Abstract
Interferon-induced transmembrane protein 3 (IFITM3) is a cellular endosome- and lysosome-localized protein that restricts numerous virus infections. IFITM3 is activated by palmitoylation, a lipid posttranslational modification. Palmitoylation of proteins is primarily mediated by zinc finger DHHC domain–containing palmitoyltransferases (ZDHHCs), but which members of this enzyme family can modify IFITM3 is not known. Here, we screened a library of human cell lines individually lacking ZDHHCs 1–24 and found that IFITM3 palmitoylation and its inhibition of influenza virus infection remained strong in the absence of any single ZDHHC, suggesting functional redundancy of these enzymes in the IFITM3-mediated antiviral response. In an overexpression screen with 23 mammalian ZDHHCs, we unexpectedly observed that more than half of the ZDHHCs were capable of increasing IFITM3 palmitoylation with ZDHHCs 3, 7, 15, and 20 having the greatest effect. Among these four enzymes, ZDHHC20 uniquely increased IFITM3 antiviral activity when both proteins were overexpressed. ZDHHC20 colocalized extensively with IFITM3 at lysosomes unlike ZDHHCs 3, 7, and 15, which showed a defined perinuclear localization pattern, suggesting that the location at which IFITM3 is palmitoylated may influence its activity. Unlike knock-out of individual ZDHHCs, siRNA-mediated knockdown of both ZDHHC3 and ZDHHC7 in ZDHHC20 knock-out cells decreased endogenous IFITM3 palmitoylation. Overall, our results demonstrate that multiple ZDHHCs can palmitoylate IFITM3 to ensure a robust antiviral response and that ZDHHC20 may serve as a particularly useful tool for understanding and enhancing IFITM3 activity.
Keywords: influenza virus, innate immunity, interferon, post-translational modification (PTM), protein palmitoylation, DHHC, IFITM, IFITM3, ZDHHC, palmitoyltransferase
Introduction
Interferon (IFN)-induced transmembrane protein 3 (IFITM3)2 is a cellular protein that blocks the membrane fusion of endocytosed viruses and thus the entry of their genomes into the cytosol (1–3). IFITM3 is particularly well characterized as a restriction factor active against influenza virus in cell culture and in vivo in both mice and humans (4–13). Additionally, infections with Chikungunya virus, Eastern equine encephalitis virus, and West Nile virus are increased in severity in IFITM3 KO mice (14, 15). IFITM3 is thought to alter membrane properties, including rigidity and curvature, providing a nonspecific mechanism for inhibiting a broad array of viruses (3, 16). We recently found that an amphipathic helix within IFITM3 is required for blocking membrane fusion mediated by viral proteins, which is consistent with the common ability of amphipathic helices to induce membrane curvature (17). Adjacent to this helix are palmitoylated cysteines (S-palmitoylated) that are required for antiviral activity (5, 9, 18–20). Palmitoylation is a 16-carbon lipid posttranslational modification that increases protein hydrophobicity and targets proteins or protein domains to lipid membranes (21). The close proximity of palmitoylation sites to the IFITM3 amphipathic helix suggests that the function of palmitoylation is to target or anchor this helix to membranes (17). Here, we examine the ability of cellular enzymes to add this critical posttranslational modification onto IFITM3.
We first reported IFITM3 palmitoylation upon its identification in a chemoproteomic screen for acylated proteins (5). Modification was mapped to three cysteines (amino acid positions 71, 72, and 105) with Cys-72 being the most important for promoting antiviral activity (5, 20). These cysteines are among the most conserved residues among all IFITM isoforms from diverse species, including bacterial IFITMs, which are thought to be primitive ancestors of the mammalian IFITMs (22). Indeed, the closely related IFITMs 1 and 2 are also palmitoylated (5, 19) as are more distantly related proteins such as IFITM5 and Syndig1, which have non-antiviral, but palmitoylation-dependent, functions in bone morphogenesis and neuronal development, respectively (23, 24). Furthermore, IFITMs from mycobacteria are palmitoylated when expressed in human cells (25). Conservation of this posttranslational modification throughout evolution further highlights its critical importance in the function of IFITM3 and other IFITMs.
S-Palmitoylation of proteins is typically mediated by members of a family of enzymes known as the zinc finger DHHC (Asp-His-His-Cys) domain–containing palmitoyltransferases (ZDHHCs) (26–28). There are at least 23 of these enzymes in mice and humans. Among the hundreds of known palmitoylated proteins, only a limited number of substrate/ZDHHC pairings have been identified to date (27). For example, ZDHHC9 is essential for palmitoylation of the Ras proto-oncogene (29), whereas ZDHHC6 is the primary palmitoyltransferase for the endoplasmic reticulum chaperone Calnexin (30). The available published data suggest that ZDHHCs have largely distinct substrate specificities (27). Through genetic studies in humans or mice, individual ZDHHCs have been implicated in neurological disorders, hair loss, and cancer (27), although which ZDHHCs play a role in immunity, including those that are involved in activating IFITM3, remain unknown (31). Here, we show that the absence of any individual human ZDHHC does not result in a defect in IFITM3 palmitoylation or the antiviral response following IFN stimulation, indicating that redundancy of endogenous ZDHHCs in mediating IFITM3 palmitoylation maintains robust IFITM3 activity. Furthermore, we screened overexpressed ZDHHCs and found that numerous members of this enzyme family are indeed able to palmitoylate IFITM3 and that ZDHHC20 is particularly adept at modifying IFITM3 and enhancing its antiviral activity. Unlike individual knock-out cells, knockdowns of additional ZDHHCs in the context of ZDHHC20 KO cells resulted in decreased IFITM3 palmitoylation, confirming a role for ZDHHC20 in palmitoylating endogenous IFITM3.
Results
Palmitoylation of human IFITM3 is not dependent on a single ZDHHC
Given the importance of palmitoylation for inhibition of influenza virus by IFITM3 (5, 18, 20), we sought to determine whether loss of any individual ZDHHC would decrease IFITM3 palmitoylation and activity. Because siRNA knockdowns of individual candidate ZDHHCs in A549 cells provided us with inconclusive results, which we presumed were due to imperfect knockdowns (Fig. S1), we purchased a commercial library of HAP1 cell lines individually lacking the human ZDHHCs 1–24 for further testing. These cells were generated using clustered regularly interspaced short palindromic repeats (CRISPR)/Cas9 technology to introduce deletions and frameshifts early in each ZDHHC gene occurring prior to the coding sequence for their catalytic DHHC motifs. Mutations in the ZDHHC genes were validated by DNA sequencing. Because confirmed antibodies against most of the ZDHHCs are not available, we spot checked specific cell lines based on the availability of well characterized antibodies for the individual ZDHHCs. Indeed, we confirmed complete knock-out of ZDHHCs 6, 7, 9, and 20, providing additional confidence in the utility of this library of cell lines (Fig. S2). Furthermore, we confirmed that IFITM3 is expressed and antivirally active in HAP1 cells by measuring IFITM3 expression and infection rates in WT and IFITM3 KO cells with and without type I IFN treatment (Fig. 1, A and B). Indeed, IFITM3 KO cells were significantly more susceptible to influenza virus infection than WT cells even after IFNβ treatment, indicating that, as expected, IFITM3 is essential for the antiviral response following IFN stimulation in HAP1 cells (Fig. 1B). We next examined endogenous IFITM3 palmitoylation in the library of ZDHHC KO cell lines after treatment with IFNβ to induce maximal IFITM3 expression. Briefly, cells were metabolically labeled with the alk-16 chemical reporter of protein palmitoylation, and immunoprecipitated IFITM3 was subjected to reaction via click chemistry with azidorhodamine to allow fluorescent visualization of palmitoylation by fluorescence gel scanning (5, 32–34). Strong IFITM3 palmitoylation signal was observed for each of the knock-out cell lines, indicating that, in agreement with our siRNA knockdown studies (Fig. S1), no individual human ZDHHC is absolutely required for IFITM3 palmitoylation (Fig. 1C). We further examined the susceptibility of the ZDHHC KO cell lines to influenza A virus infection. The individual cell lines showed similar rates of baseline infection, and treatment with IFNβ decreased infection similarly in each of the cell lines (Fig. 1D) in contrast to infection of IFITM3 KO cells in which much of the antiviral activity of IFNβ was lost (Fig. 1B). Overall, our results support the conclusion that redundancy among ZDHHCs provides the robust palmitoylation of IFITM3 required for its inhibition of influenza virus infection.
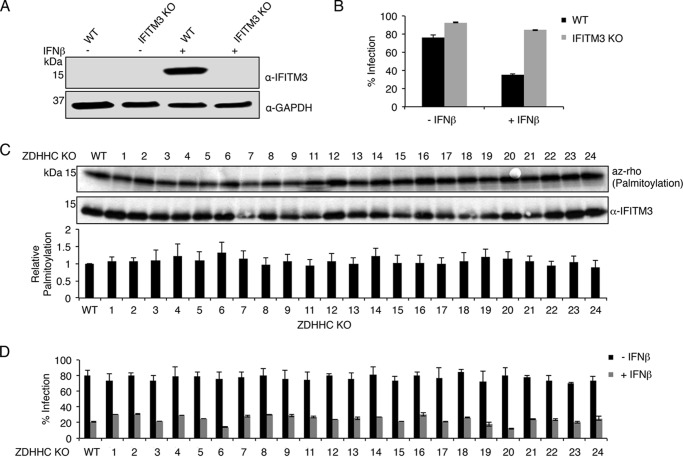
Figure 1.
Knock-out of individual human ZDHHCs does not affect IFITM3 palmitoylation or cellular susceptibility to influenza virus infection. A, IFITM3 expression in HAP1 WT and IFITM3 KO cells with or without overnight treatment with IFNβ. B, HAP1 WT and IFITM3 KO treated as in A were infected with influenza A virus (m.o.i., 1) for 24 h. Cells were stained with anti-influenza virus nucleoprotein antibodies to measure percentage of infection by flow cytometry. Average infection percentages of triplicate samples from a representative experiment of more than five experiments were graphed. Error bars represent S.D. C, the indicated HAP1 cell lines were treated with IFNβ overnight to induce expression of IFITM3. Cells were labeled for 1 h with 50 μm alk-16 and lysed prior to immunoprecipitation of IFITM3 and reaction with azidorhodamine (az-rho) via click chemistry for visualization of palmitoylation by fluorescence gel scanning. Western blotting for IFITM3 served as a loading control. The bar graph indicates normalized palmitoylation fluorescence averaged from five independent experiments. Error bars indicate S.D. of the mean. D, the indicated HAP1 cell lines were treated with IFNβ overnight or left untreated. The cells were subsequently infected with influenza A virus (m.o.i., 1) for 24 h. Cells were stained with anti-influenza virus nucleoprotein antibodies to measure percentage of infection by flow cytometry. Average infection percentages from four independent experiments, each performed in triplicate, were graphed. Error bars represent S.D.
Multiple ZDHHCs can palmitoylate IFITM3 in cells
Overexpression of individual ZDHHCs has proven to be an effective method of increasing palmitoylation of specific proteins, thereby identifying ZDHHCs than can palmitoylate proteins of interest (35–41). We thus obtained expression plasmids for murine ZDHHCs that have been used previously by multiple groups for identifying ZDHHC/substrate pairs (referred to in previous publications as DHHCs 1–23) (35–41). We measured the ability of each ZDHHC to modify IFITM3 in an overexpression screen using the alk-16 chemical reporter of protein palmitoylation to measure IFITM3 palmitoylation (5, 32–34). Quantification and averaging of IFITM3 palmitoylation levels from four independent experiments revealed that numerous ZDHHC proteins can increase IFITM3 palmitoylation in cells (Fig. 2, A and B). In this series of experiments, ZDHHCs 1, 2, 5, 6, 9, 14, 23, 24, and 25 increased IFITM3 palmitoylation by 1.7- to 3.0-fold, and ZDHHCs 3, 7, 15, and 20 increased palmitoylation by more than 3-fold. (Fig. 2, A and B) (Note that the ZDHHC numbers refer to the modern nomenclature for these proteins and that both the classic DHHC and modern ZDHHC numbering is shown in Fig. 2 to allow easy comparison with past studies using these expression constructs). Importantly, although expression levels of the ZDHHCs varied, levels of palmitoylation did not correspond simply to their unique expression levels (Fig. 2, A and B). For example, ZDHHC20 is among those enzymes with the lowest expression but was able to robustly increase IFITM3 palmitoylation (Fig. 2, A and B). Similar results were obtained when overexpressing this panel of murine ZDHHCs with human IFITM3 with ZDHHC20 showing the greatest ability to enhance human IFITM3 palmitoylation (Fig. S3). Overexpression of human ZDHHC20 with human IFITM3 similarly increased IFITM3 palmitoylation (Fig. S4). Overall, our results indicate that more than half of the ZDHHCs can modify IFITM3 in cells, suggesting a higher degree of enzyme interaction promiscuity for IFITM3 compared with other palmitoylated protein substrates reported by us (41) and others previously (27, 29, 30). For example, we previously found that only four of these constructs increased CD86 palmitoylation (41). Phylogenetic tree analysis of the murine ZDHHCs indicates that enzymes from each of the major branches of ZDHHC evolution can modify IFITM3, possibly suggesting that this activity was present in a common ancestor of the mammalian ZDHHCs (Fig. S5).
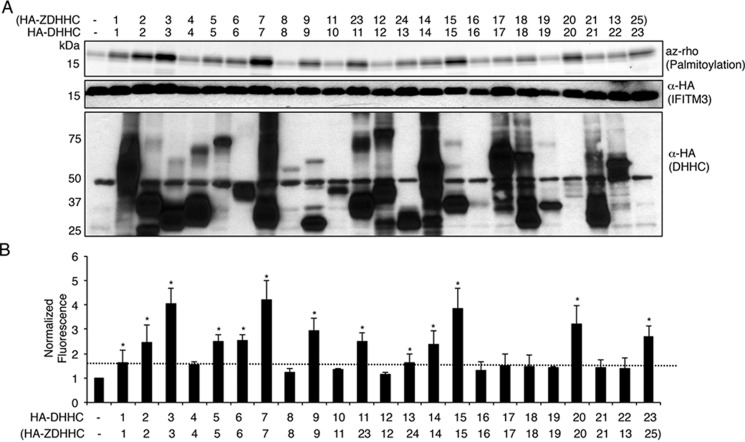
Figure 2.
Multiple ZDHHCs can palmitoylate IFITM3. A, HEK293T cells were cotransfected with murine HA-IFITM3 and the 23 HA-tagged murine DHHC constructs as indicated or control vector expressing GST (−). Parentheses indicate the modern ZDHHC nomenclature for these constructs. Cells were labeled for 1 h with 50 μm alk-16. IFITM3 was immunoprecipitated and subjected to reaction with azidorhodamine (az-rho) via click chemistry for visualization of palmitoylation by fluorescence gel scanning. Western blotting with anti-HA antibodies provided a loading control for IFITM3 and showed expression of the ZDHHCs. B, average quantitation of palmitoylation fluorescence normalized for protein loading from four independent experiments was graphed. IFITM3 palmitoylation when coexpressed with the GST control was normalized to a value of 1. Error bars indicate S.D. Asterisks indicate that palmitoylation was increased on average by greater than 1.7-fold over the control. Conditions indicated with an asterisk are also significantly different from the GST control as determined by Student's t test with p < 0.05 in all cases.
ZDHHC20 can increase antiviral activity of IFITM3
Consistent with our previous report that the palmitoylation sites on IFITM3 are not fully occupied when the protein is overexpressed in HEK293T cells (20), we observed that palmitoylation of transfected IFITM3 can be increased by co-overexpression of specific ZDHHCs (Fig. 2, A and B). Given that palmitoylation is required for IFITM3 antiviral activity (5, 20), we sought to determine whether increasing palmitoylation of IFITM3 also increases virus inhibition. We examined the ability of the four ZDHHCs that were able to most dramatically increase IFITM3 palmitoylation (ZDHHCs 3, 7, 15, and 20) for their ability to inhibit virus infections. When the enzymes were overexpressed alone in HEK293T cells, they did not alter the percentage of these cells infected with influenza A virus as compared with control cells (Fig. 3A). This result was expected given that HEK293T cells do not express detectable levels of endogenous IFITM3 (25). Alternatively, transfection of IFITM3 significantly decreased infection (Fig. 3A). Co-overexpression of ZDHHC 3, 7, or 15 with IFITM3 did not have any further effect on infection compared with IFITM3 alone, indicating that levels of palmitoylation of IFITM3 by endogenous ZDHHCs are adequate to achieve antiviral activity (Figs. 2, A and B, and 3A). However, co-overexpression of ZDHHC20 resulted in a reproducible and statistically significant enhancement in the ability of IFITM3 to restrict infection (Fig. 3A). This ability of ZDHHC20 was dependent upon its enzymatic activity as mutation of its catalytic cysteine to alanine (ZDHHC20-C156A) eliminated its enhancement of IFITM3 palmitoylation (Fig. 3B) as well as its enhancement of virus restriction (Fig. 3A). These results suggest that palmitoylation of IFITM3 by specific overexpressed ZDHHCs can provide nuanced effects on antiviral activity.
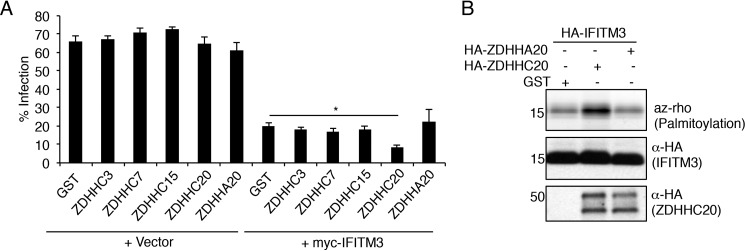
Figure 3.
ZDHHC20 overexpression increases IFITM3 antiviral activity. A, HEK293T cells were cotransfected with the indicated HA-tagged ZDHHCs or GST control plus either vector control or myc-IFITM3. ZDHHA20 refers to the ZDHHC20 C156A catalytic mutant. After overnight transfection, cells were infected with influenza virus (m.o.i., 2.5) for 6 h, collected, and stained with anti-influenza nucleoprotein, anti-myc, and anti-HA to determine percentage of infection in the transfected cells by flow cytometry. Data shown are average infection percentages from at least three independent experiments, each done in triplicate. Error bars represent S.D. of the mean. The asterisk indicates p < 0.01, Student's t test. B, HEK293T cells were transfected with the indicated plasmids and labeled for 1 h with 50 μm alk-16. Anti-HA immunoprecipitation of cell lysates followed by reaction with azidorhodamine (az-rho) via click chemistry allowed visualization of palmitoylation by fluorescence gel scanning. Western blots served as a loading control for IFITM3 and as a control for expression of both WT and mutant HA-ZDHHC20.
ZDHHCs 7 and 20 palmitoylate three cysteines within IFITM3
Cys-72 of IFITM3 has been shown to be the most important palmitoylation site for antiviral activity among the three palmitoylated cysteines within IFITM3 (20). Thus, we tested whether ZDHHCs with distinct abilities to enhance IFITM3 antiviral activity were able to palmitoylate distinct cysteines within IFITM3. Utilizing double cysteine to alanine mutants of IFITM3 in which only a single intact cysteine remained along with overexpression of ZDHHC7 or ZDHHC20, we found that both of these ZDHHCs were capable of palmitoylating all three cysteines within IFITM3 (Fig. 4, A and B). Thus, we were not able to distinguish these enzymes based on their ability to palmitoylate specific IFITM3 cysteines, although we cannot rule out subtle effects that are not distinguishable by our pulse labeling palmitoylation assay or that the individual ZDHHCs show preference for individual cysteines in the context of WT IFITM3.
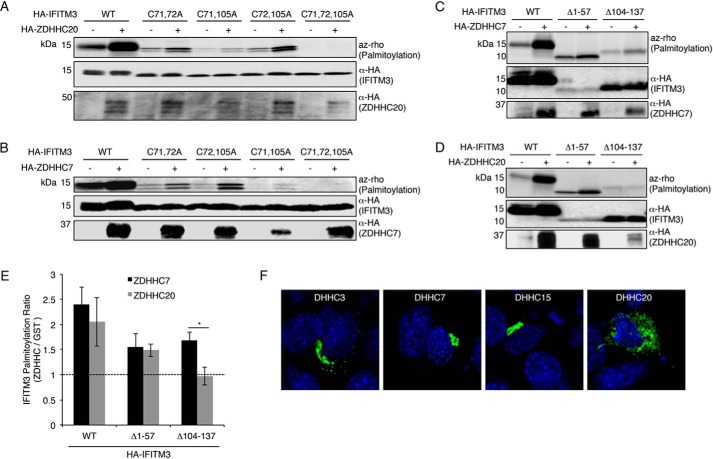
Figure 4.
ZDHHCs 20 and 7 can palmitoylate all three of IFITM3's cysteines but localize distinctly. A, HEK293Ts were cotransfected with either GST-expressing control vector or HA-ZDHHC20 and HA-IFITM3 or cysteine mutants lacking specific palmitoylation sites. Cells were then labeled with 50 μm alk-16, and immunoprecipitated IFITM3 was subjected to reaction with azidorhodamine (az-rho) via click chemistry for visualization of palmitoylation by fluorescence gel scanning. Anti-HA Western blotting confirmed comparable loading and the expression of the ZDHHCs. B, same as in A except using HA-ZDHHC7. C and D, same as in A and B except using IFITM3 constructs with the indicated truncations. E, palmitoylation signals from C and D were quantified and normalized relative to the anti-HA loading control for IFITM3 expression. For each IFITM3 construct, the normalized palmitoylation signal in the presence of the overexpressed ZDHHC was divided by the palmitoylation signal for the respective GST control. An average of results from four individual experiments is graphed. Error bars represent S.D. The asterisk indicates p < 0.0001, Student's t test. F, confocal microscopy of MEFs transfected overnight with the indicated HA-tagged ZDHHCs. Green represents HA staining (ZDHHCs), and blue represents DAPI (nuclei).
The IFITM3 C terminus is required for palmitoylation by ZDHHC20
To gain insights into the interactions between IFITM3 and distinct ZDHHCs, we examined the ability of ZDHHCs 7 and 20 to palmitoylate N-terminal and C-terminal truncation mutants of IFITM3. An N-terminal truncation mutant of IFITM3 lacking the first 57 amino acids of the protein (Δ1–57) was poorly expressed relative to full-length IFITM3 but was still robustly palmitoylated, and this palmitoylation was increased by expression of ZDHHC7 and ZDHHC20. (Fig. 4, C–E). In contrast, a C-terminal truncation mutant lacking the last 33 amino acids of the protein (Δ104–137) showed the weakest palmitoylation signal relative to its expression, and its palmitoylation signal could be increased by ZDHHC7 but not by ZDHHC20 (Fig. 4, C–E). Interestingly, in each of our experiments, we observed that both ZDHHC7 and ZDHHC20 showed weaker detection when they were coexpressed with a poorly palmitoylated mutant of IFITM3 as compared with when they were coexpressed with WT IFITM3 (Fig. 4, A–D). This may suggest that interactions with substrate proteins stabilize ZDHHCs. Overall, experiments in Fig. 4, C and D, indicate that the C terminus of IFITM3 is required for palmitoylation of IFITM3 by ZDHHC20 but not for palmitoylation by ZDHHC7, which is thought to be among the most promiscuous ZDHHCs in terms of substrate specificity (27).
ZDHHC20 colocalizes extensively with IFITM3 at LAMP1-positive compartments
We examined the localization of ZDHHCs 3, 7, 15, and 20 by confocal microscopy and found that ZDHHCs 3, 7, and 15 are localized to a perinuclear region consistent with Golgi localization previously reported for these enzymes (42), whereas ZDHHC20 is widely dispersed throughout the cell (Fig. 4F). Given the distinct cellular distributions that we observed for the different ZDHHCs, we examined colocalization of these ZDHHCs with IFITM3. IFITM3 partially localized with ZDHHCs 3, 7, and 15 and localized extensively with ZDHHC20 (Fig. 5A). Quantification of overlap between IFITM3 and these ZDHHCs in multiple cells confirmed that IFITM3 shows significantly greater colocalization with ZDHHC20 than with the other three ZDHHCs (Fig. 5B). Given that IFITM3 is well characterized to localize to lysosomes (2, 18, 25, 43, 44), we performed LAMP1, ZDHHC20, and IFITM3 costaining, and found that IFITM3 colocalizes with ZDHHC20 in LAMP1-positive lysosomal compartments (Fig. 5C). Taken together with infection data (Fig. 3), these localization results suggest that enhanced palmitoylation of IFITM3 at lysosomes may be particularly beneficial for antiviral activity.
Figure 5.
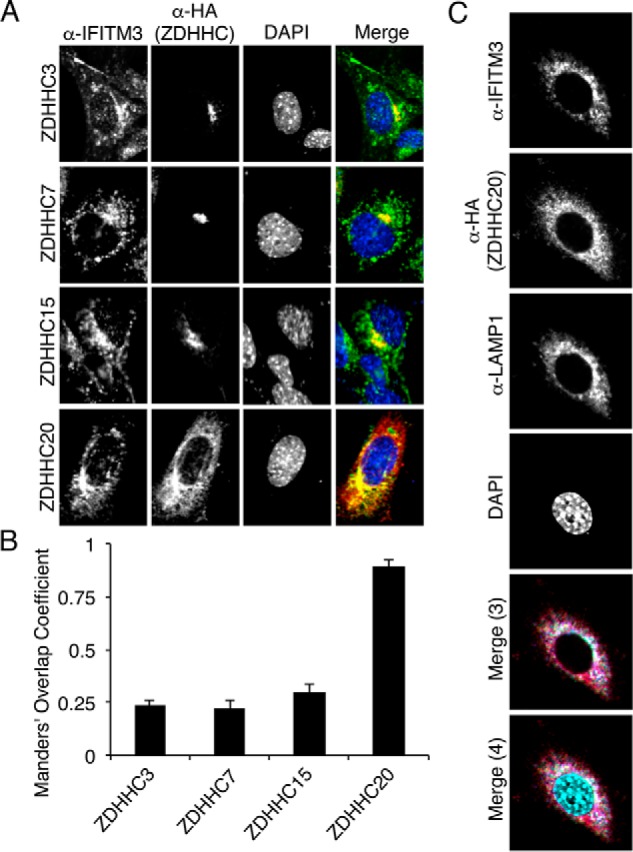
ZDHHC20 colocalizes extensively with IFITM3. A, MEFs were transfected overnight with the indicated HA-ZDHHC constructs and stimulated with IFNα for 8 h to induce IFITM3 expression. Cells were then stained with anti-HA (ZDHHC), anti-IFITM3, and DAPI for confocal microscopy imaging. B, overlap of IFITM3 with each ZDHHC was quantified from images as in A using the JACoP plugin for ImageJ software. At least 30 cells from two independent experiments were quantified and averaged. Error bars represent S.D. of the mean. Statistical significance for differences between ZDHHC20 and each of the other three ZDHHCs was determined by Student's t test, p < 0.0001. C, MEFs were transfected overnight with HA-ZDHHC20 construct and then stimulated with IFNα for 8 h to induce IFITM3 expression. Cells were then stained with anti-HA (ZDHHC), anti-IFITM3, anti-LAMP1, and DAPI for confocal microscopy imaging. Merge (3) and Merge (4) indicate three- and four-color merged images, respectively.
Depletion of ZDHHCs 3, 7, and 20 decreases IFITM3 palmitoylation in cells
Our experiments with ZDHHC KO cells indicated that more than one ZDHHC contributes to IFITM3 palmitoylation in cells (Fig. 1C). Likewise, our ZDHHC overexpression screen indicated that multiple enzymes can increase IFITM3 palmitoylation (Fig. 2). We thus posited that depletion of more than one ZDHHC would be required to observe a reduction in IFITM3 palmitoylation. Given the interesting phenotypes of ZDHHC20 in our overexpression experiments, we chose to knock down additional ZDHHCs in ZDHHC20 KO cells using siRNAs. We knocked down ZDHHC3 and ZDHHC7 based on the strong ability of these enzymes to increase IFITM3 palmitoylation when overexpressed (Fig. 2) and on previous reports suggesting that these are the most active and/or promiscuous ZDHHCs (27) and that they are ubiquitously expressed (42, 45). Knockdown of ZDHHC3 or ZDHHC7 in the absence of ZDHHC20 resulted in decreased IFITM3 palmitoylation, and this was further decreased when both enzymes were knocked down simultaneously (Fig. 6, A and B). Notably, even upon depletion of three ZDHHCs, IFITM3 palmitoylation was still detectable, indicating that additional endogenous ZDHHCs may also palmitoylate IFITM3. Nonetheless, these results indicate that multiple ZDHHCs palmitoylate endogenous IFITM3, including ZDHHCs 3, 7, and 20.
Figure 6.
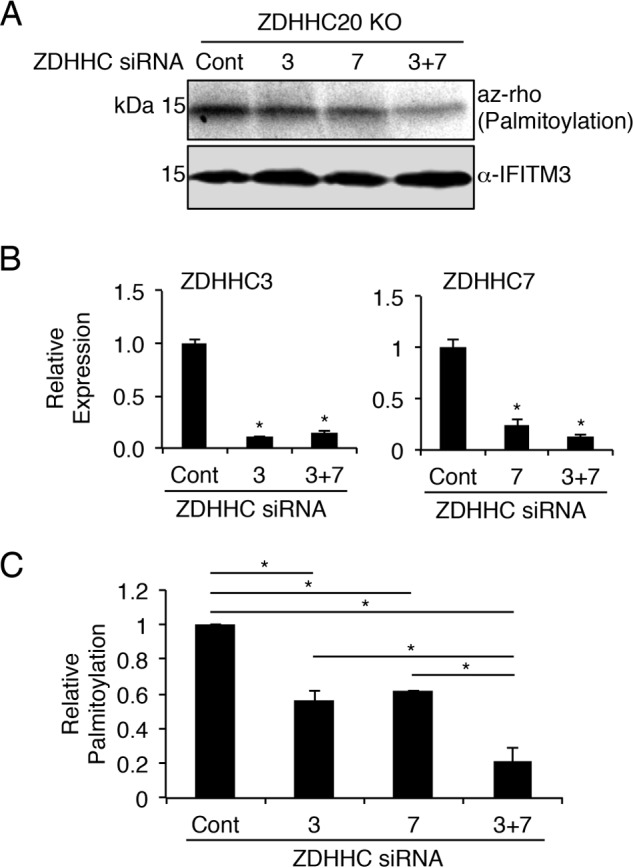
Depletion of ZDHHCs 3 and 7 in ZDHHC20 knock-out cells decreases IFITM3 palmitoylation. A–C, ZDHHC20 KO HAP1 cells were transfected with siRNAs against the indicated ZDHHCs or with negative control siRNAs. Cells were also treated with IFNβ overnight to induce expression of IFITM3. A, cells were labeled for 1 h with 50 μm alk-16 and lysed prior to immunoprecipitation of IFITM3 and reaction with azidorhodamine (az-rho) via click chemistry for visualization of palmitoylation by fluorescence gel scanning. Western blotting for IFITM3 served as a loading control. B, graphs represent knockdown efficiency of the indicated siRNAs from a representative experiment as determined using quantitative RT-PCR for the specific ZDHHC transcripts normalized to GAPDH transcript levels. The asterisk indicates p < 0.0001, Student's t test, compared with the control siRNA-treated samples. C, the bar graph indicates normalized palmitoylation fluorescence averaged from three independent experiments. Error bars indicate S.D. The asterisk indicates p < 0.0001, Student's t test. Cont, control.
Discussion
Although the activity of IFITM3 against influenza virus is negatively regulated by posttranslational modifications, including ubiquitination (6, 18), methylation (46), and phosphorylation (43, 47), palmitoylation is the only known posttranslational modification that increases IFITM3 antiviral activity (5, 18, 20, 31, 48). Thus, enzymes that add this modification are likely critical contributors to the IFN-induced immune response. Using an overexpression screen, we found an unexpected degree of redundancy among the ZDHHC enzymes for palmitoylation of IFITM3 by observing that 13 of 23 ZDHHCs increased IFITM3 palmitoylation in cells (Fig. 2, A and B). Similar published screens using the same plasmid constructs with different substrate proteins have generally revealed only a minimal number of ZDHHC candidates (35–41). For example, using these constructs in previous studies, our laboratory found that four ZDHHCs increased palmitoylation of CD86, and five ZDHHCs increased palmitoylation of Toll-like receptor 2 (41). Mapping of IFITM3-modifying enzymes onto a phylogenetic tree of the murine ZDHHCs suggests that palmitoylation of IFITM3 may be an ancient function maintained by members of nearly all of the branches of mammalian ZDHHC evolution (Fig. S5). This is further supported by previous evolutionary analysis of IFITMs suggesting that an IFITM gene was introduced into a primitive single-celled eukaryotic ancestor of mammals by horizontal gene transfer from bacteria (22). Our results may also suggest that ZDHHC redundancy as demonstrated by our KO cell experiments (Fig. 1, C and D) would safeguard against a loss of IFITM3 antiviral activity upon mutation of any single enzyme in humans.
Maintenance of redundant enzyme functionality upon gene duplication was traditionally difficult to rationalize given that non-essential genes are expected to be lost through the course of evolution. However, the “piggyback model” describes maintenance of a redundant overlapping protein function via coselection with a non-redundant function (49). The palmitoylation of IFITM3 by multiple ZDHHCs may occur through conservation of an ancestral activity, although each ZDHHC may also have unique functions, many of which remain undiscovered. For example, ZDHHC20 is able to palmitoylate IFITM3, and it is also uniquely essential among the ZDHHCs for the palmitoylation of the epidermal growth factor receptor, which was previously the only known substrate for this enzyme (50).
It is also interesting to note that we previously found that IFITM3 is not depalmitoylated to a significant extent in human cells (18) and that at least one cysteine is palmitoylated on nearly 100% of endogenous IFITM3 (20). This is in contrast to other palmitoylated proteins, such as the proto-oncogene Ras, which undergoes rapid cycles of palmitoylation and depalmitoylation that control its localization and signaling function (51). Interestingly, Ras palmitoylation is largely dependent on a single enzyme, ZDHHC9 (29), possibly suggesting that tight regulation of palmitoylation is most effectively achieved by utilizing a limited number of ZDHHCs. Likewise, the endoplasmic reticulum protein Calnexin is palmitoylated primarily by ZDHHC6 (30). Recent work shows that Calnexin palmitoylation is also precisely regulated with only a fraction of the protein being palmitoylated, allowing the palmitoylated and non-palmitoylated fractions to perform distinct functions depending on the needs of the cell (52, 53). It is tempting to speculate that evolution has largely removed functional redundancy of ZDHHCs for palmitoylating Ras and Calnexin because increasing the palmitoylation of these substrates may be detrimental to the health of cells, whereas, in contrast, robust palmitoylation of IFITM3 is beneficial for ensuring an optimal antiviral immune response.
Some ZDHHCs, such as the related proteins ZDHHCs 3 and 7, have been proposed to be highly active and promiscuous in their modification of proteins (27), consistent with their strong palmitoylation of IFITM3 (Fig. 2). Other ZDHHCs, such as ZDHHC20, are thought to have more stringent substrate selectivity. Our assays suggest that ZDHHCs 7 and 20 can both robustly modify IFITM3 and that each can modify the three individual cysteines within the IFITM3 sequence (Fig. 4, A and B). In contrast, ZDHHC20 uniquely increased antiviral activity of IFITM3 upon overexpression of both proteins (Fig. 3A). Distinct localization of ZDHHC20 with IFITM3 at lysosomes as compared with ZDHHC7 correlated with this ability (Figs. 4C and 5), although knock-out of ZDHHC20 alone did not result in decreased IFITM3 palmitoylation or activity (Fig. 1, C and D). Thus, other ZDHHCs appear to be redundant in activating endogenous IFITM3 in human cells. Indeed knockdown of ZDHHCs 3 and 7 in the context of ZDHHC20 KO resulted in a decrease in IFITM3 palmitoylation. Overall, our results indicate that ZDHHCs 3, 7, and 20 contribute to endogenous IFITM3 palmitoylation and that ZDHHC20 may serve as a tool for understanding IFITM3/ZDHHC interactions and potentially for enhancing IFITM3 antiviral activity.
Experimental procedures
Cell culture, transfections, and treatments
HEK293T cells, A549 cells, and MEFs were grown in DMEM with 10% Equafetal FBS (Atlas Biologicals). HAP1 KO cell lines were purchased from Horizon Discoveries/Life Technologies and grown in Iscove's modified Dulbecco's medium with 10% Equafetal FBS. All cells were maintained at 37 °C with 5% CO2 in a humidified incubator. For Western blotting and infection, cells were plated at 90% confluence in either 6- or 12-well plates 24 h prior to transfection with 1–4 μg of plasmids using LipoJet (Signagen Laboratories) according to the manufacturer's instructions. Human and mouse IFITM3 plasmids were described previously (5, 18, 20, 47). Expression plasmids for the murine ZDHHCs were kindly provided by Dr. Masaki Fukata (National Institute for Physiological Sciences, Japan). The human ZDHHC20 plasmid was purchased from Origene. All siRNAs were SMARTpool siRNAs from Dharmacon and were transfected for 24 h using RNAiMax transfection reagent (Life Technologies) according to the manufacturer's instructions. For some experiments, MEFs were treated with IFNα2 (EBioscience) at a 1:1000 dilution for 8 h, and HAP1 cells and A549 cells were treated with IFNβ (BEI Resources) at a 1:100 dilution overnight.
Virus infection and flow cytometry
Influenza virus strain PR8, A/PR/8/34 (H1N1), was propagated in 10-day-old embryonated chicken eggs (Charles River) for 48 h at 37 °C as described previously (54, 55). HEK293T and HAP1 cells were infected with influenza PR8 for 24 h at a multiplicity of 2.5 and 1, respectively. Infected cells were fixed in 4% paraformaldehyde for 10 min, permeabilized with 0.1% Triton X-100 for 10 min, and then blocked with 2% FBS in PBS for 10 min prior to staining. Influenza virus–infected cells were stained with either anti-influenza nucleoprotein (Abcam) primary antibody followed by mouse Alexa Fluor 647 secondary antibody (Life Technologies) or with Alexa Fluor-conjugated anti-influenza nucleoprotein produced using a 100-μg antibody labeling kit (Life Technologies). For detection of HA-ZDHHC and myc-IFITM3 cotransfected cells by flow cytometry, cells were costained with directly labeled anti-HA (Covance) and anti-myc (Clontech) antibodies. All antibodies were diluted in 0.1% Triton X-100 in PBS, and cells were stained for 20 min. Each antibody staining step was followed by three washes with 0.1% Triton X-100 in PBS to remove unbound antibody. All samples were run on a FACSCanto II flow cytometer (BD Biosciences), and results were analyzed using FlowJo software.
Metabolic labeling, immunoprecipitation, and click chemistry
All palmitoylation experiments followed a standard, previously published protocol (5, 32, 33, 56). Briefly, transfected or IFN-treated cells were metabolically labeled using 50 μm alk-16 or DMSO control in DMEM containing 2% charcoal-stripped FBS (Thermo Fisher) for 1–4 h. Cells were isolated and lysed in Brij buffer containing 50 mm triethanolamine, 150 mm NaCl, 1% Brij 97, and EDTA-free protease inhibitor (Roche Applied Science). Labeled proteins were immunoprecipitated using anti-HA- or anti-myc-agarose beads (Sigma) or anti-IFITM3 (Proteintech) with Protein G-agarose for at least 1 h at 4 °C and subsequently washed three times with lysis buffer to remove unbound proteins. This was followed by click chemistry reactions for 1 h using final concentrations of azidorhodamine (100 μm), tris(2-carboxyethyl)phosphine hydrochloride (TCEP) (1 mm), tris[(1-benzyl-1H-1,2,3-triazol-4-yl)methyl]amine (TBTA) (100 μm), and CuSO4 (1 mm) as described previously (5, 32, 33, 56). Fluorescence labeling was visualized on an Amersham Biosciences Typhoon 9410 gel scanner.
Fluorescence microscopy
For microscopy, MEFs were plated at 50% confluence on glass coverslips (Fisher Scientific) in 12-well tissue culture dishes and transfected with plasmid overnight. The next day, cells were then washed once with PBS, fixed using 4% paraformaldehyde for 10 min, permeabilized with 0.1% Triton X-100 in PBS for 10 min, and blocked in 2% FBS in PBS for 10 min. Cells were stained with primary antibodies anti-IFITM3 (Fragilis, Abcam), anti-HA (Covance), and anti-LAMP1 (Santa Cruz Biotechnology) diluted 1:1000 in 0.1% Triton X-100 in PBS for 20 min. Cells were then treated with Alexa Fluor-labeled secondary antibodies (Life Technologies) diluted 1:1000 in 0.1% Triton X-100 in PBS for 20 min. Slides were washed at least three times with 0.1% Triton X-100 in PBS between antibody staining steps. Coverslips were mounted on microscope slides with ProLong Gold antifade mountant with DAPI (Thermo Fisher) and allowed to cure overnight. Images were captured using a Fluoview FV10i confocal microscope (Olympus). Manders' overlap coefficient quantifications were performed using the JACoP plugin for ImageJ software (57).
Western blotting
For Western blotting, cells were lysed in Brij buffer, and protein samples were run on precast Criterion TGX 4–24% gels (Bio-Rad). Primary antibodies for human IFITM3 (Proteintech), HA tag (Covance), myc tag (Clontech), and GAPDH (Invitrogen), were diluted 1:1000 in 0.1% Tween 20 in PBS. Similarly, antibodies for ZDHHC6, ZDHHC7, and ZDHHC9 (Abcam) were diluted 1:500, and ZDHHC20 antibody (Sigma) was diluted 1:100. Secondary antibodies anti-mouse IgG HRP (Millipore) and anti-rabbit IgG HRP (Cell Signaling Technology) were diluted 1:10,000 in 0.1% Tween 20 in PBS.
Author contributions
J. S. Y. and H. C. H. conceived the study. J. S. Y. supervised the study. L. Z. performed and analyzed experiments in Fig. 1, C and D. J. C. H. performed experiments in Figs. 2 and 3. T. M. M. performed and analyzed experiments in Figs. 1, A and B, 4, 5, and 6 and replicated results from Figs. 1, C and D, 2, and 3. M. C. and A. D. K. performed experiments found in supporting figures and assisted in generation of plasmid reagents. T. M. M., J. S. Y., and H. C. H. analyzed data, generated figures, and wrote the manuscript. All authors reviewed the results and approved the final version of the manuscript.
Supplementary Material
This work was supported by National Institutes of Health Grant AI130110 (to J. S. Y.) and GM087544 (to H. C. H.), a Gilliam Fellowship for Advanced Study from the Howard Hughes Medical Institute (to T. M. M.), and The Ohio State University Systems and Integrative Biology Training Program funded by National Institutes of Health Grant GM068412 (to A. D. K.). The authors declare that they have no conflicts of interest with the contents of this article. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
This article contains Figs. S1–S5.
- IFITM
- interferon-induced transmembrane protein
- ZDHHC
- zinc finger DHHC domain–containing palmitoyltransferase
- MEF
- mouse embryonic fibroblast
- m.o.i.
- multiplicity of infection.
References
- 1. Desai T. M., Marin M., Chin C. R., Savidis G., Brass A. L., and Melikyan G. B. (2014) IFITM3 restricts influenza A virus entry by blocking the formation of fusion pores following virus-endosome hemifusion. PLoS Pathog. 10, e1004048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Feeley E. M., Sims J. S., John S. P., Chin C. R., Pertel T., Chen L. M., Gaiha G. D., Ryan B. J., Donis R. O., Elledge S. J., and Brass A. L. (2011) IFITM3 inhibits influenza A virus infection by preventing cytosolic entry. PLoS Pathog. 7, e1002337. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Li K., Markosyan R. M., Zheng Y. M., Golfetto O., Bungart B., Li M., Ding S., He Y., Liang C., Lee J. C., Gratton E., Cohen F. S., and Liu S. L. (2013) IFITM proteins restrict viral membrane hemifusion. PLoS Pathog. 9, e1003124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Brass A. L., Huang I. C., Benita Y., John S. P., Krishnan M. N., Feeley E. M., Ryan B. J., Weyer J. L., van der Weyden L., Fikrig E., Adams D. J., Xavier R. J., Farzan M., and Elledge S. J. (2009) The IFITM proteins mediate cellular resistance to influenza A H1N1 virus, West Nile virus, and dengue virus. Cell 139, 1243–1254 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Yount J. S., Moltedo B., Yang Y. Y., Charron G., Moran T. M., López C. B., and Hang H. C. (2010) Palmitoylome profiling reveals S-palmitoylation-dependent antiviral activity of IFITM3. Nat. Chem. Biol. 6, 610–614 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Chesarino N. M., McMichael T. M., and Yount J. S. (2015) E3 Ubiquitin Ligase NEDD4 promotes influenza virus infection by decreasing levels of the antiviral protein IFITM3. PLoS Pathog. 11, e1005095. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Everitt A. R., Clare S., Pertel T., John S. P., Wash R. S., Smith S. E., Chin C. R., Feeley E. M., Sims J. S., Adams D. J., Wise H. M., Kane L., Goulding D., Digard P., Anttila V., et al. (2012) IFITM3 restricts the morbidity and mortality associated with influenza. Nature 484, 519–523 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Bailey C. C., Huang I. C., Kam C., and Farzan M. (2012) Ifitm3 limits the severity of acute influenza in mice. PLoS Pathog. 8, e1002909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Compton A. A., Roy N., Porrot F., Billet A., Casartelli N., Yount J. S., Liang C., and Schwartz O. (2016) Natural mutations in IFITM3 modulate post-translational regulation and toggle antiviral specificity. EMBO Rep. 17, 1657–1671 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Zhang Y., Makvandi-Nejad S., Qin L., Zhao Y., Zhang T., Wang L., Repapi E., Taylor S., McMichael A., Li N., Dong T., and Wu H. (2015) Interferon-induced transmembrane protein-3 rs12252-C is associated with rapid progression of acute HIV-1 infection in Chinese MSM cohort. AIDS 29, 889–894 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Wang Z., Zhang A., Wan Y., Liu X., Qiu C., Xi X., Ren Y., Wang J., Dong Y., Bao M., Li L., Zhou M., Yuan S., Sun J., Zhu Z., et al. (2014) Early hypercytokinemia is associated with interferon-induced transmembrane protein-3 dysfunction and predictive of fatal H7N9 infection. Proc. Natl. Acad. Sci. U.S.A. 111, 769–774 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Xuan Y., Wang L. N., Li W., Zi H. R., Guo Y., Yan W. J., Chen X. B., and Wei P. M. (2015) IFITM3 rs12252 T>C polymorphism is associated with the risk of severe influenza: a meta-analysis. Epidemiol. Infect. 143, 2975–2984 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Allen E. K., Randolph A. G., Bhangale T., Dogra P., Ohlson M., Oshansky C. M., Zamora A. E., Shannon J. P., Finkelstein D., Dressen A., DeVincenzo J., Caniza M., Youngblood B., Rosenberger C. M., and Thomas P. G. (2017) SNP-mediated disruption of CTCF binding at the IFITM3 promoter is associated with risk of severe influenza in humans. Nat. Med. 23, 975–983 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Gorman M. J., Poddar S., Farzan M., and Diamond M. S. (2016) The interferon-stimulated gene Ifitm3 restricts West Nile virus infection and pathogenesis. J. Virol. 90, 8212–8225 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Poddar S., Hyde J. L., Gorman M. J., Farzan M., and Diamond M. S. (2016) The interferon-stimulated gene IFITM3 restricts infection and pathogenesis of arthritogenic and encephalitic alphaviruses. J. Virol. 90, 8780–8794 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Lin T. Y., Chin C. R., Everitt A. R., Clare S., Perreira J. M., Savidis G., Aker A. M., John S. P., Sarlah D., Carreira E. M., Elledge S. J., Kellam P., and Brass A. L. (2013) Amphotericin B increases influenza A virus infection by preventing IFITM3-mediated restriction. Cell Rep. 5, 895–908 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Chesarino N. M., Compton A. A., McMichael T. M., Kenney A. D., Zhang L., Soewarna V., Davis M., Schwartz O., and Yount J. S. (2017) IFITM3 requires an amphipathic helix for antiviral activity. EMBO Rep. 18, 1740–1751 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Yount J. S., Karssemeijer R. A., and Hang H. C. (2012) S-Palmitoylation and ubiquitination differentially regulate interferon-induced transmembrane protein 3 (IFITM3)-mediated resistance to influenza virus. J. Biol. Chem. 287, 19631–19641 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Hach J. C., McMichael T., Chesarino N. M., and Yount J. S. (2013) Palmitoylation on conserved and non-conserved cysteines of murine IFITM1 regulates its stability and anti-influenza A virus activity. J. Virol. 87, 9923–9927 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Percher A., Ramakrishnan S., Thinon E., Yuan X., Yount J. S., and Hang H. C. (2016) Mass-tag labeling reveals site-specific and endogenous levels of protein S-fatty acylation. Proc. Natl. Acad. Sci. U.S.A. 113, 4302–4307 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Blaskovic S., Blanc M., and van der Goot F. G. (2013) What does S-palmitoylation do to membrane proteins? FEBS J. 280, 2766–2774 [DOI] [PubMed] [Google Scholar]
- 22. Sällman Almén M., Bringeland N., Fredriksson R., and Schiöth H. B. (2012) The dispanins: a novel gene family of ancient origin that contains 14 human members. PLoS One 7, e31961. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Tsukamoto T., Li X., Morita H., Minowa T., Aizawa T., Hanagata N., and Demura M. (2013) Role of S-palmitoylation on IFITM5 for the interaction with FKBP11 in osteoblast cells. PLoS One 8, e75831. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Kaur I., Yarov-Yarovoy V., Kirk L. M., Plambeck K. E., Barragan E. V., Ontiveros E. S., and Díaz E. (2016) Activity-dependent palmitoylation controls SynDIG1 stability, localization, and function. J. Neurosci. 36, 7562–7568 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Melvin W. J., McMichael T. M., Chesarino N. M., Hach J. C., and Yount J. S. (2015) IFITMs from mycobacteria confer resistance to influenza virus when expressed in human cells. Viruses 7, 3035–3052 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Mitchell D. A., Vasudevan A., Linder M. E., and Deschenes R. J. (2006) Protein palmitoylation by a family of DHHC protein S-acyltransferases. J. Lipid Res. 47, 1118–1127 [DOI] [PubMed] [Google Scholar]
- 27. Greaves J., and Chamberlain L. H. (2011) DHHC palmitoyl transferases: substrate interactions and (patho)physiology. Trends Biochem. Sci. 36, 245–253 [DOI] [PubMed] [Google Scholar]
- 28. Linder M. E., and Jennings B. C. (2013) Mechanism and function of DHHC S-acyltransferases. Biochem. Soc. Trans. 41, 29–34 [DOI] [PubMed] [Google Scholar]
- 29. Swarthout J. T., Lobo S., Farh L., Croke M. R., Greentree W. K., Deschenes R. J., and Linder M. E. (2005) DHHC9 and GCP16 constitute a human protein fatty acyltransferase with specificity for H- and N-Ras. J. Biol. Chem. 280, 31141–31148 [DOI] [PubMed] [Google Scholar]
- 30. Lakkaraju A. K., Abrami L., Lemmin T., Blaskovic S., Kunz B., Kihara A., Dal Peraro M., and van der Goot F. G. (2012) Palmitoylated calnexin is a key component of the ribosome-translocon complex. EMBO J. 31, 1823–1835 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Yount J. S., Zhang M. M., and Hang H. C. (2013) Emerging roles for protein S-palmitoylation in immunity from chemical proteomics. Curr. Opin. Chem. Biol. 17, 27–33 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Charron G., Zhang M. M., Yount J. S., Wilson J., Raghavan A. S., Shamir E., and Hang H. C. (2009) Robust fluorescent detection of protein fatty-acylation with chemical reporters. J. Am. Chem. Soc. 131, 4967–4975 [DOI] [PubMed] [Google Scholar]
- 33. Yount J. S., Zhang M. M., and Hang H. C. (2011) Visualization and identification of fatty acylated proteins using chemical reporters. Curr. Protoc. Chem. Biol. 3, 65–79 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Yount J. S., Charron G., and Hang H. C. (2012) Bioorthogonal proteomics of 15-hexadecynyloxyacetic acid chemical reporter reveals preferential targeting of fatty acid modified proteins and biosynthetic enzymes. Bioorg. Med. Chem. 20, 650–654 [DOI] [PubMed] [Google Scholar]
- 35. Fukata M., Fukata Y., Adesnik H., Nicoll R. A., and Bredt D. S. (2004) Identification of PSD-95 palmitoylating enzymes. Neuron 44, 987–996 [DOI] [PubMed] [Google Scholar]
- 36. Fukata Y., Iwanaga T., and Fukata M. (2006) Systematic screening for palmitoyl transferase activity of the DHHC protein family in mammalian cells. Methods 40, 177–182 [DOI] [PubMed] [Google Scholar]
- 37. Tsutsumi R., Fukata Y., Noritake J., Iwanaga T., Perez F., and Fukata M. (2009) Identification of G protein α subunit-palmitoylating enzyme. Mol. Cell. Biol. 29, 435–447 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Sharma C., Yang X. H., and Hemler M. E. (2008) DHHC2 affects palmitoylation, stability, and functions of tetraspanins CD9 and CD151. Mol. Biol. Cell 19, 3415–3425 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Huang K., Sanders S., Singaraja R., Orban P., Cijsouw T., Arstikaitis P., Yanai A., Hayden M. R., and El-Husseini A. (2009) Neuronal palmitoyl acyl transferases exhibit distinct substrate specificity. FASEB J. 23, 2605–2615 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Pedram A., Razandi M., Deschenes R. J., and Levin E. R. (2012) DHHC-7 and -21 are palmitoylacyltransferases for sex steroid receptors. Mol. Biol. Cell 23, 188–199 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Chesarino N. M., Hach J. C., Chen J. L., Zaro B. W., Rajaram M. V., Turner J., Schlesinger L. S., Pratt M. R., Hang H. C., and Yount J. S. (2014) Chemoproteomics reveals Toll-like receptor fatty acylation. BMC Biol. 12, 91. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Ohno Y., Kihara A., Sano T., and Igarashi Y. (2006) Intracellular localization and tissue-specific distribution of human and yeast DHHC cysteine-rich domain-containing proteins. Biochim. Biophys. Acta 1761, 474–483 [DOI] [PubMed] [Google Scholar]
- 43. Jia R., Pan Q., Ding S., Rong L., Liu S. L., Geng Y., Qiao W., and Liang C. (2012) The N-terminal region of IFITM3 modulates its antiviral activity by regulating IFITM3 cellular localization. J. Virol. 86, 13697–13707 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Narayana S. K., Helbig K. J., McCartney E. M., Eyre N. S., Bull R. A., Eltahla A., Lloyd A. R., and Beard M. R. (2015) The interferon-induced transmembrane proteins, IFITM1, IFITM2, and IFITM3 inhibit hepatitis C virus entry. J. Biol. Chem. 290, 25946–25959 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Korycka J., Łach A., Heger E., Bogusławska D. M., Wolny M., Toporkiewicz M., Augoff K., Korzeniewski J., and Sikorski A. F. (2012) Human DHHC proteins: a spotlight on the hidden player of palmitoylation. Eur. J. Cell Biol. 91, 107–117 [DOI] [PubMed] [Google Scholar]
- 46. Shan Z., Han Q., Nie J., Cao X., Chen Z., Yin S., Gao Y., Lin F., Zhou X., Xu K., Fan H., Qian Z., Sun B., Zhong J., Li B., et al. (2013) Negative regulation of interferon-induced transmembrane protein 3 by SET7-mediated lysine monomethylation. J. Biol. Chem. 288, 35093–35103 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Chesarino N. M., McMichael T. M., Hach J. C., and Yount J. S. (2014) Phosphorylation of the antiviral protein interferon-inducible transmembrane protein 3 (IFITM3) dually regulates its endocytosis and ubiquitination. J. Biol. Chem. 289, 11986–11992 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Chesarino N. M., McMichael T. M., and Yount J. S. (2014) Regulation of the trafficking and antiviral activity of IFITM3 by post-translational modifications. Future Microbiol. 9, 1151–1163 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Vavouri T., Semple J. I., and Lehner B. (2008) Widespread conservation of genetic redundancy during a billion years of eukaryotic evolution. Trends Genet. 24, 485–488 [DOI] [PubMed] [Google Scholar]
- 50. Runkle K. B., Kharbanda A., Stypulkowski E., Cao X. J., Wang W., Garcia B. A., and Witze E. S. (2016) Inhibition of DHHC20-mediated EGFR palmitoylation creates a dependence on EGFR signaling. Mol. Cell 62, 385–396 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Ahearn I. M., Haigis K., Bar-Sagi D., and Philips M. R. (2011) Regulating the regulator: post-translational modification of RAS. Nat. Rev. Mol. Cell Biol. 13, 39–51 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52. Dallavilla T., Abrami L., Sandoz P. A., Savoglidis G., Hatzimanikatis V., and van der Goot F. G. (2016) Model-driven understanding of palmitoylation dynamics: regulated acylation of the endoplasmic reticulum chaperone calnexin. PLoS Comput. Biol. 12, e1004774. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Lynes E. M., Raturi A., Shenkman M., Ortiz Sandoval C., Yap M. C., Wu J., Janowicz A., Myhill N., Benson M. D., Campbell R. E., Berthiaume L. G., Lederkremer G. Z., and Simmen T. (2013) Palmitoylation is the switch that assigns calnexin to quality control or ER Ca2+ signaling. J. Cell Sci. 126, 3893–3903 [DOI] [PubMed] [Google Scholar]
- 54. Moltedo B., Li W., Yount J. S., and Moran T. M. (2011) Unique type I interferon responses determine the functional fate of migratory lung dendritic cells during influenza virus infection. PLoS Pathog. 7, e1002345. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55. Yount J. S., Kraus T. A., Horvath C. M., Moran T. M., and López C. B. (2006) A novel role for viral-defective interfering particles in enhancing dendritic cell maturation. J. Immunol. 177, 4503–4513 [DOI] [PubMed] [Google Scholar]
- 56. Charron G., Tsou L. K., Maguire W., Yount J. S., and Hang H. C. (2011) Alkynyl-farnesol reporters for detection of protein S-prenylation in cells. Mol. Biosyst. 7, 67–73 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Bolte S., and Cordelières F. P. (2006) A guided tour into subcellular colocalization analysis in light microscopy. J. Microsc. 224, 213–232 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.