table S4 tab 1) – 7 lacking any associable function. These 10 Enterococcus-unique genes do not spatially cluster on the chromosome.
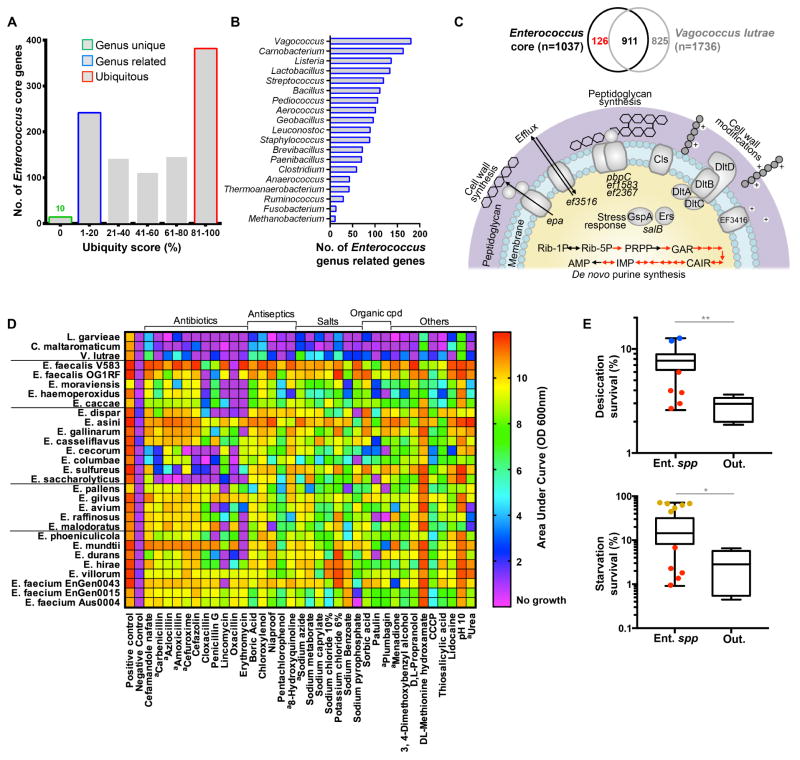
Figure 2. Genetic and phenotypic differences from outgroup species.
A: Ubiquity of 1037 Enterococcus core genes in eubacteria (BlastX against 100 representative eubacterial genomes, Table S4 tab 2), in quintiles. B: Genera of microbes harboring genes seldom found outside of enterococci (1st quintile 1 – 20 genera), ranked by the number of 1st quintile genes shared. C: Above, Venn diagram showing shared and unique genes in Enterococcus core genome and Vagococcus lutrae LBD1. Below, Diagram highlighting role of functional groups of Enterococcus core genes not occurring in Vagococcus: De novo purine biosynthesis potentially feeding into stress response (12 genes displayed with red arrows, p< 0.0001), cell wall biosynthesis and modification (15 genes, p= 0.0002) and stress response (3 genes). PRPP, phosphoribose pyrophosphate; GAR, glycinamide ribonucleotide; CAIR, carboxyaminoimidazole ribonucleotide; IMP, Inosine monophosphate; AMP, Adenosine monophosphate. D: Heatmap showing significant differences in average growth (at least 2-fold greater average growth, p < 0.01) between enterococcal and outgroup species in Biolog phenotype screen. aCompounds showing a similar pattern when tested in multiple concentrations. E: Survival to desiccation and starvation by Enterococcus spp. (Ent. spp) compared to outgroup strains V. lutrae LBD1, V. fluvialis DIV0098, C. maltaromaticum ATCC35586, L. garviae NCIMB31208 and DIV0709 (Out.). Box plots drawn using the average survival value from three independent experiments for each strain. Box plots show the median, first and third quartiles and min/max values. Colored data points show the mean values for strains from selected enterococcal phylogenetic groups (Fig. 1A).
Unique properties that distinguish enterococci from other microbes may also stem from combinations of unusual genes that are not necessarily unique. We therefore categorized the remaining Enterococcus core genes by rarity of occurrence in the 100-comparator eubacterial genome set (Table S4, tabs 1 and 2). The rarest quintile (those occurring in 1–20% of other genera), consisted of 243 core genes that exist mainly in ancestral species and related Firmicutes that live in the digestive tracts of animals (Fig. 2B). Most (68%; Fig. S2C) are of unknown function, but presumably relate to the success of these gram-positive species in mixed GI tract communities.
We next identified those core genes, and phenotypes, that specifically distinguish enterococci from ancestral lineages. V. lutrae LBD1 (isolated from the GI tract of a wild caught largemouth bass [Lebreton et al., 2013b]), lacks only 126 of the 1037 core enterococcal genes (Fig. 2C, Table S4 tab 3). Importantly, the 126 genes gained by enterococci since divergence from its last common ancestor with V. lutrae, are significantly enriched in cell wall modification (p < 0.001) and de novo purine biosynthesis (p < 0.001), functions related to stress response (Jordan et al., 2008; Gaca et al., 2012). This implies that survival in a fundamentally different environment, requiring cell wall alterations and potentially prone to environmental stress, selected for the divergence of the enterococci from Vagococcus.
To identify phenotypes that distinguish enterococci from ancestral lineages, growth of all enterococci and outgroup members was assessed under 1344 individual growth conditions (Biolog, Inc) (Fig. S2E). Enterococci as a group displayed at least 2-fold greater growth (p < 0.01) than the mean for outgroup members under 45 of the conditions tested (Fig. S2E, Table S5). Most of the 45 distinguishing phenotypes related to increased tolerance to harsh conditions: hydrogen- and ionic-bond solvents, detergents, antibiotics and other biocides (Fig. 2D). Interestingly, V. lutrae, but not Carnobacterium maltaromaticum, resists sodium azide, comparable to enterococci, and in fact grows on enterococcal selective media (bile esculin azide agar) (Isenberg et al., 1970)(Fig. S2FG). Two phylogenetically distant species, E. columbae and E. saccharolyticus, appear to have subsequently lost sodium azide resistance (Fig. S2G), raising the possibility that additional enterococcal species may be missed when using bile esculin azide agar for isolation.
All tested enterococci have global resistance to environmental stresses similar to those that now occur in modern hospitals, including to β-lactam antibiotics as well as disinfectants (Fig. 2D), prompting us to test enterococci and outgroups for survival to desiccation and starvation, conditions occurring in the hospital infection cycle. Enterococci were found more tolerant than the outgroup to both conditions (p<0.05; Fig. 2E): Both E. faecalis strains exhibited the greatest resistance to desiccation (p=0.031). E. faecium desiccation resistance was more typical of the level intrinsic to the enterococci, but species of the E. faecium group were found to be the most starvation tolerant (p=0.017). In contrast, the E. pallens group, with large genomes, showed the least tolerance for desiccation or starvation suggesting that they may derive from hosts that inhabit ecologies where regular desiccation and starvation are less selective, such as wet coastal margins.
Energy source drives enterococcal speciation
To infer changes in the host gut environment that may have driven the diversification of enterococci into various species, we performed a flux analysis (Swofford, 2001) of genes gained and lost at each node of speciation (Fig. 3A). Diversification at each node is associated with an average gain of 618 (± 304) and loss of 135 (± 103) genes, with notable exceptions. The large genome E. pallens group was precipitated by an initial gain of 1157 genes at node 18, followed by successive large gains at nodes 19 (410 genes) and 20 (408 genes). Large gains also occurred at the leaves, which is inflated by phages, mobile elements and other strain-specific determinants. For example, 1775 new genes were acquired by the E. pallens strain analyzed, which cluster in syntenic regions from which genes core to the genus are largely excluded (Fig. S2H). The E. columbae group is the exception, having undergone genome streamlining at multiple successive nodes (except for node 12, which led to the speciation of E. gallinarum and E. casseliflavus).
Figure 3. Functional classification of niche specifying genes.
A: Phylogenetic analysis using parsimony (PAUP) and minimization of the number of gain/loss events predicting flux in gene content since last common ancestor (blue = gain; red = loss) at nodes (italicized numbers 1 to 28) and leaves of the SNP-based phylogenetic tree. The size of the pie chart reflects the amplitude of total gene flux (gain + loss). B, COG functional classification of Enterococcus core genes, and niche specifying gene groups.
Genes gained (n=23,232) or lost (n=6,223) were operationally defined as niche-specifying genes (NSG) (Supplemental dataset 1). Most (63%) encode hypothetical proteins or proteins of unknown function. For 10,803 genes with ascribable putative functions, the greatest flux was in genes for carbohydrate utilization and its transcriptional control (Fig. 3B). Of carbohydrate utilization genes in flux, most (62%) code for distinct phosphotransferase transporter systems (PTS). This was consistent throughout the tree, indicating that differences in carbohydrate utilization are a common driver of species diversification since the origin of the genus (Fig. S3). These differences supplement shared central metabolic carbohydrate utilization pathways (pentose phosphate shunt, glycolysis/gluconeogenesis, Entner-Doudoroff pathway [missing in E. moraviensis, E. haemoperoxidus and E. caccae]) that allow utilization of glucose, fructose, galactose, and mannose by all species [Table S4, tab 1]. Enigmatically, although possessing the largest complement of annotated carbohydrate utilization genes, the E. pallens group ferment the fewest carbohydrates under the conditions tested (Fig. S4A), suggesting their substrates (e.g., complex polysaccharides) are not those typically fermented and tested, or that their novel carbohydrate utilization pathways are tightly regulated in ways not currently understood. Large gains and losses of genes associated with amino acid uptake and metabolism were also observed among genes in flux (12% of 10,803 annotated NSG), further supporting nutrient availability as a main driver of speciation (Fig. S3).
Host association and adaptation
Because of the emergence of enterococci as hospital pathogens, we closely examined the patterns of gene gain and loss in the evolution of M. plutonius into an overt pathogen, from its common ancestor with the most widespread generalist, E. faecalis. In becoming a pathogen, M. plutonius lost 46 PTS transporters and enzymes involved in the utilization of carbohydrates, and also lost 50 genes for the biosynthesis of all amino acids except alanine, serine and glutamine (Fig. S3 and Supplemental Dataset 1), many of which occur in predicted pathways (Paulsen et al., 2003) (Okumura et al., 2011) (Fig. 4AB). M. plutonius also acquired a pathway for the biosynthesis of a high-mannose N-linked glycan, a trait rare among eubacteria and likely to result in modification of its cell surface (Fig. 4B), possibly in a manner associated with pathogenesis (Nothaft and Szymanski, 2010). This pattern of genome reduction contrasts sharply with the recent accretion of mobile elements and resulting genome expansion by >25% of E. faecalis to become hospital pathogens (Paulsen et al., 2003).
Figure 4. Host-adaptation of enterococci.
A: Carbohydrate utilization for E. faecalis (E.fs) and M. plutonius (M.pl). Other tested carbohydrates, by Biolog phenotype microarrays, are shown on the full profiles (Fig. S4A). B: Metabolic pathways gained or lost by M. plutonius since node 2 (Fig. 3A). Each arrow represents an enzymatic step (black = ancestral enzyme; grey = no gene in Enterococcus yet associated; blue = gained; red = lost) of the KEGG pathways predicted for E. faecalis V583 and M. plutonius ATCC35311. Other key features gained by M. plutonius: restriction system type I and III (Rm-1, -3); high mannose N-linked glycan biosynthesis (Poly N-Man); Ox-P, oxidative phosphorylation. C: Representative results of pH based phenotypic screen for uric acid metabolism (color change to yellow indicates acidification, color change to red indicates uric acid metabolism by alkalinity from ammonia release). Typical result for E. faecalis or other enterococcal species (Ent. spp.) when provided no carbon source, uric acid (UA) only, or glucose (Glu) and uric acid as indicated (TnUA – representative result for each of the E. faecalis insertion mutants lacking the ability to produce NH4+ in the presence of urate). D: Competitive index for growth in vitro (BHI, to control for growth advantage or defect of the transposon mutant versus the wild-type strain in rich media), or in colonization of the GI tract of G. mellonella larvae, for E. faecalis MMH594 (WT) over the CFU of the indicated uric acid metabolism deficient mutant (Mut) recovered. E: Carbohydrate utilization for bird-associated E. cecorum (E.ce) and E. columbae (E.co). Other tested carbohydrates are shown on the full Biolog profiles (Fig. S4A). F: Metabolic pathways, gained (blue arrows) or lost (red arrows) by E. columbae since node 15 (Fig. 3A), predicted by searching the KEGG pathway map. Chemotaxis (Chem.); flagellar motility (Mot.).
In the process of becoming a widespread generalist, E. faecalis gained 335 genes in diverging from M. plutonius. A trait unique among enterococci, gained by E. faecalis after the split with M. plutonius, is the ability to metabolize uric acid (Fig. 4C), a plentiful source of carbon and nitrogen in the gut of insects and other uricotelic animals (Bursell, 1967) (Fig. S4BC). Relevant to pathogenesis, uric acid metabolism was recently shown to enhance biofilm formation by E. faecalis, requiring a selenium-dependent molybdoenzyme, xanthine dehydrogenase (EF2570) and a selenophosphate synthetase, selD (EF2567) for this activity (Srivastava et al., 2011). To identify genes involved in this pathway, and determine when they were gained, we constructed (Fig. S4D) and screened 9120 mutants in a transposon insertion library for loss of the ability to generate the pH increase from ammonia liberation from uric acid. As for the utilization of novel carbohydrates, the results of the screen indicate that the enabling event appears to stem from the gain of a unique transporter by node 4 (EF3277), which connects to pathways gained earlier and evident at node 3, including 4 genes predicted to code for selenium metabolism to provide the necessary cofactor (Srivastava et al., 2011) for xanthine dehydrogenase (EF2570) (Figure S4E).
The effect of uric acid in promoting biofilm formation had been suggested to confer increased pathogenicity to E. faecalis (Srivastava et al., 2011), but the gain of this capability appears to predate its evolution into a hospital pathogen as uric acid metabolism occurs in commensal OG1RF as well. It was therefore of interest to determine whether the ability to metabolize uric acid confers a more general advantage in the gut of a uricotelic host. The fitness of two different uric acid metabolism mutants was compared to the wild type E. faecalis parent in the GI tract of G. mellonella larvae (Fig. S4FG). Wild type E. faecalis exhibited a significant (p<0.05) colonization advantage over either mutant tested. No difference was observed in BHI, which served as a control for general growth defects in the mutants (Fig. 4D).
To further associate specific enterococcal genes with host adaptation, the genomes of E. columbae and E. cecorum – species with strong host associations – were compared. E. columbae colonizes pigeons (Baele et al., 2002), whereas E. cecorum is found in a variety of birds (Aarestrup et al., 2002). In adapting to pigeons, E. columbae gained 35 genes related to carbohydrate uptake and utilization (Fig. S3A, Fig. 4EF), which is consistent with carbohydrate metabolism driving speciation. E. columbae (but not E. cecorum) metabolizes plant sugar alcohols, pectin, all carbohydrates forming rhamnogalacturonan I (RG-I; a core constituent of plant cell walls and mucilage (Yapo, 2011)), and also carbohydrates abundant on the surface of eukaryotic cells (Fig. 4E). This correlates with the restricted diet of Columbidea birds, which feed exclusively on plant-derived material; and produce and feed their young ‘crop milk’ – a mixture of desquamated epithelial cells and fermentation products of the adult diet (Podulka, 2004). E. columbae also acquired 40 genes for flagellar assembly and chemotaxis, which may relate to novel aspects of this vertical transmission to offspring.
Origins of the enterococci
To gain insight into ecological relationships between all other enterococcal species, where definitive hosts information is much more limited, and to quantify the divergence distance between species, we calculated a) the fraction of genes shared between each genome pair, and b) the average nucleotide identity (ANI) between those shared genes (Fig. 5A and Fig. S5). Pairwise analysis allows the maximum number of genes to be included in calculating distance (Konstantinidis and Tiedje, 2005). Shared gene content identifies similarities in the ecologies occupied.
Figure 5. Calibration of enterococcal evolution.
A: Average Nucleotide Identity plot. Each dot represents a pairwise comparison of two genomes. Enterococcus spp. versus outgroup species (grey); Enterococcus spp. versus M. plutonius (red); Enterococcus spp. versus T. halophilus (green); intra-genus comparisons (black). B: Calibration of ANI scale with divergence times and ANI values calculated for E. coli versus S. typhimurium and Vibrio spp.; and A. hydrophila versus other Aeromonas spp. Larger black dots represent the mANI for E. coli versus S. typhimurium (which diverged 140 MYA); mANI for A. hydrophila versus other Aeromonas spp. (which diverged 184 MYA); and mANI for E. coli versus Vibrio spp. (which diverged 400 MYA). Published range of error for each is shown by flanking small black dots. Horizontal brown bars represent a speciation event at a given mANI value within the Enterococcus genus (corresponding to nodes in Fig. 1B phylogenetic tree) and the horizontal blue bar represents the last common ancestor (LCA) with an outgroup species. C: Fossil record corrected version of panel B, with theoretical origin of Enterococcus genus (i.e. after divergence from LCA with outgroup genus and before first Enterococcus speciation event) anchored to the initial terrestrialization of animals 425 MYA (vertical grey line).
ANI divergence between each Enterococcus species and an outgroup member (Lactococcus, Vagococcus or Carnobacterium), forms a distinct cloud on the scatter plot to the left (i.e., ANI < 63%) of that formed by all intra-Enterococcus species comparisons (Fig 5A and Fig. S5); verifying their positions outside the Enterococcus genus. In contrast, all pairwise comparisons between established Enterococcus spp. and either M. plutonius or T. halophilus, fell completely within the intra-genus cloud, again arguing for their inclusion among the enterococci.
Bacterial evolution is calibrated using, as a first approximation, molecular clocks; and because of uncertainty in those clocks, refinement based on concordance with the fossil record and other information (Ochman et al., 1999). We adapted molecular clocks previously established for estimating the divergence of E. coli from Salmonella typhimurium or the Vibrio group (Ochman and Wilson, 1987), and for estimating the time of divergence with the genus Aeromonas (Lorén et al., 2014). These clocks were chosen because they were established for microbes associated with host guts that possess generation times similar to that of enterococci. To translate these clocks, which were based on 16S rDNA and multilocus sequence comparisons, to ANI values, we calculated the corresponding mean ANI values between species used to establish those clocks: a) 3 strains of E. coli compared to 3 strains of S. typhimurium; b) 3 strains of E. coli compared to Vibrio cholerae, Vibrio parahaemolyticus, and Vibrio fischeri; and c) three strains of Aeromonas hydrophila compared to Aeromonas salmonicida, A. media and A. bivalvium (Fig. 5B). Next, to calculate a single, maximally precise genetic distance for each extant Enterococcus species from each node of speciation, we calculated the mean ANI value (mANI) for all species that connect to a given node in the phylogenetic tree (Table S6). The small standard deviation of this mANI value shows that the divergence rate for various enterococcal species is relatively constant.
This approach places the origin of the Enterococcus genus around 500 MYA (+/− 130.5 MYA; at an arbitrary midpoint between the last common ancestor shared with outgroup species and the first bifurcation within the Enterococcus genus) (Fig. 5B), in broad agreement with theoretical deduction (Van Tyne and Gilmore, 2014). Noticeably, subsequent radiations of enterococcal species occur in two distinct waves, separated by a clear gap between ANI values of 69 and 73% (Fig. 5AB).
To generate a model on the origins of enterococci that reconciles this molecular clock estimation with the fossil record of their hosts and current distribution, we took the following into consideration: 1) Enterococci are common among terrestrial life and comparatively rare in aquatic animals, which suggests that life on land fostered their proliferation and diversification; and 2) Desiccation and starvation resistance, which distinguish the enterococci from more commonly marine ancestral Vagococcus and Carnobacterium, appear to be a trait common to the original Enterococcus and now shared by members of both branches from the initial bifurcation of species within the genus. The earliest fossil record of terrestrialization by animals possessing a gut consortium is that of arthropods in the middle of the Silurian period, about 425 MYA (Benton et al., 2010). Molecular analysis suggests that arthropods may have colonized land earlier, in the late Cambrian to early Ordovician (510–471 MYA) (Rota-Stabelli et al., 2013). Placing the earliest origin of enterococci to coincide with the most conservative estimate of terrestrialization 425 MYA, based on fossil records, requires leftward compression in molecular clock estimation that is still within its estimated error (Fig. 5C).
Discussion
The 16S rDNA-based tree of life shows that enterococci emerged from among Vagococcus-like progenitors, which arose from the Carnobacteriaceae (Fig. S1B). Many Carnobacteriaceae are psychrophiles associated with marine environments and ice (Collins et al., 1987; Nicholson et al., 2012). One species, Carnobacterium pleistocenium, was revived from approximately 32,000 year old permafrost (Pikuta et al., 2005). Another species, Carnobacterium funditum sp. nov., was isolated from the frigid anoxic waters of Ace Lake Antarctica (Franzmann et al., 1991). Carnobacteria are among microbes most likely to exist in austere simulated Martian environments (Schuerger and Nicholson, 2016). Vagococci diverged from carnobacteria, and remain generally associated with marine animals (Michel et al., 2007; Svanevik and Lunestad, 2011; Buller, 2014). Prior to this work, no Vagococcus genome had been sequenced, so we determined that for V. lutrea strain LBD1, which we isolated from the gut of a line caught largemouth bass (Lebreton et al., 2013b).
Unlike C. maltaromaticum, V. lutrea is capable of growing on bile esculin azide agar (Fig. S2F). However, C. pleistocenium is capable of growing in the presence of 5% NaCl and fermenting esculin (Pikuta et al., 2005), traits preserved in the enterococci (Fig. 2D). It therefore appears that the ability to thrive in saline marine habitats predates the branching of vagococci from carnobacteria, and the ability to colonize ecologies with high levels of bile was established in the vagococci. The divergence from ancestral species with reduced genomes (Lebreton et al., 2013b, Leisner et al., 2012) indicates a heritage of obligate association with other organisms in a food web, either in marine sediments, or possibly in bile-secreting hosts in marine habitats, potentially extending back to survival in ancient ice.
We now know that over the past 15 million years, gut microbes cospeciated with hominid hosts (Moeller et al., 2016). This appears to be true on a much longer timescale, in that enterococcal speciation also parallels radiation of hosts, with the ability to utilize new carbohydrate sources among the main drivers of speciation (Fig. 3 and 4). Carbohydrate-driven enterococcal speciation is ongoing. In studying the origins of multidrug resistant hospital strains, we previously noted a split within the species E. faecium, at about the time of human urbanization, that separates commensal human strains from those of agricultural animals (from which hospital adapted lineages derive), by a distance that approximates species level distinction (Lebreton et al., 2013a; Palmer et al., 2012). Importantly, the largest blocks of difference between the genes of human commensal strains and the animal/hospital clade are operons for carbohydrate uptake and utilization (Lebreton et al., 2013a).
Rooting the emergence of enterococci commensurate with terrestrialization, conservatively about 425 MYA (Fig. 5C), has an interesting effect on the timing of nodes of divergence within the genus. Following a time of stability, reflected by a gap in new speciation, a prominent second wave of enterococcal species radiations begins at the precise time of a second wave of land animal radiation – immediately following the End Permian Extinction, 251 MYA (Benton and Twitchett, 2003) (Fig. 6). Placing the origin of the enterococci at about the time of animal terrestrialization has the effect of making diversification of enterococci highly congruent with the diversification of their terrestrial hosts (Fig. 6), a prospect strongly supported by the findings of gene flux analysis (Fig. 3 and 4). Although this model is coherent with current knowledge, specific unknowns, including exactly when the first Enterococcus emerged, and ongoing debate about the precise timing of terrestrialization of animals, remain as possible sources of error.
Figure 6. Ancient origins.
The timing of nodes divergence was calculated by mANI analysis (Fig. 5A), using a bacterial molecular clock (Fig. 5B) refined by correlation with fossil records (Fig. 5C), conservatively rooting the theoretical emergence of Enterococcus to the terrestrialization of animals by 425 MYA. Enterococcal diversity (left y-axis and tan area) represents the cumulative number of enterococcal lineages through time. Land animal diversity, computed from fossil records, (right y-axis and grey area), shows fluctuation in number of animal families over the same time scale. The position of animal classes on the time scale indicates periods of evolutionary radiations.
All of the above observations are consistent with a model that places the emergence and proliferation of enterococci in parallel with the terrestrialization and proliferation of their hosts. From aquatic animals, excreted gut flora emerge into the comparatively hospitable waterborne community (Venter et al., 2004), and settle to join complex microbial populations in food webs in sediments on the sea floor (Azam and Malfatti, 2007). Upon terrestrialization, for the first time, excreted microorganisms would experience comparative isolation, starvation, desiccation and possibly extinction. Enterococci are among the most persistent microbes of land animal excreta (Sinton et al., 2007), and are well equipped to deal with that challenge (Fig. 2E).
Although difficult to prove, the cumulative data are consistent with a model proposing the emergence of enterococci from ancestors well suited to existence in food webs in sediments (Adams Krumins et al., 2013), including those of icy marine environments; and in marine animal guts into which bile and other host factors that shape the community are secreted. The abundance of psychrophiles among carnobacteria would be consistent with their survival of the global freezes that predate the Cambrian Explosion (Hoffman and Schrag, 2000). The resistance of vagococci to bile (Fig. S2F), as well as relative sensitivity to desiccation and starvation (Fig. 2E), would position them well to live within the marine animals that initially radiated during the Cambrian Explosion. The terrestrialization of animals required, for the first time, that gut flora adapt to cycles of isolation, starvation, and desiccation, until they re-enter the food chain. Enterococci are distinguished from their ancestors, and appear to have been selected for, by virtue of having developed a hardened cell wall (Fig. 2), and the ability to cope with environmental stress (Fig. 2E) – traits that now render them resistant to denaturing solvents, disinfectants, and intrinsically, to many antibiotics (Fig. 2D). These are exactly the traits that enable them to persist in the modern hospital environment (Van Tyne and Gilmore, 2014). Thus, the emergence of enterococci as leading hospital pathogens appears to have been foreordained by events of at least 425 MYA.
STAR Methods
CONTACT FOR REAGENT AND RESOURCE SHARING
Further information and requests for resources and reagents should be directed to and will be fulfilled by the Lead Contact, Michael S. Gilmore (Michael_gilmore@meei.harvard.edu).
EXPERIMENTAL MODEL AND SUBJECT DETAILS
Collection of bacterial strains
Bacterial strains and genome sequences used in this study (Table S1) were obtained from the ATCC bacteriology collection, and other sources as indicated in the Key Resources Table. All strains were routinely grown in brain heart infusion (BHI) at 37C unless noted otherwise, and stored frozen at −80°C in BHI supplemented with 20% glycerol.
Model of colonization for the Galleria mellonella larvae
Galleria mellonella larvae of homogenous size (250–300 mg) and similar development stage (absence of grey marking in the cuticle) were selected from those obtained from a commercial provider (Petco, Cambridge - MA). G. mellonella larvae were then housed in plastic Petri dishes (5 larvae per plate) containing 10 g of pollen (YS Eco Bee Farms) for up to 8 days. Colonization of Galleria larvae by wild type E. faecalis MMH594, or the indicated mariner insertion mutants, was performed as indicated in the Method Details section.
METHOD DETAILS
Illumina and PacBio genome sequencing
Following overnight culture in BHI broth, total DNA was isolated using the Qiagen DNeasy Blood & Tissue Kit and quantified using the Qubit dsDNA HS assay. Genome sequences determined in this study were generated using both Illumina and Pacific Biosciences (PacBio) technology. PacBio libraries were generated following the manufacturer’s recommendations (Procedure and Checklist − 20 kb Template Preparation Using BluePippin Size-Selection v1 and Guidelines for Preparing Size-Selected ~20kb SMRTbell Templates v1), using the DNA Template Prep Kit 2.0 (3Kb – 10Kb) with the following modifications. For each sample, 10 μg of genomic DNA was sheared using Covaris G-tubes at 4800 RPMs. Due to the low shearing size, DNA fragments were purified, DNA damage was end-repaired, and fragments were ligated with SMRTbell sequencing adapters following the manufacturer’s recommendations (Procedure & Checklist - 2 kb Template Preparation and Sequencing v13). SMRTbell sequencing libraries were combined with sequencing primer and polymerase using DNA/Polymerase Binding Kit 2.0 and v2.0.1.2 Sample Prep Calculator. The resulting complex was subjected to PacBio sequencing followed by primary data analysis (version 2 chemistry and 2.0.1 analysis software) on a PacBio-RS instrument and following manufacturer’s recommendations. For Illumina sequencing, both jumping libraries and 180bp paired fragment libraries were prepared using commercial kits (Kapa Biosystems) as previously described (Lebreton et al., 2013a)
Genome assembly and annotation
ALLPATHS-LG assembly was run using default parameters, except that the BIG_MAP option was set to “True,” to take advantage of longer PacBio sequence reads. Statistics from the ALLPATHS-LG hybrid assemblies for all sequenced strains are shown in Table S1. NCBI accession numbers for each genome are listed in Table S1. The genomes of previously sequenced and published Enterococcus, Melissococcus, Tetragenococcus, Vagococcus, Carnobacterium or Lactobacillus species included in our phylogenetic analysis were downloaded from Genbank, bringing the total number of genomes included in the analysis to 32 (Table S1). To assure consistency and to reduce artifacts among the genomes being analyzed, all genomes, including those from Genbank, were annotated or re-annotated in uniform manner using the Broad Institute’s prokaryotic pipeline, with the Enterococcus-specific approach having been described previously (Lebreton et al., 2013a). Briefly, protein-coding genes were predicted with Prodigal (Hyatt et al., 2010), and were filtered to remove genes with >=70% overlap to tRNAs or rRNAs. Gene product names were assigned based on top BLAST hits against the SwissProt protein database (>=70% identity and >=70% query coverage), and protein family profiles were searched against TIGRfam hmmer equivalogs. Additional annotation analyses performed include PFAM, KEGG, and EC. Finally, to investigate the genomic diversity of the different enterococcal species, orthologous genes were identified in all 32 genomes using Synergy2 (http://sourceforge.net/projects/synergytwo/). Orthogroups contain orthologs, which are vertically inherited genes that likely have the same function, and also possibly paralogs, which are duplicated genes that may have different function. There were 526 single copy core orthogroups across our set of 32 strains. The presence of bacterial immunity (i.e. CRISPR/cas and restriction-modification systems), drug resistance, and prophages was determined using available online tools (CRISPRfinder: http://crispr.u-psud.fr/Server/; ResFinder: http://cge.cbs.dtu.dk/services/ResFinder/; Phast: https://cge.cbs.dtu.dk/services/Phast/).
Reconstruction of metabolic pathways
Pathway, and gene enrichment analysis within gene lists were performed using the Database for Annotation, Visualization and Integrated Discovery (DAVID, v6.7, https://david.ncifcrf.gov/). Uniprot Accession numbers were entered to create working gene lists that were analyzed with the KEGG and GOTERM_BP_FAT functions. Results were viewed using DAVID’s Functional Annotation Clustering method. Visualization and representation of pathways and gene functions, were performed by searching the KEGG Pathway Map and Brite hierarchy functions (Kanehisa and Goto, 2000) using preexisting metabolic pathway maps for strains E. faecalis V583 and M. plutonius ATCC 35311.
Phylogenetic tree and gene flow by phylogenetic analysis using parsimony
The phylogenetic tree of the Enterococcus genus was constructed by applying RAxML (Stamatakis, 2006) to a concatenated alignment of single-copy core orthogroups across all organisms (length 487713 bp). Then 1000 bootstrap iterations were calculated using RAxML’s rapid bootstrapping algorithm (Table S6). A phylogenetic analysis using parsimony was performed to reconstruct gene flows throughout the evolution of the Enterococcus genus. Evolutionary gain and loss events for each gene were assigned to positions (nodes) on the SNP based Enterococcus phylogenetic tree by parsimony, or minimization of the number of gain/loss events, using PAUP Version 3.1 (Swofford, 2001). This allowed us to estimate at which points and times in the evolution of the genus each individual gene arose, or was lost, providing a probabilistic view of the genes occurring at each stage of evolution. For a given node, gene gain/loss may represent independent events that similarly arose since divergence from the LCA, and before the next speciation (Supplemental dataset 1).
16S rDNA sequence identification in microbiome samples
Fastq files of SRA records generated by Illumina based sequencing were downloaded in October 2016, using NCBI’s SRA toolkit v2.8.0 (https://trace.ncbi.nlm.nih.gov/Traces/sra). When available, the bioprojects were chosen to sample wild animals from various classes of land or marine ecologies. Of necessity because of the low number of wild samples and despite the potential for contamination with microbes from land, data from freshwater and farm-raised fish microbiota were included to obtain a satisfactory sample size and diversity. The standard UPARSE pipeline (Edgar, 2013) was used to cluster sequences into operational taxonomic units (OTUs) and assign taxonomy. Sequences were first filtered using fastq, with a fastq_maxee parameter of 1.0. Then, filtered sequences were de-replicated and clustered using cluster_otus. Singleton clusters were removed. Taxonomy was assigned to OTUs using the utax algorithm and RDP training set v15, with confidence values for 250 nt length sequences. An OTU table was generated using the usearch_global command, with a sequence identity cutoff of 99%. The presence of Enterococcus or Vagococcus was determined uniformly for each genus: at least one OTU had to be assigned with a minimum confidence value of 0.4. If a sample did not contain any of the genera, it was removed from further analysis. Comparison of presence/absence between environments was performed using Fisher’s Exact or Chi-Square Tests for large sample size (Table S3).
Average Nucleotide Identity and temporal calibration
Orthogroups were used to determine shared gene content in pairwise genome comparisons. For a genome pair (genome A versus genome B), the total number of genes in genome A was determined, and then the number of those shared with genome B (based on shared ortholog group membership) was deduced. Percent shared gene content was calculated by dividing the number of genes common to genomes A and B by the total number of genes in genome A. To calculate percent average nucleotide identity (ANI), shared genes were aligned and the number of identical and different nucleotides determined. Percent ANI was calculated by dividing the number of identical nucleotides in shared genes by the total number of nucleotides. For each node, the mean average nucleotide identity (mANI), and standard deviation, was calculated by averaging the ANI values of species pairs clustering separately on the daughter branches of each node. As an example, mANI of Fig. 1B node 19 was calculated by averaging the ANI values of the following pairwise comparisons: E. gilvus/E. avium, E. gilvus/E. raffinosus and E. gilvus/E. malodoratus. The standard deviation of the ANI values from the obtained mANI was then calculated (Table S6).
Temporal calibration of the enterococcal ANI values was made by calculating the ANI between genomes of species employed in previous studies which dated divergence using previously only 16S rDNA sequence (the divergence of Escherichia coli and Salmonella typhimurium, estimated 120 – 160 million years ago, and E. coli and Vibrio spp. 350 – 450 MYA [Ochman and Wilson, 1987]) or by multilocus sequence comparison (various Aeromonas species from each other 167 and 202 MYA [Lorén et al., 2014]). The following genome sequences were downloaded from GenBank: A) E. coli genomes NC_000913, NC_002695 and NC_022648), which were compared to three genomes of S. typhimurium (NZ_CP014358, NZ_CP007804 and NZ_CP014356), resulting in an average ANI value of 81.2 ± 0.2 %; B) Vibrio species genomes NZ_CGHE00000000, NZ_AWNF00000000 and NZ_AHIH00000000, which were compared to the same 3 E. coli genomes generating an average ANI of 68.0 ± 1.9 %; and C) Aeromonas hydrophila genomes NZ_CP013178, NZ_CP013965 and NZ_CP011100 which were compared to genome sequences for Aeromonas salmonicida, A. media and A. bivalvium (NZ_AGVO00000000, NZ_CP007567 and NZ_CDDA00000000), which generated an average ANI of 78.9 ± 0.8 %.
Phenotype assessment
Ability to grow on commercially available enterococcal selective media (Enterococcosel® agar, BBL) and BBL CHROMagar® orientation media was investigated at 30°C or 37°C for 24h. All enterococcal species (with the exception of T. halophilus and E. italicus, for which we only possessed corresponding genome sequences) were grown on BUG+B agar plates at 30°C and analyzed by Biolog Phenotype Microarray (PM 1,2 and 9–20, inoculating fluids IF0a GN/GP and IF-10b GN/GP). Rather than using the manufacturer’s redox dye readout, optical density at 590 nm (OD590) was monitored hourly for 24 h, using a Synergy 2 microplate reader (Bio-Tek). Growth was quantified by measuring the area under the curve (AUC), which was calculated by summing individual OD590 values of each well. For the carbon metabolic profiles, an AUC with a value higher than 3.0 (compared to 2.53 ± 0.34 for the negative control well) was considered positive growth. For the identification of phenotypes shared by Enterococcus spp. but distinct from the outgroup species, we averaged the AUC values for each of 1344 conditions measured for every Enterococcus spp., and compared that to the corresponding average AUC of the outgroup species (Fig. S2E). Those conditions for which the AUC ratio was < 0.5, or > 2 were considered of possible biological relevance, and assessed by Mann–Whitney U test for statistical significance (Table S5).
To investigate resistance to desiccation and starvation for enterococcal and outgroup strains, we collected and included two additional strains to add to the genome-sequenced outgroup strains. Unlike V.lutrae LBD1 which was isolated from the intestine of a wild largemouth bass (Lebreton et al., 2013b), Vagococcus fluvialis strain DIV0098 was isolated from sediment sample collected in a salt water fish tank at the New England Aquarium and was identified by 16S sequence analysis (99.3% identity with V. fluvialis). L. garviae strain DIV0709 (100% 16S identity with L. garviae) was isolated from a bivalve mollusk (Pteriomorphia) collected in the Cape Cod Bay in Massachusetts. Bacterial cells from overnight culture in BHI broth were washed twice and suspended in PBS to a final optical density of 1.0. For the starvation experiment, 200 μl of cell suspensions were distributed in a microtiter plate that was sealed with sterile aluminum films and incubated in the dark and at room temperature. CFU counts were monitored by track dilutions on BHI agar plates at T0 and after 10 days of incubation. Survival was calculated by normalizing the surviving cells with the inoculum size for each strain. For the desiccation experiments, 100 μl of cell suspensions were distributed in a 96 well microtater plate, the inoculum were determined by CFU counts and plates were air-dried during 4 hours inside a laminar flow hood at room temperature. After 5 days of incubation in the dark, at room temperature and with a relative humidity ranging from 35 to 45%, CFU counts were monitored and percentage of survival were calculated as described above. Experiments were performed in three independent replicates for each strains and the average value was compared to other species. Statistical significance was estimated using Student’s t-test.
Fossil and phylogenetic data
Fossil occurrence data were downloaded from the most recent version of the Paleobiology Database (www.paleobiodb.org) by selecting the ‘family level’, ‘order level’, and ‘geological epochs’ options (last accessed February 25th, 2016). A total of 1,284,138 fossil occurrences (representing 8172 families and 60264 genera) were downloaded, and the data were curated by checking for synonyms among all taxa. Diversity of land animals was calculated using the “range through” method, which assumes that a family was present in all time intervals between its first and last appearance, even if not directly sampled at intervening points.
Quantitative term analysis to identify natural host associations and ecologies
NCBI PubMed and Web of Science databases (last accessed 08/2016) were searched using the string “(((((enterococcus <species>) AND <ecology>) NOT infection) NOT virulence) NOT pathogenesis)”, and results were compiled (Table S2). Host-Enterococcus associations were assigned when at least two different publications were identified. Host associations are indicated by icons included in Fig. 1.
Construction of transposon insertion mutant library in E. faecalis MMH594
A tetracycline resistance version (named pZXL5Tet) of the nisin-inducible mariner transposon harbored by delivery vector pZXL5 (Zhang et al., 2012) was generated by replacing its gentamicin resistance cassette with the tetracycline resistance gene of p3Tet (Hancock and Perego, 2004). Specifically, the PCR amplified tetracycline resistance gene was cloned into pZXL5 between the PstI and KpnI sites in the transposable element. This generated a tetracycline resistance gene suitable for selection of insertion mutants in the high-level aminoglycoside resistant E. faecalis MMH594 strain. For generation of a mutant library, the resulting modified transposon delivery vector was transformed into E. faecalis MMH594 strain as described (Shepard and Gilmore, 1995). Tetracycline-resistant electrotransformants were grown overnight at the permissive temperature of 28°C in BHI supplemented with 5 μg/mL tetracycline and 10 μg/mL chloramphenicol (to select for the delivery vector). Then, 100 μl of this overnight culture was used to inoculate 100 ml of BHI pre-warmed to the non-permissive temperature of 37°C, and supplemented with 5 μg/mL tetracycline as well as 25 ng/mL nisin (to induce the expression of the mariner transposase and, thus, transposition of the element), and grown overnight with shaking at 150 rpm. To deplete cells of the plasmid backbone, 100 μl of this culture was transferred to 100 ml of fresh pre-warmed BHI broth supplemented with 5 μg/mL tetracycline, and cultured overnight without nisin. Cells were collected by centrifugation, resuspended in 20 ml BHI containing 20% glycerol, divided into 2 ml aliquots and frozen at −80°C (MMH594 Transposon library, as listed in Key resources table).
Sequencing and determination of the complexity of the mutant pool was performed using commercially available kits (Terminal deoxylnucleotidyl Transferase [Promega], Easy-A High fidelity PCR enzyme [Agilent] and Performa spin columns [EdgeBio]) as previously described (Valentino et al., 2014)). Sequencing was performed on two separate replicates that were multiplexed and sequenced in parallel for 51 cycles in a single end sequencing reaction on a single lane of an Illumina HiSeq 2000 (Broad Institute) using a custom sequencing and standard Illumina index sequencing primers (sequences of primers pZXL5Tet TnSeq1, olj511, olj512 and olj373 are available in the Key resource table). Mapping of transposon-genome junction sequence reads to the MMH594 (https://olive.broadinstitute.org/strains/ente_faec_b594.2) genome was carried out utilizing custom scripts and programs on the Tufts University Galaxy server as described (Valentino et al., 2014). Following removal of adapter and transposon sequence, genomic sequences with a minimum read length of 25 nucleotides were aligned to the E. faecalis MMH594 genome using bowtie with its default settings. The resulting bowtie output file was then used as input for a custom script named “hopcount”. Hopcount tabulates the number of times individual insertion amplicons were re-sequenced. An excel spreadsheet was generated that indicates, for each insertion site, its position within the genome, the gene locus to which that position maps, the strand (positive vs. negative) associated with the site, as well as the number of each amplicon species read. Hopcount output was used to determine the complexity of the MMH594 transposon library (Fig. S4D), which identified 60009 unique transposon insertions in the MMH594 pool, which were evenly distributed throughout the 3.2 Mb genome.
Uric acid metabolism by E. faecalis
A carbon limited M9 (1X, Amresco) broth supplemented with 0.06 % yeast extract (Difco) was used to investigate uric acid metabolism. Anaerobic growth of E. faecalis was monitored by optical density (OD) at 590 nm following up to 96 h incubation at 37°C, in the presence or absence of added glucose (0.5% w/v) or uric acid (0.2% w/v) (Fig. S4B). Uric acid (Alfa Aesar) was solubilized as 2% solution in 0.5 N NaOH and added to the medium, which was neutralized to pH 7.2 prior to inoculation. Uric acid was measured spectrophotometrically by change in absorbance at 292 nm before and after treatment with porcine liver uricase (Sigma - U9375). NH4+ was determined enzymatically by NH4+-dependent oxidation of NADPH (measured by absorbance at 340 nm) by beef liver glutamate dehydrogenase (Sigma - AA0100). Uric acid and NH4+ in growth media were assayed after removal of cells by centrifugation. All measurements were performed in triplicate and mean values and standard errors are shown (Fig. S4C).
To screen enterococcal species and 9120 arrayed mutants from the E. faecalis MMH594 transposon library, 300 μL agar plugs were prepared in 96 microtiter plates using a medium consisting of: 1% tryptone (Difco); 0.5 % yeast extract (Difco); 0.05% glucose; 0.0012 % phenol red, and 1.5% agar; pH 7.2; with or without 0.2% (w/v) of uric acid (Mead, 1974). The agar plugs were inoculated from 37°C overnight cult ures in BHI, with a central puncture by a 96 pin replicator. Plates were incubated anaerobically at 37°C for up to 4 days, and color change for the metabolism of glucose (acidification to pH = 5.1, yellow coloration after 24 h at 37°C) and for the metabolism of uric acid (alkalini zation to pH = 9.2, red coloration after 48 to 72h at 37°C) was monitored.
Inverse PCR was performed on all E. faecalis MMH594 transposon insertion mutants unable to alkalinize the medium in the presence of urate, followed by nucleotide sequence determination from both ends, to identify the site of insertion and the gene disrupted by the transposon. We identified insertions in genes EF1390, EF1613, EF2559 (6 distinct insertions), EF2562 (2 distinct insertions), EF2570 (2 distinct insertions), EF2578, EF2579 (two distinct insertions), EF2582, EF3046 and EF3277. The genomes of mutants ΩEF2559 and Ω, with centrally located insertions, were re-sequenced to confirm that transposon insertion was the only genetic difference with the WT strain prior to further phenotypic testing.
Model of colonization for the Galleria mellonella larvae
Prior to testing, G. mellonella from our supplier was found to be naturally colonized by E. faecalis (Fig. S4F), when homogenized as previously described (Champion et al., 2009) and plated onto bile esculin azide agar (Enterococcosel®, BBL). Because this indigenous E. faecalis strain was found to be sensitive to tetracycline, larvae were orally injected, at day one and day two, with 10 μL of tetracycline (100 μg/mL) (Fig. S4F). Larvae were then allowed to feed on un-inoculated pollen for 24 hours prior to testing for decreases in natural enterococcal load (Fig. S4F). Cells from overnight cultures of wild type E. faecalis MMH594, or the indicated mariner insertion mutants, were collected by centrifugation and washed one time in PBS prior to resuspension in PBS and mixing with fresh bee pollen to obtain a final concentration of 5×107 CFU/g of pollen. G. mellonella larvae were then housed in plastic Petri dishes (5 larvae per plate) containing 10 g of the E. faecalis-pollen mix pollen for 8 days. Pollen was sampled periodically to insure equal presence of wild type and mutant. Larvae were then moved into fresh Petri dishes with 10 g of un-inoculated pollen for 2 days. Following 10 days, 10 groups of 5 live larvae were washed (90 s in 70% EtOH), homogenized, and CFU counts were performed on either BEA (to detect residual indigenous E. faecalis), or BEA supplemented with 500 μg/mL of gentamicin to confirm the colonization with the high-level gentamicin resistant E. faecalis MMH594 (Fig. S4F). To enumerate transposon insertion mutants, colonies were then tested on BEA supplemented with 5 μg/ml tetracycline. As a negative control, an arbitrarily chosen transposon mutant possessing an insertion in ef1199 of no known relevance to uric acid production or gut colonization, and phenotypically shown to metabolize uric acid, was tested in competition with WT E. faecalis MMH594 and no difference was observed (Fig. S4G).
QUANTIFICATION AND STATISTICAL ANALYSIS
Data are presented as mean ± SEM unless otherwise indicated in figure legends. Sample number (n) indicates the number of independent biological samples in each experiment. Sample numbers and experimental repeats are indicated in figures and figure legends or methods section above. Data were analyzed using the two-sided nonparametric Mann-Whitney U test of the null hypothesis of continuous data unless otherwise indicated in figure legends or method details. Differences in means were considered statistically significant at p < 0.05. Significance levels are: * p < 0.05; ** p < 0.01; *** p < 0.001; **** p < 0.0001; n.s., non-significant. Analyses were performed using the Graphpad Prism 7.0a software.
DATA AND SOFTWARE AVAILABILITY
Supplemental dataset
Accession number for whole genome sequences
L. garvieae NCIMB 31208, ASWT00000000; E. moraviensis BAA-383, ASWB00000000; E. haemoperoxidus BAA-382, ASVY00000000; E. caccae BAA-1240, ASVV00000000; E. dispar ATCC51266, ASWK00000000; E. asini ATCC700915, ASVU00000000; E. cecorum ATCC43198, ASWI00000000; E. columbae ATCC51263, ASWJ00000000; E. sulfureus ATCC49903, ASWO00000000; E. saccharolyticus ATCC43076, ASWN00000000; E. pallens BAA-351, ASWD00000000; E. gilvus BAA-350, ASWH00000000; E. avium ATCC14025, ASWL00000000; E. raffinosus ATCC49464, ASWF00000000; E. malodoratus ATCC43197, ASWA00000000; E. phoeniculicola BAA-41, ASWE00000000; E. mundtii ATCC882, ASWC00000000; E. durans ATCC6056 ASWM00000000; E. hirae ATCC8043, ASVZ00000000; and E. villorum ATCC70091, ASWG00000000.
ADDITIONAL RESOURCES
MMH594 genome sequence (https://olive.broadinstitute.org/strains/ente_faec_b594.2)
Paleobiology Database (www.paleobiodb.org)
NCBI’s SRA toolkit v2.8.0 (https://trace.ncbi.nlm.nih.gov/Traces/sra)
Synergy2 orthogrouping: http://sourceforge.net/projects/synergytwo/
CRISPRfinder: http://crispr.u-psud.fr/Server/
ResFinder: http://cge.cbs.dtu.dk/services/ResFinder/
Supplementary Material
A: 16S rDNA based phylogeny of the 49 previously known species that constitute the Enterococcus genus. The tree (Jukes-Cantor model, neighbor-joining method) was rooted using the 16S sequence of V. lutrae. Species indicated in blue were selected for sequencing and analysis. B: 16S based phylogeny of members of the Enterococcaceae and Carnobacteriaceae families (modified from existing tree of life (Yarza et al., 2008)). Closely related outgroup species of Vagococcus and Carnobacterium used in this study are indicated. Lactococcus garviae is omitted from the figure as the lineage that gives rise to it is not ancestral to the enterococci.
A: Distribution of the shared (common to all enterococci and outgroup species, and used to construct the phylogeny in figure 1B), the auxiliary and the species unique genes in study genomes. B: Gene accumulation and rarefaction curves for the genus Enterococcus. C: COG functional categories for the genus core (presence in >90 % of Enterococcus spp.), ubiquitous, genus related and genus unique genes. Ubiquitous genes are those found in 81 – 100 of the comparator genomes that span the Eubacteria (Fig. 2A), and are included for reference. D: Circular representation of the reference chromosome of E. faecalis V583, showing the location of core (n=1037), ubiquitous (n=378), shared (n=235), and unique (n=10) genes of Enterococcus spp. Mobile genetic islands predicted in E. faecalis V583 (Paulsen et al., 2003) are indicated by yellow highlight. E: Distribution of 1344 phenotype results by comparing average enterococcal species growth to average outgroup species growth. 45 conditions, represented in Fig. 2D, yielded to significantly more growth for Enterococcus spp. compared to the outgroup species. F: Growth of Vagococcus lutrae LBD1 on commercial enterococcal selective media (top, CHROMagar™; bottom, Enterococcosel™). G: Individual growth curves for all Enterococcus and outgroup species in the presence of sodium azide or chloroxylenol using the Biolog phenotypic microarrays (PM). PM plate number and well coordinates are indicated. Representative species for each phylogenetic group are shown. H: Circular map of the E. pallens BAA-351 genome showing genes of E. pallens coded in the clockwise (black) and anti-clockwise (grey) direction (outer circle) as illustrated. The 13 scaffolds of the draft genome are represented with black arrows (inner circle). Enterococcus genus core genes are shown in green, whereas the genes gained at node 18 and at the E. pallens leaf (Fig. 3A) are shown in blue.
Normalized distribution of annotated COG functional categories for niche specifying genes A. gained or B. lost at the node or leaf of each representative species (Fig. 3A). Node or leaf numbers are indicated (italicized) in front of the related species. C. gained or D. lost at the nodes of the Enterococcus phylogenetic tree (Fig. 3A).
A: Enterococcus spp. carbohydrate fermentation profiles (orange, growth; blue, no growth). Species are organized by phylogenetic tree position (Fig 1A). B: Growth of E. faecalis strain MMH594 in medium with no carbon source, uric acid or glucose. Data represent three independent experiments. Means and standard deviations are represented. C: Growth of E. faecalis MMH594 in medium supplemented with 0.5% glucose and 0.2% uric acid for 96 h at 37°C, under anaerobic conditions. Biomass (OD at 600nm, circles) Uric acid consumption (squares) and ammonia production (triangles) in the culture fluid were monitored over time. Similar phenotype was observed for negative control transposon mutant (EF1199) while both mutants in genes EF2570 and EF2559 were unable to metabolize uric acid in vitro (data not shown). D: Complexity and randomness of the mariner transposon insertion library in E. faecalis MMH594. Cumulative number of unique insertions throughout the 3.2 Mb chromosome (left panel), and distribution on the circular genomic map (right panel). E: Predicted KEGG pathway reconstruction of purine related metabolism in E. faecalis. Black arrows indicate identifiable genes associated with enzymatic activity in E. faecalis; grey arrows indicate no identifiable gene yet associated with this enzymatic activity. Genes interrupted by transposon insertion in E. faecalis mutants deficient in uric acid metabolism are indicated by locus tag numbers based on the E. faecalis V583 genome: in blue, gene gained since speciation of E. faecalis (Fig. 3A, node 4); in green, genes gained since the divergence from the last common ancestor shared with M. plutonius (Fig. 3A node 3); in black, genes shared by diverse enterococci species. F: Gut colonization of Galleria mellonella larvae. Indigenous E. faecalis before and after treatment with tetracycline (left panel). Gut colonization by E. faecalis MMH594 following one week of ad libitum feeding on pollen containing 5×107 CFU/g of pollen (right panel). G: Competitive index for growth in vitro (BHI), and for GI tract colonization of G. mellonella by wild type E. faecalis MMH594 (WT) and negative control transposon mutant (EF1199).
Each dot represents a pairwise comparison of a given species with all other species representatives in our study. Relevant comparator species are labeled.
Table S1. Strains and genomes metadata, Related to Figure 1
Table S2. Quantitative analysis of the Medline and Web of Science databases for reports of Enterococcus spp. in various ecologies, Related to Figure 1
Table S3. Prevalence of Enterococcus spp. and Vagococcus spp. in terrestrial and aquatic animals by 16S rDNA gut microbiome analysis, Related to Figure 1
Table S4. Analysis of Enterococcus spp. core genes. Related to Figure 2
Tab 1: Ubiquity score of 1037 Enterococcus core genes (>90% species) in other eubacteria.
Tab 2: Database of 100 genomes representative of families across the eubacterial domain.
Tab 3: Enterococcus core genes absent in Vagococcus lutrae LBD1.
Table S5. High-throughput phenotypic analysis using Biolog, Related to Figure 2
Supplemental dataset 1: Gene flux predicted by phylogenetic analysis using parsimony, Related to Figure 3.
Highlights.
Emergence of enterococci as hospital pathogens was foreordained by Paleozoic events
Commensal enterococci emerged from ancestors adapted to marine hosts and environments
Terrestrialization selected for a hardened cell wall leading to antibiotic resistance
Enterococcal speciation: energy source driven, parallels host radiation or extinction
Acknowledgments
The authors thank Richard Losick, Daria Van Tyne, Melissa Martin, Andrew Knoll, Eric Alm, Gregory Fournier and other members of the Gilmore Lab for helpful discussions during the development of this project and preparation of the manuscript. The authors thank Charlie Innis, Meghan May and Julia Schwartzman for the collection and/or typing of marine isolates. This project has been funded in part by DHHS/NIH/NIAID grants AI072360 and AI083214 (Harvard-wide Program on Antibiotic Resistance) (MSG); and contract HHSN272200900018C and grant U19AI110818 to the Broad Institute. The authors declare no competing financial interests.
Footnotes
Author contributions
M.S.G. conceived and planned the study. A.M.E. organized and managed genome sequencing, and contributed to project development and management. F.L. facilitated project management. F.L., A.L.M. and T.J.S. contributed expertise and performed the bioinformatic analyses. J.T.S. performed BioLog phenotypic screens. F.L. constructed the E. faecalis transposon library and performed related bioinformatic and phenotypic analyses, including intestinal colonization model of G. mellonella, and compiled all data. F.L., A.L.M., A.M.E. and M.S.G. interpreted and discussed the scientific findings. F.L. and M.S.G. wrote the paper with input from all authors.
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- Aarestrup FM, Butaye P, Witte W. Nonhuman Reservoirs of Enterococci. In: Gilmore M, editor. The Enterococci. American Society of Microbiology; 2002. pp. 55–99. [Google Scholar]
- Adams Krumins J, van Oevelen D, Martijn Bezemer T, De Deyn GB, Gera Hol WH, van Donk E, de Boer W, de Ruiter PC, Middelburg JJ, Monroy F, et al. Soil and Freshwater and Marine Sediment Food Webs: Their Structure and Function. BioScience. 2013;63:35–42. [Google Scholar]
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215:403–410. doi: 10.1016/S0022-2836(05)80360-2. [DOI] [PubMed] [Google Scholar]
- Arias CA, Murray BE. The rise of the Enterococcus: beyond vancomycin resistance. Nat Rev Microbiol. 2012;10:266–278. doi: 10.1038/nrmicro2761. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Azam F, Malfatti F. Microbial structuring of marine ecosystems. Nat Rev Microbiol. 2007;5:782–91. doi: 10.1038/nrmicro1747. Review. Erratum in: Nat Rev Microbiol. (2007). 5:966. [DOI] [PubMed] [Google Scholar]
- Baele M, Devriese LA, Butaye P, Haesebrouck F. Composition of enterococcal and streptococcal flora from pigeon intestines. J Appl Microbiol. 2002;92:348–351. doi: 10.1046/j.1365-2672.2002.01537.x. [DOI] [PubMed] [Google Scholar]
- Bailey L. Melissococcus pluton, the cause of European foulbrood of honey bees (Apis spp.) J Appl Bacteriol. 1983;55:65–69. [Google Scholar]
- Benton MJ, Twitchett RJ. How to kill (almost) all life: the end-Permian extinction event. Trends Ecol Evol. 2003;18:358–365. [Google Scholar]
- Benton MJ, Alfaro ME, Santini F, Brock C, Alamillo H, Dornburg A, Rabosky DL, Carnevale G, Harmon LJ, Alroy J, et al. The origins of modern biodiversity on land. Philos Trans R Soc Lond B Biol Sci. 2010;365:3667–3679. doi: 10.1098/rstb.2010.0269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bourgogne A, Garsin DA, Qin X, Singh KV, Sillanpaa J, Yerrapragada S, Ding Y, Dugan-Rocha S, Buhay C, Shen H, et al. Large scale variation in Enterococcus faecalis illustrated by the genome analysis of strain OG1RF. Genome Biol. 2008;9:R110. doi: 10.1186/gb-2008-9-7-r110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Borgo F, Ferrario C, Ricci G, Fortina MG. Genotypic intraspecies heterogeneity of Enterococcus italicus: data from dairy environments. J Basic Microbiol. 2013;53:20–28. doi: 10.1002/jobm.201100464. [DOI] [PubMed] [Google Scholar]
- Buller NB. Chapter 1: Aquatic Animal Species and Organism Relationship. Subchapters 1.35 Carnobacteriaceae and 1.36 Enterococcaceae. In: Buller NB, editor. Bacteria and Fungi from Fish and other Aquatic Animals: a practical identification manual. 2. CAB International; 2014. pp. 400–406. [Google Scholar]
- Bursell E. The Excretion of Nitrogen in Insects. Adv In Insect Phys. 1967;4:33–67. [Google Scholar]
- Cai J, Collins MD. Evidence for a close phylogenetic relationship between Melissococcus pluton, the causative agent of European foulbrood disease, and the genus Enterococcus. Int J Syst Bacteriol. 1994;44:365–367. doi: 10.1099/00207713-44-2-365. [DOI] [PubMed] [Google Scholar]
- Champion OL, Cooper IAM, James SL, Ford D, Karlyshev A, Wren BW, Duffield M, Oyston PCF, Titball RW. Galleria mellonella as an alternative infection model for Yersinia pseudotuberculosis. Microbiology. 2009;155:1516–1522. doi: 10.1099/mic.0.026823-0. [DOI] [PubMed] [Google Scholar]
- Cheng L, Connor TR, Sirén J, Aanensen DM, Corander J. Hierarchical and spatially explicit clustering of DNA sequences with BAPS software. Mol Biol Evol. 2013;30:1224–1228. doi: 10.1093/molbev/mst028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Collins MD, Farrow JAE, Phillips BA, Ferusu S, Jones D. Classification of Lactobacillus divergens, Lactobacillus piscicola, and Some Catalase-Negative, Asporogenous, Rod-Shaped Bacteria from Poultry in a New Genus, Carnobacterium. Int J Syst Bacteriol. 1987;37:310–316. [Google Scholar]
- Collins MD, Ash C, Farrow JA, Wallbanks S, Williams AM. 16S ribosomal ribonucleic acid sequence analyses of lactococci and related taxa. Description of Vagococcus fluvialis gen. nov., sp. nov. J. Appl. Bacteriol. 1989;67:453–460. doi: 10.1111/j.1365-2672.1989.tb02516.x. [DOI] [PubMed] [Google Scholar]
- Collins MD, Williams AM, Wallbanks S. The phylogeny of Aerococcus and Pediococcus as determined by 16S rRNA sequence analysis: description of Tetragenococcus gen. nov. FEMS Microbiol Lett. 1990;70:255–262. doi: 10.1016/s0378-1097(05)80004-7. [DOI] [PubMed] [Google Scholar]
- Courvalin P. Transfer of antibiotic resistance genes between gram-positive and gram-negative bacteria. Antimicrob Agents Chemother. 1994;38:1447–1451. doi: 10.1128/aac.38.7.1447. Review. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Edgar RC. UPARSE: Highly accurate OTU sequences from microbial amplicon reads. Nat Methods. 2013;10:996–998. doi: 10.1038/nmeth.2604. [DOI] [PubMed] [Google Scholar]
- Franzmann PD, Höpfl P, Weiss N, Tindall BJ. Psychrotrophic, lactic acid-producing bacteria from anoxic waters in Ace Lake, Antarctica; Carnobacterium funditum sp. nov. and Carnobacterium alterfunditum sp. nov. Arch Microbiol. 1991;156:255–262. doi: 10.1007/BF00262994. [DOI] [PubMed] [Google Scholar]
- Gaca AO, Abranches J, Kajfasz JK, Lemos JA. Global transcriptional analysis of the stringent response in Enterococcus faecalis. Microbiology. 2012;158:1994–2004. doi: 10.1099/mic.0.060236-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gilmore MS, Lebreton F, van Schaik W. Genomic transition of enterococci from gut commensals to leading causes of multidrug-resistant hospital infection in the antibiotic era. Curr Opin Microbiol. 2013;16:10–16. doi: 10.1016/j.mib.2013.01.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hancock LE, Perego M. Systematic inactivation and phenotypic characterization of two-component signal transduction systems of Enterococcus faecalis V583. J Bacteriol. 2004;186:7951–7958. doi: 10.1128/JB.186.23.7951-7958.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoffman PF, Schrag DP. Snowball Earth. Sci Am. 2000;282:68–75. [Google Scholar]
- Hyatt D, Chen GL, Locascio PF, Land ML, Larimer FW, Hauser LJ. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics. 2010;11:119. doi: 10.1186/1471-2105-11-119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Isenberg HD, Goldberg D, Sampson J. Laboratory studies with a selective Enterococcus medium. Appl Microbiol. 1970;20:433–436. doi: 10.1128/am.20.3.433-436.1970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jordan S, Hutchings MI, Mascher T. Cell envelope stress response in Gram-positive bacteria. FEMS Microbiol Rev. 2008;32:107–146. doi: 10.1111/j.1574-6976.2007.00091.x. [DOI] [PubMed] [Google Scholar]
- Kanehisa M, Goto S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000;28:27–30. doi: 10.1093/nar/28.1.27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Konstantinidis KT, Tiedje JM. Genomic insights that advance the species definition for prokaryotes. Proc Natl Acad Sci. 2005;102:2567–2572. doi: 10.1073/pnas.0409727102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lam MMC, Seemann T, Bulach DM, Gladman SL, Chen H, Haring V, Moore RJ, Ballard S, Grayson ML, Johnson PDR, et al. Comparative analysis of the first complete Enterococcus faecium genome. J Bacteriol. 2012;194:2334–2341. doi: 10.1128/JB.00259-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lebreton F, van Schaik W, McGuire AM, Godfrey P, Griggs A, Mazumdar V, Corander J, Cheng L, Saif S, Young S, et al. Emergence of epidemic multidrug-resistant Enterococcus faecium from animal and commensal strains. MBio. 2013a;4:e00534–13. doi: 10.1128/mBio.00534-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lebreton F, Valentino M, Duncan L, Zeng Q, Manson Mcguire A, Earl A, Gilmore M. High-Quality Draft Genome Sequence of Vagococcus lutrae Strain LBD1, Isolated from the Largemouth Bass Micropterus salmoides. Genome Announc. 2013b;1:e01087–13. doi: 10.1128/genomeA.01087-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leisner JJ, Hansen MA, Larsen MH, Hansen L, Ingmer H, Sørensen SJ. The genome sequence of the lactic acid bacterium, Carnobacterium maltaromaticum ATCC 35586 encodes potential virulence factors. Int J Food Microbiol. 2012;152:107–115. doi: 10.1016/j.ijfoodmicro.2011.05.012. [DOI] [PubMed] [Google Scholar]
- Lorén JG, Farfán M, Fusté MC, Barraclough T, Nee S, Nee S, Butlin RTMCSN, Debelle A, Kerth C, Snook R, et al. Molecular Phylogenetics and Temporal Diversification in the Genus Aeromonas Based on the Sequences of Five Housekeeping Genes. PLoS One. 2014;9:e88805. doi: 10.1371/journal.pone.0088805. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martin JD, Mundt JO. Enterococci in insects. Appl Microbiol. 1972;24:575–580. doi: 10.1128/am.24.4.575-580.1972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mead GC. Anaerobic utilization of uric acid by some group D streptococci. J Gen Microbiol. 1974;82:421–423. doi: 10.1099/00221287-82-2-421. [DOI] [PubMed] [Google Scholar]
- Michel C, Pelletier C, Boussaha M, Douet DG, Lautraite A, Tailliez P. Diversity of lactic acid bacteria associated with fish and the fish farm environment. Appl Environ Microbiol. 2007;73:2947–2955. doi: 10.1128/AEM.01852-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moeller AH, Caro-Quintero A, Mjungu D, Georgiev AV, Lonsdorf EV, Muller MN, Pusey AE, Peeters M, Hahn BH, Ochman H. Cospeciation of gut microbiota with hominids. Science. 2016;353:380–382. doi: 10.1126/science.aaf3951. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mundt JO. Occurrence of enterococci in animals in a wild environment. Appl Microbiol. 1963;11:136–140. doi: 10.1128/am.11.2.136-140.1963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nicholson WL, Krivushin K, Gilichinsky D, Schuerger AC. Growth of Carnobacterium spp. from permafrost under low pressure, temperature, and anoxic atmosphere has implications for Earth microbes on Mars. Proc Natl Acad Sci U S A. 2012;110:666–671. doi: 10.1073/pnas.1209793110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nothaft H, Szymanski CM. Protein glycosylation in bacteria: sweeter than ever. Nat Rev Microbiol. 2010;8:765–778. doi: 10.1038/nrmicro2383. [DOI] [PubMed] [Google Scholar]
- Ochman H, Wilson AC. Evolution in bacteria: Evidence for a universal substitution rate in cellular genomes. J Mol Evol. 1987;26:74–86. doi: 10.1007/BF02111283. [DOI] [PubMed] [Google Scholar]
- Ochman H, Elwyn S, Moran NA. Calibrating bacterial evolution. Proc Natl Acad Sci U S A. 1999;96:12638–12643. doi: 10.1073/pnas.96.22.12638. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Okumura K, Arai R, Okura M, Kirikae T, Takamatsu D, Osaki M, Miyoshi-Akiyama T. Complete genome sequence of Melissococcus plutonius ATCC 35311. J Bacteriol. 2011;193:4029–4030. doi: 10.1128/JB.05151-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Palmer KL, Gilmore MS. Multidrug-resistant enterococci lack CRISPR-cas. MBio. 2010;1:e00227–10. doi: 10.1128/mBio.00227-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Palmer KL, Godfrey P, Griggs A, Kos VN, Zucker J, Desjardins C, Cerqueira G, Gevers D, Walker S, Wortman J, et al. Comparative genomics of enterococci: Variation in Enterococcus faecalis, clade structure in E. faecium, and defining characteristics of E. gallinarum and E. casseliflavus. MBio. 2012;3:e00318–11. doi: 10.1128/mBio.00318-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paulsen IT, Banerjei L, Myers GSa, Nelson KE, Seshadri R, Read TD, Fouts DE, Eisen Ja, Gill SR, Heidelberg JF, et al. Role of mobile DNA in the evolution of vancomycin-resistant Enterococcus faecalis. Science. 2003;299:2071–2074. doi: 10.1126/science.1080613. [DOI] [PubMed] [Google Scholar]
- Pikuta EV, Marsic D, Bej A, Tang J, Krader P, Hoover RB. Carnobacterium pleistocenium sp. nov., a novel psychrotolerant, facultative anaerobe isolated from permafrost of the Fox Tunnel in Alaska. Int J Syst Evol Microbiol. 2005;55:473–478. doi: 10.1099/ijs.0.63384-0. [DOI] [PubMed] [Google Scholar]
- Podulka S. Handbook of bird biology (Cornell Laboratory of Ornithology) 2004. [Google Scholar]
- Raven KE, Reuter S, Gouliouris T, Reynolds R, Russell JE, Brown NM, Török ME, Parkhill J, Peacock SJ. Genome-based characterization of hospital-adapted Enterococcus faecalis lineages. Nat Microbiol. 2016;1:15033. doi: 10.1038/nmicrobiol.2015.33. [DOI] [PubMed] [Google Scholar]
- Rota-Stabelli O, Daley AC, Pisani D. Molecular timetrees reveal a Cambrian colonization of land and a new scenario for ecdysozoan evolution. 2013;23:392–398. doi: 10.1016/j.cub.2013.01.026. [DOI] [PubMed] [Google Scholar]
- Sahm DF, Kissinger J, Gilmore MS, Murray PR, Mulder R, Solliday J, Clarke B. In vitro susceptibility studies of vancomycin-resistant Enterococcus faecalis. Antimicrob Agents Chemother. 1989;33:1588–1591. doi: 10.1128/aac.33.9.1588. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schloissnig S, Arumugam M, Sunagawa S, Mitreva M, Tap J, Zhu A, Waller A, Mende DR, Kultima JR, Martin J, et al. Genomic variation landscape of the human gut microbiome. Nature. 2013;3:45–50. doi: 10.1038/nature11711. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schuerger AC, Nicholson WL. Twenty-Three Species of Hypobarophilic Bacteria Recovered from Diverse Ecosystems Exhibit Growth under Simulated Martian Conditions at 0.7 kPa. Astrobiology. 2016;16:335–347. doi: 10.1089/ast.2015.1394. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- Shankar N, Baghdayan AS, Gilmore MS. Modulation of virulence within a pathogenicity island in vancomycin-resistant Enterococcus faecalis. Nature. 2002;417:746–750. doi: 10.1038/nature00802. [DOI] [PubMed] [Google Scholar]
- Shepard BD, Gilmore MS. Electroporation and efficient transformation of Enterococcus faecalis grown in high concentrations of glycine. Methods Mol Biol. 1995;47:217–226. doi: 10.1385/0-89603-310-4:217. [DOI] [PubMed] [Google Scholar]
- Sinton LW, Braithwaite RR, Hall CH, Mackenzie ML. Survival of indicator and pathogenic bacteria in bovine feces on pasture. Appl Environ Microbiol. 2007;73:7917–7925. doi: 10.1128/AEM.01620-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stamatakis A. RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics. 2006;22:2688–2690. doi: 10.1093/bioinformatics/btl446. [DOI] [PubMed] [Google Scholar]
- Srivastava M, Mallard C, Barke T, Hancock LE, Self WT. A selenium-dependent xanthine dehydrogenase triggers biofilm proliferation in Enterococcus faecalis through oxidant production. J Bacteriol. 2011;193:1643–1652. doi: 10.1128/JB.01063-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Svanevik CS, Lunestad BT. Characterisation of the microbiota of Atlantic mackerel (Scomber scombrus) Int J Food Microbiol. 2011;151:164–170. doi: 10.1016/j.ijfoodmicro.2011.08.016. [DOI] [PubMed] [Google Scholar]
- Swofford DL. Version 4.0 beta version Phylogenetic Analysis Using Parsimony PAUP* 4.0 beta version disclaimer and user agreement. 2001 http://www.paup.sc.fsu.edu/
- Valentino MD, Foulston L, Sadaka A, Kos VN, Villet RA, Santa Maria J, Lazinski DW, Camilli A, Walker S, Hooper DC, et al. Genes Contributing to Staphylococcus aureus Fitness in Abscess- and Infection-Related Ecologies. MBio. 2014;5:e01729–14. doi: 10.1128/mBio.01729-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van Tyne D, Gilmore MS. Friend Turned Foe: Evolution of Enterococcal Virulence and Antibiotic Resistance. Annu Rev Microbiol. 2014:337–356. doi: 10.1146/annurev-micro-091213-113003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Venter JC, Remington K, Heidelberg JF, Halpern AL, Rusch D, Eisen JA, Wu D, Paulsen I, Nelson KE, Nelson W, et al. Environmental genome shotgun sequencing of the Sargasso Sea. Science. 2004;304:66–74. doi: 10.1126/science.1093857. [DOI] [PubMed] [Google Scholar]
- Wallbanks S, Martinez-Murcia AJ, Fryer JL, Phillips BA, Collins MD. 16S rRNA Sequence Determination for Members of the Genus Carnobacterium and Related Lactic Acid Bacteria and Description of Vagococcus salmoninarum sp. nov. Int J Syst Bacteriol. 1990;40:224–230. doi: 10.1099/00207713-40-3-224. [DOI] [PubMed] [Google Scholar]
- Weigel LM, Clewell DB, Gill SR, Clark NC, McDougal LK, Flannagan SE, Kolonay JF, Shetty J, Killgore GE, Tenover FC. Genetic analysis of a high-level vancomycin-resistant isolate of Staphylococcus aureus. Science. 2003;302:1569–1571. doi: 10.1126/science.1090956. [DOI] [PubMed] [Google Scholar]
- Yapo BM. Rhamnogalacturonan-I: A Structurally Puzzling and Functionally Versatile polysaccharide from Plant Cell Walls and Mucilages. Polym Rev. 2011;51:391–413. [Google Scholar]
- Yarza P, Richter M, Peplies J, Euzeby J, Amann R, Schleifer K-H, Ludwig W, Glöckner FO, Rosselló-Móra R. The All-Species Living Tree project: A 16S rRNA-based phylogenetic tree of all sequenced type strains. Syst Appl Microbiol. 2008;31:241–250. doi: 10.1016/j.syapm.2008.07.001. [DOI] [PubMed] [Google Scholar]
- Zhang X, Paganelli FL, Bierschenk D, Kuipers A, Bonten MJM, Willems RJL, van Schaik W. Genome-wide identification of ampicillin resistance determinants in Enterococcus faecium. PLoS Genet. 2012;8:e1002804. doi: 10.1371/journal.pgen.1002804. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
A: 16S rDNA based phylogeny of the 49 previously known species that constitute the Enterococcus genus. The tree (Jukes-Cantor model, neighbor-joining method) was rooted using the 16S sequence of V. lutrae. Species indicated in blue were selected for sequencing and analysis. B: 16S based phylogeny of members of the Enterococcaceae and Carnobacteriaceae families (modified from existing tree of life (Yarza et al., 2008)). Closely related outgroup species of Vagococcus and Carnobacterium used in this study are indicated. Lactococcus garviae is omitted from the figure as the lineage that gives rise to it is not ancestral to the enterococci.
A: Distribution of the shared (common to all enterococci and outgroup species, and used to construct the phylogeny in figure 1B), the auxiliary and the species unique genes in study genomes. B: Gene accumulation and rarefaction curves for the genus Enterococcus. C: COG functional categories for the genus core (presence in >90 % of Enterococcus spp.), ubiquitous, genus related and genus unique genes. Ubiquitous genes are those found in 81 – 100 of the comparator genomes that span the Eubacteria (Fig. 2A), and are included for reference. D: Circular representation of the reference chromosome of E. faecalis V583, showing the location of core (n=1037), ubiquitous (n=378), shared (n=235), and unique (n=10) genes of Enterococcus spp. Mobile genetic islands predicted in E. faecalis V583 (Paulsen et al., 2003) are indicated by yellow highlight. E: Distribution of 1344 phenotype results by comparing average enterococcal species growth to average outgroup species growth. 45 conditions, represented in Fig. 2D, yielded to significantly more growth for Enterococcus spp. compared to the outgroup species. F: Growth of Vagococcus lutrae LBD1 on commercial enterococcal selective media (top, CHROMagar™; bottom, Enterococcosel™). G: Individual growth curves for all Enterococcus and outgroup species in the presence of sodium azide or chloroxylenol using the Biolog phenotypic microarrays (PM). PM plate number and well coordinates are indicated. Representative species for each phylogenetic group are shown. H: Circular map of the E. pallens BAA-351 genome showing genes of E. pallens coded in the clockwise (black) and anti-clockwise (grey) direction (outer circle) as illustrated. The 13 scaffolds of the draft genome are represented with black arrows (inner circle). Enterococcus genus core genes are shown in green, whereas the genes gained at node 18 and at the E. pallens leaf (Fig. 3A) are shown in blue.
Normalized distribution of annotated COG functional categories for niche specifying genes A. gained or B. lost at the node or leaf of each representative species (Fig. 3A). Node or leaf numbers are indicated (italicized) in front of the related species. C. gained or D. lost at the nodes of the Enterococcus phylogenetic tree (Fig. 3A).
A: Enterococcus spp. carbohydrate fermentation profiles (orange, growth; blue, no growth). Species are organized by phylogenetic tree position (Fig 1A). B: Growth of E. faecalis strain MMH594 in medium with no carbon source, uric acid or glucose. Data represent three independent experiments. Means and standard deviations are represented. C: Growth of E. faecalis MMH594 in medium supplemented with 0.5% glucose and 0.2% uric acid for 96 h at 37°C, under anaerobic conditions. Biomass (OD at 600nm, circles) Uric acid consumption (squares) and ammonia production (triangles) in the culture fluid were monitored over time. Similar phenotype was observed for negative control transposon mutant (EF1199) while both mutants in genes EF2570 and EF2559 were unable to metabolize uric acid in vitro (data not shown). D: Complexity and randomness of the mariner transposon insertion library in E. faecalis MMH594. Cumulative number of unique insertions throughout the 3.2 Mb chromosome (left panel), and distribution on the circular genomic map (right panel). E: Predicted KEGG pathway reconstruction of purine related metabolism in E. faecalis. Black arrows indicate identifiable genes associated with enzymatic activity in E. faecalis; grey arrows indicate no identifiable gene yet associated with this enzymatic activity. Genes interrupted by transposon insertion in E. faecalis mutants deficient in uric acid metabolism are indicated by locus tag numbers based on the E. faecalis V583 genome: in blue, gene gained since speciation of E. faecalis (Fig. 3A, node 4); in green, genes gained since the divergence from the last common ancestor shared with M. plutonius (Fig. 3A node 3); in black, genes shared by diverse enterococci species. F: Gut colonization of Galleria mellonella larvae. Indigenous E. faecalis before and after treatment with tetracycline (left panel). Gut colonization by E. faecalis MMH594 following one week of ad libitum feeding on pollen containing 5×107 CFU/g of pollen (right panel). G: Competitive index for growth in vitro (BHI), and for GI tract colonization of G. mellonella by wild type E. faecalis MMH594 (WT) and negative control transposon mutant (EF1199).
Each dot represents a pairwise comparison of a given species with all other species representatives in our study. Relevant comparator species are labeled.
Table S1. Strains and genomes metadata, Related to Figure 1
Table S2. Quantitative analysis of the Medline and Web of Science databases for reports of Enterococcus spp. in various ecologies, Related to Figure 1
Table S3. Prevalence of Enterococcus spp. and Vagococcus spp. in terrestrial and aquatic animals by 16S rDNA gut microbiome analysis, Related to Figure 1
Table S4. Analysis of Enterococcus spp. core genes. Related to Figure 2
Tab 1: Ubiquity score of 1037 Enterococcus core genes (>90% species) in other eubacteria.
Tab 2: Database of 100 genomes representative of families across the eubacterial domain.
Tab 3: Enterococcus core genes absent in Vagococcus lutrae LBD1.
Table S5. High-throughput phenotypic analysis using Biolog, Related to Figure 2
Supplemental dataset 1: Gene flux predicted by phylogenetic analysis using parsimony, Related to Figure 3.