Abstract
The placenta is the extraembryonic organ that supports the fetus during intrauterine life. Although placental dysfunction results in major disorders of pregnancy with immediate and lifelong consequences for both mother and child, our knowledge of the human placenta is limited due to lack of functional experimental models1. Upon implantation, the trophectoderm of the blastocyst rapidly proliferates and generates the trophoblast, the unique cell type of the placenta. In vivo, the proliferative villous cytotrophoblast cells (VCT) differentiate into two main sub-populations: syncytiotrophoblast (SCT), the multinucleated epithelium of the villi responsible for nutrient exchange and hormone production, and extravillous trophoblast (EVT) that anchor the placenta to the maternal decidua and transform the maternal spiral arteries2. Here, we describe the generation of long-term, genetically-stable organoid cultures of trophoblast cells that can differentiate to SCT and EVT. We used HLA-typing to confirm the organoids are fetally-derived, and verified their identity against four trophoblast-specific criteria3. The cultures organise into villous-like structures, and we detected secretion of placental-specific peptides and hormones, including hCG, GDF15, PSGs and PAPPA, by mass spectrometry. The organoids also differentiate to HLA-G+ EVT that vigorously invade in 3D. Analysis of the methylome reveals the organoids closely resemble normal first-trimester placentas. This organoid model will be transformative for studying human placental development and for investigating trophoblast interactions with the local and systemic maternal environment.
To devise an organoid culture system suitable for trophoblast, we focussed on maternal and placental products that might signal to the stem/progenitor cells that reside in areas of Ki67+ VCT proliferation and/or at the base of the cytotrophoblast cell columns (CCC) that give rise to EVT (Fig.1a,b)4–9. We investigated signalling pathways between 6-8 weeks g.a when proliferation is high: WNT through β-catenin; TGFβ through SMAD2/3; MAPK through Erk1/2 and STAT3. Our findings led to empirical trials of agonists/antagonists, along with other agents, which resulted in a basal Trophoblast Organoid Medium (TOM) composed of EGF, FGF2, CHIR99021 (a WNT activator), A83-01 (a TGFβ/SMAD inhibitor) and Rspondin-1 (Extended Data Fig. 1a-d). To prepare isolates of trophoblast cells, first trimester placentas (6-9 weeks g.a) were enzymatically digested to enrich for cell clusters containing EPCAM+ cells; EPCAM marks proliferative trophoblast (Fig. 1c)10. Cell clusters were seeded into Matrigel drops and grown in TOM. Although some growth was seen, we also tested factors used in other organoid systems and/or present in the first trimester microenvironment (Extended Data Fig. 2a, Supplementary Table 1a). HGF, PGE2 and Y-27632 (a ROCK inhibitor) increase cell viability and growth and, when they are combined with TOM, there is rapid growth of cells within a week (Fig. 1d, Supplementary Table 1b). After the first passage, organoid structures appear and homogeneous trophoblast organoids are established within 2 passages (10-14 days) with an efficiency of 91% (20/22 patient samples). To confirm their fetal origin, we used microsatellite analysis and HLA typing (Extended Data Table 1). Derivation of the organoids in TOM in the absence of each factor reveals that EGF is the most important, with effects also seen with Y-27632, A83-01 and CHIR99021. We have now derived trophoblast organoids that are genetically stable after many passages (Extended Data Fig. 2b,c); three randomly selected cultures are still growing after a year (Supplementary Table 1c), are healthy and show active mitochondrial function (Extended Data Fig. 2d). Maternal epithelial cells are always detectable by flow cytometry in placental cell isolates. Nicotinamide enriches for cystic structures resembling glandular organoids at early stages of derivation (Extended Data Fig. 2a), and microsatellite analysis and HLA typing confirmed their maternal origin (Extended Data Table 2). By selecting the appropriate media, we can derive both decidual glandular and trophoblast organoid cultures from the same pregnancy (Extended Data Fig. 2e)11. This highlights the importance of verifying the maternal or fetal origin of any cultures derived from decidual or placental cell isolates.
Figure 1. Establishment of long-term organoid cultures of trophoblast from human placentas.
a, A placental villus at the maternal-fetal interface in the first trimester of pregnancy showing the different trophoblast subsets: SCT, VCT, CCC and EVT. Sources of the intrinsic and extrinsic signals that could signal to proliferative Ki67+ trophoblast cells (dark blue) are shown. b, IHC for Ki67 in early first trimester placenta (6-8 weeks g.a) compared to late first trimester (10-12 weeks g.a.). The proportion of proliferative cells is greatly reduced towards the end of first trimester and the cells are localized mostly in the CCC. Representative images from n =6 for each tissue type. Scale bars, 100μm. c, IHC for EPCAM on first trimester placenta (6-8 weeks g.a.) and cell clusters from placental digests. Experiment independently repeated twice with similar results. Arrowheads show VCT and CCC are EPCAM+ and these cells are present in the cell clumps from the placental digests. Scale bars, 50μm (placenta) and 200μm (placental digest). d, Time course for derivation of trophoblast organoids from one placental isolate. Bright field images of Matrigel drops after seeding placental digests starting from passage 0 day 0, until generation of homogenous trophoblast organoids (passage 2, day 7). For passages 0 and 1, timepoints at day 0 and 7 are shown. Passage 2, day 7 is shown together with the zoom of the boxed area in the lower panel. Experiment independently repeated for all organoid cultures with similar results. Scale bars, 500μm (Matrigel droplet images) and 200μm (zoom in on trophoblast organoids). Gestational age, g.a.; SCT, syncytiotrophoblast; VCT, villous cytotrophoblast; CCC, cytotrophoblast cell column; EVT, extravillous trophoblast; DG, decidual gland; SA, spiral artery; pl, placental; dec, decidual.
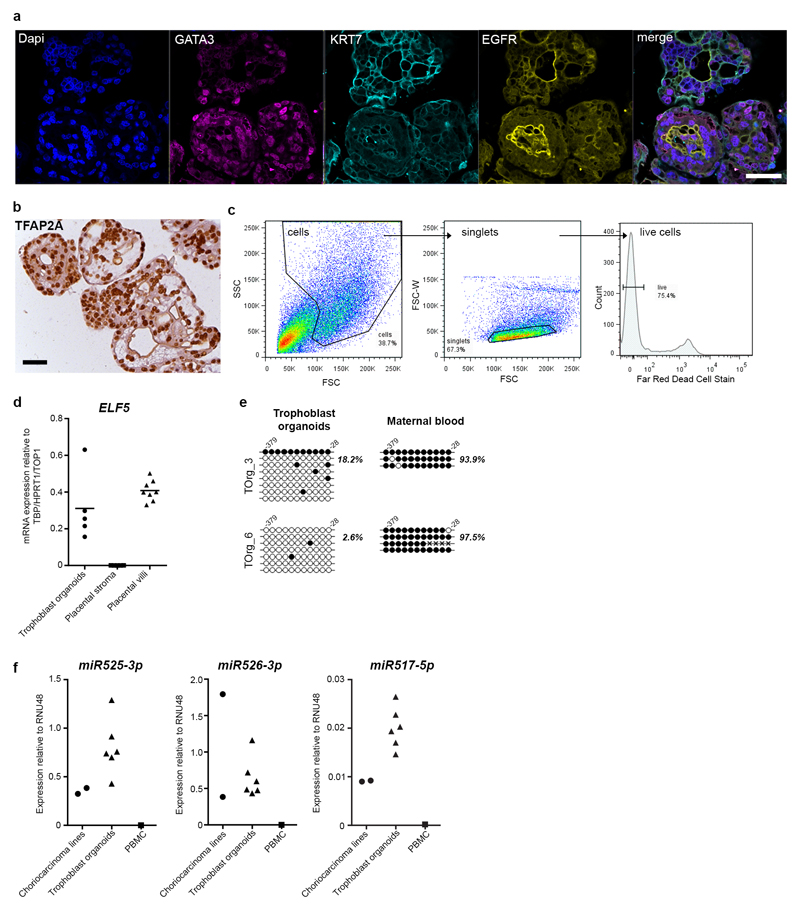
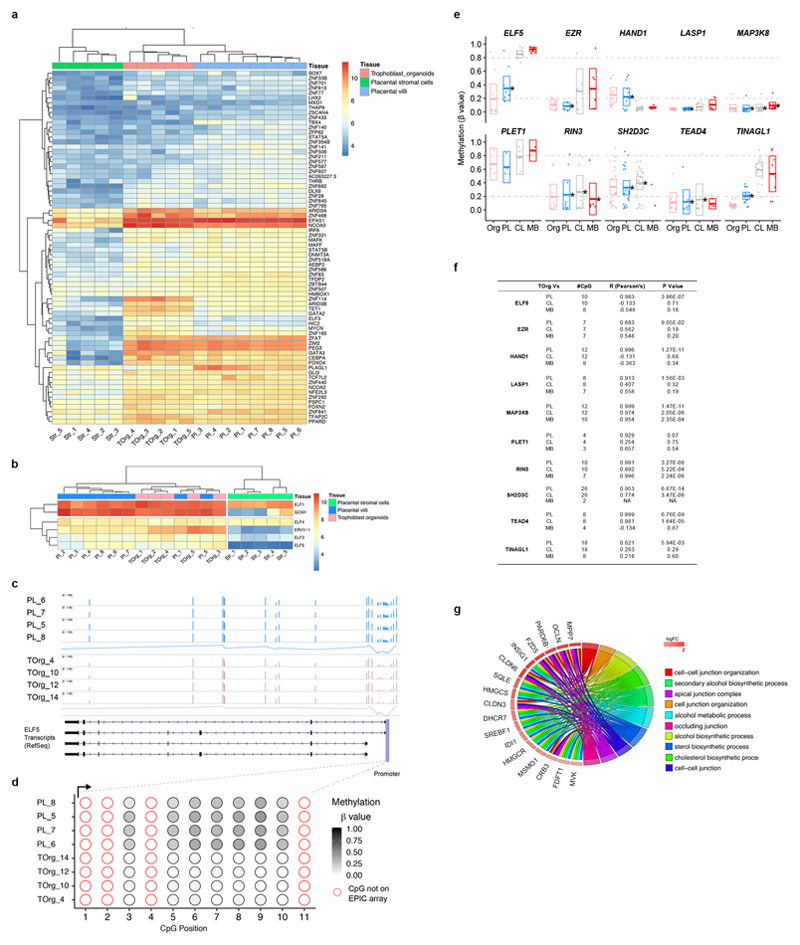
The trophoblast identity of the organoids was verified on the basis of our previously defined criteria: they express GATA3, KRT7, EGFR, TFAP2A and TFAP2C; they lack expression of HLA class I molecules; they express ELF5 and its promoter is hypomethylated; and they express microRNAs from the C19MC miRNA complex at similar or higher levels to choriocarcinoma lines, JEG-3 and JAR (Fig. 2a-d; Extended Data Fig. 3a-f; Source data Files 1-2)3. To assess how well the trophoblast organoids recapitulate their tissue of origin in an unbiased approach, we performed a microarray analysis of established organoids and compared them to first-trimester placental villi (also containing stromal, Hofbauer and endothelial cells) and cultured villous stromal cells. To check for maternal cell contamination, decidual glandular organoids were included. The results were analysed by PCA and hierarchical clustering (Fig 2e,f, Extended Data Fig. 4a,b). PC1 shows trophoblast organoids cluster closely to the placenta with enrichment for trophoblast-specific genes, CGB3, GATA3 and PSG6, compared to the stromal cells and glandular organoids. PC2 highlights epithelial genes (CLDN3, TACSTD2 and KRT23), specific for trophoblast compared to other placental cells (Extended Data Fig. 4b). IHC confirms that KRT23 is a novel trophoblast-specific keratin placental villi and in trophoblast organoids (Extended Data Fig. 4c). Comparison of differentially expressed genes (fold change≥2, adjusted P value≤0.05) between placental villi, trophoblast organoids and stromal cells highlights other genes of interest, such as PGF, CCNE1, ERBB3 and FOLR1; translation of CCNE1 in trophoblast was validated by IHC (Extended Data Fig. 4e). Imprinted genes, PEG3 and PEG10 are also highly expressed in the trophoblast organoids (Extended Data Fig. 4d). Amongst transcription factors, beside known genes, GATA3 and TFAP2C, new genes emerged such as ELF3 (Extended Data Fig. 5a,b). Genome-wide methylation analysis revealed a high degree of correlation between trophoblast organoids and first-trimester placental villi across different genomic elements compared to blood and brain (Fig. 2g). The hypomethylation of ELF5 promoter was also confirmed (Extended Data Fig. 5c,d). Analysis of the promoter regions of genes with similar methylation patterns to Elf5 in mouse TS cells shows that ELF5, EZR, TINAGL1 and LASP1 are similarly hypomethylated in placental villi and trophoblast organoids (Extended Data Fig. 5e,f)12. Gene ontology analysis of DEGs, represented by Chord plot, shows terms describing metabolic processes and cell-cell organization converging on epithelial, developmental and hormonal pathways, FZD5 (WNT signalling), INSIG1 (insulin signalling), DHCR7 (cholesterol synthesis) and OCLN (polarity)(Extended Data Fig. 5g).
Figure 2. Trophoblast organoids retain characteristic features of first trimester trophoblast in vivo, and similar transcriptome and global methylation profiles.
a, IHC for TFAP2C shows uniform expression (representative images from n=20). Scale bars, 50μm. b, FACS analysis of three trophoblast organoids (TOrg10, 12, 14) and JEG (positive control) with mAb W6/32. Gating strategy see Extended Data Fig. 3c. Experiment independently repeated 3 times. c, Bisulfite sequencing of the ELF5 promoter region of trophoblast organoids (TOrg 6) and matched maternal leukocytes (positive control). Relative % of methylated cytosine residues (filled circles) are indicated. d, qPCR analysis for miR517-3p from C19MC cluster on trophoblast organoids (n=6), JEG-3 and JAR choriocarcinoma (Chc.) lines (positive controls) and peripheral blood monocytes (PBMC). Graph shows relative expression levels to housekeeping gene RNU48. Source Data File 1. e, PCA of placental villi (n=8); trophoblast organoids derived from different placentas, TOrg_1 (passage p4), TOrg_2 (p7), TOrg_3 (p6), TOrg_4 (p4) and TOrg_5 (p6) (n=5); placental stromal cells (n=5); and decidual organoids (n=3). Analysis based on 12673 probes. Organoids cluster more closely with placental villi on PC1 axis. f, Clustered heatmap of differentially expressed genes in first trimester placental villi (n=8)(blue), trophoblast organoids (n=5)(pink) and cultured placental villous stromal cells (n=5)(green). g, Distribution of methyl-cytosine across genomic features is similar between trophoblast organoids (n=4) and placental samples (n=4). In contrast, the brain (n=1) and maternal blood (n=5) samples show distinct patterns, especially across CpG islands, gene bodies and LINE1 elements. Pearson’s correlation coefficient (R) indicated for each comparison against trophoblast organoid samples (all p values <2.2e-16). Density/violin plots are scaled to area.
Trophoblast organoids grow as complex structures closely recapitulating the structure of placental villi in vivo where VCT stain for EPCAM and CDH1 (Fig. 3a,b). The basement membrane is on the outside in contact with Matrigel with syncytial masses lining the central cavity (Extended Data Fig. 6a). Similar to in vivo, VCT cells are Ki67+ and TP63+ (Fig. 3b, Extended Data Fig. 6b,c). Following incubation with EdU, ~30-40% of cells are proliferating when the organoids are small (100-200μm diameter), with a notable decrease as they enlarge and differentiate (Extended Data Fig. 6d). Expression of SCT markers (CD71, CD46) is found inside the organoids (Extended Data Fig. 6e). These characteristic features of SCT were confirmed by electron microscopy (EM): multinucleated with abundant secretory organelles and surface microvilli (Fig. 3c). Lacunae present within the syncytial areas resemble those found in vivo (Extended Data Fig. 6f,g). GCM1 drives fusion of VCT into SCT by upregulating ERVW-1 (SYNCYTIN-1)13,14. Using qRT-PCR, we detect high levels of expression of GCM1 and ERVW-1, comparable to those in the placenta (Fig. 3d; Source Data File 2). Thus, trophoblast organoids closely mimic the villous placenta both structurally and phenotypically. SCT secretes proteins and hormones into the maternal systemic circulation, inducing maternal adaptations to pregnancy. We explored the secretory activity of trophoblast organoids through an unbiased peptidomic analysis of the organoid medium and LC MS/MS technology (Fig. 4a). Among the most abundant peptides are placental-specific PSGs and INSL4, whose functions are unknown (Extended Data Table 3, Supplementary Table 2a-h). Aldose reductase, which converts glucose to sorbitol is also detected. High concentrations of sorbital are present in first trimester placentas15. Hence, the organoids also mimic the villous placenta metabolically as well as endocrinologically. Peptides that induce physiological and metabolic adaptations during pregnancy, including hCG, KISS1 and CSH1 are all abundant as well as GDF15, implicated in hyperemesis gravidarum16. GDF15 and hCG are detected by ELISA, showing that full-length and appropriately folded hormones are secreted by trophoblast organoids (Fig. 4b,c; Source Data Files 3-4). Indeed, the ‘pregnant’ secretome of the organoids is evident using an “over-the-counter” pregnancy test kit (Fig. 4d).
Figure 3. Trophoblast organoids form complex structures resembling placental villi with formation of syncytiotrophoblast (SCT).
a, Confocal microscopy images of trophoblast organoid stained for F-actin, EPCAM, DAPI merged with phase image (representative image from n=5). The EPCAM+ cells are surrounding the organoid. Scale bar, 100μm. b, IHC for CDH1 and Ki67 of first trimester placenta and trophoblast organoids (representative images from n=6 for Ki67 and n=20 for trophoblast organoids). VCT stains positively for CDH1. Ki67 is present in the inner VCT layer in villi and the outer layer in organoids. Scale bar, 50μm. Higher power in insets scale bar, 20μm. c, Electron micrograph images of first trimester SCT compared to the centre of a trophoblast organoid. Surface microvilli (arrowheads) and multinucleated areas can be seen (arrows). Scale bars, 5μm (placenta), 1μm (trophoblast organoids, microvilli) and 2.5μm (trophoblast organoids, nuclei). Representative images from n=2. d, qPCR analysis of genes ERVW-1 and GCM1 in trophoblast organoids (n=5) compared to whole placental villi (n=8) and stromal cells isolated from the placenta (n=5). Graphs show expression levels relative to geometric mean of three housekeeping genes TBP, TOP1 and HPRT1. The mean ERVW-1 and GCM1 expression are shown for each sample group. Source Data File 2.
Figure 4. Secretome of trophoblast organoids contains placental hormones and proteins.
a, Experimental work-flow for Liquid chromatography-mass spectrometry (LC-MS) analysis of the secretome of trophoblast organoids. Supernatants from six trophoblast organoid cultures derived independently from six different placental samples were analysed: TOrg_2 (p23), TOrg_3 (p20), TOrg_5 (p6), TOrg_10 (p12), TOrg_12 (p4) and TOrg_14 (p5). b, ELISA for GDF15 secreted by trophoblast organoids (n=6). Shown is amount of GDF15 (ng/mL) produced by trophoblast organoids (between day 7-10 after passaging) in 48h. Source Data File 3. c, ELISA for HCGβ secreted by trophoblast organoids (n=5). Shown is amount of HCGβ (ng/mL) produced by trophoblast organoids (between day 7-10 after passaging) in 48h. Source Data File 4. d, Over-the-counter pregnancy stick denoting “Pregnant” after being placed into dish containing cultures of trophoblast organoids. Image reproduced with the permission of SPD Swiss Precision Diagnostics GmbH (SPD). Experiment independently repeated twice.
Human trophoblast also differentiates to EVT, a process crucial for proper placentation. EVT express HLA-G and invade decidual tissue to transform the spiral arteries17. In TOM our trophoblast organoids show only sporadic HLA-G+ cells (Extended Data Fig. 7a). Recently, long-term, 2D monolayer cultures of human trophoblast cells derived from first-trimester placentas that can differentiate into SCT and EVT were described18. By adapting their EVT differentiation protocol and culturing both our trophoblast organoids and primary villous explants in their EVT medium (EVTM), HLA-G+ cells appear that migrate out of the organoids, digest the Matrigel to form tracks, and eventually adhere to the plastic (Fig. 5a-e, Extended Data Fig. 7b, Supplementary Videos 1-6). In vivo EVT are generated at the base of CCCs where cells express ITGA210. We used flow cytometry to confirm that, after exposure of organoids to EVTM, HLA-G+ EVT appear and ITGA2+ cells disappear (Fig. 5f).
Figure 5. Generation of migratory and invasive HLA-G+ extravillous trophoblast (EVT) cells from trophoblast organoids.
a, Phase-contrast images taken across several z-stacks combined into a single image by the extended focus module from Zeiss Axiovision of a trophoblast organoid and b, a placental villous explant plated into Matrigel after 7-10 days in EVT differentiation medium (EVTM). Regions of interest in samples are boxed in white and corresponding higher power snapshots are shown with relative time-lapse intervals in yellow (h:min). Migratory cells are labelled with yellow arrowhead. In EVTM, cells from both the organoids and primary tissue show random migration. See also control image for organoid in TOM (Extended Data Fig. 7b and time-lapse videos (Supplementary Videos 1-6). Scale bars, large images in a and b 200μm; images shown from insets, 50μm. c, Phase contrast images of trophoblast organoids plated in Matrigel drop and exposed to either TOM or EVTM. Cells stream out of organoids, digesting the Matrigel and eventually adhering to the plastic only when cultured in EVTM. Scale bars, 200μm. d, Live cells growing out from trophoblast organoids in EVTM stained with HLA-G monoclonal antibody, G233. Scale bar, 50μm. e, Flow cytometry of trophoblast organoids cultured in TOM or EVTM and double-stained with monoclonal antibodies W6/32 (binds all HLA class I molecules) and MEMG9 (specific for HLA-G). In TOM virtually all cells lack HLA class I expression but become HLA-G+ after culturing in EVTM. f, Histogram showing organoids cultured in TOM or EVTM stained for ITGA2, that marks cells at the base of the cytotrophoblast cell columns. Before exposure to EVTM, 23% cells are ITGA2+ but very few are present after differentiation to HLA-G+ EVT. All experiments (a-f) have been repeated independently at least 3 times.
In summary, we describe the generation of human trophoblast organoids that grow as complex 3D structures with fusion of VCT to hCG-secreting SCT, anatomically and functionally closely resembling the villous placenta in vivo. In addition, we show differentiation to HLA-G+ EVT that vigorously invade and digest the Matrigel in 3D. Following the submission of our paper, there is a report describing the generation of trophoblast organoids from pooled patient samples; however these cannot be cultured long-term and have not been fully characterised19. Our findings complement those of Okae et al. and mean that there are now two culture systems (2D and 3D) for human trophoblast18. Our 3D model has the advantage of organising into complex structures generating both SCT and EVT. The retention of the normal villous structure will allow analysis of morphogenetic events. We anticipate that these two different models will provide valuable tools across a range of disciplines. They can be used to study maternal-fetal transmission of xenobiotics, drugs and pathogens, and the proteins and hormones derived from the SCT20. Analysis of CCC formation and EVT differentiation in vitro will allow investigation of the decidual microenvironment on trophoblast function, such as the effect of glandular histotrophic nutrition and the influence of the distinctive uterine natural killer (NK) cells21,22. Major unexplained disorders of pregnancy such as pre-eclampsia, stillbirth and fetal growth restriction have their origins in aberrant placental development in the first trimester23. Trophoblast organoids can be used to study maternal-fetal interactions following implantation, and the maternal physiological, metabolic and hormonal changes occurring during pregnancy.
Methods
Patient samples
All tissue samples used for this study were obtained with written informed consent from all participants in accordance with the guidelines in The Declaration of Helsinki 2000. Elective terminations of normal pregnancies were performed at Addenbrooke’s Hospital (6-9 weeks gestation) under ethical approval from the Cambridge Local Research Ethics Committee (04/Q0108/23). Human peripheral blood was collected from a healthy donor in a BD vacutainer following informed consent and in accordance with the ethical approval of the Human Biology Research Ethics Committee, University of Cambridge (HBREC.2016.03).
Derivation of trophoblast organoids from human placental tissue
To obtain trophoblast-enriched cell suspensions, villi from first trimester placental tissue were sequentially digested with 0.2% trypsin-250 (Pan Biotech P10-025100P)/0.02% EDTA (Sigma #E9884)/PBS then collagenase V at 1.0mg/ml (Sigma #C9263) in Hams F12/10% FBS. Both digests were pooled, washed in Advanced DMEM/F12 medium (Gibco #12634-010) and re-suspended in ~10x volume growth-factor-reduced Matrigel (Corning #356231) on ice. 25µL drops were plated per well into a 48-well culture plate (Costar #3548), set at 37°C for 15min and overlaid with 250µL Trophoblast Organoid Medium (TOM, Supplementary Table 1b). Cultures were maintained in 5% CO2 in a humidified incubator at 37°C. Medium was replaced every 2-3d. Small organoid clusters became visible by ~7d and were passaged when at least 50% had reached a diameter of 200-300µm (usually between 7-10d). Mechanical disruption was achieved with Eppendorf Explorer Plus automatic pipettes on a mix cycle of 99 rounds (x4-5), maximum speed. Organoids from the same sample were initiated and maintained in the absence of each individual component to test its importance. Frozen stocks of organoids were made in 70% TOM, 20% FBS and 10% DMSO freeze medium and stored in liquid nitrogen. A step-by-step protocol of the derivation and maintenance of human trophoblast organoid cultures can be found at Nature Protocol Exchange24.
Generation of EVT from trophoblast organoids
Trophoblast organoids were passaged and plated into 35mm dishes or ibidi μ-dishes (Thermo Scientific #81156). Differentiation was achieved through a modification of the protocol used by Okae et al18. After passaging, organoids were maintained in TOM for 3-4d and switched to EVT medium (EVTM: advanced DMEM/F12, 0.1mM 2-mercaptoethanol (Gibco #31350), 0.5% Penicillin-streptomycin, 0.3% BSA (Sigma #A8412), 1% ITS-X supplement (Gibco #51500-056), 100ng/mL NRG1 (Cell Signaling #5218SC), 7.5μM A83-01 (Tocris Biotechne #2939), 4% Knockout serum replacement (ThermoFisher #10828010). When organoids showed outgrowth of cells (typically 7-10d), the medium was changed to EVTM without NRG1 for a further 7-10d. For comparison, fresh placental villi were embedded into 300uL Matrigel in ibidi μ-dishes and grown under the same conditions.
Isolation of placental stromal cells
Placental villous stromal cells were isolated by digesting the tissue remaining after the initial trypsin/collagenase digests in 10-15mL collagenase V in Hams F12/10% FBS with gentle shaking at 37°C for 5-10min. The cell suspension was filtered through gauze, washed and pelleted. Cells were resuspended in Advanced DMEM/F12/10% FBS with additional L-Glutamine, non-essential amino acids (Gibco #11140-035) and primocin (Invivogen #ant-pm-1) and seeded into tissue culture flasks. They were cultured to 80-90% confluency and passaged once before use.
Peripheral blood monocytes (PBMC) isolation
PBMC were isolated from blood by Pancoll-based (Pan-Biotech #P04-60500) density gradient separation. PBMC viability was >95% by Trypan Blue exclusion. PBMC were resuspended in QIAzol lysis reagent (Qiagen #79306) for total RNA extraction following the supplier’s protocol.
Cell lines
Human choriocarcinoma cell lines, JEG-3 and JAR, were obtained from the American Type Culture Collection (ATCC) and cultured as previously described3.
Immunohistochemistry (IHC)
Organoids were formalin-fixed and embedded as previously described11. IHC on sections of organoids and first-trimester placentas was performed using A.Menarini heat-induced epitope retrieval buffers and Vectastain avidin-biotin-HRP reagents as previously described11. Primary antibodies (Supplementary Table 3) were replaced with equivalent concentrations of isotype-matched mouse or rabbit IgG for controls. Images were captured with a Zeiss Axiovert Z1 microscope and Axiovision imaging software SE64 V4.8.
Immunofluorescence (IF) and confocal microscopy
Trophoblast organoids were grown in 4-5 20μL Matrigel drops in 35mm ibidi μ-dishes and EdU and/or antibody labelling was performed as previously described11. EdU incubation was for 1h at 37°C in TOM containing 10μM EdU. For primary antibodies and secondary antibodies used see Supplementary Table 1d. Imaging was with the ZEISS 700 Confocal microscope and ZEN Microscope Software.
Mitotracker staining
Mitochondrial function was evaluated by Mitotracker Red-CMXRos (Thermofisher #M7512). Organoids were released from Matrigel with Cell Recovery Solution (Corning #354253) and incubated in 500nM of Mitotracker Red in TOM in suspension at 37°C for 30 mins. The organoids were washed in basal medium, resuspended and plated into a thin layer of Matrigel in ibidi μ-dishes for imaging on a ZEISS 700 Confocal microscope with ZEN Microscope Software.
Time-lapse microscopy
Trophoblast organoids or placental villous explants embedded in 300uL of Matrigel in 35mm ibidi μ-dishes were imaged in phase-contrast and across several z-stacks on a Zeiss Axiovert Z1 microscope with the multidimensional imaging function of the Axio Observer software Axiovision image software V4.8. The images were compiled into a single movie by using the extended focus wavelet function.
ELISA
Conditioned media were harvested from organoid cultures and centrifuged to remove debris and stored at -80°C until use. Human Chorionic Gonadotropin β (hCG-β) ELISA (Abcam #ab108638) was performed on 50μL supernate with 100μL sample buffer in duplicate alongside hCG-β standards following the manufacturer’s instructions. Concentration of hCG-β in the supernates was calculated from the line formula of the standard plots in Microsoft Office Excel. Supernatants were also tested with Clear&Simple Digital Pregnancy Test following manufacturer’s instructions. Image reproduced with the permission of SPD Swiss Precision Diagnostics GmbH (SPD). GDF-15 was measured by in-house electrochemiluminescence immunoassay on the MesoScale Discovery assay platform (MSD) using BioTechne DuoSet antibodies and standard (BioTechne #DY957). For further details see Supplementary Methods.
Flow cytometry
Organoids were removed from Matrigel with Cell Recovery Solution and dissociated with 0.2% trypsin 250 (Pan Biotech P10-025100P)/0.02% EDTA (Sigma #E9884) in PBS at 37°C for 5 min. Cells were washed in medium containing FBS and passed through a 40μm cell strainer (Falcon #2340). Cells were blocked with human IgG (Sigma #I4506) in DPBS (ThermoFisher Scientific #14190136)/1%FBS prior to labelling with W6/32-Alexa-488 anti-HLA-A,B,C antibody, HLA-G-PE, ITGA2-PE or isotype-matched controls (Supplementary Table 1d). LIVE/DEAD™ Fixable Far Red Dead Cell Stain (Life Technologies #L10119) was used for live/dead discrimination. Data were acquired using Cytek Development DxP8 (488/637/561). Data were analysed in FlowJo (Tree Star) and all compensation was applied digitally after acquisition.
In Situ Hybridization (ISH) Assays
ISH for LGR5 was performed on 4μm paraffin sections with RNAscope 2.0 High definition assay (Advanced Cell Diagnostics) following the manufacturer’s instructions. Briefly, tissue sections were baked at 60°C for 1h, dewaxed with xylene, cleared in 100% ethanol and air-dried prior to the standard protocol: 10min in Pretreat buffer 1, 15min in Pretreat buffer 2 and 30min at 37°C in Pretreat buffer 3 followed by incubation with LGR5 probe (#311021), positive control probe UBC (#310041) or negative control probe dapB (#310043) for 2h at 40°C. Signal was visualized with the amplification kit and DAB for 10min. Sections were dehydrated, mounted in DPX (Sigma #44581) and imaged on a Zeiss Axiovert Z1 microscope with Axiovision imaging software SE64 V4.8.
Electronic microscopy (EM)
Trophoblast organoids were directly fixed in 35mm dishes with 0.5% glutaraldehyde/0.2 M sodium cacodylate buffer (pH 7.2) for 30min and reduced with osmium tetroxide as previously described11. Ultrathin sections were examined in an FEI Tecnai G2 TEM at 80Kv. Images were acquired with MegaView III CCD and Soft Imaging Systems program. Samples from human placentas were fixed by immersion in 3% glutaraldehyde-0.3% hydrogen peroxide in 0.1mol/L 1,4-piperazine diethane sulfonic acid (PIPES) buffer (pH 7). After 2h at RT, tissue was washed for 30min in 0.1 mol/L PIPES buffer. Secondary fixation was by immersion in 1% osmium tetroxide in PIPES buffer for 1h at RT. After washing, specimens were dehydrated in graded ethanol and embedded in Araldite epoxy resin. Ultrathin sections (50nm) were cut on a Reichert-Jung Ultracut S (Reichert-Jung). Sections were counterstained with uranyl acetate, followed by lead citrate, before viewing in a Philips CM100 electron microscope (Philips Electronics).
DNA extraction and quantification
QIAamp DNA blood Mini kit (Qiagen #51104) was used to extract genomic DNA from patients’ blood for Short tandem repeat analysis, HLA tissue typing and Bisulphite sequencing. DNA was extracted from trophoblast organoids, decidual and placental tissues by digestion with ATL buffer (Qiagen #19076) and Proteinase K (Sigma #P4850), followed by purification steps with RNase A (Sigma #R6513) and Protein Precipitation Solution (Qiagen #158910). DNA was precipitated with isopropanol and washed with 70% ethanol. DNA quality and concentration were determined in a Nanodrop ND-1000 Spectrophotometer.
Comparative Genomic Hybridization (CGH) analysis
DNA from two independently derived trophoblast organoid samples at early and late passages was analysed with Agilent Sureprint G3 unrestricted CGH ISCA 8x60K array (Agilent #G4450A). DNA samples from late passage organoids were compared to early passage organoids (hybridization control). DNA was diluted to 50ng/uL and labelled using the Agilent kit following manufacturer’s instructions. Data analysis for segmentation and copy number calls was performed at a genome-wide resolution of 500kb by the default analysis method – CGH v2 from the Agilent CytoGenomics software Edition 2.5.8.11 (Build 37).
Short tandem repeat analysis and HLA typing
Microsatellite analysis was performed with the GenePrint® PowerPlex™16 System (Promega) involving fluorescent labelled multiplexed PCR amplification of 15 short tandem repeat (STR) loci and Amelogenin sex determining fragments. PCR fragment size resolution was achieved with capillary electrophoresis on a 3730XL DNA Analyzer (Applied Biosystems) before analysis of the raw data and STR allele calling with GeneMapper Versions 4 and 5 (Life technologies) fragment analysis sizing and genotyping software. All typing was performed blind. The Promega PowerPlex 16 kit was designed for Forensic testing and has a sensitivity capable of detecting down to 5% contamination. It is used by for monitoring of post-haematopoietic stem cell transplant chimerism25,26. The DNA for HLA genotyping was processed via the workflows of the EFI accredited Clinical Histocompatibility Laboratory. Low resolution typing of the HLA-A, -B and -C genes was achieved with LABType kits (One Lambda) which rely on reaction patterns observed when sequence specific DNA probes immobilised on fluorescent X-MAP polystyrene beads (Luminex) hybridise to biotin labelled multiplexed gene specific PCR amplicons. The hybrids were detected with a Liquichip 200 fluorimeter (QIAgen) and HLA allele assignment was from HLA Fusion software (One Lambda). Ultra High resolution typing of HLA-A, -B, -C, -DRB1, -DQB1 and -DPB1 was achieved with an ‘in house’ Third Generation Sequencing pipeline utilising Pacific Biosciences’ Single Molecule Real-Time DNA sequencing technology as previously described27.
Bisulphite sequencing
Approximately 300ng of DNA was used for bisulphite conversion performed with the EpiTect Bisulfite Kit (Qiagen #59110), according to the manufacturer’s protocol. The ELF5 promoter region was amplified as described previously28. PCR products were cloned and sequenced, confirming representation of distinct alleles.
Global DNA methylation analysis
Genomic DNA bisulfite (BS) was performed with the CEGX TrueMethyl kit (Cambridge Epigenetix/NuGEN) and used for microarray-based DNA methylation analysis, performed at GenomeScan (GenomeScan B.V., Leiden, The Netherlands), on the HumanMethylation850 BeadChip (Illumina) and were scanned on the Illumina iScan system. The resulting iDAT files were imported and analysed by ChAMP (v2.9.10)29,30. Samples were processed filtering for a probe detection p-value <= 0.01, probes with a bead count <3 in at least 5% of samples, no CpG and known SNPs at probe starts, probes aligning to multiple locations, and QC using the on array control probes31. Of the total probes on the array 755577 passed the filtering and QC steps. The BMIQ method was used to normalise the two probe types present on the array. Beta methylation values from the EPIC array range from 0 (unmethylated) to 1 (methylated) and are equivalent of percentage methylation32. Genomic annotations were imported from FDb.InfiniumMethylation.hg19 and IlluminaHumanMethylationEPICmanifest33. Genomic features in Fig. 2g comprise the following numbers of assayed CpGs: CpG Islands, 48799, 48799, 13576; Promoters, 151270, 151270, 88621; Gene Bodies, 280,284,280,284,117,138; LINE1, 48799, 48799, 13576. Boxplots comprise min: 1.5 x inter-quartile, lower: 1st quartile, middle: median, upper: 3rd quartile, max: 1.5 x inter-quartile range (Extended Data Figures 2G and 5e). Density/violin plots are scaled to area. LINE1 elements were downloaded as tables from UCSC Genome browser for hg1930. Maternal blood samples (normal) are taken from E-GEOD-66210.
RNA extraction, quantification and quality control
Total RNA was isolated using the miRNeasy isolation kit (Qiagen #217004) with on-column DNAse digestion (Qiagen #79254). Quantitation of RNA was performed with the Quant-iT™ RiboGreen™ RNA Assay Kit (Thermo Fisher Scientific #R11490) by measuring the intensity of fluorescence at 528nm with a Synergy HT Multi-Mode Microplate Reader (BioTek Instruments) according to manufacturer's instructions. RNA quality was assessed on the Agilent 2100 bioanalyzer (Thermo Fisher Scientific). RNA Integrity Number (RIN) of each tested sample was greater or equal to 8.
Reverse transcription and Real-time quantitative RT-PCR (qRT-PCR)
The expression of ELF5, ERVW-1 and GCM1 was analysed with Taqman Gene expression assays (Applied Biosystems). 500ng-1μg of total RNA was reverse transcribed with Superscript VILO Reverse Transcriptase (Thermo Fisher Scientific #11754050) in the presence of random hexamers and RNAse inhibitor following supplier’s instructions. qRT-PCR was performed on 7900HT Fast Real-Time PCR system (Applied Biosystems) as previously described11. Relative expression levels were normalised to the geometric mean of three housekeeping genes HPRT1, TOP1 and TBP and the 2-ΔCt method. The expression of C19MC miRNAs hsa-miR-517-5p, hsa-miR-517(a, b)-3p, hsa-miR-526b-3p, hsa-miR-525-3p and reference gene RNU48 were analysed by TaqMan miRNA assays (Applied Biosystems). 10ng of total RNA were reverse transcribed using miRNA specific stem-loop RT primers and TaqMan microRNA reverse transcription kit (Applied Biosystems #4366596) according to the supplier’s instructions. QRT-PCR assays were run with qPCRBIO Probe Mix Lo-ROX (PCR Biosystems) containing specific probes on an Eppendorf Mastercycler® RealPlex 2 instrument. Ct data were normalised to the RNU48 internal control by the 2-ΔCt method. All qPCR reactions included no-template controls and minus RT controls (–RT). For further details and for Taqman Assay IDs for each gene see Supplementary Methods.
Microarray expression profiling and data analysis
The microarray experiment was performed at Cambridge Genomic Services at University of Cambridge with a species specific Gene 2.1 ST Array Plate (Affymetrix) according to manufacturer’s instructions. Briefly, 100ng of total RNA was amplified for each sample with inline PolyA spike-in control and the WT PLUS amplification kit (Affymetrix). By using the in line hybridization controls, we successfully amplified samples with the GeneChip WT terminal labelling kit (Affymetrix). Plate arrays were processed on the GeneTitan instrument (Affymetrix) with the GeneTitan Hybridization, Wash and Stain kit (Affymetrix). Samples were hybridized to the array, washed, stained and scanned. CEL files generated were loaded in R using the oligo package from Bioconductor. The raw data were then processed after quality controls using the Robust Multichip Analysis method. The limma package (3.34.8) was used to make the comparisons, and results were corrected for multiple testing using False Discovery Rate. Microarray probes without gene identifier (ensembl gene id) were filtered out. Initial QC included PCA and MDS plots. Finally the quality of the data was assessed, and the correlation of the samples in the groups compared. Heatmaps were generated with the R package 'pheatmap' (1.0.8), which uses the Euclidean method. For the gene heatmaps, the input is the normalized intensity matrix. GO terms enrichment was obtained with R package 'clusterProfiler' (3.6.0) with function `enrichGO`, and chord plots were generated with the R package 'GOplot' (1.0.2).
LC-MS analysis of trophoblast organoid supernatants
Trophoblast organoids (day 10 after passaging) were grown in trophoblast organoid medium and supernatants collected after overnight incubation. The supernatants (500µL) and an aliquot of growth media were acidified with 50µL of 1% formic acid in water (v/v) and loaded directly onto an Oasis Prime µ-elution 96 well SPE plate (Waters #186008052) and extracted as described previously34. The eluant was evaporated under oxygen free nitrogen at 40ºC and reconstituted into 75µL of 50mM ammonium bicarbonate in water with 10mM dithiothreitol. Protein digests were prepared and analysed using an Ultimate 3000 nano LC system coupled to a Q Exactive Plus Orbitrap mass spectrometer (ThermoScientific) as described previously35. The nano LC/MS files obtained from the six different extracts were combined and searched using Peaks 8.5 software (BSI) against the human Swissprot database (Downloaded on 26-Oct-2017). A tryptic digest setting was used and precursor and product ion tolerances were set at 10 ppm and 0.05 Da respectively. The search parameters included a fixed modification of a carboxyamidomethylation on cysteine residues and variable modifications such as methionine oxidation, N-terminal pyro-glutamate, N-terminal acetylation and C-terminal amidation. A false discovery rate (FDR) value of 1% was applied at the peptide and a minimum of 1 unique peptide also was required. For further details see Supplementary Methods.
Statistics and Reproducibility
All experiments reported in this study have been reproduced with similar results derived from independent samples (tissues and organoids) from multiple patients. The number of times the experiments were repeated with independently-derived trophoblast organoid cultures are reported in figure legends as (n=) and summarized in Supplementary Table 1c. Given the descriptive nature of the work and biological variation between human samples, the experimental data points for each patient sample are shown separately unless stated otherwise. Trophoblast organoid culture protocols were independently replicated by four scientists. Statistical analyses used to analyse microarray, methylation and LC-MS data are reported in Methods above.
Extended Data
Extended Data Figure 1. Staining for signalling pathways in first trimester placenta and decidua.
a, IHC of first trimester placenta (6-8 weeks g.a.) for effectors of major signalling pathways (i) WNT signalling, non-phosphorylated (p)-β-catenin (S33/S37/T41), (ii) TGFβ signalling through p-SMAD 2 (S465/467)/Smad3 (S423/425), (iii) MAPK signalling through p-ERK 1/2 (T202/Y204) and (iv) p-STAT3 (Y705) signalling. Scale bars, 50μm. Representative images from n=8, for each antibody. BMP signalling through SMAD1/5/8 was not possible to assess by IHC. VCT and CCC displayed membrane-localised staining of non-p β-catenin, whereas p-Erk1/2 was mostly cytoplasmic in both cell types. Cytoplasmic and nuclear signals for p-Erk1/2 were detected in EVT. P-SMAD 2/3 staining also showed stronger nuclear signals in EVT, suggesting a role for TGFβ signalling in differentiation in accordance with previous report9. Phosphorylated nuclear STAT3 was detected only in EVT, again indicating involvement in their differentiation. SCT was negative for all these signals. b, Summary of findings from a. Trophoblast cells from different regions of the placenta are represented as a circle with nucleus (small inner circle). Black indicates strong staining, grey indicates faint staining and white not detected. Thicker circles indicate staining localized to cell membrane. c,In situ hybridization for LGR5 on first trimester placental villi. LGR5 transcripts are detected in VCT. Stroma is negative. Positive control probe is for UBIQUITIN (UBC). Negative control probe is for the bacterial gene dapB. Nuclei are counterstained in Haematoxylin. Images are at x10 magnification. Representative images from n=2. d, IHC for Rspondin-1 on early first trimester (6-8 weeks g.a.) and late first trimester (10-12 weeks g.a.) decidual samples. Representative images from n=2 for each tissue type. Images are at x20 magnification. SCT, syncytiotrophoblast; VCT, villous cytotrophoblast; CCC, cytotrophoblast cell columns; EVT, extravillous trophoblast; UG, uterine glands.
Extended Data Figure 2. Culture components tested for the establishment of long-term organoid cultures of trophoblast from human placentas.
a, Growth factors (HGF, PGE2, Y-27632, Nicotinamide) were added as supplements to basal trophoblast organoid medium (TOM) that contains EGF, CHIR99021, Rspondin-1, A83-01, FGF2 (Supplementary Table 1a,b). Bright field images of placental digests at passage 1, day 7. The cystic structures that appear with the addition of Nicotinamide (red asterisks) are contaminating maternal glandular organoids. Representative images from n=2. Conditions containing factors that did not show growth are not included. Scale bars in upper and lower rows, 500μm. b, Trophoblast organoid cultures at passage 2 and at passage 10 with continuous culture. Representative images from n=3. Scale bars, 500μm. c, Analysis of genetic stability of cultures (n=2) with comparative genomics hybridization (CGH) array. Shown is a representative whole-genome array CGH plot generated with Agilent Cytogenomics software. Genomic DNA from late passage (p8) trophoblast organoids is compared to genomic DNA from early passage (p2). Each spot is a single probe. Plotted are the log ratios of the average signal intensity of each probe on the Y-axis along its position on the chromosomes (1-22, X and Y) on the x-axis. A log signal ratio of 0 represents equivalent copy number the samples. No significant DNA copy number abnormalities were identified. d, Live imaging of trophoblast organoid cultures (n=2) passaged >6 months and then frozen/thawed and exposed to Mitotracker Red. Functional mitochondria are visible showing that the cells are healthy (white arrowheads). Scale bars, whole organoid 50μm; Individual cells 10μm. e, Organoids derived from the same placental cell isolate using either trophoblast organoid medium (TOM) or Decidual organoid medium (ExM) demonstrating that matched placental (fetal) and decidual (maternal) organoids can be derived from the one sample. Representative bright field images from n=3. Scale bars, 500μm.
Extended Data Figure 3. Trophoblast organoids retain characteristic features of first trimester trophoblast in vivo.
a, Representative images of positive staining for GATA3, KRT7, EGFR and DAPI on trophoblast organoids by confocal microscopy (representative image from n=3). EGFR stains both VCT and the surface of SCT as in vivo7. The basement membrane is around the outside of the organoids with formation of syncytial masses in the centre. Scale bar, 50μm. b, IHC for transcription factor TFAP2A shows uniform expression on trophoblast organoids (representative image from n=20). Scale bars, 50μm. c, Gating strategy used for flow cytometric analysis of single, live cells from trophoblast organoids. d, Quantitative RT-PCR (qPCR) analysis of ELF5 in trophoblast organoids (n=5) compared to whole placental villi (n=8) and stromal cells isolated from the placenta (n=5). Graph shows expression levels relative to the geometric mean of three housekeeping genes TBP, TOP1 and HPRT1. The mean ELF5 expression is shown for each sample group. Source Data File 2. e, Bisulfite sequencing of the ELF5 promoter region (-379bp to -28bp upstream of the transcription start site) of trophoblast organoids from two different placentas (TOrg 3, 6) and matched maternal blood leukocytes (positive control). The relative % of methylated cytosine residues (filled circles) are indicated. f, qPCR analysis for miR525-3p, miR526-3p and miR517-5p from C19MC miRNA cluster on trophoblast organoids (n=6), JEG-3 and JAR (positive controls) and peripheral blood monocytes (PBMC, low expression/negative control). Graph shows relative expression levels of each organoid culture to housekeeping gene RNU48. Source Data File 1.
Extended Data Figure 4. Hierarchical clustering of microarray data comparing placental villi, trophoblast organoids and placental stromal cells.
a, Unsupervised hierarchical clustering analysis of global gene expression profiles by microarray of first trimester placental villi (Pl)(n=8), trophoblast organoids (TOrg)(n=5), placental stromal cells (Str)(n=5) and decidual organoids (DOrg)(n=3). Analysis was based on 12673 probes. The expression profiles of trophoblast organoids cluster with first trimester placental samples whilst decidual organoids and placental stromal cells cluster in a separate tree. b, Top 20 genes contributing to PC1 and PC2 in the PCA plot from Fig. 2e. The top genes contributing to PC1 are all trophoblast-specific genes such as CGB3, GATA3 and PSG6 indicating that these genes separate the trophoblast organoid and placental villous samples from the two potentially-contaminating, non-trophoblast samples (decidual organoids and placental stroma). The top genes contributing to PC2 are epithelial genes such as CLDN3, TACSTD2 and KRT23. The organoids only contain trophoblast, but cells of the villous core (stromal, Hofbauer and endothelial cells) are also present in the placental samples. c, IHC of placental villi and trophoblast organoids stained for KRT23 showing expression in all trophoblast cells in vivo and in vitro. Experiment repeated independently 3 times. Scale bar, 50μm. Higher power in insets scale bar, 20μm. d, Clustered heatmap of differentially expressed genes between trophoblast organoids, placental villi and placental stroma with an absolute log2 fold change of 2 (adjusted p.value < 0.05). e, IHC of placental villi and trophoblast organoids stained for CCNE1 showing expression in trophoblast cells in vivo and in vitro. Scale bar, 50μm. Higher power in insets scale bar, 20μm. Experiment repeated independently 3 times.
Extended Data Figure 5. Transcription factor expression profiles of trophoblast organoids and placental villi.
a, Heatmap highlighting transcription factors from the differentially expressed genes between placental villi, trophoblast organoids and placental stromal cells. b, Heatmap of genes from the ELF family of transcription factors and syncytial genes, GCM1 and ERVW-1. ELF3 and ELF5 both show moderate expression levels across the organoids and placental samples, and very low/no expression in the stromal samples. ELF4 and ELF1 are similar in all samples. There is very high expression of ELF1 in placentas and organoids. Similarly, both ERVW-1 and especially GCM1 are expressed at higher levels in placentas and organoids in agreement with qPCR data (Fig. 3d). c, Genomic mapping of the methylation array probes to the ELF5 gene. The height of the bars indicates methylation level from 0, unmethylated to 1.0, fully methylated. d, Methylation of the ELF5 promoter shows hypomethylation in the organoid and placenta samples. e, Distribution of methyl-cytosine across the promoters of the 10 murine trophoblast gatekeeper genes12. The organoids (Org) and placenta (PL) samples show very similar methylation patterns across all 10 gene promoters that are distinct from the control/brain (CL) and maternal-blood (MB) in the majority of the genes. Boxplots comprise min: 1.5 x inter-quartile, lower: 1st quartile, middle: median, upper: 3rd quartile, max: 1.5 x inter-quartile range. Significant correlations (<0.01) are indicated (*). f, Table showing Pearson’s correlation coefficient (R), number of CpG/probes compared and p-values for e. g, Chord plot representing terms from the gene ontology analysis of upregulated genes in trophoblast organoids.
Extended Data Figure 6. Structure and proliferation in trophoblast organoids.
a, A schematic diagram of a normal placental villus in vivo compared to a trophoblast organoid. The basement membrane (BM) beneath the VCT is contiguous with the stromal villous core in vivo and with the Matrigel in vitro. The SCT contacts maternal blood in the intervillous space in vivo. SCT forms in the centre of the organoids. b, IHC for TP63 in first trimester placenta and trophoblast organoids (representative images from n=14). TP63 is expressed in VCT. Scale bar, 50μm. Higher power insets scale bar, 20μm. c, Representative images of TP63, Ki67 and DAPI staining on trophoblast organoids by confocal microscopy (n=3). Cells on the outside of the organoids are TP63+ and Ki67+. Scale bars, 20μm. d, Confocal microscopy images of trophoblast organoid stained for EdU, EPCAM and DAPI showing fewer proliferating cells (white arrowheads) as the organoids enlarge. Scale bar, 50μm. Representative images from n=3. e, IHC for markers of SCT, CD46 and CD71, in first trimester placenta and trophoblast organoids (representative images from n=20). CD46 and CD71 stain the syncytial brush border. Scale bar, 50μm. Higher power insets scale bar, 20μm. f, Carnegie stage 5b embryo (~9 days post-fertilization)from Carnegie Collection at the early lacunar stage (number 8171). Courtesy of Prof. Enders and the Centre for Trophoblast Research (https://www.trophoblast.cam.ac.uk/Resources/enders). Arrows point to examples of cavities that appear in the primitive syncytium due to fluid accumulation before the coelomic cavity and the embryo have fully developed.g, Similar cavities in placental tissue samples from first trimester (6-9 weeks g.a) and in syncytium in centre of trophoblast organoids. Boxed areas are shown at higher magnification (bottom). Scale bars, 200μm (top); 50μm (bottom). Similar morphology seen in at least 5 early placental villi and in all organoids. ICM, inner cell mass. Pr.Syn. primitive syncytium; VCT, villous cytotrophoblast; SCT, syncytiotrophoblast; Str, stromal core.
Extended Data Figure 7. Trophoblast organoids grown in TOM and EVTM.
a, Confocal image of organoid stained for F-actin, DAPI and HLA-G. A few isolated cells stain for HLA-G (white arrowheads) at the periphery of the organoid. Scale bar, 50 μm. Representative image from n= 3. b, Phase contrast images from time-lapse videos of 16h of trophoblast organoids grown in TOM when EVT differentiation does not occur (top). No invasive cells are visible. Shown for comparison is an organoid (middle) and a placental villous explant exposed to EVTM (bottom). Black arrows indicate cells migrating and arrowheads show visible tracks made as the cells invade through the Matrigel. For time-lapse videos of these cultures see Supplementary Videos 1, 4 and 5. Scale bars, 200μm.
Extended Data Table 1. Microsatellite analysis and HLA typing of organoids from placental digests in TOM show that they are of fetal origin.
a, PowerPlex16 Short Tandem Repeat (STR) genotyping of DNA from matched maternal-fetal samples (maternal blood, decidua and placenta) to identify origin of organoids. Shown in top row are the 15 STR loci analysed. Numbered STR alleles observed for each DNA at a particular locus are listed within the relevant column. As expected in most cases a maximum of two alleles were seen. In cases where there was evidence of an additional allele from a fetal derived STR haplotype the allele number appears in brackets. This is consistent with the decidua containing fetal extravillous trophoblast. The results at informative loci where fetus and trophoblast organoids match are underlined. b, HLA genotyping with Third Generation SMRT sequencing of DNA from matched maternal blood and trophoblast organoid to confirm STR analysis. HLA-A, -B, -C, -DRB1, -DQB1 and -DPB1 loci were investigated. The two HLA alleles at each locus defined at high resolution are shown for each sample. For each locus, the fetal trophoblast will share one allele with the mother but the other allele will be derived from the father and is likely to be different. Not all loci are informative as some paternal and non-inherited maternal alleles are the same.
| a | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample ID | D3S1358 | TH01 | D21S11 | D18S51 | Penta E | D5S818 | D13S317 | D7S820 | D16S539 | CSF1PO | Penta D | Amelo. | vWA | D8S1179 | FGA |
| Maternal Blood_2 | 16,17 | 7,9.3 | 28,30 | 12,14 | 15,15 | 12,13 | 10,15 | 11,11 | 12,13 | 10,10 | 9,9 | x,x | 15,18 | 12,13 | 22.2,23 |
| Decidua_2 | 16,17 | 7,9.3 | 28,30 | 12,14 | 15,15 | 12,13 | 10,15 | 11,11 | 12,13 | 10,10 | 9,9 | x,x | 15,18 | 12,13 | 22.2,23 |
| Placenta_2 | 14,16 | 6,9.3 | 28,30 | 14,15 | 12,15 | 11,13 | 10,14 | 10,11 | 12,12 | 10,10 | 9,13 | x,x | 15,17 | 13,13 | 22,23 |
| Trophoblast Organoid_2 | 14,16 | 6,9.3 | 28,30 | 14,15 | 12,15 | 11,13 | 10,14 | 10,11 | 12,12 | 10,10 | 9,13 | x,x | 15,17 | 13,13 | 22,23 |
| Maternal Blood_3 | 16,17 | 6,9 | 29,31.2 | 12,14 | 7,12 | 11,12 | 10,11 | 9,10 | 11,11 | 10,12 | 10,13 | x,x | 18,18 | 12,13 | 20,25 |
| Decidua_3 | 16,17 | 6,9 (9.3) | 29,31.2 (28) | 12,14 | 7,12 (19) | 11,12 | 10,11 (14) | 9,10 | 11,11 | 10,12 (11) | 10,13 (14) | x,x | 18,18 (19) | 12,13 | 20,25 (21) |
| Placenta_3 | 16,16 | 9,9.3 | 28,31.2 | 12,14 | 7,19 | 11,11 | 10,14 | 9,10 | 11,11 | 11,12 | 13,14 | x,x | 18,19 | 12,12 | 21,25 |
| Trophoblast Organoid_3 | 16,16 | 9,9.3 | 28,31.2 | 12,14 | 7,19 | 11,11 | 10,14 | 9,10 | 11,11 | 11,12 | 13,14 | x,x | 18,19 | 12,12 | 21,25 |
| Maternal Blood_6 | 14,15 | 7,9.3 | 30,30 | 12,16 | 16,19 | 11,11 | 11,13 | 9,12 | 11,13 | 10,11 | 9,9 | x,x | 15,15 | 10,10 | 21,26 |
| Decidua_6 | 14,15 | 7,9.3 | 30,30 | 12,16 | 16,19 | 11,11 | 11,13 | 9,12 | 11,13 | 10,11 | 9,9 | x,x | 15,15 | 10,10 | 21,26 |
| Placenta_6 | 15,17 | 6,7 | 28,30 | 16,17 | 5,16 | 10,11 | 12,13 | 9,11 | 11,13 | 10,10 | 9,13 | x,y | 15,17 | 10,11 | 24,26 |
| Trophoblast Organoid_6 | 15,17 | 6,7 | 28,30 | 16,17 | 5,16 | 10,11 | 12,13 | 9,11 | 11,13 | 10,10 | 9,13 | x,y | 15,17 | 10,11 | 24,26 |
| Maternal Blood_7 | 15,16 | 6,6 | 29,30.2 | 16,16 | 7,18 | 9,13 | 12,12 | 9,10 | 13,13 | 11,11 | 10,11 | x,x | 17,18 | 12,14 | 21,21 |
| Decidua_7 | 15,16 (17) | 6,6 | 29,30.2 | 16,16 | 7,18 | 9,13 | 12,12 (10) | 9,10 | 13,13 (10) | 11,11 | 10,11 | x,x (y) | 17,18 | 12,14 (13) | 21,21 (24) |
| Placenta_7 | 15,17 | 6,6 | 29,29 | 16,16 | 7,18 | 13,13 | 10,12 | 9,10 | 10,13 | 11,11 | 10,11 | x,y | 17,18 | 13,14 | 21,24 |
| Trophoblast Organoid_7 | 15,17 | 6,6 | 29,29 | 16,16 | 7,18 | 13,13 | 10,12 | 9,10 | 10,13 | 11,11 | 10,11 | x,y | 17,18 | 13,14 | 21,24 |
| Maternal Blood_8 | 17,18 | 7,9 | 30,30 | 16,20 | 11,12 | 12,12 | 11,12 | 9,10 | 11,14 | 11,12 | 11,13 | x,x | 15,18 | 13,14 | 22,22 |
| Decidua_8 | 17,18 (15) | 7,9 (8) | 30,30 | 16,20 (12) | 11,12 (7) | 12,12 (10) | 11,12 | 9,10 (11) | 11,14 | 11,12 | 11,13 | x,x (y) | 15,18 | 13,14 | 22,22 (28) |
| Placenta_8 | 15,18 | 7,8 | 30,30 | 12,16 | 7,12 | 10,12 | 11,11 | 9,11 | 9,14 | 11,11 | 11,11 | x,y | 15,17 | 13,14 | 22,28 |
| Trophoblast Organoid_8 | 15,18 | 7,8 | 30,30 | 12,16 | 7,12 | 10,12 | 11,11 | 9,11 | 9,14 | 11,11 | 11,11 | x,y | 15,17 | 13,14 | 22,28 |
| Maternal Blood_9 | 15,18 | 9.3,9.3 | 33.2,34.2 | 12,17 | 5,5 | 12,13 | 12,12 | 10,11 | 11,11 | 10,12 | 11,12 | x,x | 15,16 | 14,15 | 19,22 |
| Decidua_9 | 15,18 | 9.3,9.3 (10) | 33.2,34.2 (31) | 12,17 (13) | 5,5 (14) | 12,13 | 12,12 | 10,11(8) | 11,11 | 10,12 | 11,12 (10) | x,x (y) | 15,16 | 14,15 (12) | 19,22 (21) |
| Placenta_9 | 15,15 | 9.3,10 | 31,33.2 | 13,17 | 5,14 | 12,13 | 12,12 | 8,10 | 11,11 | 10,12 | 10,12 | x,y | 16,16 | 12,14 | 19,21 |
| Trophoblast Organoid_9 | 15,15 | 9.3,10 | 31,33.2 | 13,17 | 5,14 | 12,13 | 12,12 | 8,10 | 11,11 | 10,12 | 10,12 | x,y | 16,16 | 12,14 | 19,21 |
| b | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample ID | HLA-A* | HLA-A* | HLA-B* | HLA-B* | HLA-C* | HLA-C* | HLA-DRB1* | HLA-DRB1* | HLA-DQB1* | HLA-DQB1* | HLA-DPB1* | HLA-DPB1* |
| Maternal Blood_2 | 02:01:01:01/16 | 02:01:01:01/16 | 35:01:01:02/04/05/06 | 44:02:01:03 | 07:04:01:01/03 | 03:03:01:01 | 11:01:01 | 08:01/77 | 03:01:01 | 04:02:01 | 04:01:01 | 04:01:01 |
| Trophoblast Organoid_2 | 02:01:01:01/16 | 11:01:01:01 | 35:01:01:02/04/05/06 | 39:01:01:03/05 | 12:03:01:01 | 03:03:01:01 | 11:01:01 | 08:01/77 | 03:01:01 | 04:02:01 | 04:02:01 | 04:01:01 |
| Maternal Blood_3 | 02:01:01:01/16 | 24:02:01:01 | 15:01:01:01 | 40:01:02:01/04 | 03:04:01:01 | 03:03:01:01 | 04:04:01 | 01:01:01 | 03:02:01 | 05:01:01 | 04:01:01 | 16:01:01 |
| Trophoblast Organoid_3 | 02:01:01:01/16 | 24:02:01:01 | 15:01:01:01 | 15:01:01:01 | 03:04:01:01 | 03:03:01:01 | 04:04:01 | 04:01:01 | 03:02:01 | 03:02:01 | 04:01:01 | 04:01:01 |
| Maternal Blood_6 | 03:01:01:01 | 30:02:01:01 | 07:02:01:01/03 | 18:01:01:01/06 | 05:01:01:01 | 07:02:01:03 | 01:03:01 | 01:03:01 | 03:01:01 | 05:01:01 | 02:01:02/19 | 04:01:01 |
| Trophoblast Organoid_6 | 68:01:02:02 | 30:02:01:01 | 27:05:02:01 | 18:01:01:01/06 | 05:01:01:01 | 07:04:01:01/03 | 01:03:01 | 08:01/77 | 04:02:01 | 05:01:01 | 02:01:02/19 | 04:01:01 |
| Maternal Blood_7 | 32:01:01:01 | 33:01:01:01 | 14:02:01:01 | 44:02:01:01 | 05:01:01:02 | 08:02:01:01 | 01:01:01 | 01:02:01 | 05:01:01 | 05:01:01 | 13:01/107:01 | 04:01:01 |
| Trophoblast Organoid_7 | 24:02:01:04 | 33:01:01:01 | 14:02:01:01 | 44:02:01:01 | 05:01:01:02 | 08:02:01:01 | 13:01:01 | 01:02:01 | 05:01:01 | 06:03:01 | 02:01:02/19 | 04:01:01 |
| Maternal Blood_8 | 24:02:01:01 | 25:01:01:01 | 18:01:01:02/05 | 27:05:02:01 | 01:02:01:01 | 12:03:01:01 | 15:01:01 | 13:03:01 | 03:01:01 | 06:02:01 | 04:01:01 | 03:01:01 |
| Trophoblast Organoid_8 | 24:02:01:01 | 02:01:01:01/16 | 49:01:01 | 27:05:02:01 | 01:02:01:01 | 07:02:01:01/15 | 11:02:01 | 13:03:01 | 03:01:01 | 03:19:01 | 10:01:01 | 03:01:01 |
| Maternal Blood_9 | 26:01:01:01 | 03:01:01:01 | 07:02:01:01/03 | 27:05:02:01 | 02:02:01:01 | 07:02:01:03 | 04:04:01 | 04:04:01 | 03:02:01 | 03:02:01 | 04:01:01 | 02:01:02/19 |
| Trophoblast Organoid_9 | 26:01:01:01 | 02:01:01:01/16 | 44:02:01:01 | 27:05:02:01 | 02:02:01:01 | 05:01:01:02 | 04:01:01 | 04:04:01 | 03:01:01 | 03:02:01 | 04:02:01 | 02:01:02/19 |
Extended Data Table 2. Microsatellite analysis and HLA typing of organoids from placental digests derived in ExM show that they are of maternal origin.
a, PowerPlex16 STR genotyping of DNA from matched maternal-fetal samples (decidua, maternal blood, placenta) to identify origin of organoids. Shown in the top row are the 15 STR loci analysed. Numbered STR alleles observed for each DNA at a particular locus are listed within the relevant column. As expected in most cases a maximum of two alleles were seen but where there was lesser evidence of an additional allele from a fetal derived STR haplotype the allele number appears in brackets. The results at informative loci where maternal blood, decidua and organoids match are underlined. b, HLA genotyping with LABType Reverse SSO of DNA from 4 pregnancies with matched decidua, blood, placenta and organoids. Each pair of HLA alleles defined, at low resolution, for each sample at a particular locus are listed within the relevant column. The maternal origin of the organoids is clear from HLA-A, -B and -C alleles in 3 of the 4 pregnancies. The data was inconclusive for pregnancy 2 but all 4 organoids are clearly maternal in origin when using both methods (see above).
| a | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample ID | D3S1358 | TH01 | D21S11 | D18S51 | Penta E | D5S818 | D13S317 | D7S820 | D16S539 | CSF1PO | Penta D | Amelo. | vWA | D8S1179 | FGA |
| Maternal Blood_1 | 15, 15 | 7, 9 | 30, 30 | 14, 17 | 11, 12 | 12, 13 | 11, 12 | 8, 8 | 11, 12 | 12, 12 | 9, 12 | xx | 17, 19 | 13, 15 | 22, 25 |
| Decidua_1 | 15, 15 | 7, 9 | 30, 30 | 14, 17 | 11, 12 | 12, 13 | 11, 12 | 8, 8 | 11, 12 | 12, 12 | 9, 12 | xx | 17, 19 | 13, 15 | 22, 25 |
| Placenta_1 | 15, 15 | 7, 8 | 29, 30 | 17, 17 | 10, 12 | 11, 12 | 11, 12 | 8, 10 | 12, 12 | 10, 12 | 10, 12 | xx | 16, 17 | 13, 15 | 22, 25 |
| Decidual Organoid_1 | 15, 15 | 7, 9 | 30, 30 | 14, 17 | 11, 12 | 12, 13 | 11, 12 | 8, 8 | 11, 12 | 12, 12 | 9, 12 | xx | 17, 19 | 13, 15 | 22, 25 |
| Maternal Blood_2 | 16, 17 | 6, 9.3 | 28, 28 | 15, 19 | 12, 12 | 11, 12 | 8, 11 | 12, 14 | 12, 12 | 10, 11 | 11, 12 | xx | 16, 17 | 12, 15 | 20, 21 |
| Decidua_2 | 16, 17 | 6, 9.3 | 28, 28 | 15, 19 | 12, 12 | 11, 12 | 8, 11 | 12, 14 | 12, 12 | 10, 11 | 11, 12 | xx | 16, 17 | 12, 15 | 20, 21 |
| Placenta_2 | 16, 17 | 6, 8 | 28, 31.2 | 15, 19 | 12, 15 | 11, 12 | 8, 9 | 10, 12 | 12, 13 | 10, 11 | 11, 12 | xx | 16, 17 | 10, 15 | 20, 23 |
| Decidual Organoid_2 | 16, 17 | 6, 9.3 | 28, 28 | 15, 19 | 12, 12 | 11, 12 | 8, 11 | 12, 14 | 12, 12 | 10, 11 | 11, 12 | xx | 16, 17 | 12, 15 | 20, 21 |
| Maternal Blood_3 | 14, 17 | 6, 6 | 28, 31.2 | 12, 12 | 7, 13 | 12, 12 | 11, 12 | 10, 12 | 11, 13 | 10, 11 | 9, 10 | xx | 15, 17 | 13, 13 | 23, 24 |
| Decidua_3 | 14, 17 | 6, 6 | 28, 31.2 | 12, 12, (15) | 7, 13 | 12, 12, (13) | 11, 12 | 10, 12, (9) | 11, 13 | 10, 11, (12) | 9, 10, (13) | xx (y) | 15, 17 | 13, 13, (14) | 23, 24, (22) |
| Placenta_3 | 17, 17 | 6, 6 | 28, 28 | 12, 15 | 7, 12 | 12, 13 | 11, 11 | 9, 10 | 13, 13 | 11, 12 | 10, 13 | xy | 15, 17 | 13, 14 | 22, 24 |
| Decidual Organoid_3 | 14, 17 | 6, 6 | 28, 31.2 | 12, 12 | 7, 13 | 12, 12 | 11, 12 | 10, 12 | 11, 13 | 10, 11 | 9, 10 | xx | 15, 17 | 13, 13 | 23, 24 |
| Maternal Blood_4 | 15, 15 | 9.3, 9.3 | 30, 32.2 | 18, 22 | 12, 14 | 11, 12 | 8, 12 | 10, 12 | 12, 13 | 9, 10 | 12, 16 | xx | 15, 16 | 13, 13 | 19, 22 |
| Decidua_4 | 15, 15 | 9.3, 9.3 | 30, 32.2 | 18, 22 | 12, 14 | 11, 12 | 8, 12 | 10, 12 | 12, 13 | 9, 10 | 12, 16 | xx | 15, 16 | 13, 13 | 19, 22 |
| Placenta_4 | 15, 16 | 7, 9.3 | 30, 30 | 14, 18 | 11, 12 | 11, 12 | 12, 14 | 9, 12 | 13, 14 | 10, 10 | 9, 16 | xx | 16, 16 | 13, 14 | 20, 22 |
| Decidual Organoid_4 | 15, 15 | 9.3, 9.3 | 30, 32.2 | 18, 22 | 12, 14 | 11, 12 | 8, 12 | 10, 12 | 12, 13 | 9, 10 | 12, 16 | xx | 15, 16 | 13, 13 | 19, 22 |
| b | |||
|---|---|---|---|
| Sample ID | HLA-A | HLA-B | HLA-C |
| Maternal Blood_1 | 01,02 | 41,52 | 12,17 |
| Decidua_1 | 01,02 | 41,52 | 12,17 |
| Placenta_1 | 01,02 | 27,52 | 01,12 |
| Decidual Organoid_1 | Fail x2 | 41,52 | 12,17 |
| Maternal Blood_2 | 02,02 | 07,44 | 05,07 |
| Decidua_2 | 02,02 | 07,44 | 05,07 |
| Placenta_2 | 02,02 | 07,44 | 05,07 |
| Decidual Organoid_2 | 02,02 | 07,44 | 05,07 |
| Maternal Blood_3 | 25,29 | 18,44 | 12,16 |
| Decidua_3 | Fail x2 | 18,44 | 12,16 |
| Placenta_3 | 23,25 | 18,44 | 04,12 |
| Decidual Organoid_3 | 25,29 | 18,44 | 12,16 |
| Maternal Blood_4 | 23,24 | 13,44 | 04,04 |
| Decidua_4 | 23,24 | 13,44 | 04,04 |
| Placenta_4 | 11,24 | 13,40 | 02,04 |
| Decidual Organoid_4 | 23,24 | 13,44 | 04,04 |
Extended Data Table 3. Liquid chromatography-mass spectrometry (LC-MS) analysis of the secretome of trophoblast organoids.
Supernatants from independently-derived trophoblast organoid cultures from six different placental samples were analysed: TOrg_2 (p23), TOrg_3 (p20), TOrg_5 (p6), TOrg_10 (p12), TOrg_12 (p4) and TOrg_14 (p5). Table shows the glycoproteins, classical peptides/protein hormones, placental specific peptides and proteins enriched in placental tissue identified in the secretome data by LC-MS. The -10lgP value is the statistical significance assigned to a peptide/protein match by the PEAKS software37. Coverage% refers to the proportion of the primary amino acid sequence of each protein that is identified in the experiment. #Peptides refers to the number of peptide matches assigned to a protein, whilst #Unique refers to the number of peptides that are assigned solely to that protein group. #Spec refers to the number of peptide MS/MS spectra matched against a particular protein. The columns also indicate whether these are unique products produced by the placenta and/or whether they are products produced by other tissues but highly enriched in the placenta (among the top 10 organs producing that protein based on RNA expression levels). Tissue location data was compared to data from the Human Protein Atlas, https://www.proteinatlas.org/. All protein identification data are in Supplementary Table 2.
| Accession | Description | -10lgP | Coverage (%) | Peptides | Unique | Spec | Placental specific | Placental enriched |
|---|---|---|---|---|---|---|---|---|
| Glycoproteins | ||||||||
| Q16557|PSG3_HUMAN | Pregnancy-specific beta-1-glycoprotein 3 GN=PSG3 | 272.19 | 28 | 10 | 1 | 25 | Yes | |
| P11465|PSG2_HUMAN | Pregnancy-specific beta-1-glycoprotein 2 GN=PSG2 | 259.79 | 34 | 8 | 2 | 30 | Yes | |
| Q00888|PSG4_HUMAN | Pregnancy-specific beta-1-glycoprotein 4 GN=PSG4 | 236.62 | 21 | 7 | 1 | 17 | Yes | |
| P11464|PSG1_HUMAN | Pregnancy-specific beta-1-glycoprotein 1 GN=PSG1 | 228.57 | 21 | 8 | 1 | 15 | Yes | |
| Classical peptides/hormones | ||||||||
| P01233|CGHB_HUMAN | Choriogonadotropin subunit beta GN=CGB | 340.98 | 48 | 20 | 20 | 218 | Yes | |
| P01215|GLHA_HUMAN | Glycoprotein hormones alpha chain GN=CGA | 167.67 | 44 | 3 | 3 | 33 | Yes | |
| Q15726|KISS1_HUMAN | Metastasis-suppressor KiSS-1 GN=KISS1 | 268.26 | 62 | 11 | 11 | 59 | Yes | |
| Q99988|GDF15_HUMAN | Growth/differentiation factor 15 GN=GDF15 | 236.31 | 34 | 7 | 7 | 42 | Yes | |
| Q14641|INSL4_HUMAN | Early placenta insulin-like peptide GN=INSL4 | 154.72 | 27 | 2 | 2 | 4 | Yes | |
| P35318|ADML_HUMAN | Adrenomedullin GN=ADM | 114.05 | 10 | 1 | 1 | 8 | Yes | |
| P0DML2|CSH1_HUMAN | Chorionic somatomammotropin hormone 1 GN=CSH1 | 85.82 | 6 | 1 | 1 | 1 | Yes | |
| P01344|IGF2_HUMAN | Insulin-like growth factor II GN=IGF2 | 59.42 | 4 | 1 | 1 | 1 | Yes | |
| P01303|NPY_HUMAN | Pro-neuropeptide Y GN=NPY | 50.4 | 8 | 1 | 1 | 1 | unknown | unknown |
| Placental specific peptides (not hormones) | ||||||||
| P35556|FBN2_HUMAN | Fibrillin-2 GN=FBN2 | 157.65 | 1 | 2 | 2 | 3 | Yes | |
| Q9BXP8|PAPP2_HUMAN | Pappalysin-2 GN=PAPPA2 | 124.88 | 1 | 1 | 1 | 1 | Yes | |
| O60829|PAGE4_HUMAN | P antigen family member 4 GN=PAGE4 | 112.65 | 37 | 2 | 2 | 2 | Yes | |
| P14061|DHB1_HUMAN | Estradiol 17-beta-dehydrogenase 1 GN=HSD17B1 PE=1 SV=3 | 83.47 | 5 | 1 | 1 | 1 | Yes | |
| P02771|FETA_HUMAN | Alpha-fetoprotein GN=AFP PE=1 SV=1 | 74.38 | 2 | 2 | 1 | 19 | Yes | |
| Q96GT9|XAGE2_HUMAN | X antigen family member 2 GN=XAGE2 | 62.57 | 23 | 1 | 1 | 1 | Yes | |
| Proteins enriched in placental tissue | ||||||||
| P09486|SPRC_HUMAN | SPARC GN=SPARC | 310.3 | 59 | 13 | 13 | 29 | Yes | |
| Q12805|FBLN3_HUMAN | EGF-containing fibulin-like extracellular matrix protein 1 GN=EFEMP1 | 270.65 | 25 | 7 | 7 | 17 | Yes | |
| P15121|ALDR_HUMAN | Aldose reductase GN=AKR1B1 | 264.8 | 49 | 8 | 8 | 21 | Yes | |
| P80723|BASP1_HUMAN | Brain acid soluble protein 1 GN=BASP1 | 234.76 | 36 | 4 | 4 | 13 | Yes | |
| P02751|FINC_HUMAN | Fibronectin GN=FN1 | 233.91 | 5 | 7 | 7 | 7 | Yes | |
| O95633|FSTL3_HUMAN | Follistatin-related protein 3 GN=FSTL3 | 222.26 | 29 | 5 | 5 | 27 | Yes | |
| P23142|FBLN1_HUMAN | Fibulin-1 GN=FBLN1 | 218.6 | 10 | 5 | 5 | 10 | Yes | |
| P10451|OSTP_HUMAN | Osteopontin GN=SPP1 | 178.67 | 32 | 5 | 5 | 5 | Yes | |
| P14543|NID1_HUMAN | Nidogen-1 GN=NID1 | 162.21 | 4 | 4 | 4 | 21 | Yes | |
| Q14118|DAG1_HUMAN | Dystroglycan GN=DAG1 | 148.59 | 5 | 2 | 2 | 7 | Yes | |
| P27487|DPP4_HUMAN | Dipeptidyl peptidase 4 GN=DPP4 | 115.06 | 3 | 1 | 1 | 1 | Yes | |
| P21980|TGM2_HUMAN | Protein-glutamine gamma-glutamyltransferase 2 GN=TGM2 | 106.05 | 6 | 2 | 2 | 2 | Yes | |
| P24593|IBP5_HUMAN | Insulin-like growth factor-binding protein 5 GN=IGFBP5 | 100.63 | 5 | 1 | 1 | 1 | Yes | |
| P09601|HMOX1_HUMAN | Heme oxygenase 1 GN=HMOX1 | 100.13 | 8 | 1 | 1 | 1 | Yes | |
| Q15582|BGH3_HUMAN | Transforming growth factor-beta-induced protein ig-h3 GN=TGFBI | 99.46 | 3 | 1 | 1 | 1 | Yes | |
| O76061|STC2_HUMAN | Stanniocalcin-2 GN=STC2 | 94.72 | 8 | 1 | 1 | 2 | Yes | |
| P05111|INHA_HUMAN | Inhibin alpha chain GN=INHA | 80.59 | 5 | 1 | 1 | 2 | Yes | |
| P25815|S100P_HUMAN | Protein S100-P GN=S100P | 70.28 | 40 | 2 | 2 | 2 | Yes | |
| P17936|IBP3_HUMAN | Insulin-like growth factor-binding protein 3 GN=IGFBP3 | 58.22 | 6 | 1 | 1 | 1 | Yes | |
Supplementary Material
Acknowledgements
The authors are grateful to patients for donating tissue for research. We thank D. Moore and staff at the Addenbrookes Hospital, Cambridge; J. Bauer, E. Clemente, E. Farnell, C. Reitter at Department of Pathology; K. Burling and Core Biochemical Assay Laboratory staff at NIHR Cambridge Biomedical Research Centre; I. Simonic at Medical Genetics Laboratory, Cambridge University Hospital; T.C-Davies, H.W.Yung, B.D. Simons, B.K. Koo, M. Huch, Y. W. Loke and all members of the Moffett and Burton labs. This work was funded by Centre for Trophoblast Research, University of Cambridge and Medical Research Council (MR/L020041/1). A.Moffett holds a Wellcome Trust joint investigator award (200841/Z/16/Z). M.Y.Turco received funding from E.U. 7th Framework Programme for research, technological development and demonstration (PIEF-GA-2013-629785) and now holds a Royal Society Dorothy Hodgkin Fellowship. F. Reimann and F. Gribble hold Wellcome Trust joint investigator awards (106262/Z/14/Z and 106263/Z/14/Z), and funding from MRC-Metabolic Diseases Unit (MRC_MC_UU_12012/3), NIHR-Cambridge BRC and AZ/MedImmune.
Footnotes
Data availability. Microarray data for Figure 2, Extended Data Figures 4 and 5 have been deposited in the ArrayExpress database at EMBL-EBI under accession number E-MTAB-6683 (https://www.ebi.ac.uk/arrayexpress/experiments/E-MTAB-6683). Illumina EPIC Methylation array data for Figure 2 and Extended Data Figure 5 have been deposited in the ArrayExpress database at EMBL-EBI under accession number E-MTAB-7204. (https://www.ebi.ac.uk/arrayexpress/experiments/E-MTAB-7204). Code used to analyse microarray data and EPIC array samples is available at https://github.com/CTR-BFX/2018-Turco-Moffett. The mass spectrometry proteomics data for Extended Data Table 3 and Supplementary Table 2 have been deposited to the ProteomeXchange Consortium via the PRIDE36 partner repository (www.ebi.ac.uk/pride/archive/) with the dataset identifier PXD009118 and 10.6019/PXD009118. All source data for graphs are in Supplementary Table 3. All other data supporting the findings of this study are available from the corresponding authors upon reasonable request.
Author Contributions M.Y.T. and L.G. designed and carried out all the core experiments and analysed the data; R.K. performed LCS-MS analysis; R.S.H. and M.P. performed microarray and DNA methylation analysis; A.McW. and S.G.E.M. performed and analysed the STR and HLA typing; M.Ho. performed electron microscopy and confocal analysis; R.F. and H.S. assisted with IHC and ELISA experiments; L.E. performed qPCR experiments; A.S, F.G, F.R, M.He. and S.O. provided intellectual input to the study; M.He. performed the ELF5 methylation experiment; G.J.B. and A.M. assisted in analysis and interpretation of results and jointly supervised the project. M.Y.T, G.J.B. and A.M. wrote the manuscript.
Competing interests The authors declare no competing interests.
References
- 1.Burton GJ, Jauniaux E. What is the placenta? Am J Obs Gynecol. 2015;213:S6 e1–S6 8. doi: 10.1016/j.ajog.2015.07.050. [DOI] [PubMed] [Google Scholar]
- 2.Hamilton WJ, Boyd JD. Development of the human placenta in the first three months of gestation. J Anat. 1960;94:297–328. [PMC free article] [PubMed] [Google Scholar]
- 3.Lee CQ, et al. What Is Trophoblast? A Combination of Criteria Define Human First-Trimester Trophoblast. Stem Cell Reports. 2016;6:257–272. doi: 10.1016/j.stemcr.2016.01.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Kunath T, et al. Developmental differences in the expression of FGF receptors between human and mouse embryos. Placenta. 2014;35:1079–1088. doi: 10.1016/j.placenta.2014.09.008. [DOI] [PubMed] [Google Scholar]
- 5.Paiva P, et al. Human chorionic gonadotrophin regulates FGF2 and other cytokines produced by human endometrial epithelial cells, providing a mechanism for enhancing endometrial receptivity. Hum Reprod. 2011;26:1153–1162. doi: 10.1093/humrep/der027. [DOI] [PubMed] [Google Scholar]
- 6.Hempstock J, Cindrova-davies T, Jauniaux E, Burton GJ. Endometrial glands as a source of nutrients, growth factors and cytokines during the first trimester of human pregnancy : A morphological and immunohistochemical study. 2004;14:1–14. doi: 10.1186/1477-7827-2-58. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Mühlhauser J, et al. Differentiation and proliferation patterns in human trophoblast revealed by c-erbB-2 oncogene product and EGF-R. J Histochem Cytochem. 1993;41:165–73. doi: 10.1177/41.2.8093455. [DOI] [PubMed] [Google Scholar]
- 8.Sonderegger S, Husslein H, Leisser C, Knofler M. Complex expression pattern of Wnt ligands and frizzled receptors in human placenta and its trophoblast subtypes. Placenta. 2007;28(Suppl A):S97–102. doi: 10.1016/j.placenta.2006.11.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Haider S, Kunihs V, Fiala C, Pollheimer J, Knofler M. Expression pattern and phosphorylation status of Smad2/3 in different subtypes of human first trimester trophoblast. Placenta. 2017;57:17–25. doi: 10.1016/j.placenta.2017.06.003. [DOI] [PubMed] [Google Scholar]
- 10.Lee CQE, et al. A niche of trophoblast progenitor cells identified by integrin alpha2 is present in first trimester human placentas. Development. 2018 doi: 10.1242/dev.162305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Turco MY, et al. Long-term, hormone-responsive organoid cultures of human endometrium in a chemically defined medium. Nat Cell Biol. 2017;19:568–577. doi: 10.1038/ncb3516. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Cambuli F, et al. Epigenetic memory of the first cell fate decision prevents complete ES cell reprogramming into trophoblast. Nat Commun. 2014;5:5538. doi: 10.1038/ncomms6538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Yu C, et al. GCMa regulates the syncytin-mediated trophoblastic fusion. J Biol Chem. 2002;277:50062–50068. doi: 10.1074/jbc.M209316200. [DOI] [PubMed] [Google Scholar]
- 14.Frendo JL, et al. Direct involvement of HERV-W Env glycoprotein in human trophoblast cell fusion and differentiation. Mol Cell Biol. 2003;23:3566–3574. doi: 10.1128/MCB.23.10.3566-3574.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Jauniaux E, Hempstock J, Teng C, Battaglia FC, Burton GJ. Polyol Concentrations in the Fluid Compartments of the Human Conceptus during the First Trimester of Pregnancy : Maintenance of Redox Potential in a Low Oxygen Environment. 2005;90:1171–1175. doi: 10.1210/jc.2004-1513. [DOI] [PubMed] [Google Scholar]
- 16.Petry CJ, et al. GDF15 Concentrations in Maternal Serum Associated with Vomiting in Pregnancy: the Cambridge Baby Growth Study. bioRxiv. 2017 doi: 10.1101/221267. [DOI] [Google Scholar]
- 17.Pijnenborg R, Vercruysse L, Hanssens M. The Uterine Spiral Arteries In Human Pregnancy: Facts and Controversies. Placenta. 2006;27:939–958. doi: 10.1016/j.placenta.2005.12.006. [DOI] [PubMed] [Google Scholar]
- 18.Okae H, et al. Derivation of Human Trophoblast Stem Cells. Cell Stem Cell. 2018;22:50–63 e6. doi: 10.1016/j.stem.2017.11.004. [DOI] [PubMed] [Google Scholar]
- 19.Haider S, et al. Self-Renewing Trophoblast Organoids Recapitulate the Developmental Program of the Early Human Placenta. Stem cell reports. 2018;11:537–551. doi: 10.1016/j.stemcr.2018.07.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Robbins JR, Bakardjiev AI. Pathogens and the placental fortress. Curr Opin Microbiol. 2012;15:36–43. doi: 10.1016/j.mib.2011.11.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Burton GJ, Jauniaux E, Charnock-Jones DS. The influence of the intrauterine environment on human placental development. Int J Dev Biol. 2010;54:303–312. doi: 10.1387/ijdb.082764gb. [DOI] [PubMed] [Google Scholar]
- 22.Moffett A, Colucci F. Co-evolution of NK receptors and HLA ligands in humans is driven by reproduction. Immunol Rev. 2015;267:283–297. doi: 10.1111/imr.12323. [DOI] [PubMed] [Google Scholar]
- 23.Smith GC. First-trimester determination of complications of late pregnancy. JAMA. 2010;303:561–562. doi: 10.1001/jama.2010.102. [DOI] [PubMed] [Google Scholar]
- 24.Turco, et al. Establishment and differentiation of long-term trophoblast organoid cultures from the human placenta. Nat Protocol Exchange. doi: 10.1038/s41596-020-0381-x. [DOI] [PubMed] [Google Scholar]
- 25.Nollet F, et al. Standardisation of multiplex fluorescent short tandem repeat analysis for chimerism testing. Bone Marrow Transplant. 2001;28:511–518. doi: 10.1038/sj.bmt.1703162. [DOI] [PubMed] [Google Scholar]
- 26.Mossallam GI, et al. Comparison of variable number tandem repeat and short tandem repeat genetic markers for qualitative and quantitative chimerism analysis post allogeneic stem cell transplantation. J Egypt Natl Canc Inst. 2005;17:103–13. [PubMed] [Google Scholar]
- 27.Turner TR, et al. Single molecule real-time DNA sequencing of HLA genes at ultra-high resolution from 126 International HLA and Immunogenetics Workshop cell lines. HLA. 2018;91:88–101. doi: 10.1111/tan.13184. [DOI] [PubMed] [Google Scholar]
- 28.Hemberger M, et al. ELF5-enforced transcriptional networks define an epigenetically regulated trophoblast stem cell compartment in the human placenta. Hum Mol Genet. 2010;19:2456–2467. doi: 10.1093/hmg/ddq128. [DOI] [PubMed] [Google Scholar]
- 29.Morris TJ, et al. ChAMP: 450k Chip Analysis Methylation Pipeline. Bioinformatics. 2014;30:428–430. doi: 10.1093/bioinformatics/btt684. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Aryee MJ, et al. Minfi: a flexible and comprehensive Bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinformatics. 2014;30:1363–1369. doi: 10.1093/bioinformatics/btu049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Zhou W, et al. Comprehensive characterization, annotation and innovative use of Infinium DNA methylation BeadChip probes. Nucleic Acids Res. 2016;45 doi: 10.1093/nar/gkw967. gkw967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Teschendorff AE, et al. A beta-mixture quantile normalization method for correcting probe design bias in Illumina Infinium 450 k DNA methylation data. Bioinformatics. 2013;29:189–196. doi: 10.1093/bioinformatics/bts680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Fortin J-P, et al. Preprocessing, normalization and integration of the Illumina HumanMethylationEPIC array with minfi. Bioinformatics. 2016;33 doi: 10.1093/bioinformatics/btw691. btw691. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Karolchik D, et al. The UCSC Table Browser data retrieval tool. Nucleic Acids Res. 2004;32:493D–496. doi: 10.1093/nar/gkh103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Kay RG, et al. M F. Liquid chromatography/mass spectrometry based detection and semi-quantitative analysis of INSL5 in human and murine tissues. Rapid Commun Mass Spectrom. 2017;31:1963–1973. doi: 10.1002/rcm.7978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Vizcaino JA, et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 2016;44 doi: 10.1093/nar/gkw880. 11033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Zhang J, et al. PEAKS DB: de novo sequencing assisted database search for sensitive and accurate peptide identification. Mol Cell Proteomics. 2012;11 doi: 10.1074/mcp.M111.010587. M111.010587. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.