Abstract
In genetic screens aimed at understanding drug resistance mechanisms in chronic myeloid leukemia cells, inactivation of the cullin 3 adapter protein-encoding leucine zipper like transcription regulator 1 (LZTR1) gene led to enhanced mitogen-activated protein kinase (MAPK) pathway activity and reduced sensitivity to tyrosine kinase inhibitors. Knockdown of the Drosophila LZTR1 orthologue CG3711 resulted in a RAS-dependent gain-of-function phenotype. Endogenous human LZTR1 associates with the main RAS isoforms. Inactivation of LZTR1 led to decreased ubiquitination and enhanced plasma membrane localization of endogenous KRAS (V-Ki-ras2 Kirsten rat sarcoma viral oncogene homolog). We propose that LZTR1 acts as a conserved regulator of RAS ubiquitination and MAPK pathway activation. Because LZTR1 disease mutations failed to revert loss-of-function phenotypes, our findings provide a molecular rationale for LZTR1 involvement in a variety of inherited and acquired human disorders.
Chronic myeloid leukemia (CML) is characterized by the expression of the constitutively active oncogenic tyrosine kinase fusion BCR-ABL (1). The proliferation and survival of BCR-ABL+ CML cells depends on the activation state of key cellular signaling networks including the mitogen-activated protein kinase (MAPK) pathway (1). The development of the tyrosine kinase inhibitor (TKI) imatinib has provided a successful targeted therapeutic however limited by the development of resistance (2).
Genetic screens in the near-haploid human CML cell line KBM-7 allow unbiased identification of candidate genes affecting inhibitor resistance (3). We performed comprehensive haploid genetic screening with six TKIs in clinical use or under evaluation (Fig. S1A). Retroviral gene-trap mutagenized cells were exposed to the TKIs at concentrations corresponding to half maximal inhibitory concentration (IC50) to IC70 dosage (Fig. S1B), resistant cell populations were collected after selection and genomic gene-trap insertions identified by deep sequencing. Each screen resulted in enrichment of disruptive insertions in 5 to 18 different genes (Fig. 1A, Fig. S1C-G, S2A, and Table S1). We identified a recurrent set (≥ 4 screens) of six genes (NF1, WT1, PTPN1, PTPN12, LZTR1, BAP1; “TOP6” set) (Fig. 1B) with overrepresentation of disruptive genomic gene-trap integrations strongly indicating a selective advantage upon drug treatment (Fig. S2B).
Figure 1. Haploid genetic screens identify gene knockouts promoting BCR-ABL inhibitor resistance.
(A) Circos plot of the haploid genetic screen in the CML cell line KBM-7 upon treatment with ponatinib. Each dot represents a mutagenized gene identified in the resistant cell population, the dot size corresponds to the number of independent insertions identified per gene and the distance from the circos plot center indicates the significance of enrichment compared to an unselected control data set. Hits with a false discovery rate (FDR) adjusted P-value lower than 10-4 are labeled by gene name. (B) Bubble plot depicting the “TOP6” set of genes identified in four or more of the six haploid screens upon treatment with 1st, 2nd and 3rd generation BCR-ABL inhibitors. The size of each bubble corresponds to the number of independent insertions per gene and the color gradient depicts the FDR adjusted P-value of enrichment significance. (C) Multi-color competition assay (MCA)-derived fold change of cell populations after imatinib treatment of KBM-7Cas9 CML cells transduced with sgRNAs targeting the “TOP6” genes or sgRen.208 (targeting Renilla luciferase) as negative control. sgRNA-infected cell populations were mixed in a 1:1 ratio, treated with increasing drug concentrations and analyzed by flow cytometry after 14 days. Data are shown as mean value ± s.d. of at least two independent experiments (n ≥ 2) performed in duplicates. sgRNAs are labeled by gene name followed by the genomic targeting sequence position numbered according to the sequence position on the corresponding mRNA.
We used a lentiviral CRISPR/Cas9 multi-color competition assay (MCA)-based coculture system to evaluate gene-mediated drug resistance effects. SpCas9 expressing KBM-7 (KBM-7Cas9) cells were infected with lentiviral single guide RNA (sgRNA) vectors co-expressing reporter fluorophores enabling tracing of knockout and control cell populations by flow cytometry in the same well (Fig. S3A). Mixed sgRen (targeting Renilla luciferase) control cell populations expressing fluorescent proteins (GFP+ or mCherry+) did not show any preferential outgrowth of resistant cells upon 14 days of TKI treatment (Fig. 1C and Fig. S3H). In contrast, KBM-7Cas9 GFP+ cells harboring sgRNAs that target the TOP6 genes showed decreased amount of cognate protein (Fig. S3B-G) and demonstrated enhanced cell survival and outgrowth in the presence of imatinib (Fig. 1C) and rebastinib (Fig. S3H). Thus, we functionally validated the TOP 6 genes as important for drug action in BCR-ABL+ CML cells.
Although NF1, PTPN1 and PTPN12 share the ability to modulate MAPK pathway activation and WT1 as well as BAP1 function through transcriptional regulation (Fig. S2C), we could not deduce any mechanistic explanation for the role of leucine zipper like transcription regulator 1 (LZTR1) in enhanced CML cell survival from the existing literature (4, 5). To exclude cell line-specific effects, we confirmed that loss of LZTR1 expression induced resistance to imatinib and rebastinib in other CML cell lines (Fig. S4A-C). Although we identified significant LZTR1 enrichment only in four of the genetic screens, KBM-7Cas9 sgLZTR1 cells exhibited various degrees of resistance against all tested BCR-ABL TKIs (Fig. S3I). We used a CRISPR/Cas9-based domain scanning strategy to test whether both N-terminal Kelch domains and C-terminal Broad-Complex, Tramtrack, and Bric a brac (BTB) and partial BACK domains are essential for the drug resistance phenotype (6–8). All protein domain-targeting sgRNAs showed efficient indel formation (Fig. S3J) and induced resistant outgrowth of targeted cell populations exposed to rebastinib, indicating that the entire protein is functionally required (Fig. S3K). To determine whether LZTR1 exerts its function only in a CML specific context, we infected FLT3-ITD+ acute myeloid leukemia (AML) MV4-11Cas9 cells with LZTR1-targeting sgRNAs (Fig. S4D). FLT3 inhibitor treatment led to outgrowth of resistant cells underlining a more general role for LZTR1 in the drug response of hematopoietic cancers driven by different tyrosine kinases (Fig. S4E-G).
KBM-7Cas9 CML cells infected with distinct sgRNAs targeting LZTR1 displayed enhanced phosphorylation of MAPK kinase 1 (MEK1) and -2 and extracellular signal-regulated kinase 1 (ERK1) and -2, indicative of augmented MAPK pathway activation (Fig. 2A). By contrast, global tyrosine phosphorylation, as well as phosphorylation of AKT (at S473 and T308), the protein kinase S6K1, ribosomal protein S6 and the direct BCR-ABL substrate signal transducer and activator of transcription 5 (STAT5) remained unchanged (Fig. S4 J and K). Additional CML (K-562 and LAMA-84) and AML (MV4-11) cell lines had similarly enhanced MAPK pathway activation under normal growth conditions as well as, in the case of CML cells, upon increasing concentration of imatinib treatment (Fig. 2A, Fig. S4 H and I, and Fig. S5A-C). LZTR1 full-length cDNA complementation in K-562Cas9 sgLZTR1 cells reverted both enhanced MEK and ERK phosphorylation as well as TKI resistance (Fig. 2B and C). Treatment of K-562Cas9 sgLZTR1 cells with the clinically approved inhibitor of MEK1 and -2 trametinib reverted enhanced activation of ERK1 and -2 and pharmacologically counteracted the drug resistance phenotype (Fig. 2D and E). Furthermore, cDNA expression of constitutively active Mek1D218, D222 (Mek1 DD) in K-562rtTA3 cells led to enhanced phosphorylation of ERK1 and -2 and reduced imatinib sensitivity (Fig. S6A and B). Activation of the MAPK pathway could also be inferred by the gene expression and transcription factor enrichment signature obtained with RNA sequencing experiments in KBM-7Cas9 sgLZTR1 compared to sgRen cells (Fig. S5D and E and Table S2). Altogether, the data established a causal role for enhanced MAPK pathway activation in the resistance of CML cells towards TKI therapy as elicited by loss of LZTR1 function.
Figure 2. Loss of LZTR1 enhances MAPK pathway activation.
(A) Phosphorylation of MEK and ERK in KBM-7Cas9 and K-562Cas9 CML cells transduced with the indicated sgRNAs. (B) Immunoblot analysis of MEK and ERK phosphorylation as well as LZTR1 expression in sgRen.208-expressing K-562Cas9 CML cells transduced with empty vector, and sgLZTR1.466-expressing cells transduced with empty vector or LZTR1-cDNA-containing MSCV retrovirus. Quantification of MEK and ERK phosphorylation is shown next to the corresponding immunoblots. (C) Competitive proliferation assay (MCA) of K-562Cas9 sgRen.208 cells transduced with empty vector and sgLZTR1.466 cells transduced with empty vector or LZTR1 cDNA after treatment with increasing concentrations of imatinib for 14 days. (D) Phosphorylation of MEK and ERK in K-562Cas9 cells expressing sgLZTR1.466 and treated with increasing concentrations of trametinib for 3 hours. (E) Changes in cell populations measured by MCA of K-562Cas9 CML cells expressing sgRen.208 or sgLZTR1.466 after 14 days of treatment with increasing concentrations of imatinib alone or in combination with trametinib. Immunoblot results in (A, B and D) are representative of at least two independent biological experiments (n ≥ 2). MCA data in (C, E) are shown as mean value ± s.d. of at least two independent experiments (n ≥ 2). DMSO treatment served as negative control.
As BTB domain-containing proteins serve as adaptor proteins for the cullin 3 (CUL3) E3 ubiquitin ligase complex enabling specific substrate recognition and ubiquitination (9), we tested whether loss of CUL3 expression could mimic the observed LZTR1 loss-of-function phenotype. Indeed, K-562Cas9 sgCUL3 cells demonstrated enhanced MAPK pathway activation and, in contrast to sgLZTR1 cells, increased phosphorylation of AKT (Fig. S4K). However, sgRNAs targeting CUL3 had a pronounced antiproliferative effect both in KBM-7 Cas9 and K-562Cas9 cells (Fig. S6C and D) therefore providing a potential explanation why CUL3 was not detected in our genetic screens (Fig. S2A).
In contrast to the CML cell lines, depletion of LZTR1 with CRISPR sgRNAs in HeLa, human embryonic kidney (HEK)-293T or HAP1 cells did not increase MAPK pathway activation under comparable culture conditions (Fig. S7A and C). However, after serum stimulation of cells cultured without serum, HEK293TCas9 sgLZTR1 cells showed a more pronounced activation of MEK and ERK than in control cells (Fig. S7B). Similarly, in HAP1 cells, a non-hematopoietic derivative of the KBM-7 cell line (10), phorbol-12-myristat-13-acetat (PMA) treatment led to enhanced MAPK pathway activation in the absence of LZTR1 compared with that in wild-type (WT) cells (Fig. S7C). Whereas HAP1Cas9 sgCUL3 cells exhibited increased phosphorylation of ERK1, ERK2 and AKT (Fig. S7F), loss of LZTR1 expression in HAP1Cas9 altered only the MAPK pathway as identified by means of pathway array-based assessment of kinase activation (Fig. S7D-E and Table S3).
Genetic studies have identified LZTR1 mutations in glioblastoma (GBM) (11), schwannomatosis (SWNMT) (8) and Noonan syndrome (NS) (5), a developmental syndrome which is part of the larger group of RASopathies characterized by mutations in components of the RAS-MAPK pathway (12). Identification of NF1 and LZTR1 loss-of-function-induced MAPK pathway activation in our haploid resistance screens combined with human LZTR1 mutations in NS indicated that LZTR1 might directly regulate guanosine triphosphatases (GTPases) of the RAS family.
Drosophila wing vein formation and eye development serve as excellent in vivo readouts for RAS signaling (13–15). CG3711 encodes the Drosophila orthologue of mammalian LZTR1, which contains a unique N-terminal domain (amino acids 1-184) that is only found in Drosophila. This is followed by the highly conserved remaining part of the protein (54% sequence identity) (Fig. S8A-C). Systemic depletion of CG3711 with RNA interference using act5C-Gal4 yielded viable flies; however, the majority of wings of these flies displayed wing vein defects characterized by extra veins and vein tissue (Fig. 3A and B and Fig. S8D). This phenotype closely resembles a gain-of-function increase of RAS-MAPK signaling (15) and could be rescued by a decrease in abundance of RAS via dRas (Drosophila Ras) heterozygosity (Fig. 3C). Drosophila R7 photoreceptor induction requires RAS function (13, 14). We used a mild dominant negative version, RasV12 C40, which although locked in the active 5′-triphosphate (GTP)-bound state does not activate MAPK signaling (16). RasV12 C40 expression in the developing eyes (via the sevenless/sev-Gal4 expression system) led to a frequent loss of the R7 photoreceptor (∼30% of ommatidia display R7 loss, and some also lost other R-cells) (Fig. S8E and H). Because RasV12 C40 is constitutively active, it also causes defects in ommatidial rotation (16, 17), serving as internal control. When both sev-Gal4 driven CG3711 RNAi and RasV12 C40 were expressed, the loss of R7 phenotype was almost completely suppressed (Fig. S8F and H). The ommatidial rotation defects were enhanced rather than suppressed because these involve MAPK-independent RAS signaling (16) (Fig. S8F). Expression of CG3711 RNAi in the eye alone did not induce phenotypic changes (Fig. S8G and H). Thus, RAS is crucial for the phenotypes induced by depletion of CG3711.
Figure 3. LZTR1 modulates MAPK pathway activation through RAS regulation.
(A) Morphology of adult wings from act5C-Gal4, UAS-wIR (act>wIR for short) and act>CG3711IR #1 RNAi fly lines. (B) Quantification of act>wIR and act>CG3711IR RNAi lines as percentage of wings with ectopic wing vein formation. P-value for RNAi #1 and #3 in the wing is <0.0001 (****) and for #2 is 0.0255. (C) Quantification of act>wIR and act>CG3711IR RNAi lines alone or in a Rase2F/+ background as percentage of wings with ectopic wing vein formation. For statistical assessment, partial extra vein and extra vein formation have been combined. P-value for both RNAi line comparisons in the wing is <0.0001 (****). (D) FLAG immunoprecipitates (IP) and whole cell extracts (WCE) from K-562rtTA3 cells expressing FLAG-BirA* tagged GFP or KRAS4A WT, ΔHVR, C180S or C186S after 48 hours of doxycycline treatment were immunoblotted for the indicated proteins. (E) Confocal microscopy of HAP1 WT cells and HAP1 cells with endogenously FLAG-tagged KRAS transduced with sgRen.208, sgLZTR1.620, sgLZTR1.466, sgCUL3.852 or sgCUL3.1396 and stained with anti-FLAG. Scale bar in all panels is 10μm. (G) Tandem ubiquitin binding domain (TUBE)-based purifications of ubiquitinated proteins and whole cell extracts (WCE) from HAP1 WT and endogenously FLAG-tagged KRAS cells transduced with sgRen.208 or sgLZTR1.466 were analyzed by immunoblotting with the indicated antibodies. WT, wild type.
To explore possible interactions of LZTR1 with RAS, we used a BirA* fusion protein-based proximity biotinylation-dependent (BioID) proteomic approach, which allows identification of weak interaction partners. We expressed KRAS4A, KRAS4B, NRAS and HRAS as N-terminal FLAG-BirA* fusion proteins in K-562rtTA3 CML cells. This led to enhanced MEK and ERK activation demonstrating that the BirA* tag does not generally interfere with RAS signaling (Fig. S9A). We identified between 153 and 258 proteins in proximity of or interacting with human RAS proteins, among them several known interactors and pathway components (Fig. S9B-F and Table S4). A common set of 123 proteins repeatedly purified with all four RAS proteins, with gene ontology analysis showing a significant enrichment in components associated with plasma membrane and Golgi apparatus as well as the MAPK pathway (Fig. S9G-H). We identified LZTR1 among the most significant interactors of all four RAS proteins (Fig. S9B-F). Intersection of the 123 common interactors identified by BioID with a fluorescence-activated cell sorting (FACS)-based haploid genetic pathway screen for the identification of gene knockout alleles influencing phosphorylation of ERK1 and -2 (18) revealed LZTR1 as the only common RAS interactor in our proteomic data with a negative regulatory function within the RAS-MAPK pathway (Fig. S10A).
BioID proximity biotinylation-based and FLAG tag-based co-immunoprecipitation experiments confirmed the interaction of all four RAS proteins with endogenous LZTR1 (Fig. S10B and C). Only the four main RAS isoforms interacted specifically with LZTR1, whereas six related RAS family GTPases showed no interaction (Fig. S11A). Nine other canonical positive RAS-MAPK pathway regulators also failed to interact with endogenous LZTR1 (Fig. S11B). Immunoprecipitations with extracts derived from LZTR1 knockout cells confirmed the specificity of the interaction (Fig. S11C). The C-terminal post-translational acylations of KRAS proteins appeared to be required for the interaction with LZTR1 (Fig. 3D and Fig. S11D). Thus, proper membrane associated localization mediated by this region may be required for specific interaction with LZTR1.
Stably expressed full-length hemagglutinin (HA)-tagged LZTR1 protein showed a speckled and vesicular staining pattern in HeLa, HEK293T and HAP1 cells whereas all domain mutants had a homogenous cytoplasmic distribution (Fig. S12A-D). Full-length LZTR1 displayed an overlapping localization within speckled and vesicular compartments with transiently transfected V5-tagged CUL3 in HeLa cells (Fig. S13A). The LZTR1-stained speckles further overlapped with the autophagosome marker LC3B fused to mCherry (Fig. S13B), but we failed to detect LZTR1 localization with marker proteins of the Golgi, lysosome, peroxisome or early and late endosome compartments (Fig. S13C). Isoform-specific antibodies that recognize endogenous human RAS isoforms are not available (19). We therefore endogenously tagged KRAS (Fig. S14A), and confirmed specificity of detection in immunofluorescence and immunoblotting by genetic inactivation of the tagged genomic allele (Fig. S14 B to D). KRAS localized to a large number of small-punctate structures, likely membrane-containing small vesicles (Fig. 3E and Fig. S14D). Inactivation of endogenous LZTR1 led to an increased RAS signal, particularly at the periphery of cells, at regions of cell-to-cell contacts (Fig. 3E and Fig. S14D). This phenotype appeared to be dependent on the CUL3 E3 ligase complex, because genetic inactivation of CUL3 led to similar phenotypic changes (Fig. 3E and Fig. S14E).
We also assessed interaction of endogenously tagged KRAS with exogenous LZTR1 in HAP1 cells (Fig. S15A). Exogenous LZTR1 localized to fewer, larger, and more oblong structures than endogenous KRAS (Fig. S15B). A fraction of the LZTR1-stained structures also contained KRAS. Treatment with the cullin neddylation inhibitor MLN4924 caused clustering of LZTR1-containing structures that appeared to surround particles strongly stained with KRAS (Fig. S15B, MLN4924 panel). Caution is required for the interpretation of these patterns because LZTR1 was overexpressed, and KRAS was at endogenous concentration. However, the images could represent a trapped proteostatic process involving the two proteins.
We tested whether LZTR1 affected ubiquitination of RAS proteins by acting as a substrate adaptor for the CUL3 E3 ligase complex (11, 20). Ubiquitination of RAS proteins is known, but little is known of its consequences (21–25). Co-expression of HA-ubiquitin with either one of the four RAS isoforms alone only resulted in a basal state of ubiquitination, but addition of MYC-tagged CUL3 and V5-tagged LZTR1 led to increased ubiquitination of RAS proteins (Fig. S16A-D). LZTR1-deletion mutants that lack either one or both C-terminal BTB domains or the N-terminal Kelch domain failed to induce an equivalent degree of ubiquitination of KRAS4A (Fig. S16A). KRAS4A required the presence of its C-terminal hypervariable (HVR) and the farnesylated cysteine 186 for LZTR1-mediated ubiquitination (Fig. S16E). LZTR1-induced ubiquitination was unchanged in cells treated with a proteasome inhibitor but could be blocked by cullin neddylation inhibition, which is consistent with the observed subcellular localization changes (Fig. S16F). Specificity for LZTR1 in the process was further underscored by the failure of two prominent CUL3 adaptors, Kelch-like ECH-associated protein 1 (KEAP1) and Speckle-type POZ protein (SPOP), to cause a comparable ubiquitination (Fig. S16G). We used HAP1 cells bearing endogenously tagged KRAS and tandem ubiquitin binding domain (TUBE) reagents, to capture endogenously ubiquitinated proteins and evaluate the contribution of endogenous LZTR1 (26). In the absence of any stimulation, immunoprecipitation with TUBE purified several proteins that reacted with pan-ubiquitin antibodies (Fig. 3F, third panel from the top, and Fig. S16H). The same precipitates contained proteins reacting with pan-RAS antibodies (Fig. 3F, second panel from the top). Last, FLAG antibodies detected a protein not visible in the whole cell extract (Fig. 3F, first panel from the top, and Fig. S16H). TUBE-mediated immunoprecipitation from corresponding cells in which LZTR1 had been genetically inactivated did not contain amounts of RAS detectable with these antibodies (Fig. 3F, first and second panel from the top, and Fig. S16H). The signal obtained with pan-ubiquitin antibodies was unchanged (Fig. 3F, third panel from the top, and Fig. S16H). Thus, human LZTR1 appears to mediate ubiquitination of endogenous human KRAS and possibly other RAS proteins.
If a main function of LZTR1 is to regulate RAS, then loss of RAS function should compensate for loss of LZTR1 activity as suggested in the fruit fly epistatic analysis, thereby representing an ideal experimental setup to assess the disease-associated LZTR1 mutations by their dependency on RAS activity. We confirmed that loss of LZTR1 function enhanced RAS activity in K-562 cells (Fig. S17A). We then performed CRISPR/Cas9-based double knockout MCA experiments (Fig. S17B-D). Single sgLZTR1 and sgLZTR1-sgRen double-infected cells were resistant to imatinib compared to control sgRen cells (Fig. S17E-F). sgRNAs targeting KRAS abolished cellular outgrowth, whereas sgRNAs targeting NRAS, HRAS and RIT1 failed to do so (Fig. S17E-F). sgLZTR1-sgKRAS cells had reduced MEK phosphorylation comparable to that of sgRen cells, whereas sgNRAS and sgHRAS cells maintained enhanced MAPK pathway activation (Fig. S17G). KRAS inactivation led to a strong antiproliferative phenotype in KBM-7Cas9 cells and, to a lesser extent, K-562Cas9 cells (Fig. S18 A and B), indicating that the selective requirement of KRAS in mediating LZTR1-induced enhanced MAPK pathway activation might represent a prerequisite of KRAS for BCR-ABL signaling in CML cells. We additionally used a fibronectin-fold-based monobody, NS1, which bears low nanomolar affinity for the dimerization interface of both KRAS and HRAS, and is able to interfere with their signaling activities (27). Cells stably expressing the NS1 monobody did not show increased activation of the MAPK pathway or drug resistance associated with loss of LZTR1 function (Fig. S18 C and D).
More than 50 different mutations have been mapped to the human LZTR1 gene in various diseases (5, 28) (8, 11, 29, 30) (Fig. S19A-B and Table S5). To establish a mechanistic link between mutations that affect human diseases and the biochemical processes described here, we focused on LZTR1 missense mutations identified in GBM (11), NS (5) and SWNMT (8) (Fig. S20B). Mutation-bearing LZTR1 cDNAs were tested for their ability to complement K-562 CML cells deficient in endogenous LZTR1 (Fig. S20A). In contrast to WT LZTR1, the disease-associated mutations failed to reduce MAPK pathway activation, despite being expressed at comparable or higher amount (Fig. 4A-C). Similarly, all missense mutations apart from the GBM-associated T288I and R810W failed to restore sensitivity to imatinib treatment (Fig. S20C-E). LZTR1 mutations within the Kelch domain partially retained their localization pattern in comparison to the WT protein, whereas mutations in the BTB/BACK domains mislocalized to the cytoplasm (Fig. S21 A and B). In agreement, expression of LZTR1 Kelch domain mutations identified in GBM, NS and SWNMT in K-562 WT cells had a resistance-mediating effect in MCA assays in cells treated with imatinib, whereas BTB/BACK domain mutations did not (Fig. S22A-C). Furthermore, in contrast to the LZTR1 WT protein, the two mutants R198G and G248R identified in GBM and NS failed to induce a similar increase in ubiquitination on KRAS4A (Fig. S22D). These findings provide functional evidence that human LZTR1 missense mutations identified in GBM, NS and SWNMT represent loss-of-function towards ubiquitination and inhibition of RAS activity.
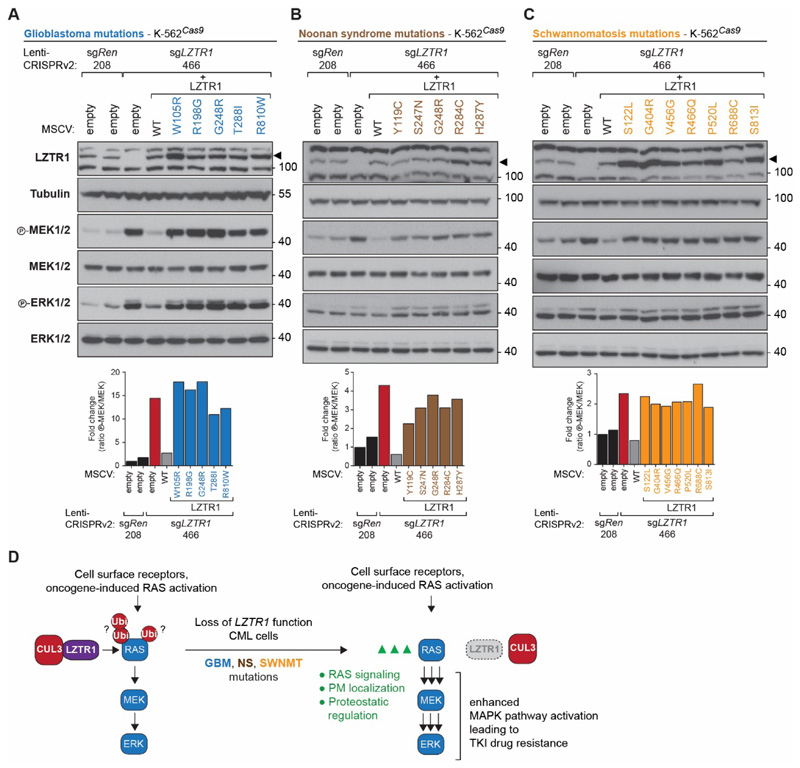
Figure 4. LZTR1 disease missense mutations fail to rescue the loss-of-function phenotype.
(A-C) Immunoblotting for MEK and ERK phosphorylation as well as LZTR1 expression of K-562Cas9 sgRen.208-expressing cells retrovirally transduced with empty vector, and sgLZTR1.466-expressing cells transduced with either empty vector, LZTR1 WT, or LZTR1 mutations identified in GBM (blue) (A), NS (brown) (B), or SWNMT (orange) (C). The LZTR1 G248R mutation has been identified in both GBM and NS. Immunoblot results are representative of at least two independent biological experiments (n ≥ 2) and quantification of MEK phosphorylation in the displayed blots is shown. (D) Mechanistic model of CUL3-LZTR1-mediated RAS ubiquitination and enhanced MAPK pathway activation and BCR-ABL inhibitor drug resistance induced by loss of LZTR1 function.
Together with the accompanying manuscript on the role of LZTR1 in diseases driven by the dysregulation of RAS ubiquitination and signaling (Steklov, Pandolfi, Baietti et al. SCIENCE 2018) our work illustrates the importance of an additional, underappreciated layer of RAS regulation (Fig. 4D).
Supplementary Material
One Sentence Summary.
Genetic screens for BCR-ABL TKI resistance identify LZTR1 as mediator of RAS ubiquitination and regulator of MAPK pathway activation.
Acknowledgments
We are grateful to all members of the Superti-Furga laboratory for help, discussions and advice, E. Girardi and E. Salzer for critically reading the manuscript, and P. Májek for feedback on the bioinformatic analysis of BioID proteomic experiments. Supported by the Austrian Academy of Sciences, ERC grants (i-FIVE 250179 and Game of Gates 695214) and Austrian Science Fund grant (FWF SFB F4711) to G.S.-F, EMBO long-term fellowship to M.R. (ALTF 1346-2011) and G.I.V. (ALTF 1543-2012), Austrian Science Fund grant (FWF SFB F4702) to R.K., NIH grants R01 EY013256 and GM102811 to M.M, and funding from the Cancer Genomics Center (CGC.nl), KWF grant NKI 2015-7609 and the European Research Council (ERC) Starting Grant (ERC-2012-StG 309634) to T.R.B.. We thank the Biomedical Sequencing Facility for advice on Illumina sequencing and the Core Facility Imaging of the Medical University of Vienna for technical support. T.R.B. is cofounder and SAB member of Haplogen GmbH and cofounder and managing director of Scenic Biotech. G.I.V. is cofounder and employee of Allcyte GmbH. G.S.-F. is cofounder and shareholder of Allcyte GmbH and Haplogen GmbH. Deep sequencing datasets have been deposited in the NCBI Sequence Read Archive under the accession number SRP157890 and BioID mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository under the accession number PXD011280. The KBM-7 cell line is available from T.R.B. under a materials transfer agreement with the Netherlands Cancer Institute.
Footnotes
Author contributions:
J.W.B. and G.S-F. conceived the study based on discussions with T.R.B.. J.W.B., F.K., and M.P. performed research, G.M.C. and M.M. designed and performed Drosophila experiments. L.X.H., G.I.V., A.F., M.R. and G.W. generated reagents and provided scientific insight. F. S. analyzed haploid genetic screening data, created circos plots and the graphical display of insertion sites. V.S. performed RNA sequencing and enrichment analysis and compiled cBioPortal mutation data. K.P. and A.C.M. performed proteomic sample analysis. V.A.B. and T.R.B. provided reagents and gave experimental advice. R.K. supervised sequencing data analysis and gave experimental advice. J.W.B., G.M.C., F.K., G.I.V., L.X.H., F.S., M.M. and G.S-F. analyzed and interpreted the data. J.W.B., M.M. and G.S-F. wrote the paper.
Competing financial interests:
The authors declare no competing financial interest.
References
- 1.Ren R. Mechanisms of BCR–ABL in the pathogenesis of chronic myelogenous leukaemia. Nat Rev Cancer. 2005;5:172–183. doi: 10.1038/nrc1567. [DOI] [PubMed] [Google Scholar]
- 2.O'Hare T, Zabriskie MS, Eiring AM, Deininger MW. Pushing the limits of targeted therapy in chronic myeloid leukaemia. Nat Rev Cancer. 2012;12:513–526. doi: 10.1038/nrc3317. [DOI] [PubMed] [Google Scholar]
- 3.Carette JE, et al. Global gene disruption in human cells to assign genes to phenotypes by deep sequencing. Nature Biotechnology. 2011;29:1–7. doi: 10.1038/nbt.1857. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Simanshu DK, Nissley DV, McCormick F. RAS Proteins and Their Regulators in Human Disease. Cell. 2017;170:17–33. doi: 10.1016/j.cell.2017.06.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Yamamoto GL, et al. Rare variants in SOS2 and LZTR1 are associated with Noonan syndrome. J Med Genet. 2015;52:413–421. doi: 10.1136/jmedgenet-2015-103018. [DOI] [PubMed] [Google Scholar]
- 6.Canning P, et al. Structural basis for Cul3 protein assembly with the BTB-Kelch family of E3 ubiquitin ligases. J Biol Chem. 2013;288:7803–7814. doi: 10.1074/jbc.M112.437996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Nacak TG, Leptien K, Fellner D, Augustin HG, Kroll J. The BTB-kelch protein LZTR-1 is a novel Golgi protein that is degraded upon induction of apoptosis. J Biol Chem. 2006;281:5065–5071. doi: 10.1074/jbc.M509073200. [DOI] [PubMed] [Google Scholar]
- 8.Piotrowski A, et al. Germline loss-of-function mutations in LZTR1 predispose to an inherited disorder of multiple schwannomas. Nat Genet. 2014;46:182–187. doi: 10.1038/ng.2855. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Xu L, et al. BTB proteins are substrate-specific adaptors in an SCF-like modular ubiquitin ligase containing CUL-3. Nature. 2003;425:316–321. doi: 10.1038/nature01985. [DOI] [PubMed] [Google Scholar]
- 10.Carette JE, et al. Ebola virus entry requires the cholesterol transporter Niemann-Pick C1. Nature. 2011;477:340–343. doi: 10.1038/nature10348. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Frattini V, et al. The integrated landscape of driver genomic alterations in glioblastoma. Nat Genet. 2013;45:1141–1149. doi: 10.1038/ng.2734. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Tidyman WE, Rauen KA. The RASopathies: developmental syndromes of Ras/MAPK pathway dysregulation. Curr Opin Genet Dev. 2009;19:230–236. doi: 10.1016/j.gde.2009.04.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Therrien M, et al. KSR, a novel protein kinase required for RAS signal transduction. Cell. 1995;83:879–888. doi: 10.1016/0092-8674(95)90204-x. [DOI] [PubMed] [Google Scholar]
- 14.Karim FD, et al. A screen for genes that function downstream of Ras1 during Drosophila eye development. Genetics. 1996;143:315–329. doi: 10.1093/genetics/143.1.315. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Oishi K, et al. Phosphatase-defective LEOPARD syndrome mutations in PTPN11 gene have gain-of-function effects during Drosophila development. Hum Mol Genet. 2009;18:193–201. doi: 10.1093/hmg/ddn336. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Gaengel K, Mlodzik M. Egfr signaling regulates ommatidial rotation and cell motility in the Drosophila eye via MAPK/Pnt signaling and the Ras effector Canoe/AF6. Development. 2003;130:5413–5423. doi: 10.1242/dev.00759. [DOI] [PubMed] [Google Scholar]
- 17.Brown KE, Freeman M. Egfr signalling defines a protective function for ommatidial orientation in the Drosophila eye. Development. 2003;130:5401–5412. doi: 10.1242/dev.00773. [DOI] [PubMed] [Google Scholar]
- 18.Brockmann M, et al. Genetic wiring maps of single-cell protein states reveal an off-switch for GPCR signalling. Nature. 2017;546:307–311. doi: 10.1038/nature22376. [DOI] [PubMed] [Google Scholar]
- 19.Waters AM, et al. Evaluation of the selectivity and sensitivity of isoform- and mutation-specific RAS antibodies. Sci Signal. 2017;10:eaao3332. doi: 10.1126/scisignal.aao3332. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Bennett EJ, Rush J, Gygi SP, Harper JW. Dynamics of cullin-RING ubiquitin ligase network revealed by systematic quantitative proteomics. Cell. 2010;143:951–965. doi: 10.1016/j.cell.2010.11.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Jura N, Scotto-Lavino E, Sobczyk A, Bar-Sagi D. Differential modification of Ras proteins by ubiquitination. Molecular Cell. 2006;21:679–687. doi: 10.1016/j.molcel.2006.02.011. [DOI] [PubMed] [Google Scholar]
- 22.Yan H, Jahanshahi M, Horvath EA, Liu H-Y, Pfleger CM. Rabex-5 ubiquitin ligase activity restricts Ras signaling to establish pathway homeostasis in Drosophila. Curr Biol. 2010;20:1378–1382. doi: 10.1016/j.cub.2010.06.058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Sasaki AT, et al. Ubiquitination of K-Ras enhances activation and facilitates binding to select downstream effectors. Sci Signal. 2011;4:ra13. doi: 10.1126/scisignal.2001518. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Baker R, et al. Differences in the regulation of K-Ras and H-Ras isoforms by monoubiquitination. J Biol Chem. 2013;288:36856–36862. doi: 10.1074/jbc.C113.525691. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Rose CM, et al. Highly Multiplexed Quantitative Mass Spectrometry Analysis of Ubiquitylomes. Cell Syst. 2016;3:395–403.e4. doi: 10.1016/j.cels.2016.08.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Hjerpe R, et al. Efficient protection and isolation of ubiquitylated proteins using tandem ubiquitin-binding entities. EMBO Rep. 2009;10:1250–1258. doi: 10.1038/embor.2009.192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Spencer-Smith R, et al. Inhibition of RAS function through targeting an allosteric regulatory site. Nat Chem Biol. 2017;13:62–68. doi: 10.1038/nchembio.2231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Johnston JJ, et al. Autosomal recessive Noonan syndrome associated with biallelic LZTR1 variants. Genet Med. 2018 doi: 10.1038/gim.2017.249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Gröbner SN, et al. The landscape of genomic alterations across childhood cancers. Nature. 2018;555:321–327. doi: 10.1038/nature25480. [DOI] [PubMed] [Google Scholar]
- 30.Cancer Genome Atlas Research Network. Electronic address: wheeler@bcm.edu, Cancer Genome Atlas Research Network. Saksena G. Comprehensive and Integrative Genomic Characterization of Hepatocellular Carcinoma. Cell. 2017;169:1327–1341.e23. doi: 10.1016/j.cell.2017.05.046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Sanjana NE, Shalem O, Zhang F. Improved vectors and genome-wide libraries for CRISPR screening. Nat Methods. 2014;11:783–784. doi: 10.1038/nmeth.3047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Shalem O, et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science. 2014;343:84–87. doi: 10.1126/science.1247005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Montague TG, Cruz JM, Gagnon JA, Church GM, Valen E. CHOPCHOP: a CRISPR/Cas9 and TALEN web tool for genome editing. Nucleic Acids Research. 2014;42:W401–7. doi: 10.1093/nar/gku410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Doench JG, et al. Rational design of highly active sgRNAs for CRISPR-Cas9-mediated gene inactivation. Nature Biotechnology. 2014;32:1262–1267. doi: 10.1038/nbt.3026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Lackner DH, et al. A generic strategy for CRISPR-Cas9-mediated gene tagging. Nat Commun. 2015;6:10237. doi: 10.1038/ncomms10237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Lampson BL, et al. Rare codons regulate KRas oncogenesis. Curr Biol. 2013;23:70–75. doi: 10.1016/j.cub.2012.11.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Bigenzahn JW, et al. An Inducible Retroviral Expression System for Tandem Affinity Purification Mass-Spectrometry-Based Proteomics Identifies Mixed Lineage Kinase Domain-like Protein (MLKL) as an Heat Shock Protein 90 (HSP90) Client. Mol Cell Proteomics. 2016;15:1139–1150. doi: 10.1074/mcp.O115.055350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Dull T, et al. A third-generation lentivirus vector with a conditional packaging system. J Virol. 1998;72:8463–8471. doi: 10.1128/jvi.72.11.8463-8471.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Schambach A, et al. Lentiviral vectors pseudotyped with murine ecotropic envelope: increased biosafety and convenience in preclinical research. Exp Hematol. 2006;34:588–592. doi: 10.1016/j.exphem.2006.02.005. [DOI] [PubMed] [Google Scholar]
- 40.Fellmann C, et al. An optimized microRNA backbone for effective single-copy RNAi. Cell Rep. 2013;5:1704–1713. doi: 10.1016/j.celrep.2013.11.020. [DOI] [PubMed] [Google Scholar]
- 41.Couzens AL, et al. Protein interaction network of the mammalian Hippo pathway reveals mechanisms of kinase-phosphatase interactions. Sci Signal. 2013;6:rs15. doi: 10.1126/scisignal.2004712. [DOI] [PubMed] [Google Scholar]
- 42.Sakuma T, Nakade S, Sakane Y, Suzuki K-IT, Yamamoto T. MMEJ-assisted gene knock-in using TALENs and CRISPR-Cas9 with the PITCh systems. Nat Protoc. 2016;11:118–133. doi: 10.1038/nprot.2015.140. [DOI] [PubMed] [Google Scholar]
- 43.Jae LT, et al. Virus entry. Lassa virus entry requires a trigger-induced receptor switch. Science. 2014;344:1506–1510. doi: 10.1126/science.1252480. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Krzywinski M, et al. Circos: an information aesthetic for comparative genomics. Genome Res. 2009;19:1639–1645. doi: 10.1101/gr.092759.109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Brinkman EK, Chen T, Amendola M, van Steensel B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Research. 2014;42:e168. doi: 10.1093/nar/gku936. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Schindelin J, et al. Fiji: an open-source platform for biological-image analysis. Nat Methods. 2012;9:676–682. doi: 10.1038/nmeth.2019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Rappsilber J, Ishihama Y, Mann M. Stop and go extraction tips for matrix-assisted laser desorption/ionization, nanoelectrospray, and LC/MS sample pretreatment in proteomics. Anal Chem. 2003;75:663–670. doi: 10.1021/ac026117i. [DOI] [PubMed] [Google Scholar]
- 48.Olsen JV, et al. Parts per million mass accuracy on an Orbitrap mass spectrometer via lock mass injection into a C-trap. Mol Cell Proteomics. 2005;4:2010–2021. doi: 10.1074/mcp.T500030-MCP200. [DOI] [PubMed] [Google Scholar]
- 49.Kersey P, Hermjakob H, Apweiler R. VARSPLIC: alternatively-spliced protein sequences derived from SWISS-PROT and TrEMBL. Bioinformatics. 2000;16:1048–1049. doi: 10.1093/bioinformatics/16.11.1048. [DOI] [PubMed] [Google Scholar]
- 50.Colinge J, Masselot A, Giron M, Dessingy T, Magnin J. OLAV: towards high-throughput tandem mass spectrometry data identification. Proteomics. 2003;3:1454–1463. doi: 10.1002/pmic.200300485. [DOI] [PubMed] [Google Scholar]
- 51.Teo G, et al. SAINTexpress: improvements and additional features in Significance Analysis of INTeractome software. J Proteomics. 2014;100:37–43. doi: 10.1016/j.jprot.2013.10.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Mellacheruvu D, et al. The CRAPome: a contaminant repository for affinity purification-mass spectrometry data. Nat Methods. 2013;10:730–736. doi: 10.1038/nmeth.2557. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.R Core Team. R: A Language and Environment for Statistical Computing. 2016 (available at https://www.R-project.org)
- 54.Heberle H, Meirelles GV, da Silva FR, Telles GP, Minghim R. InteractiVenn: a web-based tool for the analysis of sets through Venn diagrams. BMC Bioinformatics. 2015;16:169. doi: 10.1186/s12859-015-0611-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Huang DW, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4:44–57. doi: 10.1038/nprot.2008.211. [DOI] [PubMed] [Google Scholar]
- 56.Tomlinson A, Bowtell DD, Hafen E, Rubin GM. Localization of the sevenless protein, a putative receptor for positional information, in the eye imaginal disc of Drosophila. Cell. 1987;51:143–150. doi: 10.1016/0092-8674(87)90019-5. [DOI] [PubMed] [Google Scholar]
- 57.Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550. doi: 10.1186/s13059-014-0550-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Marbach D, et al. Tissue-specific regulatory circuits reveal variable modular perturbations across complex diseases. Nat Methods. 2016;13:366–370. doi: 10.1038/nmeth.3799. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.