Significance
Reversible ubiquitination regulates cellular protein abundance. The receptor-like kinase (RLK) RGFR1 plays an important role in regulating root meristem development. Upon ligand binding, activated RGFR1 is ubiquitinated and eventually degraded by the 26S proteasome. In this study, we found that the ubiquitin-specific proteases UBP12 and UBP13 are components of the RGFR1 signaling pathway. UBP13 interacts directly with RGFR1 and can counteract the ligand-induced ubiquitination and degradation of RGFR1. Our discovery provides insight into the dynamic regulation of cell-surface RLKs, which may serve to fine-tune the sensitivity of plant cells to extracellular stimuli.
Keywords: RGF1, RGFR1, PLT1/2, UBP12/13, root meristem
Abstract
ROOT MERISTEM GROWTH FACTOR (RGF) 1 is an important peptide hormone that regulates root growth. Upon binding to its receptor, RGFR1, RGF1 regulates the expression of two transcription factors, PLETHORA 1 and 2 (PLT1/2), to influence root meristem development. Here, we show that the ubiquitin-specific proteases UBP12 and UBP13 are positive regulators of root meristem development and that UBP13 interacts directly with RGF1 receptor (RGFR1) and its close homolog RGFR2. The ubp12,13 double-mutant root is completely insensitive to exogenous applied RGF1. Consistent with this result, RGF1-induced ubiquitination and turnover of RGFR1 protein were accelerated in ubp12,13-mutant plants but were delayed in transgenic plants overexpressing UBP13. Genetic analysis showed that PLT2 or RGFR1 overexpression partially rescued the short-root phenotype and the reduced cortical root meristem cell number in ubp12,13 plants. Together, our results demonstrate that UBP12/13 are regulators of the RGF1–RGFR1–PLT1/2 signaling pathway and that UBP12/13 can counteract RGF1-induced RGFR1 ubiquitination, stabilize RGFR1, and maintain root cell sensitivity to RGF1.
During development, the cells of a multicellular organism are exposed to numerous extracellular signaling molecules produced by themselves or other cells. These signaling molecules are generally perceived by various receptors to initiate signal transduction pathways that alter gene expression and specify cell fate. The types and the concentration of extracellular signaling molecules are changing constantly to meet the demands of different developmental stages, and signal-transduction pathways associated with these signaling molecules must be efficiently switched on and off accordingly. Inactivation of a signaling pathway often involves the removal of the ligand by degradation or sequestration; cells may also be desensitized to extracellular signaling molecules via receptor reduction or inactivation. Of these mechanisms, ubiquitination-mediated receptor degradation has been shown to be one of the most efficient ways to reduce the number of receptor molecules in a cell and decrease the sensitivity of cells to extracellular signals. Ubiquitination is a common type of posttranslational modification in which a 8.6-kDa ubiquitin molecule is attached to the lysine residue of a protein via the sequential activation of a ubiquitin-activating enzyme (E1), a ubiquitin-conjugating enzyme (E2), and a ubiquitin ligase (E3) (1). It has been shown that a number of plant receptors, including FLAGELLIN-SENSING 2 (FLS2), LYSIN MOTIF RECEPTOR KINASE 5 (LYK5), BRASSINOSTEROID INSENSITIVE 1 (BRI1), and CORONATINE INSENSITIVE 1, can be ubiquitinated (2–5). Ubiquitination directs these receptor molecules for degradation by the 26S proteasome or internalization by endocytosis, thereby attenuating cellular responses to extracellular signals.
Similar to other posttranslational modifications (e.g., phosphorylation and methylation), protein ubiquitination is reversible. Ubiquitinated proteins can be deubiquitinated by deubiquitinating enzymes (DUBs), allowing them to resume their biological functions. The Arabidopsis genome encodes ∼64 candidate DUBs, which can be further classified into five subfamilies: ubiquitin-specific proteases (UBPs), ubiquitin C-terminal hydrolases, ovarian tumor proteases, Machado–Joseph domain proteases, and JAB1/MPN/MOV34 domain proteases (6). These DUBs play essential roles in regulating many cellular processes; however, compared with the studies of protein ubiquitination in the literature, the molecular functions of DUBs are mostly unclear. Therefore identifying the direct targets of DUBs, especially those involved in plant developmental signaling, will help decode the mechanisms by which reversible ubiquitination regulates the growth and development of plants.
In this study we found that the simultaneous knockout of two closely related ubiquitin-specific proteases, UBP12 and UBP13, in Arabidopsis resulted in pleiotropic phenotypes, including an extremely short root and growth arrest at an early stage in young seedlings. An examination of ubp12,13 double-mutant root showed that the number of meristematic cells in the cortex was greatly reduced. Yeast two-hybrid assays demonstrated that UBP13 interacts directly with the peptide hormone ROOT MERISTEM GROWTH FACTOR 1 (RGF1)-perceiving receptor RGFR1, also known as RGF1 INSENSITIVE1 (RGI1). By immunoblotting, we found that the degradation of RGFR1 was greatly enhanced in ubp12,13-mutant plants but delayed in UBP13-overexpressing transgenic plants. Furthermore, the ubp12,13-mutant root was insensitive to RGF1, and RGFR1 overexpression partially recovered the short root and reduced cortical root meristem cell-number phenotypes in ubp12,13 plants. Together, our findings demonstrate that UBP12/UBP13 are regulators of the RGF1-signaling pathway in Arabidopsis roots. UBP12/UBP13 may help stabilize RGFR1 in Arabidopsis roots and maintain root cell sensitivity to RGF1.
Results
Simultaneously Knocking Out the Expression of UBP12 and UBP13 Reduced the Number of Cortical Root Meristem Cells.
Previous studies showed that UBP12 and UBP13 play important roles in regulating plant innate immunity, photoperiodic flowering, and jasmonic acid (JA) signaling (7–9). To uncover additional functions of UBP12 and UBP13 in plant development, we isolated transfer DNA (T-DNA) insertion mutants for these two genes (Fig. S1A). The confirmed single mutants were crossed to generate the ubp12,13 double mutant. Although it was reported that homozygous ubp12,13 double-mutant plants could not be produced (7), we successfully obtained ubp12,13 double mutants from the segregated progeny of ubp12+/−ubp13−/− or ubp12−/−ubp13−/+ plants, albeit at a relatively low frequency (∼16%). The isolated ubp12,13 double mutant was confirmed by semiquantitative RT-PCR, which showed a lack of UBP12 and UBP13 transcripts (Fig. S1B).
Similar to previous reports, no obvious growth phenotype was observed for ubp12 or ubp13 single-mutant plants. In comparison, the ubp12,13 double mutant was very small; it stopped growing after 4 wk and eventually died (Fig. S1 C and D). Therefore, the ubp12,13 double mutant could be maintained only as heterozygous ubp12+/−ubp13−/− or ubp12−/−ubp13+/− plants. Similar to the growth defect in the aerial parts of the plant, the root of the ubp12,13 mutant was very short; on average, the root length in 7-d-old ubp12,13 mutant plants was around 27% of that in Columbia-0 (Col-0) control plants (Fig. S2). Confocal microscopic examinations of propidium iodide (PI)-stained roots showed that ubp12,13 had a disordered stem cell niche and reduced number of cortical cells in the meristem, which might help explain the short root phenotype of the double mutant (Fig. S2).
To confirm that the reduced root growth and reduced cortical root meristem cell number in the ubp12,13 mutant were indeed caused by knocking out UBP12 and UBP13 expression, full-length UBP13 cDNA was cloned into the β-estradiol–inducible vector pER8-YFP and transformed into ubp12−/+ubp13−/− mutant plants. Triple-homozygous seedlings from the T3 generation were analyzed. β-Estradiol (30 μM)–induced UBP13 expression in ubp12,13 plants could significantly recover the short root phenotype and increase the number of cortical root meristem cells in the double mutant (Fig. S3). Taken together, these results suggest that UBP12 and UBP13 play critical roles in regulating root meristem development.
UBP12 and UBP13 Regulate PLETHORA 1 and PLETHORA 2 Expression.
Maintenance of the size of the root meristem in Arabidopsis is reportedly regulated via several interdependent genetic pathways, including the CLE40–CRINKLY4/CLAVATA1–WUSCHEL-RELATED HOMEOBOX 5 (WOX5) pathway (10, 11), the Auxin–RGF1–PLETHORA 1/2 (PLT1/2) pathway (12–14), and the SHORTROOT (SHR)–SCARECROW (SCR) pathway (15, 16). To investigate whether any of these known root meristem maintenance-related pathways were associated with the decreased root meristem size observed in the ubp12,13 mutant, we crossed ubp12+/−ubp13−/− mutant plants with a number of transgenic plants harboring various root development markers, including pSCR:SCR-GFP, pSHR:SHR-GFP, pWOX5:GFP, QC184, J0571, pCO2:YFP, pPLT1:YFP, pPLT2:YFP, pPLT1:PLT1-YFP, and pPLT2:PLT2-YFP. Microscopic examinations of F3-segregated triple-homozygous plant roots showed that the expression patterns of pSCR:SCR-GFP, pSHR:SHR-GFP, J0571, and pCO2:YFP were similar in ubp12,13 double-mutant and Col-0 plants (Fig. S4). Corresponding to its disordered stem cell niche phenotype, the expression patterns of pWOX5:GFP and QC184 in the ubp12,13 mutant were expanded from central quiescent cells to the surrounding cells (Fig. S4). Most strikingly, the expression of pPLT1:YFP, pPLT2:YFP, pPLT1:PLT1-YFP, and pPLT2:PLT2-YFP was barely detectable in ubp12,13 plants (Fig. 1A), suggesting that UBP12 and UBP13 play essential roles in regulating PLT1 and PLT2 expression.
Fig. 1.
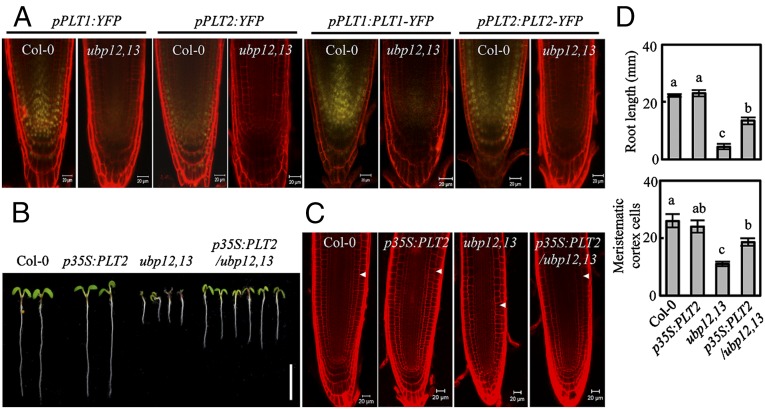
UBP12 and UBP13 regulate PLT1 and PLT2 expression. (A) The expression of pPLT1:YFP, pPLT1:PLT1-YFP, pPLT2:YFP, and pPLT2:PLT2-YFP in wild-type (Col-0) or ubp12,13-mutant roots. (Scale bars: 20 μm.) (B) Seven-day-old wild-type, ubp12,13-mutant, and p35S:PLT2-expressing seedlings. (Scale bar: 1 cm.) (C) Confocal microscopy images of PI-stained roots from plants shown in B. (Scale bars: 20 μm.) White arrowheads mark the cortex transition zones. (D) Quantification of root length and the cortical root meristem cell number for the plants shown in B and C. Error bars represent ±SD, n > 15.
PLT1 and PLT2 expression in Arabidopsis is critical for root meristem development. Simultaneous knockout of PLT1 and PLT2 results in Arabidopsis roots with a reduced cortical meristem size (12, 17). The similar reduced root cortical meristem size phenotype in the ubp12,13 and plt1plt2 mutants and the PLT1 and PLT2 expression defects in ubp12,13 plants prompted us to test whether UBP12/UBP13 regulate PLT1/PLT2-mediated root meristem development. PLT2 overexpression partially rescued the short root and the reduced cortical root meristem cell number phenotypes in the ubp12,13 double mutant (Fig. 1 B–D), suggesting that UBP12/UBP13 act upstream of the PLT1/PLT2 pathway to regulate root meristem size.
UBP13 Interacts with the RGF1 Receptors RGFR1 and RGFR2.
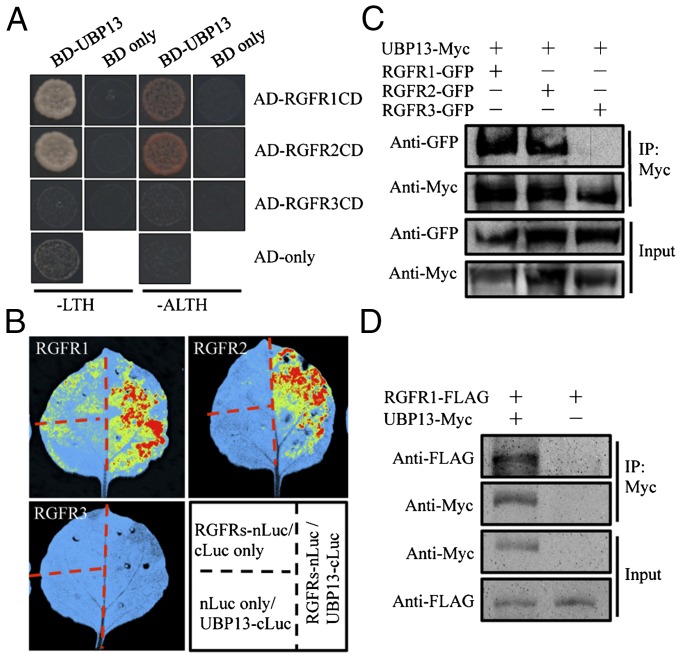
To identify UBP12/UBP13 substrates that could regulate PLT1/PLT2-mediated root meristem development, a yeast two-hybrid screen was performed using UBP13 as bait. The screen yielded 318 positive clones, corresponding to 161 potential UBP13-interacting proteins. Notably, this list included a fragment of cytoplasmic domain of the leucine-rich repeat (LRR)-type receptor-like kinase (RLK) RGFR1. It was recently discovered that RGFR1 and its close homologous proteins (RGFR2, RGFR3, RGFR4, and RGFR5) could serve as the receptors for peptide hormone RGF1 (17–19). RGF1 binding stimulates the phosphorylation and polyubiquitination of RGFR1, which could regulate the expression of PLT1 and PLT2 to influence root growth and root meristem size (17). Interestingly, we found that UBP13 could interact specifically with the cytoplasmic domains of RGFR1 and RGFR2, but not RGFR3, in the yeast two-hybrid assay (Fig. 2A). These results were validated in vivo using a firefly luciferase complementation assay (Fig. 2B). Additionally, RGFR1 and RGFR2, but not RGFR3, could be coimmunoprecipitated with UBP13 from tobacco leaves cotransformed with p35S:RGFRs-GFP and p35S:UBP13-Myc or double-transgenic Arabidopsis plants expressing p35S:RGFR1-FLAG and p35S:UBP13-Myc (Fig. 2 C and D).
Fig. 2.
UBP13 interacts with RGFR1 and RGFR2 but not with RGFR3. (A) Yeast two-hybrid assay shows that UBP13 interacts directly with the cytoplasmic domain (RGFRsCD) of RGFR1 and RGFR2 but not of RGFR3. (B–D) Firefly luciferase complementation assay (B) and coimmunoprecipitation (C and D) using transiently transformed Nicotiana benthamiana leaves (B and C) or transgenic Arabidopsis plants (D) show that UBP13 interacts with RGFR1 and RGFR2, but not RGFR3.
RGFR1 Stability Is Regulated by UBP13.
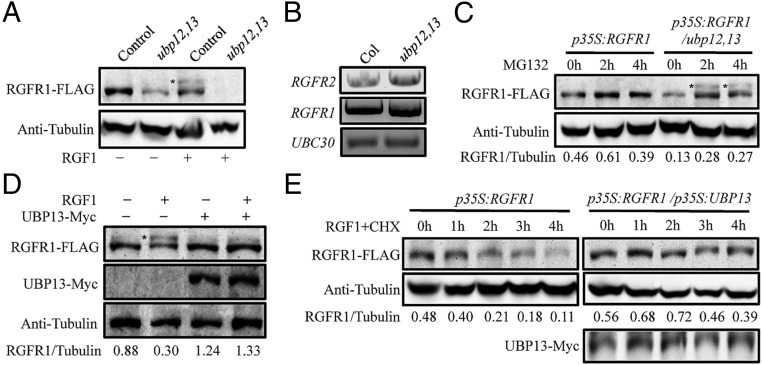
Previous studies showed that UBP12 and UBP13 have protein deubiquitination activity in vitro (8). To investigate whether UBP13 can mediate the deubiquitination of RGFR1, we crossed p35S:RGFR1-FLAG transgenic plants with ubp12(−/+)13(−/−) mutants and harvested F3-segregated triple-homozygous mutants for analysis. Immunoblots showed that the protein abundance of RGFR1-FLAG was greatly reduced in ubp12,13 double mutants (Fig. 3A). Similar to the result reported previously (17), 30 min of treatment with 20 μM RGF1 promoted the ubiquitination of RGFR1-FLAG and at the same time decreased the level of RGFR1-FLAG protein in the wild-type background. In comparison, the RGF1 treatment reduced RGFR1-FLAG protein to an undetectable level in the ubp12,13 mutant (Fig. 3A). To eliminate the possibility that the reduced RGFR1-FLAG protein level in the ubp12,13 mutant was caused by transcriptional regulation, the transcript levels of RGFR1 and RGFR2 in the ubp12,13 mutant were examined. Fig. 3B and Fig. S5 show that the transcripts of RGFR1 and RGFR2 in the ubp12,13 mutant were slightly higher than those in the Col-0 control, suggesting that the reduced RGFR1-FLAG protein level in the ubp12,13 mutant was due to translational or posttranslational regulation. We next treated seedlings with a 26S proteasome inhibitor, MG132, and found that the treatment increased RGFR1-FLAG abundance in both Col-0 and ubp12,13-mutant plants (Fig. 3C). In comparison, a significant amount of MG132-stabilized RGFR1-FLAG was ubiquitinated in the ubp12,13 mutant, but not in Col-0 plants, supporting the hypothesis that UBP12 and UBP13 regulate the deubiquitination of RGFR1.
Fig. 3.
UBP12 and UBP13 modulate RGFR1 stability. (A) The RGFR1 protein level decreased in ubp12,13 double-mutant plants. p35S:RGFR1-FLAG–expressing seedlings were treated with or without 20 μM RGF1 for 30 min. (B) RT-PCR analyses of the RGFR1 and RGFR2 transcript levels in the ubp12,13 mutant. (C) MG132 treatment stabilized RGFR1. Seedlings were treated with 50 μM MG132 for the indicated time. (D and E) UBP13 overexpression inhibited the RGF1-induced ubiquitination and degradation of RGFR1. Seedlings were treated with 20 μM RGF1 for 30 min (D) or with 20 μM RGF1 and 200 μM CHX for the indicated time (E). Asterisks in A, C, and D denote ubiquitinated RGFR1-FLAG. RGFR1/tubulin represents the signal ratio of RGFR1-FLAG to tubulin.
We also investigated whether increasing the expression of UBP13 would have any direct effect on RGFR1-FLAG stability. p35S:RGFR1-FLAG/p35S:UBP13-Myc double-homozygous transgenic plants, which were generated by genetic crosses, were treated with or without 20 μM RGF1 for 30 min and were subjected to total protein extraction for immunoblotting. UBP13 overexpression slightly increased the abundance of RGFR1-FLAG protein in the absence of RGF1 and decreased the RGF1-induced ubiquitination of RGFR1-FLAG (Fig. 3D). Furthermore, 1-wk-old p35S:RGFR1-FLAG or p35S:RGFR1-FLAG/p35S:UBP13-Myc transgenic seedlings were treated with 20 μM RGF1 and 200 μM cycloheximide (CHX) for various time intervals, and changes in RGFR1-FLAG abundance over time were examined. As shown in Fig. 3E, RGF1 treatment reduced the RGFR1-FLAG level when new protein synthesis was blocked by CHX. At 4 h after RGF1 treatment the amount of RGFR1-FLAG was reduced to a barely detectable level. In UBP13-overexpressing transgenic plants, RGF1 and CHX cotreatment could still trigger the turnover of RGFR1-FLAG, but the speed was much slower (Fig. 3E). Taken together, these results suggest that UBP13 regulates the deubiquitination of RGFR1 and that it helps maintain RGFR1-FLAG at a relatively stable level in vivo.
UBP12 and UBP13 Regulate Root Meristem Development Through the RGF1–RGFR1 Signaling Pathway.
To investigate whether UBP12/UBP13 are involved in the RGF1 signaling pathway, we examined the response of ubp12,13-mutant plants to exogenous RGF1. Consistent with previous reports, RGF1 treatment induced a significant wavy root phenotype and increased the PLT2 protein level as well as the number of cortical root meristem cells in wild-type seedlings (Fig. 4 A, C, and E and Fig. S6). In contrast, root growth, PLT2 protein level, and the cortical root meristem cell number in ubp12,13 mutant were nearly the same for seedlings grown in the presence or absence of RGF1, suggesting that the ubp12,13 mutant was insensitive to RGF1 (Fig. 4 B, D, and E and Fig. S6). Given that the protein abundance of RGFR1 is regulated by UBP12/UBP13, we wondered whether the insensitivity of ubp12,13 plants to RGF1 was due to the insufficient amount of RGFR1 in the mutant. To test this hypothesis, we overexpressed RGFR1-FLAG in ubp12,13 plants and found that increasing the abundance of RGFR1 partially rescued ubp12,13 mutant’s short root phenotype and significantly increased the number of cortical root meristem cells (Fig. 4 F–H). These results demonstrate that UBP12 and UBP13 play critical roles in maintaining RGFR1 abundance and that they regulate root meristem development through the RGF1–RGFR1 signaling pathway.
Fig. 4.
UBP12 and UBP13 function in the RGF1-signaling pathway to regulate root meristem formation. (A–D) Wild-type (A and C) and ubp12,13-mutant (B and D) plants were grown in the presence or absence of 100 nM RGF1 for 7 d (A and B); the roots were then stained with PI and observed by confocal microscopy (C and D). (Scale bars: 20 μm in C and D.) (E) Quantification of the cortical root meristem cell numbers for the plants shown in C and D. Error bars represent SD (n > 20). (F) Seven-day-old wild-type, p35S:RGFR1-FLAG, ubp12,13, and p35S:RGFR1-FLAG–expressing ubp12,13 seedlings. (G) Confocal microscopic images of PI-stained roots of plants shown in F. (H) Quantitation of root length and number of cortical root meristem cells in the plants shown in F and G. Error bars represent ± SD (n > 20). White arrowheads in C, D, and G mark the junction between the meristematic and elongation zones.
Discussion
Given the importance of roots for water and nutrient absorption and transport, root growth is tightly regulated by various phytohormones as well as by a number of secreted peptide hormones (20, 21). These substances coordinately control root growth by regulating cell division in the meristematic region and cell elongation in the transition and elongation zones. Among these root growth regulators, RGFs are polypeptides containing 13 amino acids and a sulfated tyrosine in their mature form and are mostly expressed in the root meristem (14). The binding of RGF1 to RGFR1–5 initiates a signal-transduction pathway that regulates the expression of the downstream transcription factors PLT1/PLT2 and controls root meristem development (14, 17–19). RGF1 binding may also promote the polyubiquitination and degradation of RGFR1 (17).
In this study, we showed that simultaneously knocking out the expression of UBP12 and UBP13 in Arabidopsis caused a short-root phenotype characterized by a significantly reduced number of cortical cells in the root meristem and that this phenotype could be recovered by the inducible expression of UBP13 in ubp12,13 double-mutant plants. Several lines of evidence indicate that UBP12/UBP13 regulate root meristem development via RGF1–RGFR1–PLT1/PLT2 signaling. First, PLT1/PLT2 expression was decreased to an undetectable level in ubp12,13 plants, and PLT2 overexpression partially rescued the short-root and reduced cortical root meristem cell number phenotypes in ubp12,13-mutant plants. Second, yeast two-hybrid, firefly luciferase complementation, and coimmunoprecipitation assays all demonstrated that UBP13 could interact directly with RGFR1 and RGFR2 and that the in vivo expression levels of UBP12/UBP13 negatively regulate RGF1-induced RGFR1 ubiquitination and degradation. Third, the ubp12,13 mutant was insensitive to RGF1, and RGFR1 overexpression could partially rescue the meristem developmental defect of the mutant. Fourth, histochemical staining of pUBP13:β-glucuronidase (GUS) transgenic seedlings showed that UBP13 is expressed in the root cortical meristem region (Fig. S7). Together, our results not only reveal a key role for UBP12/UBP13 in regulating root meristem development but also demonstrate a mechanism whereby UBP12/UBP13 regulate RGFR1 abundance. The balance between RGF1-induced RGFR1 ubiquitination and degradation and the deubiquitination of RGFR1 by UBP12/UBP13 may regulate RGFR1 protein dynamics, affect the sensitivity of root cells to RGF1, and modify root growth and development.
In Arabidopsis, more than 600 RLKs have been identified (22), and several have been reported to be ubiquitinated. For example, FLS2 and LYK5 are receptors for bacterial flagellin and fungal chitooligosaccharides, respectively, and both initiate immune signaling in Arabidopsis. Flagellin binding promotes the interaction of FLS2 with the E3 ubiquitin ligase PLANT U-BOX 12 (PUB12) as well as its close homolog PUB13. PUB12/PUB13 ubiquitinate FLS2, targeting it for degradation by the 26S proteasome (2). Interestingly, PUB13 is also the E3 ligase for LYK5. Unlike FLS2, PUB13 interacts with LYK5 in the absence of chitooligosaccharides, resulting in the turnover of LYK5. Chitooligosaccharides stimulate the dissociation of PUB13 from LYK5, promote LY5 accumulation, and increase cellular sensitivity to the ligand (3). BRI1 is the receptor for phytohormone brassinosteroids (BRs). It was found previously that K63 of BRI1 is polyubiquitinated in vivo. This ubiquitination is not regulated by BR binding, but it promotes BRI1 internalization (4). Together, these findings demonstrate that the ubiquitination of RLKs reduces the number of receptors on the cell surface and attenuates cellular sensitivity to extracellular signaling molecules. Multiple studies in animal cells have shown that ubiquitinated receptor kinases on the plasma membrane can be deubiquitinated by DUBs, reversing the turnover of those receptors and resetting cellular sensitivity to extracellular signals (23). Our study of the deubiquitination of RGFR1 by UBP13 demonstrates that DUBs-mediated receptor kinase stabilization also exists in plants; thus, our work provides additional information on the dynamic regulation of cell-surface RLKs in planta. Future identification of the DUBs that are responsible for deubiquitinating various plant RLKs will enhance our understanding of the ways in which the on-and-off signals mediated by these RLKs are fine-tuned.
Of the 27,416 protein-coding genes in Arabidopsis [The Arabidopsis Information Resource (TAIR )10], around 6% encode components of the protein ubiquitination and degradation system, including two E1s, at least 37 E2s, and more than 1,400 E3s (24, 25). In comparison, only 64 potential deubiquitination proteases were identified in a sequence analysis (6). As the substrate specificity of this system is mostly determined by E3, the difference in the numbers of DUBs and E3s suggests that DUBs are multifunctional; each DUB could regulate the deubiquitination of different target proteins during different cellular processes. Indeed, studies have found that reducing the expression of UBP12/UBP13 by RNAi or T-DNA insertion resulted in increased resistance to Pseudomonas syringae pv. tomato DC3000, early flowering, and a shortened circadian period; however, the direct targets of UBP12/UBP13 were not identified in these studies (7, 8). Additionally, it was found that UBP12/UBP13 could interact with the plant-specific polycomb group (PcG) protein LIKE HETEROCHROMATIN PROTEIN 1 (LHP1) and regulate autonomous endosperm development (26). Jeong et al. (9) discovered that UBP12/UBP13 can deubiquitinate MYC2, the central transcription factor in the JA signaling pathway, preventing it from being degraded and maintaining cellular sensitivity to JA. In the present study we found that RGFR1 is a substrate of UBP12/UBP13. We also described the pleiotropic growth phenotypes of the ubp12,13-null mutant, including extremely slow aerial organ growth and death at 4 wk, which cannot be explained by the currently known targets of UBP12/UBP13. Future identification of additional UBP12/UBP13 substrates will help decoding the mechanisms of UBP12/UBP13-mediated protein deubiquitination that regulate plant growth and development.
Materials and Methods
Plant Growth Conditions.
Seeds were surface-sterilized in 75% ethanol for 3 min followed by three quick rinses with 100% ethanol. The seeds were completely dried in a sterile hood before planting on medium containing 1/2 Murashige and Skoog (MS) basal salts, 1% sucrose, and 0.5% Gelzan (Caisson Laboratories, Inc.). Seeds were vernalized at 4 °C in the dark for 2 d and then were grown vertically at 22 °C under long-day conditions (∼100 μmol⋅m−2⋅s−1, 16 h light/8 h dark) for 7 d before using for microscopic observation, immunoblotting, or the gene-expression assay, unless indicated otherwise. For the root growth assays, seedlings were grown on medium with or without 100 nM synthesized sulfonated RGF1 for 7 d before images were acquired. Root length was measured using ImageJ software. For statistic analysis, one-way ANOVA analysis was performed; statistically significant differences are indicated by different lowercase letters (P < 0.05).
Microscopic Imaging.
One-week-old Arabidopsis roots were mounted in 10 μg/mL of PI (Sigma-Aldrich) and were observed directly under a confocal laser-scanning microscope (LSM 710; Zeiss).
Protein Interaction Assays.
cDNAs encoding full-length protein sequences of UBP13 or cytoplasmic domains of RGFR1, RGFR2, or RGFR3 were cloned into the pGBKT7 or pGADT7 vectors. Yeast two-hybrid assays were performed according to a standard protocol. For firefly luciferase complementation assays, cDNAs encoding full-length protein sequences of UBP13, RGFR1, RGFR2, and RGFR3 were cloned into pCAMBIA-cLuc or pCAMBIA-nLuc, respectively. Firefly luciferase complementation assays were performed as described previously (27). To analyze the interaction among UBP13, RGFR1, and RGFR2 in vivo, 1-wk-old Arabidopsis seedlings expressing p35S:RGFR1-FLAG or p35S:RGFR1-FLAG/p35S:UBP13-Myc were used. Alternatively, tobacco leaves were coinfiltrated with Agrobacterium tumefaciens strain GV3101 harboring p35S:RGFR1-GFP, p35S:RGFR2-GFP, p35S:RGFR3-GFP, or p35S:UBP13-Myc expression vectors. Thirty-six hours after the infiltration, the leaves were harvested for immunoprecipitation. Protein coimmunoprecipitation assays were performed as reported previously (28).
Immunoblotting.
Seven-day-old seedlings were submerged in double-distilled H2O containing 50 μM MG132 or 20 μM sulfonated RGFl with or without 200 μM CHX for the indicated time. The proteins were separated by SDS/PAGE and blotted onto nitrocellulose membranes. RGFRl-FLAG or UBP13-Myc was detected using monoclonal anti-FLAG or anti-Myc antibodies, respectively (Sigma-Aldrich).
Supplementary Material
Acknowledgments
We thank Dr. Chengwei Yang for providing the pPLT1:PLT1-YFP and pPLT2:PLT2-YFP marker lines; Dr. Lizhen Tao for providing the pPLT1:YFP, pPLT2:YFP, QC184, and pSCR:SCR-GFP marker lines; Dr. Chunming Liu for providing the pCO2:YFP, J0571, pWOX5:GFP, and pSHR:SHR-GFP marker lines; and Dr. Xiaofeng Cao for providing pUBP13:GUS transgenic plants. This study was supported by National Natural Science Foundation of China Grants 31770308 (to Y.S.) and 91417313 (to W.T.) and Human Resource Department of Hebei Province, China Grants E2011100004 (to W.T.) and E2013100004 (to Y.S.).
Footnotes
The authors declare no conflict of interest.
This article is a PNAS Direct Submission.
This article contains supporting information online at www.pnas.org/lookup/suppl/doi:10.1073/pnas.1714177115/-/DCSupplemental.
References
- 1.Hershko A, Ciechanover A. The ubiquitin system for protein degradation. Annu Rev Biochem. 1992;61:761–807. doi: 10.1146/annurev.bi.61.070192.003553. [DOI] [PubMed] [Google Scholar]
- 2.Lu D, et al. Direct ubiquitination of pattern recognition receptor FLS2 attenuates plant innate immunity. Science. 2011;332:1439–1442. doi: 10.1126/science.1204903. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Liao D, et al. Arabidopsis E3 ubiquitin ligase PLANT U-BOX13 (PUB13) regulates chitin receptor LYSIN MOTIF RECEPTOR KINASE5 (LYK5) protein abundance. New Phytol. 2017;214:1646–1656. doi: 10.1111/nph.14472. [DOI] [PubMed] [Google Scholar]
- 4.Martins S, et al. Internalization and vacuolar targeting of the brassinosteroid hormone receptor BRI1 are regulated by ubiquitination. Nat Commun. 2015;6:6151. doi: 10.1038/ncomms7151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Yan J, et al. The Arabidopsis F-box protein CORONATINE INSENSITIVE1 is stabilized by SCFCOI1 and degraded via the 26S proteasome pathway. Plant Cell. 2013;25:486–498. doi: 10.1105/tpc.112.105486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Isono E, Nagel M-K. Deubiquitylating enzymes and their emerging role in plant biology. Front Plant Sci. 2014;5:56. doi: 10.3389/fpls.2014.00056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Ewan R, et al. Deubiquitinating enzymes AtUBP12 and AtUBP13 and their tobacco homologue NtUBP12 are negative regulators of plant immunity. New Phytol. 2011;191:92–106. doi: 10.1111/j.1469-8137.2011.03672.x. [DOI] [PubMed] [Google Scholar]
- 8.Cui X, et al. Ubiquitin-specific proteases UBP12 and UBP13 act in circadian clock and photoperiodic flowering regulation in Arabidopsis. Plant Physiol. 2013;162:897–906. doi: 10.1104/pp.112.213009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Jeong JS, Jung C, Seo JS, Kim JK, Chua NH. The deubiquitinating enzymes UBP12 and UBP13 positively regulate MYC2 levels in jasmonate responses. Plant Cell. 2017;29:1406–1424. doi: 10.1105/tpc.17.00216. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Stahl Y, Wink RH, Ingram GC, Simon R. A signaling module controlling the stem cell niche in Arabidopsis root meristems. Curr Biol. 2009;19:909–914. doi: 10.1016/j.cub.2009.03.060. [DOI] [PubMed] [Google Scholar]
- 11.Stahl Y, et al. Moderation of Arabidopsis root stemness by CLAVATA1 and ARABIDOPSIS CRINKLY4 receptor kinase complexes. Curr Biol. 2013;23:362–371. doi: 10.1016/j.cub.2013.01.045. [DOI] [PubMed] [Google Scholar]
- 12.Aida M, et al. The PLETHORA genes mediate patterning of the Arabidopsis root stem cell niche. Cell. 2004;119:109–120. doi: 10.1016/j.cell.2004.09.018. [DOI] [PubMed] [Google Scholar]
- 13.Mähönen AP, et al. PLETHORA gradient formation mechanism separates auxin responses. Nature. 2014;515:125–129. doi: 10.1038/nature13663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Matsuzaki Y, Ogawa-Ohnishi M, Mori A, Matsubayashi Y. Secreted peptide signals required for maintenance of root stem cell niche in Arabidopsis. Science. 2010;329:1065–1067. doi: 10.1126/science.1191132. [DOI] [PubMed] [Google Scholar]
- 15.Sabatini S, Heidstra R, Wildwater M, Scheres B. SCARECROW is involved in positioning the stem cell niche in the Arabidopsis root meristem. Genes Dev. 2003;17:354–358. doi: 10.1101/gad.252503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Helariutta Y, et al. The SHORT-ROOT gene controls radial patterning of the Arabidopsis root through radial signaling. Cell. 2000;101:555–567. doi: 10.1016/s0092-8674(00)80865-x. [DOI] [PubMed] [Google Scholar]
- 17.Ou Y, et al. RGF1 INSENSITIVE 1 to 5, a group of LRR receptor-like kinases, are essential for the perception of root meristem growth factor 1 in Arabidopsis thaliana. Cell Res. 2016;26:686–698. doi: 10.1038/cr.2016.63. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Shinohara H, Mori A, Yasue N, Sumida K, Matsubayashi Y. Identification of three LRR-RKs involved in perception of root meristem growth factor in Arabidopsis. Proc Natl Acad Sci USA. 2016;113:3897–3902. doi: 10.1073/pnas.1522639113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Song W, et al. Signature motif-guided identification of receptors for peptide hormones essential for root meristem growth. Cell Res. 2016;26:674–685. doi: 10.1038/cr.2016.62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Lee Y, Lee W-S, Kim S-H. Hormonal regulation of stem cell maintenance in roots. J Exp Bot. 2013;64:1153–1165. doi: 10.1093/jxb/ers331. [DOI] [PubMed] [Google Scholar]
- 21.Delay C, Imin N, Djordjevic MA. Regulation of Arabidopsis root development by small signaling peptides. Front Plant Sci. 2013;4:352. doi: 10.3389/fpls.2013.00352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Shiu SH, Bleecker AB. Receptor-like kinases from Arabidopsis form a monophyletic gene family related to animal receptor kinases. Proc Natl Acad Sci USA. 2001;98:10763–10768. doi: 10.1073/pnas.181141598. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.McCann AP, Scott CJ, Van Schaeybroeck S, Burrows JF. Deubiquitylating enzymes in receptor endocytosis and trafficking. Biochem J. 2016;473:4507–4525. doi: 10.1042/BCJ20160826. [DOI] [PubMed] [Google Scholar]
- 24.Vierstra RD. The ubiquitin-26S proteasome system at the nexus of plant biology. Nat Rev Mol Cell Biol. 2009;10:385–397. doi: 10.1038/nrm2688. [DOI] [PubMed] [Google Scholar]
- 25.Wang F, Deng XW. Plant ubiquitin-proteasome pathway and its role in gibberellin signaling. Cell Res. 2011;21:1286–1294. doi: 10.1038/cr.2011.118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Derkacheva M, et al. H2A deubiquitinases UBP12/13 are part of the Arabidopsis polycomb group protein system. Nat Plants. 2016;2:16126. doi: 10.1038/nplants.2016.126. [DOI] [PubMed] [Google Scholar]
- 27.Chen H, et al. Firefly luciferase complementation imaging assay for protein-protein interactions in plants. Plant Physiol. 2008;146:368–376. doi: 10.1104/pp.107.111740. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Wang R, et al. The brassinosteroid-activated BRI1 receptor kinase is switched off by dephosphorylation mediated by cytoplasm-localized PP2A B’ subunits. Mol Plant. 2016;9:148–157. doi: 10.1016/j.molp.2015.10.007. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.