Significance
The roles of tumor-infiltrating leukocytes in mediating cancer progression are well recognized, but a multidimensional analysis of the entire cancer immune system is lacking. Here, we dissected the cancer-immune landscape in hepatocellular carcinoma (HCC) across tumor, nontumor, and peripheral blood cells using time-of-flight mass cytometry, multiplex immunofluorescence tissue staining, and NanoString analysis. We identified various immune subsets that were enriched in the tumor microenvironment and their potential impact on the tumor immunity based on their detailed phenotypes. This study has validated the concept of a cancer-immune gradient and demonstrated in primary HCC that immune-cell subsets become progressively suppressive as they traverse the nontumor to tumor microenvironment. These data have opened avenues for the design of immunotherapeutics in HCC.
Keywords: CyTOF, tumor microenvironment, regulatory T cells, resident memory T cells, hepatocellular carcinoma
Abstract
The recent development of immunotherapy as a cancer treatment has proved effective over recent years, but the precise dynamics between the tumor microenvironment (TME), nontumor microenvironment (NTME), and the systemic immune system remain elusive. Here, we interrogated these compartments in hepatocellular carcinoma (HCC) using high-dimensional proteomic and transcriptomic analyses. By time-of-flight mass cytometry, we found that the TME was enriched in regulatory T cells (Tregs), tissue resident memory CD8+ T cells (TRMs), resident natural killer cells (NKRs), and tumor-associated macrophages (TAMs). This finding was also validated with immunofluorescence staining on Foxp3+CD4+ and PD-1+CD8+ T cells. Interestingly, Tregs and TRMs isolated from the TME expressed multiple markers for T-cell exhaustion, including PD-1, Lag-3, and Tim-3 compared with Tregs and TRMs isolated from the NTME. We found PD-1+ TRMs were the predominant T-cell subset responsive to anti–PD-1 treatment and significantly reduced in number with increasing HCC tumor progression. Furthermore, T-bet was identified as a key transcription factor, negatively correlated with PD-1 expression on memory CD8+ T cells, and the PD-1:T-bet ratio increased upon exposure to tumor antigens. Finally, transcriptomic analysis of tumor and adjacent nontumor tissues identified a chemotactic gradient for recruitment of TAMs and NKRs via CXCR3/CXCL10 and CCR6/CCL20 pathways, respectively. Taken together, these data confirm the existence of an immunosuppressive gradient across the TME, NTME, and peripheral blood in primary HCC that manipulates the activation status of tumor-infiltrating leukocytes and renders them immunocompromised against tumor cells. By understanding the immunologic composition of this gradient, more effective immunotherapeutics for HCC may be designed.
The immune system has a critical role in determining cancer pathogenesis and clinical fate (1, 2), and breakthroughs in the development of cancer immunotherapeutics have revolutionized the way we view and treat cancers (3). Numerous clinical trials on cancer immunotherapeutics are ongoing—particularly those assessing the efficacy of immune-checkpoint inhibitors (4–7). Despite encouraging results from clinical trials, fundamental differences in responsiveness both between different cancer types and within the same cancer type have been reported (8). The majority of research performed to date has focused on definitive cellular populations or individual molecules, and although this approach has been informative, it has not provided us with the knowledge as to which relevant immune functions are pivotal in certain cancers compared with others.
Hepatocellular carcinoma (HCC) is one of the most common cancers among males and the second most common cause of cancer-associated deaths globally (9). Limited treatment options and a high mortality rate pose a pressing need for the discovery of new therapeutic interventions (10). To date, immunotherapeutics based on immune-checkpoint inhibitors have generated promising results in some patients with HCC, but the majority do not respond well to treatment (11). Several previous studies on the tumor microenvironment (TME) have shown its crucial role in tumor progression and disease prognosis (12–15), but furthering our understanding of the cancer-immune landscape remains a challenge.
In this study, we hypothesized that mechanistically and clinically relevant immune signatures are profoundly shaped by the microenvironment in which immune cells reside, and that interactions between the microenvironment and immune cells are dynamic along a “cancer-immune gradient” that encompasses the TME, the nontumor microenvironment (NTME), and the peripheral blood (PB). Certain immune subsets [such as memory CD8+ T and T-regulatory (Treg) cells] may assume different phenotypes and functions depending on the microenvironment that they infiltrate and, therefore these differences may be exploited in different pharmacologic ways and in a tissue context-dependent manner. By determining the cancer-immune gradient for particular cancers, such as HCC, we can develop novel therapeutics that more precisely exploit the key pathologic molecules specific to the TME.
To accurately delineate the cancer-immune gradient in HCC, we first compared tumor-infiltrating leukocytes (TILs) to nontumor infiltrating leukocytes (NILs) and PB mononuclear cells (PBMCs) using time-of-flight mass cytometry (CyTOF). We identified multiple TIL subsets with increasingly suppressive phenotypes as they advanced from the PB and NTME to the TME. This was also validated with multiplex immunofluorescence tissue staining on two important immunosuppressive subsets: Foxp3+CD4+ Treg and PD-1+ CD8+ T cells. Specifically, tumor-infiltrating memory CD8+ T cells [resident memory (TRM) and effector memory (TEM)] showed high expression of multiple exhaustion makers, particularly, programmed death 1 (PD-1), an immune-exhaustion marker that is known to permit cancer immune evasion (16). Our in vitro studies found that these cells were the key subsets responsive to anti–PD-1 (monoclonal antibody) treatment. Elevated expression of PD-1 by TRMs was associated with down-regulation of the transcription factor T-bet upon increasing tumor-antigen exposure. Finally, we examined the general immune gene expression profile of the TME versus the NTME by transcriptomic analysis of 800 pan-cancer immune genes. By this method, we found that the TME hosts chemokines that likely recruit exhausted and potentially immunosuppressive TIL subsets.
Taken together, this unique in-depth understanding of the interface between systemic and tumor-specific immunity in HCC based on the combination of high-dimensional proteomic and transcriptomic analyses has identified axes as to how the TME might shape the composition and activation status of TILs. These findings will open up new avenues for future immunotherapeutic design.
Methods
Patients.
Tumor and adjacent nontumor liver tissues and PBMCs were collected from 54 patients with HCC who underwent curative resection at Singapore General Hospital. Patient consent was obtained according to the guidelines of the SingHealth Central Institutional Review Board. Fresh samples were obtained from 28 patients where TILs or NILs were freshly isolated using enzymatic digestion with 100 units/mL collagen IV (Thermo Fisher Scientific) for 30 min at 37 °C as previously described (17) and PBMCs were isolated from preoperative blood samples collected on the same day of surgical resection by layering over Ficoll-Paque Plus (GE Healthcare), as previously described (12). Fresh patient samples (n = 28) were selected for subsequent analyses based on immune-cell availability and as far as possible, equal distribution of viral status and tumor staging. Full clinical and demographic information of the patients and samples used for various experiments is summarized in SI Appendix, Table S1. Matched tumor and nontumor formalin-fixed paraffin-embedded (FFPE) tissue sections were obtained from an additional 26 HCC patients (SI Appendix, Table S2) for validation of CyTOF data using multiplex immunofluorescence tissue staining as described below.
CyTOF.
TILs, PBMCs, and NILs were obtained from patients with HCC (n = 14, 14, and 7, respectively) as described above, and processed for CyTOF analysis. A panel of 35 antibodies that encompassed a broad range of immune subsets was used (SI Appendix, Table S3) together with a combination of three anti-CD45 antibody barcodes for simultaneous sample processing, as previously described (18). The antibodies were either conjugated in-house according to the manufacturer’s instructions (Fluidigm) or purchased preconjugated directly from the supplier (Fluidigm). The cells were thawed and rested overnight in complete RPMI medium supplemented with 10% FBS, 1% penicillin/streptomycin/glutamine, and 10 mM Hepes at 37 °C. The cells were then either stimulated for 6 h with 150 ng/mL PMA (phorbol myristate acetate; Sigma) and 100 ng/mL ionomycin (Sigma) or unstimulated and exposed to 3 μg/mL Brefeldin A (eBiosience) and 2 μM monesin (BioLegend) during the final 4 h of the incubation. Next, the cells were stained with cisplatin (Fludigm) to identify live/dead cells and incubated with metal-conjugated surface-membrane antibodies. The cells were then fixed in 1.6% paraformaldehyde and permeablized in 100% methanol to permit staining with intracellular metal-conjugated antibodies. Finally, the cells were labeled with an iridium-containing DNA intercalator before analysis on a CyTOF-II mass cytometer (Fluidigm). The signal was bead normalized using EQ Four Element Calibration Beads (EQ Beads, 201078, Fluidigm) according to manufacturer’s instructions (19). The generated files underwent auto debarcoding and filtering for live/dead cells and DNA using R Studio (version 0.98.1073). The files were then analyzed using in-house enhanced automatic classification of cellular expression by nonlinear stochastic embedding (ACCENSE) software based on the combination of Barnes–Hut stochastic neighbor embedding (SNE) nonlinear dimension reduction algorithm and a k-means clustering algorithm (20) after down-sampling to 10,000 CD45+ immune cells from each sample for fair comparison across samples. Nodes that were significantly enriched in TILs, NILs, or PBMCs were identified using the Kruskal–Wallis rank sum test. Some of the data were independently validated using FlowJo software (version 10.0.7, Tree Star). Both 2D and 3D heat maps were plotted based on all significant nodes using R Studio for data visualization.
Multiplex Immunofluorescence Tissue Staining by Vectra.
FFPE sections from 26 HCC patients, who underwent curative resection from 1991 to 2009, were obtained from the Department of Anatomical Pathology, Division of Pathology, Singapore General Hospital (SI Appendix, Table S2). Two cores of tumor and nontumor FFPE tissues from each patient were stained with Opal Multiplex Immunohistochemistry Detection Kit and images were acquired using a Vectra 3.0 Pathology Imaging System Microscope (Perkin-Elmer) as described previously (21). Antibodies used were: anti-CD8 (DAKO, clone C8/144B), anti-CD4 (Abcam, clone EPR6855), anti–PD-1 (Abcam, clone NAT105), and anti-Foxp3 (Abcam, clone 236A/E7). Detection dye for each antibody was: Opal690 dye (CD8), Opal540 dye (CD4), Opal650 dye (PD-1), and Opal520 dye (Foxp3). DAPI were used as a nuclear counterstain. Quantification of positively stained cells was performed on the whole 1-mm core (area = 0.785 mm2) and the average values of two cores from each patient sample were calculated and shown as the number of cells per square millimeter.
In Vitro Cellular Coculture and Cytokine Production Assay.
For detection of the T-bet:PD-1 ratio in PBMCs upon exposure to tumor antigens, autologous tumor-dissociated cells and PBMCs from eight HCC specimens were thawed and rested overnight. A total of 0.5 × 106 dissociated tumor cells (representing the tumor antigen) or PBMCs (as a control) were then subjected to X-ray irradiation (RS-2000, Radsource) at 200 Gy and PBMCs (representing feeder cells) were irradiated at 30 Gy, as previously described (22). The irradiated cells were mixed with 1 × 106 nonirradiated PBMCs (n = 8) and analyzed for the T-bet:PD-1 ratio on days 0, 1, 2, 3, and 6 of coculture by flow cytometry (BD LSRFortessa X-20, BD Biosciences). The cells were stained with anti-human CD45, CD3, CD4, CD8, CD45RO, T-bet, CD103, PD-1 (BioLegend), CD56 (BD Biosciences), and blue-fluorescent reactive dye (Life Technologies) for live/dead cell staining.
To examine the response of TILs to anti–PD-1 treatment in vitro, similarly irradiated TILs and PBMCs were mixed with 1 × 106 nonirradiated TILs that had been exposed to 10 μg/mL anti-human PD-1 antibody (clone J116, eBioscience) or mouse IgG1, κ-isotype control (eBioscience) for 18 h (n = 8). The cells were exposed to Brefeldin A and monesin (BioLegend) during the final 4 h of the incubation. Production of TNFα and IFNγ (BioLegend), cell-surface marker expression and live/dead cell staining (as described above) in nonirradiated TILs was measured by flow cytometry (BD LSRFortessa X-20, BD Biosciences). Postacquisition analysis was performed for both experiments using FlowJo software (version X.0.7, Tree Star).
NanoString Analysis.
Unsorted tumor and adjacent nontumor liver tissues were collected upon resection from 20 patients with HCC. Tissues were lyzed using Tissuelyzer-II (Qiagen) and the lysates were processed using the mirVana miRNA Isolation Kit (Ambion) to extract total RNA. Quantification of RNA was performed using the Quant-iT RiboGreenRNA assay kit (Invitrogen). The samples were then analyzed using the nCounter platform (NanoString Technologies) using a combination of the prebuilt nCounter PanCancer Immune Profiling code set (consisting of 770 genes) and an additional 30 customized compatible probes from the nCounter Panel-Plus code set. The raw data were normalized against housekeeping genes using the NanoStringNorm R package and log transformed, before analyzing by Student’s paired t test and generating heat maps for visualization.
Statistical Analyses.
For CyTOF data, the nonparametric Kruskal–Wallis rank sum test was used to identify nodes that were differentially present among three groups and the Student’s paired t test was used to analyze NanoString data to identify significantly expressed genes. Statistical analyses of the FlowJo data were performed using a paired or unpaired Student’s t test, unpaired Mann–Whitney u test, and Spearson’s correlation test, as indicated (GraphPad Prism V.6.0f).
Results
TME, NTME, and PB Exhibit a Unique Immune Landscape Defined by Specific Immune Subsets.
We first designed a CyTOF assay that would permit a deep interrogation of the TME, NTME, and PB immune landscapes in tissues/blood extracted from patients with HCC (Fig. 1A). To this aim, we designed a panel of 35 surface membrane and intracellular immune markers that encompassed a broad range of immune subsets (SI Appendix, Table S3) to examine the global phenotypes of TILs, NILs, and PBMCs from patients (n = 7) with HCC (SI Appendix, Table S1). The cells were prestimulated for 6 h with PMA and ionomycin, as the differences between the TME, NTME, and PB compartments were enhanced upon activation.
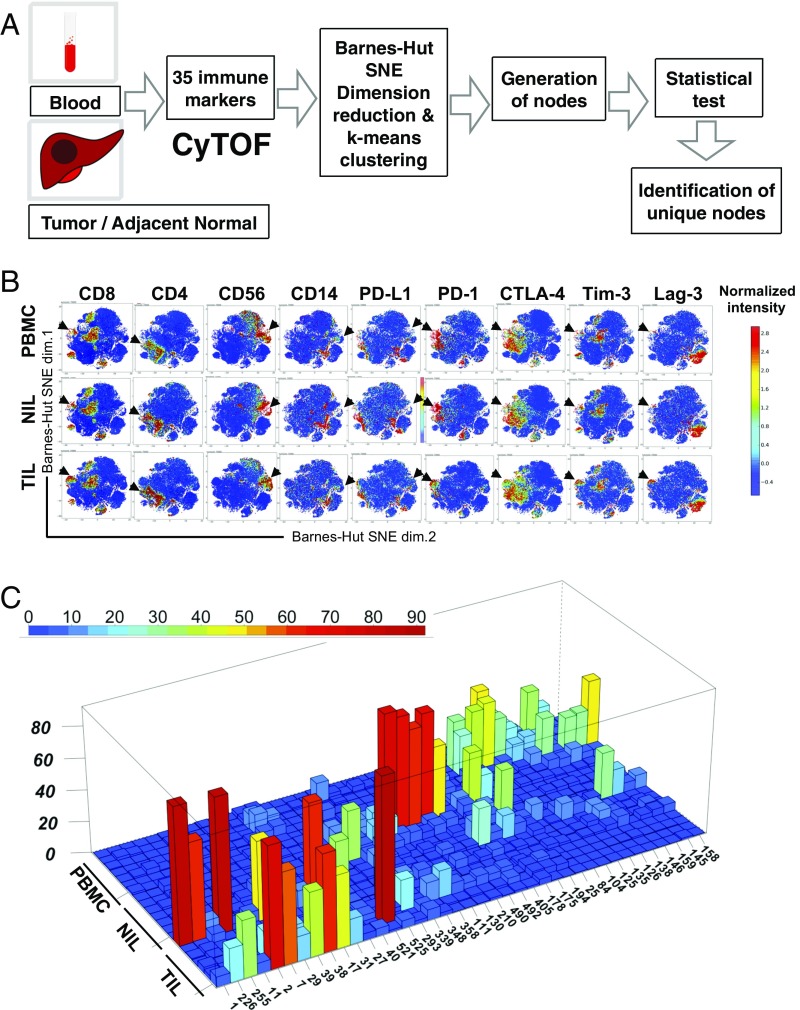
Fig. 1.
High-dimensional analysis with Barnes–Hut SNE identified differentially enriched immune subsets in TILs, NILs, and PBMCs. (A) Time-of-flight mass cytometry (CyTOF) pipeline from data acquisition, dimension reduction, and clustering to node generation. The resulting nodes, clustered by similarity in their immune phenotypes, were subjected to statistical testing to identify significantly enriched nodes from a given group (TIL, NIL, or PBMC). (B) The 2D cellular t-SNE plots of CyTOF data from PBMCs, NILs, and TILs as gated on: CD8, CD4, CD56, CD14, PD-L1, PD-1, CTLA-4, Tim-3, and Lag-3. Each dot represents one single cell. Arrows showed distinct differences in TILs. (C) A 3D illustration of node percentages in either PBMC, NIL, or TIL compartments as grouped into node ID and major immune subsets. n = 7 from each compartment.
Data files generated from the CyTOF analysis for individual samples were subjected to a dimension reduction process based on the Barnes–Hut SNE algorithm and then followed by cell clustering based on the k-means clustering algorithm using in-house enhanced ACCENSE software (20). Following this step, the cells formed distinct clusters (representing immune subsets) based on similarities in immune-marker expression. These nodes were further analyzed by Kruskal–Wallis rank sum test to identify those that were statistically different among the three cellular compartments (combined node frequency >70% in one of the compartments at P < 0.05) (Fig. 1A). The expressions of selective markers were shown with 2D t-SNE plots for lineage markers, such as CD8, CD4, CD56, and CD14, as well as exhaustion markers PD-L1, PD-1, CTLA-4/CD152, Tim-3, and Lag-3 (Fig. 1B). Differences in node frequencies can be appreciated across three different compartments (SI Appendix, Fig. S1A). With the statistical analysis, we identified nodes that showed distinct distributions across TIL, NIL, and PBMC compartments (Fig. 1C). These data imply that the immune landscapes of the TME, NTME, and PB in HCC are unique in terms of their cellular composition and would therefore respond differentially to immunotherapeutic intervention.
By this unbiased and unsupervised method, 15 TIL, eight NIL, and 11 PBMC-enriched nodes were identified, and three and two nodes were enriched in both TIL/NIL and NIL/PBMC, respectively (SI Appendix, Fig. S1B). We further analyzed the full 35-immune marker expression profile of each of the statistically significant nodes as identified above in a 2D heat map to identify phenotypic differences or similarities between the nodes (Fig. 2A). Generally, we found that the immune subsets within the TILs and NILs were phenotypically more similar to each other than the immune subsets derived from the PBMCs. Classification of these phenotypically similar nodes identified the immune subsets that were enriched in each compartment. TIL-enriched subsets were identified as: TRMs that were CD103+CD45RO+CD8+PD-1+CD152/CTLA-4+Lag-3+Tim-3+; TEMs that were PD-1+Lag-3+CD152+Tim-3+CD45RO+CCR7−; Tregs that were CD4+Foxp-3+CD152 +IL10+CD45RO+PD-1+; and CXCR3+ natural killer (NK) cells that were either resident (CD103+) NKR cells or nonresident (CD103−) NK cells that were GzmB−. The NIL-enriched subsets were: PD-1− TEM; GzmB+ NK cells; and CD8+CD56+ NKT cells. The PBMC-enriched subsets were: naive CD8+ T cells and CD45RA+CD4+ T cells.
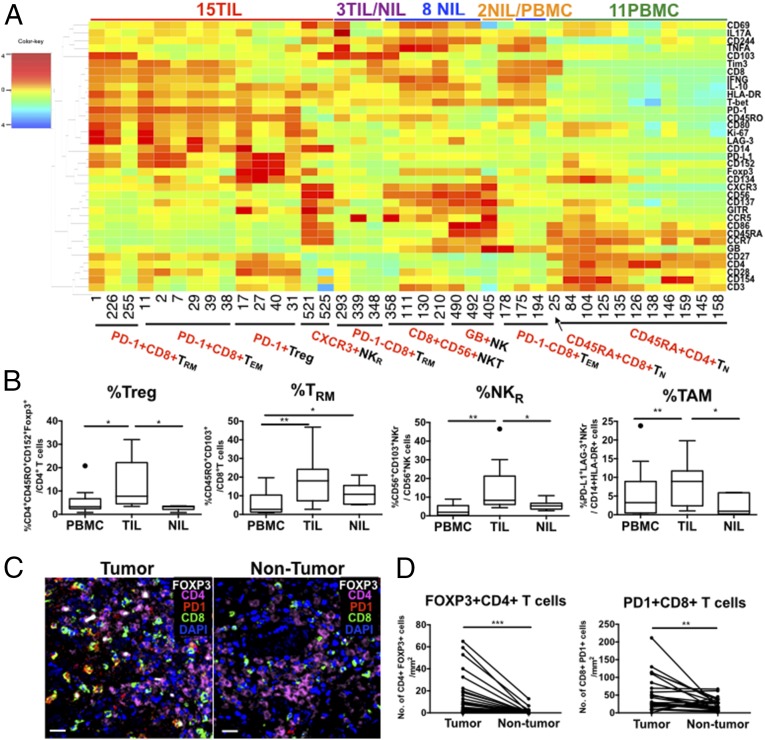
Fig. 2.
High-dimensional Barnes–Hut SNE analysis defines the distribution of specific immune subsets in TILs, NILs, and PBMCs. (A) A 2D heat map showing differential expression of 35 immune markers by TIL-enriched (red bar), NIL-enriched (blue bar), PBMC-enriched (green bar), TIL/NIL-enriched (purple bar), or NIL/PBMC-enriched (orange bar) nodes. Immune subsets were categorized based on their marker expression. (B) Percentage of Tregs, TRMs, NKRs, and TAMs in PBMCs (n = 12), TILs (n = 12), or NILs (n = 7). Data represent the means ± SD and were analyzed by paired Student’s t test, **P < 0.01 and *P < 0.05. (C) Representative images from multiplex immunofluorescence tissue staining for CD8 (green), PD-1 (red), CD4 (magenta), Foxp3 (white), and DAPI (blue) on tumor and nontumor FFPE tissues. (Scale bar, 20 μm.) (D) Quantification of the number of Foxp3+CD4+ Treg and PD-1+CD8+ T cells per square millimeter in tumor versus nontumor tissues from n = 26 paired HCC samples. Paired Student’s t test. **P < 0.01 and ***P < 0.001.
Binary analysis of the resting or unstimulated cells confirmed the composition of the TIL-enriched immune subsets described above and identified an additional TIL-enriched population: CD14+HLA-DR+PD-L1+Lag-3+ tumor-associated macrophages (TAMs). These cells were largely eliminated upon PMA and ionomycin stimulation (SI Appendix, Fig. S3A). These data were validated by manual gating using FlowJo for these specific immune subsets in the three compartments with additional TIL and PBMC samples available (SI Appendix, Fig. S2A). Of note, the overall lower number of NIL cases was due to the limited availability of large enough surgical nontumor tissue from which immune cells could be isolated for CyTOF analysis. Indeed, we observed that Tregs, NKRs, and TAMs were significantly enriched in TILs, whereas TRMs were enriched in both TILs and NILs (Fig. 2B and SI Appendix, Fig. S3B). Data validation was performed using multiplex immunofluorescence tissue staining on an additional 26 paired tumor and nontumor FFPE samples (SI Appendix, Table S2) and demonstrated that indeed the number of Foxp3+CD4+ Tregs and PD-1+CD8+ T cells per area was higher in tumor versus nontumor compartments (Fig. 2 C and D).
Tregs Assume a More Immunosuppressive Phenotype When in Contact with the TME.
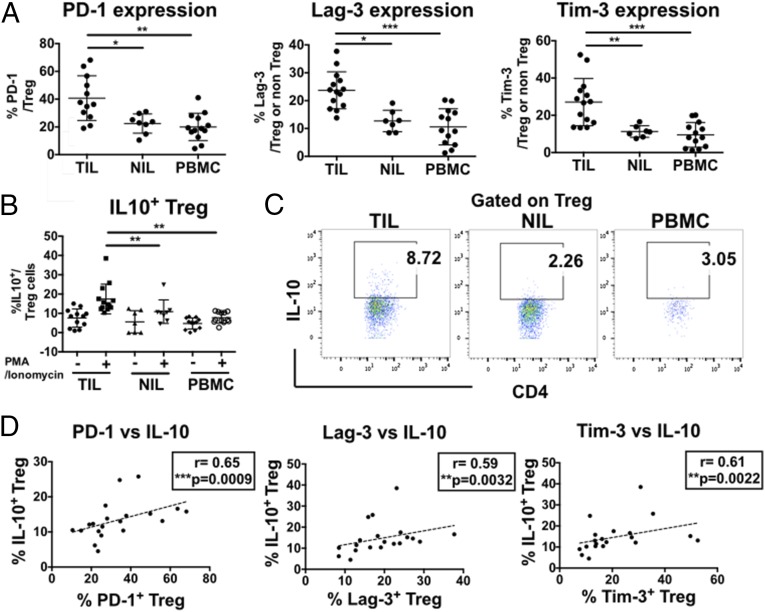
We next aimed to address whether the cellular subsets enriched in the TILs exhibited divergent phenotypes when exposed to different microenvironments. First, we focused on Tregs, given their well-established correlation with poor prognosis in various cancers, including HCC (23–25). Recent studies have also demonstrated the expression of exhaustion markers on Tregs, in particular PD-1, and their effect on Treg function (26, 27). We then examined whether the Tregs expressed different immune exhaustion markers between the TIL, NIL, and PBMC groups. Interestingly, we observed that a higher percentage of Tregs expressed multiple exhaustion markers in those derived from the TIL compared with the NIL and PBMC compartments: PD-1 (TIL, 40.7 ± 16.1%; NIL, 22.5 ± 6.9%; and PBMC, 20.0 ± 9.8%), Lag-3 (TIL, 23.7 ± 6.6%; NIL, 12.7 ± 3.9%; and PBMC, 10.6 ± 6.5%), and Tim-3 (TIL, 27.1 ± 12.7; NIL, 11.3 ± 3.1%; and PBMC, 9.5 ± 6.7%) (Fig. 3A). In addition, we found that upon stimulation with PMA and ionomycin, Tregs from the TILs expressed significantly more IL-10 than Tregs from NILs or PBMCs (Fig. 3 B and C). Representative plots of IL-10 expression in Treg populations of the TILs, NILs, and PBMCs clearly demonstrated a higher percentage of IL-10+ Tregs derived from the TILs upon stimulation (Fig. 3C). The suppressive phenotype of Tregs from TILs was also demonstrated by its ability to suppress T-cell proliferation (SI Appendix, Fig. S4).
Fig. 3.
The TME is enriched with more exhausted Tregs compared with the NTME. (A) Percentage of PD-1–, Lag-3–, or Tim-3–expressing Treg cells from TILs versus NILs or PBMCs. ***P < 0.001, **P < 0.01; *P < 0.05, paired Student’s t test. (B) Percentage of IL10+ Tregs from TILs, NILs, or PBMCs with or without 6-h PMA and ionomycin stimulation. **P < 0.01, paired Student’s t test. (C) Representative plots showing expression of IL-10 pregated on CD4+Foxp3+CTLA-4+ Tregs in TILs versus NILs and PBMCs from one HCC patient HEP178. (A–C) TILs (n = 12) versus NILs (n = 7–8) and PBMCs (n = 12). Data represent the means ± SD. (D) Correlation between expression of exhaustion markers: PD-1, Lag-3, and Tim-3 with IL-10 on Tregs from TILs and NILs. n = 21. P values and correlation coefficients (r) were calculated with Spearman’s correlation test. ***P < 0.001 and **P < 0.01.
Finally, we examined whether high IL-10 expression (indicative of a suppressive phenotype) was associated with exhaustion-marker expression. Indeed, we observed that the expression of the exhaustion markers PD-1, Lag-3, and Tim-3 correlated with the expression of IL-10 in Tregs (Fig. 3D). These data support the concept of an immune-cancer gradient, whereby the same cell type (i.e., Tregs) can assume different phenotypes and functions, depending on their residing microenvironment. Specifically, we found that Tregs exhibited an increased immunosuppressive behavior in the TME.
CD8+ TRMs and TEMs Exhibit an Enhanced T-Cell–Exhausted State in the TME.
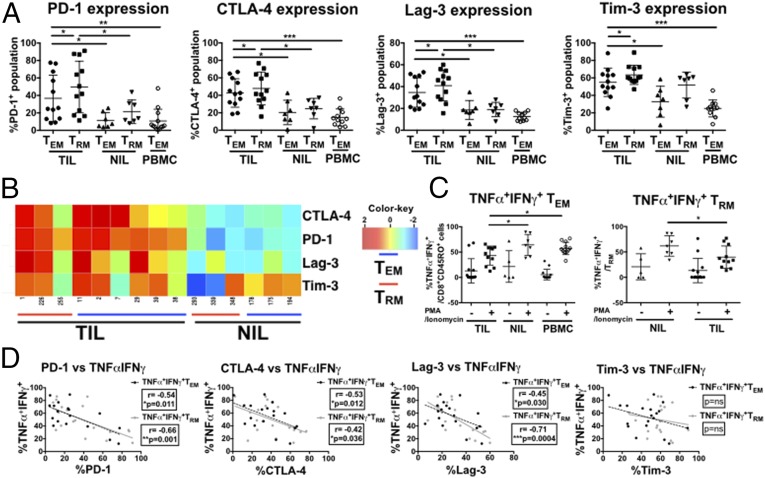
We next explored the cancer-immune gradient between the TME and the NTME and focused on analyzing their common T-cell subsets, TEM and TRM. We examined the expression of multiple exhaustion markers on these immune subsets by manual gating for PD-1, CTLA-4, Lag-3, and Tim-3 and observed a significantly higher percentage of PD-1+CTLA-4+ and Lag-3+ CD8+ TEM and TRM cells in the TILs compared with the NILs (Fig. 4A). Conversely, a significantly higher percentage of Tim-3+ cells was only observed in TEMs from the TILs versus NILs (Fig. 4A). A 2D heat map illustration of the expression level of exhaustion markers on TEM or TRM cells also showed a generally higher expression of these markers from those infiltrating TME versus NTME (Fig. 4B).
Fig. 4.
Exhaustion marker and cytokine expression by memory CD8+ from TILs, NILs, and PBMCs. (A) Percentage of PD-1–, CTLA-4–, Lag-3–, or Tim-3–expressing CD8+CD45RO+CD103− TEMs and CD8+CD45RO+CD103+ TRMs in TILs (n = 12) versus NILs (n = 7) or PBMCs (n = 12). Data represent the means ± SD and were analyzed by paired Student’s t test, ***P < 0.001, **P < 0.01, and *P < 0.05. (B) A 2D heat map representing the expression of exhaustion markers by TEM (red line) or TRM (blue line) nodes in TILs or NILs. (C) Percentage of TNFα and IFNγ-expressing TEMs and TRMs in TILs (n = 12) versus NILs (n = 5–7) or PBMCs (n = 12) with or without 6-h PMA and ionomycin stimulation. Data represent the means ± SD and were analyzed by paired Student’s t test, *P < 0.05. (D) Correlation between percentage of PD-1+, CTLA-4+, Lag-3+, or Tim-3+ TEMs and TRMs versus TNFα+IFNγ+ TEMs and TRMs in TILs and NILs upon 6-h PMA and ionomycin stimulation. n = 19 each. P values and correlation coefficients (r) were calculated with Spearman’s correlation test. ***P < 0.001, **P < 0.01, and *P < 0.05.
We next examined the responsiveness of the TEM and TRM cells expressing exhaustion markers to ex vivo immune activation with PMA and ionomycin. Upon activation, a lower percentage of TNFα+IFNγ++ TEM and TRM cells were identified in the TILs compared with the NILs (Fig. 4C). We then examined the association between the expression of exhaustion markers with the expression of TNFα and IFNγ on these cells. Indeed, we observed an inverse correlation between the expressions of PD-1, CTLA-4, and Lag-3 with the expression of TNFα+IFNγ++ on TEM and TRM cells upon PMA and ionomycin stimulation (Fig. 4D), indicating a link between the expression of these exhaustion markers and the functional competency of these cells. The correlation between the expression of Tim-3 and the cytokines produced by either TEM or TRM cells was, however, not significant (Fig. 4D), perhaps suggesting a less critical role for Tim-3 on the functional competency of these cells. These data further strengthen the hypothesis of a cancer-immune gradient and demonstrate the generation of an increasingly immunosuppressive phenotype from the PB to the TME in HCC. Further, these data demonstrate that exhaustion markers are widely expressed by multiple T-cell subsets enriched in the TME, including Tregs, CD8+ TRMs, and TEMs.
PD-1–Expressing CD45RO+CD8+ TILs in the TME Are Responsive to Anti–PD-1 Treatment and Mediate Tumor Progression.
An immediate question arising from these data is how two distinct and specific adjacent microenvironments (TME and NTME) can shape T-cell function. To address this question, we decided to analyze the possible link between the expression of the transcription factor T-bet and PD-1 on TEMs and TRMs from the TILs and NILs, as previous data had found that T-bet can suppress PD-1 expression in mice (28, 29). In our human HCC samples, we observed a higher percentage of T-bet−-expressing TEMs and TRMs isolated from the NILs compared with the TILs (Fig. 5A). An inverse correlation between T-bet and PD-1 was also found in the same cells from both the TILs and NILs (Fig. 5B). These data suggest that T-bet may control PD-1 expression when exposed to the TME.
Fig. 5.
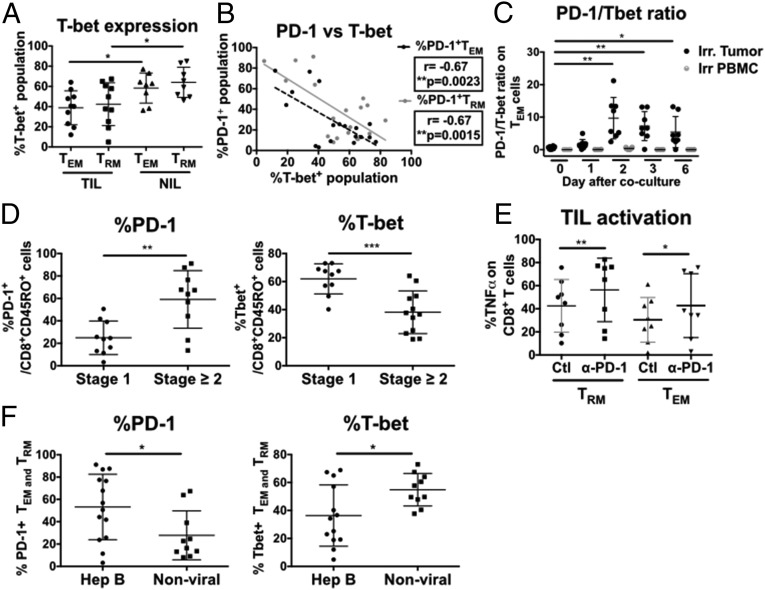
T-bet is the critical transcription factor that correlates with down-regulation of PD-1 upon tumor antigen exposure and tumor progression. (A) Percentage of T-bet–expressing TEMs and in TILs (n = 12) versus NILs (n = 7). Data represent the means ± SD and were analyzed by paired Student’s t test. *P < 0.05. (B) Correlation between percentage of PD-1+ TEMs and TRMs versus T-bet+ TEMs and TRMs in TILs and NILs upon 6-h PMA and ionomycin stimulation. n = 19 each, P values and correlation coefficients (r) were calculated with Spearman’s correlation test. **P < 0.01. (C) Ratios of percentages of PD-1+ versus T-bet+ on CD8+CD45RO+ TEM cells from PBMCs upon coculture with irradiated autologous tumor (n = 8) or PBMC (n = 6) and feeder cells. Data represent the means ± SD and were analyzed by paired Student’s t test with reference to day 0. **P < 0.01 and *P < 0.05. (D) Percentage of PD-1+ or T-bet+ TEMs and TRMs in stage 1 versus stages 2–4. Data represent the means ± SD and were analyzed by unpaired Student’s t test. n = 10 each. ***P < 0.001 and **P < 0.01. (E) Percentage of TNFα-expressing TEMs and TRMs from TILs after 18 h cocultured with autologous irradiated tumor cells with anti–PD-1 antibody (α–PD-1) or with isotype control mouse IgG1,κ-antibody (Ctl). n = 8. Data represent the means ± SD and were analyzed by paired Student’s t test with reference to control. **P < 0.01 and *P < 0.05. (F) Percentage of PD-1+ or T-bet+ TEMs and TRMs in HBV-infected (n = 12) versus nonvirally infected (n = 10) HCC patients. Data represent the means ± SD. P values were calculated using unpaired Student’s t test. *P < 0.05.
To test this hypothesis, we cocultured PBMCs from patients with HCC that exhibited low PD-1:T-bet ratio (at day 0, Fig. 5C) with irradiated autologous tumor samples (as a source of tumor antigens) and feeder cells, as previously described (22). After 48-h exposure to the irradiated tumor sample, the ratio of PD-1 versus T-bet expression on TEM cells from the PBMCs was enhanced compared with day 0, and this ratio was maintained until day 6 (Fig. 5C). Of note, the same experiment performed using irradiated autologous PBMCs showed no increase in PD-1/T-bet ratio, demonstrating the specificity of this up-regulation of PD-1 when exposed to tumor antigen (Fig. 5C). We hypothesized that this phenomenon may have a clinically relevant implication on tumor progression and thus analyzed the percentage of PD-1+ and T-bet+ CD8+CD45RO+ cells in early stage (stage 1) versus advanced stage (stage ≥2) tumors. Strikingly, we observed that the percentages of PD-1+ memory CD8+ T cells were significantly increased in the later-staged HCC tumors compared with the early-staged tumors, whereas the percentages of T-bet+ memory CD8+ T cells were significantly decreased in the later-staged tumors (Fig. 5D). Importantly, these cellular subsets from the TILs were also responsive to anti–PD-1 treatment, as determined by the production of TNFα upon anti–PD-1 treatment in vitro (Fig. 5E).
We also explored the balance between T-bet and PD-1 expression in patients with HCC and concomitant chronic hepatitis B virus (HBV) infection. Higher expression of PD-1 and correspondingly lower expression of T-bet was detected in patients infected with HBV compared with uninfected patients (Fig. 5F), indicating an enhanced level of TIL exhaustion in chronic HBV infection. Overall, these in vitro data directly indicate that these described TIL subsets (PD-1+CD8+ TRMs and TEMs) are the primary target for anti–PD-1 immunotherapy, which has important implications for the design of anti–PD-1 treatments in HCC.
TME Shapes the Immune Landscape and Determines the Immune Subsets That Accumulate in the Tumor.
Our data so far have shown that the TME is infiltrated with phenotypically more immunosuppressive/exhausted T cells, compared with the NTME and PB, thus indicating the critical role of the TME in shaping the phenotypes of TILs. To further support the crucial role of TME in shaping the immune landscape of HCC, we analyzed the RNA expression of 800 cancer-immune genes from unsorted tumor and nontumor samples using NanoString analysis. We found overall an higher expression of cancer-associated genes (including BIRC5, CDK1, TTK, ITGA6, PBK, and SPA17, and melanoma-associated antigen genes) in the TME compared with the NTME, which validated the purity of our samples (Fig. 6A). We also found that the TME expressed lower levels of multiple inflammatory cytokines and chemokines (including IL1B, IL6, IL8, TNF, GZMH, GZMK, CXCL12, CXCL14, CXCL1, CXCL16, CCL21, and CCL19) (Fig. 6A), which is indicative of a more immunosuppressive TME.
Fig. 6.
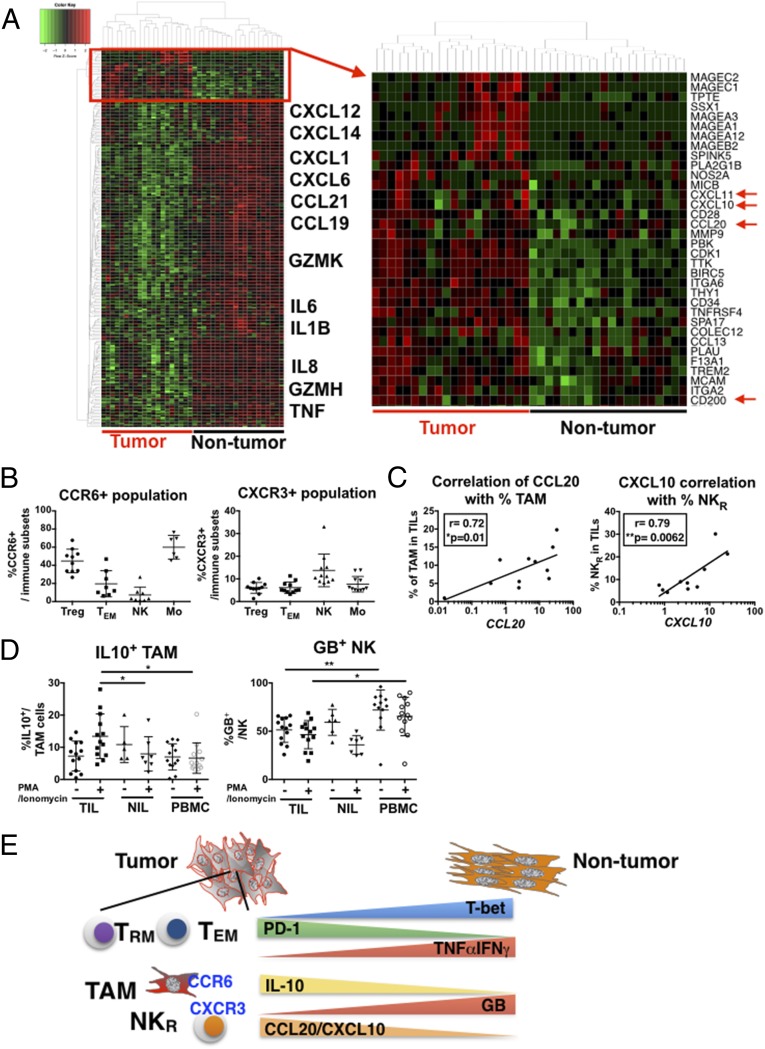
The tumor microenvironment shapes and dictates TIL infiltration in HCC. (A) Heat map showing NanoString analysis data from unsorted tumor versus adjacent nontumor tissues. n = 20 paired tissues. Box shows enriched genes in TME versus NTME with the magnified image on the Right. (B) Expression of CCR6 and CXCR3 on Tregs (CD4+Foxp3+), TEMs (CD8+CD45RO+), NKs (CD56+), or monocytes (CD14+HLA-DR+) from PBMCs of HCC patients. n = 6–10. Data represent the means ± SD. (C) Correlation of RNA expression of CCL20 to percentage of TAMs, and CXCL10 to percentage of NKRs in TILs. n = 12. P values and correlation coefficients (r) were calculated with Spearman’s correlation test. *P < 0.05 and **P < 0.01. (D) Percentage of IL-10+ TAMs and granzyme B (GB)+ NKs from TILs (n = 12) versus NILs (n = 5–7) or PBMCs (n = 12) with or without 6-h PMA and ionomycin stimulation. Data represent the means ± SD and were analyzed by paired Student’s t test. *P < 0.05 and **P < 0.01. (E) Working hypothesis model showing tumor-enriched chemokines: CCL20 attracting CCR6+ TAMs, which produce high levels of IL-10; whereas CXCL10 attracts CXCR3+ NKRs that express low levels of granzyme B (GB) within the tumor site. Lower expression level of T-bet, and a reciprocal higher expression level of PD-1, and lower production of TNFα and IFNγ was observed on TEMs and TRMs infiltrating the tumor.
Several chemokines (including CCL20, CXCL10, and CXCL11) were highly expressed in the TME compared with the NTME (Fig. 6A, arrows). CCL20 is of particular interest as it is a ligand for CCR6 that is known to be expressed by Treg (30), CD8+ TEM (31, 32), Th-17 cells (33), B cells, immature dendritic cells (DCs) (34), and tumor-promoting macrophages (35). We also observed high CD200 expression levels in the TME (Fig. 6A, arrow), which has known roles in tumor immune evasion via interactions with TAMs (36) and in suppressing antitumor immunity (37).
Given the enrichment of CCL20, CXCL10, and CXCL11 chemokines in the TME, we examined the protein expression of their respective receptors—CCR6 for CCL20, CXCR3 for CXCL10, and CXCL11 on the TILs. First, we examined by flow cytometry the protein expression of CCR6 and CXCR3 on PBMCs from patients with HCC, based on the hypothesis that these PBMC subsets were the potential target for chemotaxis along the chemokine-gradient toward the TME. A higher proportion of CCR6+ macrophages/monocytes (defined as CD14+HLA-DR+ cells) and CXCR3+CD56+ NK cells were observed compared with other immune subsets (Fig. 6B). By combining the chemokine expression level with the RNA expression analysis and CyTOF data, we identified a correlation between the RNA expression level of CCL20 with the percentage of intratumoral TAMs and the expression level of CXCL10 with the percentage of intratumoral NKRs (Fig. 6C). In addition, TIL-residing NK cells expressed a lower level of granzyme B and the TAMs expressed higher levels of IL-10 (Fig. 6D). These data again indicate that tumor-enriched chemokines attract more suppressive immune subsets to the TME.
Taking all these data together, we propose that the primary HCC immune landscape might hinge on the following: two key chemokines, CCL20 and CXCL10, that may induce the accumulation of two immune-cell subsets, TAM and NKR, which express high levels of IL-10 and low levels of granzyme B in the TME (Fig. 6E); and the transcription factor T-bet, which may be the coordinating factor controlling the expression of PD-1 and the corresponding production of TNFα and IFNγ by TEM and TRM cells along an immunosuppressive gradient from NILs to TILs (Fig. 6E).
Discussion
The vital contribution of the immune surveillance system to cancer progression has been swiftly translated to the clinical setting with the introduction of immunotherapeutics to treat various cancers (3, 4). However, the marked variability in patient (and cancer-type) responsiveness to such treatments, including in HCC, renders the urgent need for an advanced understanding of the interaction between the host immune system and tumor-cell survival (5). In this study, we dissected the immune compartments within the TME, NTME, and PB in patients with HCC to define the cancer-immune gradient and delineate the phenotypes and functions of the infiltrating immune subsets. This holistic approach permitted a broad understanding of HCC as a systemic disease whereby the immune response is differentially manipulated according to various factors (such as exposure to tumor antigens and differential expression of intratumoral chemokines) present in the cancer and noncancer microenvironments.
This study uses CyTOF for deep phenotyping of the immune-cell subsets that accumulate in the TME, NTME, and PB in patients with HCC. We found that the cellular subsets that represented the tumor tissue exhibited an exhausted and immunosuppressive phenotype, as evidenced by the enhanced expression of multiple exhaustion markers (PD-1, CD152, and Lag-3), accumulation of IL-10–expressing immunosuppressive Treg and TAM subsets, and low expression of inflammatory cytokines (TNFα, IFNγ, and granzyme B) by tumor-infiltrating T and NK cells. The role of exhaustion markers (particularly PD-1) on Treg function remains controversial, as it was demonstrated in malignant gliomas that PD-1hi Treg was dysfunctional and produced IFNγ (26). Others have described that PD-1 expression is correlated with more suppressive Treg phenotypes in chronic viral infection (27). Our data, however, have clearly demonstrated a correlation between Treg suppression and exhaustion-marker expression.
Despite their phenotypically exhausted and suppressive nature, some of the TIL-enriched immune subsets remained functionally competent upon ex vivo stimulation. For instance, CD8+ TRMs and TEMs from the TILs expressed TNFα and IFNγ upon activation with PMA and ionomycin, and their overall percentage was reduced in advanced-stage tumors. These data imply that these cell subtypes are critical for tumor progression. This observation is consistent with previous reports that high abundance of CD45RO+ memory-CD8+ T cells and TRMs are associated with good prognosis in several types of cancer (1, 38). We found that the PD-1+ TRMs and TEMs could also be activated with anti–PD-1 treatment ex vivo and hence may be candidate immune cells that respond to immune-checkpoint blockade. These data have important clinical relevance as the relative presentation of these critical immune subsets could be considered as a biomarker for potential clinical response against anti–PD-1 therapy.
The majority of cases of HCC reportedly arise from chronic viral hepatitis infection (39, 40). Furthermore, chronic HBV infection has been linked to a marked increase in exhaustion-marker expression, including PD-1 on T cells (41). Due to this increase, concerns with regards to possible toxicity of immunotherapeutics in HCC have been raised, as the application of check-point blockage will very likely illicit an off-target immune response to the remaining noncancerous liver tissues infected with HBV. Interestingly, our data suggest that such toxicity will not be a major concern, due to the identification of preferentially higher expression of PD-1 by TILs than NILs. Even though we validated that T cells from patients infected with HBV exhibited higher PD-1 expression, the percentage of PD-1+ T cells remains higher in TILs versus NILs, even in this group of HCC patients. This concept is again consistent with the hypothesis of a cancer-immune gradient, particularly regarding the TRMs and TEMs from the TILs that showed an increasingly exhaustive profile from the adjacent NTME to the TME. This increase in exhaustion was induced upon exposure to tumor antigens, thus indicating an important role for the TME in shaping and maintaining an exhausted and immunosuppressive microenvironment. The approach used in this study has important implications for the design of novel immunotherapeutics in HCC, as the expression profile of various exhaustion markers could be analyzed on multiple TIL or NIL subsets simultaneously to select the most effective therapeutics with the least potential for adverse effects. For instance, our detailed phenotypic data have provided insight into the potential for combined immunotherapeutic treatments in HCC, such as the combination of PD-1 and CTLA-4 or Lag-3 blockade based on their differential expression levels in TILs versus NILs.
This study also demonstrated an inverse relationship between T-bet and PD-1 expression in TILs and NILs in the cancer setting, suggesting a role for T-bet in modulating PD-1 expression. Indeed, exposure of patient PBMCs to tumor antigens resulted in an up-regulation of PD-1 and a corresponding down-regulation of T-bet in TEM cells. This paper reports the relationship between T-bet and PD-1 in HCC, although a similar association has been previously described during chronic virus infection (28), including chronic HBV infection (42). Consistently, we also observed higher PD-1 and lower T-bet expression in TILs from patients with concurrent HCC and HBV infections compared with those without HBV infection. Our previous characterization of HCC immune gene signatures demonstrated that TBX21 (T-bet) was an immune gene associated with a good prognosis in HCC (12). Taking this previous observation and our current data together, we propose that the inverse relationship between T-bet and PD-1 is one of the mechanisms for the evolution from chronic infection and inflammation to tumor development and cancer progression.
Finally, we explored how the TME might shape the relative composition and function of infiltrating immune subsets. NanoString data comparing the TME versus NTME revealed enhanced expression of CCL20, CXCL10, and CXCL11 in the TME. Next we detected the expression of the respective chemokine receptors CCR6 on TAM and CXCR3 on NK cells. Production of these chemokines correlated with an increased percentage of TAMs and NKRs in tumors. Most importantly, both of these immune subsets exhibited an immunosuppressive phenotype in the TME, characterized by high IL-10 and low granzyme B expression. These findings are consistent with previous reports that the accumulation of TAMs is associated with poor prognosis in cancer (43).
One of the biggest challenges in this study design was the limited number of cases from adjacent nontumor liver tissue, which was typically smaller in size and thus provided a limited number of immune cells that could be isolated for CyTOF analysis. Therefore, in the majority of the analyses, the NILs exhibited the lowest number of cases compared with TILs or PBMCs. Despite this caveat, most of the data were independently validated by other in vitro assays or by conventional flow cytometry, and in all cases, paired statistical tests were performed to validate the findings from individual patients rather than pooling. Most importantly, the unbiased and nonsupervised nature of our CyTOF analysis using statistical tests to screen and guide subsequent discovery is a method of data analysis showcased by our study, which to the best of our knowledge has not been previously shown in other reported CyTOF data analysis pipelines.
In summary, our data provide evidence of a dynamic immune gradient that is dominated by immune subsets that are progressively more exhausted and immunosuppressive. Our holistic approach has emphasized two main points: (i) immune suppression is microenvironment specific, as immune competency is generally maintained in the periphery or in the adjacent NTME with chronic inflammation; and (ii) the relevant immune subsets follow a chemokine gradient that is expressed in the tumor, and the cellular phenotypes are modified upon exposure to the TME. This chemokine gradient preferentially recruits and/or retains TAMs and NKRs in the tumor. These data may be readily translated into the context of immune monitoring and accurate cancer staging, and may be applied to the design of novel therapeutic targets toward the cancer itself.
Supplementary Material
Acknowledgments
The authors thank Win Htut Oo, Fiona Ni Ni Moe, and Phang Su Ting (National Cancer Centre, Singapore) for coordination of the patient sample collection and assistance in obtaining patient consent; Dr. Davide Lucchesi, Dr. Lakshmi Ramakrishna, and Dr. Chin Teck Ng (SingHealth Translational Immunology and Inflammation Centre, STIIC) for their contributions in scientific discussion; and Insight Editing London for language editing of this manuscript prior to submission. This work was supported by the National Medical Research Council (NMRC), Singapore (MOHIAFCAT2001, TCR15Jun006, CIRG16may048, NMRC/CG RIE2015, NMRC/STaR/020/2013, NMRC/MOHIAFCAT2/005/2015, CIRg13nov032, and NMRC/MOHIAFCAT1-6003), Duke University–National University of Singapore Medical School, STIIC, and Biomedical Research Council (BMRC) (BMRC-EDB IAF: IAF311020 and SPF2014/005).
Footnotes
The authors declare no conflict of interest.
This article is a PNAS Direct Submission.
This article contains supporting information online at www.pnas.org/lookup/suppl/doi:10.1073/pnas.1706559114/-/DCSupplemental.
References
- 1.Fridman WH, Pagès F, Sautès-Fridman C, Galon J. The immune contexture in human tumours: Impact on clinical outcome. Nat Rev Cancer. 2012;12:298–306. doi: 10.1038/nrc3245. [DOI] [PubMed] [Google Scholar]
- 2.Makarova-Rusher OV, Medina-Echeverz J, Duffy AG, Greten TF. The yin and yang of evasion and immune activation in HCC. J Hepatol. 2015;62:1420–1429. doi: 10.1016/j.jhep.2015.02.038. [DOI] [PubMed] [Google Scholar]
- 3.Couzin-Frankel J. Breakthrough of the year 2013. Cancer immunotherapy. Science. 2013;342:1432–1433. doi: 10.1126/science.342.6165.1432. [DOI] [PubMed] [Google Scholar]
- 4.Errico A. Immunotherapy: PD-1-PD-L1 axis: Efficient checkpoint blockade against cancer. Nat Rev Clin Oncol. 2015;12:63. doi: 10.1038/nrclinonc.2014.221. [DOI] [PubMed] [Google Scholar]
- 5.Greten TF, Wang XW, Korangy F. Current concepts of immune based treatments for patients with HCC: From basic science to novel treatment approaches. Gut. 2015;64:842–848. doi: 10.1136/gutjnl-2014-307990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Hodi FS, et al. Improved survival with ipilimumab in patients with metastatic melanoma. N Engl J Med. 2010;363:711–723. doi: 10.1056/NEJMoa1003466. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Hong YP, Li ZD, Prasoon P, Zhang Q. Immunotherapy for hepatocellular carcinoma: From basic research to clinical use. World J Hepatol. 2015;7:980–992. doi: 10.4254/wjh.v7.i7.980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Callahan MK, Wolchok JD. At the bedside: CTLA-4- and PD-1-blocking antibodies in cancer immunotherapy. J Leukoc Biol. 2013;94:41–53. doi: 10.1189/jlb.1212631. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Ferlay J, et al. Cancer incidence and mortality worldwide: Sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2015;136:E359–E386. doi: 10.1002/ijc.29210. [DOI] [PubMed] [Google Scholar]
- 10.El-Serag HB. Hepatocellular carcinoma. N Engl J Med. 2011;365:1118–1127. doi: 10.1056/NEJMra1001683. [DOI] [PubMed] [Google Scholar]
- 11.Kudo M. Immune checkpoint blockade in hepatocellular carcinoma. Liver Cancer. 2015;4:201–207. doi: 10.1159/000367758. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Chew V, et al. Chemokine-driven lymphocyte infiltration: An early intratumoural event determining long-term survival in resectable hepatocellular carcinoma. Gut. 2012;61:427–438. doi: 10.1136/gutjnl-2011-300509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Chew V, et al. Inflammatory tumour microenvironment is associated with superior survival in hepatocellular carcinoma patients. J Hepatol. 2010;52:370–379. doi: 10.1016/j.jhep.2009.07.013. [DOI] [PubMed] [Google Scholar]
- 14.Hernandez-Gea V, Toffanin S, Friedman SL, Llovet JM. Role of the microenvironment in the pathogenesis and treatment of hepatocellular carcinoma. Gastroenterology. 2013;144:512–527. doi: 10.1053/j.gastro.2013.01.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Schmidt N, Flecken T, Thimme R. Tumor-associated antigen specific CD8(+) T cells in hepatocellular carcinoma: A promising target for immunotherapy. OncoImmunology. 2014;3:e954919. doi: 10.4161/21624011.2014.954919. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Blank C, Gajewski TF, Mackensen A. Interaction of PD-L1 on tumor cells with PD-1 on tumor-specific T cells as a mechanism of immune evasion: Implications for tumor immunotherapy. Cancer Immunol Immunother. 2005;54:307–314. doi: 10.1007/s00262-004-0593-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Han A, Glanville J, Hansmann L, Davis MM. Linking T-cell receptor sequence to functional phenotype at the single-cell level. Nat Biotechnol. 2014;32:684–692. doi: 10.1038/nbt.2938. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Lai L, Ong R, Li J, Albani S. A CD45-based barcoding approach to multiplex mass-cytometry (CyTOF) Cytometry A. 2015;87:369–374. doi: 10.1002/cyto.a.22640. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Finck R, et al. Normalization of mass cytometry data with bead standards. Cytometry A. 2013;83:483–494. doi: 10.1002/cyto.a.22271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Shekhar K, Brodin P, Davis MM, Chakraborty AK. Automatic Classification of Cellular Expression by Nonlinear Stochastic Embedding (ACCENSE) Proc Natl Acad Sci USA. 2014;111:202–207. doi: 10.1073/pnas.1321405111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Garnelo M, et al. Interaction between tumour-infiltrating B cells and T cells controls the progression of hepatocellular carcinoma. Gut. 2017;66:342–351. doi: 10.1136/gutjnl-2015-310814. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Carluccio S, et al. Generation of tumor-specific cytotoxic T-lymphocytes from the peripheral blood of colorectal cancer patients for adoptive T-cell transfer. J Cell Physiol. 2015;230:1457–1465. doi: 10.1002/jcp.24886. [DOI] [PubMed] [Google Scholar]
- 23.Chen KJ, et al. Intratumoral regulatory T cells alone or in combination with cytotoxic T cells predict prognosis of hepatocellular carcinoma after resection. Med Oncol. 2012;29:1817–1826. doi: 10.1007/s12032-011-0006-x. [DOI] [PubMed] [Google Scholar]
- 24.Wilke CM, Wu K, Zhao E, Wang G, Zou W. Prognostic significance of regulatory T cells in tumor. Int J Cancer. 2010;127:748–758. doi: 10.1002/ijc.25464. [DOI] [PubMed] [Google Scholar]
- 25.Shin JI, Ha SJ. Regulatory T cells-an important target for cancer immunotherapy. Nat Rev Clin Oncol. 2014;11:307. doi: 10.1038/nrclinonc.2013.208-c1. [DOI] [PubMed] [Google Scholar]
- 26.Lowther DE, et al. 2016. PD-1 marks dysfunctional regulatory T cells in malignant gliomas. JCI Insight 1(5)
- 27.Park HJ, et al. PD-1 upregulated on regulatory T cells during chronic virus infection enhances the suppression of CD8+ T cell immune response via the interaction with PD-L1 expressed on CD8+ T cells. J Immunol. 2015;194:5801–5811. doi: 10.4049/jimmunol.1401936. [DOI] [PubMed] [Google Scholar]
- 28.Kao C, et al. Transcription factor T-bet represses expression of the inhibitory receptor PD-1 and sustains virus-specific CD8+ T cell responses during chronic infection. Nat Immunol. 2011;12:663–671. doi: 10.1038/ni.2046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Taylor A, et al. Glycogen synthase kinase 3 inactivation drives T-bet-mediated downregulation of co-receptor PD-1 to enhance CD8(+) cytolytic T cell responses. Immunity. 2016;44:274–286. doi: 10.1016/j.immuni.2016.01.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Yamazaki T, et al. CCR6 regulates the migration of inflammatory and regulatory T cells. J Immunol. 2008;181:8391–8401. doi: 10.4049/jimmunol.181.12.8391. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Kondo T, Takata H, Takiguchi M. Functional expression of chemokine receptor CCR6 on human effector memory CD8+ T cells. Eur J Immunol. 2007;37:54–65. doi: 10.1002/eji.200636251. [DOI] [PubMed] [Google Scholar]
- 32.Rivino L, et al. CCR6 is expressed on an IL-10-producing, autoreactive memory T cell population with context-dependent regulatory function. J Exp Med. 2010;207:565–577. doi: 10.1084/jem.20091021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Acosta-Rodriguez EV, et al. Surface phenotype and antigenic specificity of human interleukin 17-producing T helper memory cells. Nat Immunol. 2007;8:639–646. doi: 10.1038/ni1467. [DOI] [PubMed] [Google Scholar]
- 34.Schutyser E, Struyf S, Van Damme J. The CC chemokine CCL20 and its receptor CCR6. Cytokine Growth Factor Rev. 2003;14:409–426. doi: 10.1016/s1359-6101(03)00049-2. [DOI] [PubMed] [Google Scholar]
- 35.Boyle ST, Faulkner JW, McColl SR, Kochetkova M. The chemokine receptor CCR6 facilitates the onset of mammary neoplasia in the MMTV-PyMT mouse model via recruitment of tumor-promoting macrophages. Mol Cancer. 2015;14:115. doi: 10.1186/s12943-015-0394-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Liao KL, Bai XF, Friedman A. The role of CD200-CD200R in tumor immune evasion. J Theor Biol. 2013;328:65–76. doi: 10.1016/j.jtbi.2013.03.017. [DOI] [PubMed] [Google Scholar]
- 37.Kretz-Rommel A, et al. CD200 expression on tumor cells suppresses antitumor immunity: New approaches to cancer immunotherapy. J Immunol. 2007;178:5595–5605. doi: 10.4049/jimmunol.178.9.5595. [DOI] [PubMed] [Google Scholar]
- 38.Djenidi F, et al. CD8+CD103+ tumor-infiltrating lymphocytes are tumor-specific tissue-resident memory T cells and a prognostic factor for survival in lung cancer patients. J Immunol. 2015;194:3475–3486. doi: 10.4049/jimmunol.1402711. [DOI] [PubMed] [Google Scholar]
- 39.Kremsdorf D, Soussan P, Paterlini-Brechot P, Brechot C. Hepatitis B virus-related hepatocellular carcinoma: Paradigms for viral-related human carcinogenesis. Oncogene. 2006;25:3823–3833. doi: 10.1038/sj.onc.1209559. [DOI] [PubMed] [Google Scholar]
- 40.Nakamoto Y, Guidotti LG, Kuhlen CV, Fowler P, Chisari FV. Immune pathogenesis of hepatocellular carcinoma. J Exp Med. 1998;188:341–350. doi: 10.1084/jem.188.2.341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Ye B, et al. T-cell exhaustion in chronic hepatitis B infection: Current knowledge and clinical significance. Cell Death Dis. 2015;6:e1694. doi: 10.1038/cddis.2015.42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Fan R, et al. T-bet expression in CD8+ T cells associated with chronic hepatitis B virus infection. Virol J. 2016;13:14. doi: 10.1186/s12985-016-0473-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Zhang QW, et al. Prognostic significance of tumor-associated macrophages in solid tumor: A meta-analysis of the literature. PLoS One. 2012;7:e50946. doi: 10.1371/journal.pone.0050946. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.