Abstract
Elevated tryptophan (Trp) catabolism in the tumor microenvironment (TME) can mediate immune suppression by upregulation of IFNg-inducible indoleamine 2,3-dioxygenase (IDO1) and/or ectopic expression of the predominantly liver-restricted enzyme tryptophan 2,3-dioxygenase (TDO)1–5. Whether these effects are due to Trp depletion in the TME or are mediated by the accumulation of the IDO1/TDO product Kynurenine (Kyn) remains controversial5–13. Here we show that administration of a pharmacologically optimized enzyme (PEGylated Kynureninase) that degrades Kyn into immunologically inert, non-toxic and readily-cleared metabolites inhibits tumor growth. Enzyme treatment is associated with a marked increase in the tumor infiltration and proliferation of polyfunctional CD8+ lymphocytes. We show that PEG-Kynureninase administration has substantial therapeutic effects when combined with approved checkpoint inhibitors or with a cancer vaccine for the treatment of large B16-F10 melanoma, 4T1 breast carcinoma or CT26 colon carcinoma tumors. PEG-Kynureninase mediates prolonged depletion of Kyn in the TME and reverses the modulatory effects of IDO1/TDO upregulation in the tumor microenvironment.
Introduction
Upregulation of Trp catabolism by IDO1 and/or TDO represents one of the most studied and clinically validated pathways for immune suppression in tumors. These enzymes catalyze the oxidation of Trp to N-formyl L-kynurenine, which is rapidly converted by formamidases to Kyn. Elevated concentrations of Kyn and higher plasma Kyn/Trp ratios are frequently observed in advanced stage cancer patients and correlate with poor prognoses8. In addition to IDO1/TDO, Kyn can also be generated by the IDO2 isozyme; however, the role of IDO2 in cancer remains uncertain9. IDO1/TDO expression in the TME has been linked to the induction of numerous tolerogenic immune phenotypes including the suppression of effector T-cell activation, enhanced infiltration of myeloid-derived suppressor cells (MDSCs), B cell dysfunction and promotion of tumor angiogenesis3. IDO1 is upregulated in numerous malignancies in response to local inflammatory responses that arise from CD8+ T-cell infiltration and from crosstalk with other immunosuppressive pathways10,11. IDO1 expression has been shown to mediate resistance to cancer immunotherapy with αCTLA4 and αPD-1 antibodies and to inhibit CAR T-cell function in murine models12–14. Furthermore, Kyn synthesis has also been reported to promote tumorigenesis in a manner independent of immunological effects, by activating β-catenin and Akt/PI3K signaling15. Small molecule inhibitors of IDO1 are currently being evaluated in several clinical trials
Mechanistically, early studies suggested that the immunosuppressive effect of Trp catabolism on T-cells is a consequence of the reduction in the local concentration of Trp and the ensuing induction of general starvation responses through activation of the general control nonderepressible 2 kinase (GCN2) and/or suppression of the mTOR pathways3 (Fig. 1a). Elevated Trp catabolism also results in increased concentration of Kyn locally within the TME and systemically, when tumor burden is high. Kyn has been shown to be an AhR ligand16,17, whose activation and nuclear translocation induce a number of immunosuppressive phenotypes in T-cells, including reprogramming of Th17 cells into Tregs16. Furthermore, a recent report revealed that in addition to its direct effect on AhR, in vitro Kyn is slowly converted non-enzymatically to byproducts that serve as high-affinity (sub-nM) AhR agonists18. Downstream metabolites of the Trp catabolism/Kyn pathway, notably 3′OH-kynurenine (3′OH-Kyn) and 3′OH-anthranilic acid (3′OH-AA) are also AhR ligands; however, their concentration, at least in the serum of healthy humans, is 10–50 fold lower than that of Kyn19. The relative significance of Trp starvation responses versus Kyn accumulation in the TME as the driver for immunosuppression has been debated for almost 20 years. While the view that localized Trp starvation is primarily responsible for the impaired T-cell function in tumors remains prevalent, quantitative arguments and recent experimental evidence have begun casting doubts on this hypothesis4–7.
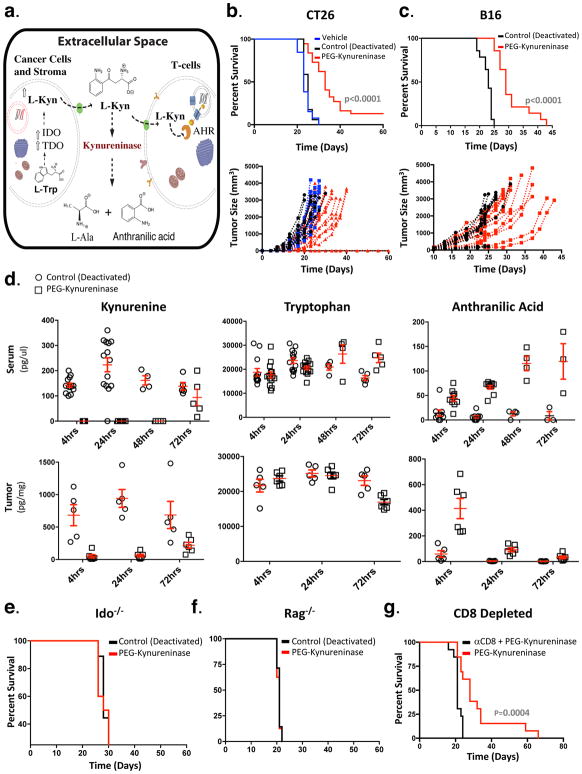
Figure 1. Kynureninase administration reduces tumor growth by depleting kynurenine from the tumor microenvironment.
(a) Schematic of the proposed therapeutic mechanism of Kynureninase administration. (b) Mice were treated with vehicle control (PBS, n = 13) heat-deactivated control enzyme (20 mg/kg, n = 17) or PEG-Kynureninase (20 mg/kg, n = 37) four days after CT26 tumor implantation and administered every 72 hours thereafter for a total of 6 doses. Graph shows survival data compiled from three independent experiments. (c) Ten days after tumor challenge, mice with large, established B16-F10 tumors (104 mm3 mean +/− 9.5 mm3) were treated with PEG-Kynureninase (20 mg/kg) or heat-deactivated control enzyme (20 mg/kg) every 72 hours for a total of 6 doses. Graph shows survival data from three independent experiments (n = 14 mice per group). (d) Mice bearing large B16-F10 tumors (198 mm3 mean +/− 6.8 mm3) were treated with a single dose of PEG-Kynureninase (20 mg/kg) or heat-deactivated control enzyme. Serum and tumor samples were collected for LC-MS/MS metabolomics analysis at times indicated from three independent experiments, with each symbol representing data from individual mice and the red bars representing the mean +/− SEM (tumor analysis of all metabolites: n = 5 for deactivated enzyme control and n = 6 for PEG-Kynureninase treated groups at each time point; serum Kynurenine and Tryptophan: n = 13 per group at 4 hours, n = 14 mice per group at 24 hours, n = 4 mice per group at 48 hours and n = 5 mice per group at 72 hours; serum Anthranilic Acid: n = 9 mice per group at 4 hours and 24 hours, n = 4 mice per group at 48 hours and n = 3 mice per group at 72 hours). (e) Ido1−/− (n = 10 per group) or (f) Rag−/− (n = 8 per group) mice bearing established B16-F10 tumors (>89 mm3) were treated with PEG-Kynureninase (20 mg/kg) or heat-deactivated control enzyme (20 mg/kg) on day 10 and then every 72 hours thereafter for a total of 6 doses. Graphs show representative survival data from one of two independent experiments which had similar results. (g) Mice with established B16-F10 tumors were administered either CD8 depleting or isotype control antibodies and treated with PEG-Kynureninase (20 mg/kg) every 72 hours for a total of six doses staring on day 10. Graph depicts cumulative survival data compiled from two independent experiments with similar results (n = 13 mice per group). Statistical significance of treated mice compared to (b and c) control groups or (g) CD8 depleted mice was determined by Log-rank (Mantel-Cox) test.
Eliminating the extracellular pool of Kyn in a manner that does not affect the consumption of Trp by IDO1/TDO could help to unequivocally assess whether immunosuppression is driven primarily by Kyn accumulation in the TME or by Trp starvation-induced mechanisms. We hypothesized that such selective removal of the extracellular Kyn pool may be accomplished through the systemic administration of an appropriate, pharmacologically optimized enzyme capable of degrading Kyn into non-toxic and immunologically inert metabolites (Fig. 1a). From a therapeutic standpoint, enzyme treatment could offer several important advantages relative to the small molecule IDO1 inhibitors currently in clinical evaluation, including the ability to treat not only cancers in which IDO1 is upregulated, but also tumors expressing TDO1, or both, along with other pharmacologically favorable features, as discussed below.
Of the three enzymes that use Kynurenine as a substrate, namely Kynurenine Monooxygenase, Kynurenine Aminotransferase and Kynureninase, only the latter has biochemical and pharmacological properties (including cofactor requirements, ability to function in the relatively high redox environment of serum and interstitial fluid, biophysical stability and enzyme products that are unlikely to lead to adverse neurophysiological or immunological effects) to be a candidate for in vivo administration (Supplementary Fig. 1)20,21. The Homo sapiens Kynureninase catalyzes the degradation of Kyn with slow kinetics, having instead a strong preference for 3′OH-Kyn, the product of Kynurenine Monooxygenase (kcat/KM of 1.0×102 M−1s−1 and 3.7 ×104 M−1s−1 for Kyn and 3′OH-Kyn, respectively, Supplementary Table 1). Other eukaryotic Kynureninases also preferentially degrade 3′OH-Kyn with similar kinetics (Supplementary Table 1)22. In initial experiments we expressed H. sapiens Kynureninase in E.coli, purified it to near homogeneity (>95%) with low endotoxin content and then conjugated it to 5 kDa PEG-NHS ester for prolonged systemic retention23. PEGylation did not impair the catalytic activity of the enzyme. However, peritumoral administration of the PEGylated H. sapiens Kynureninase to mice bearing established B16-F10 tumors had no effect on tumor growth or mean survival (data not shown). We reasoned that the lack of efficacy of the H. sapiens Kynureninase was a consequence of its poor kinetics with Kyn as a substrate. To address this problem, we expressed and characterized several prokaryotic Kynureninases which bioinformatically were predicted to have preferential activity for Kyn over 3′OH-Kyn. (Supplementary Table 1). The P. fluorescens and M. paludis enzymes which have kcat/KM values 760× and 570× greater, respectively, relative to the human enzyme were expressed, purified to near homogeneity and PEGylated as above (Supplementary Fig. 2). In contrast to the H. sapiens Kynureninase, administration of either of the highly catalytically active P. fluorescens or M. paludis enzymes resulted in strong anti-tumor effects in multiple murine tumor models, as discussed below.
The addition of bacterial PEG-Kynureninases completely reversed Kyn-induced cell death of activated mouse T-cells in vitro (Supplementary Fig. 3). Previous studies have shown similar effects of Kyn on activated human T-cells24. For in vivo experiments mice were treated with either active P. fluorescens PEG-Kynureninase or heat-deactivated enzyme as a control. Heat deactivated, PEGylated protein, rather than mutated inactive enzyme was used as a control to properly account for the effects of protein batch-to-batch variation and for the presence of trace impurities. Dose-finding studies established that maximal efficacy was observed when the enzyme was administered peritumorally at 20 mg/kg weight (Supplementary Fig. 4). Pharmacokinetic analysis revealed that PEG-Kynureninase was maintained at >10 μg/ml for at least 70 hours after administration (Supplementary Fig. 5). As shown in Figure 1b, administration of active enzyme in mice bearing palpable syngeneic CT26 colon carcinoma tumors (Day 4), which express IDO1, led to a nearly 45% improvement in mean survival (23 versus 33 days) relative to animals receiving vehicle alone or deactivated enzyme. Of note, ~15% of the animals had complete, durable responses accompanied by long-lasting and tumor-specific immunity as demonstrated in re-challenge experiments (Supplementary Fig. 6a). We further observed that the effect of PEG-Kynureninase on animals bearing CT26 tumors was statistically indistinguishable from a group that received αPD-1 (Supplementary Fig. 7).
Next, we evaluated the effect of PEG-Kynureninase administration on large tumors (>100 mm3, Day 10) in the extensively studied B16-F10 murine melanoma model. B16-F10 cells do not express IDO1, however tumor growth potently induces the upregulation of IDO1 in the TME5,10. As a monotherapy, administration of PEG-Kynureninase resulted in statistically significant hindrance of tumor growth and increased mean survival over the control group (Fig. 1c, p<0.0001). Time course LC-MS/MS metabolomics analysis showed that within 8 hours following enzyme administration, the serum concentration of Kyn was reduced by over 100-fold, to below the limit of detection (<1 pg/μl, Fig. 1d and Supplementary Table 2) and remained undetectable for at least 48 hours. The Kyn concentration in tumor samples was markedly elevated (~4x) relative to serum, as expected given the upregulation of IDO1 in the TME in the B16-F10 model (Fig. 1d). Following PEG-Kynureninase administration, the tumor levels of Kyn were substantially reduced, to <5% of the values observed in animals treated with deactivated enzyme. Consistent with the hypothesis presented above, Trp levels in B16-F10 tumors were comparable to those in the serum19 and were unaffected by treatment with enzyme (Fig. 1d). The concentration of anthranilic acid, the product of enzymatic degradation of Kyn increased in the serum and transiently in tumor samples. Treatment with PEG-Kynureninase did not affect the tumor level of downstream metabolites of the Kynurenine pathway that are known AhR ligands, namely 3′OH-Kyn, 3′OH-AA and Quinolinic acid (Supplementary Fig. 8). The concentration of these metabolites remained within or below the physiological range for humans19.
The effect of PEG-Kynureninase on the growth of B16-F10 tumors was completely abolished in Ido1−/− mice (Fig. 1e), which is consistent with previous studies using IDO inhibitors25,26. To test the role of the host immune system versus tumor-cell intrinsic effects 27 in the therapeutic function of PEG-Kynureninase, we used NOD-Scid IL2rγc−/− (NSG) and Rag−/− mice. The former strain lacks functional lymphocytes and is defective in innate immune function, while the latter lacks only adaptive immunity. In both mouse strains, enzyme treatment had no effect on the growth of B16-F10 tumors (Fig. 1f and Supplementary Fig. 9a). More specifically, depletion of CD8+ cells abrogated the anti-tumor effect of enzyme treatment (Fig. 1g) resulting in a median survival (21 days) similar to that of untreated mice (23 days) in Figure 1c. Furthermore, Kyn had no direct effect on the growth of tumor cells in vitro (Supplementary Fig. 9b). Collectively, these results indicate that the action of Kynureninase is dependent on the presence of CD8+ cytotoxic T-cells.
Following treatment of C57BL/6 mice bearing large B16-F10 tumors (>100 mm3, Day 10) we observed elevated infiltration of CD8+ cells (Fig. 2a). One defining feature of an effective antitumor immune response is the presence of infiltrating effector CD8+ T-cells into the interior of the tumor as opposed to the margin28. Immunofluorescence examination revealed that in control tumors, CD8+ T-cells localized predominantly at the tumor margins. However, the CD8+ T-cell interior:margin ratio increased 2-fold in tumors from PEG-Kynureninase treated animals (0.39 for control and 0.79 for enzyme treated group (Fig. 2b and Supplementary Fig. 10). We also observed a two-fold increase in CD8+ tumor infiltrating lymphocytes (TIL) expressing the proliferation marker Ki67 (Fig. 2c).
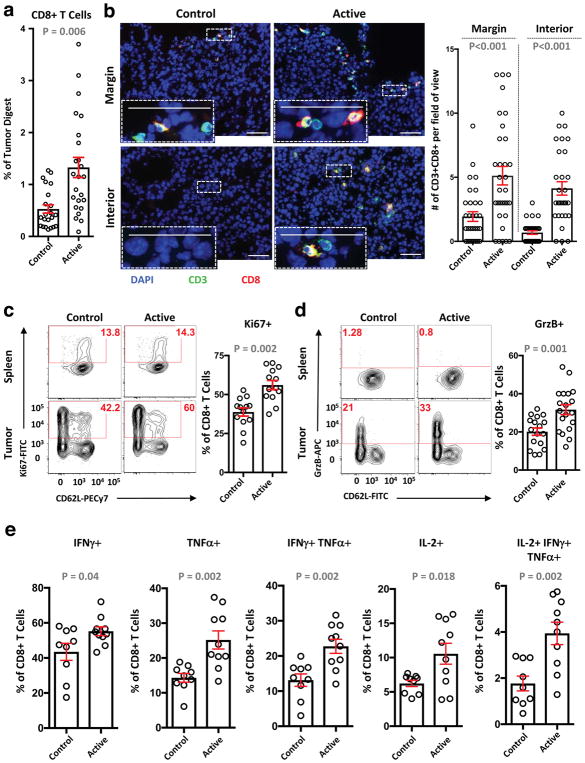
Figure 2. Kynureninase treatment increases the frequency of effector CD8+ TIL.
(a–e) Mice with large B16-F10 tumors (125 mm3 mean +/− 8 mm3, day 10) were treated with two doses of PEG-Kynureninase (20 mg/kg) or heat-deactivated control enzyme administered 72 hours apart. Tumors and spleens were analyzed 24 hours after the second treatment. CD8+ and CD4+ TIL in flow cytometric analysis were identified by surface expression of CD45 and T-cell markers. (a) Percent viable CD8+ TIL in tumor digests (n = 22 deactivated control and n = 24 PEG-Kynureninase compiled from seven independent experiments). (b) Shown are representative images of the margin and interior of frozen tumor sections along with insets from enlargements of the regions indicated by white dotted lines. Graph depicts the collective quantification of CD3+CD8+ cells from 10 fields view for both along the margin and interior of each tumor (n = 3 mice per group, data compiled from two independent experiments with similar results). Scale bars = 50 μm. (c and d) Representative flow cytometric plots of CD8+ TIL and splenic CD8+ T-cells evaluated for expression of (c) Ki67 (n = 13 per group, compiled from four independent experiments) and (d) Granzyme B (n = 16 deactivated control and n = 18 PEG-Kynureninase, compiled from six independent experiments). (e) Production of cytokines by CD8+ TIL after ex vivo stimulation (n = 9 deactivated control and n = 10 PEG-Kynureninase, compiled from three independent experiments). (c–e) Graphs depict cumulative tumor analysis with each symbol representing results from individual tumors and the red bars indicating the mean +/− SEM. Statistical significance was determined by unpaired t-test (two-tailed) for all analyses.
T-cell polyfunctionality reflects a high degree of activation in the TME and correlates with protective antitumor responses in animal models and, importantly, with favorable clinical outcomes in patients29. We observed that enzyme treatment resulted in an increased fraction (2-fold) of CD8+ TIL expressing Granzyme B capable of producing IFNγ, TNFα or IL-2 (Fig. 2d, e and Supplementary Fig. 11). We also observed a smaller increase in CD4+ TIL capable of producing both IFNγ and TNFα (Supplementary Fig. 12, P = 0.04). We did not observe any change in the proportion or proliferation status (Ki67+ cells) of CD4+Foxp3+ Tregs in tumors from treated animals (Supplementary Fig. 12) and only a small effect on MDSCs, which did not reach statistical significance (Supplementary Fig. 13). Similar to these findings, no changes in the frequency of tumor infiltrating CD4+Foxp3+ Tregs were observed after treatment with IDO1 inhibitors in multiple tumor models14,30,31. PEG-Kynureninase treatment resulted in a slight decrease in the percentage of T-cells in the tumor draining lymph node (TdLN) but no change in T-cell activation or the frequency of CD4+Foxp3+ T-cells, CD8+ or MDSCs was observed (Supplementary Fig. 14). Likewise, IDO1 inhibitors had been shown to exert minimal effects on T-cell activation in peripheral organs12,14,30. Collectively, our results demonstrate that PEG-Kynureninase affects CD8+ T-cells in the tumor but not in the periphery.
Earlier studies have documented that small molecules that inhibit or otherwise modulate the immunosuppressive effects associated with IDO1 expression (e.g. indoximod) synergize well with αPD-1 checkpoint inhibition therapy in murine models of melanoma and glioblastoma14,30. Combination treatment of mice bearing large (~125 mm3) B6-F10 tumors with αPD-1+PEG-Kynureninase resulted in 45% complete responses compared to ~10% in the group treated with αPD-1 alone (Fig. 3a). Neither CD8+ nor CD4+ TILs in PEG-Kynureninase-treated animals showed statistically significant differences in PD-1 expression indicating that depletion of Kyn in the TME does not impact T-cell PD-1 status (Supplementary Fig. 15). To determine the durability of the therapeutic response and the induction of immunological memory, we re-challenged the surviving mice 2 months after PEG-Kynureninase/αPD-1 therapy cessation with B16-F10 tumor cells in the opposite flank. We found that 100% of the long-term surviving mice rejected the tumor (Supplementary Fig. 16a).
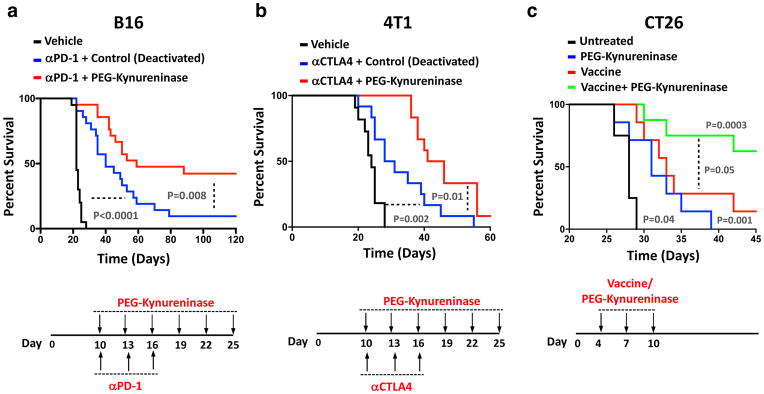
Figure 3. Combination of PEG-Kynureninase with other immunotherapies for the treatment of large established tumors in multiple tumor models.
(a) Survival data for mice with large, established (117 mm3 mean +/− 6.5 mm3) B16-F10 tumors treated with vehicle control (n = 10), αPD-1 + deactivated enzyme (n = 21), or αPD-1 + PEG-Kynureninase (n = 21). Dosing schedule as shown (αPD-1: 10 mg/kg/dose, PEG-Kynureninase: 20 mg/kg/dose). Results are compiled from two independent experiments with similar results (n = 21 treated groups and n = 10 vehicle alone). (b) Survival data of mice bearing large (110 mm3 mean +/− 8 mm3) 4T1 tumors treated with vehicle control (n = 10), αCTLA4+deactivated enzyme (n = 12) or αCTLA4+PEG-Kynureninase (n = 12). Dosing schedule as shown (αCTLA4: 10 mg/kg/dose, PEG-Kynureninase: 20 mg/kg/dose). Graph depicts data from one experiment. (c) Mice challenged with 5 × 105 CT26 cells were treated with Gp96-Ig vaccine (1 × 106 CT26 cells transfected with Gp96-Ig), PEG-Kynureninase (20 mg/kg) or both as shown, with two doses of PEG-Kynureninase on day 10. Graph depicts representative data from one of two independent experiments with similar results (n = 4 untreated mice and n = 7 mice per treatment groups). Statistical significance of treatment groups in comparison to either control groups or between treatment groups (indicated by dashed lines) were determined by Log-rank (Mantel-Cox) test.
We next examined the impact of combination therapy using immune checkpoint antibodies together with PEG-Kynureninase in mice bearing tumors that express IDO1. In mice bearing CT26 tumors, αPD-1+PEG-Kynureninase treatment resulted in complete tumor rejection and immunity to re-challenge with CT26 tumor cells in 50% of the animals (Supplementary Fig. 6b and 7). The 4T1 breast carcinoma model is relevant to human breast cancer owing to its poor immunogenicity, aggressiveness and propensity for metastasis. 4T1 tumor cells express high levels of IDO1 and form very aggressive tumors27. Consistent with previous reports32, animals bearing large 4T1 tumors and treated with αCTLA4 antibody showed a transient decrease in tumor growth but no overall survival benefit. In contrast, PEG-Kynureninase in combination with αCTLA4 showed a ~45% increase in median survival compared to αCTLA4 with deactivated enzyme, with 8% of the mice completely rejecting the tumors and becoming immune to re-challenge (Fig. 3b and Supplementary Fig. 16b). Finally, we evaluated whether PEG-Kynureninase could be combined with a clinically relevant cancer vaccine. The ImPACT vaccine platform comprises engineered tumor cell lines expressing the heat-shock chaperone protein gp96 fused to an IgG-Fc domain. The gp96-Ig binds to endogenous tumor antigens and is secreted to the extracellular space to effectively stimulate cross-presentation, eliciting potent CD8+ T-cell responses33. Mice bearing CT26 tumors received either mitomycin-treated ImPACT CT26 vaccine cells together with inactive enzyme, PEG-Kynureninase alone or combination therapy of ImPACT + PEG-Kynureninase as shown in Figure 3c. ImPACT or PEG-Kynureninase alone had a comparable effect on mean survival relative to vehicle, whereas the combination resulted in 60% tumor-free survival.
Discussion
By employing Kynureninase administration to selectively degrade Kyn in a manner that does not impact Trp catabolism, we provide strong mechanistic evidence using multiple tumor types that accumulation of Kyn in the TME is critical for immunosuppression by the Trp catabolism pathway. This conclusion is consistent with recent literature meta-analyses4–7 and experimental data reported by Sonner and coworkers showing that deletion of GCN2, which had been proposed to be activated by IDO1-mediated depletion of Trp in the TME3, does not affect anti-tumor T-cell responses in the B16 model5.
Of note, our data also demonstrate that pharmacologically optimized PEG-Kynureninase is a promising immunotherapeutic approach for cancer therapy. Multiple small molecule inhibitors of IDO1 are undergoing clinical evaluation and IDO1+TDO dual inhibitors were reported several years ago but are not yet in clinical development1. The most advanced IDO1 inhibitor, epacadostat showed promising efficacy in phase II studies in combination with immune-oncology drugs2. However, a very recent (ECHO301) phase III clinical trial in melanoma failed to show improved efficacy compared to anti-PD1 treatment alone. Kynureninase offers several distinct pharmacological advantages over small molecule inhibitors: First, unlike the small molecule inhibitors of IDO1 currently in the clinic, PEG-Kynureninase eliminates Kyn from the TME irrespective of its source, that is, regardless of whether it is produced by IDO1, TDO or by tumors expressing both enzymes (Supplementary Fig. 17). While dual inhibitors of IDO1 and TDO are in preclinical development, the complete blockade of Trp catabolism is likely to have a series of undesired side effects including among others: (a) substantially elevated concentrations of Trp, which in turn can be carcinogenic34 and also have neurological effects due to the ensuing increase in the serotonin synthesis flux20, 21; (b) the blocking of the synthesis of downstream metabolites of Kynurenine, including neuroprotective compounds such as kynurenic acid and also the production of NAD and niacin, especially in the liver, which constitutively expresses TDO and plays a dominant role in Trp catabolism.
Second, PEG-Kynureninase eliminates Kyn completely and for prolonged periods (Fig. 1d). The complete depletion of the extracellular Kyn pool, in contrast to the effect of IDO1 inhibitors, which only partially reduce systemic levels of Kyn (as they do not impact the constitutive flux originating from the TDO pathway), could lead to the elimination of Kyn spontaneous byproducts that were recently shown to be potent AhR agonists18. The more favorable pharmacokinetics of PEG-Kynureninase (Supplementary Fig. 5) may be particularly advantageous for combination therapy with i.v. administered immune checkpoint antibodies. Along these lines we observed that administration of PEG-Kynureninase exhibited a comparable or stronger anti-tumor effect than clinical-stage small molecule inhibitors of IDO1, most notably in combination therapy with αPD-1 (Supplementary Fig. 18).
Third, most IDO1 inhibitors under clinical investigation (and also published pre-clinical TDO inhibitors) structurally resemble Trp and have various degrees of AhR agonistic activity at physiologically relevant concentrations35. Thus, unlike PEG-Kynureninase, which disrupts the Trp catabolism-AhR axis by virtue of its ability to degrade extracellular Kyn, small molecule inhibitors of IDO1/TDO can result in drug-induced, low level AhR activation, a process that could induce certain immunosuppressive phenotypes 3,16,36.
Fourth, while experience with tyrosine kinase inhibitors suggests that the use of small molecule inhibitors of enzymes that are important for cancer growth frequently results in the acquisition of mutations that enable drug-resistance37, such mechanisms are clearly not relevant for PEG-Kynureninase, whose function is to degrade an immunosuppressive metabolite. Finally, we note that we observed no evidence of toxicities or overt signs of autoimmunity such as weight loss, organ size or vitiligo with PEG-Kynureninase as monotherapy or with the combination therapy regimens tested.
Because the bacterial enzymes used here to demonstrate the therapeutic effect of Kynureninase administration could pose an immunogenicity risk, we are engineering a human enzyme with the requisite Kyn degradation kinetics and high stability in vivo for optimal pharmacodynamic properties as desired for clinical development. While the possibility that abrogation of the suppressive effects IDO pathway by PEG-Kynureninase could stimulate the elicitation of anti-drug antibodies (ADAs) in humans38, the use of a therapeutic agent comprising a human enzyme that is heavily PEGylated and thus shielded from immune recognition is expected to minimize immunogenicity concerns. Kynurenine, and especially its downstream products, have been established to have a plethora of both neurotoxic but also neuroprotective effects (e.g. Kynureninic acid)21. Numerous studies have suggested that an elevated Kyn/Trp ratio in serum is associated with various neuropathologies and conversely a reduction in circulating Kyn could be beneficial for mental health21. Nonetheless, given the complex roles of the Kyn pathway in neurophysiology, the consequences of depleting peripheral Kyn in the brain following enzyme therapy will need to be evaluated.
Materials and Methods
Cloning of Kynureninase genes
Genes encoding 8 different Kynureninase (KYNase) enzymes of bacterial or mammalian origin were synthesized as Integrated DNA Technologies (IDT) gblocks using E.coli optimized codons and an N-terminal 6X-Histidine affinity tag. Full-length Kynureninase genes of Homo sapiens (Hs-KYNase), Mus musculus (Mm-KYNase), Mucilaginibacter Paludis (Mp-KYNase), Cesiribacter andamanesis (Ca-KYNase), Fulvivirga imtechensis (Fi-KYNase), Cyclobacterium marinum (Cm-KYNase), Chlamydophila pecorum (Cp-KYNase), Acinetobacter Baumannii (Ab-KYNase) and Pseudomonas fluorescens (Pf-KYNase), were inserted into a pet28a vector backbone (see Supplementary Table 1).
Protein expression and purification for initial kinetic characterization
For enzyme expression and purification39 proteins were expressed in either BL21(DE3*) E. coli or in the BL21 derivative strain, C41(DE3)40. E. coli transformants were plated on LB agar plates containing 50 μg/mL kanamycin. After overnight growth at 37ºC, single colonies were used to inoculate 3 mL starter cultures of Difco™ LB Broth, Lennox (Becton, Dickinson, and Company) plus 50 μg/mL kanamycin and grown overnight at 37ºC. Subsequently, 1 mL of the starter culture was used to inoculate 500mL of Difco™ Terrific Broth (Becton, Dickinson, and Company) + 50μg/mL kanamycin. 500mL cultures were grown to an OD600 = 1.0 at 37°C, before overnight induction with 0.5 μM Isopropyl β-D-1-thiogalactopyranoside (Fisher) at 25°C. Cells were pelleted by centrifugation and then re-suspended in 25 mL of lysis buffer containing 100 mM sodium phosphate pH 8.0, 1 mM pyridoxal-5-phosphate monohydrate (PLP, Fisher), 300mM NaCl, 1 mM phenylmethylsulfonyl fluoride (PMSF, Sigma), 25 mM imidazole (Fisher), 0.1 % Tween 20 (Sigma) and 25 U/mL Universal Nuclease (Pierce). Resuspended cells were lysed with a FRENCH® Press cell disruptor (Thermo Electron Corporation) at 1,500 psi, and solids were removed by centrifugation at 20,000 × g for 1 h. All subsequent steps were performed at 25°C unless otherwise stated. Lysate supernatant was filtered through a 0.22 μm syringe filter and applied via gravity flow to an Ni-NTA (Qiagen) column pre-equilibrated with wash buffer containing 300 mM NaCl, 100 mM sodium phosphate pH 8.0, 25 mM imidazole and 0.1 % Tween 20. The column was washed with 20 column volumes (CV) of buffer containing 300 mM NaCl, 50 mM sodium phosphate pH 8.0, 25 mM imidazole and 0.1% Tween 20. The protein was eluted from the column by adding 5 CV of buffer containing 300 mM NaCl, 100 mM sodium phosphate pH 8.0, 5 mM PLP and 300 mM imidazole. To ensure full loading of PLP, the eluted protein was then incubated at 37°C for 1–2 h in the elution buffer. Purified protein was then dialyzed against 50 mM Tris-HCl pH 8.5 overnight at 4°C to remove excess PLP and imidazole. For long-term storage, glycerol was added to a final concentration of 15 % v/v, and the protein solution was flash frozen in liquid nitrogen and stored at −80°C.
Kinetic analysis of Kynureninase enzymes
A range of substrate concentration from 0 to 1 mM and enzyme (0.01 to 1 μM) were used to determine Michaelis-Menten kinetic parameters for the kynureninase enzymes against both kynurenine (Kyn, Sigma) and 3-Hydroxy-DL-kynurenine (DL-3-OH-Kyn Sigma). Reactions were initiated by adding 80 μl of substrate solution to 20 μL of enzyme solution in Dulbelcco’s phosphate buffered saline pH = 7.4 (DPBS). Enzymatic activity was evaluated by monitoring substrate absorbance over time using a BioTek Synergy HT 96-well plate spectrometer. Absorbance at 365 nm was monitored for Kyn degradation, and at 373 nm for DL-3-OH-Kyn. Initial rates were determined from the linear region of the reaction progress curves with <10% of substrate conversion. Michaelis-Menten parameters, kcat and KM, were determined using KaleidaGraph (Synergy software).
Preparation of PEGylated enzyme for animal studies
Pf-Kynureninase and Mp-Kynureninase were expressed from 8 L cultures grown and induced as described above. Cells were pelleted by centrifugation and then re-suspended in 800mL of buffer containing 50 mM sodium phosphate pH 8.0, 1 mM PLP, 300 mM NaCl, 1 mM PMSF, 25 mM imidazole, 0.1 % Tween 20, and 25 U/m Universal Nuclease. Resuspended cells were lysed by performing three sequential passes through a Nano DeBee homogenizer (B.E.E. International, Inc.) at 30,000 psi. Solids were removed from the lysed cells by centrifugation at 20,000 x g for 30 minutes at 4°C, and the lysate was passed through at 5 μM filter. To ensure low endotoxin levels, glass columns pre-cleaned with 1 M NaOH were used for column preparations for all purifications. 17 mL Ni-NTA resin was pre-equilibrated with buffer containing 50 mM sodium phosphate pH 8.0, 0.1 mM PLP, 300mM NaCl, 1 mM PMSF, 25 mM imidazole, 0.1 % Tween 20, and 25 U/mL Universal Nuclease in a cold room at 4°C. Filtered lysate was then passed over the pre-equilibrated column using a peristaltic pump at a flow rate of 3 mL/min. The resin was then washed with 5 CV of buffer containing 50 mM sodium phosphate pH 8.0, 0.1 mM PLP, 300mM NaCl, and 25 mM imidazole, followed by 2.5 L of endotoxin-free GIBCO® PBS pH 7.4 (DPBS, Thermo Fisher) with 0.1 % Triton™ X-114 (Sigma) + .1 mM PLP at a flow rate of 5 mL/min, followed by 70 mL of endotoxin-free PBS. Enzymes were eluted with 200 mL 0.22 μM filtered elution buffer containing 100 mM sodium phosphate, pH 8.0, 300mM NaCl, and 300 mM imidazole, and PLP was added to a concentration of 1 mM. After incubation at 37°C for 1 h, enzymes were buffer exchanged with 100 mM sodium phosphate pH 8.5 using a Pall Corporation Minimate™ Tangential Flow Filtration dia-filtration system (10K capsule), and then concentrated to 5 mg/L by centrifugation at 4000g at 4°C in Vivaspin Turbo 15 10,000 MWCO ultrafiltration units (Sartorius, Goettingen, Germany). PLP was added to a concentration of 1 mM and enzymes were incubated at room temperature for 30 minutes. Enzymes were then PEGylated using a 100-fold molar excess of Methoxy-PEG-CO(CH2)2COO-NHS, MW 5,000 Da (NOF America Corporation) powder which was added directly to the concentrated enzyme solution and stirred slowly for 0.5–1 h at room temperature. Then dia-filtration with a 30K capsule was used to remove excess Methoxy-PEG-CO(CH2)2COO-NHS and to buffer exchange into endotoxin-free DPBS. The PEGylated Kynureninase protein was further concentrated by centrifugation at 4000g at 4°C in Vivaspin Turbo 15 10,000 MWCO ultrafiltration units. Glycerol was added to the concentrated enzyme such that the final concentrations of PEGylated enzyme and glycerol were 10 mg/mL and 15% v/v, respectively. PEGylated-Kynureninase was then 0.22 μM sterile filtered and flash frozen in liquid nitrogen and stored at −80°C. After at least 24 h, a small portion of enzyme was thawed and analyzed for catalytic activity. Endotoxin levels were determined using the Limulus Amebocyte Lysate (LAL) Chromogenic Endotoxin Quantitation Kit (Pierce). Purity and extent of PEGylation were determined both by SDS-PAGE on a 4–20% gradient gel and size exclusion chromatography (SEC). The latter was performed on an Amersham/Pharmacia P-900 FPLC fitted with a Sephadex S-200 10/300 column (GE Healthcare). Samples were run in PBS at an initial concentration of 1 mg/mL with an injection volume of 1 mL.
Standards and reagents
Formic acid (FA), methanol (MeOH), dimethylsulfoxid (DMSO) and acetonitrile (ACN) of analytical grade were from Fisher Scientific (Pittsburgh, PA, USA). Water was of ultrapure grade (EMD Millipore Co., Billerica, MA, USA). Deuterated standards {Fumaric acid (2,3-D2, 98% purity), DL-Glutamic acid (2,4,4-D3, 98% purity), L-Tryptophan (indole-D5, 98% purity) and Nicotinic acid (D4, 98% purity)} were from Cambridge Isotope Laboratories (Cambridge Isotope Laboratories Inc., Tewksbury, MA, USA). Deuterated Anthranilic acid (3,4,5,6-D4, 98% purity), Kynurenic acid (3,4,6,7,8-D5, 98% purity), Kynurenine (D4, 98% purity), Quinolinic acid (4,5,6-D3, 99% purity), 3-Hydroxy-DL-kynurenine (5,8,8-D3, 98% purity), 3-Hydroxyanthranilic acid (4,6-D2, 98% purity) were from Buchem (Buchem BV, Apeldoorn, The Nederlands). Deuterated Picolinic acid (99% purity) was from Acros Organics (Thermo Fisher Sci, NJ, USA). Commercial calibration solutions for the MS device were from Thermo Scientific (Thermo Fisher Sci., San José, CA, USA).
Liquid chromatography-electrospray/high resolution-mass spectrometry (HPLC-ESI/HRMS) analysis
Chromatographic separations were performed on a Thermo Scientific Accela HPLC system equipped with a quaternary pump, vacuum degasser and an open autosampler with a temperature controller. Separation of metabolites was achieved through a 150 mM × 2.1 mM i.d., 4 μM particle size Synergi Hydro-RP C18 column (Phenomenex Inc, Torrance, CA, USA). Separation conditions were: solvent A, water/FA (99.8:0.2); solvent B, ACN; separation gradient, initially 1% B, held for 2 minutes and then linear 30–80% B in 8 minutes, washing with 98% B for 5 minutes and column equilibration with 1% B for 15 minutes; flow rate, 0.25 mL/min; injection volume, 5μl; autosampler temperature, 6°C; column temperature, 22°C.
Mass spectrometry analysis was carried out on a Thermo Scientific (Thermo Fisher Sci., Bremen, Germany, EU) Q Exactive benchtop Orbitrap detector loading an electrospray (ESI) source simultaneously operating in fast negative/positive polarity switching ionization mode. The detector was run in full scan MS analysis under the following conditions: spray voltage, 4.0 kV; capillary temperature, 300°C; sheath gas, 55 (arbitrary units); auxiliary gas, 30 (arbitrary units); microscans, 1; AGC target, 1e6; maximum injection time, 100 ms; mass resolution, 70,000; considered m/z range, 115–450. To ensure the accuracy of the MS analysis, the detector was calibrated through the commercial calibration solutions provided by the manufacturer followed by a customized adjustment for small molecular masses. Thus, masses at m/z 87.00877 (Pyruvic acid); 117.01624 (D2-Fumaric acid); 149.06471 (D3-Glutamic acid); 265.14790 (Sodium dodecyl sulfate) and 514.288441 (Sodium taurocholate) were used for the negative ionization mode whereas masses at m/z 74.09643 (n-Butylamine), 138.06619 (Caffeine fragment), 195.08765 (Caffeine) and 524.26496 (Met-Arg-Phe-Ala pentapeptide, MRFA) adjusted the mass accuracy of the positive ionization mode. The MS detector was calibrated before the analysis of each batch of samples, keeping the mass tolerance below 3 ppm during the time of this research. The LC-MS platform of analysis was controlled by a PC loading the software Xcalibur v. 2.2 SP1.48 (Thermo Scientific, San Jose, CA, USA). Detailed chromatographic and MS properties of determined metabolites are summarized in Supplementary Table 2.
LC-MS data analysis
Raw LC-MS data was automatically processed by SIEVE v. 2.2.58 SP2 (Thermo Fisher Sci., San Jose, CA, USA) loading the following settings: frame parameter adjustment, m/z range 115–450; frame time width, automatic; m/z width, 10 ppm; considered running time, 15 minutes; Intensity threshold, 10,000 minimum counts; minimum scans across the peak, 5; signal-to-noise background, 3; an in-house database listing all the target metabolites of the Kynurenine pathway metabolism was loaded using 3 ppm as mass tolerance to perform the positive assignments; max RT shift, 0.2 min; MaxCharge, 1; Peak algorithm to peak extraction, PPD; Peak integration algorithm, ICIS (previously optimized using Xcalibur and a real sample as reference); DBlookup method, FRAMEMZ. Absolute quantification was performed using the spiked known amount of their respective deuterated counterparts through the formula:
Where Wx and Ax are, respectively, the weight and the area under the peak corresponding to the identified metabolite; Wis and Ais are, respectively, the weight of added deuterated internal standard and the area under its corresponding peak; and Fx,is is the relative response factor of the metabolite against its respective deuterated counterpart (1, in all cases). Results were normalized with respect to the measured volume and weight of serum and tissue samples, respectively. In contrast, relative quantification was done through the signals from metabolites and the closest deuterated standard eluted according to the formula:
Absolute quantitation was carried out by spiking samples with an internal standards solution containing an accurate weight of all the deuterated standards mentioned above, with the exception of D2-Fumaric and D3-Glutamic acids which were first solubilized in water/MeOH (50:50) with 2% of DMSO.
To analyze tumor tissue concentrations of tryptophan catabolism metabolites, C57B6L/6j inbred mice were inoculated with 5 × 104 B16-F10 cells on the right flank. Once tumors 25–35mm2, (~12 days after inoculation), mice were then treated with one dose of 20 mg/kg Pf-KYNase pt.. Tumors were excised at the indicated time points, snap frozen in liquid nitrogen and stored at −80°C. Tissue aliquots were weighed and placed in 2 mL tissue homogenization tubes, mixed with beads. After which, 0.5 mL of chilled water/MeOH (50:50) was added. Samples were homogenized in a Precellys-24 cryohomogenizer (Bertin Technologies, Saint Quentin en Yvelines, France) at −5 °C at 5000 rpm during two cycles of 20 s. Lysates were recovered and poured into glass vials containing 0.5 mL of chilled chloroform. Homogenization tubes were washed with an additional 0.5 mL of chilled water/MeOH (50:50). Glass vials were stirred at 1,500 rpm for 3 minutes, centrifuged at 5,000 rpm for 20 minutes at 4 °C and the methanolic supernatants were poured in Eppendorf tubes to be finally desiccated until dehydrated in a Centrivap vacuum concentrator at 4 °C (Labconco, Kansas City, MO, USA). Samples were re-suspended in 15 μL of the internal standards solution and 135 μL of ultrapure water. The resulting 150 μL solutions were filtered and stored at −80 °C until LC-MS analysis.
To analyze serum concentrations of tryptophan catabolism metabolites, serum from the tumor bearing mice described in the paragraph above was collected following a terminal bleed. Blood was clotted at room temperature for 10 min and then centrifuged at 500 × g for 5 minutes. The serum fraction was drawn off and centrifuged a second time at 1000 × g for 5 minutes, and then stored at −80°C until use. 10 μL of samples were diluted with 15 μL of the internal standards solution (incorporating a weight range of 7.3–8.5 ng) and 125 μL of water. The resulting 150 μL were stirred and subsequently cleaned from solids and proteins through a Nanosep 3K ultra centrifugal filter (Pall Co. Port Washington, NY, USA) at 8,000 rpm for 2 h at 4 °C. The resultant filtrates were poured into the LC vials and stored at −80 °C until LC-MS analysis.
In vitro T-cell and tumor cell line assays
Naïve CD8+ lymphocytes were magnetically isolated from single-cell suspensions prepared from spleens of tumor-free C57B6L/6j mice using the EasySep™ Mouse Naïve CD8+ T Cell Isolation Kit (Stemcellï Technologies). Enriched naïve CD8+ T cells (2 × 105) were stimulated in U-bottom 96-well plates pre-coated with 2 μg/mL of anti-CD3 (17A2, Ebioscience) and anti-CD28 (37.51, Ebioscience) in the presence or absence of the indicated concentrations of Kyn, Kynureninase (Mp-KYNase, 1 μM) or vehicle control (10% DMSO in PBS). After being stimulated for 48 h, cells were transferred to new wells and rested for 24 h. Cells were then analyzed for viability by Annexin-V (Biolegend) and propidium iodide (PI) (Enzo Life Sciences) staining. Proliferation was determined by CFSE dilution. Cells were cultured at 37°C in complete RPMI media composed of Roswell Park Memorial Institute 1640 medium (RPMI 1640) supplemented with 10% FBS, 1× non-essential amino acids, 55μM β-mercaptoethanol, 2 mM GlutaMAX, 1× penicillin-streptomycin-glutamine (100 U/mL penicillin, 100 μg/mL streptomycin, 292 μg/mL glutamine) and 1mM sodium pyruvate (all from GIBCO®).
In vitro enzymatic activity on human cell lines
SKOV3 human ovary adenocarcinoma (IDO1+) and A172 human glioblastoma (TDO+) cell lines were obtained from ATCC and cultured in McCoy’s 5A (10% FBS) and Dulbecco’s Modified Eagle’s Medium (DMEM, 10% FBS) culture media, respectively. Cell lines were incubated at 37°C in 96 well plates with either the IDO1 inhibitor INCB024360 (Epacadostat, Selleck) at 2 μM for SKOV3 cells and 20 μM for A172 cells; the TDO inhibitor 680C91 (WuXi Apptec) at 20 μM for all cells; or Kynureninase (Mp-KYNase) at 0.36 mg/ml for all cells. In addition, A172 cells were stimulated with recombinant human IFNγ (100 ng/mL, Sigma) to induce IDO1 expression to determine the effects on Kynureninase, INCB024360 and 680C91 on Kyn levels made by IDO1/TDO dual expressing cells. After 48 h, cell supernatants were collected, snap frozen in liquid nitrogen and stored at −80°C. Kynurenine concentrations of the supernatant were then determined by LC/MS. McCoy’s 5A, DMEM and FBS were all purchased from GIBCO®.
Animal experiments
C57BL/6J, BALB/cJ, Ido1−/−, Rag1−/− and NOD-Scid IL2rγc−/− (NSG) mice were purchased from Jackson Laboratories (Bar Harbor, Maine, USA). All mice were maintained in pathogen-free, ventilated cages with irradiated food and autoclaved water at The University of Texas at Austin (UT Austin) or WuXi AppTec. Experimental procedures were approved by the Institutional Animal Care and Use Committee (IACUC) at UT Austin or WuXi AppTec. Mice were monitored daily and euthanized by CO2 asphyxiation and cervical dislocation prior to any signs of distress.
Pharmacokinetic analysis of PEG-Pf-Kynase in vivo
BALB/cJ mice were treated intravenously with a single 20 mg/kg dose of PEG-Kynureninase and then sacrificed 1, 2 or 3 days after treatment to collect samples for analysis using a sandwich ELISA for detecting the concentrations of PEG-Kynureninase. Briefly, 96-well Maxisorp™ plates (Thermo Fisher) were coated at 4°C overnight with 5 μg/mL of anti-PEG antibody (Abcam PEG-B-47 - ab51257) suspended in PBS. After washing five times with PBS + 0.05% Tween 20 (PBST), the plate was blocked with 1% PBS-Milk (PBSM) for 1 h at room temperature and then washed with PBST. Serum diluted 1:100 in PBSM was added to the plate for 1 h at room temperature and then washed with PBST. Subsequently, 100 μL (0.83μg/ml) of HRP-labeled anti-PEG antibody (Abcam PEG-B-47 - ab202640) diluted 600-fold in PBSM were added for 1 h at room temperature, followed by washing with PBST. Plates were developed by adding 100 μL of 1-Step™ Ultra TMB-ELISA Substrate Solution (Thermo, 34028), incubating at room temperature for 15 minutes, then adding 100 μL of 0.3 M sulfonic acid, and then measuring the absorbance at 450 nm. A standard curve was generated from known PEGylated Kynureninase concentrations and used to convert absorbance to enzyme concentration.
Tumor cell culture and vaccine cell line preparation
Murine B16-F10 melanoma (CRL-6475), 4T1 metastatic mammary carcinoma (CRL-2539), and CT26 colon carcinoma (CRL-2638) cell lines were purchased from the American Type Culture Collection (ATCC), cultured according to manufacturer’s directions and tested for mycoplasma using the Universal Mycoplasma test kit (ATCC). For vaccination studies, CT26 mouse colon carcinoma cells used for transfection were maintained in Iscove’s Modified Dulbecco’s Media (IMDM) supplemented with glutamine and 10% Fetal Bovine Serum. The CT26-gp96-Ig (ImPACT, Heat Biologics, North Carolina) cell line was generated by transfecting CT26 cells with a DNA plasmid expressing murine gp96-Ig using Effectene transfection reagents (Qiagen Hilden, Germany) according to manufacturer’s directions. Transfectants were selected with G418 and a single cell clone secreting a high-level of gp96-Ig was isolated by limiting dilution. Gp96-Ig levels were determined by ELISA. All cells were cultured at 37°C with 5% CO2.
Tumor models and therapy
C57B6L/6j inbred males and females aged 6–8 weeks were inoculated with 5 × 104 B16-F10 cells into the right flank. BALB/c mice were inoculated with 5 × 104 of CT26 cells into the right flank or with 5 × 104 4T1 cells into the mammary fat pad. Mice were treated with peritumoral injections of PEG-Kynureninase (20 mg/kg) or deactivated PEG-Kynureninase (heat-deactivated for 10 minutes at 100°C). For CD8 depletion studies, mice were anti-CD8 (10mg/kg, clone YTS 169.4, Bio X Cell) or isotype control (10mg/kg, clone LTF-2, Bio X Cell) were administered 1 day prior to treatment with PEG-Kynureninase and then every 7th day for a total of 3 doses. Anti-CTLA4 (10 mg/kg, clone UC10-4F10-11, Bio X Cell) or anti–PD-1 (10 mg/kg, clone RMP1-14, Bio X Cell) were administered by intraperitoneal injections as indicated. Epacadostat was administered by oral lavage twice daily (100mg/kg) and NLG919 was administered in the drinking water (3mg/ml). Tumor size was tracked three times weekly for the duration of the experiment using calipers. Mice were sacrificed once tumor sizes reached 4000 mm3.
For cancer vaccine efficacy studies, first tumors were established by injecting 5 × 105 CT26 cells into the right flank of BALB/c mice. Vaccination was performed by intraperitoneal injection of 1 × 106 CT26-Gp96-Ig cells (described above) pre-treated with mitomycin-C (10 μg/mL) at 37°C in 5% CO2 for 2–3 h administered on days 4, 7 and 10. PEG-Kynureninase or deactivated enzyme was injected peritumorally one time on days 4 and 7 and two times on day 10.
Tumor immune infiltrate phenotyping
Mice with established tumors (~125 mm3) were administered two peritumoral injections of PEG-Pf-Kynureninase three days apart and necropsied 24 h after the second dose. Tumors, spleens, tumor draining lymph nodes (TdLN) and contralateral inguinal lymph nodes (iLN) were processed to obtain single cell suspensions by mechanical digestion and treated with red blood cell lysis buffer (BioLegend). Cells were then strained though a 40 μm filter and stained with the following fluorescently labeled antibodies toward surface proteins: CD3ε (145-2C11), TCRβ (H57-597), CD5 (53-7.3), CD45.2 (104), CD8 (53-6.7), CD4 (GK1.5), CD25 (PC61.5), PD-1 (29.F1A12), Gr1 (RB6-8C5), CD11b (M1/70) and CD62L (MEL-14). For all intracellular protein analysis, cells were co-incubated with a fixable viability dye (Tonbo Biosciences) during surface staining, which were then fixed and permeabilized using either the Foxp3 Staining Buffer (Ebioscience) or BD Cytofix/Cytoperm™ kit according to the manufactures’ instructions. Permeabilized cells were then incubated with fluorescently labeled antibodies toward Ki67 (SolA15), Granzyme B (GB11) or Foxp3 (FJK-16s).
To determine cytokine production, 1 × 107 cells from tumor digests were cultured in 1 mL of complete RPMI media (described above) and stimulated at 37°C with phorbol myristate acetate (PMA)/Ionomycin (Ebioscience) in 48 well plates for 1 h, at which point Brefeldin A (Ebioscience) added. Cells were stimulated for an additional 4 h. After stimulation, cells were surface stained with fluorescently labeled antibodies and viability dye, and then permeabilized using the BD Cytofix/Cytoperm™ kit. Intracellular cytokine production was determined using fluorescently labeled antibodies toward TNFα (TN3-19.12), IFNγ (XMG1.2) and IL-2 (JES6-5H4). All stains were done in flow cytometry wash buffer (PBS containing 1% FBS and 0.5 mM EDTA) at 4°C for 30 minutes. All antibodies were purchased from Ebioscience, BioLegend or Tonbo Biosciences. An LSRFortessa instrument (BD Biosciences) was used for flow cytometric data acquisition and analyzed with FlowJo (Treestar).
Immunostaining and microscopy
Immunostaining of B16-F10 tumors was performed on 7 μm acetone-fixed cryosections according to standard protocols. The tissues were incubated with fluorophore-conjugated anti-CD3 (17A2) and anti-CD8 (53-6.7) antibodies at 4°C followed by nuclear DAPI counterstain. Images were acquired using an Axio Imager A1 microscope (Zeiss,), with an AxioCam MRm camera (Zeiss), Zeiss EC Plan-Apochromat 10x/0.45 and Plan-Apochromat 20x/0.8 objectives with AxioVision 4.8 software (Zeiss). Adobe Photoshop v12.0 (Adobe) was used for overlays of fluorescent channels, and to adjust exposure uniformly across the images. For quantitation, ten random, non-overlapping regions along the margin and interior were imaged for each tumor. The number of CD3+CD8+ nucleated cells was then counted for each image. Sections stained with DAPI only were used as controls and the treatment group that each tumor belonged to was unknown at the time to the individual doing the imaging and counting.
Statistical analysis
Statistical significance (p-value) was determined by unpaired student unpaired (two-tailed) t test and Log-rank (Mantel Cox) test. Significant p-values are denoted by *P<.05, **P<.01, ***P<.001 and ****P<.0001. All statistical analysis was performed using GraphPad Prism software (GraphPad).
Data Availability
The data that support the findings of this study are available from the corresponding author upon reasonable request. Further details of experimental design and reagents can also be found in the Nature Research Life Sciences Reporting Summary linked to this article.
Supplementary Material
Acknowledgments
We are grateful to Nora Ashoura for assistance with various aspects of this work, data interpretation and/or comments on the manuscript. This work was supported by funding from NIH 1 RO1 CA189623 (E.S. and G.G.), the Cancer Prevention and Research Institute of Texas grant DP150061 (E.S., L.I.R.E. and G.G.) and Kyn Therapeutics (G.G. and E.S.). Support from the American Cancer Society was provided to T.A.T. (Postdoctoral Fellowship 126584-PF-14-216-01-TBF), N.M. (Postdoctoral Fellowship 123506-PF-13-354-01-CDD) and J.B. (Postdoctoral Fellowship 128252-PF-15-143-01-CDD). M.D. acknowledges support by a post-doctoral fellowship from the Cancer Prevention and Research Institute of Texas (RP140108).
Footnotes
Author Contributions
T.A.T., K.C.G., N.M and M.D. designed and performed key experiments; J. B., C. L., A. Q., B.T., Y.T., W-C.L., C.S.K., K.F. and M.S.Y. expressed, cloned, characterized and prepared enzymes for in vitro and in vivo studies; B.T. and Y.K. developed and performed PK assays; J.D.D., M.M., M.Z., G.F., K.M., S.C. and T.H.S. designed and performed certain in vivo and in vitro experiments; E.S. and S.T. performed metabolomics analyses; T.A.T., K.C.G., M.D., J.D.D., L.I.R.E, G.G. and E.S. interpreted data; G.G., T.A.T., K.C.G. and E.S. wrote the manuscript.
Competing Financial Interests
The authors declare competing financial interests: details are available in the online version of the paper.
References
- 1.Cheong JE, Sun L. Targeting the IDO1/TDO2-KYN-AhR Pathway for Cancer Immunotherapy - Challenges and Opportunities. Trends Pharmacol Sci. 2018;39:307–325. doi: 10.1016/j.tips.2017.11.007. [DOI] [PubMed] [Google Scholar]
- 2.Prendergast GC, Malachowski WP, Duhadaway JB, Muller AJ. Discovery of IDO1 Inhibitors: From Bench to Bedside. Cancer Research. 2017;77:6795–6811. doi: 10.1158/0008-5472.CAN-17-2285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Munn DH, Mellor AL. Indoleamine 2,3-dioxygenase and tumor-induced tolerance. 2007;117:1147–1154. doi: 10.1172/JCI31178. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Badawy AAB, Namboodiri AMA, Moffett JR. The end of the road for the tryptophan depletion concept in pregnancy and infection. Clin Sci. 2016;130:1327–1333. doi: 10.1042/CS20160153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Sonner JK, et al. The stress kinase GCN2 does not mediate suppression of antitumor T cell responses by tryptophan catabolism in experimental melanomas. Oncoimmunology. 2016;5:1–12. doi: 10.1080/2162402X.2016.1240858. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Van de Velde LA, et al. Stress Kinase GCN2 Controls the Proliferative Fitness and Trafficking of Cytotoxic T Cells Independent of Environmental Amino Acid Sensing. Cell Rep. 2016;17:2247–2258. doi: 10.1016/j.celrep.2016.10.079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Platten M, Wick W, Van den Eynde BJ. Tryptophan catabolism in cancer: beyond IDO and tryptophan depletion. Cancer Research. 2012;72:5435–5440. doi: 10.1158/0008-5472.CAN-12-0569. [DOI] [PubMed] [Google Scholar]
- 8.Puccetti P, et al. Accumulation of an endogenous tryptophan-derived metabolite in colorectal and breast cancers. PLoS ONE. 2015;10:e0122046. doi: 10.1371/journal.pone.0122046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Fatokun AA, Hunt NH, Ball HJ. Indoleamine 2,3-dioxygenase 2 (IDO2) and the kynurenine pathway: characteristics and potential roles in health and disease. Amino Acids. 2013;45:1319–1329. doi: 10.1007/s00726-013-1602-1. [DOI] [PubMed] [Google Scholar]
- 10.Spranger S, et al. Up-regulation of PD-L1, IDO, and T(regs) in the melanoma tumor microenvironment is driven by CD8(+) T cells. Sci Transl Med. 2013;5:200ra116–200ra116. doi: 10.1126/scitranslmed.3006504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Mondanelli G, et al. A Relay Pathway between Arginine and Tryptophan Metabolism Confers Immunosuppressive Properties on Dendritic Cells. Immunity. 2017;46:233–244. doi: 10.1016/j.immuni.2017.01.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Holmgaard RB, Zamarin D, Munn DH, Wolchok JD, Allison JP. Indoleamine 2,3-dioxygenase is a critical resistance mechanism in antitumor T cell immunotherapy targeting CTLA-4. Journal of Experimental Medicine. 2013;210:1389–1402. doi: 10.1084/jem.20130066. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Ninomiya S, et al. Tumor indoleamine 2,3-dioxygenase (IDO) inhibits CD19-CAR T cells and is downregulated by lymphodepleting drugs. Blood. 2015;125:3905–3916. doi: 10.1182/blood-2015-01-621474. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Spranger S, et al. Mechanism of tumor rejection with doublets of CTLA-4, PD-1/PD-L1, or IDO blockade involves restored IL-2 production and proliferation of CD8(+) T cells directly within the tumor microenvironment. J Immunother Cancer. 2014;2:3. doi: 10.1186/2051-1426-2-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Thaker AI, et al. IDO1 metabolites activate β-catenin signaling to promote cancer cell proliferation and colon tumorigenesis in mice. Gastroenterology. 2013;145:416–25. e1–4. doi: 10.1053/j.gastro.2013.05.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Mezrich JD, et al. An interaction between kynurenine and the aryl hydrocarbon receptor can generate regulatory T cells. The Journal of Immunology. 2010;185:3190–3198. doi: 10.4049/jimmunol.0903670. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Opitz CA, et al. An endogenous tumour-promoting ligand of the human aryl hydrocarbon receptor. Nature. 2011;478:197–203. doi: 10.1038/nature10491. [DOI] [PubMed] [Google Scholar]
- 18.Seok SH, et al. Trace derivatives of kynurenine potently activate the aryl hydrocarbon receptor (AHR) Journal of Biological Chemistry. 2018;293:1994–2005. doi: 10.1074/jbc.RA117.000631. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Theofylaktopoulou D, et al. A community-based study on determinants of circulating markers of cellular immune activation and kynurenines: the Hordaland Health Study. Clin Exp Immunol. 2013;173:121–130. doi: 10.1111/cei.12092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Vécsei L, Szalárdy L, Fülöp F, Toldi J. Kynurenines in the CNS: recent advances and new questions. Nat Rev Drug Discov. 2012;12:1–19. doi: 10.1038/nrd3793. [DOI] [PubMed] [Google Scholar]
- 21.Cervenka I, Agudelo LZ, Ruas JL. Kynurenines: Tryptophan’s metabolites in exercise, inflammation, and mental health. Science. 2017;357:eaaf9794. doi: 10.1126/science.aaf9794. [DOI] [PubMed] [Google Scholar]
- 22.Phillips RS. Structure and mechanism of kynureninase. Arch Biochem Biophys. 2014;544:69–74. doi: 10.1016/j.abb.2013.10.020. [DOI] [PubMed] [Google Scholar]
- 23.Stone E, et al. Strategies for optimizing the serum persistence of engineered human arginase I for cancer therapy. J Control Release. 2012;158:171–179. doi: 10.1016/j.jconrel.2011.09.097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Frumento G, et al. Tryptophan-derived catabolites are responsible for inhibition of T and natural killer cell proliferation induced by indoleamine 2,3-dioxygenase. J Exp Med. 2002;196:459–468. doi: 10.1084/jem.20020121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Hou DY, et al. Inhibition of indoleamine 2,3-dioxygenase in dendritic cells by stereoisomers of 1-methyl-tryptophan correlates with antitumor responses. Cancer Research. 2007;67:792–801. doi: 10.1158/0008-5472.CAN-06-2925. [DOI] [PubMed] [Google Scholar]
- 26.Banerjee T, et al. A key in vivo antitumor mechanism of action of natural product-based brassinins is inhibition of indoleamine 2,3-dioxygenase. Oncogene. 2008;27:2851–2857. doi: 10.1038/sj.onc.1210939. [DOI] [PubMed] [Google Scholar]
- 27.Levina V, Su Y, Gorelik E. Immunological and Nonimmunological Effects of Indoleamine 2,3-Dioxygenase on Breast Tumor Growth and Spontaneous Metastasis Formation. Journal of Immunology Research. 2012;2012:1–12. doi: 10.1155/2012/173029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Pagès F, et al. In situ cytotoxic and memory T cells predict outcome in patients with early-stage colorectal cancer. Journal of Clinical Oncology. 2009;27:5944–5951. doi: 10.1200/JCO.2008.19.6147. [DOI] [PubMed] [Google Scholar]
- 29.Yuan J, et al. CTLA-4 blockade enhances polyfunctional NY-ESO-1 specific T cell responses in metastatic melanoma patients with clinical benefit. Proc Natl Acad Sci USA. 2008;105:20410–20415. doi: 10.1073/pnas.0810114105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Wainwright DA, et al. Durable therapeutic efficacy utilizing combinatorial blockade against IDO, CTLA-4, and PD-L1 in mice with brain tumors. Clin Cancer Res. 2014;20:5290–5301. doi: 10.1158/1078-0432.CCR-14-0514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Holmgaard RB, Zamarin D, Lesokhin A, Merghoub T, Wolchok JD. Targeting myeloid-derived suppressor cells with colony stimulating factor-1 receptor blockade can reverse immune resistance to immunotherapy in indoleamine 2,3-dioxygenase-expressing tumors. EBioMedicine. 2016;6:50–58. doi: 10.1016/j.ebiom.2016.02.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Grosso JF, Jure-Kunkel MN. CTLA-4 blockade in tumor models: an overview of preclinical and translational research. Cancer Immun. 2013;13:5. [PMC free article] [PubMed] [Google Scholar]
- 33.Schreiber TH, Raez L, Rosenblatt JD, Podack ER. Tumor immunogenicity and responsiveness to cancer vaccine therapy: the state of the art. Semin Immunol. 2010;22:105–112. doi: 10.1016/j.smim.2010.02.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Chung KT, Gadupudi GS. Possible roles of excess tryptophan metabolites in cancer. Environ Mol Mutagen. 2011;52:81–104. doi: 10.1002/em.20588. [DOI] [PubMed] [Google Scholar]
- 35.Moyer BJ, et al. Indoleamine 2,3-dioxygenase 1 (IDO1) inhibitors activate the aryl hydrocarbon receptor. Toxicol Appl Pharmacol. 2017;323:74–80. doi: 10.1016/j.taap.2017.03.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Terness P, et al. Inhibition of allogeneic T cell proliferation by indoleamine 2,3-dioxygenase-expressing dendritic cells: mediation of suppression by tryptophan metabolites. 2002;196:447–457. doi: 10.1084/jem.20020052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Sierra JR, Cepero V, Giordano S. Molecular mechanisms of acquired resistance to tyrosine kinase targeted therapy. Mol Cancer. 2010;9:75. doi: 10.1186/1476-4598-9-75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Matino D, et al. IDO1 suppresses inhibitor development in hemophilia A treated with factor VIII. J Clin Invest. 2015;125:3766–3781. doi: 10.1172/JCI81859. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Koushik SV, Sundararaju B, McGraw RA, Phillips RS. Cloning, sequence, and expression of kynureninase from Pseudomonas fluorescens. Arch Biochem Biophys. 1997;344:301–308. doi: 10.1006/abbi.1997.0220. [DOI] [PubMed] [Google Scholar]
- 40.Miroux B, Walker JE. Over-production of proteins in Escherichia coli: mutant hosts that allow synthesis of some membrane proteins and globular proteins at high levels. J Mol Biol. 1996;260:289–298. doi: 10.1006/jmbi.1996.0399. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The data that support the findings of this study are available from the corresponding author upon reasonable request. Further details of experimental design and reagents can also be found in the Nature Research Life Sciences Reporting Summary linked to this article.